Abstract
Fast exhaustion of fossil fuel stocks, as well as problems allied with air pollution, has created a worldwide attention in searching for alternative, renewable lignocellulosic macromolecule–based sources of energy. Bioethanol is one of the valuable substitutes produced by fermentation process. It significantly reduces the consumption of fossil-based fuels and thereby the net carbon dioxide emission. The looming requirement for replacing the fuels based on fossils with a more eco-friendly renewable solution has created much attention in finding out abundant and cheap resources for biofuel production. The utilization of easily available, cheaper, and renewable lignocellulosics would make bioethanol more competitive than fossil fuels. Novel substrates, strain improvements, limited byproducts, product tolerance, and fermentation conditions have been drawing researchers’ attention to increase bioethanol productivity. The present paper discusses the bioethanol production from lignocellulosic-based renewable resources. It also focuses on present challenges and prospects for efficient bioethanol production.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
The most abundant and sustainable asset in the biosphere is lignocellulosic biomass. Its photosynthetic formation reached more than 200 billion tons consistently [1]. Biomass holds stored energy, which can be utilized as a fuel energy source. In the core, sun-powered energy is caught by the biomass (plants) through the cycle called photosynthesis in which light energy is converted into chemical energy that can be later exploited as fuel. The chemical energy which is fixed or stored in the form of carbohydrate molecules gets synthesized from carbon dioxide and water molecules in the plant leaves [2]. Agricultural remnants such as fruits and vegetables are the organic content–enriched materials, not perfectly utilized and sometimes end up as polluters of the environment. These underutilized parts can be valorized into bioethanol production. It can be produced by the fermentation of any sugar-containing raw materials. A successful bioconversion of these carbohydrate resources is considered the most valuable phase for bioethanol production [3].
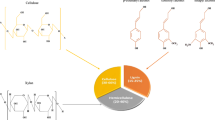
Renewable resource–based ethanol may improve energy accessibility, reduce air pollution, and lessen the atmospheric CO2 accumulation [4, 5]. Many problems which arose due to the consumption of fossil fuels, viz. global warming, environmental pollution, and economic depletion, compel the researchers to find renewable, sustainable, and eco-friendly alternatives. However, confronts undergo in converting lignocellulosic biomass into cost-effective and energy-efficient sustainable biofuels [6]. Renewable energy has been categorized into various fuels, viz. solar, wind power, biomass, geothermal, and tidal energy. Biomass-based energy shares a major portion of renewable energy. Biomass feedstock used for bioenergy is grouped into agricultural, forestry, industry, garden residues, and food residues [7, 8]. First-generation bioethanol (1G) was mainly produced from food crops such as corn and sugarcane; therefore, increased production can indirectly create a worldwide food scarcity. Therefore, it is an urgent need to develop lignocellulosic bioresource–based second-generation bioethanol (2G) production. The highly explored feedstocks for potential ethanol production are agricultural (crop residues), forestry wastes (mill residues), corn, sugarcane, sugar beet, pulp and paper, municipal solid wastes, and microalgae, etc. [4, 7, 9,10,11]. Bioconversion is rather challenging due to the complex structural veracity of plant cell wall materials which is integrally designed to resist microbial degradation [12]. Non-wood lignocellulosic biomass is abundantly available, low cost, and easy to process and consists of a short growth and harvest period, all these properties make it much useful for biomass conversion technology [13]. The major components in lignocellulosic biomass are cellulose, hemicellulose, and lignin. Cellulose- and hemicellulose-based sugars are the most useful feedstocks for future bioethanol production [14]. Lignin is a complex polyphenolic polymer made up of coniferyl, sinapyl, and p-coumaryl alcohol–based phenyl propanoid units. Lignin creates structural stability and rigidity of plant cell walls and hinders the enzymatic hydrolysis of biomass [15]. The production of ethanol from lignocellulosic comprises three major steps: pretreatment, hydrolysis, and fermentation [16, 17].
Pretreatment of the material is necessary to alter the size, structure, and chemical constituents of the biomass, loosen the cellulose fibers from the matrix of lignin, and thereby enhance the accessibility of the enzymes to the substrate. Several pretreatment strategies have been studied, including chemical, physical, and biological strategies. Recent approaches in pretreatment technology may improve saccharification efficiency and thereby reduce the overall production cost [9, 14, 16]. Several phenol-based inhibitory compounds are formed due to lignin degradation under the pretreatment process. Such inhibitory compounds badly affect the enzymatic hydrolysis as well as fermentation process in terms of cell growth and sugar metabolism [15, 18]. Cellulosic bioethanol production mostly depends upon the bioconversion of rigid fibers into fermentable sugars. Therefore, necessary pretreatment processes, viz. physicochemical and biochemical, are essential to improve the accessibility or porosity of recalcitrant lignocellulosic [19, 20].
Lignocelluloses have played a vital role in the production of fermentable sugars for the manufacturing of bio-commodities [21]. The yeast Saccharomyces cerevisiae has long been used as an efficient agent for ethanol production at the laboratory as well as industrial level with high efficiency, thus considered as the world’s premier industrial microorganisms in terms of exploitation and applications. Yeast better tolerates under a wide range of pH, ethanol, and inhibitory compounds as compared to other ethanol producers [3, 22]. Zymomonas mobilis is also acting as a promising alternative for ethanol production because of its high glucose uptake and high ethanol tolerance. The key enzymes for ethanol fermentation are alcohol dehydrogenase and pyruvate decarboxylase [23].
The pentoses basically D-xylose and D-arabinose produced from hemicellulose hydrolysis are not directly uptake by Saccharomyces strains; therefore, genetic modification is required. Candida shehatae is capable of co-fermenting pentoses and hexoses for ethanol production [4]. Scientists are working upon designing organisms through genetic engineering tools to integrate desired enzymes into a single organism. Studies on designer cellulosomes and microbial consortia development relating to consolidated bioprocessing are exciting to overcome the issue of appropriate lignocellulose conversions [24, 25]. Regardless of the goal of nonstarch source–based 21 billion gallons of biofuels by 2022, only 142 million gallons of lignocellulosic-based biofuel was produced in 2015 in the USA [5]. The present review paper discusses the utilization of various raw materials in bioethanol production. It also focuses on the various recent approaches and different aspects used for enhancing bioethanol production.
2 Microorganisms involved in bioethanol production
A conventional and traditional player of industrial bioethanol production is S. cerevisiae, due to its high productivity, high inhibitor tolerance, and ease to genetically engineer. Its toughness enables it to handle tough industrial conditions. Novel pathway introduction and cellular process optimization by metabolic engineering are making its wide range of applications [26, 27]. Several thermotolerant ethanologenic species, i.e., Clostridium thermocellum, C. thermohydrosulfuricum, C. thermosaccharolyticum, Caldicellulosiruptor sp., Thermotoga sp., Thermoanaerobium brockii, Thermoanaerobacter ethanolicus, T. thermo-hydrosulfuricus, T. mathranii, Thermoanaerobacter BG1L1, T. pentosaceus, Thermoanaerobacter sp. DBT-IOC-X2 etc., have been identified as potential ethanol producers.
Since Saccharomyces cerevisiae is unable to ferment pentose-based sugars, the search for pentose-fermenting microorganisms could be an option for better utilization. A yeast strain identified as Meyerozyma guilliermondii utilized pentoses [28,29,30,31,32]. Glycerol-resistant mutant strains of Enterobacter aerogenes ATCC 13048 are also used for ethanol production [33]. Debaryomyces nepalensis, an osmotolerant yeast, utilized both hexoses and pentoses [34].
Co-cultivation of S. stipitis and S. cerevisiae is used for the utilization of mixed C6/C5 sugars [35]. Functional rumen bacterial consortia (FRBC) is also used for production [36]. Consortia of cellulolytic Bacillus sp. THLA0409 and ethanolic Klebsiella oxytoca THLC0409, Saccharomyces cerevisiae OVB 11 and Pichia stipitis NCIM 3498, and Saccharomyces cerevisiae and Candida tropicalis as well as Saccharomyces cerevisiae MTCC 173 and Zymomonas mobilis MTCC 2428 were found satisfactorly [23, 37,34,39].
Xylose utilizable Saccharomyces cerevisiae strain NAPX37 was used [40]. N. crassa strain utilized both hexose and pentose sugars [41]. Spathaspora passalidarum is a non-Saccharomyces yeast also used in bioethanol production [42]. A new yeast strain of Clavispora sp. NRRL Y-50464 is used which utilizes cellobiose [43]. Other strains such as probiotic yeast Saccharomyces boulardii and Clostridium ragsdalei were also found effective [44, 45]. After exhaustive evaluation, three main bioethanol producers, i.e., Saccharomyces cerevisiae, Zymomonas mobilis, and Escherichia coli, have emerged. Engineered bacterial and yeast strains have been constructed with better features through metabolic and genetic engineering that are advantageous for ethanol production [46].
3 Enzymes involved in biomass hydrolysis
There are several enzymes that have been utilized in the bioconversion process. Thermophilic bacteria–based thermostable enzymes play an important role in efficient biomass hydrolysis [47]. Laccases (E.C. 1.10.3.2) (benzenediol, oxygen oxidoreductase, or p-diphenol oxidase) belonging to the oxidoreductase class are used in breaking the plant’s lignocellulosic wall and responsible for degrading the complex polyphenol structure that constitutes lignin. After laccases, lignin peroxidase (LiP) and manganese peroxidase (MnP) are the most significant ligninolytic enzymes. They belong to the heme proteins because they have the protoporphyrin IX as a prosthetic group. Lignin peroxidase (LiP) (E.C. 1.11.1.14) can catalyze and degrade a wide number of aromatic structures such veratryl alcohol (3,4-dimethoxybenzyl) and methoxybenzenes. LiP oxidizes aromatic rings moderately activated by electron-donating substitutes [48].
Glyoxal oxidases (GLOX) are a type of extracellular H2O2-generating peroxidases mainly oxidizing aldehydes generated during lignin and carbohydrate degradation [49]. Aryl-alcohol oxidase (AAO) is providing H2O2 needed by ligninolytic peroxidases for lignin degradation. Thermostable endoglucanase (EndoI) produced by the thermophilic fungus Thermoascus aurantiacus was also found effective [50]. Zhang et al. [51] reported that permeases are responsible for incorporating various essential nutrients and excluding harmful products. Family 1 carbohydrate-binding modules (CBHI) enhance saccharification rates by cloning and expressing CBHI CBM from T. harzianum (CBMCBHI) into Escherichia coli by small ubiquitin-like modifier (SUMO) [52].
Deesterification of the plant biomass is achieved by carbohydrate esterases to make it more approachable for the hydrolytic enzymes such as cellulases, hemicellulases, ligninolytic, and pectinases [53]. A thermostable laccase was produced from Thermus sp. 2.9 for the delignification of Eucalyptus biomass [54]. A new thermo and solvent-stable xylanase was extracted from Bacillus oceanisediminis strain SJ3 by three-phase partitioning [55].
To enhance the activity and thermal stability as well as flexibility of proteins, disulfide bonds present in Trichoderma reesei–based endoglucanase II have been eliminated by site-directed mutagenesis. Replacement of Cysteine99 with valine and Cysteine323 with histidine caused the elimination of two disulfide bonds [56]. To improve the degradation efficiency of cellulases, endoglucanase (Endo5), exoglucanase (Exo5), and different carbohydrate-binding modules (CBMs) were fused to yield several bifunctional cellulases, containing Endo5-2CBM-Exo5, Endo5-CBM3b-Exo5, and Endo5-CBM28-Exo5 [57].
Laccase derived from Trametes maxima IIPLC-32 is used for the detoxification of phenolic inhibitors [58]. Escherichia coli–based laccase CueO was fused with the Dockerin domain of a cellulosome system and finally assembled with the scaffoldin miniCbpA to make a laccase–miniCbpA complex with increased laccase activity [59]. Lytic polysaccharide monooxygenases (LPMOs) have recently been shown to significantly enhance the degradation of recalcitrant polysaccharides. The copper-containing LPMOs utilize electrons to oxidatively cleave polysaccharides [60]. Chimeric thermostable GH7 cellobiohydrolases in Saccharomyces cerevisiae were engineered along with overexpressed glucose-tolerant β-glucosidase [61]. Expressing glycosyl hydrolases in the lignocellulosic-based feedstock is an approving alternative for its utilization. Yeast having laccase with ABTS was effective for direct fermentation of cellulosic materials [5, 62].
4 Raw materials used in bioethanol production
Lignocellulose is a renewable structural component of all plants. Lignocellulose consists of three major components (cellulose, hemicellulose, and lignin) linked by non-covalent forces as well as covalent crossed connections. Lignin acts as a barrier for any treatment due to strong networking with both hemicelluloses and cellulose, therefore prevents the penetration of lignocellulolytic enzymes used for bioconversion [4, 19].
There are various raw materials, which have been used as a potential source for bioethanol production, viz. wheat straw [63,60,65], baggasse [66], corn cobs [67,64,69], pineapple waste [70], Jabon wood [71], cotton waste biomass [72, 73], banana residue [74], old newspapers waste [75], acid hydrolyzate of rice water waste [76], rice husk [77, 78], rice straw [79,76,77,82], rice bran [83], rice hull [84], seed cake [85], dairy industry effluents (cheese whey) [86, 87], domestic food waste [88], papaya peel (PP) [89], spent tea waste (STW), spent wash [90]. sorghum bagasse [91, 92], and sugarcane leaves [93].
Other waste biomasses used are kitchen wastes [94], castor plant [95], potato pulp [96], algal hydrolysates [97], solid digestate from anaerobic digestion [98], oat hulls [99], Arundo donax biomass [100], loblolly pine [101], distillery stillage [102], banana peels [103], Prosopis juliflora [104], olive tree biomass (OTB) [105], jackfruit outer rind [106], Jerusalem artichoke [107], coffee residue waste (CRW) [108], coffee pulp [109], Jerusalem artichoke (Helianthus tuberosus L.) tubers [110], napier grass [111], aloe vera leaf rind [112], waste newspaper hydrolysates [113], safflower plant [114], and palm wood [115].
Other resources are, viz. citrus peel [116], globe artichoke crop residues [117], date waste [118], sorghum milling waste [119], switchgrass [120], Miscanthus grass [121], Conocarpus erectus leaves [122], pineapple fruit peel [123], duckweeds [124], seaweed biomass [125, 126], grape winery waste [127], horticultural waste [128], synthesis gas [129], sweet sorghum juice [130], sugi pulp [131], cow manure [132], cotton gin trash (CGT) [133], high rate algal pond (HRAP) [134], Isoberlinia doka sawdust [19], Ulva fasciata seaweeds [135], palm date [136], orange peel [137], pomegranate peels [138], anaerobic digested sludge (ADS) [139], okara (soybean residue) [140], spent maitake culture medium (SMCM) [9], palm kernel cake [141], kans grass biomass [142], carob solid waste [143], cardoon (Cynara cardunculus) and rockrose (Cistus ladanifer) [144], lemon peels [145], cogongrass [146], Salicornia bigelovii (halophyte plant) [147], cotton stalk [148], and water hyacinth biomass [149].
Scientists have also utilized rapeseed and corn stalks for bioethanol production [150]. Turkish hazelnut husk into lignocellulosic ethanol was investigated [151]. The application of ultrasonic treated sweet lime peel for bioethanol production was also investigated [152]. Stillage from distiller’s dried grains with solubles (DDGS), a by-product of the bioethanol industry, has also been used for bioethanol production [153, 154]. Aloe vera rind (AVR) obtained after extraction of the gel was used for bioethanol production [155]. Hydrolyzed spirulina biomass along with molasses serves as a substrate in bioethanol production [156]. Raw oil palm leaves were used as a substrate for ethanol production [157]. Partially delignified cellulignin (PDCL) was studied for biothanol production [158]. Olive mill waste (OMW), a semisolid waste generated from olive oil production, acts as an attractive substrate for bioethanol production [159]. Hemicellulose fraction of palm fiber is used as a source of sugars for the production of bioethanol by Scheffersomyces stipites [160].
Acid-hydrolyzed broth of banana pseudostem is a potential candidate for ethanol production [161]. Fermented rice noodle wastewater was investigated for ethanol production under an entrapped yeast cell sequencing batch reactor (ECSBR) [162]. Supplementation fruit pulps (mango, banana, and sapota), 4% fruit pulp/puree, enhanced ethanol production (up to 83.1%) in very high-gravity (VHG) fermentation [163]. Switchgrass is a potential source of renewable biomass for conversion to bioethanol [164]. Hardwood spent sulphite liquors (HSSLs) were used in bioethanol production [165]. Soft drinks industry–based wastewater was examined as a media source for bioethanol fermentation [166, 167]. Bioethanol production from autohydrolyzed green coconut shell was also investigated [168]. A wild-growing glucose-rich (56.7% glucose content) brown seaweed species Laminaria digitata was used as the feedstock for bioethanol production [169]. Corn hybrids with high stalk sugar content or “sugarcorn” were used for bioethanol production [170]. Gelidium latifolium was selected as a potential resource [171]. Recycled paper sludge was valorized by a bioethanol production process with cellulase recycling [172].
Bioethanol was also produced from the macroalgae Sargassum spp. [173]. Breadfruit starch hydrolysate (BFSH) was used as the sole carbon source for bioethanol production [174]. Water hyacinth (Eichhornia crassipes), a fast-growing aquatic weed, was used for bioethanol production [175]. De-oiled Pongamia pinnata seed cake acts as a promising feedstock for ethanol production [176]. Kappaphycus alvarezii biorefineries are used for bioethanol production [177]. Bioethanol was also produced from the delignified coconut fiber [178]. Eastern gamagrass C4 perennial grass was used as an alternative cellulosic feedstock for bioethanol production [179].
Waste cotton materials were used as a substrate for bioethanol production [180]. Bioethanol was also produced from Parthenium hysterophorus biomass [181]. The potential of Brachiaria mutica (Para grass) for bioethanol production from Loktak Lake was investigated by Sahoo et al. [182]. Bioethanol production from Japanese bamboo was studied by Singh et al. [183]. Kitchen waste was utilized for bioethanol production [184]. Saccharum biomasses (Saccharum munja and sugarcane bagasse) were used for bioethanol production [185].
Bioethanol production from black tea waste biomass was investigated by Priharto et al. [186]. Ethanol was also produced from rice straw hydrolysate by non-conventional yeasts [187]. Empty fruit bunch was considered a substrate for bioethanol [188]. Oil palm frond juice was used as a renewable source [189].
Salicornia sinus-persica, a succulent halophyte, was used as a raw material for bioethanol production [190]. Eucalyptus sawdust was used for ethanol production [191]. Sulla (Hedysarum coronarium L.) was used as a potential feedstock for biofuel and protein [192]. High solid SSCF of alkaline-pretreated corncob is also used for ethanol production by recombinant Zymomonas mobilis CP4 [193].
Seaweed biomass (Kappaphycus alvarezii, red algal biomass) was utilized by marine yeast for bioethanol production [194]. Bioethanol was also produced from seaweeds (Laminaria digitata, Ulvalactuca, and Dilsea carnosa) [195]. N-Acetyl-d-glucosamine (GlcNAc) was used for ethanol production by Scheffersomyces stipitis strains [196]. Acacia mangium, Paraserianthes falcataria wood, and Elaeis guineensis trunk were investigated for ethanol production by Kaida et al. [197]. Microalgae (Microcystis aeruginosa) were used for efficient bioethanol production. The valorization of coffee byproducts for bioethanol production was studied by Dadi et al. [198]. Eco-friendly processes were studied for soft drink industry wastewater reuse as a growth medium for Saccharomyces cerevisiae–based bioethanol production [167].The utilization of various raw materials for bioethanol production is also illustrated in Table 1.
5 Fermentation strategies used
A number of fermentation strategies have been already used for the effective and economical production of bioethanol such as separate hydrolysis and fermentation (SHF), simultaneous saccharification and fermentation (SSF), and simultaneous saccharification and co-fermentation (SSCF). The new approaches such as simultaneous isomerisation and fermentation (SIF) of xylose and simultaneous isomerisation and co-fermentation (SICF) of a glucose/xylose mixture were carried out by Saccharomyces cerevisiae in the presence of xylose isomerase [303]. Simultaneous saccharification and fermentation of waste wheat–rye bread were investigated and achieved a final ethanol concentration of 128.01 g/L [304]. Semi-simultaneous saccharification and fermentation (SSSF) of ethanol production has also been investigated [305]. Dilute phosphoric acid and steam-based pretreatment of Eucalyptus benthamii for biofuel production was carried out under liquefaction plus simultaneous saccharification and co-fermentation (L+SSCF) process [306]. Biomass was subjected to simultaneous pretreatment and saccharification (SPS) using a cocktail of hydrolytic and oxidizing enzymes for bioethanol production [307]. Bioethanol production was improved by Scheffersomyces stipitis using retentostat extractive fermentation at high xylose concentration [308]. Bioethanol production from pretreated mango stem bark after maceration (MSBAM) was evaluated. The highest yield (84.5%) was obtained under pre-saccharification followed by simultaneous saccharification and fermentation (PSSF) process [309]. Damaged rice grains were used for bioethanol production using presacchararification step followed by simultaneous saccharification and fermentation (SSF) by using waste brewer’s yeast [310]. Sequential fermentation by Saccharomyces cerevisiae and Pichia stipitis improved bioethanol production [311]. A fluidized bed fermenter under a magnetic field was also used for bioethanol production [312].
6 Strategies used for improvement in ethanol production
6.1 Engineered biomass for efficient utilization
The volatile matter and fixed carbon influenced the biological conversion process of the fuel. Woody biomass has a much higher fixed carbon content as compared to LCB. The biomass fuel efficacy does depend not only on the proximate and ultimate analysis but also on the atomic ratio of H/C (hydrogen/carbon) and O/C (oxygen/carbon). The lower the ratio, the higher the energy content. The material with a relatively low O/C ratio has more energy density and higher heating value. LCB feedstocks are primarily composed of carbohydrate polymer and a lower concentration of proteins, acids, salts, and minerals [2].
The structural configuration, arrangement, and chemical composition of wood cell walls have directly affected the hydrolysis process of biofuel production. The understanding of the construction patterns and nature of the cell wall is the key point of second-generation biofuels, which has been done by glycome profiling [313]. Control of phase transition between vegetative to reproductive may also improve biomass yield with reducing lignin content. Delayed floral initiation may be used as a convenient tool for improving biomass quantity and quality [314]. For improved saccharification efficiency, the genetic variability of cell wall degradability has been accessed [315].
Quantitative trait locus (QTL) analysis was used to determine the zones of the phenotypic variation of chemical traits in the genome of interest. Signals for QTLs were assigned to G-lignin and S-lignin, and the ratio between them determines the cellulose, hemicellulose, and water contents. QTL mapped onto chromosomes V, X, XI, and XVI signifies that the saccharification process is under the influence of genetic impact. There may be opportunities to improve the breeding programs for willows for increasing enzymatic saccharification yields and biofuel production [316, 317]. Altering the “Glycomic Code” of cell wall polysaccharides may improve bioenergy production efficiency. The identification of pointrons (hydrolysis resistant) possibly transformed into pexons (available for enzyme attack) is important so that walls would become susceptible for hydrolysis [318].
Cell wall engineering was carried out by heterologous expression of cell wall–degrading enzymes for better conversion of lignocellulosic biomass. Cell wall–hydrolyzing enzymes alter the structural arrangements of the cell wall and reduce cell wall rigidity [319]. The various interactions in the cell wall architecture based on acetyl and phenolic linkages as well as polysaccharide–polysaccharide linkage play a vital role in the development of efficient bioethanol production [320]. Cell wall modification may enhance saccharification [321].
The wall structure of sengon (Paraserianthes falcataria) has been modified through overexpression of poplar cellulase in the cell walls. The overexpression caused a decrement in xyloglucan bound to the walls [322]. To determine lignocellulosic biomass biodegradability, cellulose nanowhiskers gel, lignocellulosic-based xylan matrix, and synthetic lignin were constructed. Application of these materials indicates that the lignocellulosic utilization depended strongly on the xylan Ara/Xyl ratio and the cellulose crystallinity [323].
6.2 Pretreatment approaches used
Pretreatment of raw materials has often been found useful to improve its digestibility and accessibility for microbial attack (by removing core and noncore lignin fractions). It results in enlargement of the inner surface area of substrate particles, accomplished by partial solubilization and/or degradation of hemicellulose and lignin [324, 325]. The goal of any pretreatment process is to alter or remove structural and compositional impediments by breaking the lignin seal, thereby separating the carbohydrates from the lignin matrix as well as disrupting the crystalline structure of cellulose. An efficient pretreatment must free the highly crystalline structure of cellulose and extend the amorphous areas [326,323,328]. Various pretreatment methods such as physical, chemical, physicochemical, and biological methods are described in Table 2.
Pulsed-electric field (PEF) pretreatment exposes the cellulose present in the biomass by creating the pores in the cell membrane, thereby allowing the entry of agents that will break the cellulose into constituent sugars. Under this treatment, biomass is subjected to a sudden burst of high voltage between 5.0 and 20.0 kV/cm for short durations (nano to milliseconds) [330]. Another method is by using a deep eutectic solvent (a fluid generally composed of two or three cheap and safe components) that are capable of self-association, often through hydrogen bond interactions, to form a eutectic mixture with a melting point lower than that of each component. Choline chloride (ChCl)–based deep eutectic solvents (DES) are used for the pretreatment process [331].
H2O2 (0.5% v/v)-based pretreatment at 121°C for 30 min was used as an effective method for the conversion of waste office paper and newspaper into fermentable sugars and after bioethanol [332]. Retama raetam biomass was pretreated by thermo-mechanical process for effective utilization [333]. Soaking-assisted and thermal-pretreated cassava peel waste was investigated for bioethanol production [334]. Bioethanol was also produced from Calliandra calothyrsus shrub pretreated under hot water–based hydrothermal explosion [335]. Acid impregnation-steam explosion was used as a pretreatment method for ethanol production from oil palm empty fruit bunch (EFB) [336]. A series of ionic liquids including conventional, protic, and brønsted acidic type ionic liquids were evaluated as a source for the pretreatment of Taiwan grass [337]. Sonication after acid hydrolysis enhanced the total reducing sugar (TRS) extraction from sugarcane bagasse [338]. Cactus (CAC) and green and mature coconut shells were pretreated by NaClO2–C2H4O2 and sequential NaClO2–C2H4O2/autohydrolysis for their better utilization [339].
The combined effect of ionic liquid (1-butyl-3-methylimidazolium chloride, [BMIM]Cl) and radiation (Tungsten–Halogen) on hydrolysis of waste papaya epidermis was explored by Chatterjee et al. [340]. Chitosan-coated magnetic nanoparticle (C-MNP)–based immobilized laccase was applied for agave biomass pretreatment [341]. Ionic liquid 1-ethyl-3-methylimidazolium acetate pretreated with the Agave tequilana bagasse has shown promising results [342]. The modified thermochemical disk refining pretreatment (TCDRP) has a greater effect on agricultural biomass and hardwood (white birch) utilization [343].
Microwave-assisted lignin solubilized in protic ionic compounds containing 2,3,4,5-tetraphenyl-1H-imidazolium and inorganic anions was used as pretreatment [344]. For effective biomass utilization, mild photocatalyzed and catalyzed green oxidation of lignin was investigated. Lignins showed some mineralization when irradiated in the presence of H5[PMo10V2O40] × H2O (POM-1), K5[Ru(H2O)PW11O39] (POM-2), K4[SiW12O40]8H2O (POM-3), and TiO2 [345]. Dielectric barrier discharge (DBD) plasma can be used for the pretreatment of lignocellulosic materials. Such plasmas are sources of highly reactive species (radicals, ozone, atoms, ions, and excited molecules) [346]. Low-moisture anhydrous ammonia (LMAA) was used for the pretreatment of napier grass (Pennisetum purpureum Schumach) [347].
Electron beam irradiation–based biodegradation (EBIBB) was used for rice straw depolymerizations [348]. Microwave-assisted chemical pretreatment of miscanthus was studied [349]. Alkaline wet oxidation (WO) was used for the pretreatment of wheat straw, resulting in a formation of hemicellulose-rich hydrolysate and a cellulose-rich solid fraction [350].
Hydrothermal carbonization (HTC) is used as a method of pretreatment for efficient biomass utilization [351]. Phosphoric acid (1% w/w)–based steam explosion is used for olive tree pruning utilization [352]. Lower S/G ratio, as well as reduced number of phenolic OH group in acid-pretreated lignin, may affect simultaneous saccharification and fermentation [353]. S/G/H ratios of lignin fractions are important to rationalize the differences among the feedstock behavior [354]. Holocellulase immobilized on iron oxide (Fe2O3) nanoparticles improved hydrolysis of paddy straw. It has also been suggested that magnetic enzyme nanoparticle complexes (MENC) showed better immobilization efficiency (60–80%) for different enzymes [355]. Membrane-based separation increased the ethanol yields during fermentation which suggests the importance of separation after pretreatment [356]. The effect of ultrasound on enzymatic hydrolysis of a newspaper was investigated. The combined effect of ultrasound and enzymes lowers the diffusion-limiting barrier to enzyme/substrate binding to increase the reaction rate [357].
High solid loading hydrolysis (HSLEH) of sugarcane bagasse (SCB) pretreated by low-temperature aqueous ammonia soaking (AAS) was performed to obtain high concentrations of glucose [358]. Surfactant-assisted ionic liquid 1-ethyl-3-methylimidazolium acetate ([EMIM]OAc)–based pretreatment of beech wood waste was used for enhanced ethanol production [359]. WO (wet oxidation) and BM (ball milling)–based pretreatment of the macroalgae Chaetomorpha linum showed the highest ethanol yield [360].
Acid-catalyzed choline acetate ionic liquid pretreatment of paperboard mill sludge was used for effective production [361]. Zirconium metal–based organic framework (MOF)–assisted hydrothermally pretreated Platanus × acerifolia exfoliating bark was used for bioethanol production with promising results such as altered the morphology and higher the porosity and surface area [362].
Eucalyptus globulus and Nothofagus pumilio residues were pretreated with ionic liquids (IL) such as 1-N-ethyl-3-methylimidazolium chloride (C2mimCl) and 1-N-ethyl-3-methylimidazolium acetate (C2minOAc) for efficient conversion [363]. Steam-exploded and N-methylmorpholine-N-oxide (NMMO)–treated pinewood was investigated for biofuel production [364]. Two different acid-functionalized magnetic nanoparticles (MNPs), i.e., alkylsulfonic acid (Fe3O4-MNPs@[Si@AS) and butylcarboxylic acid (Fe3O4-MNPs@Si@BCOOH), were synthesized and evaluated for the pretreatment of sugarcane bagasse. Both Fe3O4-MNPs@Si@AS and Fe3O4-MNPs@Si@BCOOH showed the maximum amount of sugar (xylose) liberated, i.e., 18.83 g/L and 18.67 g/L, respectively [365].
A sequentially two-stage pretreatment process of autohydrolysis and alkaline extraction was carried out for effective bamboo utilization [366]. Phosphoric acid along with hydrogen peroxide (HP) pretreatment was employed on wheat straw for ethanol conversion [367]. Acidic ionic liquids (AILs) are a type of IL that has emerged as very attractive pretreatment solvents for biomass utilization [368]. Enhancement of enzymatic digestibility of Miscanthus biomass was improved by chemical-based electron beam irradiation [369]. The saccharification process was improved by gamma irradiation [370]. High pressure–assisted alkali pretreatment (HPAP) of cotton stalk led to the highest reducing sugar and ethanol yields (271.70 mg g−1 and 45.53%, respectively) [371].
Bioethanol production from ultrasonic irradiated waste newspaper by Saccharomyces cerevisiae was investigated by Preeti et al. [372]. Microwave irradiation accelerated pine cones act as a potential feedstock for bioethanol production [373]. Microwave-assisted ionic liquid–based catalytic conversion of non-edible lignocellulosic sunn hemp fibers to bioethanol was reported [374]. Bermuda grass, reed, and rapeseed were pretreated with phosphoric acid–acetone for ethanol production [375]. Two-stage pretreatment of Eucalyptus woody biomass with alkaline sulphonation and steam was carried out to enhance its enzymatic digestibility for bioethanol production [376]. Cellulase-bound magnetic nanoparticles were used as nanobiocatalyst for the hydrolysis of Sesbania aculeate biomass [377]. Microwave-assisted acid hydrolysis (H2SO4 and HCl with >0.5 mol/L) to produce bioethanol from sago pith waste (SPW) was studied [378]. The application of nano-biocatalyst (NiO) in simultaneous saccharification and fermentation of potato peel waste meaningfully enhanced bioethanol production (>65%) [379].
The nanofiltration (NF) and reverse osmosis (RO) membranes were chosen to evaluate their sugar rejection and inhibitor removal performance [380]. NaIO4 + H2SO4 and electron beam irradiation (EBI) pretreatment was used in the process to enhance the efficiency of straw conversion [381]. Alkali metal salt along with orthophosphoric acid was used for the pretreatment of microwave-assisted biomass to enhance sugar and bioethanol generation [382]. Organosolv pretreatment removes lignin from the biomass and makes the sugars available for conversion [383].
Pretreatment of sugarcane bagasse with liquid hot water (LHW) and aqueous ammonia (AA) showed better performance in terms of hemicellulose solubilization and lignin removal [384]. Ozonation of lignocellulosic waste (municipal trimmings) acts as an energetically suitable pretreatment method [385]. Two-stage dilute acid pretreatment was performed for effective utilization of Loblolly Pine [386]. Pycnoporus cinnabarinus–based laccase-mediator 1-hydroxybenzotriazole (HBT) was found effective for pretreatment of wheat straw [387]. The utilization of blue agave bagasse was enhanced under a combined extrusion–saccharification process [388]. Oxidative depolymerization along with acidic hydrolysis has consistently been used for the pretreatment of lignocellulosic biomass (wheat straw, sawdust, and lignin), which makes it possible to obtain a high content of soluble organic compounds in the hydrolysate (44–94 g COD/L) and to enhance the concentration of reducing sugars from 1 to 36% [389]. Peracetic acid–ionic liquid pretreatment was used for utilization of seaweed waste biomass from the carrageenan industry for bioethanol production [390]. During the pretreatment process, various toxic compounds may be generated that cause strong inhibition on cell growth and the metabolic capacity of fermenting strains. These are furan aldehydes, 2-furylaldehyde (furfural), and 5-hydroxymethyl-2-furaldehyde (HMF) produced by the degradation of pentose and hexose sugars respectively [391]. Inhibitory effects of phenolic components of spruce hydrolysates, viz. homovanilyl alcohol, vanillin, syringic acid, vanillic acid, gallic acid, dihydroferulic acid, p-coumaric acid, hydroquinone, ferulic acid, homovanillic acid, 4-hydroxybenzoic acid, 4-hydroxy-3-methoxycinnamaldehyde, and vanillylidenacetone, were investigated on the cell growth of Saccharomyces cerevisiae and it was observed that 4-hydroxy-3-methoxycinnamaldehyde was found to be the most toxic that inhibits the growth even at a very low concentration at 1.8 mM [392].
6.3 Genetic engineering aspects
For the holistic development of interesting microbes used in bioethanol production, genetic engineering could be playing a vital role. The host organism generally used for bioethanol production may not be tolerant of certain conditions such as temperature, pH, and ethanol stresses. Therefore, the host organisms used for bioethanol production need to be genetically engineered to make an effective and efficient condition for ethanol production. Recently, several genome engineering techniques have been developed. These techniques include (a) CRISPR/Cas system, (b) nuclease-based TALEN system, (c) zinc finger domain–based ZFN system, (d) meganuclease system, and (e) oligonucleotide-based YOGE system. Protein engineering studies as well as whole genome sequencing of bioethanol producers suggest that alteration of one or more nucleotides can bring out large changes in the direction of improved bioethanol production [393].
A lot of attention has also been focused on genetically engineered strains that can efficiently utilize both glucose and pentoses, and convert them to ethanol. Metabolic strategies seek to generate efficient biocatalysts (bacteria and yeast) for the bioconversion of most hemicellulosic sugars to products such as ethanol [394]. The biochemical production capacity of E. coli has been enhanced by the combinatorial application of recent approaches, viz. metabolic engineering, systems biology, synthetic biology, and evolutionary engineering [395].
CRISPR/Cas9 is used to disrupt the alcohol dehydrogenase-2 gene in Saccharomyces cerevisiae via complete deletion of the gene and introduction of a frameshift mutation in the ADH2 locus for improved ethanol yield [396]. For better utilization of xylose, metabolically engineered Saccharomyces cerevisiae was produced by integrating xylitol dehydrogenase gene (XYL2) into the chromosome [397]. A putative thermostable endoglucanase gene was inserted into a pET21 vector and transformed in E. coli BL21 for expression [398]. Bioethanol from lignocellulosic biomass requires robust Saccharomyces cerevisiae strains with improved tolerance capacity for toxic compounds. Genes (ADH6, HAA1, or PMA1) involved in detoxification and tolerance to inhibitors have been recognized. Overexpressing genes encode the transcription factor (YAP1) and the mitochondrial NADH-cytochrome b5 reductase (MCR1) for faster hexose catabolism [399].
Redox imbalance is the major challenge in the recombinant strains expressing S. stipitis XR-XDH pathway–based xylose-metabolizing cells, because xylose reductase prefers NADPH, whereas xylitol dehydrogenase strictly utilizes NAD+, leading to the accumulation of NADP+ and NADH. The ratio of NADP+/NADPH directly influences the activity of glucose-6-phosphate dehydrogenase and thereby affects sugar utilization [400]. Acremonium cellulolyticus was transformed (YKX1) by the β-xylosidase gene driven by the cellobiohydrolase Ι (cbh1) promoter by the protoplast-polyethyleneglycol (PEG) method. Now YKX1 can produce a higher amount of β-xylosidase [401].
Cloning of novel bacterial xylanases from lignocellulose-enriched compost metagenomic libraries was performed for the complete hydrolysis of lignocellulosic biomass into fermentable sugars [402]. To develop multiple stress tolerance (high-temperature and osmotic stress) in Saccharomyces cerevisiae, intracellular osmolyte glycerol production was quickly induced by osmotic shock due to overexpression of GPD1 and GPD2 genes encoding isoenzymes of NAD-dependent glycerol 3-phosphate dehydrogenase under the regulation of high osmolarity glycerol (HOG) pathway [403].
Tolerance for ethanol and heat stresses in Saccharomyces cerevisiae strains are important for industrial ethanol production. Genes accountable for ethano-thermotolerance were identified by transposon mutagenesis in Saccharomyces cerevisiae. Seven responsible genes (CMP2, IMD4, SSK2, PPG1, DLD3, PAM1, and MSN2) were identified. Knockout mutants of seven individual genes were ethanol tolerant whereas three of them (SSK2, PPG1, and PAM1) were tolerant to heat. The genes identified under this investigation may be helpful in the development of industrial yeast strains [404].
In another investigation, stress tolerance and the performance of ethanol fermentation of the four euploid strains were compared. Triploid showed a higher fermentation rate even in the presence of lignocellulosic hydrolysate–based inhibitors [405]. The thermotolerant Kluyveromyces marxianus is a potential candidate for high-temperature ethanol fermentation. At high temperatures, mitochondrial respiration is stimulated, leading to more reactive oxygen species (ROS) formation and lowered ratio of reduced NADH/oxidized NAD+ [406]. Overexpressed SNARE genes increased heterologous cellulase secretion in Saccharomyces cerevisiae. Soluble N-ethylmaleimide-sensitive factor attachment receptor proteins (SNAREs) play an important role in yeast protein-trafficking [407]. Adaptive evolution of xylose-fermenting Saccharomyces cerevisiae strains was performed with δ-integration of different xylA genes of the fungus Orpinomyces sp. and bacterium Prevotella ruminicola, thereby constructing two industrial S. cerevisiae strains, O7 and P5 [408].
A rumen metagenomic DNA fragment (Csd4) expressed in Escherichia coli MS04 improves ethanol fermentation. Csd4 acts as a saccharification enhancer to reduce the enzymatic load and operating time required for cellulose deconstruction [409]. A recombinant Saccharomyces cerevisiae strain was transformed with xylose reductase (XR) and xylitol dehydrogenase (XDH) genes from Pichia stipites; increment in ethanol production may be due to cofactor imbalance between NADPH-preferring XR and NAD+-dependent XDH [410]. Sometimes high hydrostatic pressure activates gene expression that leads to enhancement in ethanol production [411].
Proline acts as an osmotic stress protectant in yeast. Proline-accumulated S. cerevisiae cells were constructed by disrupting the PUT1 gene. Engineered strains revealed higher tolerance to many stresses, viz. freezing, desiccation, oxidation, and ethanol as well [412]. The efficient fermentation of glucose and xylose can improve by a two-stage transcriptional reprogramming (TSTR) strategy. The TSTR strategy improves ethanol production efficiency [376]. The thermotolerant methylotrophic yeast Hansenula polymorpha can ferment xylose, glucose, and cellobiose at elevated temperatures. Recombinant alcohol dehydrogenase 1 of H. polymorpha (HpADH1) overexpressed in Escherichia coli exhibited much higher catalytic efficiency for ethanol production [413]. The ethanol fermentation ability of the thermotolerant yeast Kluyveromyces marxianus (able to utilize glucose, mannose, galactose, xylose, and arabinose) was examined. It was found that KmGAL1 and KmXYL1 genes are responsible for sugar utilization [414].
Clustered regularly interspaced short palindromic repeats (CRISPR)–associated protein (CRISPR-Cas) technology with targeted genome editing exhibits a more precise and accurate gene knockout and knock-in system as compared to zinc finger nucleases (ZFN) and transcription activator-like effector nucleases (TALEN) [415]. Improvement in multiple stress tolerance capacity of yeast strain RPR39 by sequential mutagenesis (ethyl methane sulfonate, N-methyl-N′-nitro-N-nitrosoguanidine, near and far ultraviolet radiations) for enhanced bioethanol production [416]. Activation of β-glucosidase expression system in the multiple stress-tolerant (acid, ethanol and thermo) yeast Issatchenkia orientalis MF-121 strain for efficient ethanol production [417]. Formic acid–tolerant recombinant yeast strains were constructed by upregulation of formate dehydrogenase genes (FDH1 and FDH2) [418].
Furfural is one of the major inhibitors generated during bioethanol fermentation. Enhanced furfural tolerance of Tn 2 may be deliberated by the combined effect of lesser ROS (reactive oxygen sp) accumulation (early event) and an efficient detoxification of furfural (late event) [419]. For better biomass utilization, the endoglucanase I and II genes (egI or Cel7B and egII or Cel5A) of Trichoderma reesei QM6a were successfully cloned and expressed in Saccharomyces cerevisiae under the transcriptional control of the yeast ENO1 promoter and terminator sequences [420]. Co-expression of a cellobiose phosphorylase and lactose permease allows intracellular cellobiose utilization by Saccharomyces cerevisiae [421]. Scheffersomyces stipitis strain expressing xylose reductase–xylitol dehydrogenase (XR-XDH) pathway under adaptive evolution treatment affects sulfur amino acid biosynthesis and redox stress as well. These findings provide new insights for engineered bioethanol-producing strain through reverse metabolic engineering [422]. Neotermes koshunensis (termite) secretes endogenous β-glucosidase in the salivary glands and this was successfully expressed in Aspergillus oryzae [423].
A xylose-metabolizing yeast was constructed by the integration of XI overexpression cassettes into the genome of the Saccharomyces cerevisiae MT8-1 strain. Knockout of GRE3 (a gene encoding nonspecific aldose reductase) of the host yeast strain improved ethanol productivity [424]. A recombinant of xylose and cellooligosaccharide-assimilating yeast strain has been constructed by integrating genes responsible for the expressions of xylose reductase and xylitol dehydrogenase from Pichia stipitis, and xylulokinase from Saccharomyces cerevisiae as well as β-glucosidase from Aspergillus acleatus on the cell surface [425]. Improvement in tolerance of Saccharomyces cerevisiae for hot-compressed water-treated cellulose by expression of ADH1 [426]. Pyruvate decarboxylase (PDC) of Gluconobacter oxydans was considered to be a suitable candidate for heterologous expression in the thermophile Geobacillus thermoglucosidasius for ethanol production [427]. A combination of UV mutagenesis and protoplast fusion was used to construct strains with improved stress performance [428]. One major barrier to the economic conversion of biomass to ethanol is the inhibitory compound such as furfural and 5-hydroxymethylfurfural (HMF). Ethanologenic yeasts undergo a genomic adaptation process during the adaptation phase for various inhibitors [429].
SUMOylation acts as a novel potential mechanism to reduce the multiple inhibitory effects of fermentation inhibitors by regulating the lag phase [430]. More oleic acid in the plasma membrane contributes to the acetic acid tolerance of yeast [431]. Ethanol production from xylose is improved by mating recombinant xylose-fermenting Saccharomyces cerevisiae strains. Xylose-fermenting, haploid, yeast cells of the opposite mating type were hybridized to produce a diploid strain hiding two sets of xylose-assimilating genes encoding xylose reductase, xylitol dehydrogenase, and xylulokinase resulting in improvements in fermentation ability [432]. Zymobacter palmae directly fermented cellulosic materials by co-expressing foreign endoglucanase and β-glucosidase genes [433].
Engineered microbes with vgb/VHb could be useful in enhancing bioethanol production [434]. Ethanol is directly produced at high temperature using the thermotolerant yeast Kluyveromyces marxianus displaying cellulolytic enzymes. The strain was genetically engineered to display Trichoderma reesei endoglucanase and Aspergillus aculeatus β-glucosidase on the cell surface [435]. Genetically engineered Zymomonas mobilis efficiently produced bioethanol from the hydrolysate of wood biomass containing glucose, mannose, and xylose as major sugar components [436]. Pyruvate decarboxylase (pdc) and alcohol dehydrogenase II (adhII), from Zymomonas mobilis, were heterologously expressed in the gram-positive bacterium Streptomyces lividans TK24 [437]. A novel endoglucanase encoding gene was cloned from Alicyclobacillus vulcanalis and expressed in E. coli for hydrolysis [438]. A recombinant S. cerevisiae strain is constructed displaying phytase on the cell surface which could improve ethanol production performance and effectively reduce the discharge of phosphorus [439].
A novel agglutinin expression system is constructed as well as immobilization β-glucosidase1 on the surface of wild-type Saccharomyces cerevisiae Y5 exhibiting a strong bioethanol fermentation capacity [440]. A novel aldehyde reductase encoded by YML131W from Saccharomyces cerevisiae gives tolerance to furfural derived from lignocellulosic biomass conversion [441]. The expression of cellulolytic genes is elicited using a recombinant endoxylanase from Trichoderma harzianum IOC-3844 [442]. Engineering of cellulolytic Saccharomyces cerevisiae strains is a promising way for lignocellulosic ethanol production [443]. Komeshu et al. used genetically engineered microbes for bioethanol production [444]. The activities and thermostabilities of the four PPP enzymes (transaldolase: TAL1, transketolase: TKL1, ribose-5-phosphate ketol-isomerase: RKI1, and d-ribulose-5-phosphate 3-epimerase: RPE1) can affect the efficiency of cellulosic ethanol production. Strains that overexpressed S. cerevisiae TKL1 exhibited the highest rate of xylose consumption [445].
Disruption of the alkaline phosphatase gene PHO13 enhances ethanol production by a strain expressing the xylose reductase (XR) and xylitol dehydrogenase (XDH) gene [446]. S. cerevisiae strain co-expressing genes for several cell surface cellulases and the cellodextrin transporter was constructed to improving the efficiency of direct ethanol fermentation [447]. A novel xylose-fermenting yeast strain, FSC1, was developed for ethanol production by intergeneric hybridization between S. cerevisiae and Candida intermedia mutants by using a protoplast fusion technique [448]. Ethanolic xylose fermentation is controlled by the XR activity. Xylose transport also plays an important role in ethanol production [449]. A novel Clostridium thermocellum cellulolytic recombinant cellulase is expressed in Escherichia coli cells [23].
Overexpression or deletion of genes enhances acetic acid tolerance. Strains overexpressing ASC1 and GND1 displayed enhanced tolerance to acetic acid [450]. Multifunctional β-glucosidase/β-xylosidase/α-arabinosidase (Bgxa1) is found as an interesting candidate for the saccharification of lignocellulosic material [451]. Overexpression of GRE2 gene from Scheffersomyces stipitis to Saccharomyces cerevisiae as an aldehyde reductase contributes tolerance to aldehyde inhibitors produced from lignocellulosic biomass. GRE2 can reduce furfural to FM and reduce hydroxymethyl furfural to FDM [452]. Expression of dehydrin gene from Arctic Cerastium arcticum increases abiotic stress tolerance and fermentation capacity of a genetically engineered Saccharomyces cerevisiae [453]. It has been demonstrated that S. cerevisiae has the ability of in situ detoxification of aldehydes (furan, aliphatic, and phenolic) to their corresponding less toxic alcohols by the action of NAD(P)H-dependent aldehyde reductases [454].
Heterologous genes for xylose utilization were introduced into an industrial Saccharomyces cerevisiae [455]. Overexpression of PMA1 enhances tolerance to various types of stress and constitutively activates the SAPK pathways in Saccharomyces cerevisiae [456]. Restitution of the NAD+/NADH redox balance plays a vital role in ethanol stress response [457]. Multiple gene-mediated NAD(P)H-dependent aldehyde reduction is a mechanism of in situ detoxification of furfural and 5-hydroxymethylfurfural by Saccharomyces cerevisiae [458]. Overexpression of native Saccharomyces cerevisiae exocytic SNARE genes increased cellulase secretion. SNAREs (soluble NSF [N-ethylmaleimide-sensitive factor attachment receptor proteins) are required for fusion events under intracellular membrane transport and facilitate protein trafficking between the various membrane-enclosed organelles and the plasma membrane [407]. Co-expression of TAL1 and ADH1 in recombinant Saccharomyces cerevisiae improves ethanol production from lignocellulosic hydrolysates [459].
Deletion of the PHO13 gene in Saccharomyces cerevisiae improves ethanol production from lignocellulosic hydrolysate in the presence of acetic and formic acids, and furfural [460]. PRS3, RPB4, and ZWF1 were identified as key genes for yeast tolerance to lignocellulose-derived inhibitors or multiple stresses [461]. Ethanol production is improved through decreased glycerol synthesis in Saccharomyces cerevisiae by metabolic and genetic engineering approaches. Glycerol production was hindered by the deletion of the most important GPD genes involved in glycerol production [462]. The Msn2 overexpression of various antioxidant enzyme genes in microbial strain showed tolerance to oxidative stress during ethanol production [463].
A recombinant S. cerevisiae strain (SK-NY), overexpressing GRE3-encoded NADPH-dependent aldose reductase and NADP+-dependent xylulokinase, was constructed for efficient bioethanol production [464]. The redox balance between xylose reductase (XR) and xylitol dehydrogenase (XDH) is an important parameter for effective xylose fermentation. Xylitol accumulation is reduced and ethanol production is improved by reversing the dependency of XDH from NAD+ to NADP+ [465]. Efficient xylose-fermenting Saccharomyces cerevisiae is constructed through a synthetic isozyme system of xylose reductase from Scheffersomyces stipites. The xylose-metabolic genes (XYL1, XYL2 and XYL3) from Scheffersomyces stipitis have been engineered into S. cerevisiae [466].
Ethanol production from xylose in the presence of acetic acid is improved by the overexpression of HAA1 gene and the deletion of PHO13 gene in Saccharomyces cerevisiae [467]. Improved sucrose metabolism by overexpressing invertase is an attractive strategy to improve ethanol yields. The promoter and 5′ coding sequences of SUC2 are engineered, resulting in (94%) cytosolic localization of invertase [468]. Co-consumption of multiple sugars can be attained by modulating phosphotransferase system (PTS); this may be improved by amplifying the non-PTS pathway genes such as galP and glk [469]. The xylose utilization capability of Saccharomyces cerevisiae was enhanced by applying the concept of inverse metabolic engineering to identify the factors involved in improving xylose utilization. It has been observed that deletion of molecular chaperone-encoding genes HSP26, SSA1, and HSP104 facilitates the protein folding of xylose isomerase and enhancing xylose isomerase activity [470]. Saitoh et al. [471] constructed the triple auxotrophic strain OC2-HUT and introduced cell surface–displaying β-glucosidase (BGL) gene and a xylose-assimilating gene to generate the final strain OC2-ABGL4Xyl for efficient ethanol production. Improved xylose isomerase activity, upregulation of glycolysis and glutamate synthesis enzymes, and downregulation of trehalose and glycogen synthesis altogether contribute to the effective xylose utilization by the strain [472].
Co-fermentation of cellulose/xylan was investigated by the engineered industrial yeast strain OC-2 displaying both β-glucosidase and β-xylosidase [473]. Saccharomyces cerevisiae strain was engineered for xylose assimilation by the constitutive overexpression of the Orpinomyces xylose isomerase, S. cerevisiae xylulokinase, and the Pichia stipitis SUT1 sugar transporter genes [474]. To enhance heterologous cellulase protein production in yeast, a plasmid embracing the endoglucanase gene from Clostridium thermocellum (Ctcel8A) was used to transform a homozygous diploid yeast [475]. Saccharomyces cerevisiae strain was engineered with a three-plasmid SUMO yeast expression system by utilizing the portable small ubiquitin-like modifier (SUMO) vector set combined with the efficient endogenous yeast protease Ulp1 [476]. Transaldolase and transketolase are the key enzymes responsible for non-oxidative pentose phosphate pathway–based xylose utilization in recombinant Saccharomyces cerevisiae. Overexpression of TAL1 (transaldolase gene) and TKL1 (transketolase gene) increases the flux from the pentose phosphate pathway into the glycolytic pathway [477].
Improvement in cellulase production can also be done by modifying regulator expression in T. harzianum [478]. PfMig188, a catabolically derepressed engineered strain of the hyper-cellulolytic fungus Penicillium funiculosum NCIM1228, was investigated. Results demonstrated that the PfMig188 secretome has relatively broad substrate specificity and acts as an efficient substitute for T. reesei–based secretomes for diverse biomass saccharification [479].
Deletion of the HXK2 gene (a moonlighting protein) in Saccharomyces cerevisiae enables mixed sugar fermentation of glucose and galactose (major sugar components of red seaweeds) in oxygen-limited conditions [480]. Overexpression of native PSE1 and SOD1 genes under the transcriptional control of the constitutive PGK1 promoter in Saccharomyces cerevisiae improved heterologous cellulase secretion. The effect of these genes on heterologous protein secretion of three cellulases—an exoglucanase encoded by cel6A of Neocallimastix patriciarum, a β-glucosidase encoded by cel3A of Saccharomycopsis fibuligera, and an endoglucanase encoded by cel7B of Trichoderma reesei—was investigated by integrating the PGK1P/T–PSE1 and PGK1P/T–SOD1 cassettes into S. cerevisiae strains to produce the relevant cellulases [481]. A novel β-glucosidase gene encoding a protein (BglA) of 446 amino acid, belonging to the glycoside hydrolase family 1 (GH1), was cloned from a hyperthermophilic bacterium Thermotoga naphthophila RKU-10T and overexpressed in Escherichia coli BL21CodonPlus. All these significant features make BglA an appropriate candidate for biotechnological and industrial applications [482].
6.4 Attachment on cell surface aspects
Yeast cell surface engineering enables more than 100 enzymes to be displayed on the surface of a yeast cell. The displaying yeast can be used as a whole-cell biocatalyst without requiring enzyme separation and purification processes [483]. Construction of a new system for endoglucanases exhibiting carbohydrate-binding modules using yeast cell surface engineering. Saccharomyces cerevisiae BY4741 (Δ sed1) exhibiting 3 cellulases (Trichoderma reesei endoglucanase II [EG], T. reesei cellobiohydrolase II [CBH], and Aspergillus aculeatus β-glucosidase I [BG]) was constructed by yeast cell surface engineering [484]. For making more efficient bioethanol production, the endoglucanase gene endo753 of Aspergillus flavus NRRL3357 was introduced on the cell surface of Saccharomyces cerevisiae EBY100 strain by the C-terminal fusion using Aga2p protein as an anchor attachment tag [485].
Heterologous cellulolytic enzymes were expressed on the Z. palmae cell surface by cell surface display motif of the Pseudomonas ice nucleation protein N-terminal anchoring [486]. Yeast strain was engineered by codisplaying several hemicellulolytic enzymes on the surface of xylose-utilizing Saccharomyces cerevisiae cells [487]. A recombinant was developed by expressing three cellulases from Clostridium cellulolyticum—endoglucanase (Cel5A), exoglucanase (Cel9E), and β-glucosidase—on the surface of the Escherichia coli LY01 [488].
6.5 By checking bacterial contamination
The presence of bacterial contaminants reduces alcoholic fermentation. Antibiotics are currently used to control contamination, but their residues may be detected; therefore the antimicrobial activity of the natural compounds such as hops extract, 4-hydroxybenzoic acid, nisin Z, and lysozyme were investigated and found their great potential for the substitute of antibiotics used conventionally in the ethanol industry [489]. Bioethanol fermentation is usually suppressed by lactic acid bacteria (LAB), thereby leading to a decrease in bioethanol yield. Nisin-loaded P4VP microspheres were added into the simulated contaminative fermentation system for controlling the L. plantarum contamination in bioethanol fermentation [490].
Bacteriophage can be used as a potential alternative agent for controlling Lactobacillus plantarum contamination during bioethanol production. Moreover, increased concentrations of monounsaturated fatty acids due to bacteriophage treatment might lead to more membrane fluidity and promote the cell viability of S. cerevisiae [491]. Bacteriocins, bacteriophages, and beneficial bacteria are used as a non-conventional antimicrobial agents to reduce bacterial contamination in the bioethanol industry [492]. Brettanomyces/Dekkera bruxellensis–based microbial contamination of ethanol fermentation has been controlled by saccharomycin (biocide composed of antimicrobial peptides) secreted by Saccharomyces cerevisiae [493]. Transcriptional profile (genes with significantly repressed or induced expression) of a bioethanol production contaminant can provide information on antimicrobials, to combat yeast contamination during industrial bioethanol production [494].
6.6 To develop ethanol and acetate tolerance
Alcohol toxicity is a more serious problem for bioproduction using bacteria. Alcohols interact directly with the lipid bilayer because of their amphiphilicity and thereby, membrane fluidity is altered. These changes in fluidity increase membrane permeability and induce conformational changes in membrane proteins. Ethanol-induced membrane induces the expression of heat-shock and phage-shock proteins. Transcriptomic analyses identified important roles of the groESL chaperone system and the global regulator of sporulation in alcohol tolerance [495]. Higher ethanol concentration in Saccharomyces cerevisiae leads to cell growth inhibition and ultimately cell death. Several mechanisms, viz. changes in gene expression, membrane composition, and increment in chaperone proteins, help stabilize other denatured proteins [496].
Ethanol and acetate accumulation under the fermentation process affects ethanol yield by stressing the metabolic capabilities of the microorganisms. Such conditions can be regulated by overexpression of the iron-sulfur cluster (ISC) in the E. coli KO11 strain [497, 498]. Green tea polyphenols (GTP) enhance the ethanol tolerance of S. cerevisiae may be due to the significantly differentially expression of large amounts of genes related to the cell wall, cell membrane, basic metabolism, and redox regulation [499]. Amend effect of Cyclocarya paliurus (C. paliurus) triterpenoids on S. cerevisiae under the ethanol stress was explored. It has been observed that the treatment of triterpenoids enhances ethanol tolerance of S. cerevisiae [500].
Bacterial signals of N-acyl homoserine lactones induce the changes of S. cerevisiae morphology, thus making it more ethanol tolerant. Bacterial signals QSMs (quorum signal molecules) of N-acyl homoserine lactones induce the changes of morphology and ethanol tolerance in Saccharomyces cerevisiae. Microbes communicate with each other using chemical signal molecules, termed autoinducers (AI) or quorum sensing molecules (QSM). When the signal molecules accumulate a threshold, the communicating microbes begin to alter gene expression and therefore behavior in response. Saccharomyces cerevisiae, exposed to short-chain 3-OC6-HSL and long-chain C12-HSL, showed obvious changes in morphology and ethanol tolerance [501]. Issatchenkia orientalis, a non-Saccharomyces yeast that can resist a wide variety of environmental stresses (ethanol stress), has potential use in bioethanol production [502]. The metabolic differences of diploid (α/a) and haploid (α, a) yeasts in response to ethanol stress were recently studied. It was found that the haploid genotype being more susceptible to ethanol stress as compared to diploid may be due to its higher content of protective metabolites including polyols [503]. Antiseptics such as hydrogen peroxide, potassium metabisulfite, and 3,4,4-trichlorocarbanilide have been shown to inhibit and control bacterial contamination in ethanol fermentations [504].
Stress-tolerant Saccharomyces cerevisiae strains are developed by metabolic engineering through cell flocculation and zinc supplementation [505]. Metabolically engineered Saccharomyces cerevisiae strain showed tolerance for acetic and formic acids. Improved activities of transaldolase (TAL) and formate dehydrogenase (FDH) through metabolic engineering successfully deliberated resistance to weak acids in a recombinant xylose-fermenting Saccharomyces cerevisiae strain [506]. Plasma membrane proteins Yro2 and Mrh1 are required for acetic acid tolerance in Saccharomyces cerevisiae [507]. Creation of yeasts with acid tolerance was successful using yeast cell surface engineering. The cell wall of Saccharomyces cerevisiae plays a crucial role in the biophysical characteristics of the cell surface. The modification of the cell wall property is an important factor for adaptation under a stressful environment. A novel peptide, Scr35, that provides acid tolerance in yeasts was obtained [508].
Expression of a salt-induced 2-Cys peroxiredoxin from Oryza sativa improves stress tolerance in the recombinant yeast Saccharomyces cerevisiae. Peroxiredoxins (Prxs) are a thiol-specific antioxidant enzymes that are seriously involved in cell defense and protect cells from oxidative damage [509]. Adaptive laboratory evolution (ALE) was used for the development of furfural and acetic acid–tolerant strain [510]. Rpn4 and proteasome-mediated yeast resistance to ethanol includes regulation of autophagy. It has been suggested that Rpn4 affects the autophagic system activity upon ethanol stress through the PRB1 regulation [511]. One approach to alleviate the inhibition problem is to use genetic engineering to introduce increased tolerance by overexpression of Saccharomyces cerevisiae Pad1p. The overexpressing transformants showed approximately tenfold higher activity [512].
Acetate is an effective agent for the prevention of bacterial contamination, but it negatively affects the fermentation ability of S. cerevisiae. Overexpression of the organic acid–tolerant HAA1 gene, which encodes a transcriptional activator, could be a useful molecular breeding method for acetate-tolerant yeast strains [513].
7 Future prospects
To develop a successful and commercially viable technology for bioethanol production, novel strategies for generating enzyme cocktails for lignocellulose hydrolysis in biorefineries would be developed by enzyme engineering, reconstitution of enzyme mixtures, and bioprospecting for superior enzymes. The current situation warrants the need for integrated research and development of lignocellulosic biomass utilization [514]. Understanding of the complexity of lignocellulose feedstock, their chemical compositions, and the knowledge of pretreatment methods are required [19].
Functional metagenomic strategies for the finding of novel enzymes for biomass hydrolysis and biofuel production such as new-generation sequencing and mining the metagenome are becoming more efficient [515]. In recent years, post-genomic approaches such as metabolomics in combination with other omics such as genomics, chemogenomics, transcriptomics, and proteomics were studied to boost the use of systems metabolic engineering tools in industrial settings. It provides insights of the mechanisms and interactions of genes and allowed to better understand under severe environments, overexpression and downregulation of multiple genes, and construction of synthetic regulatory proteins and other components such as ethanol and acetic acid tolerance [495, 498, 516,513,518].
Permeases are directly involved in the utilization of and regulatory response to nutrient sources. Permease regulatory mechanisms on yeast metabolic engineering provide important insights for the elimination of harmful substances in S. cerevisiae [519]. S. cerevisiae is frequently challenged by bacterial contamination and a combination of lignocellulosic inhibitors formed during the pretreatment so it can be checked [520]. Required robust Saccharomyces cerevisiae strains with improved capacity to cope with the toxic compounds formed during the biomass pretreatment, among which are 5-hydroxymethylfurfural (HMF), furfural, weak organic acids, and phenolic compounds generated [399].
Recently, global transcription machinery engineering (gTME) has been applied as an effective technique to enhance the target specific phenotype of microbes for enhanced ethanol production. The gTME uses random mutagenesis libraries of global transcription factors generated by error-prone PCR to reprogram transcription and obtain specific phenotypes. Ep-PCR is a fast and cheap molecular biology method for the random mutation in a particular piece of DNA [495, 521]. Microorganisms have great potential for the engineering and/or incorporation of complete metabolic pathways for the over production of value-added chemicals. Despite the wide capability of microorganisms, there is still a lack of knowledge about the metabolic networks responsible for such processes. Metabolomics are used for microbial strain selection and engineering novel biochemical pathways strictly responsible for efficient biomass conversion [503, 522].
Xylose acts as the second most prevalent sugar after glucose in lignocellulosic biomass utilization; therefore, extensive research efforts have been made and still needed to introduce heterologous genes for xylose metabolism into S. cerevisiae. For this reason, detailed studies about naturally xylose-fermenting yeasts species (Scheffersomyces stipitis or Pachisolen tannophilus), comparative genomics, and evolutionary analysis are needed as effective approaches to determine the limiting steps in pentose metabolism. Overexpression of genes encoding enzymes of non-oxidative pentose phosphate pathway (PPP) and replacement of a small amount of enzymes of xylose metabolism, as well as isolation of xylose transporters were pointed out as crucial factors for the adequate function of this pathway [522, 523].
The combination of metabolomics, fluxomics, and synthetic biology is used as a powerful tool for prospecting novel metabolic routes [524, 525]. Designer cellulosomes (also known as chimeras) unlike native cellulosomes are artificial constructs, composed of chimeric scaffoldin and enzymes with cohesins and dockerins of divergent specificities, thus providing interdomain flexibility in the enzyme complex for effective utilization [21]. Recent trends in ligninolytic green biotechnology by immobilization engineering processes suggest the potential industrial applications of ligninolytic enzymes in various sectors of the modern industry [526].
Nanoparticles are gaining increasing interest among researchers due to their exquisite properties. They are also being explored in biofuels to improve the performance of bioethanol production. Different types of nanomaterials (metallic, nanofibers, and nanotubes) have been used and they can effectively suppress inhibitory compounds under certain conditions [527]. Metabolic engineering is used of microorganisms for biofuel production. Metabolically engineered yeasts in surface displaying various hydrolytic enzymes appear to hold the greatest potential. The bacterium Zymomonas mobilis metabolically engineered to make bioethanol from pentose sugars is already being commercialized [51, 415, 528].
Chromosomal integration of genetic material is the preferred method to overcome gene loss that may occur by homologous recombination. The use of CRISPR/Cas9 provides a marker-free genome-editing tool and thereby, it should open a new avenue in creating microbial biorefineries for enhanced bioethanol production by engineering the microbial genomes for desired traits such as enhancing the biofuel tolerance, inhibitor tolerance, and thermotolerance as well as modifying the cellulases and hemicelluloses enzymes [415, 529].
However, more research on the metabolic pathways, regulation of end-product formation, and construction of genetically engineered thermophilic/thermotolerant microorganisms with high tolerance to ethanol is required for industrial fermentations. Evolutionary engineering is another approach for development. It uses laboratory evolution for selecting industrially relevant traits. By integrating whole genome sequencing, bioinformatics, classical genetics, and genome-editing techniques, evolutionary engineering has also become a powerful approach for the identification and reverse engineering of molecular mechanisms that underlie industrially relevant traits [524].
Adaptive laboratory evolution (ALE) is a powerful tool for analyzing phenotypic and genotypic changes during bacterial evolution. In this approach, cells are cultured under a selective environment for many generations, leading to adaptive evolution [495]. DNA microarrays can also be used in detecting transcription factor binding sites and single-nucleotide polymorphisms. Target genes for genetic manipulation should be identified to confer useful phenotypes, such as stress tolerance and high fermentation activity, and to improve the production of the target product [530]. It is possible to evaluate the effects of specific mutations on alcohol tolerance using genome-editing technology [495].
Genome shuffling is an efficient way to improve complex traits or phenotypes under the control of multiple genes. Genome shuffling is the best way for strain improvement in very high-gravity (VHG) fermentation [531, 532]. A better perception of the yeast adaptation under multiple stresses is of critical importance to develop strategies to improve yeast robustness and bioconversion capacity from lignocellulosic biomass [533]. Design and engineer a strain with a high secretory phenotype, bioethanol-specific stressors, including tolerance to products formed during hydrolysis of lignocellulosic substrates [534]. Industrial yeast strains with better xylose fermentation ability and stress tolerance are important for economical lignocellulosic bioethanol technology [535]. Isolated strains of Saccharomyces cerevisiae from different sources display extensive genetic and phenotypic diversity. To better understand how genomic changes influence phenotypes is more important for developing strategies. Whole genome sequencing was carried out based on single-nucleotide variations and small insertions/deletions/annotations in the genome. Phylogenetic analysis also recommended the unique genes, obtained through horizontal gene transfer from other species. RNA-Seq analysis also suggests that sometimes unique genes are not functional due to unidentified intron sequences [376].
As an efficient ethanol-producing bacterium, Zymomonas mobilis has created special attention due to several properties, viz. high sugar uptake, ethanol yield, and tolerance. Different metabolic engineering strategies have been used to create a new metabolic pathway for Z. mobilis to broaden its application range [51]. Expressing hydrolase in the lignocellulosic feedstock is a favorable alternative, due to the large availability of biomass [5]. The mixed feedstock approach to lignocellulosic ethanol production has shown that their use can bring about significant cost savings as compared to single feedstock utilization [536]. Stress tolerance in industrial yeast strains is an important point for cost-effective bioprocessing. It optimizes microbial systems to adapt under environmental stresses and thus has a huge power of the creation of robust stress-tolerant yeasts [537]. The future of lignocellulosic biomass would be based on improvements of plant biomass, metabolic engineering of ethanol production pathway and hydrolytic enzyme-producing microorganisms, and the fullest exploitation of waste biomass and process integration of the individual steps involved in bioethanol production [538].
8 Conclusions
Lignocellulosic biomass, as a waste material, offers an attractive alternative for its valorization into valuable products. However, the recalcitrance of these materials and the inability of microorganisms to efficiently ferment each sugar present as well as lignocellulosic hydrolysates still lower the production of bioethanol. The viability of lignocellulosic material for ethanol production has been still searched around the world depending upon its availability. More attention is required on the development of sustainable and scalable fuels from renewable biomass, viz. agricultural and industrial residues, as a means to curb global warming. There is an intense emphasis on lowering the costs of renewable bioethanol production by overcoming the challenges connected to high substrate costs, limited microbial capacity, stress tolerance, low titers, and low production rates.
References
Ali N, Zhang Q, Liu Z et al (2020) Emerging technologies for the pretreatment of lignocellulosic materials for bio-based products. Appl Microbiol Biotechnol 104:455–473. https://doi.org/10.1007/s00253-019-10158-w
Singh YD (2019) Cellulosic bioethanol production from Eragrostis airoides Nees grass collected from Northeast India. SN Appl Sci 1:889. https://doi.org/10.1007/s42452-019-0952-z
Verma N, Bansal MC, Kumar V (2011) Pea peel waste: a lignocellulosic waste and its utility in cellulase production by Trichoderma reesei under solid state cultivation. Bioresources 6(2):1505–1519
Bhatia L, Johri S, Ahmad R (2012) An economic and ecological perspective of ethanol production from renewable agro waste: a review. AMB Express 2:65. https://doi.org/10.1186/2191-0855-2-65
Xiao Y, Poovaiah C, Coleman HD (2016) Expression of glycosyl hydrolases in lignocellulosic feedstock: an alternative for affordable cellulosic ethanol production. Bioenerg Res 9:1290–1304
Khan MI, Lee MG, Shin JH et al (2017) Pretreatment optimization of the biomass of Microcystis aeruginosa for efficient bioethanol production. AMB Express 7:19. https://doi.org/10.1186/s13568-016-0320-y
Dantur KI, Enrique R, Welin B et al (2015) Isolation of cellulolytic bacteria from the intestine of Diatraea saccharalis larvae and evaluation of their capacity to degrade sugarcane biomass. AMB Express 5:15. https://doi.org/10.1186/s13568-015-0101-z
Luque R, Clark JH (2013) Valorisation of food residues: waste to wealth using green chemical technologies. Sustain Chem Process 1:10. https://doi.org/10.1186/2043-7129-1-10
Shimoda T, Shirouchi T, Suzuki A et al (2012) Storage of maitake mushroom (Grifola frondosa) culture medium after harvesting fruit bodies is an effective pretreatment for ethanol conversion. J Wood Sci 58:342–351. https://doi.org/10.1007/s10086-012-1254-x
Favaro L, Basaglia M, Casella S (2012) Processing wheat bran into ethanol using mild treatments and highly fermentative yeasts. Biomass Bioenergy 46:605–617
Miezah K, Obiri-Danso K, Kádár Z et al (2017) Municipal solid waste management in a low income economy through biogas and bioethanol production. Waste Biomass Valor 8:115–127. https://doi.org/10.1007/s12649-016-9566-5
Kumar AK, Parikh BS, Singh SP et al (2015) Use of combined UV and chemical mutagenesis treatment of Aspergillus terreus D34 for hyper-production of cellulose-degrading enzymes and enzymatic hydrolysis of mild-alkali pretreated rice straw. Bioresour Bioprocess 2:35. https://doi.org/10.1186/s40643-015-0062-8.
Verma N, Kumar V, Bansal MC (2020) Comparative view on microbial consumption of agro-based lignocellulosic waste biomass in sustainable production of cellulases. Biomass Conv Bioref. https://doi.org/10.1007/s13399-020-00617-0
Reis ALS, de Fátima Rodrigues de Souza R, Baptista Torres RRN et al (2014) Oxygen-limited cellobiose fermentation and the characterization of the cellobiase of an industrial Dekkera/Brettanomyces bruxellensis strain. SpringerPlus 3:38. https://doi.org/10.1186/2193-1801-3-38
Bagewadi ZK, Mulla SI, Ninnekar HZ (2017) Optimization of laccase production and its application in delignification of biomass. Int J Recycl Org Waste Agric 6:351–365. https://doi.org/10.1007/s40093-017-0184-4
Verma N, (2010) Bioethanol from biomass: a review. 1, Issue : 2, 245-256.
Koti S, Govumoni SP, Gentela J et al (2016) Enhanced bioethanol production from wheat straw hemicellulose by mutant strains of pentose fermenting organisms Pichia stipitis and Candida shehatae. SpringerPlus 5:1545. https://doi.org/10.1186/s40064-016-3222-1
Verma N, Kumar V, Bansal MC (2019) Utility of starchy, lignocellulosics and cellulosics hydrolysates on cellulase production under liquid state fermentation. Waste Dispos Sustain Energy 1:289–299. https://doi.org/10.1007/s42768-019-00019-7
Madu JO, Fabunmi TE, Agboola BO (2019) Effects of chemical treatments on the bioethanol yield and composition of Isoberlinia doka waste. SN Appl Sci 1:216. https://doi.org/10.1007/s42452-019-0223-z
Bak JS (2015) Downstream optimization of fungal-based simultaneous saccharification and fermentation relevant to lignocellulosic ethanol production. SpringerPlus 4:47. https://doi.org/10.1186/s40064-015-0825-x
Obeng EM, Adam SNN, Budiman C et al (2017) Lignocellulases: a review of emerging and developing enzymes, systems, and practices. Bioresour Bioprocess 4:16. https://doi.org/10.1186/s40643-017-0146-8
Eardley J, Timson DJ (2020) Yeast cellular stress: impacts on bioethanol production. Fermentation 6:109. https://doi.org/10.3390/fermentation6040109
Das SP, Deka D, Ghosh A et al (2013) Scale up and efficient bioethanol production involving recombinant cellulase (Glycoside hydrolase family 5) from Clostridium thermocellum. Sustain Chem Process 1:19. https://doi.org/10.1186/2043-7129-1-19
Olguin-Maciel E, Singh A, Chable-Villacis R, Tapia-Tussell R, Ruiz HA (2020) Consolidated bioprocessing, an innovative strategy towards sustainability for biofuels production from crop residues: an overview. Agronomy 10:1834. https://doi.org/10.3390/agronomy10111834
Pana L, Heb M, Wu B, Wang Y, Hu G, Maa K (2019) Simultaneous concentration and detoxification of lignocellulosic hydrolysates by novel membrane filtration system for bioethanol production. J Clean Prod 227(1):1185–1194
Matsushika A, Inoue H, Kodaki T et al (2009) Ethanol production from xylose in engineered Saccharomyces cerevisiae strains: current state and perspectives. Appl Microbiol Biotechnol 84:37–53. https://doi.org/10.1007/s00253-009-2101-x
Nielsen J, Larsson C, Maris A, Pronk J (2013) Metabolic engineering of yeast for production of fuels and chemicals. Curr Opin Biotechnol. 24(3):398–404
Ziegler I., Zahra D et al. (2011) Steam explosion pretreatment of willow grown on phytomanaged soils for bioethanol production. Ind Crops Prod 140.
Sittijundaa S, Faria A et al (2013) Ethanol production from glucose and xylose by immobilized Thermoanaerobacter pentosaceus at 70 °C in an up-flow anaerobic sludge blanket (UASB) reactor. Bioresour Technol 143:598–607
Gonçalves FA, Ruiz HA, Santos ESD, Teixeira JA, Macedo GRD (2016) Bioethanol production by Saccharomyces cerevisiae, Pichia stipitis and Zymomonas mobilis from delignified coconut fibre mature and lignin extraction according to biorefinery concept. Renew Energy 94:353–365, ISSN 0960-1481. https://doi.org/10.1016/j.renene.2016.03.045
Georgieva TI, Ahring BK (2007) Evaluation of continuous ethanol fermentation of dilute-acid corn stover hydrolysate using thermophilic anaerobic bacterium Thermoanaerobacter BG1L1. Appl Microbiol Biotechnol 77:61–68. https://doi.org/10.1007/s00253-007-
Martini C, Tauk-Tornisielo SM, Codato CB et al (2016) A strain of Meyerozyma guilliermondii isolated from sugarcane juice is able to grow and ferment pentoses in synthetic and bagasse hydrolysate media. World J Microbiol Biotechnol 32:80. https://doi.org/10.1007/s11274-016-2036-1
Nwachukwu R, Shahbazi A, Wang L et al (2012) Bioconversion of glycerol to ethanol by a mutant Enterobacter aerogenes. AMB Express 2:20. https://doi.org/10.1186/2191-0855-2-20
Kumdam H, Narayana Murthy S, Gummadi SN (2013) Production of ethanol and arabitol by Debaryomyces nepalensis: influence of process parameters. AMB Express 3:23. https://doi.org/10.1186/2191-0855-3-23
Unrean P, Khajeeram S (2015) Model-based optimization of Scheffersomyces stipitis and Saccharomyces cerevisiae co-culture for efficient lignocellulosic ethanol production. Bioresour Bioprocess 2:41. https://doi.org/10.1186/s40643-015-0069-1
Hoa C-Y, Chang JJ, Lin JJ et al (2011) Establishment of functional rumen bacterial consortia (FRBC) for simultaneous biohydrogen and bioethanol production from lignocellulose. Int J Hydrog Energy 36(19):12168–12176
Trana D-T, PoleI Y, Lin CW (2013) Developing co-culture system of dominant cellulolytic Bacillus sp. THLA0409 and dominant ethanolic Klebsiella oxytoca THLC0409 for enhancing ethanol production from lignocellulosic materials. J Taiwan Inst Chem Eng 44(5):762–769
Yadav KS, Naseeruddin GS et al (2011) Bioethanol fermentation of concentrated rice straw hydrolysate using co-culture of Saccharomyces cerevisiae and Pichia stipites. Bioresource Technol 102(11):6473–6478
Subsamrana K, Mahakhan P, Vichitphan K et al (2019) Potential use of vetiver grass for cellulolytic enzyme production and bioethanol production., Biocatalysis and Agricultural. Biotechnology. 17:261–268
Li Y, Mitsumasu K, Gou Z et al (2016) Xylose fermentation efficiency and inhibitor tolerance of the recombinant industrial Saccharomyces cerevisiae strain NAPX37. Appl Microbiol Biotechnol 100:1531–1542. https://doi.org/10.1007/s00253-015-7167-z
Dogaris I, Mamma D, Kekos D (2013) Biotechnological production of ethanol from renewable resources by Neurospora crassa: an alternative to conventional yeast fermentations? Appl Microbiol Biotechnol 97:1457–1473. https://doi.org/10.1007/s00253-012-4655-2
Collograi KC, da Costa AC, Ienczak JL (2019) Effect of contamination with Lactobacillus fermentum I2 on ethanol production by Spathaspora passalidarum. Appl Microbiol Biotechnol 103:5039–5050. https://doi.org/10.1007/s00253-019-09779-y
Liu Y, Zhou H, Wang L, Wang S, Fan L (2016) Improving Saccharomyces cerevisiae growth against lignocellulose-derived inhibitors as well as maximizing ethanol production by a combination proposal of γ-irradiation pretreatment with in situ detoxification. Chem Eng J 287(1):302–312
Hull CM, Loveridge EJ, Donnison IS et al (2014) Co-production of bioethanol and probiotic yeast biomass from agricultural feedstock: application of the rural biorefinery concept. AMB Express 4:64. https://doi.org/10.1186/s13568-014-0064-5
Saxena J, Tanner RS (2012) Optimization of a corn steep medium for production of ethanol from synthesis gas fermentation by Clostridium ragsdalei. World J Microbiol Biotechnol 28:1553–1561. https://doi.org/10.1007/s11274-011-0959-0
Zaldivar J, Nielsen J, Olsson L (2001) Fuel ethanol production from lignocellulose: a challenge for metabolic engineering and process integration. Appl Microbiol Biotechnol 56:17–34. https://doi.org/10.1007/s002530100624
Chang T, Yao S (2011) Thermophilic, lignocellulolytic bacteria for ethanol production: current state and perspectives. Appl Microbiol Biotechnol 92:13–27. https://doi.org/10.1007/s00253-011-3456-3
Plácido J, Capareda S (2015) Ligninolytic enzymes: a biotechnological alternative for bioethanol production. Bioresour Bioprocess 2:23. https://doi.org/10.1186/s40643-015-0049-5
Daou M, Faulds CB (2017) Glyoxal oxidases: their nature and properties. World J Microbiol Biotechnol 33:87. https://doi.org/10.1007/s11274-017-2254-1
Mamma D, Hatzinikolaou D, Kekos D et al (2009) Adsorption of major endoglucanase from Thermoascus aurantiacus on cellulosic substrates. World J Microbiol Biotechnol 25:781. https://doi.org/10.1007/s11274-008-9949-2
Zhang K, Lu X, Li Y et al (2019) New technologies provide more metabolic engineering strategies for bioethanol production in Zymomonas mobilis. Appl Microbiol Biotechnol 103:2087–2099. https://doi.org/10.1007/s00253-019-09620-6
Mello BL, Polikarpov I (2014) Family 1 carbohydrate binding-modules enhance saccharification rates. AMB Express 4:36. https://doi.org/10.1186/s13568-014-0036-9
Kameshwar S, Qin W (2018) Structural and functional properties of pectin and lignin–carbohydrate complexes de-esterases: a review. Bioresour Bioprocess 5:43. https://doi.org/10.1186/s40643-018-0230-8
Navas LE, Martínez FD, Taverna ME et al (2019) A thermostable laccase from Thermus sp. 2.9 and its potential for delignification of Eucalyptus biomass. AMB Express 9:24. https://doi.org/10.1186/s13568-019-0748-y
Boucherba N, Gagaoua M, Bouanane-Darenfed A et al (2017) Biochemical properties of a new thermo- and solvent-stable xylanase recovered using three phase partitioning from the extract of Bacillus oceanisediminis strain SJ3. Bioresour Bioprocess 4:29. https://doi.org/10.1186/s40643-017-0161-9
Zadeh AA et al (2018) Disulfide bonds elimination of endoglucanase II from Trichoderma reesei by site-directed mutagenesis to improve enzyme activity and thermal stability: an experimental and theoretical approach. Int J Biol Macromol 120(Part B,D):1572–1580
Liab X, Xia J, Zhu X et al (2019) Construction and characterization of bifunctional cellulases: caldicellulosiruptor-sourced endoglucanase, CBM, and exoglucanase for efficient degradation of lignocellulose. Biochem Eng J 151(15):107363
Kumar S, Chakrabortya PS (2018) Microwave-assisted ionic liquid-mediated rapid catalytic conversion of non-edible lignocellulosic Sunn hemp fibres to biofuels. Bioresource Technol 253:85–93
Eun J, Seung H et al (2014) Enzymatic degradation of lignocellulosic biomass by continuous process using laccase and cellulases with the aid of scaffoldin for ethanol production. Process Biochem 49(8):1266–1273
Frommhagen M, Westphal AH, Hilgers R, Koetsier MJ, Hinz SWA, Visser J, Gruppen H, Berkel WJHV, Kabel MA (2018) Quantification of the catalytic performance of C1-cellulose-specific lytic polysaccharide monooxygenases. Appl Microbiol Biotechnol 102:1281–1295
Yang F, Yang X, Li Z et al (2015) Overexpression and characterization of a glucose-tolerant β-glucosidase from T. aotearoense with high specific activity for cellobiose. Appl Microbiol Biotechnol 99:8903–8915. https://doi.org/10.1007/s00253-015-6619-9
Nakanishi A, Bae J, Kuroda K et al (2012) Construction of a novel selection system for endoglucanases exhibiting carbohydrate-binding modules optimized for biomass using yeast cell-surface engineering. AMB Express 2:56. https://doi.org/10.1186/2191-0855-2-56
Irfan M, Asghar U, Nadeem M et al (2016) Statistical optimization of saccharification of alkali pretreated wheat straw for bioethanol production. Waste Biomass Valor 7:1389–1396. https://doi.org/10.1007/s12649-016-9540-2
Hasanly A, Khajeh Talkhoncheh M, Karimi Alavijeh M (2018) Techno-economic assessment of bioethanol production from wheat straw: a case study of Iran. Clean Techn Environ Policy 20:357–377. https://doi.org/10.1007/s10098-017-1476-0
Wu M, Liu H, Guo J et al (2018) Enhanced enzymatic hydrolysis of wheat straw by two-step pretreatment combining alkalization and adsorption. Appl Microbiol Biotechnol 102:9831–9842. https://doi.org/10.1007/s00253-018-9335-4
Ire FS, Ezebuiro V, Ogugbue CJ (2016) Production of bioethanol by bacterial co-culture from agro-waste-impacted soil through simultaneous saccharification and co-fermentation of steam-exploded bagasse. Bioresour Bioprocess 3:26. https://doi.org/10.1186/s40643-016-0104-x
Arumugam A, Malolan VV, Ponnusami V (2020) Contemporary pretreatment strategies for bioethanol production from corncobs: a comprehensive review. Waste Biomass Valor. https://doi.org/10.1007/s12649-020-00983-w
Prajapati V, Trivedi U, Patel KC (2015) Bioethanol production from the raw corn starch and food waste employing simultaneous saccharification and fermentation approach. Waste Biomass Valor 6:191–200. https://doi.org/10.1007/s12649-014-9338-z
Saliu BK, Sani A (2012) Bioethanol potentials of corn cob hydrolysed using cellulases of Aspergillus niger and Penicillium decumbens. EXCLI J 11:468–479
Conesa C, Seguí L, Fito P (2018) Hydrolytic performance of Aspergillus niger and Trichoderma reesei cellulases on lignocellulosic industrial pineapple waste intended for bioethanol production. Waste Biomass Valor 9:1359–1368. https://doi.org/10.1007/s12649-017-9887-z
Wistara NJ, Pelawi R, Fatriasari W (2016) The effect of lignin content and freeness of pulp on the bioethanol productivity of Jabon wood. Waste Biomass Valor 7:1141–1146. https://doi.org/10.1007/s12649-016-9510
Ranjithkumar M, Ravikumar R, Sankar MK et al (2017) An effective conversion of cotton waste biomass to ethanol: a critical review on pretreatment processes. Waste Biomass Valor 8:57–68. https://doi.org/10.1007/s12649-016-9563-8
Sasaki C, Kiyokawa A, Asada C et al (2019) Glucose and valuable chemicals production from cotton waste using hydrothermal method. Waste Biomass Valor 10:599–607. https://doi.org/10.1007/s12649-017-0084-x
Guerrero AB, Aguado PL, Sánchez J et al (2016) GIS-based assessment of banana residual biomass potential for ethanol production and power generation: a case study. Waste Biomass Valor 7:405–415. https://doi.org/10.1007/s12649-015-9455-3
Bilal M, Asgher M, Iqbal HMN et al (2017) Enhanced bio-ethanol production from old newspapers waste through alkali and enzymatic delignification. Waste Biomass Valor 8:2271–2281. https://doi.org/10.1007/s12649-017-9871-7
Hatami-manesh M, Younesi H, Bahramifar N et al (2019) Fermentative production of ethanol from acid hydrolyzate of rice water waste using Saccharomyces cerevisiae: experimental and kinetic studies. Waste Biomass Valor. https://doi.org/10.1007/s12649-019-00697-8
Abbas A, Ansumali S (2010) Global potential of rice husk as a renewable feedstock for ethanol biofuel production. Bioenerg Res 3:328–334. https://doi.org/10.1007/s12155-010-9088-0
Tabata T, Yoshiba Y, Takashina T et al (2017) Bioethanol production from steam-exploded rice husk by recombinant Escherichia coli KO11. World J Microbiol Biotechnol 33:47. https://doi.org/10.1007/s11274-017-2221-x
Sharma S, Nandal P, Arora A (2019) Ethanol production from NaOH pretreated rice straw: a cost effective option to manage rice crop residue. Waste Biomass Valor 10:3427–3434. https://doi.org/10.1007/s12649-018-0360-4
Sindhu R, Binod P, Janu KU et al (2012) Organosolvent pretreatment and enzymatic hydrolysis of rice straw for the production of bioethanol. World J Microbiol Biotechnol 28:473–483. https://doi.org/10.1007/s11274-011-0838-8
Takano M, Hoshino K (2018) Bioethanol production from rice straw by simultaneous saccharification and fermentation with statistical optimized cellulase cocktail and fermenting fungus. Bioresour Bioprocess 5:16. https://doi.org/10.1186/s40643-018-0203-y
Phuong NTM, Hoang PH, Dien LQ et al (2017) Optimization of sodium sulfide treatment of rice straw to increase the enzymatic hydrolysis in bioethanol production. Clean Techn Environ Policy 19:1313–1322. https://doi.org/10.1007/s10098-016-1329-2
Tiwari S, Jadhav SK, Tiwari KL (2015) Bioethanol production from rice bran with optimization of parameters by Bacillus cereus strain McR-3. Int J Environ Sci Technol 12:3819–3826. https://doi.org/10.1007/s13762-014-0746-1
Dagninoab EP et al (2013) Optimization of the acid pretreatment of rice hulls to obtain fermentable sugars for bioethanol production. Ind Crops Prod 42:363–368
Bateni H, Bateni F, Karimi K (2017) Effects of oil extraction on ethanol and biogas production from Eruca sativa seed cake. Waste Biomass Valor 8:1897–1905. https://doi.org/10.1007/s12649-016-9731-x
Kasmi M (2018) Biological processes as promoting way for both treatment and valorization of dairy industry effluents. Waste Biomass Valor 9:195–209. https://doi.org/10.1007/s12649-016-9795-7
Samara C et al (2019) Optimization of bioethanol production from cheese whey using Kluyveromyces marxianus URM 7404. BIocatal Agric Biotechnol 20:101182
Sotiropoulos A, Malamis D, Loizidou M (2015) Dehydration of domestic food waste at source as an alternative approach for food waste management. Waste Biomass Valor 6:167–176. https://doi.org/10.1007/s12649-014-9343-2
Pathak PD, Mandavgane SA, Kulkarni BD (2019) Waste to wealth: a case study of papaya peel. Waste Biomass Valor 10:1755–1766. https://doi.org/10.1007/s12649-017-0181-x
Fito J, Tefera N, Van Hulle SWH (2019) Sugarcane biorefineries wastewater: bioremediation technologies for environmental sustainability. Chem Biol Technol Agric 6:6. https://doi.org/10.1186/s40538-019-0144-5
Goshadroua A, Karimia K, Taherzadeh MJ (2011) Bioethanol production from sweet sorghum bagasse by Mucor hiemalis. Ind Crop Prod 34(1):1219–1225
Dogaris I, Gkounta O, Mamma D et al (2012) Bioconversion of dilute-acid pretreated sorghum bagasse to ethanol by Neurospora crassa. Appl Microbiol Biotechnol 95:541–550. https://doi.org/10.1007/s00253-012-4113-1
Jutakanokea R et al (2012) Sugarcane leaves: pretreatment and ethanol fermentation by Saccharomyces cerevisiae. Biomass Bioenergy 39:283–289
Nihan O, Cekmecelioglu UD (2011) Cost-effective approach to ethanol production and optimization by response surface methodology. Waste Manag 31(4):636–643
Bateni H, Karimiac K, Zamaniac A, Benakashanic F (2014) Castor plant for biodiesel, biogas, and ethanol production with a biorefinery processing perspective. Appl Energy 136(31):14–22
Gao MT, Yano S, Sakanishi HK (2012) Production of ethanol from potato pulp: Investigation of the role of the enzyme from Acremonium cellulolyticus in conversion of potato pulp into ethanol. Process Biochem 47(12):2110–2115
Chena R, Benjamin D et al (2014) Effects of algal hydrolyzate as reaction medium on enzymatic hydrolysis of lignocelluloses. Biomass and Bioenergy. Volume 67:72–78
Teater C, Yue Z, MacLellan J, Liu Y, Liao W (2011) Assessing solid digestate from anaerobic digestion as feedstock for ethanol production. Bioresour Technol 102(2):1856–1862
Ekaterina A et al (2017) Dilute nitric-acid pretreatment of oat hulls for ethanol production. Biochem Eng J 126(15):118–125
Loaces I, Schein S, Noya F (2017) Ethanol production by Escherichia coli from Arundo donax biomass under SSF SHF or CBP process configurations and in situ production of a multifunctional glucanase and xylanase. Bioresour Technol 224:307–313
Frederick WJ Jr, Lien SJ et al (2008) Production of ethanol from carbohydrates from loblolly pine: a technical and economic assessment. Bioresour Technol 99(11):5051–5057
Mikulski D, Kłosowski G (2018) Efficiency of dilute sulfuric acid pretreatment of distillery stillage in the production of cellulosic ethanol. Bioresource Technology. Volume 268:424–433
Sarkar D, Prajapati S, Poddar K, Sarkar A (2019) Production of ethanol by Enterobacter sp. EtK3 during fruit waste biotransformation. Int Biodeterior Biodegradation 145:104795
Naseeruddin S, Desai S, Raoa V (2017) Ethanol production from lignocellulosic substrate Prosopis juliflora. Renew Energy. 103:701–707
Carlos J et al (2017) Combined acid/alkaline-peroxide pretreatment of olive tree biomass for bioethanol production. Bioresour Technol 239:326–335
Sreena CP, Sebastian D (2019) Jackfruit outer rind: a sustainable feedstock for fermentable sugar production using recombinant endoglucanase from Bacillus subtilis MU S1. Environ Technol Innov 16:100448
Susana M et al (2018) Evaluation of Jerusalem artichoke as a sustainable energy crop to bioethanol: energy and CO2eq emissions modeling for an industrial scenario. Energy. 150(1):468–481
Anh Q et al (2017) Development of an integrated process to produce d-mannose and bioethanol from coffee residue waste. Bioresour Technol 244(Part 1):1039–1048
Gurram R, Al-Shannag M, Knapp S et al (2016) Technical possibilities of bioethanol production from coffee pulp: a renewable feedstock. Clean Techn Environ Policy 18:269–278. https://doi.org/10.1007/s10098-015-1015-9
Liava V, Karkanis A, Danalatos N, Tsiropoulos N (2021) Cultivation practices, adaptability and phytochemical composition of Jerusalem artichoke (Helianthus tuberosus L.): a weed with economic value. Agronomy 11:914. https://doi.org/10.3390/agronomy11050914
Liu Y-K et al (2017) Production of bioethanol from Napier grass via simultaneous saccharification and co-fermentation in a modified bioreactor. J Biosci Bioeng 124(2):184–188
Rajeswari G, Jacob S (2020) Deciphering the aloe vera leaf rind as potent feedstock for bioethanol through enzymatic delignification and its enhanced saccharification. Ind Crop Prod 143:111876
Byadgi SA, Kalburgi PB (2016) Production of bioethanol from wastenews paper. Procedia Environ Sci 35(2016):555–562
Khounania Z, Nazemi F, Shafieic M et al (2019) Techno-economic aspects of a safflower-based biorefinery plant co-producing bioethanol and biodiesel. Energy Convers Manag 201(1):112184
Sathendra ER et al (2019) Bioethanol production from palm wood using Trichoderma reesei and Kluveromyces marxianus. Bioresour Technol 271:345–352
Indulekha J, Gokul Siddarth MS, Kalaichelvi P, Arunagiri A (2017) Characterization of citrus peels for bioethanol production. In: Mohan BR, Srinikethan G, Meikap B (eds) Materials, energy and environment engineering. Springer, Singapore. https://doi.org/10.1007/978-981-10-2675-1_1
Pesce GR, Fernandes MC, Mauromicale G (2020) Globe artichoke crop residues and their potential for bioethanol production by dilute acid hydrolysis. Biomass Bioenergy 134:105471
Taghizadeh-Alisaraei A, Motevali A, Ghobadian B (2019) Ethanol production from date wastes: adapted technologies, challenges, and global potential. Renew Energy 143:1094–1110. https://doi.org/10.1016/j.renene.2019.05.048
Amina M et al (2019) The development of a biorefining strategy for the production of biofuel from sorghum milling waste. Biochem Eng J 150:107288
Brown C., Griggs T., Holaskova I., Skousen J, Switchgrass Biofuel Production on Reclaimed Surface Mines
Purdy SJ, Cunniff J, Maddison AL, Jones EL, Barraclough T, Castle M, Davey CL, Jones CM, Shield I, Gallagher J, Donnison I, Clifton-Brown J (2015) Seasonal carbohydrate dynamics and climatic regulation of senescence in the perennial grass. Miscanthus, Bioenerg Res 8:28–41
Rehman O, Shahid A, Liu C et al (2019) Optimization of low-temperature energy-efficient pretreatment for enhanced saccharification and fermentation of Conocarpus erectus leaves to produce ethanol using Saccharomyces cerevisiae. Biomass Conv Bioref. https://doi.org/10.1007/s13399-019-00529-8
Casabar JT, Unpaprom Y, Ramaraj R (2019) Fermentation of pineapple fruit peel wastes for bioethanol production. Biomass Conv Bioref 9:761–765. https://doi.org/10.1007/s13399-019-00436-y
Verma R, Suthar S (2015) Utility of duckweeds as source of biomass energy: a review. Bioenerg Res 8:1589–1597. https://doi.org/10.1007/s12155-015-9639-5
Park HR, Jung KA, Lim S et al (2014) Quantitative sustainability assessment of seaweed biomass as bioethanol feedstock. Bioenerg Res 7:974–985. https://doi.org/10.1007/s12155-014-9430-z
Hessami MJ, Phang S, Salleh A et al (2018) Evaluation of tropical seaweeds as feedstock for bioethanol production. Int J Environ Sci Technol 15:977–992. https://doi.org/10.1007/s13762-017-1455-3
Zacharof MP (2017) Grape winery waste as feedstock for bioconversions: applying the biorefinery concept. Waste Biomass Valor 8:1011–1025
Xin F, Zhang H, Wong W (2013) Bioethanol production from horticultural waste using crude fungal enzyme mixtures produced by solid state fermentation. Bioenerg Res 6:1030–1037
Saxena J, Tanner RS (2011) Effect of trace metals on ethanol production from synthesis gas by the ethanologenic acetogen, Clostridium ragsdalei. J Ind Microbiol Biotechnol 38(4):513–521. https://doi.org/10.1007/s10295-010-0794-6
Laopaiboon L, Laopaiboon P (2012) Ethanol production from sweet sorghum juice in repeated-batch fermentation by Saccharomyces cerevisiae immobilized on corncob. World J Microbiol Biotechnol 28:559–566. https://doi.org/10.1007/s11274-011-0848-6
Shimokawa T, Shibuya H, Ikeda T et al (2013) Ethanol production from sugi pulp under simultaneous saccharification and fermentation using a cocktail enzyme of T. reesei and A. tubingensis produced by solid-state fermentation. J Wood Sci 59:171–178. https://doi.org/10.1007/s10086-012-1307-1
Yan Q, Liu X, Wang Y et al (2018) Cow manure as a lignocellulosic substrate for fungal cellulase expression and bioethanol production. AMB Express 8:190. https://doi.org/10.1186/s13568-018-0720-2
Plácido J, Capareda S (2014) Analysis of alkali ultrasonication pretreatment in bioethanol production from cotton gin trash using FT-IR spectroscopy and principal component analysis. Bioresour Bioprocess 1:23. https://doi.org/10.1186/s40643-014-0023-7
El-Mekkawi SA, Abdo SM, Samhan FA et al (2019) Optimization of some fermentation conditions for bioethanol production from microalgae using response surface method. Bull Natl Res Cent 43:164. https://doi.org/10.1186/s42269-019-0205-8
Offei F, Mensah M, Kemausuor F (2019) Cellulase and acid-catalysed hydrolysis of Ulva fasciata, Hydropuntia dentata and Sargassum vulgare for bioethanol production. SN Appl Sci 1:1469. https://doi.org/10.1007/s42452-019-1501-5
Fennouche I, Khellaf N, Djelal H et al (2019) An effective acid pretreatment of agricultural biomass residues for the production of second-generation bioethanol. SN Appl Sci 1:1460. https://doi.org/10.1007/s42452-019-1517-x
Jha P, Singh S, Raghuram M et al (2019) Valorisation of orange peel: supplement in fermentation media for ethanol production and source of limonene. Environ Sustain 2:33–41. https://doi.org/10.1007/s42398-019-00048-2
Demiray E, Karatay SE, Dönmez G (2019) Improvement of bioethanol production from pomegranate peels via acidic pretreatment and enzymatic hydrolysis. Environ Sci Pollut Res 26:29366–29378. https://doi.org/10.1007/s11356-019-06020-1
Bashiri R, Farhadian M, Asadollahi MA et al (2016) Anaerobic digested sludge: a new supplementary nutrient source for ethanol production. Int J Environ Sci Technol 13:763–772. https://doi.org/10.1007/s13762-015-0925-8
Vong WC, Lim XY, Liu S (2017) Biotransformation with cellulase, hemicellulase and Yarrowia lipolytica boosts health benefits of okara. Appl Microbiol Biotechnol 101:7129–7140. https://doi.org/10.1007/s00253-017-8431-1
Chandel AK, Singh OV, Narasu L, Rao V (2011) Bioconversion of Saccharum spontaneum (wild sugarcane) hemicellulosic hydrolysate into ethanol by mono and co-cultures of Pichia stipitis NCIM3498 and thermotolerant Saccharomyces cerevisiae-VS3. New Biotechnol 28(6):593–599
Singh LK, Majumde CB, Ghosh S (2014) Development of sequential-co-culture system (Pichia stipitis and Zymomonas mobilis) for bioethanol production from Kans grass biomass. Biochem Eng J. 82:150–157
Bahry H et al (2017) Valorization of carob waste: definition of a second-generation bioethanol production process. Bioresour Technol 235:25–34
Fernandes MC, Ferro MD, Paulino AFC, Chaves HT, Evtuguin DV, Xavier AMRB (2018) Comparative study on hydrolysis and bioethanol production from cardoon and rockrose pretreated by dilute acid hydrolysis. Ind Crop Prod 111:633–641, ISSN 0926-6690. https://doi.org/10.1016/j.indcrop.2017.11.037
Boluda-Aguilar M, López-Gómez A (2013) Production of bioethanol by fermentation of lemon (Citrus limon L.) peel wastes pretreated with steam explosion. Ind Crops and Prod. 41:188–197
Baskar G et al (2016) Sesbania aculeate biomass hydrolysis using magnetic nanobiocomposite of cellulase for bioethanol production. Renew Energy 98:23–28
Mathias DJ, Kumar S, Rangarajan V (2019) An investigation on citrus peel as the lignocellulosic feedstock for optimal reducing sugar synthesis with an additional scope for the production of hydrolytic enzymes from the aqueous extract waste. Biocatal Agric Biotechnol 20:101259
Uyan M, Alptekin FM, Bastabak B et al (2019) Combined biofuel production from cotton stalk and seed with a biorefinery approach. Biomass Conv Bioref. https://doi.org/10.1007/s13399-019-00427-z
Pothiraj C, Arumugam R, Gobinath M (2014) Sustaining ethanol production from lime pretreated water hyacinth biomass using mono and co-cultures of isolated fungal strains with Pichia stipitis. Bioresour Bioprocess 1:27. https://doi.org/10.1186/s40643-014-0027-3
Xu C, Xia T, Wang J et al (2020) Selectively desirable rapeseed and corn stalks distinctive for low-cost bioethanol production and high-active biosorbents. Waste Biomass Valor. https://doi.org/10.1007/s12649-020-01026-0
Sayar NA, Pinar O, Kazan D et al (2019) Bioethanol production from Turkish hazelnut husk process design and economic evaluation. Waste Biomass Valor 10:909–923. https://doi.org/10.1007/s12649-017-0103-y
John I, Pola J, Appusamy A (2019) Optimization of ultrasonic assisted saccharification of sweet lime peel for bioethanol production using Box–Behnken method. Waste Biomass Valor 10:441–453. https://doi.org/10.1007/s12649-017-0072-1
Pena R, Lú-Chau TA, Lema JM (2012) Use of white-rot fungi for valorization of stillage from bioethanol production. Waste Biomass Valor 3:295–303. https://doi.org/10.1007/s12649-012-9130-x
Dania A et al (2014) Towards integrated biorefinery from dried distillers grains: Selective extraction of pentoses using dilute acid hydrolysis. Biomass Bioenergy 71:178–186
Rajeswari G, Arutselvy B, Jacob S (2019) Delignification of aloe vera rind by mild acid associated microwave pretreatment to persuade enhanced enzymatic saccharification. Waste Biomass Valor. https://doi.org/10.1007/s12649-019-00830-7
Cardias BB, Trevisol TC, Bertuol GG et al (2020) Hydrolyzed spirulina biomass and molasses as substrate in alcoholic fermentation with application of magnetic fields. Waste Biomass Valor. https://doi.org/10.1007/s12649-020-00966-x
Ezeilo UR, Wahab RA, Tin LC et al (2019) Fungal-assisted valorization of raw oil palm leaves for production of cellulase and xylanase in solid state fermentation media. Waste Biomass Valor. https://doi.org/10.1007/s12649-019-00653-6
Barcelos CA, Maeda RN, Betancur GJV et al (2013) The essentialness of delignification on enzymatic hydrolysis of sugar cane bagasse cellulignin for second generation ethanol production. Waste Biomass Valor 4:341–346. https://doi.org/10.1007/s12649-012-9137-3
Herrero ML, Vallejo MD, Sardella MF et al (2015) Acid pretreatment of two phase olive mill waste to improve bioavailable sugars: conditions optimization using response surface methodology. Waste Biomass Valor 6:37–44. https://doi.org/10.1007/s12649-014-9336-1
Brito PL, de Azevedo Ferreira CM, Silva AFF et al (2018) Hydrolysis, detoxification and alcoholic fermentation of hemicellulose fraction from palm press fiber. Waste Biomass Valor 9:957–968. https://doi.org/10.1007/s12649-017-9882-4
Linzmeyer P, Ramlow H, Souza O et al (2019) Influence of neutralizing agents on the recovery of ethanol from banana pseudostem broth by pervaporation. Waste Biomass Valor. https://doi.org/10.1007/s12649-019-00773-z
Siripattanakul-Ratpukdi S (2012) Ethanol production potential from fermented rice noodle wastewater treatment using entrapped yeast cell sequencing batch reactor. Appl Water Sci 2:47–53. https://doi.org/10.1007/s13201-011-0024-z
Lebaka VR, Ryu H, Wee Y (2014) Effect of fruit pulp supplementation on rapid and enhanced ethanol production in very high gravity (VHG) fermentation. Bioresour Bioprocess 1:22. https://doi.org/10.1186/s40643-014-0022-8
Bowman MJ et al (2012) Liquid chromatography–mass spectrometry investigation of enzyme-resistant xylooligosaccharide structures of switchgrass associated with ammonia pretreatment, enzymatic saccharification, and fermentation. Bioresour Technol 110:437–447
Susana R et al (2013) Advances in ethanol production from hardwood spent sulphite liquors. Process Biochem 48(2):272–282
Miguel A et al (2013) Wastewater from the soft drinks industry as a source for bioethanol production. Bioresource Technol 136:140–147
Kasmi M, Chatti A, Hamdi M et al (2016) Eco-friendly process for soft drink industries wastewater reuse as growth medium for Saccharomyces cerevisiae production. Clean Techn Environ Policy 18:2265–2278. https://doi.org/10.1007/s10098-016-1144-9
Avelino F et al (2015) Bioethanol production from coconuts and cactus pretreated by autohydrolysis. Ind Crop Prod 77(23):1–12
Hou X, HøegHansen J, Bjerre AB (2015) Integrated bioethanol and protein production from brown seaweed Laminaria digitata. Bioresource Technol 197:310–317
Gomez R et al (2018) Bioethanol and biobutanol production from sugarcorn juice. Biomass Bioenergy 108:455–463
Meinita MDN, Marhaeni B, Winanto T, Setyaningsih D, Hong YK (2015) Catalytic efficiency of sulfuric and hydrochloric acids for the hydrolysis of Gelidium latifolium (Gelidiales, Rhodophyta) in bioethanol production. J Ind Eng Chem 27:108–114
Gomes D, Domingues L, Gama M (2016) Valorizing recycled paper sludge by a bioethanol production process with cellulase recycling. Bioresour Technol 216:637–644
Borinesa MG, de Leon RL, Cuelloc JL (2013) Bioethanol production from the macroalgae Sargassum spp. Bioresour Technol 138:22–29
Betiku E, Taiwo AE (2015) Modeling and optimization of bioethanol production from breadfruit starch hydrolyzate vis-à-vis response surface methodology and artificial neural network. Renew Energy 74:87–94
Das SP, Gupta A, Das D, Goyal A (2016) Enhanced bioethanol production from water hyacinth (Eichhornia crassipes) by statistical optimization of fermentation process parameters using Taguchi orthogonal array design. Int Biodeterior Biodegradation 109:174–184
Muktham RK, Andrew S, Suresh B, Bhargava K, Bankupalli S (2016) Bioethanol production from non-edible de-oiled Pongamia pinnata seed residue-optimization of acid hydrolysis followed by fermentation. Ind Crop Prod 94(30):490–497
Ulises I et al (2017) Chemical, structural, and ultrastructural analysis of waste from the carrageenan and sugar-bioethanol processes for future bioenergy generation. Biomass Bioenergy 107:233–243
Avelino F et al (2016) Bioethanol production by Saccharomyces cerevisiae, Pichia stipitis and Zymomonas mobilis from delignified coconut fibre mature and lignin extraction according to biorefinery concept. Renew Energy 94:353–365
Xumeng GV et al (2012) Eastern gamagrass as an alternative cellulosic feedstock for bioethanol production. Process Biochem 47(2):335–339
Nikolića S et al (2017) Production of bioethanol from pre-treated cotton fabrics and waste cotton materials. Carbohydr Polym. 164:136–144
Mohan SS et al (2016) Bioethanol production through separate hydrolysis and fermentation of Parthenium hysterophorus biomass. Renew Energy 86:1317–1323
Sahoo D, Ummalyma SB, Okram AK, Sukumaran RK, George E, Pandey A (2017) Potential of Brachiaria mutica (Para grass) for bioethanol production from Loktak Lake. Bioresour Technol 242:133–138. https://doi.org/10.1016/j.biortech.2017.03.047
Singh N et al (2018) Bioethanol production potential of a novel thermophilic isolate Thermoanaerobacter sp. DBT-IOC-X2 isolated from Chumathang hot spring. Biomass Bioenergy 116:122–130
Thi L et al (2019) Aqueous acidified ionic liquid pretreatment for bioethanol production and concentration of produced ethanol by pervaporation. J Ind Eng Chem 69(25):57–65
Cardona CA, Quintero JA, Paz IC (2010) Production of bioethanol from sugarcane bagasse: status and perspectives. Bioresour Technol 101(13):4754–4766. https://doi.org/10.1016/j.biortech.2009.10.097
Priharto N et al (2020) Fast pyrolysis with fractional condensation of lignin-rich digested stillage from second-generation bioethanol production. J Anal Appl Pyrolysis 145:104756
Nandal P, Sharma S, Arora A (2020) Bioprospecting non-conventional yeasts for ethanol production from rice straw hydrolysate and their inhibitor tolerance. Renew Energy 147(Part 1):1694–1703
Dahnuma D et al (2015) Comparison of SHF and SSF processes using enzyme and dry yeast for optimization of bioethanol production from empty fruit bunch. Energy Procedia 68:107–116
Soplah S et al (2015) Fresh oil palm frond juice as a renewable, non-food, non-cellulosic and complete medium for direct bioethanol production. Ind Crop Prod 63:357–361
Alassali A, Cybulska I, Galvan AR et al (2017) Wet fractionation of the succulent halophyte Salicornia sinus-persica, with the aim of low input (water saving) biorefining into bioethanol. Appl Microbiol Biotechnol 101:1769–1779. https://doi.org/10.1007/s00253-016-8049-8
Guigou MD, Cebreiros F, Cabrera MN et al (2017) Bioethanol production from Eucalyptus grandis hemicellulose recovered before kraft pulping using an integrated biorefinery concept. Biomass Conv Bioref 7:191–197. https://doi.org/10.1007/s13399-016-0218-6
Amato G, Giambalvo D, Frenda AS et al (2016) Sulla (Hedysarum coronarium L.) as potential feedstock for biofuel and protein. Bioenerg Res 9:711–719. https://doi.org/10.1007/s12155-016-9715-5
Su R, Ma Y, Qi W et al (2013) Ethanol production from high-solid SSCF of alkaline-pretreated corncob using recombinant Zymomonas mobilis CP4. Bioenerg Res 6:292–299. https://doi.org/10.1007/s12155-012-9256-5
Khambhaty Y, Upadhyay D, Kriplani Y et al (2013) Bioethanol from macroalgal biomass: utilization of marine yeast for production of the same. Bioenerg Res 6:188–195. https://doi.org/10.1007/s12155-012-9249-4
Kostas ET., White DA., Cook DJ. Bioethanol production from UK seaweeds: investigating variable pre-treatment and enzyme hydrolysis parameters. BioEnergy Res
Inokuma K, Hasunuma T, Kondo A (2016) Ethanol production from N-acetyl-d-glucosamine by Scheffersomyces stipitis strains. AMB Express 6:83. https://doi.org/10.1186/s13568-016-0267-z
Kaida R, Kaku T, Baba K et al (2009) Enhancement of saccharification by overexpression of poplar cellulase in sengon. J Wood Sci 55:435–440. https://doi.org/10.1007/s10086-009-1048-y
Dadi D, Beyene A, Simoens K et al (2018) Valorization of coffee byproducts for bioethanol production using lignocellulosic yeast fermentation and pervaporation. Int J Environ Sci Technol 15:821–832. https://doi.org/10.1007/s13762-017-1440-x/
Zhang M, Shukla P, Ayyachamy M et al (2010) Improved bioethanol production through simultaneous saccharification and fermentation of lignocellulosic agricultural wastes by Kluyveromyces marxianus 6556. World J Microbiol Biotechnol 26:1041–1046. https://doi.org/10.1007/s11274-009-0267-0
Quevedo-Hidalgo B, Monsalve-Marín F, Narváez-Rincón PC et al (2013) Ethanol production by Saccharomyces cerevisiae using lignocellulosic hydrolysate from Chrysanthemum waste degradation. World J Microbiol Biotechnol 29:459–466. https://doi.org/10.1007/s11274-012-1199-7
John I, Pola J, Thanabalan M et al (2019) Bioethanol production from musambi peel by acid catalyzed steam pretreatment and enzymatic saccharification: optimization of delignification using Taguchi Design. Waste Biomass Valor. https://doi.org/10.1007/s12649-018-00565-x
Messaoudi Y, Smichi N, Bouachir F et al (2019) Fractionation and biotransformation of lignocelluloses-based wastes for bioethanol, xylose and vanillin production. Waste Biomass Valor 10:357–367. https://doi.org/10.1007/s12649-017-0062-3
Sowatad A, Todhanakasem T (2020) Bioethanol production by repeated batch using immobilized yeast cells on sugarcane bagasse. Waste Biomass Valor 11:2009–2016. https://doi.org/10.1007/s12649-018-0534-0
Farah Amani AH, Toh SM, Tan JS et al (2018) The efficiency of using oil palm frond hydrolysate from enzymatic hydrolysis in bioethanol production. Waste Biomass Valor 9:539–548. https://doi.org/10.1007/s12649-017-0005-z
Manivannan A, Narendhirakannan RT (2015) Bioethanol production from aquatic weed water hyacinth (Eichhornia crassipes) by yeast fermentation. Waste Biomass Valor 6:209–216. https://doi.org/10.1007/s12649-015-9347-6
Loizidou M, Alamanou DG, Sotiropoulos A et al (2017) Pilot scale system of two horizontal rotating bioreactors for bioethanol production from household food waste at high solid concentrations. Waste Biomass Valor 8:1709–1719. https://doi.org/10.1007/s12649-017-9900-6
Chilari D, Dimos K, Georgoula G et al (2017) Bioethanol production from alkali-treated cotton stalks at high solids loading applying non-isothermal simultaneous saccharification and fermentation. Waste Biomass Valor 8:1919–1929. https://doi.org/10.1007/s12649-016-9818-4
Bona D, Vecchiet A, Pin M et al (2018) The biorefinery concept applied to bioethanol and biomethane production from manure. Waste Biomass Valor 9:2133–2143. https://doi.org/10.1007/s12649-017-9981-2
Alamanou DG, Malamis D, Mamma D et al (2015) (2015) Bioethanol from dried household food waste applying non-isothermal simultaneous saccharification and fermentation at high substrate concentration. Waste Biomass Valor 6:353–361. https://doi.org/10.1007/s12649-015-9355-6
Jutakridsada P, Saengprachatanarug K, Kasemsiri P et al (2019) Bioconversion of Saccharum officinarum leaves for ethanol production using separate hydrolysis and fermentation processes. Waste Biomass Valor 10:817–825. https://doi.org/10.1007/s12649-017-0104-x
Bhatia L, Johri S (2016) FTIR analysis and optimization of simultaneous saccharification and fermentation parameters for sustainable production of ethanol from peels of Ananas cosmosus by Mucor indicus MTCC 4349. Waste Biomass Valor 7:427–438. https://doi.org/10.1007/s12649-015-9462-4
Selvakumar P, Kavitha S, Sivashanmugam P (2019) Optimization of process parameters for efficient bioconversion of thermo-chemo pretreated Manihot esculenta Crantz YTP1 stem to ethanol. Waste Biomass Valor 10:2177–2191. https://doi.org/10.1007/s12649-018-0244-7
Yuan Z, Wei W, Li G et al (2019) High titer ethanol production from combined alkaline/alkaline hydrogen peroxide pretreated bamboo at high solid loading. Waste Biomass Valor. https://doi.org/10.1007/s12649-019-00638-5
Nutongkaew T, Prasertsan P, Leamdum C et al (2020) Bioconversion of oil palm trunk residues hydrolyzed by enzymes from newly isolated fungi and use for ethanol and acetic acid production under two-stage and simultaneous fermentation. Waste Biomass Valor 11:1333–1347. https://doi.org/10.1007/s12649-019-00678-x
Vaid S, Bajaj BK (2017) Production of ionic liquid tolerant cellulase from Bacillus subtilis G2 using agroindustrial residues with application potential for saccharification of biomass under one pot consolidated bioprocess. Waste Biomass Valor 8:949–964. https://doi.org/10.1007/s12649-016-9626-x
Njoku SI, Iversen JA, Uellendahl H et al (2013) Production of ethanol from hemicellulose fraction of cocksfoot grass using Pichia stipitis. Sustain Chem Process 1:13. https://doi.org/10.1186/2043-7129-1-13
Biswas R, Uellendahl H, Ahring BK (2013) Conversion of C6 and C5 sugars in undetoxified wet exploded bagasse hydrolysates using Scheffersomyces (Pichia) stipitis CBS6054. AMB Express 3:42. https://doi.org/10.1186/2191-0855-3-42
Ding C, Wang X, Li M (2019) Evaluation of six white-rot fungal pretreatments on corn stover for the production of cellulolytic and ligninolytic enzymes, reducing sugars, and ethanol. Appl Microbiol Biotechnol 103:5641–5652. https://doi.org/10.1007/s00253-019-09884-y
Lezinou V, Christokopoulos P, Li LW et al (1995) Study of a single and mixed culture for the direct bio-conversion of sorghum carbohydrates to ethanol. Appl Microbiol Biotechnol 43:412–415. https://doi.org/10.1007/BF00218442
Nikolić S, Mojović L, Rakin M et al (2011) Utilization of microwave and ultrasound pretreatments in the production of bioethanol from corn. Clean Techn Environ Policy 13:587–594. https://doi.org/10.1007/s10098-011-0366-0
Awedem Wobiwo F, Chaturvedi T, Boda M et al (2019) Bioethanol potential of raw and hydrothermally pretreated banana bulbs biomass in simultaneous saccharification and fermentation process with Saccharomyces cerevisiae. Biomass Conv Bioref 9:553–563. https://doi.org/10.1007/s13399-018-00367-0
Kuloyo OO, du Preez JC, García-Aparicio MP et al (2014) Opuntia ficus-indica cladodes as feedstock for ethanol production by Kluyveromyces marxianus and Saccharomyces cerevisiae. World J Microbiol Biotechnol 30:3173–3183. https://doi.org/10.1007/s11274-014-1745-6
Dasgupta D, Suman SK, Pandey D et al (2013) Design and optimization of ethanol production from bagasse pith hydrolysate by a thermotolerant yeast Kluyveromyces sp. IIPE453 using response surface methodology. SpringerPlus 2:159. https://doi.org/10.1186/2193-1801-2-159
Chen M, Kaur P, Dien B et al (2013) Use of tropical maize for bioethanol production. World J Microbiol Biotechnol 29:1509–1515. https://doi.org/10.1007/s11274-013-1317-1
Islam ZU, Klykov SP, Yu Z et al (2018) Fermentation of detoxified acid-hydrolyzed pyrolytic anhydrosugars into bioethanol with Saccharomyces cerevisiae 2.399. Appl Biochem Microbiol 54:58–70. https://doi.org/10.1134/S0003683818010143
Stobienia M, Kalschne DL, Peron-Schlosser B et al (2020) Evaluation of ultrasound waves on S. cerevisiae stimulation in the bioethanol production from rice bran. Bioenerg Res. https://doi.org/10.1007/s12155-019-10088-5
Peinado S, Mateo S, Sánchez S et al (2019) Effectiveness of sodium borohydride treatment on acid hydrolyzates from olive-tree pruning biomass for bioethanol production. Bioenerg Res 12:302–311. https://doi.org/10.1007/s12155-019-09979-4
Sharma D, Sud A, Bansal S et al (2018) Endocellulase production by Cotylidia pannosa and its application in saccharification of wheat bran to bioethanol. Bioenerg Res 11:219–227. https://doi.org/10.1007/s12155-017-9890-z
Murari CS, da Silva DCMN, Schuina GL et al (2019) Bioethanol production from dairy industrial coproducts. Bioenerg Res 12:112–122. https://doi.org/10.1007/s12155-018-9949-5
Lin Y, Zhao Y, Ruan X et al (2019) The potential of constructed wetland plants for bioethanol production. Bioenerg Res. https://doi.org/10.1007/s12155-019-10065-y
Kaur A, Kuhad RC (2019) Valorization of rice straw for ethanol production and lignin recovery using combined acid-alkali pre-treatment. Bioenerg Res 12:570–582
Corbin KR, Betts NS, van Holst N et al (2016) Low-input fermentations of agave tequilana leaf juice generate high returns on ethanol yields. Bioenerg Res 9:1142–1154. https://doi.org/10.1007/s12155-016-9755-x
Hijosa-Valsero M, Garita-Cambronero J, Paniagua-García AI et al (2019) Tomato waste from processing industries as a feedstock for biofuel production. Bioenerg Res 12:1000–1011. https://doi.org/10.1007/s12155-019-10016-7
Yasuda M, Takenouchi Y, Nitta Y et al (2015) Italian ryegrass (Lolium multiflorum Lam) as a high-potential bio-ethanol resource. Bioenerg Res 8:1303–1309. https://doi.org/10.1007/s12155-015-9582-5
Tahir B, Mezori HA (2020) Bioethanol production from Quercus aegilops using Pichia stipitis and Kluyveromyces marxianus. Biomass Conv Bioref. https://doi.org/10.1007/s13399-020-00704-2
Slathia PS, Raina N, Kiran A et al (2020) Dilute acid pretreatment of pine needles of Pinus roxburghii by response surface methodology for bioethanol production by separate hydrolysis and fermentation. Biomass Conv Bioref 10:95–106. https://doi.org/10.1007/s13399-019-00433-1
Sarbishei S, Goshadrou A, Hatamipour MS (2020) Mild sodium hydroxide pretreatment of tobacco product waste to enable efficient bioethanol production by separate hydrolysis and fermentation. Biomass Conv Bioref. https://doi.org/10.1007/s13399-020-00644-x
Rana QUA, Khan MAN, Irfan M et al (2019) Starved Spirodela polyrhiza and Saccharomyces cerevisiae: a potent combination for sustainable bioethanol production. Biomass Conv Bioref. https://doi.org/10.1007/s13399-019-00540-z
Niju S, Nishanthini T, Balajii M (2020) Alkaline hydrogen peroxide-pretreated sugarcane tops for bioethanol production—a process optimization study. Biomass Conv Bioref 10:149–165. https://doi.org/10.1007/s13399-019-00524-z
Sangkharak K, Chookhun K, Numreung J et al (2019) Utilization of coconut meal, a waste product of milk processing, as a novel substrate for biodiesel and bioethanol production. Biomass Conv Bioref. https://doi.org/10.1007/s13399-019-00456-8
Guigou M, Cabrera MN, Vique M et al (2019) Combined pretreatments of eucalyptus sawdust for ethanol production within a biorefinery approach. Biomass Conv Bioref 9:293–304. https://doi.org/10.1007/s13399-018-0353-3
Germec M, Turhan I (2018) Ethanol production from acid-pretreated and detoxified rice straw as sole renewable resource. Biomass Conv Bioref 8:607–619. https://doi.org/10.1007/s13399-018-0310-1
Palacios AS, Ilyina A, Ramos-González R et al (2019) Ethanol production from banana peels at high pretreated substrate loading: comparison of two operational strategies. Biomass Conv Bioref. https://doi.org/10.1007/s13399-019-00562-7
Guo J, Wang Y, Cheng J et al (2020) Enhancing enzymatic hydrolysis and fermentation efficiency of rice straw by pretreatment of sodium perborate. Biomass Conv Bioref. https://doi.org/10.1007/s13399-020-00668-3
Manzanares P, Ballesteros I, Negro MJ et al (2012) Biological conversion of forage sorghum biomass to ethanol by steam explosion pretreatment and simultaneous hydrolysis and fermentation at high solid content. Biomass Conv Bioref 2:123–132. https://doi.org/10.1007/s13399-012-0040-8
Granados-Arvizu J, Melo-Sabogal D, Amaro-Reyes A et al (2019) Corn pericarp pretreated with dilute acid: bioconversion of sugars in the liquid fraction to ethanol and studies on enzymatic hydrolysis of the solid fraction. Biomass Conv Bioref. https://doi.org/10.1007/s13399-019-00534-x
Camargo D, Sene L (2014) Production of ethanol from the hemicellulosic fraction of sunflower meal biomass. Biomass Conv Bioref 4:87–93. https://doi.org/10.1007/s13399-013-0096-0
Choi G, Kang H, Moon S (2009) Repeated-batch fermentation using flocculent hybrid, Saccharomyces cerevisiae CHFY0321 for efficient production of bioethanol. Appl Microbiol Biotechnol 84:261–269. https://doi.org/10.1007/s00253-009-1946-3/
Miranda JR, Passarinho PC, Gouveia L (2012) Bioethanol production from Scenedesmus obliquus sugars: the influence of photobioreactors and culture conditions on biomass production. Appl Microbiol Biotechnol 96:555–564. https://doi.org/10.1007/s00253-012-4338-z
Hu N, Yuan B, Sun J et al (2012) Thermotolerant Kluyveromyces marxianus and Saccharomyces cerevisiae strains representing potentials for bioethanol production from Jerusalem artichoke by consolidated bioprocessing. Appl Microbiol Biotechnol 95:1359–1368. https://doi.org/10.1007/s00253-012-4240-8
Takagi T, Sasaki Y, Motone K et al (2017) Construction of bioengineered yeast platform for direct bioethanol production from alginate and mannitol. Appl Microbiol Biotechnol 101:6627–6636. https://doi.org/10.1007/s00253-017-8418-y
Du C, Li Y, Zhao X et al (2019) The production of ethanol from lignocellulosic biomass by Kluyveromyces marxianus CICC 1727-5 and Spathaspora passalidarum ATCC MYA-4345. Appl Microbiol Biotechnol 103:2845–2855. https://doi.org/10.1007/s00253-019-09625-1
Sasaki K, Tsuge Y, Kawaguchi H et al (2017) Sucrose purification and repeated ethanol production from sugars remaining in sweet sorghum juice subjected to a membrane separation process. Appl Microbiol Biotechnol 101:6007–6014. https://doi.org/10.1007/s00253-017-8316-3
Naseeha A, Chohan GS, Aruwajoye Y, Sewsynker-Sukai EB, Kana G (2020) Valorisation of potato peel wastes for bioethanol production using simultaneous saccharification and fermentation: process optimization and kinetic assessment. Renew Energy 146:1031–1040
Ben I et al (2020) Evaluation of the non-conventional yeast strain Wickerhamomyces anomalus (Pichia anomala) X19 for enhanced bioethanol production using date palm sap as renewable feedstock. Renew Energy 154:71–81
Marthinus W et al (2019) Application of industrial amylolytic yeast strains for the production of bioethanol from broken rice. Bioresour Technol 294:122222
Cripwell R (2015) Utilisation of wheat bran as a substrate for bioethanol production using recombinant cellulases and amylolytic yeast. Appl Energy 160(15):610–617
Mohapatraa S, Mishra C, Behera S, Thatoic H (2017) Application of pretreatment, fermentation and molecular techniques for enhancing bioethanol production from grass biomass – a review. Renew Sust Energ Rev 78:1007–1032
Tan L et al (2020) Production of bioethanol from unwashed-pretreated rapeseed straw at high solid loading. Bioresour Technol 303:122949
Kumar P, Kumar V, Kumar S, Singh J, Kumar P (2020) Bioethanol production from sesame (Sesamum indicum L.) plant residue by combined physical, microbial and chemical pretreatments. Bioresour Technol 297:122484. https://doi.org/10.1016/j.biortech.2019.122484
Luiza A et al (2020) Simultaneous saccharification and fermentation of Spirulina sp. and corn starch for the production of bioethanol and obtaining biopeptides with high antioxidant activity/. Bioresour Technol 301:122698
Prasad S, Kumar S, Yadav KK, Choudhry J, Kamyab H, Bach QV, Sheetal KR, Kannojiya S, Gupta N (2020) Screening and evaluation of cellulytic fungal strains for saccharification and bioethanol production from rice residue. Energy 190:116422
Azevedo A et al (2017) Life cycle assessment of bioethanol production from cattle manure. J Clean Prod 162(20):1021–1030
Kumar N, Chatterjee S, Hemalatha M, Althuri A, Min B, Kim SH, Mohan SV (2020) Deoiled algal biomass derived renewable sugars for bioethanol and biopolymer production in biorefinery framework. Bioresour Technol 296:122315
Chu Q et al (2018) Two-stage pretreatment with alkaline sulphonation and steam treatment of Eucalyptus woody biomass to enhance its enzymatic digestibility for bioethanol production. Energy Convers Manag 175(1):236–245
Gil LS, Maupoey PF (2018) An integrated approach for pineapple waste valorisation. Bioethanol production and bromelain extraction from pineapple residues. J Clean Prod 172(20):1224–1231
Saravanan KK et al (2018) Evaluation of the saccharification and fermentation process of two different seaweeds for an ecofriendly bioethanol production., Biocatalysis and Agricultural Biotechnology. Volume 14:444–449
Sindhu R et al (2011) Dilute acid pretreatment and enzymatic saccharification of sugarcane tops for bioethanol production. Bioresour Technol 102(23):10915–10921
Dong C, Wang Y, Zhang H, Leu SY (2018) Feasibility of high-concentration cellulosic bioethanol production from undetoxified whole Monterey pine slurry. Bioresour Technol 250:102–109
Silva FL et al (2018) Valorization of an agroextractive residue—Carnauba straw—for the production of bioethanol by simultaneous saccharification and fermentation (SSF). Renew Energy 127:661–669
Yuan Z, Wen Y, Li G (2018) Production of bioethanol and value added compounds from wheat straw through combined alkaline/alkaline-peroxide pretreatment. Bioresour Technol 259:228–236
Pilavtepe M, Celiktas MS, Sargin S, Yesil-Celiktas O (2013) Transformation of Posidonia oceanica residues to bioethanol. Ind Crop Prod 51:348–354
Yan J, Wei Z, Wang Q, He M, Li S, Irbis C (2015) Bioethanol production from sodium hydroxide/hydrogen peroxide-pretreated water hyacinth via simultaneous saccharification and fermentation with a newly isolated thermotolerant Kluyveromyces marxianu strain. Bioresour Technol 193:103–109
Tanaka K et al (2019) Production of high-concentration bioethanol from cassava stem by repeated hydrolysis and intermittent yeast inoculation. Int Biodeterior Biodegradation 138:1–7
Coban I, Sargin S, Celiktas MS, Yesil-Celiktas O (2012) Bioethanol production from raffinate phase of supercritical CO2 extracted Stevia rebaudiana leaves. Bioresour Technol 120:52–59
Favaro L et al (2017) Production of bioethanol from multiple waste streams of rice milling. Bioresour Technol 244(Part 1):151–159
Sheikh MMI et al (2013) Production of bioethanol from waste money bills – a new cellulosic material for biofuels. Food Bioprod Process 91(1):60–65
Wu FC et al (2014) Sequential acid and enzymatic hydrolysis in situ and bioethanol production from Gracilaria biomass. Bioresour Technol 156:123–131
Raghavi S, Sindhu R, Binod P, Gnansounou E, Pandey A (2016) Development of a novel sequential pretreatment strategy for the production of bioethanol from sugarcane trash. Bioresour Technol 199:202–210
Prajapati BP, Kumar U, Rahul J, Suryawanshi K, Kango N (2020) Sugarcane bagasse saccharification using Aspergillus tubingensis enzymatic cocktail for 2G bio-ethanol production. Renew Energy 152:653–663
Malik K, Salama ES, Kim TH, Li X (2020) Enhanced ethanol production by Saccharomyces cerevisiae fermentation post acidic and alkali chemical pretreatments of cotton stalk lignocellulose. Int Biodeterior Biodegrad 147:104869
Okuda N, Ninomiya K, Takao M, Katakura Y, Shioya S (2007) Microaeration enhances productivity of bioethanol from hydrolysate of waste house wood using ethanologenic Escherichia coli KO11. J Biosci Bioeng 103(4):350–357
Vučurović VM, Razmovski RN (2012) Sugar beet pulp as support for Saccharomyces cerivisiae immobilization in bioethanol production. Ind Crop Prod 39:128–134
Choi IS et al (2015) A low-energy, cost-effective approach to fruit and citrus peel waste processing for bioethanol production. Appl Energy 140:65–74
Hashem M, Darwish SMI (2010) Production of bioethanol and associated by-products from potato starch residue stream by Saccharomyces cerevisiae. Biomass Bioenergy 34(7):953–959
Gupta A, Das SP, Ghosh A, Choudhary R, Das D, Goyal A (2014) Bioethanol production from hemicellulose rich Populus nigra involving recombinant hemicellulases from Clostridium thermocellum., Bioresource Technology. Volume 165:205–213
Kuhad RC, Gupta R, Khasa YP, Singh A (2010) Bioethanol production from Lantana camara (red sage): pretreatment, saccharification and fermentation. Bioresour Technol 101(21):8348–8354
Yeh R-H et al (2016) Bioethanol production from pretreated Miscanthus floridulus biomass by simultaneous saccharification and fermentation. Biomass Bioenergy 94:110–116
Alam MA et al (2019) Process optimization for the production of high-concentration ethanol with Scenedesmus raciborskii biomass. Bioresour Technol 294:122219
Todhanakasem T et al (2014) Biofilm production by Zymomonas mobilis enhances ethanol production and tolerance to toxic inhibitors from rice bran hydrolysate. New Biotechnol 31(5):451–459
Prasad S, Malav MK, Kumar S, Singh A, Pant D, Radhakrishnan S (2018) Enhancement of bio-ethanol production potential of wheat straw by reducing furfural and 5-hydroxymethylfurfural (HMF). Bioresource Technol Rep 4:50–56
Lee JV, Li P, Lee J, Ryu HJ, Oh KK (2013) Ethanol production from Saccharina japonica using an optimized extremely low acid pretreatment followed by simultaneous saccharification and fermentation. Bioresour Technol 127:119–125
Rijal D, Vancov T, McIntosh S, Ashwath N, Grant A, Stanley GA (2016) Process options for conversion of Agave tequilana leaves into bioethanol. Ind Crop Prod 84:263–272
Sevgi ED, Karatay E, Dönmez G (2018) Evaluation of pomegranate peel in ethanol production by Saccharomyces cerevisiae and Pichia stipites. Energy. 159:988–994
Inokuma K, Takano M, Hoshino K (2013) Direct ethanol production from N-acetylglucosamine and chitin substrates by Mucor species. Biochem Eng J 72(15):24–32
Okamoto K, Nitta Y, Maekawa N, Yanase H (2011) Direct ethanol production from starch, wheat bran and rice straw by the white rot fungus Trametes hirsute., Enzyme and Microbial. Technology. 48(3):273–277
Yuan Z, Wei W, Li G et al (2020) High titer ethanol production from combined alkaline/alkaline hydrogen peroxide pretreated bamboo at high solid loading. Waste Biomass Valor 11:2795–2805. https://doi.org/10.1007/s12649-019-00638-5
Nishimura H et al (2017) Production of ethanol from a mixture of waste paper and kitchen waste via a process of successive liquefaction, presaccharification, and simultaneous saccharification and fermentation. Waste Manag 67:86–94
Nair RKB et al (2015) Dilute phosphoric acid pretreatment of wheat bran for enzymatic hydrolysis and subsequent ethanol production by edible fungi Neurospora intermedia. Ind Crop Prod 69:314–323
Siwarasak P, Pajantagate P, Prasertlertrat K (2012) Use of Trichoderma reesei RT-P1 crude enzyme powder for ethanol fermentation of sweet sorghum fresh stalks. Bioresour Technol 107:200–204
Zhao X, Dong L, Chen L, Liu D (2013) Batch and multi-step fed-batch enzymatic saccharification of formiline-pretreated sugarcane bagasse at high solid loadings for high sugar and ethanol titers. Bioresour Technol 135:350–356
Rezania S, Din F, Taib MS, Mohamad S, Dahalan FA, Kamyab H, Darajeh N, Ebrahimi SS (2018) Ethanol production from water hyacinth (Eichhornia crassipes) using various types of enhancers based on the consumable sugars. Waste Biomass Valor 9(6):939–946
Chandrakant P, Bisaria V (2000) Simultaneous bioconversion of glucose and xylose to ethanol by Saccharomyces cerevisiae in the presence of xylose isomerase. Appl Microbiol Biotechnol 53:301–309. https://doi.org/10.1007/s002530050025
Pietrzak W, Kawa-Rygielska J (2015) Simultaneous saccharification and ethanol fermentation of waste wheat–rye bread at very high solids loading: effect of enzymatic liquefaction conditions. Fuel. 147:236–242
Shen J, Agblevorb FA (2010) Modeling semi-simultaneous saccharification and fermentation of ethanol production from cellulose. Biomass Bioenergy 34(8):1098–1107
Castroa E et al (2014) Optimization of dilute-phosphoric-acid steam pretreatment of Eucalyptus benthamii for biofuel production. Appl Energy 125(15):76–83
Sudha S et al (2015) Simultaneous pretreatment and saccharification: Green technology for enhanced sugar yields from biomass using a fungal consortium. Bioresour Technol 179:50–57
Farias D, Ibraim D, Atala P, Maugeri F (2017) Improving bioethanol production by Scheffersomyces stipitis using retentostat extractive fermentation at high xylose concentration. Biochem Eng J 121:171–180
Carrillo-Nieves D et al (2017) Process alternatives for bioethanol production from mango stem bark residues. Bioresour Technol 239:430–436
Mihajlovski K et al (2018) Valorization of damaged rice grains: optimization of bioethanol production by waste brewer’s yeast using an amylolytic potential from the Paenibacillus chitinolyticus CKS1. Fuel. 224:591–599
José D et al (2018) Comparison of two pretreatments methods to produce second-generation bioethanol resulting from sugarcane bagasse. Ind Crop Prod 122:414–421
Dussán KJ, Justo OR, Perez VH et al (2019) Bioethanol production from sugarcane bagasse hemicellulose hydrolysate by immobilized S. shehatae in a fluidized bed fermenter under magnetic field. Bioenerg Res 12:338–346. https://doi.org/10.1007/s12155-019-09971-y
Salazar MM, Grandis A, Pattathil S et al (2016) Eucalyptus cell wall architecture: clues for lignocellulosic biomass deconstruction. Bioenerg Res 9:969–979. https://doi.org/10.1007/s12155-016-9770-y
Tadege M, Chen F, Murray J et al (2015) Control of vegetative to reproductive phase transition improves biomass yield and simultaneously reduces lignin content in Medicago truncatula. Bioenerg Res 8:857–867. https://doi.org/10.1007/s12155-014-9565-y
Duceppe M, Bertrand A, Pattathil S et al (2012) Assessment of genetic variability of cell wall degradability for the selection of alfalfa with improved saccharification efficiency. Bioenerg Res 5:904–914. https://doi.org/10.1007/s12155-012-9204-4
Brereton NJB, Pitre FE, Hanley SJ et al (2010) QTL mapping of enzymatic saccharification in short rotation coppice willow and its independence from biomass yield. Bioenerg Res 3:251–261. https://doi.org/10.1007/s12155-010-9077-3
Pawar PMA, Schnürer A, Mellerowicz EJ, Rönnberg-Wästljung AC (2018) QTL mapping of wood FT-IR chemotypes shows promise for improving biofuel potential in short rotation coppice willow (Salix spp.). BioEnergy Res 11:351–363
Buckeridge MS, de Souza AP (2014) Breaking the “Glycomic Code” of cell wall polysaccharides may improve second-generation bioenergy production from biomass. Bioenerg Res 7:1065–1073. https://doi.org/10.1007/s12155-014-9460-6
Turumtay H (2015) Cell wall engineering by heterologous expression of cell wall-degrading enzymes for better conversion of lignocellulosic biomass into biofuels. Bioenerg Res 8:1574–1588. https://doi.org/10.1007/s12155-015-9624-z
de Souza AP, Leite DCC, Pattathil S et al (2013) Composition and structure of sugarcane cell wall polysaccharides: implications for second-generation bioethanol production. Bioenerg Res 6:564–579. https://doi.org/10.1007/s12155-012-9268-1
Cook C, Francocci F, Cervone F, Bellincampi D, Bolwell PG, Ferrari S, Devoto A (2015) Combination of pretreatment with white rot fungi and modification of primary and secondary cell walls improves saccharification. Bioenerg Res 8:175–186
Kaida R, Kaku T, Baba K et al (2009) Enzymatic saccharification and ethanol production of Acacia mangium and Paraserianthes falcataria wood, and Elaeis guineensis trunk. J Wood Sci 55:381–386. https://doi.org/10.1007/s10086-009-1038-0
Yücel Y, Göycıncık S (2015) Optimization and modelling of process conditions using response surface methodology (RSM) for enzymatic saccharification of spent tea waste (STW). Waste Biomass Valor 6:1077–1084. https://doi.org/10.1007/s12649-015-9395-y
Pandey A, Soccol CR, Nigam P, Soccol VT (2000) Biotechnological potential of agro-industrial residues, I: sugarcane bagasse. Bioresource Technol 74:69–80
Talebnia F, Karakashev D, Angelidaki I (2010) Production of bioethanol from wheat straw: an overview on pretreatment, hydrolysis and fermentation. Bioresour Technol 101:4744–4753
Kumakura M, Kojima T, Koetsu I (1982) Pretreatment of lignocellulosic wastes by combination of irradiation and mechanical crushing. Biomass 2:299–300
Mathew GM, Sukumaran RK, Singhania RR, Pandey A (2008) Progress in research on fungal cellulases for lignocellulosic degradation. JSIR 67:898–907
Mosier N, Wyman C, Dale B, Elander R, Lee YY, Holtzapple M, Ladisch M (2005) Features of promising technologies for pretreatment of lignocellulosic biomass. Bioresour Technol 96:673–686
Balat M, Balat H, Oz C (2008) Progress in bioethanol processing. Prog Energy Combust Sci 34:551–573
Kumar AK, Sharma S (2017) Recent updates on different methods of pretreatment of lignocellulosic feedstocks: a review. Bioresour Bioprocess 4:7. https://doi.org/10.1186/s40643-017-0137-9
Jhansi LK et al (2019) Natural deep eutectic solvents (DES) for fractionation of waste lignocellulosic biomass and its cascade conversion to value-added bio-based chemicals. Biomass Bioenergy 120:417–425
Annamalai N, Al Battashi H, Anu SN et al (2020) Enhanced bioethanol production from waste paper through separate hydrolysis and fermentation. Waste Biomass Valor 11:121–131. https://doi.org/10.1007/s12649-018-0400-0
Smichi N, Messaoudi Y, Gelicus A et al (2020) Enzymatic hydrolysis of instant controlled pressure drop pretreated retama raetam for bioethanol production. Waste Biomass Valor 11:187–200. https://doi.org/10.1007/s12649-018-0366-y
Aruwajoye GS, Faloye FD, Kana EG (2019) Process optimisation of enzymatic saccharification of soaking assisted and thermal pretreated cassava peels waste for bioethanol production. Waste Biomass Valor. https://doi.org/10.1007/s12649-018-00562-0
Adaganti SY, Yaliwal VS, Kulkarni BM et al (2014) Factors affecting bioethanol production from lignocellulosic biomass (Calliandra calothyrsus). Waste Biomass Valor 5:963–971. https://doi.org/10.1007/s12649-014-9305-8
Siramon P, Punsuvon V, Vaithanomsat P (2018) Production of bioethanol from oil palm empty fruit bunch via acid impregnation-steam explosion pretreatment. Waste Biomass Valor 9:1407–1414. https://doi.org/10.1007/s12649-017-9924-y
Merino O, Almazán V, Martínez-Palou R et al (2017) Screening of ionic liquids for pretreatment of Taiwan grass in Q-tube minireactors for improving bioethanol production. Waste Biomass Valor 8:733–742. https://doi.org/10.1007/s12649-016-9612-3
Bhattacharyya S, Datta S, Bhattacharjee C (2012) Sonication boost the total reducing sugar (TRS) Extraction from sugarcane bagasse after dilute acid hydrolysis. Waste Biomass Valor 3:81–87. https://doi.org/10.1007/s12649-011-9078-2
Gonçalves FA, Ruiz HA, dos Santos ES et al (2019) Valorization, comparison and characterization of coconuts waste and cactus in a biorefinery context using NaClO2–C2H4O2 and sequential NaClO2–C2H4O2/autohydrolysis pretreatment. Waste Biomass Valor 10:2249–2262. https://doi.org/10.1007/s12649-018-0229-6
Chatterjee S, Barman S, Chakraborty R (2019) Combined effects of ionic liquid and tungsten–halogen radiation on heterogeneous hydrolysis kinetics of waste papaya epidermis for production of total reducing sugar. Waste Biomass Valor 10:1845–1855. https://doi.org/10.1007/s12649-018-0220-2
Sánchez-Ramírez J, Martínez-Hernández JL, López-Campos RG et al (2018) Laccase validation as pretreatment of agave waste prior to saccharification: free and immobilized in superparamagnetic nanoparticles enzyme preparations. Waste Biomass Valor 9:223–234. https://doi.org/10.1007/s12649-016-9774-z
Equihua-Sánchez M, Barahona-Pérez LF (2019) Physical and chemical characterization of Agave tequilana bagasse pretreated with the ionic liquid 1-ethyl-3-methylimidazolium acetate. Waste Biomass Valor 10(5):1285–1294
Chen J, Adjallé K, Lai TT et al (2019) Effect of mechanical pretreatment for enzymatic hydrolysis of woody residues, corn stover and alfalfa. Waste Biomass Valor. https://doi.org/10.1007/s12649-019-00856-x
Merino O, Cerón-Camacho R, Luque R, Martínez-Palou R(2019) Microwave-assisted lignin solubilization in protic ionic compounds containing 2,3,4,5-tetraphenyl-1H-imidazolium and inorganic anions. Waste Biomass Valor.
Tonucci L, Coccia F, Bressan M et al (2012) Mild photocatalysed and catalysed green oxidation of lignin: a useful pathway to low-molecular-weight derivatives. Waste Biomass Valor 3:165–174. https://doi.org/10.1007/s12649-011-9102-6
Miranda FS, Rabelo SC, Pradella JGC et al (2019) Plasma in-liquid using non-contact electrodes: a method of pretreatment to enhance the enzymatic hydrolysis of biomass. Waste Biomass Valor. https://doi.org/10.1007/s12649-019-00824-5
Yasuda M, Nagai H, Takeo K et al (2014) Bio-ethanol production through simultaneous saccharification and co-fermentation (SSCF) of a low-moisture anhydrous ammonia (LMAA)-pretreated napiegrass (Pennisetum purpureum Schumach). SpringerPlus 3:333. https://doi.org/10.1186/2193-1801-3-333
Bak JS (2014) Process evaluation of electron beam irradiation-based biodegradation relevant to lignocellulose bioconversion. SpringerPlus 3:487. https://doi.org/10.1186/2193-1801-3-487
Zhu Z, Macquarrie DJ, Simister R et al (2015) Microwave assisted chemical pretreatment of Miscanthus under different temperature regimes. Sustain Chem Process 3:15. https://doi.org/10.1186/s40508-015-0041-6
Klinke H, Thomsen A, Ahring B (2001) Potential inhibitors from wet oxidation of wheat straw and their effect on growth and ethanol production by Thermoanaerobacter mathranii. Appl Microbiol Biotechnol 57:631–638. https://doi.org/10.1007/s002530100825
Gupta D, Mahajani SM, Garga A (2019) Effect of hydrothermal carbonization as pretreatment on energy recovery from food and paper wastes. Bioresour Technol 285:121329
Negro MJ et al (2014) Ethanol production from glucose and xylose obtained from steam exploded water-extracted olive tree pruning using phosphoric acid as catalyst. Bioresour Technol 153:101–107
Lim WS et al (2013) Structural properties of pretreated biomass from different acid pretreatments and their effects on simultaneous saccharification and ethanol fermentation. Bioresour Technol 139:214–219
Prasad Y et al (2013) Effect of different pretreatments on delignification pattern and enzymatic hydrolysability of miscanthus, oil palm biomass and typha grass. Bioresource Technol 135:82–88
Kumar A, Singh S, Tiwari R, Goel R, Naina L (2017) Immobilization of indigenous holocellulase on iron oxide (Fe2O3) nanoparticles enhanced hydrolysis of alkali pretreated paddy straw. Int J Biol Macromol 96:538–549
Sasaki K et al (2015) Mechanical milling and membrane separation for increased ethanol production during simultaneous saccharification and co-fermentation of rice straw by xylose-fermenting Saccharomyces cerevisiae. Bioresour Technol 185:263–268
Preeti B, Parag S, Gogate R (2015) Ultrasound-assisted bioethanol production from waste newspaper. Ultrason Sonochem 27:37–45
Kanga KE, Chung DP, Kima Y, Chung BW, Choia GW (2015) High-titer ethanol production from simultaneous saccharification and fermentation using a continuous feeding system. Fuel. 145:18–24
Goshadrou A, Lefsrud M (2017) Synergistic surfactant-assisted [EMIM]OAc pretreatment of lignocellulosic waste for enhanced cellulose accessibility to cellulose. Carbohydr Polym 166:104–113
Jensena NS et al (2013) Pretreatment of the macroalgae Chaetomorpha linum for the production of bioethanol – comparison of five pretreatment technologies. Bioresour Technol 140:36–42
Farghalyae A et al (2017) Bioethanol production from paperboard mill sludge using acid-catalyzed bio-derived choline acetate ionic liquid pretreatment followed by fermentation process. Energy Convers Manag 145:255–264
Ahmed IN et al (2019) Zirconium based metal-organic framework in-situ assisted hydrothermal pretreatment and enzymatic hydrolysis of Platanus X acerifolia exfoliating bark for bioethanol production. Bioresour Technol 280:213–221
Elena M et al (2016) Second generation bioethanol from Eucalyptus globulus Labill and Nothofagus pumilio: Ionic liquid pretreatment boosts the yields. Ind Crop Prod 80:148–155
Khoshnevisan B et al (2016) Biogas and bioethanol production from pinewood pre-treated with steam explosion and N-methylmorpholine-N-oxide (NMMO): a comparative life cycle assessment approach. Energy 114:935–950
Ingle AP, Rafael R, Silvio P, da Silva S (2020) Pretreatment of sugarcane bagasse using two different acid-functionalized magnetic nanoparticles: a novel approach for high sugar recovery. Renew Energy 150:957–964
Yuana Z, Wenb Y (2017) Evaluation of an integrated process to fully utilize bamboo biomass during the production of bioethanol. Bioresour Technol 236:202–211
Qiu J (2018) Bioethanol production from wheat straw by phosphoric acid plus hydrogen peroxide (PHP) pretreatment via simultaneous saccharification and fermentation (SSF) at high solid loadings. Bioresour Technol 268:355–362
Mehmood A et al (2019) Acidic ionic liquids: promising and cost-effective solvents for processing of lignocellulosic biomass. J Mol Liq 287:110943
Yanga SJ et al (2015) Enhancement of enzymatic digestibility of Miscanthus by electron beam irradiation and chemical combined treatments for bioethanol production. Chem Eng J 275:227–234
Choi MYJ, Lee JW, Park DH (2012) Improvement of saccharification process for bioethanol production from Undaria sp. by gamma irradiation. Radiation Physics Chem 81(8):999–1002
Wanga M et al (2016) Bioethanol production from cotton stalk: a comparative study of various pretreatments. Fuel. 184(15):527–532
Subhedar PB, Babu NR, Gogate PR (2015) Intensification of enzymatic hydrolysis of waste newspaper using ultrasound for fermentable sugar production. Ultrason Sonochem 22:326–332
Victora A, Pulidindi IN, Gedankena A (2015) Assessment of holocellulose for the production of bioethanol by conserving Pinus radiata cones as renewable feedstock. J Environ Manag 162:215–220
Kumar S et al (2018) Potential of Trametes maxima IIPLC-32 derived laccase for the detoxification of phenolic inhibitors in lignocellulosic biomass prehydrolysate. Int Biodeterior Biodegradation 133:1–8
Saadiah H et al (2017) Feasibility of using kitchen waste as future substrate for bioethanol production: a review. Renew Sust Energ Rev 74:671–686
Zhang X, Wang J, Zhang W et al (2018) Optimizing the coordinated transcription of central xylose-metabolism genes in Saccharomyces cerevisiae. Appl Microbiol Biotechnol 102:7207–7217. https://doi.org/10.1007/s00253-018-9172-5
Stolarskia MJ et al (2015) Lignocellulosic biomass from short rotation woody crops as a feedstock for second-generation bioethanol production. Ind Crop Prod 75(Part B):66–75
Bañuelosa JA, Hernández IV, Guerra-Balcázar M, Arjona N (2018) Production, characterization and evaluation of the energetic capability of bioethanol from Salicornia Bigelovii as a renewable energy source. Renew Energy 123:125–134
Bua J et al (2019) Co-production of high-gravity bioethanol and succinic acid from potassium peroxymonosulfate and deacetylation sequentially pretreated sugarcane bagasse by simultaneous saccharification and co-fermentation. Energy Convers Manag 186:131–139
Ulaganathan K, Goud S, Reddy M, Kayalvili U (2017) Genome engineering for breaking barriers in lignocellulosic bioethanol production. Renew Sust Energ Rev 74:1080–1107
Guo X, Li H, Yan H et al (2019) Production of organic carboxylic acids by hydrothermal conversion of electron beam irradiation pretreated wheat straw. Biomass Conv Bioref. https://doi.org/10.1007/s13399-019-00471-9
Bhardwaj N, Kumar B, Verma P (2020) Microwave-assisted pretreatment using alkali metal salt in combination with orthophosphoric acid for generation of enhanced sugar and bioethanol. Biomass Conv Bioref. https://doi.org/10.1007/s13399-020-00640-1
da Silva ARG, Errico M, Rong B (2018) Evaluation of organosolv pretreatment for bioethanol production from lignocellulosic biomass: solvent recycle and process integration. Biomass Conv Bioref 8:397–411. https://doi.org/10.1007/s13399-017-0292-4
Lv S, Yu Q, Zhuang X et al (2013) The influence of hemicellulose and lignin removal on the enzymatic digestibility from sugarcane bagasse. Bioenerg Res 6:1128–1134. https://doi.org/10.1007/s12155-013-9297-4
Rosen Y, Mamane H, Gerchman Y (2019) Short ozonation of lignocellulosic waste as energetically favorable pretreatment. Bioenerg Res 12:292–301. https://doi.org/10.1007/s12155-019-9962-3
Sannigrahi P, Ragauskas AJ, Miller SJ (2008) Effects of two-stage dilute acid pretreatment on the structure and composition of lignin and cellulose in loblolly pine. Bioenerg Res 1:205–214. https://doi.org/10.1007/s12155-008-9021-y
Rencoret J, Pereira A, del Río JC et al (2016) Laccase-mediator pretreatment of wheat straw degrades lignin and improves saccharification. Bioenerg Res 9:917–930. https://doi.org/10.1007/s12155-016-9745-z
Montiel C, Hernández-Meléndez O, Vivaldo-Lima E et al (2016) Enhanced bioethanol production from blue agave bagasse in a combined extrusion–saccharification process. Bioenerg Res 9:1005–1014. https://doi.org/10.1007/s12155-016-9747-x
Gladchenko MA, Gaydamaka SN, Murygina VP et al (2019) Anaerobic conversion of lignocellulose to materials for biofuel production: volatile fatty acids and ethanol. Appl Biochem Microbiol 55:756–764. https://doi.org/10.1134/S0003683819070020//////
Agung U, Wijayantac T, Gotode M, Kamiyade N (2015) Great potency of seaweed waste biomass from the carrageenan industry for bioethanol production by peracetic acid–ionic liquid pretreatment. Biomass Bioenergy 81:63–69
Yang Y, Hu M, Tang Y et al (2018) Progress and perspective on lignocellulosic hydrolysate inhibitor tolerance improvement in Zymomonas mobilis. Bioresour Bioprocess 5:6. https://doi.org/10.1186/s40643-018-0193-9
Adeboye PT, Bettiga M, Olsson L (2014) The chemical nature of phenolic compounds determines their toxicity and induces distinct physiological responses in Saccharomyces cerevisiae in lignocellulose hydrolysates. AMB Express 4:46. https://doi.org/10.1186/s13568-014-0046-7
Wirawana F et al (2020) Continuous cellulosic bioethanol co-fermentation by immobilized Zymomonas mobilis and suspended Pichia stipitis in a two-stage process. Appl Energy 266:114871
Aristidou A, Penttilä M (2000) Metabolic engineering applications to renewable resource utilization. Curr Opin Biotechnol 11(2):187–198
Chena X et al (2013) Metabolic engineering of Escherichia coli: a sustainable industrial platform for bio-based chemical production. Biotechnol Adv 31(8):1200–1223
Xue T, Liu K, Chen D et al (2018) Improved bioethanol production using CRISPR/Cas9 to disrupt the ADH2 gene in Saccharomyces cerevisiae. World J Microbiol Biotechnol 34:154. https://doi.org/10.1007/s11274-018-2518-4
Guo C, Jiang N (2013) Physiological and enzymatic comparison between Pichia stipitis and recombinant Saccharomyces cerevisiae on xylose fermentation. World J Microbiol Biotechnol 29:541–547. https://doi.org/10.1007/s11274-012-1208-x
Tambalo RD, Raymundo AK, Grunden AM (2020) Thermostable endoglucanase gene derived by amplification from the genomic DNA of a cellulose-enriched mixed culture from mudspring water of Mt. Makiling, Laguna, Philippines. World J Microbiol Biotechnol 36:51. https://doi.org/10.1007/s11274-020-02825-2
Wallace-Salinas V, Signori L, Li Y et al (2014) Re-assessment of YAP1 and MCR1 contributions to inhibitor tolerance in robust engineered Saccharomyces cerevisiae fermenting undetoxified lignocellulosic hydrolysate. AMB Express 4:56. https://doi.org/10.1186/s13568-014-0056-5
Zeng R, Hu Q, Yin X et al (2016) Cloning a novel endo-1,4-β-d-glucanase gene from Trichoderma virens and heterologous expression in E. coli. AMB Express 6:108. https://doi.org/10.1186/s13568-016-0282-0
Kanna M, Yano S, Inoue H et al (2011) Enhancement of β-xylosidase productivity in cellulase producing fungus Acremonium cellulolyticus. AMB Express 1:15. https://doi.org/10.1186/2191-0855-1-15
Ellilä S, Bromann P, Nyyssönen M et al (2019) Cloning of novel bacterial xylanases from lignocellulose-enriched compost metagenomic libraries. AMB Express 9:124. https://doi.org/10.1186/s13568-019-0847-9
Kitichantaropas Y, Boonchird C, Sugiyama M et al (2016) Cellular mechanisms contributing to multiple stress tolerance in Saccharomyces cerevisiae strains with potential use in high-temperature ethanol fermentation. AMB Express 6:107. https://doi.org/10.1186/s13568-016-0285-x
Kim H, Kim N, Yang J et al (2011) Identification of novel genes responsible for ethanol and/or thermotolerance by transposon mutagenesis in Saccharomyces cerevisiae. Appl Microbiol Biotechnol 91:1159–1172. https://doi.org/10.1007/s00253-011-3298-z
Zhang K, Fang Y, Gao K et al (2017) Effects of genome duplication on phenotypes and industrial applications of Saccharomyces cerevisiae strains. Appl Microbiol Biotechnol 101:5405–5414. https://doi.org/10.1007/s00253-017-8284-7
Fu X, Li P, Zhang L et al (2019) Understanding the stress responses of Kluyveromyces marxianus after an arrest during high-temperature ethanol fermentation based on integration of RNA-Seq and metabolite data. Appl Microbiol Biotechnol 103:2715–2729. https://doi.org/10.1007/s00253-019-09637-x
Van Zyl JHD, Den Haan R, Van Zyl WH (2014) Over-expression of native Saccharomyces cerevisiae exocytic SNARE genes increased heterologous cellulase secretion. Appl Microbiol Biotechnol 98:5567–5578. https://doi.org/10.1007/s00253-014-5647-1
Li Y, Zeng W, Gou M et al (2017) Transcriptome changes in adaptive evolution of xylose-fermenting industrial Saccharomyces cerevisiae strains with δ-integration of different xylA genes. Appl Microbiol Biotechnol 101:7741–7753. https://doi.org/10.1007/s00253-017-8494-z
Loaces I, Amarelle V, Muñoz-Gutierrez I et al (2015) Improved ethanol production from biomass by a rumen metagenomic DNA fragment expressed in Escherichia coli MS04 during fermentation. Appl Microbiol Biotechnol 99:9049–9060. https://doi.org/10.1007/s00253-015-6801-0
Matsushika A, Watanabe S, Kodaki T et al (2008) Expression of protein engineered NADP+-dependent xylitol dehydrogenase increases ethanol production from xylose in recombinant Saccharomyces cerevisiae. Appl Microbiol Biotechnol 81:243–255. https://doi.org/10.1007/s00253-008-1649-1
Bravim F, Lippman SI, da Silva LF et al (2013) High hydrostatic pressure activates gene expression that leads to ethanol production enhancement in a Saccharomyces cerevisiae distillery strain. Appl Microbiol Biotechnol 97:2093–2107. https://doi.org/10.1007/s00253-012-4356-x/
Takagi H (2008) Proline as a stress protectant in yeast: physiological functions, metabolic regulations, and biotechnological applications. Appl Microbiol Biotechnol 81:211–223. https://doi.org/10.1007/s00253-008-1698-5
Suwannarangsee S, Oh D, Seo J et al (2010) Characterization of alcohol dehydrogenase 1 of the thermotolerant methylotrophic yeast Hansenula polymorpha. Appl Microbiol Biotechnol 88:497–507. https://doi.org/10.1007/s00253-010-2752-7//
Rodrussamee N, Lertwattanasakul N, Hirata K et al (2011) Growth and ethanol fermentation ability on hexose and pentose sugars and glucose effect under various conditions in thermotolerant yeast Kluyveromyces marxianus. Appl Microbiol Biotechnol 90:1573–1586. https://doi.org/10.1007/s00253-011-3218-2
Shanmugam S, Ngo HH, Wu YR (2020) abc., Advanced CRISPR/Cas-based genome editing tools for microbial biofuels production: a review. Renew Energy. 149:1107–1119
Kumari R, Pramanik K (2012) Improvement of multiple stress tolerance in yeast strain by sequential mutagenesis for enhanced bioethanol production. J Biosci Bioeng 114(6):622–629
Kitagawa T, Tokuhiro K, Sugiyama H et al (2010) Construction of a β-glucosidase expression system using the multistress-tolerant yeast Issatchenkia orientalis. Appl Microbiol Biotechnol 87:1841–1853. https://doi.org/10.1007/s00253-010-2629-9
Hasunuma T, Sung K, Sanda T et al (2011) Efficient fermentation of xylose to ethanol at high formic acid concentrations by metabolically engineered Saccharomyces cerevisiae. Appl Microbiol Biotechnol 90:997–1004. https://doi.org/10.1007/s00253-011-3085-x
Kim H, Kim N, Kim W et al (2012) Insertion of transposon in the vicinity of SSK2 confers enhanced tolerance to furfural in Saccharomyces cerevisiae. Appl Microbiol Biotechnol 95:531–540. https://doi.org/10.1007/s00253-012-4022-3
Almeida JRM, Röder A, Modig T et al (2008) NADH- vs NADPH-coupled reduction of 5-hydroxymethyl furfural (HMF) and its implications on product distribution in Saccharomyces cerevisiae. Appl Microbiol Biotechnol 78:939–945. https://doi.org/10.1007/s00253-008-1364-y
Sadie CJ, Rose SH, den Haan R, van Zyl WH (2011) Co-expression of a cellobiose phosphorylase and lactose permease enables intracellular cellobiose utilisation by Saccharomyces cerevisiae. Appl Microbiol Biotechnol 90(4):1373–1380. https://doi.org/10.1007/s00253-011-3164-z
Zeng W, Tang Y, Gou M et al (2017) Comparative transcriptomes reveal novel evolutionary strategies adopted by Saccharomyces cerevisiae with improved xylose utilization capability. Appl Microbiol Biotechnol 101:1753–1767. https://doi.org/10.1007/s00253-016-8046-y
Uchima CA, Tokuda G, Watanabe H et al (2011) Heterologous expression and characterization of a glucose-stimulated β-glucosidase from the termite Neotermes koshunensis in Aspergillus oryzae. Appl Microbiol Biotechnol 89:1761–1771. https://doi.org/10.1007/s00253-010-2963-y
Tanino T, Hotta A, Ito T et al (2010) Construction of a xylose-metabolizing yeast by genome integration of xylose isomerase gene and investigation of the effect of xylitol on fermentation. Appl Microbiol Biotechnol 88:1215–1221. https://doi.org/10.1007/s00253-010-2870-2
Katahira S, Mizuike A, Fukuda H et al (2006) Ethanol fermentation from lignocellulosic hydrolysate by a recombinant xylose- and cellooligosaccharide-assimilating yeast strain. Appl Microbiol Biotechnol 72:1136–1143. https://doi.org/10.1007/s00253-006-0402-x
Jayakody LN, Horie K, Hayashi N et al (2012) Improvement of tolerance of Saccharomyces cerevisiae to hot-compressed water-treated cellulose by expression of ADH1. Appl Microbiol Biotechnol 94:273–283. https://doi.org/10.1007/s00253-012-3918-2
Van Zyl LJ, Taylor MP, Eley K, Tuffin M, Cowan DA (2014) Engineering pyruvate decarboxylase-mediated ethanol production in the thermophilic host Geobacillus thermoglucosidasius. Appl Microbiol Biotechnol 98(3):1247–1259. https://doi.org/10.1007/s00253-013-5380-1
Hou X, Yao S (2012) Improved inhibitor tolerance in xylose-fermenting yeast Spathaspora passalidarum by mutagenesis and protoplast fusion. Appl Microbiol Biotechnol 93:2591–2601. https://doi.org/10.1007/s00253-011-3693-5
Liu ZL (2006) Genomic adaptation of ethanologenic yeast to biomass conversion inhibitors. Appl Microbiol Biotechnol 73:27–36. https://doi.org/10.1007/s00253-006-0567-3
Jayakody LN, Kadowaki M, Tsuge K et al (2015) SUMO expression shortens the lag phase of Saccharomyces cerevisiae yeast growth caused by complex interactive effects of major mixed fermentation inhibitors found in hot-compressed water-treated lignocellulosic hydrolysate. Appl Microbiol Biotechnol 99:501–515. https://doi.org/10.1007/s00253-014-6174-9
Zheng D, Liu T, Chen J et al (2013) Comparative functional genomics to reveal the molecular basis of phenotypic diversities and guide the genetic breeding of industrial yeast strains. Appl Microbiol Biotechnol 97:2067–2076. https://doi.org/10.1007/s00253-013-4698-z
Kato H, Suyama H, Yamada R et al (2012) Improvements in ethanol production from xylose by mating recombinant xylose-fermenting Saccharomyces cerevisiae strains. Appl Microbiol Biotechnol 94:1585–1592. https://doi.org/10.1007/s00253-012-3914-6
Kojima M, Okamoto K, Yanase H (2013) Direct ethanol production from cellulosic materials by Zymobacter palmae carrying Cellulomonas endoglucanase and Ruminococcus β-glucosidase genes. Appl Microbiol Biotechnol 97:5137–5147. https://doi.org/10.1007/s00253-013-4874-1
Sanny T, Arnaldos M, Kunkel SA et al (2010) Engineering of ethanolic E. coli with the Vitreoscilla hemoglobin gene enhances ethanol production from both glucose and xylose. Appl Microbiol Biotechnol 88:1103–1112. https://doi.org/10.1007/s00253-010-2817-7
Yanase S, Hasunuma T, Yamada R et al (2010) Direct ethanol production from cellulosic materials at high temperature using the thermotolerant yeast Kluyveromyces marxianus displaying cellulolytic enzymes. Appl Microbiol Biotechnol 88:381–388. https://doi.org/10.1007/s00253-010-2784-z
Yanase H, Miyawaki H, Sakurai M et al (2012) Ethanol production from wood hydrolysate using genetically engineered Zymomonas mobilis. Appl Microbiol Biotechnol 94:1667–1678. https://doi.org/10.1007/s00253-012-4094-0.///
Lee JS, Chi W, Hong S et al (2013) Bioethanol production by heterologous expression of Pdc and AdhII in Streptomyces lividans. Appl Microbiol Biotechnol 97:6089–6097. https://doi.org/10.1007/s00253-013-4951-5
Boyce A, Walsh G (2015) Characterisation of a novel thermostable endoglucanase from Alicyclobacillus vulcanalis of potential application in bioethanol production. Appl Microbiol Biotechnol 99:7515–7525. https://doi.org/10.1007/s00253-015-6474-8
Chen X, Xiao Y, Shen W et al (2016) Display of phytase on the cell surface of Saccharomyces cerevisiae to degrade phytate phosphorus and improve bioethanol production. Appl Microbiol Biotechnol 100:2449–2458. https://doi.org/10.1007/s00253-015-7170-4
Yang F, Cao M, Jin Y et al (2013) Construction of a novel а-agglutinin expression system on the surface of wild-type Saccharomyces cerevisiae Y5 and genetic immobilization of β-glucosidase1. Bioenerg Res 6:1205–1211. https://doi.org/10.1007/s12155-013-9312-9
Li X, Yang R, Ma M et al (2015) A novel aldehyde reductase encoded by YML131W from Saccharomyces cerevisiae confers tolerance to furfural derived from lignocellulosic biomass conversion. Bioenerg Res 8:119–129. https://doi.org/10.1007/s12155-014-9506-9
Generoso WC, Malagó-Jr W, Pereira-Jr N et al (2016) Triggering the expression of cellulolytic genes using a recombinant endoxylanase from Trichoderma harzianum IOC-3844. Bioenerg Res 9:931–941. https://doi.org/10.1007/s12155-016-9748-9
Feng C, Zou S, Liu C et al (2016) Ethanol production from acid- and alkali-pretreated corncob by endoglucanase and β-glucosidase co-expressing Saccharomyces cerevisiae subject to the expression of heterologous genes and nutrition added. World J Microbiol Biotechnol 32:86
Komesu A, Oliveira J, Neto JM, Penteado ED, Diniz AAR, Martins LHDS (2020) Chapter 10 - Xylose fermentation to bioethanol production using genetic engineering microorganisms, Editor(s): Arindam Kuila, Vinay Sharma, Genetic and metabolic engineering for improved biofuel production from lignocellulosic biomass, Elsevier, 143-154, ISBN 9780128179536, https://doi.org/10.1016/B978-0-12-817953-6.00010-5.
Kobayashi Y, Sahara T, Ohgiya S et al (2018) Systematic optimization of gene expression of pentose phosphate pathway enhances ethanol production from a glucose/xylose mixed medium in a recombinant Saccharomyces cerevisiae. AMB Express 8:139. https://doi.org/10.1186/s13568-018-0670-8
Bamba T, Hasunuma T, Kondo A (2016) Disruption of PHO13 improves ethanol production via the xylose isomerase pathway. AMB Express 6:4. https://doi.org/10.1186/s13568-015-0175-7
Yamada R, Nakatani Y, Ogino C et al (2013) Efficient direct ethanol production from cellulose by cellulase- and cellodextrin transporter-co-expressing Saccharomyces cerevisiae. AMB Express 3:34. https://doi.org/10.1186/2191-0855-3-34
Kahar P, Tanaka SA (2014) xylose-fermenting yeast hybridized by intergeneric fusion between Saccharomyces cerevisiae and Candida intermediamutants for ethanol production. Sustain Chem Process 2:17. https://doi.org/10.1186/s40508-014-0017-y
Olofsson K, Runquist D, Hahn-Hägerdal B et al (2011) A mutated xylose reductase increases bioethanol production more than a glucose/xylose facilitator in simultaneous fermentation and co-fermentation of wheat straw. AMB Express 1:4. https://doi.org/10.1186/2191-0855-1-4
Lee Y, Nasution O, Choi E et al (2015) Transcriptome analysis of acetic-acid-treated yeast cells identifies a large set of genes whose overexpression or deletion enhances acetic acid tolerance. Appl Microbiol Biotechnol 99:6391–6403. https://doi.org/10.1007/s00253-015-6706-y
Gruninger RJ, Gong X, Forster RJ et al (2014) Biochemical and kinetic characterization of the multifunctional β-glucosidase/β-xylosidase/α-arabinosidase, Bgxa1. Appl Microbiol Biotechnol 98:3003–3012. https://doi.org/10.1007/s00253-013-5191-4
Wang X, Ma M, Liu ZL et al (2016) GRE2 from Scheffersomyces stipitis as an aldehyde reductase contributes tolerance to aldehyde inhibitors derived from lignocellulosic biomass. Appl Microbiol Biotechnol 100:6671–6682. https://doi.org/10.1007/s00253-016-7445-4
Kim I, Kim Y, Yoon H (2013) Expression of salt-induced 2-Cys peroxiredoxin from Oryza sativa increases stress tolerance and fermentation capacity in genetically engineered yeast Saccharomyces cerevisiae. Appl Microbiol Biotechnol 97:3519–3533. https://doi.org/10.1007/s00253-012-4410-8
Wang H, Li Q, Kuang X et al (2018) Functions of aldehyde reductases from Saccharomyces cerevisiae in detoxification of aldehyde inhibitors and their biotechnological applications. Appl Microbiol Biotechnol 102:10439–10456. https://doi.org/10.1007/s00253-018-9425-3
Zaldivar J, Borges A, Johansson B et al (2002) Fermentation performance and intracellular metabolite patterns in laboratory and industrial xylose-fermenting Saccharomyces cerevisiae. Appl Microbiol Biotechnol 59:436–442. https://doi.org/10.1007/s00253-002-1056-y
Lee Y, Nasution O, Lee YM et al (2017) Overexpression of PMA1 enhances tolerance to various types of stress and constitutively activates the SAPK pathways in Saccharomyces cerevisiae. Appl Microbiol Biotechnol 101:229–239. https://doi.org/10.1007/s00253-016-7898-5
Stanley D, Chambers PJ, Stanley GA, Borneman A, Fraser S (2010) Transcriptional changes associated with ethanol tolerance in Saccharomyces cerevisiae. Appl Microbiol Biotechnol 88(1):231–239. https://doi.org/10.1007/s00253-010-2760-7
Lewis Liu Z, Moon J, Andersh BJ et al (2008) Multiple gene-mediated NAD(P)H-dependent aldehyde reduction is a mechanism of in situ detoxification of furfural and 5-hydroxymethylfurfural by Saccharomyces cerevisiae. Appl Microbiol Biotechnol 81:743–753
Hasunuma T, KuIsmail KS, Nambu Y, Kondo A (2014) Co-expression of TAL1 and ADH1 in recombinant xylose-fermenting Saccharomyces cerevisiae improves ethanol production from lignocellulosic hydrolysates in the presence of furfural. J Biosci Bioeng 117(2):165–169
Fujitomia K, Sanda T, Hasunum T, Kondo A (2012) Deletion of the PHO13 gene in Saccharomyces cerevisiae improves ethanol production from lignocellulosic hydrolysate in the presence of acetic and formic acids, and furfural. Bioresour Technol 111:161–166
Joana T et al (2015) Contribution of PRS3, RPB4 and ZWF1 to the resistance of industrial Saccharomyces cerevisiae CCUG53310 and PE-2 strains to lignocellulosic hydrolysate-derived inhibitors. Bioresour Technol 191:7–16
Pooya M et al (2019) Progress toward improving ethanol production through decreased glycerol generation in Saccharomyces cerevisiae by metabolic and genetic engineering approaches. Renew Sust Energ Rev 115:109353
Sasano Y, Watanabe D, Ukibe K et al (2012) Overexpression of the yeast transcription activator Msn2 confers furfural resistance and increases the initial fermentation rate in ethanol production. J Biosci Bioeng 113(4):451–455
Mohammad S, Khattab R, Saimura M, Kodaki T (2013) Boost in bioethanol production using recombinant Saccharomyces cerevisiae with mutated strictly NADPH-dependent xylose reductase and NADP+-dependent xylitol dehydrogenase. J Biotechnol 165(3–4):153–156
Mohammad S, Khattab R, Saimura M, Kodaki T (2014) Efficient bioethanol production by overexpression of endogenous Saccharomyces cerevisiae xylulokinase and NADPH-dependent aldose reductase with mutated strictly NADP+-dependent Pichia stipitis xylitol dehydrogenase. Process Biochem 4911:1838–1842
Joa JH, Park YC, Jin YS, Seo JH (2017) Construction of efficient xylose-fermenting Saccharomyces cerevisiae through a synthetic isozyme system of xylose reductase from Scheffersomyces stipites. Bioresour Technol 241:88–94
Sakihama Y, Hasunuma T, Kondo A (2015) Improved ethanol production from xylose in the presence of acetic acid by the overexpression of the HAA1 gene in Saccharomyces cerevisiae. J Biosci Bioeng 119(3):297–302
Basso TO et al (2011) Engineering topology and kinetics of sucrose metabolism in Saccharomyces cerevisiae for improved ethanol yield. Metab Eng 13(6):694–703
Yao R, Shimizu K (2013) Recent progress in metabolic engineering for the production of biofuels and biochemicals from renewable sources with particular emphasis on catabolite regulation and its modulation. Process Biochem 48(9):1409–1417
Hou J, Jiao C, Peng B, Shen Y, Bao X (2016) Mutation of a regulator Ask10p improves xylose isomerase activity through up-regulation of molecular chaperones in Saccharomyces cerevisiae. Metab Eng 38:241–250
Saitoh S, Hasunuma T, Tanaka T, Kondo A (2010) Co-fermentation of cellobiose and xylose using beta-glucosidase displaying diploid industrial yeast strain OC-2. Appl Microbiol Biotechnol 87(5):1975–1982. https://doi.org/10.1007/s00253-010-2714-0
Shen Y, Chen X, Peng B et al (2012) An efficient xylose-fermenting recombinant Saccharomyces cerevisiae strain obtained through adaptive evolution and its global transcription profile. Appl Microbiol Biotechnol 96:1079–1091. https://doi.org/10.1007/s00253-012-4418-0
Saitoh S, Tanaka T, Kondo A (2011) Co-fermentation of cellulose/xylan using engineered industrial yeast strain OC-2 displaying both β-glucosidase and β-xylosidase. Appl Microbiol Biotechnol 91:1553–1559. https://doi.org/10.1007/s00253-011-3357-5
Madhavan A, Tamalampudi S, Srivastava A et al (2009) Alcoholic fermentation of xylose and mixed sugars using recombinant Saccharomyces cerevisiae engineered for xylose utilization. Appl Microbiol Biotechnol 82:1037–1047. https://doi.org/10.1007/s00253-008-1818-2
Kitagawa T et al (2011) Identification of genes that enhance cellulase protein production in yeast. J Biotechnol 151(2):194–203
Hughes SR et al (2009) Engineered Saccharomyces cerevisiae strain for improved xylose utilization with a three-plasmid SUMO yeast expression system. Plasmid 61(1):22–38
Matsushika A et al (2012) Characterization of non-oxidative transaldolase and transketolase enzymes in the pentose phosphate pathway with regard to xylose utilization by recombinant Saccharomyces cerevisiae. Enzym Microb Technol 51(1):10,16–10,25
Silva P et al (2017) The relation between xyr1 overexpression in Trichoderma harzianum and sugarcane bagasse saccharification performance. J Biotechnol 246(20):24–32
Ogunyewo OA et al (2020) Engineered Penicillium funiculosum produces potent lignocellulolytic enzymes for saccharification of various pretreated biomasses. Process Biochem 92:49–60
Bae Y-H et al (2014) Deletion of the HXK2 gene in Saccharomyces cerevisiae enables mixed sugar fermentation of glucose and galactose in oxygen-limited conditions. Process Biochem 49(4):547–553
Kroukamp H et al (2013) Overexpression of native PSE1 and SOD1 in Saccharomyces cerevisiae improved heterologous cellulase secretion. Appl Energy 102:150–156
Akram F et al (2016) Cloning with kinetic and thermodynamic insight of a novel hyperthermostable β-glucosidase from Thermotoga naphthophila RKU-10T with excellent glucose tolerance. J Mol Catal B Enzym 124:92–104
Khaw TS, Katakura Y, Koh J et al (2006) Evaluation of performance of different surface-engineered yeast strains for direct ethanol production from raw starch. Appl Microbiol Biotechnol 70:573–579. https://doi.org/10.1007/s00253-005-0101-z
Nakanishi A, Bae JG, Fukai K et al (2012) Effect of pretreatment of hydrothermally processed rice straw with laccase-displaying yeast on ethanol fermentation. Appl Microbiol Biotechnol 94:939–948. https://doi.org/10.1007/s00253-012-3876-8
Gao G, Mao R, Xiao Y et al (2017) Efficient yeast cell-surface display of an endoglucanase of Aspergillus flavus and functional characterization of the whole-cell enzyme. World J Microbiol Biotechnol 33:114. https://doi.org/10.1007/s11274-016-2182-5
Kojima M, Akahoshi T, Okamoto K et al (2012) Expression and surface display of Cellulomonas endoglucanase in the ethanologenic bacterium Zymobacter palmae. Appl Microbiol Biotechnol 96:1093–1104. https://doi.org/10.1007/s00253-012-4424-2
Sakamoto T, Hasunum T, Hori Y, Yamada R, Kondo A (2012) Direct ethanol production from hemicellulosic materials of rice straw by use of an engineered yeast strain codisplaying three types of hemicellulolytic enzymes on the surface of xylose-utilizing Saccharomyces cerevisiae cells. J Biotechnol 158(4):203–210
Ryu S, Karim MN (2011) A whole cell biocatalyst for cellulosic ethanol production from dilute acid-pretreated corn stover hydrolyzates. Appl Microbiol Biotechnol 91:529–542. https://doi.org/10.1007/s00253-011-3261-z
Casa-Villegas M, Marín-Navarro J, Polaina J (2017) Synergies in coupled hydrolysis and fermentation of cellulose using a Trichoderma reesei enzyme preparation and a recombinant Saccharomyces cerevisiae strain. World J Microbiol Biotechnol 33:140. https://doi.org/10.1007/s11274-017-2308-4
Li M, Hu H, Chen Z et al (2018) Using drug-loaded pH-responsive poly(4-vinylpyridine) microspheres as a new strategy for intelligent controlling of Lactobacillus plantarum contamination in bioethanol fermentation. World J Microbiol Biotechnol 34:146. https://doi.org/10.1007/s11274-018-2533-5
Cui F, Zhang R, Liu H et al (2015) Metabolic responses to Lactobacillus plantarum contamination or bacteriophage treatment in Saccharomyces cerevisiae using a GC–MS-based metabolomics approach. World J Microbiol Biotechnol 31:2003–2013. https://doi.org/10.1007/s11274-015-1949-4
Ceccato-Antonini SR (2018) Conventional and nonconventional strategies for controlling bacterial contamination in fuel ethanol fermentations. World J Microbiol Biotechnol 34:80. https://doi.org/10.1007/s11274-018-2463-2
Branco P, Sabir F, Diniz M et al (2019) Biocontrol of Brettanomyces/Dekkera bruxellensis in alcoholic fermentations using saccharomycin-overproducing Saccharomyces cerevisiae strains. Appl Microbiol Biotechnol 103:3073–3083. https://doi.org/10.1007/s00253-019-09657-7
Lourencetti NMS, Wolf IR, Lacerda MPF et al (2018) Transcriptional profile of a bioethanol production contaminant Candida tropicalis. AMB Express 8:166. https://doi.org/10.1186/s13568-018-0693-1
Horinouchi T, Maeda T, Furusawa C (2018) Understanding and engineering alcohol-tolerant bacteria using OMICS technology. World J Microbiol Biotechnol 34(11):157. https://doi.org/10.1007/s11274-018-2542-4
Ding J, Huang X, Zhang L et al (2009) Tolerance and stress response to ethanol in the yeast Saccharomyces cerevisiae. Appl Microbiol Biotechnol 85:253–263. https://doi.org/10.1007/s00253-009-2223-1
Martínez-Alcantar L, Díaz-Pérez AL, Campos-García J (2019) The isc gene cluster expression ethanol tolerance associated improves its ethanol production by organic acids flux redirection in the ethanologenic Escherichia coli KO11 strain. World J Microbiol Biotechnol 35:189. https://doi.org/10.1007/s11274-019-2769-8
Geng P, Zhang L, Shi GY (2017) Omics analysis of acetic acid tolerance in Saccharomyces cerevisiae. World J Microbiol Biotechnol 33:94. https://doi.org/10.1007/s11274-017-2259-9
Cheng L, Zhang X, Zheng X et al (2019) RNA-seq transcriptomic analysis of green tea polyphenols regulation of differently expressed genes in Saccharomyces cerevisiae under ethanol stress. World J Microbiol Biotechnol 35:59. https://doi.org/10.1007/s11274-019-2639-4
Chen Y, Zhang X, Zhang M et al (2018) A transcriptome analysis of the ameliorate effect of Cyclocarya paliurus triterpenoids on ethanol stress in Saccharomyces cerevisiae. World J Microbiol Biotechnol 34:182. https://doi.org/10.1007/s11274-018-2561-1
Ren G, Ma A, Liu W et al (2016) Bacterial signals N-acyl homoserine lactones induce the changes of morphology and ethanol tolerance in Saccharomyces cerevisiae. AMB Express 6:117. https://doi.org/10.1186/s13568-016-0292-y
Miao Y, Xiong G, Li R et al (2018) Transcriptome profiling of Issatchenkia orientalis under ethanol stress. AMB Express 8:39. https://doi.org/10.1186/s13568-018-0568-5
Abdelnur PV, Caldana C, Martins MCM (2014) Metabolomics applied in bioenergy. Chem Biol Technol Agric 1:22. https://doi.org/10.1186/s40538-014-0022-0
Haris S, Fang C, Bastidas-Oyanedel J et al (2018) Natural antibacterial agents from arid-region pretreated lignocellulosic biomasses and extracts for the control of lactic acid bacteria in yeast fermentation. AMB Express 8:127. https://doi.org/10.1186/s13568-018-0654-8
Cheng C, Zhang M, Xue C, Bai F, Zhao X (2017) Development of stress tolerant Saccharomyces cerevisiae strains by metabolic engineering: new aspects from cell flocculation and zinc supplementation. J Biosci Bioeng 123(2):141–146
Sanda T, Hasunum T, Matsuda F, Kondo A (2011) Repeated-batch fermentation of lignocellulosic hydrolysate to ethanol using a hybrid Saccharomyces cerevisiae strain metabolically engineered for tolerance to acetic and formic acids. Bioresour Technol 102(17):7917–7924
Takabatake A, Kawazoe N, Izawa S (2015) Plasma membrane proteins Yro2 and Mrh1 are required for acetic acid tolerance in Saccharomyces cerevisiae. Appl Microbiol Biotechnol 99:2805–2814. https://doi.org/10.1007/s00253-014-6278-2
Matsui K, Kuroda K, Ueda M (2009) Creation of a novel peptide endowing yeasts with acid tolerance using yeast cell-surface engineering. Appl Microbiol Biotechnol 82:105–113. https://doi.org/10.1007/s00253-008-1761-2//
Kim I, Kim H, Kim Y et al (2013) Expression of dehydrin gene from Arctic Cerastium arcticum increases abiotic stress tolerance and enhances the fermentation capacity of a genetically engineered Saccharomyces cerevisiae laboratory strain. Appl Microbiol Biotechnol 97:8997–9009. https://doi.org/10.1007/s00253-013-4729-9
Shui Z, Qin H, Wu B et al (2015) Adaptive laboratory evolution of ethanologenic Zymomonas mobilis strain tolerant to furfural and acetic acid inhibitors. Appl Microbiol Biotechnol 99:5739–5748. https://doi.org/10.1007/s00253-015-6616-z
Bubis JA, Spasskaya DS, Gorshkov VA et al (2020) Rpn4 and proteasome-mediated yeast resistance to ethanol includes regulation of autophagy. Appl Microbiol Biotechnol. https://doi.org/10.1007/s00253-020-10518-x
Larsson S, Nilvebrant N, Jönsson L (2001) Effect of overexpression of Saccharomyces cerevisiae Pad1p on the resistance to phenylacrylic acids and lignocellulose hydrolysates under aerobic and oxygen-limited conditions. Appl Microbiol Biotechnol 57:167–174. https://doi.org/10.1007/s002530100742
Inaba T, Watanabe D, Yoshiyama Y et al (2013) An organic acid-tolerant HAA1-overexpression mutant of an industrial bioethanol strain of Saccharomyces cerevisiae and its application to the production of bioethanol from sugarcane molasses. AMB Express 3:74. https://doi.org/10.1186/2191-0855-3-74
Mohanram S, Amat D, Choudhary J et al (2013) Novel perspectives for evolving enzyme cocktails for lignocellulose hydrolysis in biorefineries. Sustain Chem Process 1:15. https://doi.org/10.1186/2043-7129-1-15
Wang H, Hart DJ, An Y (2019) Functional metagenomic technologies for the discovery of novel enzymes for biomass degradation and biofuel production. Bioenerg Res 12:457–470. https://doi.org/10.1007/s12155-019-10005-w
Zhao J, Wang G, Chu J et al (2020) Harnessing microbial metabolomics for industrial applications. World J Microbiol Biotechnol 36:1. https://doi.org/10.1007/s11274-019-2775-x
Sagt CM (2013) Systems metabolic engineering in an industrial setting. Appl Microbiol Biotechnol 97(6):2319–2326. https://doi.org/10.1007/s00253-013-4738-8
Vemuri GN, Aristidou AA (2005) Metabolic engineering in the -omics era: elucidating and modulating regulatory networks. Microbiol Mol Biol Rev 69(2):197–216. https://doi.org/10.1128/MMBR.69.2.197-216.2005
Zhang P, Chen Q, Fu G et al (2019) Regulation and metabolic engineering strategies for permeases of Saccharomyces cerevisiae. World J Microbiol Biotechnol 35:112. https://doi.org/10.1007/s11274-019-2684-z
Narayanan V, Sànchez i Nogué V, van Niel EWJ et al (2016) Adaptation to low pH and lignocellulosic inhibitors resulting in ethanolic fermentation and growth of Saccharomyces cerevisiae. AMB Express 6:59. https://doi.org/10.1186/s13568-016-0234-8
El-Rotail AAMM, Zhang L, Li Y et al (2017) A novel constructed SPT15 mutagenesis library of Saccharomyces cerevisiae by using gTME technique for enhanced ethanol production. AMB Express 7:111. https://doi.org/10.1186/s13568-017-0400-7
Bilal M et al (2018) Metabolic engineering and enzyme-mediated processing: a biotechnological venture towards biofuel production – A review. Renew Sustain Energy Rev 82(Part 1):436–447
Moreno AD, Carbone A, Pavone R, Olsson L, Geijer C (2019) Evolutionary engineered Candida intermedia exhibits improved xylose utilization and robustness to lignocellulose-derived inhibitors and ethanol. Appl Microbiol Biotechnol 103:1405–1416
Kaku T, Kaida R, Baba K et al (2011) Improvement of fermentable sugar yields of mangium through transgenic overexpression of xyloglucanase. J Wood Sci 57:545–548. https://doi.org/10.1007/s10086-011-1180-3
Clomburg JM, Gonzalez R (2010) Biofuel production in Escherichia coli: the role of metabolic engineering and synthetic biology. Appl Microbiol Biotechnol 86:419–434. https://doi.org/10.1007/s00253-010-2446-1
Asgher M, Shahid M, Kamal S, Iqbal MNH (2014) Recent trends and valorization of immobilization strategies and ligninolytic enzymes by industrial biotechnology. J Mol Catal B, Enzym 101:56–66
Patrick T et al (2019) Application of nanoparticles in biofuels: an overview. Fuel 237:380–397
Majidian P, Tabatabaei M, Zeinalabedini M, Naghshbandi MN (2017) Metabolic engineering of microorganisms for biofuel production. Renew Sust Energ Rev 82:3863–3885
Stovicek V, Borja GM, Forster J, Borodina I (2015) EasyClone 2.0: expanded toolkit of integrative vectors for stable gene expression in industrial Saccharomyces cerevisiae strains. J Ind Microbiol Biotechnol 42(11):1519–1531. https://doi.org/10.1007/s10295-015-1684-8
Weber C, Farwick A, Benisch F et al (2010) Trends and challenges in the microbial production of lignocellulosic bioalcohol fuels. Appl Microbiol Biotechnol 87:1303–1315. https://doi.org/10.1007/s00253-010-2707-z
Liu J, Ding W, Zhang G et al (2011) Improving ethanol fermentation performance of Saccharomyces cerevisiae in very high-gravity fermentation through chemical mutagenesis and meiotic recombination. Appl Microbiol Biotechnol 91:1239–1246. https://doi.org/10.1007/s00253-011-3404-2
Zheng D, Chen J, Zhang K et al (2014) Genomic structural variations contribute to trait improvement during whole-genome shuffling of yeast. Appl Microbiol Biotechnol 98:3059–3070. https://doi.org/10.1007/s00253-013-5423-7
Cunha JT, Romaní A, Costa CE et al (2019) Molecular and physiological basis of Saccharomyces cerevisiae tolerance to adverse lignocellulose-based process conditions. Appl Microbiol Biotechnol 103:159–175. https://doi.org/10.1007/s00253-018-9478-3
Davison SA, den Haan R, van Zyl WH (2016) Heterologous expression of cellulase genes in natural Saccharomyces cerevisiae strains. Appl Microbiol Biotechnol 100:8241–8254. https://doi.org/10.1007/s00253-016-7735-x/////
Li H, Shen Y, Wu M et al (2016) Engineering a wild-type diploid Saccharomyces cerevisiae strain for second-generation bioethanol production. Bioresour Bioprocess 3:51. https://doi.org/10.1186/s40643-016-0126-4
Oke MA, Suffian M, Annuar M, Simarani K (2016) Mixed feedstock approach to lignocellulosic ethanol production—prospects and limitations. Bioenerg Res 9:1189–1203
Swamy KBS, Zhou N (2019) Experimental evolution: its principles and applications in developing stress-tolerant yeasts. Appl Microbiol Biotechnol 103:2067–2077. https://doi.org/10.1007/s00253-019-09616-2
Chandel AK, Singh OV (2011) Weedy lignocellulosic feedstock and microbial metabolic engineering: advancing the generation of ‘biofuel’. Appl Microbiol Biotechnol 89:1289–1303. https://doi.org/10.1007/s00253-010-3057-6
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Verma, N., Kumar, V. Microbial conversion of waste biomass into bioethanol: current challenges and future prospects. Biomass Conv. Bioref. 13, 6419–6456 (2023). https://doi.org/10.1007/s13399-021-01824-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13399-021-01824-z