Abstract
We describe a procedure by which filter paper is digested with a cellulolytic enzyme preparation, obtained from Trichoderma reesei cultivated under solid state fermentation conditions and then fermented by a recombinant Saccharomyces cerevisiae strain. The yeast strain produces a β-glucosidase encoded by the BGL1 gene from Saccharomycopsis fibuligera that quantitatively and qualitatively complements the limitations that the Trichoderma enzyme complex shows for this particular activity. The supplemental β-glucosidase activity fuels the progression of cellulose hydrolysis and fermentation by decreasing the inhibitory effects caused by the accumulation of cellobiose and glucose. Fermentation of filter paper by this procedure yields ethanol concentrations above 70 g/L.
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cellulose, because of its chemical composition, abundance in nature and the fact that it does not have a direct use as food, is the primary substrate for second generation bioethanol. Standard, industrially implemented processes, to produce ethanol from cellulolytic substrates, comprise an enzymatic step, in which commercial enzymes are used to render fermentable sugar (i.e. glucose) and a fermentative step carried out by the yeast Saccharomyces cerevisiae. In a widely disseminated but oversimplified vision, enzymatic hydrolysis of cellulose is considered to be carried out by the combined action of endo and exo glucanases, that yield cellobiose and glucose as major products, and β-glucosidase, which splits cellobiose into glucose. However, cellulose digestion by highly efficient cellulolytic microorganisms is a considerably more complex process. In addition to the action of different type of enzymes that act coordinately, there are also non-catalytic proteins that play essential roles. For instance, in bacterial cellulosomes the association of catalytic (cellulolytic enzymes) and non-catalytic (scaffoldins) proteins assure an arrangement of the enzymatic machinery that maximizes the efficiency of the cellulolytic process (Artzi et al. 2017). Recent studies with the fungus Trichoderma reesei (Hypocrea jecorina), from which most commercial cellulolytic enzyme preparations are derived, show intricate complexities in both composition and function of the cellulolytic machinery (Lehmann et al. 2016; Jalak et al. 2012). Proteomic analysis reveals the importance of non-catalytic, accessory enzymes, such as cellulose induced proteins and swollenin, in cellulolysis (Gupta et al. 2016; Herpoël-Gimbert et al. 2008).
The industrial production of ethanol from cellulosic substrates makes use of high amounts of enzymes, required for yielding fermentable sugar (glucose). Cellulose digestion and fermentation can be carried out in a single step in simultaneous saccharification and fermentation processes or sequentially, as separate hydrolysis and fermentation steps (Paulova et al. 2015). An approach proposed to eliminate the cost burden of enzymes has been the construction of engineered S. cerevisiae strains that co-express various genes coding for cellulolytic enzymes (Davison et al. 2016; Liu et al. 2016; Matano et al. 2012; Yamada et al. 2011; Wen et al. 2010; Tsai et al. 2009; van Zyl et al. 2007; Fujita et al. 2004). However, because of the aforementioned inherent complexity of enzymatic hydrolysis of cellulose, this approach has clear limitations and in consequence the cellulose degrading capability of the engineered yeast strains is not as efficient as the enzyme complexes produced by Trichoderma or other fungi. Additionally concomitant forced expression of multiple genes can overstress the physiology of the host yeast.
The aim of this work was to set up a cost-effective procedure for ethanol production from cellulose. This has been carried out by coupling hydrolysis using crude Trichoderma enzymes and fermentation with a selected yeast strain. It is known that Trichoderma cellulolytic complex is deficient in β-glucosidase activity (Lehmann et al. 2016; Sørensen et al. 2013; Nieves et al. 1998). Structural and functional features of the GH1 and GH3 β-glucosidases responsible for the hydrolysis of cellobiose have been characterized in detail (Sanz-Aparicio et al. 1998; Pozzo et al. 2010). Based on this information, we choose a S. cerevisiae strain that expresses extracellular β-glucosidase from Saccharomycopsis fibuligera, and has been shown to ferment cellobiose with high efficiency (Tang et al. 2013; Gurgu et al. 2011).
Materials and methods
Microbial strains and culture media
Trichoderma reesei (H. jecorina) CECT2415 (identical to ATCC 56764 and NRRL 11236) was obtained from Colección Española de Cultivos Tipo (http://www.uv.es/cect). Recombinant S. cerevisiae T500 that expresses a β-glucosidase gene from S. fibuligera and its parental strain BY4741 have been described (Gurgu et al. 2011; Marín-Navarro et al. 2011).
Culture media used were: YPD: 1% yeast extract, 2% peptone, 2% glucose; YP2P and YP5P: same composition as YPD but with 2 or 5% filter paper (RM13054252, 73 g/m2, Filtros Anoia, Barcelona Spain), cut into 25 mm2 squares, instead of glucose, respectively (% refers to weight/volume).
Trichoderma cellulolytic enzyme complex preparation
Trichoderma precultures (2 mL) were prepared in YP2P medium supplemented with 0.5% starch and 0.5 g/L Tween 80, and incubated under agitation for 72 h, at 30 °C. For submerged fermentation, 25 mL of YP5P with 0.5% starch medium were inoculated with the preculture and incubated at 30 °C with aeration by vigorous shaking (200 rpm) for several days. The cultures were centrifuged and the supernatant used as enzyme source. For solid state fermentation, 5 g of filter paper squares in a 250 mL Erlenmeyer flask were soaked with 8 mL of YP supplemented with 0.5% starch, inoculated with 2 mL of preculture, and incubated at 30 °C for several days. To extract the enzymes from the solid state fermentation, the mycelium grown in the flask was thoroughly mixed with 8 mL of extracting solution (1% yeast extract, 2% peptone) and squeezed. The recovered slurry was clarified by centrifugation, filtered through a 220 nm membrane and then used as enzyme source.
The filter paper assay for cellulose hydrolysis and enzyme assays
Enzymatic digestion of filter paper was assayed by measuring reducing sugars with the dinitrosalicylic (DNS) acid method, as described by Zhang et al. (2009). Two assays, hydrolysis of the chromogenic p-nitrophenyl β-d-glucopyranoside (PNPG) substrate and hydrolysis of cellobiose, were used to monitor β-glucosidase activity. PNPG hydrolysis was assayed as described (Arrizubieta and Polaina 2000). Hydrolysis of cellobiose was carried out under the same conditions, but the activity was determined by measuring the amount of free glucose with a commercial kit (Sigma-Aldrich, St Louis MO, USA).
Production of glucose, cellobiose and other soluble sugars resulting from enzymatic digestion of filter paper by the action of Trichoderma enzymes was analyzed by ion exchange chromatography using a CarboPac PA-100 column in a HPLC instrument equipped with a pulsed amperometric detector (Dionex, Thermo Fisher Scientific, Waltham, MA, USA). Mono, di, tri and tetrasaccharides: glucose, cellobiose, cellotriose and cellotetraose (Sigma-Aldrich, St Louis MO, USA) were used as standards.
For determination of Km, dialyzed samples of β-glucosidase from either, S. cerevisiae T500 culture supernatant or T. reesei solid state fermentation enzyme preparation, were incubated with cellobiose at different concentrations in the range 0.5–20 mM, at 37 °C, pH 5.5. Glucose released in these reactions was measured by HPLC and used to calculate values of initial velocity, which were plotted versus substrate concentration. To calculate Km values, the data were adjusted to a rectangular hyperbola.
Inhibition of β-glucosidase activity by glucose was measured using as enzyme source samples of the Trichoderma enzyme cocktail or the supernatant of a S. cerevisiae T500 culture, dialyzed against citrate–phosphate buffer, pH 5.5, or phosphate buffer, pH 7.0.
Fermentations
Precultures of recombinant S. cerevisiae T500 were grown overnight at 30 °C in liquid YPD medium supplemented with the antibiotic geneticin (G418) at 100 µg/mL to select for the maintenance of the plasmid harboring the gene coding for S. fibuligera β-glucosidase. Fermentation vessels were prepared as follows. The cells (S. cerevisiae) from a 10 mL preculture were collected by centrifugation and added to the mixture of digested paper and Trichoderma enzymes obtained from the solid state fermentation, making a final volume of 10 mL. This mixture was transferred to a 14 mL tube, to which 0.5 g of filter paper squares were added. The tube was then closed, leaving a minimum headspace, and incubated at 30 °C. Successive paper loads were added to the fermentation tube. Samples were removed at different intervals. Ethanol was measured using an enzymatic kit (Thermo Scientific) and residual glucose and cellobiose by HPLC.
Results
Production of Trichoderma enzyme preparation and sugar profile resulting from its action on cellulose
We compared the strength of the cellulolytic enzyme complex recovered after growing T. reesei ATCC 56764 under two different culture conditions: Fig. 1A shows that solid state fermentation gave a cellulolytic cocktail whose activity is about ten times higher than the maximal activity obtained from the supernatant of submerged fermentation. Regardless of genetic and physiological considerations about differences in the synthesis or secretion of enzymes between submerged and solid state fermentations (Barrios-González 2012), the second procedure makes enzyme recovery easier. In addition to measuring cellulose digestion, we determined β-glucosidase activity present in the T. reesei cocktail. The profile of β-glucosidase production (Fig. 1B) was similar to that of cellulase activity. Submerged cultures showed a slow increase of β-glucosidase activity for about 10 days after the inoculation and then decreased. The activity of solid state cultures continued to increase for at least 14–16 days. On the basis of these results, cellulolytic enzymes produced by solid state fermentation were used for further experiments.
Production of cellulolytic enzymes by T. reesei under submerged fermentation (blue circles) or solid state fermentation (orange squares) conditions. A Cellulase activity, expressed in Filter Paper Units (FPU)/mL. B β-glucosidase activity. Error bars indicate standard deviation of duplicates. (Color figure online)
Hydrolysis of cellulose (5% filter paper) by the T. reesei enzyme cocktail (2 FPU/mL), after 8 h of incubation at 50 °C, yielded glucose and cellobiose as the only two significant products. (Fig. 2). The presence of cellobiose, although in lesser amount than glucose, confirms previous reports that β-glucosidase is the limiting cellulolytic activity in the extract (Lehmann et al. 2016; Nieves et al. 1998). Kinetics of cellobiose and glucose release from cellulose by the action of T. reesei enzymes was monitored for 48 h, using filter paper at different concentrations (2.5, 5 and 10%) as the substrate (Fig. 3). In all cases glucose concentration increased steadily up to different values, proportional to the initial amount of cellulose. In contrast, cellobiose concentration quickly reached a maximum, proportional to cellulose amount, which was maintained for some time, probably as a result of an equilibrium in which the velocity of cellobiose release by cellobiohydrolases matched that of hydrolysis by β-glucosidases. Cellobiose concentration at this point represented a low but significant fraction (around 10%) of the initial amount of cellulose. Since cellobiose is a potent inhibitor of cellobiohydrolases, its presence must significantly restrict cellulose hydrolysis. According to our data, digestion at 5% paper leads to an accumulation of cellobiose of over 15 mM after 20 h, much higher than the concentration of inhibitor that reduces enzyme activity by 50% (IC50) reported for T. reesei Cel7A cellobiohydrolase: 0.4–0.7 mM (Teugjas and Väljamäe 2013a). Only after prolonged incubation times cellobiose approaches complete hydrolysis.
Analysis of the soluble sugar profile. A Sugars released by the action of the enzyme cocktail (2 FPU/mL) with 5% filter paper, after 8 h at 50 °C. B Sugars present in the T. reesei enzyme cocktail obtained by solid state fermentation, before incubation with the substrate. C Commercial sugar standards used as control: (1) glucose, (2) cellobiose, (3) cellotriose, (4) cellotetraose
Time-course analysis of soluble sugars released from cellulose digestion with the T. reesei enzyme preparation. Filter paper at different concentrations (2.5%, pink squares; 5%, blue circles; 10%, green triangles) was incubated at 50 °C with the enzyme cocktail (2 FPU/mL) for different times. Reaction was stopped by heating at 95 °C for 10 min. A Cellobiose production, B glucose production. (Color figure online)
Comparison of β-glucosidase activity in the Trichoderma enzyme preparation with that produced by S. cerevisiae T500
We have analyzed and compared the properties of the β-glucosidase activity present in the Trichoderma enzyme cocktail with that secreted by the recombinant yeast, by two criteria: their performance at different concentrations of cellobiose and their inhibition by glucose. Both enzymes behaved similarly at different concentrations of cellobiose in the range 5–400 mM. Although T. reesei genome has ten gene sequences encoding β-glucosidases, only one of these enzymes, Bgl I, also termed Cel 3A, is released to the culture medium among the proteins that constitute the cellulolytic enzyme complement (Guo et al. 2016a; Karkehabadi et al. 2014). Therefore, the observed β-glucosidase activity determined for Trichoderma must reflect the action of this enzyme, for which we have determined a Km value for cellobiose of 2.07 ± 0.5 mM, not significantly different from the 1.9 mM reported in the literature (Woodward and Arnold 1981). The Km for cellobiose of the yeast enzyme was slightly lower: 1.74 ± 0.08 mM (experimental data for Km determinations are given as supplementary material, Figure S1).
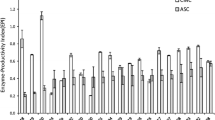
Results of the inhibitory effect of glucose presented in Fig. 4 show that at glucose concentration in the range 5–75 mM, about the same at which glucose is accumulated during the fermentation, the S. fibuligera β-glucosidase produced by recombinant S. cerevisiae T500, with PNPG as the substrate, is significantly less sensitive to inhibition by glucose than the Trichoderma β-glucosidase. The inhibitory effect of glucose when measured with cellobiose as the substrate could only be reliably assayed at low glucose concentration, since at higher concentrations the relative amount of glucose (inhibitor) and glucose (product) could not be estimated. At 1 mM, glucose inhibited cellobiose hydrolysis by both enzymes similarly, ca. 25%.
Inhibition of β-glucosidase activity by glucose. Activities of dialyzed samples of S. fibuligera (blue) and T. reesei (orange) β-glucosidases were assayed with PNPG as the substrate at pH 5.5 (solid bars) or pH 7 (dotted bars), in the presence of increasing concentrations of glucose. Value of activity without added glucose, at pH 5.5, was taken as 100%. Error bars indicate standard deviation of triplicates. (Color figure online)
Cellulose conversion into ethanol
Cellulose fermentation was carried out with filter paper digested with the T. reesei cellulolytic cocktail. Two fermenting S. cerevisiae strains were assayed: β-glucosidase producing recombinant T500 and its untransformed parental strain BY4741. As the initial 5% load of paper present in the medium was consumed, additional paper loads were added, as indicated in Fig. 5. Transient cellobiose accumulation (ca. 5 mM) was observed with the control strain about 24 h after each paper load, whereas T500 strain hydrolyzed cellobiose more efficiently. We also observed that paper degradation proceeded more rapidly with strain T500. Accordingly, ethanol yield obtained in the fermentation with T500 was significantly (35%) higher than that obtained with the control strain. These results indicate, in agreement with our previous observation, that even a small cellobiose accumulation becomes a limiting factor for both cellulolysis progression and ethanol production. Overall fermentation yields in terms of paper conversion to ethanol were 61% and 45% for T500 and the parental strain, respectively. The experimental design used here, in which we have coupled the cellulolytic machinery produced by T. reesei, with fermentation by a β-glucosidase producing S. cerevisiae strain, yields considerably higher ethanol concentrations (about 7%) than any previous report using recombinant yeast strains that express one or several cellulolytic enzymes.
Fermentation of filter paper by S. cerevisae T500, expressing the S. fibuligera BGL1 gene (orange squares), and parental strain BY4741 (blue circles), coupled to cellulose hydrolysis by T. reesei enzymes. A Ethanol, B cellobiose, and C glucose, were monitored during the fermentation. Error bars indicate standard deviation of duplicates. Additions of filter paper to the fermentation vessels (0.5 g, filled triangles; 0.25 g, open triangle) were carried out at the times indicated on the top of the figure. (Color figure online)
Discussion
The aim of this work was to work out a simple, basic, reliable and cost-effective model system for the conversion of cellulose into ethanol. In situ production of cellulolytic enzymes has obvious advantages over the addition of commercial ones. Beyond economic considerations, the composition of the enzyme complex can be tailored, monitored and manipulated at will. Even using a non-engineered T. reesei strain, the cellulolytic activity of the resulting enzyme preparation was enough to achieve an ethanol yield of about 7%, considerably higher than the minimal 4.5–5.0% threshold considered to make distillation profitable (Lennartsson et al. 2014; Viikari et al. 2012). As a first objective, we studied the conditions for in situ production of a cellulolytic enzyme complex sufficiently powerful to drive fermentation to an ethanol concentration higher than 5%. Different reports show that fungal cultivation under solid state fermentation conditions sharply increases the production yield of enzymes and other metabolites, compared to alternative submerged fermentation (Barrios-González 2012). We compared the performance of T. reesei CECT 2415 under either cultivation conditions and found that solid state fermentation gave an enzyme cocktail with the required properties for efficient cellulose digestion.
According to genome annotations, T. reesei has ten gene sequences encoding β-glucosidases, two of which belong to family GH1 and the other eight to family GH3, but only one of these enzymes, Bgl I (Cel3A), is released to the culture medium and therefore becomes part of the cellulolytic enzyme cocktail (Guo et al. 2016a). Diverse reports show that the action of this single β-glucosidase represents a bottleneck for the production of cellulosic ethanol (Lehmann et al. 2016; Sørensen et al. 2013; Nieves et al. 1998). Accordingly, cellulolytic strength of the Trichoderma enzyme cocktail was increased by supplementation with exogenous β-glucosidases (Del Pozo et al. 2012). One of the reasons that explain this effect is that cellobiose is a potent inhibitor of cellobiohydrolase, with a Ki ca. 1.5 mM (Gruno et al. 2004). As we have shown, cellobiose accumulates during the action of T. reesei enzyme cocktail (up to 15 mM on 5% paper). Therefore an increase in the amount of β-glucosidase is expected to improve cellulose digestion. Inhibition of β-glucosidase by glucose is another factor that needs to be considered since glucose accumulates during cellulolysis up to a concentration of 70 mM (Fig. 3). Different studies have addressed this problem either looking for natural variants of the enzyme less prone to glucose inhibition (Teugjas and Väljamäe 2013b), or by engineering one of T. reesei β-glucosidases (Guo et al. 2016b). Improvement of glucose inhibition has also been achieved by directed evolution of an Aspergillus niger β-glucosidase expressed in S. cerevisiae (Larue et al. 2016).
In this work we have used the enzyme encoded by the S. fibuligera BGL1 gene, produced by S. cerevisiae during the fermentation process, as a source of supplemental β-glucosidase for cellulolysis. This enzyme is expressed and secreted under control of a constitutive promoter and the STA1 gene secretion signal (Marín-Navarro et al. 2011) and, as it was shown previously, confers to S. cerevisiae the capability of utilizing cellobiose very efficiently (Gurgu et al. 2011). Our results show that with PNPG as the substrate, this enzyme is significantly more resistant to inhibition by glucose that the β-glucosidase (Cel3A) supplied by the Trichoderma enzyme cocktail (Fig. 4). Although we cannot assure that this is also true for cellobiose, comparative analysis of the three-dimensional structures of T. reesei Cel 3 A and S. fibuligera Bgl I (see Supplementary material, Figure S2) suggests that the relative affinity of glucose for its binding site would be higher for Cel 3A which would therefore be more sensitive than Bgl I to the inhibitory effect of the monosaccharide. Because of the coupled hydrolysis fermentation setup used here, the inhibitory effect exerted by high glucose concentration values attained during fermentation is counteracted by continuous production of supplemental β-glucosidase and a very active consumption of glucose by the yeast. Overall, the supply of cellobiase activity provided by the fermenting yeast represents a cheaper and easier solution than supplementation of the Trichoderma enzyme preparation with a commercial enzyme.
Conclusion
This communication describes a simple and economical procedure by which cellulose is converted into ethanol with high efficiency. Digestion of filter paper with a crude T. reesei enzyme preparation coupled to fermentation by a S. cerevisiae strain expressing β-glucosidase from S. fibuligera, is synergic for various reasons: (i) supplementary β-glucosidase activity relieves cellobiohydrolase inhibition by cellobiose improving cellulose digestion; (ii) functionally, the yeast β-glucosidase is complementary to that of Trichoderma and (iii) continual production of β-glucosidase during glucose fermentation counteracts the inhibitory effect of glucose. Consequently, this procedure allows ethanol yields higher than 7%, a level that can render distillation profitable.
References
Arrizubieta MJ, Polaina J (2000) Increased thermal resistance and modification of the catalytic properties of a β-glucosidase by random mutagenesis and in vitro recombination. J Biol Chem 275:28843–28848
Artzi L, Bayer EA, Moraïs S (2017) Cellulosomes: bacterial nanomachines for dismantling plant polysaccharides. Nat Rev Microbiol 2:83–95
Barrios-González J (2012) Solid-state fermentation: physiology of solid medium, its molecular basis and applications. Process Biochem 47:175–185
Davison SA, den Haan R, van Zyl WH (2016) Heterologous expression of cellulase genes in natural Saccharomyces cerevisiae strains. Appl Microbiol Biotechnol 100:8241–8254
Del Pozo MV, Fernández-Arrojo L, Gil-Martínez J, Montesinos A, Chernikova TN, Nechitaylo TY, Waliszek A, Tortajada M, Rojas A, Huws SA, Golyshina OV, Newbold CJ, Polaina J, Ferrer M, Golyshin PN (2012) Microbial β-glucosidases from cow rumen metagenome enhance the saccharification of lignocellulose in combination with commercial cellulase cocktail. Biotechnol Biofuels 5:73
Fujita Y, Ito J, Ueda M, Fukuda H, Kondo A (2004) Synergistic saccharification, and direct fermentation to ethanol, of amorphous cellulose by use of an engineered yeast strain codisplaying three types of cellulolytic enzyme. Appl Environ Microbiol 70:1207–1712
Gruno M, Väljamäe P, Pettersson G, Johansson G (2004) Inhibition of the Trichoderma reesei cellulases by cellobiose is strongly dependent on the nature of the substrate. Biotechnol Bioeng 86:504–511
Guo B, Sato N, Biely P, Amano Y, Nozaki K (2016a) Comparison of catalytic properties of multiple β-glucosidases of Trichoderma reesei. Appl Microbiol Biotechnol 100:4959–4968
Guo B, Amano Y, Nozaki K (2016b) Improvements in glucose sensitivity and stability of Trichoderma reesei β-glucosidase using site-directed mutagenesis. PLoS One 11(1):e0147301
Gupta V, Steindorff AS, de Paula RG, Silva-Rocha R, Mach-Aigner AR, Mach RL, Silva RN (2016) The post-genomic era of Trichoderma reesei: what’s next? Trends Biotechnol 34:970–982
Gurgu L, Lafraya A, Polaina J, Marín-Navarro J (2011) Fermentation of cellobiose to ethanol by industrial Saccharomyces strains carrying the β-glucosidase gene (BGL1) from Saccharomycopsis fibuligera. Bioresour Technol 102:5229–5236
Herpoël-Gimbert I, Margeot A, Dolla A, Jan G, Mollé D, Lignon S, Mathis H, Sigoillot JC, Monot F, Asther M (2008) Comparative secretome analyses of two Trichoderma reesei RUT-C30 and CL847 hypersecretory strains. Biotechnol Biofuels 1:18
Jalak J, Kurašin M, Teugjas H, Väljamäe P (2012) Endo-exo synergism in cellulose hydrolysis revisited. J Biol Chem 287:28802–28815
Karkehabadi S, Helmich KE, Kaper T, Hansson H, Mikkelsen NE, Gudmundsson M, Piens K, Fujdala M, Banerjee G, Scott-Craig JS, Walton JD, Phillips GNJ, Sandgren M (2014) Biochemical characterization and crystal structures of a fungal family 3 beta-glucosidase, Cel3A from Hypocrea jecorina. J Biol Chem 289:31624–31637
Larue K, Melgar M, Martin VJ (2016) Directed evolution of a fungal β-glucosidase in Saccharomyces cerevisiae. Biotechnol Biofuels 9:52
Lehmann L, Rønnest NP, Jørgensen CI, Olsson L, Stocks SM, Jørgensen HS, Hobley T (2016) Linking hydrolysis performance to Trichoderma reesei cellulolytic enzyme profile. Biotechnol Bioeng 113:1001–1010
Lennartsson PR, Erlandsson P, Taherzadeh MJ (2014) Integration of the first and second generation bioethanol processes and the importance of by-products. Bioresour Technol 165:3–8
Liu Z, Ho SH, Sasaki K, den Haan R, Inokuma K, Ogino C, van Zyl W, Hasunuma T, Kondo A (2016) Engineering of a novel cellulose-adherent cellulolytic Saccharomyces cerevisiae for cellulosic biofuel production. Sci Rep 6:24550
Marín-Navarro J, Gurgu L, Alamar S, Polaina J (2011) Structural and functional analysis of hybrid enzymes generated by domain shuffling between Saccharomyces cerevisiae (var. diastaticus) Sta1 glucoamylase and Saccharomycopsis fibuligera Bgl1 β-glucosidase. Appl Microbiol Biotechnol 89:121–130
Matano Y, Hasunuma T, Kondo A (2012) Display of cellulases on the cell surface of Saccharomyces cerevisiae for high yield ethanol production from high-solid lignocellulosic biomass. Bioresour Technol 108:128–133
Nieves RA, Ehrman CI, Adney WS, Elander RT, Himmel ME (1998) Survey and analysis of commercial cellulase preparations suitable for biomass conversion to ethanol. World J Microbiol Biotechnol 14:301–304
Paulova L, Patakova P, Branska B, Rychtera M, Melzoch K (2015) Lignocellulosic ethanol: technology design and its impact on process efficiency. Biotechnol Adv 33:1091–1107
Pozzo T, Pasten JL, Karlsson EN, Logan DT (2010) Structural and functional analyses of β-glucosidase 3B from Thermotoga neapolitana: a thermostable three-domain representative of glycoside hydrolase 3. J Mol Biol 397:724–739
Sanz-Aparicio J, Hermoso JA, Martinez-Ripoll M, Lequerica JL, Polaina J (1998) Crystal structure of β-glucosidase A from Bacillus polymyxa: insigths into the catalytic activity of family 1 glycosyl hydrolases. J Mol Biol 275:491–502
Sørensen A, Lübeck M, Lübeck PS, Ahring BK (2013) Fungal beta-glucosidases: a bottleneck in industrial use of lignocellulosic materials. Biomolecules 3:612–631
Tang H, Hou J, Shen Y, Xu L, Yang H, Fang X, Bao X (2013) High β-glucosidase secretion in Saccharomyces cerevisiae improves the efficiency of cellulase hydrolysis and ethanol production in simultaneous saccharification and fermentation. J Microbiol Biotechnol 23:1577–1585
Teugjas T, Väljamäe P (2013a) Product inhibition of cellulases studied with 14 C-labeled cellulose substrates. Biotechnol Biofuels 6:104
Teugjas H, Väljamäe P (2013b) Selecting β-glucosidases to support cellulases in cellulose saccharification. Biotechnol Biofuels 6:105
Tsai SL, Oh J, Singh S, Chen R, Chen W (2009) Functional assembly of minicellulosomes on the Saccharomyces cerevisiae cell surface for cellulose hydrolysis and ethanol production. Appl Environ Microbiol 75:6087–6093
van Zyl WH, Lynd LR, den Haan R, McBride JE (2007) Consolidated bioprocessing for bioethanol production using Saccharomyces cerevisiae. Adv Biochem Eng Biotechnol 108:205–235
Viikari L, Vehmaanperä J, Koivula A (2012) Lignocellulosic ethanol: from science to industry. Biomass Bioenerg 46:13–24
Wen F, Sun J, Zhao H (2010) Yeast surface display of trifunctional minicellulosomes for simultaneous saccharification and fermentation of cellulose to ethanol. Appl Environ Microbiol 76:1251–1260
Woodward J, Arnold SL (1981) The Inhibition of β-Glucosidase activity in Trichoderma reesei C30 cellulase by derivatives and isomers of glucose. Biotechnol Bioeng 23:1553–1562
Yamada R, Taniguchi N, Tanaka T, Ogino C, Fukuda H, Kondo A (2011) Direct ethanol production from cellulosic materials using a diploid strain of Saccharomyces cerevisiae with optimized cellulase expression. Biotechnol Biofuels 4:8
Zhang YH, Hong J, Ye X (2009) Cellulase assays. Methods Mol Biol 581:213–231
Acknowledgements
This work was funded by Grants EU H2020–634486-INMARE, and BIO2013-48779-C4-3-R from Spain’s ‘Secretaría de Estado de Investigación, Desarrollo e Innovación’. MC-V was supported by a SENESCYT predoctoral fellowship from the Government of República del Ecuador. We thank Lola Abolafio for technical assistance and Simon Gough for critical reading of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Casa-Villegas, M., Marín-Navarro, J. & Polaina, J. Synergies in coupled hydrolysis and fermentation of cellulose using a Trichoderma reesei enzyme preparation and a recombinant Saccharomyces cerevisiae strain. World J Microbiol Biotechnol 33, 140 (2017). https://doi.org/10.1007/s11274-017-2308-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-017-2308-4