Abstract
It is of utmost importance to construct industrial xylose-fermenting Saccharomyces cerevisiae strains for lignocellulosic bioethanol production. In this study, two xylose isomerase-based industrial S. cerevisiae strains, O7 and P5, were constructed by δ-integration of the xylose isomerase (XI) gene xylA from the fungus Orpinomyces sp. and from the bacterium Prevotella ruminicola, respectively. The xylose consumption of the strains O7 and P5 at 48-h fermentation was 17.71 and 26.10 g/L, respectively, in synthetic medium with xylose as the sole sugar source. Adaptive evolution further improved the xylose fermentation capacity of the two strains to 51.0 and 28.9% in average, respectively. The transcriptomes of these two strains before and after evolution were analyzed using RNA-Seq. The expression levels of the genes involved in cell integrity, non-optimal sugar utilization, and stress response to environment were significantly up-regulated after evolution and did not depend on the origin of xylA; the expression levels of the genes involved in transmembrane transport, rRNA processing, cytoplasmic translation, and other processes were down-regulated. The expression of genes involved in central carbon metabolism was fine-tuned after the evolution. The analysis of transcription factors (TFs) indicated that most of the genes with significant differential expression were regulated by the TFs related to cell division, DNA damage response, or non-optimal carbon source utilization. The results of this study could provide valuable references for the construction of efficient xylose-fermenting XI strains.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Saccharomyces cerevisiae is regarded as particularly suited for bioethanol production owing to its high efficiency of hexose sugar fermentation, excellent tolerance to ethanol and lignocellulosic pretreatment inhibitors, ability of high-density fermentations (Buijs et al. 2013; Mabee and Saddler 2010; Matsushika et al. 2009; Zhou et al. 2014), as well as resistance to bacterial contamination as it can ferment at relative low pH (Matsushika et al. 2009). However, native S. cerevisiae is unable to ferment xylose, the second most abundant sugar present in lignocellulosic hydrolysate.
Two different xylose metabolic pathways have been successfully introduced into S. cerevisiae by heterologous expression: the xylose reductase (XR) and xylitol dehydrogenase (XDH) (XR-XDH) pathway and the xylose isomerase (XI) pathway (Kim et al. 2013; Matsushika et al. 2009). The hindrance of the XR-XDH pathway is redox imbalance because XR catalyzes the reduction of xylose to xylitol using preferably NADPH, whereas XDH catalyzes the oxidation of xylitol to xylulose strictly using NAD+, leading to the accumulation of xylitol and reduced ethanol yields (Li et al. 2016a). The XI pathway has received wide attention during the recent years, because xylose is directly converted to xylulose by XI without a coenzyme and no intermediate is produced (Diao et al. 2013). However, in view of recent findings, although the ethanol yield of XI strains was reported as high as 0.46 g/g consumed xylose in synthetic medium (Demeke et al. 2013), the xylose consumption rate is still lower than that of the XR-XDH strains, especially for xylose and glucose co-fermentation, in which xylose fermentation cannot start before the glucose is almost completely consumed (Vilela et al. 2015). In lignocellulosic hydrolysates, the xylose consumption was even worse (Ko et al. 2016). Therefore, rational engineering strategies, such as improving the xylose transport, promoting the conversion of xylose to xylulose, and increasing the flux of the pentose phosphate pathway (PPP), have been used to improve the xylose fermentation capacity of XI strains (Demeke et al. 2013; Diao et al. 2013; Lee et al. 2014; Vilela et al. 2015). Evolutionary engineering is another effective strategy to improve global cell metabolism and improve the xylose fermentation efficiency of XI strains (Demeke et al. 2013; Diao et al. 2013; Hector et al. 2013; Lee et al. 2014; Madhavan et al. 2009; Smith et al. 2014; Vilela et al. 2015; Zhou et al. 2012). However, the mechanism of adaptive evolution for XI strains is not well understood. Although there are some recent studies focusing on the regulation of genes involved in central carbon metabolism during evolution (Qi et al. 2015; Vilela et al. 2015), there are no reports focusing on whole genome transcriptional changes of XI strains before and after evolution.
In heterologous expression of xylose isomerase, selecting a suitable host strain is extremely important to ensure a high XI activity in S. cerevisiae. Most of the studies reported used laboratory yeast strains as hosts for the expression of XI. However, it is considered that pathways optimized in laboratory strains may not be applicable in industrial strains owing to the different metabolic backgrounds (Feng and Zhao 2013a). Additionally, many studies used plasmids to express the heterologous protein, which is unstable because of the loss or alteration of the plasmid during strain growth or fermentation (Diao et al. 2013; Zhang et al. 1996). The construction of a stable industrial strain with high xylose fermentation efficiency is of utmost importance for lignocellulosic bioethanol production.
In a previous study, the haploid S. cerevisiae strain YC-8 was generated from the industrial strain NAPX37 (Li et al. 2016a). It served as the host strain for the XI expression. Two xylA genes, one from the fungus Orpinomyces sp. and the other from the bacterium Prevotella ruminicola, with or without codon optimization, were expressed in YC-8 through multi-copy plasmid expression. After evolution, strains introduced with the codon-optimized xylA genes showed higher XI activities and better xylose-fermenting capacities (Li et al. 2016b). In the present study, these two codon-optimized genes were integrated into the genome of YC-8 by using a δ-integration method. The resulted strains, O7 and P5, showed much higher XI activities and xylose fermentation rates than the strains with plasmid expression of XI (Li et al. 2016b). After evolution, the xylose fermentation rates of the strains O7 and P5 were further improved. The genetic basis underlying the evolved phenotype was investigated by comparing the global gene expressions before and after evolution. Transcription factors (TFs) of the differentially expressed genes were analyzed to reveal the underlying mechanism of the evolution. The aim of this study is to provide references for the construction of an efficient xylose-fermenting XI strain.
Materials and methods
Strains and culture conditions
All S. cerevisiae strains used in the study are listed in Table S1 (supplementary material). Xylose utilization strains were routinely cultured in Yeast Peptone (YP) medium (20 g/L peptone, 10 g/L yeast extract) containing 20 g/L xylose. Strain YC-8 was cultured in YP medium containing 20 g/L glucose. Yeast Nitrogen Base (YNB) medium (1.7 g/L yeast nitrogen base without amino acids) containing 20 g/L xylose (2% YNBX) and 40 mg/L uracil was used for selection during strain construction and adaption. YP medium containing 40 g/L xylose (4% YPX) or 40 g/L xylose and 60 g/L glucose (10% YPDX) were used for batch fermentation. Escherichia coli DH5α (Takara, Dalian, China) was used for plasmid preparation and cultured in LB/Amp medium (10 g/L tryptone, 5 g/L yeast extract, 5 g/L sodium chloride, and 0.1 mg/mL ampicillin, pH 7.0). Solid media was prepared by adding 1.5% (w/v) agar.
Strain construction
The codon-optimized xylA from Orpinomyces sp. (OrxylA2) (accession no. KY630521) and P. ruminicola (PrxylA2) (accession no. KY630522) were multi-integrated into the yeast genome by using a δ-integration method (Tanino et al. 2010), and the TDH3 promoter (PTHD3) was used to control the gene expressions. The fragments PTHD3-OrxylA2-TTDH3 and PTHD3-PrxylA2-TTDH3 were amplified from the plasmids pRS426-OXYLA2 and pRS426-PXYLA2 (Supplementary Table S1), respectively, using the primer pair δ-XI-F/δ-XI-R harboring the sequences for homologous recombination (Supplementary Table S1). The fragments were inserted into the δ region of the host strain YC-8 by homologous recombination, and the transformants were checked for their growth and xylose fermentation capacities. Those with good xylose fermentation capacities were analyzed to determine the copy number of the xylA gene through quantitative polymerase chain reaction (PCR) using the primer pairs q-OXYL2-F/q-OXYL2-R and q-PXYL2-F/q-PXYL2-R for strains O7 and P5, respectively (Supplementary Table S1). The ACT1 gene was selected as the internal control to calculate the relative copy number of xylA gene (xylA copy numbers divided by ACT1 copy numbers). Two strains named O7 and P5, one with OrxylA2 and the other with PrxylA2, having similar relative copy numbers (1.17), were selected for the following adaption (Fig. 1).
Strain adaption
Strain adaption under microaerobic condition was performed at 30 °C in 300-mL flasks (with cotton plug) containing 100 mL 2% YNBX medium and mixed using a HS-6DN magnetic stirrer (AS ONE, Osaka, Japan) at a speed of 200 rpm. When the cells grew to the exponential phase, the cultures were inoculated into fresh medium with a starting OD660 of 0.05. This adaption was carried out for 60 days. Single cells were isolated from the final cultures and screened by growth and xylose fermentation capacity.
Xylose fermentation assays
Yeast cultures were pre-cultivated under aerobic conditions at 30 °C for 16 h using 5% YPD medium in a rotary shaker with a shaking speed of 160 rpm. Cells were harvested by centrifugation, washed twice with distilled water, and then inoculated into fermentation medium with an initial OD660 of 0.8 (approximately 0.2 g dry cell weight (DCW)/L). Fermentation was performed under microaerobic condition in 300-mL flasks (with cotton plug) with 100 mL of 4% YPX medium or 10% YPDX medium, at 35 °C for 48 h in a rotary shaker with an agitation speed of 120 rpm. Fermentation in 4% YPX was performed for 24 h and the xylose isomerase activities were assayed according to the method described previously (Li et al. 2016b). Bagasse hydrolysate was prepared to evaluate the xylose fermentation capacity of the strains. Bagasse was pretreated by using the dilute H2SO4 method described by Wang et al. (2015) with minor modifications: 50 g (dry weight) bagasse was mixed with 50 g of 5% (w/v) H2SO4. Tap water (450 mL) was added into the pretreatment reaction system. Pretreatment was performed at 140 °C for 10 min in a pressure-resistant glass reactor (TEM-V; Taiatsu Techno, Osaka, Japan). After pretreatment, solids and liquid were separated and the liquid, which mainly contained xylose and inhibitors, was used for fermentation. The concentrations of acetic acid, formic acid, furfural, and 5-hydroxymethylfurfural (5-HMF), typical inhibitors in the lignocellulose hydrolysate (Palmqvist and Hahn-Hägerdal 2000), were 45.6, 0.87, 0.147, and 0.027 mM, respectively. Glucose and xylose in the hydrolysate were adjusted to approximately 60 and 40 g/L, respectively. Concentrations of other sugars in the hydrolysate were low: 2.27 g/L of arabinose, 0.78 g/L of galactose, 0.13 g/L of cellobiose, 0.11 g/L of rhamnose, and 0.05 g/L of mannose. The hydrolysate was adjusted to pH 5.0 using NaOH before fermentation. The fermentation was carried out at 35 °C for 48 h. All experiments were performed in duplicate. Analysis of variance (ANOVA) was performed, and the results were expressed as mean values. A Duncan test was conducted to examine significant differences among experimental mean values (P < 0.05).
The concentration of sugars was determined by high performance liquid chromatography (HPLC) (LC-10AD VP, Shimadzu, Kyoto, Japan) equipped with a fluorescence detector (RF-10AXL) as previously described (Tang et al. 2006). Ethanol concentrations were measured by gas chromatography (GC 353B, GL Sciences, Tokyo, Japan) with a flame ionization detector (FID) and isopropanol was used as the internal standard (Tang et al. 2006). Xylitol and glycerol were assayed by HPLC (SCL-10A VP, Shimadzu, Kyoto, Japan) equipped with an AMINEX HPX-87H column (300 × 7.8 mm) (Bio-Rad, Hercules, USA) and a RID-10A refractive index detector (Shimadzu, Kyoto, Japan) as previously described (Li et al. 2014). Furfural, 5-HMF, acetic acid, and formic acid concentrations were determined using HPLC (LC-10AD VP, Shimadzu, Kyoto, Japan) (Wang et al. 2015).
RNA preparation and sequencing
Total RNA was extracted at 24 h of the fermentation process in 4% YPX using the Yeast RNAiso Kit (TaKaRa, Dalian, China). Yeast cells were harvested by centrifugation at 8000×g and 4 °C for 2 min. RNA concentration was measured using Qubit® RNA Assay Kit in Qubit® 2.0 Fluorimeter (Life Technologies, CA, USA). RNA purity was checked using the NanoPhotometer® spectrophotometer (IMPLEN, CA, USA). RNA integrity was assessed using the RNA Nano 6000 Assay Kit of the Agilent Bioanalyzer 2100 system (Agilent Technologies, CA, USA). Each RNA sample used for sequencing was the mixture of two biological replicates.
RNA-Seq libraries were constructed and sequenced at Sangon Biotech (Shanghai, China). Briefly, a total amount of 3 μg RNA per sample was used as an input material for the RNA sample preparations. Sequencing libraries were generated using the NEBNext® Ultra™ RNA Library Prep Kit for Illumina® (NEB, MA, USA) following the manufacturer’s recommendations and index codes were added to assign unique sequences to each sample. Briefly, mRNA was purified from total RNA using poly-T oligo-attached magnetic beads. Fragmentation was carried out using divalent cations under elevated temperature in NEBNext First Strand Synthesis Reaction Buffer (5X). First strand cDNA was synthesized using random hexamer primers and M-MuLV Reverse Transcriptase. Second strand cDNA synthesis was subsequently performed using DNA polymerase I and RNase H. Remaining overhangs were converted into blunt ends via exonuclease/polymerase activities. After adenylation of the 3′ ends of the DNA fragments, NEB Next Adaptors with hairpin loop structure were ligated to prepare them for hybridization. The cDNA library fragments were purified with AMPure XP system (Beckman Coulter, Beverly, USA) to select fragments of 150~200 bp in length. A volume of 3 μL USER Enzyme (NEB, MA, USA) was mixed with the size-selected, adaptor-ligated cDNA at 37 °C for 15 min followed by 5 min at 95 °C before PCR. The PCR was performed using the Phusion High-Fidelity DNA polymerase, Universal PCR primers, and the Index (X) Primer. Finally, PCR products were purified (AMPure XP system) and library quality was assessed with the Agilent Bioanalyzer 2100 system. The cDNA libraries were sequenced using the Illumina Hiseq 2500 platform. The raw sequence data of O7, O7E15, O7E49, P5, P5E22, P5E35, P5E47, and P5E49 can be obtained through the NCBI accession numbers SRR5251643, SRR5251642, SRR5251641, SRR5251640, SRR5251639, SRR5251638, SRR5251637, and SRR5251636, respectively.
Transcriptome analysis
Clean raw data were obtained by removing the reads containing adapter, poly-N, and low quality (Zhao et al. 2015). The sequence files in FASTQ format were analyzed using the Phred software (http://www.phred.org). Reads Per Kilobase per Million (RPKM) was used for a relative assessment of gene expression levels. Differentially expressed genes (DEGs) were identified by the DESeq R package (1.10.1), the assembled transcripts between control and experimental group were compared using Cuffdiff, with cutoff P value set as 0.05, and the absolute value of log2(fold change) ≥ 1 was set as the threshold for the significance of differential expression (Feng and Zhao 2013b). The gene ontology (GO) analysis was performed by using generic GO term mapper developed by Princeton University (http://go.princeton.edu/cgi-bin/GOTermMapper). TFs of the genes with significant differential expression were searched in the YEASTRACT database (http://www.yeastract.com/), and only transcription regulators with direct or indirect documented evidence were considered (Feng and Zhao 2013b).
Results
Strain construction and evolution
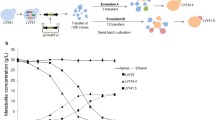
For efficient expression of xylose isomerase in industrial S. cerevisiae, the codon-optimized xylA (Li et al. 2016b) from Orpinomyces sp. (OrxylA2) and P. ruminicola (PrxylA2) were inserted by using a δ-integration method. The strains expressing different xylA with the best growth characteristics and approximately the same xylA copy numbers were selected to achieve commensurate catabolic levels. The selected strains, expressing OrxylA2 and PrxylA2, were named O7 and P5, respectively (Fig. 1).
The strains O7 and P5 were subjected to evolutionary engineering using repeated batch cultivation. After 30 transfers (60 days totally), the cell growth and the xylose consumption rates of both strains became stable and consistent. Mutant strains were isolated using 2% YPX plates. The growth and fermentation capacities of these isolated strains were compared using 2% YPX medium and 4% YPX medium, respectively. The two best mutant strains, O7E15 and O7E49, originating from strain O7, and the four best mutant strains, P5E22, P5E35, P5E47, and P5E49, originating from strain P5, were selected for further study.
Xylose fermentation of original and evolved strains
The xylose fermentation results of O7 and its evolved strains O7E15 and O7E49, which expressed codon-optimized xylA from Orpinomyces sp., are shown in supplementary Fig. S1 (A) and Table 1. As no ethanol was produced in the absence of sugar, the ethanol detected was completely produced from xylose. After 24-h fermentation, xylose consumption of the evolved strains was higher than that of the original strain O7 (Supplementary Fig. S1 (A) and Table 1). The XI activity of strain O7 was 1.372 IU/mg-protein, and evolution improved XI activity of 1.08- and 1.03-fold for strains O7E15 and O7E49, respectively (Fig. 2). As the fermentation continued, the two evolved strains showed faster xylose consumption rates than the original strain. After 48 h of fermentation, strain O7 consumed 17.71 g/L xylose, which was significantly lower than the evolved strains O7E15 (29.66 g/L) and O7E49 (23.83 g/L). Concomitantly, the ethanol production of the evolved strains O7E15 (12.99 g/L) and O7E49 (9.63 g/L) was higher than of the original strain O7 (7.01 g/L) (Supplementary Fig. S1 (A) and Table 1). Ethanol yields of strains O7E15 and O7E49 were 0.438 and 0.404 g/g consumed xylose, respectively, which are 1.11- and 1.02-fold higher than the original strain O7 (Table 1).
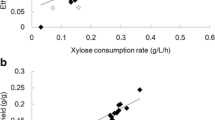
The xylose consumption and cell growth of the strain P5, which expressed the xylA gene from P. ruminicola, and its evolved strains (Supplementary Fig. S1 (B) and Table 1) were similar to those of strain O7 and its evolved strains. After 24 h of fermentation, all four evolved strains consumed more xylose than the original strain P5. XI activity of the strain P5 was 0.841 IU/mg-protein, and evolution improved XI activity 1.14-, 1.01-, 1.32, and 1.33-fold for strains P5E22, P5E35, P5E47, and P5E49, respectively. After 48 h of fermentation, strain P5 consumed 26.10 g/L xylose and produced 10.45 g/L ethanol, which were significantly higher than that of the strain O7. Four evolved strains, P5E22, P5E35, P5E47, and P5E49, consumed 35.24, 32.55, 33.71, and 33.11 g/L xylose, respectively, and the xylose consumption rates were 1.35-, 1.25-, 1.29-, and 1.27-fold higher than that of the original strain P5. The ethanol yield of strains P5E22, P5E47, and P5E49 was 0.428, 0.436, and 0.438 g/g consumed xylose, respectively, which were 1.07-, 1.09-, and 1.10-fold higher than that of the original strain P5. However, the ethanol yield of the evolved strain P5E35 was a little lower than that of strain P5 (Table 1). XI activities of strains P5E22, P5E47, and P5E49, but not strain P5E35, were higher than that of the original strain P5. The correlation between XI activity and xylose consumption rate was weak (Fig. 2 and Table 1), which indicated that some other factors may contribute to the improved xylose-fermenting capacity.
There were differences in cell growth, xylose consumption, and ethanol production among the strains O7 and P5 and their evolved strains. The correlation between the cell growth (OD660) and ethanol production (ethanol concentration) was not significant (Supplementary Fig. S2 (A)). However, there was an interrelationship between the cell yield and ethanol yield; lower cell yield resulted in higher ethanol yield (Supplementary Fig. S2 (B) and Table 1).
Glucose and xylose co-fermentation
Strain O7 and its evolved strains all consumed 49.61 g/L glucose in the first 8 h of fermentation (Supplementary Fig. S3 and Table 2). For strain O7, only 4.36 g/L xylose was consumed in the first 8-h fermentation. About 19.74 g/L xylose was consumed after 48-h fermentation and about 29.93 g/L ethanol was produced with 0.432 g/g of ethanol yield based on the total consumed sugar. The xylose fermentation capacity of the evolved strains O7E15 and O7E49 was better than that of strain O7. In the first 8 h of fermentation, strains O7E15 and O7E49 consumed 6.25 and 6.59 g/L xylose, respectively, and after 48-h fermentation, each of the strains consumed more than 27 g/L xylose, which were significantly higher than that of strain O7. The ethanol production rate of the evolved strains O7E15 and O7E49 was 1.12- and 1.05-fold higher than that of the original strain. However, during the 48-h fermentation, the ethanol yield of the evolved strain O7E49 was lower than that of the original strain O7. The main reason was that the higher ratio of consumed xylose was directed to cell growth in the strain O7E49 compared to that in the original strain O7. The cell yield based on xylose consumption of O7E49 was 0.370, which was nearly 2.4-fold higher than those of strains O7 and O7E15 (Table 2).
Strain P5 and its evolved strains all consumed 49.22 g/L glucose in the first 8 h of fermentation (Supplementary Fig. S4 and Table 2). The strain P5 consumed only approximately 2 g/L xylose during the first 8 h of fermentation. After 48-h fermentation, the strain P5 consumed 19.39 g/L xylose and the ethanol yield based on total consumed sugar was 0.411 g/g. Xylose consumption of the evolved strains P5E22, P5E35, P5E47, and P5E49 ranged from 21.56 to 26.33 g/L, which was 1.22-, 1.26-, 1.11-, and 1.35-fold higher than that of the original strain P5, respectively. The ethanol yields based on the total consumed sugar of all the evolved strains were greater than that of strain P5 (Table 2). The strain P5 presented an ethanol yield of only 0.191 g/g-xylose (Table 2) when it fermented xylose during the co-fermentation (YPDX medium). This yield was lower than the ethanol yield of the same strain when fermenting xylose as the sole sugar (YPX medium) (Table 1). Not only the xylose consumption rate but also the ethanol yield from xylose of strain P5 was severely repressed by glucose during co-fermentation, which was different from that of the other four evolved strains (Tables 1 and 2).
Lignocellulosic hydrolysate fermentation
The xylose fermentation performance of the evolved strains O7E15 and P5E49 was better than that of the other evolved strains; therefore, they and their original strains O7 and P5 were evaluated using bagasse hydrolysate (Supplementary Fig. S5 and Table 3). During the first 6-h fermentation, all the strains assimilated 57.27 g/L glucose but limited xylose. After 48-fermentation, the evolved strains O7E15 and P5E49 consumed 28.30 and 23.18 g/L xylose, respectively, which were 1.37- and 1.05-fold higher than those of the original strains O7 and P5, respectively. In total, 35.57 and 33.89 g/L ethanol was produced by the strains O7E15 and P5E49, respectively. The ethanol yields based on the total consumed sugar were 0.411 and 0.412 g/g, respectively, which were slightly higher than the original strains (Table 3).
Transcriptional differences of the strains before and after adaptive evolution
Total RNA samples were extracted after the 24-h fermentation when xylose was used as the sole sugar source. The RNA-Seq method was used to analyze global gene expression and to explore the molecular mechanism of strain evolution in xylose medium.
Compared to the original strain O7, a total of 63 genes showed significantly different expression levels (fold change ≥ 2) in both the evolved strains O7E15 and O7E49, of which 21 genes were up-regulated and 42 genes were down-regulated (Fig. 3a and Supplementary Tables S2 and S3). An analysis of GO identified that the up-regulated genes were involved mostly in cell assembly, such as meiotic cell cycle, cell wall organization or biogenesis, and mitochondrion organization (Fig. 4a), as well as substance transport and metabolism, such as transmembrane transport, cellular amino acid metabolic process, and carbohydrate metabolic process. The emphasis on these biological processes might explain the improved cell growth and fermentation ability of the evolved strains in xylose medium. As shown in Fig. 4a, down-regulated genes were involved mainly in the biological process of transmembrane transport, rRNA processing, ribosomal large subunit biogenesis, ribosomal small subunit biogenesis, and more. Some genes involved in the cellular amino acid metabolic process, such as ACO2, ARO2, and MIS1, were down-regulated. Previous studies indicated that the loss of function of ACO2 and MIS1 could increase the ethanol yield from glucose (Rossouw et al. 2013) and improve cell growth of S. cerevisiae in non-fermenting sugar (Shannon and Rabinowitz 1988), respectively.
For strain P5, four evolved strains, P5E22, P5E35, P5E47, and P5E49, were selected for transcriptome comparison. Genes with significant differential expression (fold change ≥ 2) in at least three strains were analyzed. In total, 33 genes were significantly up-regulated and 14 genes were down-regulated (Fig. 3b and Supplementary Tables S4 and S5). GO analysis showed that most of the significantly up-regulated genes were involved in cell growth (such as chromatin organization, cytoskeleton organization, cell wall organization or biogenesis, and peptidyl-amino acid modification), cell self-repair (DNA repair and cellular response to DNA damage stimulus), and intracellular metabolism (such as nucleobase-containing small molecule metabolic process, carbohydrate metabolic process, monocarboxylic acid metabolic process). Most of the significantly down-regulated genes were involved in cellular translation (such as cytoplasmic translation, mRNA processing, and rRNA processing), transmembrane transport, and lipid metabolic process.
Table 4 summarizes the changes of genes involved in the xylose metabolic pathway before and after evolution. Most of the genes involved in xylose metabolic pathways were down-regulated after evolution, but the changes were not significant. Among the genes for sugar transportation, HXT5 and GAL2, which may be involved in xylose transportation (Hamacher et al. 2002), had the same regulation trends in the strains O7 and P5 during evolution, and both were up-regulated but not significantly (< 2-fold). Compared to the original strain O7, expressions of PFK1, ENO1, and PYK2 were up-regulated in the evolved strains (O7E15 and O7E49). Compared to the original strain P5, expressions of TKL1, PFK1, FBA1, and PYK2 were up-regulated in all the evolved strains (P5E22, P5E35, P5E47, and P5E49), but only the changed expression level of FBA1 was significant (> 2-fold). Two genes involved in glycolysis, PFK1 and PYK2, were up-regulated (< 2-fold) in all the evolved strains of O7 and P5.
Although strains O7 and P5 were engineered from the same original strain using the same chromosome integration method, and the evolutionary processes were synchronous, gene expression in the evolved strains was different. Only ENA1, SLZ1, and PEX18 were significantly (> 2-fold) up-regulated, and RPL23A and RPS22A were significantly (> 2-fold) down-regulated in all evolved strains of O7 and P5.
Transcription factor analysis
Analyses of TFs were conducted to unravel the regulation mechanism of gene expression changes. Nine TFs regulated more than 60% of the significantly up-regulated genes and 11 TFs controlled more than 60% of the significantly down-regulated genes in evolved strains of O7 (Fig. 5a). Eight TFs regulated more than 60% of the significantly up-regulated genes and 10 TFs controlled more than 60% of the significantly down-regulated genes in evolved strains of P5 (Fig. 5b). Notably, regardless of the origin of xylA genes, Sfp1p and Ace2p were most the important TFs regulating the majority of the up- and down-regulated genes (Fig. 5). As shown in Supplementary Table S6, Sfp1p and Ace2p are involved in cell division and Sfp1p is also involved in nutrient assimilation, stress, and DNA damage response. Tec1p and Ste12p regulated more than 70% of the up-regulated genes but regulated less than 60% of down-regulated genes in the evolved strains of O7 and P5. Tec1p is involved in Ty1-mediated gene activation, and Ste12p plays a role in signaling pathways in carbon metabolism (Supplementary Table S6); therefore, they might be important regulators for improved xylose metabolism. Cst6p, which is involved in the utilization of non-optimal carbon sources and chromosome stability, regulated more than 60% of the down-regulated genes in the evolved strains of O7 and P5. Sin3p, which plays a role in the maintenance of chromosomal integrity, controlled more than 60% of the down-regulated genes in the evolved strains of P5. The TFs active in gene regulation might indicate that evolutionary engineering improved the cellular metabolism, cell integrity, and stress response ability of the evolved strains and thereby enhanced the cell growth and fermentation in xylose medium.
Discussion
In the present study, two codon-optimized xylA genes, one from the fungus Orpinomyces sp. and the other from the bacterium P. ruminicola, with the strong promoter P TDH3 were integrated into the industrial S. cerevisiae host strain YC-8 through δ-integration, and the strains O7 and P5 were constructed, respectively. Evolutionary engineering further improved the xylose fermentation capacity of the strains O7 and P5. This capacity of the evolved strains of O7 and P5 was higher than those of the laboratory strains reported in many previous reports (Vilela et al. 2015; Lee et al. 2014; Madhavan et al. 2009; Zhou et al. 2012; Hector et al. 2013; Qi et al. 2015). Although the ethanol production rates were still slower than those of the industrial strains reported in recent studies (Diao et al. 2013; Demeke et al. 2013), fermentation in the present study utilized much lower inoculum size and haploid strains with unregulated PPP pathway.
Different original xylA genes resulted in different xylose fermentation capacities, as suggested by the fermentation results of strains P5 and O7 (Tables 1, 2, and 3). A notable difference was found when fermenting YPX with xylose as sole sugar source, and the strain P5 had 47.3% higher xylose consumption rate than strain O7 (Table 1). However, the difference in xylose consumption became less obvious when fermenting mixed sugar, mainly due to the stronger repression on xylose consumption of the strain P5 by coexisting glucose (Tables 2 and 3). These results suggested that the origin of xylA genes affected not only the xylose consumption but also the response to glucose repression.
A different evolutionary effect was found among strains evolved from strains O7 and P5. All of the evolved strains studied here showed an improved xylose consumption rate; however, some of them maintained the same ethanol yields as the original strains and others showed higher ethanol yields than the original strains (Tables 1 and 2). The increase of ethanol yield was due to the reduced cell yield after evolution (Supplementary Fig. S2 (B)), which suggests the adjustment of carbon metabolism from cell growth to ethanol production during evolution.
The molecular mechanisms underlying the improved phenotype as a result of evolution were investigated at the genome-wide transcriptional level. The results showed that the transcription of genes involved in central carbon metabolism did not change significantly after evolution. While the non-fermentable carbon source-inducible transporter gene HXT5 and the broad substrate sugar transporters GAL2 were up-regulated, the heterologous xylA gene and XKS1 encoding for xylulokinase were down-regulated, in all evolved strains from O7 and P5 (Table 4). The down-regulation of xylA did not reduce XI activity. Contrarily, XI activities of all the evolved strains were improved compared to that of the original strains (Fig. 2). Additionally, compared to the original strains, most of the genes in the PPP and glycolysis pathway of the evolved strains were down-regulated (Table 4). In the glycolysis pathway, PGK1 and PYK2 showed differential regulation after evolution. PGK1 and PYK2, encoding the 3-phosphoglycerate kinase and pyruvate kinase, respectively, are involved in ATP production (Larsson et al. 2000). The expression change of these genes indicated that the xylose metabolic pathway had been improved for better balance during the evolutionary process, similar to the results of the previous study in which evolutionary engineering fine-tuned gene expression in the xylose pathway in XI strains (Qi et al. 2015).
In the evolved strains of O7, significantly up-regulated genes were mainly involved in meiotic cell cycle, transmembrane transport, sporulation, carbohydrate metabolic process, and many more (Supplementary Table S2). Among these genes, VPS13 regulates phosphatidylinositol-4-phosphate (PtdIns(4)P) levels in the prospore membrane, and loss of VPS13 could lead to morphological defects of the membrane (Park et al. 2015). SLZ1 can be induced in extremely nutrient-poor conditions (Agarwala et al. 2012). The up-regulation of this gene was mainly due to the minimal medium (YNBX) used during adaption evolution. The genes ENA1, MIM2, PDR5, PEX18, and STL1 are involved in transmembrane transport. ENA1 encodes P-type ATPases, its expression is low under standard growth conditions (Nakayama et al. 2004), but rapidly increased after exposure to saline, osmotic, or alkaline stresses (Nakayama et al. 2004; Platara et al. 2006). STL1 encodes the glycerol/H+ symporter, which plays an essential role for signaling (Ferreira and Lucas 2007). MIM2 is a novel central player in the biogenesis of mitochondrial outer membrane proteins, the defects of Mim2 would lead to slow cell growth (Dimmer et al. 2012). PDR5 encodes for the yeast ATP-binding cassette transporter, which is repressed during growth on non-fermentable carbon sources in S. cerevisiae (Mamnun et al. 2004). The overexpression of these genes could explain why the growth and fermentation of the evolved strains in xylose medium were improved. Significantly down-regulated genes in the evolved strains of O7 were mainly involved in the bioprocess of rRNA processing, ribosomal small/large subunit biogenesis, cellular amino acid metabolic process, and many more (Supplementary Table S3). The significant down-regulation of these genes was beneficial to improve the growth and xylose fermentation. For example, ODC2 encodes mitochondrial transport proteins and the high-copy number of ODC2 would suppress cell growth (Tibbetts et al. 2002). ACO2 encodes for the aconitase isoforms 2 and its deletions would lead to increasing ethanol yields from glucose and fructose as sources (Rossouw et al. 2013). MIS1 encodes for the mitochondrial C1-THF synthase. Shannon and Rabinowitz (1988) reported that S. cerevisiae strains lacking a functional MIS1 gene were viable and could grow in medium containing a non-fermentable carbon source.
Most of the up-regulated genes in the evolved strains of P5 also benefit cell growth and fermentation in xylose. The overexpression of the ACS1 gene, which encodes acetyl-coenzyme A synthase, could promote the metabolism of the tricarboxylic acid (TCA) cycle (Kratzer and Schüller 1995). The overexpression of SPS100, YPK2, MPS3, and MYO1 could promote cell assembly during growth and reproduction (Destruelle et al. 1994; Law and Segall 1988; Jaspersen et al. 2002; Tolliday et al. 2003). SPS100 could promote spore wall maturation (Destruelle et al. 1994). The putative protein kinase gene YPK2 (with YPK1) is essential to cell growth (Law and Segall 1988). Some genes involved in cell stress resistance were also up-regulated after evolution (Supplementary Table S4). BDF1 is involved in salt resistance (Fu et al. 2013); GDB1 plays a role in salt and thermo resistance (Teste et al. 2000), and ENA1 is involved in osmotic stress resistance. More importantly, the genes RPH1 and SNF2, which are needed for DNA damage repair and Ty transcription (Kim et al. 2002; Happel et al. 1991), respectively, were also up-regulated after evolution. The up-regulation of these genes indicated that the multi-copies of xylA inserted into the δ region might result in chromosome structural instability of the strain, though there is no report on this to date. Significantly down-regulated genes in the evolved strains of P5 were mainly involved in transmembrane transport, cytoplasmic translation, mRNA/rRNA processing, and more (Supplementary Table S5). For example, RPL30 is involved in mRNA and rRNA processing (Macías et al. 2008). ERG28 plays a role in ergosterol biosynthesis; its down-regulation might negatively affect the tolerance of the strains to vanillin, a typical inhibitor of the lignocellulose hydrolysate (Palmqvist and Hahn-Hägerdal 2000).
As shown in Supplementary Table S2~S5, different sets of genes were differentially expressed in the O7 and P5 evolutionary strains, compared to their respective original strains. Because the xylA integration and strain evolution of O7 and P5 were synchronized, it could be deduced that the difference in the gene regulations was at least partly caused by the different origin of the xylA genes. However, three genes, ENA1, SLZ1, and PEX18, were significantly up-regulated in all the evolved strains of O7 and P5. ENA1 and PEX18 are involved in transmembrane transport; SLZ1 is involved in sporulation (Supplementary Tables S2 and S4). Besides, two genes, RPL23A and RPS22A which encode ribosomal protein of the large subunit and small subunit, respectively, were significantly down-regulated in all the evolved strains of O7 and P5 (Supplementary Tables S3 and S5). There are no reports to date about the interrelationship of the functions of these five genes and the xylose utilization. As they were enriched in all the evolved strains, it is worthy to further study these genes.
No matter the origin of xylA genes expressed, the bioprocesses of cell assembly, cell integrity, non-optimal sugar utilization, and stress response to the environment were improved in all the evolved strains of O7 and P5. As xylose is not a fermentable sugar for S. cerevisiae and it is recognized by S. cerevisiae as a stress factor (Zeng et al. 2016), it is reasonable that the up-regulation of non-optimal sugar utilization and stress response would benefit the xylose fermentation in the evolved strains. Heterologous protein expression might lead to metabolic burden (van Rensburg et al. 2012), and the integration position in the chromosome, gene copy number, and promoter strength is closely connected with the phenotype of the strains (Flagfeldt et al. 2009). From the transcriptomic results of the present study, the heterologous overexpression of the xylA gene in the δ region might result in metabolic burden and chromosome instability. The up-regulation of bioprocesses of cell assembly and cell integrity might contribute partly to the improved xylose fermentation of the evolved strains. The results of TF analysis supported the above speculation to some extent. The TFs Sfp1p and Ace2p, which are related to cell division and DNA damage response (Supplementary Table S2), regulated more than 84% of the genes with significant differential expression in the evolved strains of O7 and P5, respectively (Fig. 5). Besides Sfp1p and Ace2p, Tec1p and Ste12p regulated most of the up-regulated genes in evolved strains. The two TFs cooperate with each other to regulate genes specific for haploid invasive growth (Supplementary Table S6). Notably, Ste12p was reported to be involved in signaling pathways of carbon metabolism in S. cerevisiae (Feng and Zhao 2013a). A recent study suggested that the S. cerevisiae strains lack sensing and signaling of xylose (Brink et al. 2016). Gcn4p, which regulated nearly 80% of the up-regulated and down-regulated genes in the evolved strains of O7 and P5, is related to amino acid biosynthesis. Feng and Zhao (2013a) reported that Sfp1p, Ste12p, and Gcn4p were very important for xylose metabolism regulations in XR-XDH S. cerevisiae strains. It could be suggested that these TFs also played a critical role in the evolved XI strains in the present study, though their contributions to the xylose fermentation need further investigation.
In conclusion, the XI strains O7 and P5, which expressed xylA originating from Orpinomyces sp. and P. ruminicola, respectively, showed good xylose fermentation capacity. Evolutionary engineering was effective for the improvement of xylose fermentation capacity of the both strains. After evolution, the expression of genes involved in central carbon metabolism was fine-tuned. Gene ontology and transcription factors analysis indicated that genes with significant differential expression were mainly involved in chromosome stability, non-optimal sugar utilization, stress response to environment, and more. These results could provide references for constructing efficient xylose-fermenting XI strains for lignocellulosic bioethanol production.
References
Agarwala SD, Blitzblau HG, Hochwagen A, Fink GR (2012) RNA methylation by the MIS complex regulates a cell fate decision in yeast. PLoS Genet 8(6):e1002732
Brink DP, Borgström C, Tueros FG, Gorwa-Grauslund MF (2016) Real-time monitoring of the sugar sensing in Saccharomyces cerevisiae indicates endogenous mechanisms for xylose signaling. Microb Cell Factories 15:183
Buijs NA, Siewers V, Nielsen J (2013) Advanced biofuel production by the yeast Saccharomyces cerevisiae. Curr Opin Chem Biol 17(3):480–488
Demeke MM, Dietz H, Li Y, Foulquié-Moreno MR, Mutturi S, Deprez S, Abt TD, Bonini BM, Lidén G, Dumortier F (2013) Development of a D-xylose fermenting and inhibitor tolerant industrial Saccharomyces cerevisiae strain with high performance in lignocellulose hydrolysates using metabolic and evolutionary engineering. Biotechnol Biofuel 6(5):89–89
Destruelle M, Holzer H, Klionsky DJ (1994) Identification and characterization of a novel yeast gene: the YGP1 gene product is a highly glycosylated secreted protein that is synthesized in response to nutrient limitation. Mol Cell Biol 14(4):2740–2754
Diao L, Liu Y, Qian F, Yang J, Yu J, Sheng Y (2013) Construction of fast xylose-fermenting yeast based on industrial ethanol-producing diploid Saccharomyces cerevisiae by rational design and adaptive evolution. BMC Biotechnol 13(1):1–9
Dimmer KS, Papić D, Schumann B, Sperl D, Krumpe K, Walther DM, Rapaport D (2012) A crucial role for Mim2 in the biogenesis of mitochondrial outer membrane proteins. J Cell Sci 125(Pt 14):3464–3473
Feng X, Zhao H (2013a) Investigating host dependence of xylose utilization in recombinant Saccharomyces cerevisiae strains using RNA-seq analysis. Biotechnol Biofuels 6(1):96
Feng X, Zhao H (2013b) Investigating glucose and xylose metabolism in Saccharomyces cerevisiae and Scheffersomyces stipitis via 13C metabolic flux analysis. AICHE J 59(9):3195–3202
Ferreira C, Lucas C (2007) Glucose repression over Saccharomyces cerevisiae glycerol/H+ symporter gene STL1 is overcome by high temperature. FEBS Lett 581(9):1923–1927
Flagfeldt DB, Siewers V, Huang L, Nielsen J (2009) Characterization of chromosomal integration sites for heterologous gene expression in Saccharomyces cerevisiae. Yeast 26(10):545–551
Fu J, Hou J, Liu L, Chen L, Wang M, Shen Y, Zhang Z, Bao X (2013) Interplay between BDF1 and BDF2 and their roles in regulating the yeast salt stress response. FEBS J 280(9):1991–2001
Happel AM, Swanson MS, Winston F (1991) The SNF2, SNF5 and SNF6 genes are required for ty transcription in Saccharomyces cerevisiae. Genetics 128(1):69–77
Hamacher T, Becker J, Gardonyi M, Hahn-Hägerdal B, Boles E (2002) Characterization of the xylose-transporting properties of yeast hexose transporters and their influence on xylose utilization. Microbiol-SGM 148:2783–2788
Hector RE, Dien BS, Cotta MA, Mertens JA (2013) Growth and fermentation of D-xylose by Saccharomyces cerevisiae expressing a novel D-xylose isomerase originating from the bacterium Prevotella ruminicola TC2-24. Biotechnol Biofuels 6(1):1–12
Jaspersen SL, Giddings TH, Mark W (2002) Mps3p is a novel component of the yeast spindle pole body that interacts with the yeast centrin homologue Cdc31p. J Cell Biol 159(6):945–956
Kim EM, Jang YK, Park SD (2002) Phosphorylation of Rph1, a damage-responsive repressor of PHR1 in Saccharomyces cerevisiae, is dependent upon Rad53 kinase. Nucleic Acids Res 30(3):643–648
Kim SR, Park YC, Jin YS, Seo JH (2013) Strain engineering of Saccharomyces cerevisiae for enhanced xylose metabolism. Biotechnol Adv 31(6):851–861
Ko JK, Um Y, Woo HM, Kim KH, Lee SM (2016) Ethanol production from lignocellulosic hydrolysates using engineered Saccharomyces cerevisiae harboring xylose isomerase-based pathway. Bioresour Technol 209:290–296
Kratzer S, Schüller HJ (1995) Carbon source-dependent regulation of the acetyl-coenzyme a synthetase-encoding gene ACS1 from Saccharomyces cerevisiae. Gene 161(1):75–79
Larsson C, Påhlman IL, Gustafsson L (2000) The importance of ATP as a regulator of glycolytic flux in Saccharomyces cerevisiae. Yeast 16(9):797–809
Law DT, Segall J (1988) The SPS100 gene of Saccharomyces cerevisiae is activated late in the sporulation process and contributes to spore wall maturation. Mol Cell Biol 8(2):912–922
Lee SM, Jellison T, Alper HS (2014) Systematic and evolutionary engineering of a xylose isomerase-based pathway in Saccharomyces cerevisiae for efficient conversion yields. Biotechnol Biofuels 7(1):122
Li YC, Gou ZX, Liu ZS, Tang YQ, Akamatsu T, Kida K (2014) Synergistic effects of TAL1 over-expression and PHO13 deletion on the weak acid inhibition of xylose fermentation by industrial Saccharomyces cerevisiae strain. Biotechnol Lett 36(10):2011–2021
Li YC, Mitsumasu K, Gou ZX, Min G, Tang YQ, Li GY, Wu XL, Akamatsu T, Taguchi H, Kida K (2016a) Xylose fermentation efficiency and inhibitor tolerance of the recombinant industrial Saccharomyces cerevisiae strain NAPX37. Appl Microbiol Biotechnol 100(3):1531–1542
Li YC, Li GY, Gou M, Xia ZY, Tang YQ, Kida K (2016b) Functional expression of xylose isomerase in flocculating industrial Saccharomyces cerevisiae strain for bioethanol production. J Biosci Bioeng 121(6):685–691
Mabee W, Saddler J (2010) Bioethanol from lignocellulosics: status and perspectives in Canada. Bioresour Technol 101(13):4806–4813
Macías S, Bragulat M, Tardiff DF, Vilardell J (2008) L30 binds the nascent RPL30 transcript to repress U2 snRNP recruitment. Mol Cell 30(6):732–742
Madhavan A, Tamalampudi S, Ushida K, Kanai D, Katahira S, Srivastava A, Fukuda H, Bisaria VS, Kondo A (2009) Xylose isomerase from polycentric fungus Orpinomyces: gene sequencing, cloning, and expression in Saccharomyces cerevisiae for bioconversion of xylose to ethanol. Appl Microbiol Biotechnol 82(6):1067–1078
Mamnun YM, Schüller C, Kuchler K (2004) Expression regulation of the yeast PDR5 ATP-binding cassette (ABC) transporter suggests a role in cellular detoxification during the exponential growth phase. FEBS Lett 559(1–3):111–117
Matsushika A, Inoue H, Kodaki T, Sawayama S (2009) Ethanol production from xylose in engineered Saccharomyces cerevisiae strains: current state and perspectives. Appl Microbiol Biotechnol 84(1):37–53
Nakayama H, Yoshida K, Shinmyo A (2004) Yeast plasma membrane Ena1p ATPase alters alkali-cation homeostasis and confers increased salt tolerance in tobacco cultured cells. Biotechnol Bioeng 85(7):776–789
Palmqvist E, Hahn-Hägerdal B (2000) Fermentation of lignocellulosic hydrolysates. II: inhibitors and mechanisms of inhibition. Bioresour Technol 74(1):25–33
Park JS, Halegoua S, Kishida S, Neiman AM (2015) A conserved function in phosphatidylinositol metabolism for mammalian Vps13 family proteins. PLoS One 10(4):e0124836
Platara M, Ruiz A, Serrano R, Palomino A, Moreno F, Ariño J (2006) The transcriptional response of the yeast Na+-ATPase ENA1 gene to alkaline stress involves three main signaling pathways. J Biol Chem 281(48):36632–36642
Qi X, Zha J, Liu GG, Zhang W, Li BZ, Yuan YJ (2015) Heterologous xylose isomerase pathway and evolutionary engineering improve xylose utilization in Saccharomyces cerevisiae. Front Microbiol 6:1165
Rossouw D, Heyns EH, Setati ME, Bosch S, Bauer FF (2013) Adjustment of trehalose metabolism in wine Saccharomyces cerevisiae strains to modify ethanol yields. Appl Environ Microbiol 79(17):5197–5207
Shannon KW, Rabinowitz JC (1988) Isolation and characterization of the Saccharomyces cerevisiae MIS1 gene encoding mitochondrial C1-tetrahydrofolate synthase. J Biol Chem 263(16):7717–7725
Smith J, Van Rensburg E, Görgens JF (2014) Simultaneously improving xylose fermentation and tolerance to lignocellulosic inhibitors through evolutionary engineering of recombinant Saccharomyces cerevisiae harbouring xylose isomerase. BMC Biotechnol 14(1):1–17
Tang YQ, An M, Liu K, Nagai S, Shigematsu T, Morimura S, Kida K (2006) Ethanol production from acid hydrolysate of wood biomass using the flocculating yeast Saccharomyces cerevisiae strain KF-7. Process Biochem 41(4):909–914
Tanino T, Hotta A, Ito T, Ishii J, Yamada R, Hasunuma T, Ogino C, Ohmura N, Ohshima T, Kondo A (2010) Construction of a xylose-metabolizing yeast by genome integration of xylose isomerase gene and investigation of the effect of xylitol on fermentation. Appl Microbiol Biotechnol 88(5):1215–1221
Teste MA, Enjalbert B, Parrou JL, François JM (2000) The Saccharomyces cerevisiae YPR184w gene encodes the glycogen debranching enzyme. FEMS Microbiol Lett 193(1):105–110
Tibbetts AS, Sun Y, Lyon NA, Ghrist AC, Trotter PJ (2002) Yeast mitochondrial oxodicarboxylate transporters are important for growth on oleic acid. Arch Biochem Biophys 406(1):96–104
Tolliday N, Pitcher M, Li R (2003) Direct evidence for a critical role of myosin II in budding yeast cytokinesis and the evolvability of new cytokinetic mechanisms in the absence of myosin II. Mol Biol Cell 14(2):798–809
van Rensburg E, den Haan R, Smith J, van Zyl WH, Görgens JF (2012) The metabolic burden of cellulase expression by recombinant Saccharomyces cerevisiae Y294 in aerobic batch culture. Appl Microbiol Biotechnol 96(1):197–209
Vilela LD, de Araujo VPG, Paredes RD, Bon EPD, Torres FAG, Neves BC, Eleutherio ECA (2015) Enhanced xylose fermentation and ethanol production by engineered Saccharomyces cerevisiae strain. AMB Express 5(1):16
Wang G, Tan L, Sun ZY, Gou ZX, Tang YQ, Kida K (2015) Production of bioethanol from rice straw by simultaneous saccharification and fermentation of whole pretreated slurry using Saccharomyces cerevisiae KF-7. Environ Prog Sustain Energy 34(2):582–588
Zeng W-Y, Tang Y-Q, Gou M, Xia Z-Y, Kida K (2016) Transcriptomes of a xylose-utilizing industrial flocculating Saccharomyces cerevisiae strain cultured in media containing different sugar sources. AMB Express 6:51
Zhang Z, Moo-Young M, Chisti Y (1996) Plasmid stability in recombinant Saccharomyces cerevisiae. Biotechnol Adv 14(4):401–435
Zhao Q, Pan L, Ren Q, Hu D (2015) Digital gene expression analysis in hemocytes of the white shrimp Litopenaeus vannamei in response to low salinity stress. Fish Shellfish Immun 42(2):400–407
Zhou H, Cheng JS, Wang BL, Fink GR, Stephanopoulos G (2012) Xylose isomerase overexpression along with engineering of the pentose phosphate pathway and evolutionary engineering enable rapid xylose utilization and ethanol production by Saccharomyces cerevisiae. Metab Eng 14(6):611–622
Zhou YJ, Buijs NA, Siewers V, Nielsen J (2014) Fatty acid-derived biofuels and chemicals production in Saccharomyces cerevisiae. Front Bioeng Biotechnol 2:32
Acknowledgements
This work was supported by the National Natural Science Foundation of China (31170093) and the Talent Project for Science and Technology Innovation of Sichuan Province (2017RZ0021).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have declared no conflict of interest, and this article does not contain any studies with human participants or animals performed by any of the authors.
Electronic supplementary material
ESM 1
(PDF 1328 kb).
Rights and permissions
About this article
Cite this article
Li, YC., Zeng, WY., Gou, M. et al. Transcriptome changes in adaptive evolution of xylose-fermenting industrial Saccharomyces cerevisiae strains with δ-integration of different xylA genes. Appl Microbiol Biotechnol 101, 7741–7753 (2017). https://doi.org/10.1007/s00253-017-8494-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-017-8494-z