Abstract
Background and Objective
The In Vivo Mechanistic Static Model (IMSM) is a powerful method used to predict the magnitude of drug–drug interactions (DDIs) mediated by cytochromes. The objective of this study was to extend the IMSM paradigm to DDIs mediated by efflux transporters and cytochromes.
Methods
First, a generic model for this kind of interaction was devised. A flexible approach was then developed to estimate the characteristic parameters [the contribution ratios (CRs) and inhibition or induction potencies (IXs)] from clinical data by non-linear regression. Next, this approach was applied to the DDIs mediated by P-glycoprotein (P-gp) and cytochrome P450 (CYP) 3A4/3A5 in a large set of victim drugs and interactors. Lastly, the model and associated parameters were used to identify the DDIs most at risk of overexposure.
Results
A total of 25 substrates and 26 interactors (three inducers, 23 inhibitors) could be considered in the regression analysis. The number of observations [area under the plasma concentration–time curve ratios or renal clearance ratios (Robs)] was 138. Fifty CRs and 57 IXs were estimated. The proportions of predictions within 0.67- to 1.5-fold Robs and within 0.5- to 2-fold Robs were 79% and 93% for the internal validation and 76% and 88% for the external validation, respectively. The median fold error was 0.98 (the ideal value is 1) and the interquartile range of the fold error was 0.36. The relative standard error of parameter estimates was a maximum of 15%.
Conclusions
The IMSM approach was successfully extended to DDIs mediated by P-gp and CYP3A4/3A5. The method revealed good predictive performances by internal and external validation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Pharmacokinetic drug–drug interactions mediated by membrane transporters and cytochromes may result in large variations of drug exposure. |
A new approach is presented to predict the variation of exposure due to interactions mediated by P-glycoprotein and cytochrome P450 3A4/3A5. |
This approach was applied to a large set of substrates and inhibitors or inducers. The predictive performances were suitable for clinical applications. |
1 Introduction
Pharmacokinetic drug–drug interactions (DDIs) are well-known to be a major factor of inter-individual variability in drug exposure [1]. As a result, drug agencies have issued guidelines to evaluate the potential DDIs during drug development [2, 3]. The strongest interactions are mediated by cytochromes and membrane transporters. Much effort has been put into devising quantitative methods to predict the impact of DDIs on drug exposure in order to reduce the experimental burden, duration, and cost of drug development. Currently, the favored approach relies on in vitro–in vivo extrapolation, using a physiologically based pharmacokinetic (PBPK) model [4, 5]. This approach is mainly used for metabolic, cytochrome-mediated DDIs. However, the importance of DDIs mediated by membrane transporters is now widely recognized [6]. These interactions may involve influx transporters such as organic anion transporter polypeptides (OATPs) [7,8,9] and efflux transporters such as P-glycoprotein [P-gp, also known as multidrug resistance 1 (MDR1) coded by the ABCB1 gene] [10] or breast cancer resistance protein (BCRP, also known as ABCG2) [11], among others.
The PBPK approach allows complete description of the pharmacokinetics of the interaction, but this remains complex, time-consuming, and requires determination of the empirical scaling factors for application to clinical data [12, 13]. As a result, the PBPK approach is mainly used in the context of drug development. Besides this approach, when estimation of the variation of drug exposure in plasma at steady state is the major goal (such as in clinical settings), a static model is an attractive approach because of its simplicity. Static models for transporter-mediated DDIs have previously been described [14,15,16]. All of these approaches rely on some in vitro–in vivo extrapolations to estimate the parameters of the model.
Another approach to static models for predicting DDIs has been developed, which is based solely on clinical data. For this reason, the method is called the In vivo Mechanistic Static Model (IMSM). One feature of the IMSM is that the victim drugs and interactors are characterized by dimensionless parameters [the contribution ratios (CRs) and the inhibition or induction potencies (IXs)] that are easy to interpret. This approach was introduced by Ohno et al. [17, 18] to predict cytochrome P450 (CYP) 3A4-mediated DDIs. We extended the method to other cytochromes [19,20,21,22], the combined impact of DDIs and cytochrome polymorphism [23, 24], and, recently, the prediction of the inter-individual variability of DDIs [25]. The predictions made using the IMSM have proven to be accurate in a large dataset of more than 600 DDIs [26] and also for DDIs involving inhibitory metabolites [27].
In this study, the IMSM paradigm was extended to describe DDIs mediated by efflux transporters and cytochromes. First, a generic model for this kind of interaction was devised. A flexible approach was then developed to estimate the characteristic parameters (the CRs and IXs) from clinical data. Next, this new approach was applied to the DDIs mediated by P-gp and CYP3A4 in a large set of victim drugs and interactors. Lastly, the model and associated parameters were used to identify the DDIs with high risk of overexposure.
2 Methods
2.1 Generic Model
The goal of our model is to describe the impact of DDIs mediated by a single efflux membrane transporter and one or several cytochromes quantitatively. The victim drug, which is a substrate of the transporter and possibly of the cytochromes, is assumed to be administered by the oral route. The interactor, inhibitor, and/or inducer of the transporter, and possibly of the cytochromes, is also given by the oral route. The impact of the interaction is mainly characterized by the variation in exposure [area under the plasma concentration–time curve (AUC)] of the substrate in plasma. In this study, we do not attempt to model the variation of exposure in tissues (e.g., in brain).
The transporter is assumed to be present in the gut wall, liver (on the apical canalicular membrane), and kidney (on the apical membrane), resulting in drug secretion in the gut lumen, bile, and urine. The cytochromes are assumed to be located in the gut wall and liver. For simplicity here, we consider a single cytochrome (CYP3A4/A5) in the equations, but extension of the model to drugs metabolized by several cytochromes is straightforward.
A semi-physiologic, compartmental model is used to describe the pharmacokinetics of the victim drug. The two parts of the model are shown in Fig. 1a, b.
a, b Representation of the semi-physiologic compartmental model of drug absorption (a) and clearance (b). CLiEH enterohepatic cycle with an intrinsic clearance, CLg clearance of the gut, CLih-cyp clearance of the drug in the liver by cytochromes, CLih-other clearance of the drug in the liver by other metabolism, CLr renal clearance, CLg-cyp clearance of the gut due to cytochromes, CLg-other clearance of the gut due to other metabolism, CLg-pgp clearance of the gut due to P-glycoprotein, FaFg fraction of drug that escapes from the gut, fub unbound drug concentration in blood, Qg blood flow rate in gut blood, Qh blood flow rate in liver blood, PSg permeability surface product of the gut, PSh permeability surface product of hepatocytes, R dosing rate
The first part is the absorption model, which is composed of two compartments: the lumen of the gastrointestinal tract combined with the intracellular space of the enterocytes and the gut blood. The second part is the clearance (CL) model, which is composed of four compartments: the liver blood, the intracellular space of hepatocytes, the bile compartment, and the blood of the systemic circulation. To derive the model, we assumed that the victim drug is given as a continuous zero-order input at rate R (e.g., in mg/h) in the lumen and that the system is at steady state. However, the final equations do not rely on this assumption.
The drug in compartment 1 may disappear with an apparent clearance of the gut (CLg), which is due to CYP3A4 metabolism (CLg-cyp), elimination by the lumen flow of the drug that has been effluxed from the enterocyte to the lumen and not reabsorbed (CLg-pgp), and elimination by the lumen flow before absorption (CLg-other). The drug in compartment 1 is transferred by passive diffusion to the gut blood with a gut permeability surface product PSg (which has a clearance dimension). The concentration of drug in gut blood is C2, and the blood flow rate is Qg. The output rate of the drug from compartment 2 is Qg.C2, which is by definition equal to FaFg.R, where FaFg is the fraction of drug that escapes from the gut.
In the liver blood, part of the drug is transported into the systemic circulation with a blood flow rate Qh, while the other part is eliminated by secretion into the bile or metabolism. The level of permeability of drug through the hepatocyte membrane is assumed to be high and there is no active transport of drug between liver blood and hepatocyte intracellular water. Hence, the concentrations in liver blood and the intracellular space of hepatocytes are in rapid equilibrium. The rate of elimination from liver blood is fub.C3 (drug concentration in compartment 3).CLih (total hepatic intrinsic clearance), where fub is the unbound drug concentration in blood. The drug in the liver may undergo metabolism by cytochromes (CLih-cyp) and an enterohepatic cycle with an intrinsic clearance (CLiEH), which is the sum of CLih-pgp (the intrinsic clearance for efflux of the parent drug in bile) and CLih-mr (the intrinsic clearance for metabolites that may be recycled, e.g., glucuronides, plus efflux by other transporters). The intact drug eliminated by active efflux and recycling metabolites goes into the bile compartment and a fraction (FaFg) returns to the liver blood. From the systemic circulation (‘blood’ compartment), the drug may return to the liver or eventually be eliminated by the renal route (renal clearance [CLr]). The pharmacokinetics of the victim drug are assumed to be linear, i.e., the clearances are independent of the substrate dose.
As shown in “Appendix A”, the drug concentration in blood (C4) may be calculated by solving the following system of equations:
where Rg is the ratio of gut clearance (CLg) to the apparent gut permeability surface product (PSg-app) escaping from the gut, and V3 and V4 are the volume of compartments 3 and 4, respectively. This parameter is actually a tuning parameter that accounts for the fact that a high level of extraction may be reached in the gut, although the number of cytochromes and transporters is much less than in the liver. A high extraction (low FaFg) may be understood as high Rg, i.e., a low PSg-app compared with CLg. In this system of equations, the parameters (Rg, fub, CLih, Qh, and CLr) are fixed in accordance with the literature.
In DDI studies, non-compartmental analysis is frequently applied. Estimates of total apparent clearance (CL/F) and possibly CLr (drug amount in urine/AUC) are obtained in each arm of the studies (subjects who receive the victim drug alone and subjects who receive the victim drug and the interactor). Hence, the ratio of AUCs (RAUC = AUC*/AUC) and the ratio of CLr values (RCLr = CLr*/CLr) may be obtained (where * denotes the parameter value for the substrate combined with the interactor). These metrics of the interaction have been expressed as a function of a number of parameters characteristic of the substrate (the CRs) and the perpetrator (the IXs). The CR of each pathway that may be impacted by the interaction are defined as follows:
where CRg-cyp and CRg-pgp are the contributions of CYP and Pgp to gut clearance, CRh-cyp, CRh-pgp and CRh-mr are the contributions of CYP, Pgp and recycling metabolites to hepatic intrinsic clearance, and CRr-pgp is the contribution of Pgp secretion to renal clearance. Each CR may range from 0 to 1. In addition, the sum of CRs in the gut and the sum of CRs in the liver must each be less than or equal to 1.
The characteristic parameters of the interactor are defined as follows:
-
IXg-cyp and IXh-cyp are the inhibition or induction potency of the interactor on CYP metabolism in the gut and the liver, respectively.
-
IXg-pgp, IXh-pgp, and IXr-pgp are the inhibition or induction potency of the interactor on the efflux transporter in the gut, liver, and kidney, respectively.
For inhibitors, each IX may range from 0 (no inhibition) to − 1 (complete inhibition). For inducers, each IX may range from 0 (no induction) to ∞ in theory. In practice, IX estimates of the most potent inducers have been ≤ 10.
In case of interaction, the value of the parameters is modified as follows:
The drug concentration in blood in case of interaction C4* can be calculated by solving the system of Eqs. (2a)–(2c) with the parameters modified according to Eqs. (3a)–(3d). The predicted AUC ratio is C4*/C4, although it is not computed exactly in this way (see “Appendix A”).
This procedure may be used to estimate the characteristic parameters by regression, using data from clinical studies of DDIs, and then to predict the magnitude of DDIs that have not been studied so far.
2.2 Estimation Method
For a given DDI, there are up to ten parameters (five CRs and five IXs) involved in the AUC ratio. Hence, it is not possible to estimate these parameters from a single clinical study; it is necessary to combine the results of several clinical studies involving a number of victim drugs and interactors. The parameters (the CRs and IXs) can then be estimated by regression. Even in this case, the number of datapoints will be hardly any larger than the number of parameters to be estimated, eventually leading to a large uncertainty regarding parameter estimates. To overcome this difficulty, we propose the following approach.
First, we use all available data, i.e., the AUC ratios (Eq. 1a–1c) and the CLr ratios (Eq. 3a–3d).
Second, the parameters are estimated by weighted non-linear least squares. The weights are defined as the inverse of the expected standard error of the AUC ratio (see “Appendix B”).
Third, all the constraints of the parameters, defined earlier, are accounted for in the regression in order to improve the consistency (avoid unrealistic values) and the identifiability of the model. The constraints are enforced by penalization in the objective function.
Fourth, some parameters are fixed using available data for drug interactions involving only cytochromes but not transporters (see Sect. 2.3).
The objective function to be minimized with respect to the parameters is finally as follows:
where \(\hat{P}\) represents the current estimates of CRs and IXs, Rpred and Robs are the predicted and observed AUC and CLr ratios, ns is the number of observations (number of AUC ratios and number of CLr ratio), and pen is the value of the penalty if some constraints are violated (see “Appendix B”). The expected standard error of Robs is calculated according to our previous work [25] (see “Appendix B”).
To minimize the objective function, the simplex algorithm is used; this is a local optimizer, based on a derivative-free method. This choice was made because the algorithm works well even with a complex set of constraints such as ours. The model was coded in Matlab® R2014b (MathWorks, Natick, MA, USA) and the function fminsearch was used for the simplex minimization of the objective function.
A confidence interval for the parameter estimates was obtained using a parametric bootstrap of the observations (see “Appendix B”).
2.3 Application to P-Glycoprotein
To evaluate and illustrate our approach, we considered the DDIs mediated by P-glycoprotein and CYP3A4/3A5.
A literature search was conducted in PubMed using the key words P-glycoprotein, drug interaction, and pharmacokinetics, with the results restricted to human studies in the period 1980–2017. After elimination of irrelevant or incomplete studies, a set of substrates and interactors was defined. Interactions studies with midazolam and alprazolam, which are CYP3A4/A5 substrates but not P-gp substrates, were also included in order to increase the precision of the estimates of IX of some interactors acting on CYP3A4/A5 and P-gp. Midazolam and alprazolam were chosen because (1) their F and Fg (1) their bioavailability (F) and bioavailability in the gut (Fg) are known, allowing the contribution of the gut and the liver to their pharmacokinetics to be disentangled; (2) a wide range of interaction studies have been published with each of them; and (3) their pharmacokinetics are appropriate for our purpose. Midazolam is highly metabolized by CYP3A4 in the gut (Fg = 0.51) and in the liver, while alprazolam is weakly metabolized by CYP3A4 in the gut (Fg = 0.94) but highly metabolized in the liver [28,29,30]. The interactions with midazolam and alprazolam allowed estimation of the IXg-cyp and IXh-cyp values for several interactors by root finding in Eq. (40) (see “Appendix A”).
To estimate the CRg-pgp, we need at least one interactor whose IRg-pgp is known or fixed. Itraconazole IRg-pgp was fixed to - 0.99. The reasons for this are that (1) itraconazole is responsible for the strongest known interaction with aliskiren, a P-gp substrate; (2) inhibition of P-gp in the gut was deemed to be the unique mechanism for this interaction; and (3) the concentration of drug producing 50% inhibition (IC50) of itraconazole for P-gp is 1.4 mg/L [31], a value far lower than the expected concentration of itraconazole in the gut lumen after a 200 mg dose.
The data were divided into two parts: the learning dataset was made up of the most informative studies with respect to the parameters to be estimated, given the assumed mechanisms of interaction; and the validation dataset consisted of the rest of the published data, involving a substrate and an interactor whose CRs and IXs had been estimated in the learning study.
The CRs and IXs were estimated as described earlier using the learning set. The goodness of fit was evaluated by several criteria. First, a plot of Rpred versus Robs was examined and visually compared with the identity line. Second, the proportions of predictions within 0.67- to 1.5-fold Robs and within 0.5- to 2-fold Robs were calculated. Third, the median and the interquartile range of Rpred/Robs, denoted as median fold error (MFE) and interquartile fold error (IQFE), were computed. Fourth, the confidence intervals of the parameter estimates were examined to ensure that all parameters were estimated with a good precision [the relative standard error (RSE) should be less than 30%]. Last, an external validation was carried out using the validation dataset. The Rpred was compared with Robs for a range of interactions not included in the learning dataset.
A sensitivity analysis was conducted to assess the impact of fixing the pharmacokinetic parameters (Rg, fub, CLih, Qh, and CLr) to a single common value for all predictions with a given substrate. Because all of the pharmacokinetic parameters are derived from the oral clearance and the fraction of dose absorbed for the substrate given alone, the impact of the variation of these two parameters on the predicted AUC ratio was evaluated by simulation under different assumptions regarding the CR and IX values.
3 Results
Following the literature search, some data had to be excluded before analysis. The fexofenadine–carbamazepine interaction [32] was excluded because the interaction was deemed to be due in part to the transporter multidrug resistance protein 2 (MRP2) in addition to P-gp. The interactions of verapamil as a substrate with atorvastatin [33] and lovastatin [34] were discarded because the pharmacokinetics of verapamil are strongly non-linear.
A total of 25 substrates (including midazolam and alprazolam) and 26 interactors (three inducers, 23 inhibitors) were retained in the regression analysis. The number of observations ns was 138, with 123 AUC ratios and 15 CLr ratios. The total number of parameters in the model was (25 + 26) × 5 = 255. The number of parameters to be estimated was 107 (50 CRs and 57 IXs). The rest of the parameters were fixed either to zero (e.g., for midazolam and alprazolam, CRg-pgp = CRh-pgp = CRr-pgp = 0) or, for CRh-cyp and IXh-cyp, to the value determined previously [17, 18, 35]. The data are shown in Table 1. The fixed parameters are shown in Electronic Supplementary Material Table 1. In Table 1, comparison of the AUC ratio with the elimination half-life (t½) ratio provides information on the site of the interaction. Indeed, the AUC ratio is equal to (CL/CL*) × (F*/F), while the t½ ratio is equal to (CL/CL*) if the volume of distribution is not altered by the interaction. If the AUC ratio is different from unity while the t½ ratio is close to unity, then the site of interaction is the gut, not the liver.
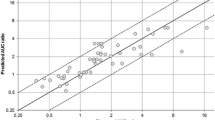
The goodness-of-fit plots are shown in Fig. 2. These plots did not show any particular trend, as expected, but there is an outlier at (Rpred 16, Robs 70) corresponding to the tacrolimus–telaprevir interaction. The same plot in log–log scale with the 2-fold prediction error limits confirms the lack of trend, but there are ten values outside these 2-fold error limits (one outside the 3-fold error limits). The outliers are described in Table 2.
a Predicted ratio (RAUC or RCLr) versus observed ratio for the learning set. The dashed line is the identity line (y = x). b Same plot as in a but the axes range is 0–18. c Predicted ratio (RAUC or RCLr) versus observed ratio in log–log scale. The continuous line is the identity line (y = x), the lower dashed line is y = 0.5x, and the upper dashed line is y = 2x. d Predicted ratio (RAUC or RCLr) versus observed ratio for the validation set. The dashed line is the identity line (y = x). An outlier at (Robs = 13.1, Rpred = 7.1) is not in the figure. AUC area under the plasma concentration–time curve, CLr renal clearance, RAUC ratio of AUCs, RCLr ratio of CLr values, Rpred predicted AUC and CLr ratio, Robs observed AUC and CLr ratio
The proportions of predictions within 0.67- to 1.5-fold Robs and within 0.5- to 2-fold Robs were 79% and 93%, respectively. The MFE was 0.98 (the ideal value is 1). The interquartile range of the fold error was 0.36. Regarding the confidence interval of the parameter estimates, the highest RSE was 15% for CRs and 10% for IXs (data not shown). Hence, the model was deemed acceptable.
The parameter estimates are shown in Tables 3 (CRs) and 4 (IXs). The substrates with a CRg-pgp greater than 0.9 are digoxin and edoxaban. The substrates with the highest CRh-pgp are colchicine (0.60) and fexofenadine (0.66). The substrates with a major elimination by tubular secretion are dabigatran (CRk-pgp = 0.5) and celiprolol (0.50). The major inhibitors of gut efflux are telaprevir, boceprevir, itraconazole, cyclosporine (ciclosporin), and lopinavir/ritonavir (IXg-pgp < − 0.9). The strongest inhibitors of efflux in bile are ketoconazole and ritonavir (IXh-pgp < − 0.9). Finally, cyclosporine and ritonavir are major inhibitors of tubular secretion (IXr-pgp < - 0.9).
The results of the external validation on 17 DDIs are shown in Fig. 2d and Table 5. The MFE was 0.90 (the ideal value is 1) while 76% and 88% of the predicted values were within 0.67- to 1.5-fold and within 2-fold of the observed values, respectively.
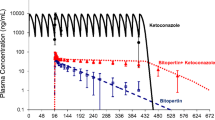
Some examples of predictions of the AUC ratio for several associations of drugs are shown in Fig. 3.
The results of the sensitivity analysis are shown in Electronic Supplementary Material Figs. 1 and 2. The main result is that the predicted AUC ratio is weakly sensitive to a small variation of the oral clearance and the fraction of dose absorbed in most regions of the plot. Hence, the choice of the reference values of the fixed parameters is not critical.
4 Discussion
In this study, a general model for analyzing and predicting DDIs mediated by an efflux transporter and a cytochrome was derived and evaluated. This a static (i.e., steady-state) mechanistic model, in the framework of CRs and IXs calculated solely from clinical data. This model, which we propose to call the ‘second-generation model’, is an improvement over our first-generation model, which did not (1) separate the intestine and the liver and (2) consider the impact of P-gp [35]. Although the model was applied only to P-gp as an efflux transporter, it could also apply to other transporters with the same pattern of actions such as BCRP. Likewise, we considered a single cytochrome (CYP3A4/A5) in this study, but the extension to a model with several cytochromes is straightforward. Extension to multiple transporters would be more problematic because the number of parameters to estimate would be very large. Incorporation of the impact of transporter or cytochrome polymorphism in this framework is also quite simple; see Tod et al. [24] for a description of how to proceed. Consideration of the impact of polymorphism is useful to increase the identifiability of the model when the genetic variations have a strong impact on the activity of the transporter or cytochromes.
Relying solely on in vivo data for the computations has the advantage that (1) there is no need to extrapolate from in vitro data and to make all the assumptions associated with this approach [36]; and (2) the impact of all the molecular species generated by the interactor (enantiomers, metabolites) is accounted for in the estimation of IXs [27]. The drawback is that for a new substrate or interactor, clinical data are required to be able to compute the CRs or IXs before other DDIs can be predicted. In addition, in vitro data, although not used in the computations, remain useful to ascertain the mechanisms of the interaction, or to fix some parameters at zero before applying the model.
The CR–IX framework is attractive because its parameters have an immediate physiological or pharmacological interpretation. Especially for inhibitors, the value of IX is more informative than an inhibition constant (Ki) value, because the Ki by itself cannot be interpreted if the concentration of the inhibitor facing the enzyme or the transporter is unknown. However, the CRs (for the transporter) and IXs depend on the dose of the substrate and interactor, respectively. As the dose increases, the CRs might decrease (if the transporter is saturated) while the IXs increase (in absolute value). For example, the CRg-pgp of atorvastatin was 0.66 at 20 mg but 0.28 at 40 mg, while the CRg-pgp of cyclosporine was 0.52 at 10 mg but 0.29 at 100 mg. Another advantage of the CR–IX framework is that any kind of inhibition (competitive or not) is handled in the same way. On the other hand, the IXs are estimated at steady state (e.g., for inducers or mechanism-based inhibitors of cytochromes), and thus the time course of the interaction cannot be described.
Despite its assumptions (for the substrate: linear pharmacokinetics, no active influx, rapid equilibrium between blood and intracellular water, no variation of binding in blood, no variation of blood-to-plasma ratio; for the interactor: the value of IX is independent of the substrate) and approximations (the CRh-mr is fixed and the recycling metabolites are assumed to be completely back transformed to the parent drug), the goodness-of-fit criteria and predictive performance metrics did not invalidate the model. Of note, our approach performed better than the in vitro–in vitro extrapolation method using digoxin as a probe for P-gp-mediated DDIs [37].
Using such a model requires an appropriate method for estimating the parameters. We proposed a flexible strategy that allows all available data to be treated at once and to enforce a complex set of constraints on the parameters to be estimated. Depending on how informative the data at hand is, the parameter estimates might be precise or not. If some parameter estimates are imprecise, more data should be incorporated. In our case, the addition of data obtained with selective substrates of intestinal and hepatic CYP3A4 (alprazolam and midazolam), which are not substrates of P-gp, improved the precision of the estimated potency of several mixed interactors [e.g., ketoconazole, clarithromycin, rifampicin (rifampin)]. As a result, the confidence interval of the parameter estimates expressed as RSE was quite narrow (see “Results”).
Treating a large amount of data all at once might reveal some outliers, which requires close examination. An outlier might sometimes suggest that the mechanisms of the interaction are not those assumed. For example, an additional transporter might be involved. Alternatively, some subjects might have a genetic variant resulting in different activity of the transporter or the CYPs. Lastly, the assumption of independence of IX with respect to the substrate may be violated. In particular, the DDIs with tacrolimus are not well-predicted with this model; this issue with tacrolimus has been observed by others [38, 39]. The interactions with tacrolimus are better predicted when the Ki values of the inhibitors are measured with tacrolimus as the substrate [39]. In our paradigm, this is equivalent to considering that the IXs of the inhibitors are specific to tacrolimus. In other words, the interactions with tacrolimus are poorly predicted with our approach, which should not be used in this case.
Despite these encouraging results, we do not pretend that all estimates obtained in this study are definitive. When new data become available, some estimates might be revised. For atorvastatin, some CYP3A4 metabolism was found (CRh-cyp = 0.27 and 0.23 from two different studies), although it is considered that the hepatic clearance of atorvastatin depends only on OATP influx at microdose [40].
From a practical point of view, the interactions involving efflux transporters at risk of overexposure are those involving (1) a substrate with a high CR for efflux combined with a strong efflux inhibitor; or (2) a substrate with mixed elimination (P-gp plus cytochrome) combined with a strong mixed inhibitor. According to the model and the parameter values (Tables 3 and 4), the following interactions, which have not been studied so far, are expected to exhibit the highest RAUC: aliskiren–telaprevir 6.5, venetoclax–ritonavir 5.3, venetoclax–telaprevir 5.9, venetoclax–itraconazole 5.3, naloxegol–lopinavir/ritonavir 15.6.
A limitation of the model is that only the impact of the interaction on exposure in plasma is described. However, efflux transporters such as P-gp and BCRP are also present at the blood–brain barrier. Inhibition of these transporters at the blood–brain barrier leads to a higher drug concentration in the brain parenchyma [41,42,43,44]. This increased penetration in the brain may have beneficial or adverse consequences that cannot be predicted solely by the RAUC in plasma. This is a limitation that can seldom be overcome, because measurements of drug concentration in the brain are generally not available in clinical trials. Hence, establishing a predictive model seems out of reach. Another limitation is that the description of the renal elimination of the drug is simplistic. Renal tubular secretion is taken into account through its contribution to CLr by the parameter CRr-pgp, and the interaction on tubular secretion is described through the potency of the interactor IXr-pgp; however, additional mechanisms of interaction due to modifications of renal blood flow, glomerular filtration rate, and tubular reabsorption, if present, are not taken into account.
5 Conclusions
In this study, the IMSM approach was successfully extended to DDIs mediated by P-gp and CYP3A4/3A5. The method revealed good predictive performances by internal and external validation, with the exception of tacrolimus. However, because clinical data are required to train the algorithm, the method cannot be applied at the stage of preclinical drug development. The IMSM approach is more useful during the clinical phase of development and in hospital settings, e.g., in the context of prescription analysis to optimize treatments. Currently, the IMSM approach is implemented for CYP-mediated DDIs on the free DDI-Predictor website (https://www.ddi-predictor.org). Extension to incorporate P-gp-mediated DDIs will occur in the future.
References
Neuvonen PJ. Towards safer and more predictable drug treatment–reflections from studies of the first BCPT Prize awardee. Basic Clin Pharmacol Toxicol. 2012;110(3):207–18.
Food and Drug Administration. Guidance for industry: drug interaction studies—study design, data analysis, implications for dosing, and labeling recommendations. Rockville: FDA; 2012.
European Medicines Agency. Guideline on the investigation of drug interactions. London: Committee for Medical Products for Human Use (CHMP); 2012.
Rostami-Hodjegan A, Tucker G. ‘In silico’ simulations to assess the ‘in vivo’ consequences of ‘in vitro’ metabolic drug–drug interactions. Drug Discov Today Technol. 2004;1(4):441–8.
Chu X, Korzekwa K, Elsby R, Fenner K, Galetin A, Lai Y, et al. Intracellular drug concentrations and transporters: measurement, modeling, and implications for the liver. Clin Pharmacol Ther. 2013;94(1):126–41.
International Transporter C, Giacomini KM, Huang SM, Tweedie DJ, Benet LZ, Brouwer KL, et al. Membrane transporters in drug development. Nat Rev Drug Discov. 2010;9(3):215–36.
Niemi M, Pasanen MK, Neuvonen PJ. Organic anion transporting polypeptide 1B1: a genetically polymorphic transporter of major importance for hepatic drug uptake. Pharmacol Rev. 2011;63(1):157–81.
Zhou Y, Yuan J, Li Z, Wang Z, Cheng D, Du Y, et al. Genetic polymorphisms and function of the organic anion-transporting polypeptide 1A2 and its clinical relevance in drug disposition. Pharmacology. 2015;95(3–4):201–8.
Yu J, Zhou Z, Tay-Sontheimer J, Levy RH, Ragueneau-Majlessi I. Intestinal drug interactions mediated by OATPs: a systematic review of preclinical and clinical findings. J Pharm Sci. 2017;106(9):2312–25.
Wessler JD, Grip LT, Mendell J, Giugliano RP. The P-glycoprotein transport system and cardiovascular drugs. J Am Coll Cardiol. 2013;61(25):2495–502.
Mao Q, Unadkat JD. Role of the breast cancer resistance protein (BCRP/ABCG2) in drug transport—an update. AAPS J. 2015;17(1):65–82.
Sayama H, Komura H, Kogayu M. Application of hybrid approach based on empirical and physiological concept for predicting pharmacokinetics in humans—usefulness of exponent on prospective evaluation of predictability. Drug Metab Dispos. 2013;41(2):498–507.
Guest EJ, Rowland-Yeo K, Rostami-Hodjegan A, Tucker GT, Houston JB, Galetin A. Assessment of algorithms for predicting drug–drug interactions via inhibition mechanisms: comparison of dynamic and static models. Br J Clin Pharmacol. 2011;71(1):72–87.
Yoshida K, Maeda K, Sugiyama Y. Transporter-mediated drug–drug interactions involving OATP substrates: predictions based on in vitro inhibition studies. Clin Pharmacol Ther. 2012;91(6):1053–64.
Hu ZY. Disposition pathway-dependent approach for predicting organic anion-transporting polypeptide-mediated drug–drug interactions. Clin Pharmacokinet. 2013;52(6):433–41.
Varma MV, Lin J, Bi YA, Rotter CJ, Fahmi OA, Lam JL, et al. Quantitative prediction of repaglinide–rifampicin complex drug interactions using dynamic and static mechanistic models: delineating differential CYP3A4 induction and OATP1B1 inhibition potential of rifampicin. Drug Metab Dispos. 2013;41(5):966–74.
Ohno Y, Hisaka A, Suzuki H. General framework for the quantitative prediction of CYP3A4-mediated oral drug interactions based on the AUC increase by coadministration of standard drugs. Clin Pharmacokinet. 2007;46(8):681–96.
Ohno Y, Hisaka A, Ueno M, Suzuki H. General framework for the prediction of oral drug interactions caused by CYP3A4 induction from in vivo information. Clin Pharmacokinet. 2008;47(10):669–80.
Tod M, Goutelle S, Clavel-Grabit F, Nicolas G, Charpiat B. Quantitative prediction of cytochrome P450 (CYP) 2D6-mediated drug interactions. Clin Pharmacokinet. 2011;50(8):519–30.
Castellan AC, Tod M, Gueyffier F, Audars M, Cambriels F, Kassai B, et al. Quantitative prediction of the impact of drug interactions and genetic polymorphisms on cytochrome P450 2C9 substrate exposure. Clin Pharmacokinet. 2013;52(3):199–209.
Goutelle S, Bourguignon L, Bleyzac N, Berry J, Clavel-Grabit F, Tod M. In vivo quantitative prediction of the effect of gene polymorphisms and drug interactions on drug exposure for CYP2C19 substrates. AAPS J. 2013;15(2):415–26.
Gabriel L, Tod M, Goutelle S. Quantitative prediction of drug interactions caused by CYP1A2 inhibitors and inducers. Clin Pharmacokinet. 2016;55(8):977–90.
Tod M, Goutelle S, Gagnieu MC, Genophar IIWG. Genotype-based quantitative prediction of drug exposure for drugs metabolized by CYP2D6. Clin Pharmacol Ther. 2011;90(4):582–7.
Tod M, Nkoud-Mongo C, Gueyffier F. Impact of genetic polymorphism on drug–drug interactions mediated by cytochromes: a general approach. AAPS J. 2013;15(4):1242–52.
Tod M, Bourguignon L, Bleyzac N, Goutelle S. A model for predicting the interindividual variability of drug–drug interactions. AAPS J. 2017;19(2):497–509.
Tod M, Pierrillas PB, Bourguignon L, Goutelle S. Comparison of the static in vivo approach to a physiologically based pharmacokinetic approach for metabolic drug–drug interactions prediction. Int J Pharmacokinet. 2016;1(1):25–34. https://doi.org/10.4155/ipk.16.2.
Tod M, Goutelle S, Bourguignon L, Bleyzac N. Quantitative prediction of drug-drug interactions involving inhibitory metabolites by physiologically based pharmacokinetic models: is it worth it? CPT Pharmacomet Syst Pharmacol. 2017;6(4):226.
Gertz M, Davis JD, Harrison A, Houston JB, Galetin A. Grapefruit juice–drug interaction studies as a method to assess the extent of intestinal availability: utility and limitations. Curr Drug Metab. 2008;9(8):785–95.
Gertz M, Harrison A, Houston JB, Galetin A. Prediction of human intestinal first-pass metabolism of 25 CYP3A substrates from in vitro clearance and permeability data. Drug Metab Dispos. 2010;38(7):1147–58.
Kadono K, Akabane T, Tabata K, Gato K, Terashita S, Teramura T. Quantitative prediction of intestinal metabolism in humans from a simplified intestinal availability model and empirical scaling factor. Drug Metab Dispos. 2010;38(7):1230–7.
Wang EJ, Lew K, Casciano CN, Clement RP, Johnson WW. Interaction of common azole antifungals with P glycoprotein. Antimicrob Agents Chemother. 2002;46(1):160–5.
Akamine Y, Miura M, Yasui-Furukori N, Kojima M, Uno T. Carbamazepine differentially affects the pharmacokinetics of fexofenadine enantiomers. Br J Clin Pharmacol. 2012;73(3):478–81.
Choi DH, Shin WG, Choi JS. Drug interaction between oral atorvastatin and verapamil in healthy subjects: effects of atorvastatin on the pharmacokinetics of verapamil and norverapamil. Eur J Clin Pharmacol. 2008;64(5):445–9.
Choi DH, Chung JH, Choi JS. Pharmacokinetic interaction between oral lovastatin and verapamil in healthy subjects: role of P-glycoprotein inhibition by lovastatin. Eur J Clin Pharmacol. 2010;66(3):285–90.
Loue C, Tod M. Reliability and extension of quantitative prediction of CYP3A4-mediated drug interactions based on clinical data. AAPS J. 2014;16(6):1309–20.
Li R, Barton HA, Varma MV. Prediction of pharmacokinetics and drug–drug interactions when hepatic transporters are involved. Clin Pharmacokinet. 2014;53(8):659–78.
Fenner KS, Troutman MD, Kempshall S, Cook JA, Ware JA, Smith DA, et al. Drug–drug interactions mediated through P-glycoprotein: clinical relevance and in vitro-in vivo correlation using digoxin as a probe drug. Clin Pharmacol Ther. 2009;85(2):173–81.
Chapron B, Risler L, Phillips B, Collins C, Thummel K, Shen D. Reversible, time-dependent inhibition of CYP3A-mediated metabolism of midazolam and tacrolimus by telaprevir in human liver microsomes. J Pharm Pharm Sci. 2015;18(1):101–11.
Oda K, Yamano K, Otsuka Y. In vitro–in vivo extrapolations to evaluate the effect of concomitant drugs on tacrolimus (FK506) exposure. Biopharm Drug Dispos. 2015;36(5):265–74.
Maeda K, Ikeda Y, Fujita T, Yoshida K, Azuma Y, Haruyama Y, et al. Identification of the rate-determining process in the hepatic clearance of atorvastatin in a clinical cassette microdosing study. Clin Pharmacol Ther. 2011;90(4):575–81.
de Vries NA, Zhao J, Kroon E, Buckle T, Beijnen JH, van Tellingen O. P-glycoprotein and breast cancer resistance protein: two dominant transporters working together in limiting the brain penetration of topotecan. Clin Cancer Res. 2007;13(21):6440–9.
Tang SC, de Vries N, Sparidans RW, Wagenaar E, Beijnen JH, Schinkel AH. Impact of P-glycoprotein (ABCB1) and breast cancer resistance protein (ABCG2) gene dosage on plasma pharmacokinetics and brain accumulation of dasatinib, sorafenib, and sunitinib. J Pharmacol Exp Ther. 2013;346(3):486–94.
Durmus S, Sparidans RW, van Esch A, Wagenaar E, Beijnen JH, Schinkel AH. Breast cancer resistance protein (BCRP/ABCG2) and P-glycoprotein (P-GP/ABCB1) restrict oral availability and brain accumulation of the PARP inhibitor rucaparib (AG-014699). Pharm Res. 2015;32(1):37–46.
Henin E, Honorat M, Guitton J, Di Pietro A, Payen L, Tod M. Pharmacokinetic interactions in mice between irinotecan and MBL-II-141, an ABCG2 inhibitor. Biopharm Drug Dispos. 2017;38(5):351–62.
Rebello S, Compain S, Feng A, Hariry S, Dieterich HA, Jarugula V. Effect of cyclosporine on the pharmacokinetics of aliskiren in healthy subjects. J Clin Pharmacol. 2011;51(11):1549–60.
Vaidyanathan S, Camenisch G, Schuetz H, Reynolds C, Yeh CM, Bizot MN, et al. Pharmacokinetics of the oral direct renin inhibitor aliskiren in combination with digoxin, atorvastatin, and ketoconazole in healthy subjects: the role of P-glycoprotein in the disposition of aliskiren. J Clin Pharmacol. 2008;48(11):1323–38.
Rebello S, Leon S, Hariry S, Dahlke M, Jarugula V. Effect of verapamil on the pharmacokinetics of aliskiren in healthy participants. J Clin Pharmacol. 2011;51(2):218–28.
Tapaninen T, Backman JT, Kurkinen KJ, Neuvonen PJ, Niemi M. Itraconazole, a P-glycoprotein and CYP3A4 inhibitor, markedly raises the plasma concentrations and enhances the renin-inhibiting effect of aliskiren. J Clin Pharmacol. 2011;51(3):359–67.
Vaidyanathan S, Valencia J, Kemp C, Zhao C, Yeh CM, Bizot MN, et al. Lack of pharmacokinetic interactions of aliskiren, a novel direct renin inhibitor for the treatment of hypertension, with the antihypertensives amlodipine, valsartan, hydrochlorothiazide (HCTZ) and ramipril in healthy volunteers. Int J Clin Pract. 2006;60(11):1343–56.
Frost CE, Byon W, Song Y, Wang J, Schuster AE, Boyd RA, et al. Effect of ketoconazole and diltiazem on the pharmacokinetics of apixaban, an oral direct factor Xa inhibitor. Br J Clin Pharmacol. 2015;79(5):838–46.
Frost C, Shenker A, Gandhi MD, Pursley J, Barrett YC, Wang J, et al. Evaluation of the effect of naproxen on the pharmacokinetics and pharmacodynamics of apixaban. Br J Clin Pharmacol. 2014;78(4):877–85.
Vakkalagadda B, Frost C, Byon W, Boyd RA, Wang J, Zhang D, et al. Effect of rifampin on the pharmacokinetics of apixaban, an oral direct inhibitor of factor Xa. Am J Cardiovasc Drugs. 2016;16(2):119–27.
Lee JE, van Heeswijk R, Alves K, Smith F, Garg V. Effect of the hepatitis C virus protease inhibitor telaprevir on the pharmacokinetics of amlodipine and atorvastatin. Antimicrob Agents Chemother. 2011;55(10):4569–74.
Mazzu AL, Lasseter KC, Shamblen EC, Agarwal V, Lettieri J, Sundaresen P. Itraconazole alters the pharmacokinetics of atorvastatin to a greater extent than either cerivastatin or pravastatin. Clin Pharmacol Ther. 2000;68(4):391–400.
Kantola T, Kivisto KT, Neuvonen PJ. Effect of itraconazole on the pharmacokinetics of atorvastatin. Clin Pharmacol Ther. 1998;64(1):58–65.
Summary of product characteristics: KALETRA (lopinavir/ritonavir). http://ec.europa.eu/health/documents/community-register/2001/200103204167/anx_4167_en.pdf. Accessed 24 Aug 2018.
Lilja JJ, Backman JT, Laitila J, Luurila H, Neuvonen PJ. Itraconazole increases but grapefruit juice greatly decreases plasma concentrations of celiprolol. Clin Pharmacol Ther. 2003;73(3):192–8.
Garg V, van Heeswijk R, Lee JE, Alves K, Nadkarni P, Luo X. Effect of telaprevir on the pharmacokinetics of cyclosporine and tacrolimus. Hepatology. 2011;54(1):20–7.
Hulskotte E, Gupta S, Xuan F, van Zutven M, O’Mara E, Feng HP, et al. Pharmacokinetic interaction between the hepatitis C virus protease inhibitor boceprevir and cyclosporine and tacrolimus in healthy volunteers. Hepatology. 2012;56(5):1622–30.
Lu C, Hatsis P, Berg C, Lee FW, Balani SK. Prediction of pharmacokinetic drug–drug interactions using human hepatocyte suspension in plasma and cytochrome P450 phenotypic data. II. In vitro–in vivo correlation with ketoconazole. Drug Metab Dispos. 2008;36(7):1255–60.
Florea NR, Capitano B, Nightingale CH, Hull D, Leitz GJ, Nicolau DP. Beneficial pharmacokinetic interaction between cyclosporine and itraconazole in renal transplant recipients. Transpl Proc. 2003;35(8):2873–7.
Davis MW, Wason S. Effect of steady-state atorvastatin on the pharmacokinetics of a single dose of colchicine in healthy adults under fasted conditions. Clin Drug Investig. 2014;34(4):259–67.
Terkeltaub RA, Furst DE, Digiacinto JL, Kook KA, Davis MW. Novel evidence-based colchicine dose-reduction algorithm to predict and prevent colchicine toxicity in the presence of cytochrome P450 3A4/P-glycoprotein inhibitors. Arthritis Rheum. 2011;63(8):2226–37.
Summary of product characteristics: PRADAXA (dabigatran). http://www.ema.europa.eu/docs/en_GB/document_library/EPAR_-_Product_Information/human/000829/WC500041059.pdf. Accessed 24 Aug 2018.
Delavenne X, Ollier E, Basset T, Bertoletti L, Accassat S, Garcin A, et al. A semi-mechanistic absorption model to evaluate drug–drug interaction with dabigatran: application with clarithromycin. Br J Clin Pharmacol. 2013;76(1):107–13.
Hartter S, Sennewald R, Nehmiz G, Reilly P. Oral bioavailability of dabigatran etexilate (Pradaxa®) after co-medication with verapamil in healthy subjects. Br J Clin Pharmacol. 2013;75(4):1053–62.
Rengelshausen J, Goggelmann C, Burhenne J, Riedel KD, Ludwig J, Weiss J, et al. Contribution of increased oral bioavailability and reduced nonglomerular renal clearance of digoxin to the digoxin–clarithromycin interaction. Br J Clin Pharmacol. 2003;56(1):32–8.
Mahgoub AA, El-Medany AH, Abdulatif AS. A comparison between the effects of diltiazem and isosorbide dinitrate on digoxin pharmacodynamics and kinetics in the treatment of patients with chronic ischemic heart failure. Saudi Med J. 2002;23(6):725–31.
Summary of product characteristics: MULTAQ (dronedarone). http://www.ema.europa.eu/docs/en_GB/document_library/EPAR_-_Product_Information/human/001043/WC500044534.pdf. Accessed 24 Aug 2018.
Martin P, Gillen M, Millson D, Oliver S, Brealey C, Elsby R, et al. Effects of fostamatinib on the pharmacokinetics of digoxin (a P-glycoprotein substrate): results from in vitro and phase I clinical studies. Clin Ther. 2015;37(12):2811–22.
Jalava KM, Partanen J, Neuvonen PJ. Itraconazole decreases renal clearance of digoxin. Ther Drug Monit. 1997;19(6):609–13.
Pedersen KE, Christiansen BD, Klitgaard NA, Nielsen-Kudsk F. Effect of quinidine on digoxin bioavailability. Eur J Clin Pharmacol. 1983;24(1):41–7.
Greiner B, Eichelbaum M, Fritz P, Kreichgauer HP, von Richter O, Zundler J, et al. The role of intestinal P-glycoprotein in the interaction of digoxin and rifampin. J Clin Invest. 1999;104(2):147–53.
Ding R, Tayrouz Y, Riedel KD, Burhenne J, Weiss J, Mikus G, et al. Substantial pharmacokinetic interaction between digoxin and ritonavir in healthy volunteers. Clin Pharmacol Ther. 2004;76(1):73–84.
Penzak SR, Shen JM, Alfaro RM, Remaley AT, Natarajan V, Falloon J. Ritonavir decreases the nonrenal clearance of digoxin in healthy volunteers with known MDR1 genotypes. Ther Drug Monit. 2004;26(3):322–30.
Wyen C, Fuhr U, Frank D, Aarnoutse RE, Klaassen T, Lazar A, et al. Effect of an antiretroviral regimen containing ritonavir boosted lopinavir on intestinal and hepatic CYP3A, CYP2D6 and P-glycoprotein in HIV-infected patients. Clin Pharmacol Ther. 2008;84(1):75–82.
Mueller SC, Uehleke B, Woehling H, Petzsch M, Majcher-Peszynska J, Hehl EM, et al. Effect of St John’s wort dose and preparations on the pharmacokinetics of digoxin. Clin Pharmacol Ther. 2004;75(6):546–57.
Garg V, Chandorkar G, Farmer HF, Smith F, Alves K, van Heeswijk RP. Effect of telaprevir on the pharmacokinetics of midazolam and digoxin. J Clin Pharmacol. 2012;52(10):1566–73.
Teng R, Butler K. A pharmacokinetic interaction study of ticagrelor and digoxin in healthy volunteers. Eur J Clin Pharmacol. 2013;69(10):1801–8.
Johansson S, Read J, Oliver S, Steinberg M, Li Y, Lisbon E, et al. Pharmacokinetic evaluations of the co-administrations of vandetanib and metformin, digoxin, midazolam, omeprazole or ranitidine. Clin Pharmacokinet. 2014;53(9):837–47.
Verapamil—FDA prescribing information. https://www.accessdata.fda.gov/drugsatfda_docs/label/2016/018925s010lbl.pdf. Accessed 24 Aug 2018.
Hedman A, Angelin B, Arvidsson A, Beck O, Dahlqvist R, Nilsson B, et al. Digoxin–verapamil interaction: reduction of biliary but not renal digoxin clearance in humans. Clin Pharmacol Ther. 1991;49(3):256–62.
Parasrampuria DA, Mendell J, Shi M, Matsushima N, Zahir H, Truitt K. Edoxaban drug–drug interactions with ketoconazole, erythromycin, and cyclosporine. Br J Clin Pharmacol. 2016;82(6):1591–600.
Mendell J, Zahir H, Matsushima N, Noveck R, Lee F, Chen S, et al. Drug–drug interaction studies of cardiovascular drugs involving P-glycoprotein, an efflux transporter, on the pharmacokinetics of edoxaban, an oral factor Xa inhibitor. Am J Cardiovasc Drugs. 2013;13(5):331–42.
Kovarik JM, Kalbag J, Figueiredo J, Rouilly M, Frazier OL, Rordorf C. Differential influence of two cyclosporine formulations on everolimus pharmacokinetics: a clinically relevant pharmacokinetic interaction. J Clin Pharmacol. 2002;42(1):95–9.
Kovarik JM, Beyer D, Bizot MN, Jiang Q, Shenouda M, Schmouder RL. Effect of multiple-dose erythromycin on everolimus pharmacokinetics. Eur J Clin Pharmacol. 2005;61(1):35–8.
Kovarik JM, Beyer D, Bizot MN, Jiang Q, Shenouda M, Schmouder RL. Blood concentrations of everolimus are markedly increased by ketoconazole. J Clin Pharmacol. 2005;45(5):514–8.
Kovarik JM, Beyer D, Bizot MN, Jiang Q, Allison MJ, Schmouder RL. Pharmacokinetic interaction between verapamil and everolimus in healthy subjects. Br J Clin Pharmacol. 2005;60(4):434–7.
Shimizu M, Uno T, Sugawara K, Tateishi T. Effects of single and multiple doses of itraconazole on the pharmacokinetics of fexofenadine, a substrate of P-glycoprotein. Br J Clin Pharmacol. 2006;62(3):372–6.
Shon JH, Yoon YR, Hong WS, Nguyen PM, Lee SS, Choi YG, et al. Effect of itraconazole on the pharmacokinetics and pharmacodynamics of fexofenadine in relation to the MDR1 genetic polymorphism. Clin Pharmacol Ther. 2005;78(2):191–201.
Tateishi T, Miura M, Suzuki T, Uno T. The different effects of itraconazole on the pharmacokinetics of fexofenadine enantiomers. Br J Clin Pharmacol. 2008;65(5):693–700.
van Heeswijk RP, Bourbeau M, Campbell P, Seguin I, Chauhan BM, Foster BC, et al. Time-dependent interaction between lopinavir/ritonavir and fexofenadine. J Clin Pharmacol. 2006;46(7):758–67.
Kharasch ED, Bedynek PS, Walker A, Whittington D, Hoffer C. Mechanism of ritonavir changes in methadone pharmacokinetics and pharmacodynamics: II. Ritonavir effects on CYP3A and P-glycoprotein activities. Clin Pharmacol Ther. 2008;84(4):506–12.
Hamman MA, Bruce MA, Haehner-Daniels BD, Hall SD. The effect of rifampin administration on the disposition of fexofenadine. Clin Pharmacol Ther. 2001;69(3):114–21.
Wang Z, Hamman MA, Huang SM, Lesko LJ, Hall SD. Effect of St John’s wort on the pharmacokinetics of fexofenadine. Clin Pharmacol Ther. 2002;71(6):414–20.
Dresser GK, Schwarz UI, Wilkinson GR, Kim RB. Coordinate induction of both cytochrome P4503A and MDR1 by St John’s wort in healthy subjects. Clin Pharmacol Ther. 2003;73(1):41–50.
Saruwatari J, Yasui-Furukori N, Niioka T, Akamine Y, Takashima A, Kaneko S, et al. Different effects of the selective serotonin reuptake inhibitors fluvoxamine, paroxetine, and sertraline on the pharmacokinetics of fexofenadine in healthy volunteers. J Clin Psychopharmacol. 2012;32(2):195–9.
Talavera Pons S, Boyer A, Lamblin G, Chennell P, Chatenet FT, Nicolas C, et al. Managing drug–drug interactions with new direct-acting antiviral agents in chronic hepatitis C. Br J Clin Pharmacol. 2017;83(2):269–93.
Hachad H, Ragueneau-Majlessi I, Levy RH. A useful tool for drug interaction evaluation: the University of Washington Metabolism and Transport Drug Interaction Database. Hum Genom. 2010;5(1):61–72.
Kim TE, Lee H, Lim KS, Lee S, Yoon SH, Park KM, et al. Effects of HM30181, a P-glycoprotein inhibitor, on the pharmacokinetics and pharmacodynamics of loperamide in healthy volunteers. Br J Clin Pharmacol. 2014;78(3):556–64.
Niemi M, Tornio A, Pasanen MK, Fredrikson H, Neuvonen PJ, Backman JT. Itraconazole, gemfibrozil and their combination markedly raise the plasma concentrations of loperamide. Eur J Clin Pharmacol. 2006;62(6):463–72.
Misaka S, Miyazaki N, Yatabe MS, Ono T, Shikama Y, Fukushima T, et al. Pharmacokinetic and pharmacodynamic interaction of nadolol with itraconazole, rifampicin and grapefruit juice in healthy volunteers. J Clin Pharmacol. 2013;53(7):738–45.
Bui K, She F, Zhou D, Butler K, Al-Huniti N, Sostek M. The effect of quinidine, a strong P-glycoprotein inhibitor, on the pharmacokinetics and central nervous system distribution of naloxegol. J Clin Pharmacol. 2016;56(4):497–505.
Mueck W, Kubitza D, Becka M. Co-administration of rivaroxaban with drugs that share its elimination pathways: pharmacokinetic effects in healthy subjects. Br J Clin Pharmacol. 2013;76(3):455–66.
Bottiger Y, Sawe J, Brattstrom C, Tollemar J, Burke JT, Hass G, et al. Pharmacokinetic interaction between single oral doses of diltiazem and sirolimus in healthy volunteers. Clin Pharmacol Ther. 2001;69(1):32–40.
Cohen EE, Wu K, Hartford C, Kocherginsky M, Eaton KN, Zha Y, et al. Phase I studies of sirolimus alone or in combination with pharmacokinetic modulators in advanced cancer patients. Clin Cancer Res. 2012;18(17):4785–93.
Zimmerman JJ. Exposure–response relationships and drug interactions of sirolimus. AAPS J. 2004;6(4):e28.
Floren LC, Bekersky I, Benet LZ, Mekki Q, Dressler D, Lee JW, et al. Tacrolimus oral bioavailability doubles with coadministration of ketoconazole. Clin Pharmacol Ther. 1997;62(1):41–9.
Nara M, Takahashi N, Miura M, Niioka T, Kagaya H, Fujishima N, et al. Effect of itraconazole on the concentrations of tacrolimus and cyclosporine in the blood of patients receiving allogeneic hematopoietic stem cell transplants. Eur J Clin Pharmacol. 2013;69(6):1321–9.
Giessmann T, May K, Modess C, Wegner D, Hecker U, Zschiesche M, et al. Carbamazepine regulates intestinal P-glycoprotein and multidrug resistance protein MRP2 and influences disposition of talinolol in humans. Clin Pharmacol Ther. 2004;76(3):192–200.
Westphal K, Weinbrenner A, Zschiesche M, Franke G, Knoke M, Oertel R, et al. Induction of P-glycoprotein by rifampin increases intestinal secretion of talinolol in human beings: a new type of drug/drug interaction. Clin Pharmacol Ther. 2000;68(4):345–55.
Schwarz UI, Hanso H, Oertel R, Miehlke S, Kuhlisch E, Glaeser H, et al. Induction of intestinal P-glycoprotein by St John’s wort reduces the oral bioavailability of talinolol. Clin Pharmacol Ther. 2007;81(5):669–78.
Schwarz UI, Gramatte T, Krappweis J, Berndt A, Oertel R, von Richter O, et al. Unexpected effect of verapamil on oral bioavailability of the beta-blocker talinolol in humans. Clin Pharmacol Ther. 1999;65(3):283–90.
Agarwal SK, Salem AH, Danilov AV, Hu B, Puvvada S, Gutierrez M, et al. Effect of ketoconazole, a strong CYP3A inhibitor, on the pharmacokinetics of venetoclax, a BCL-2 inhibitor, in patients with non-Hodgkin lymphoma. Br J Clin Pharmacol. 2017;83(4):846–54.
Agarwal SK, Hu B, Chien D, Wong SL, Salem AH. Evaluation of rifampin’s transporter inhibitory and CYP3A inductive effects on the pharmacokinetics of venetoclax, a BCL-2 inhibitor: results of a single- and multiple-dose study. J Clin Pharmacol. 2016;56(11):1335–43.
Damkier P, Hansen LL, Brosen K. Effect of diclofenac, disulfiram, itraconazole, grapefruit juice and erythromycin on the pharmacokinetics of quinidine. Br J Clin Pharmacol. 1999;48(6):829–38.
Kaukonen KM, Olkkola KT, Neuvonen PJ. Itraconazole increases plasma concentrations of quinidine. Clin Pharmacol Ther. 1997;62(5):510–7.
Damkier P, Hansen LL, Brosen K. Rifampicin treatment greatly increases the apparent oral clearance of quinidine. Pharmacol Toxicol. 1999;85(6):257–62.
Gorski JC, Jones DR, Haehner-Daniels BD, Hamman MA, O’Mara EM Jr, Hall SD. The contribution of intestinal and hepatic CYP3A to the interaction between midazolam and clarithromycin. Clin Pharmacol Ther. 1998;64(2):133–43.
Zimmermann T, Yeates RA, Laufen H, Scharpf F, Leitold M, Wildfeuer A. Influence of the antibiotics erythromycin and azithromycin on the pharmacokinetics and pharmacodynamics of midazolam. Arzneimittelforschung. 1996;46(2):213–7.
Lam YW, Alfaro CL, Ereshefsky L, Miller M. Pharmacokinetic and pharmacodynamic interactions of oral midazolam with ketoconazole, fluoxetine, fluvoxamine, and nefazodone. J Clin Pharmacol. 2003;43(11):1274–82.
Backman JT, Kivisto KT, Olkkola KT, Neuvonen PJ. The area under the plasma concentration–time curve for oral midazolam is 400-fold larger during treatment with itraconazole than with rifampicin. Eur J Clin Pharmacol. 1998;54(1):53–8.
Chung E, Nafziger AN, Kazierad DJ, Bertino JS Jr. Comparison of midazolam and simvastatin as cytochrome P450 3A probes. Clin Pharmacol Ther. 2006;79(4):350–61.
Kirby BJ, Collier AC, Kharasch ED, Whittington D, Thummel KE, Unadkat JD. Complex drug interactions of HIV protease inhibitors 1: inactivation, induction, and inhibition of cytochrome P450 3A by ritonavir or nelfinavir. Drug Metab Dispos. 2011;39(6):1070–8.
Summary of product characteristics: OLYSIO (simeprevir). http://www.ema.europa.eu/docs/en_GB/document_library/EPAR_-_Product_Information/human/002777/WC500167867.pdf. Accessed 24 Aug 2018.
Wang Z, Gorski JC, Hamman MA, Huang SM, Lesko LJ, Hall SD. The effects of St John’s wort (Hypericum perforatum) on human cytochrome P450 activity. Clin Pharmacol Ther. 2001;70(4):317–26.
Summary of product characteristics: Incivo, INN-telaprevir. 2013. http://www.ema.europa.eu/docs/en_GB/document_library/EPAR_-_Product_Information/human/002313/WC500115529.pdf. Accessed 24 Aug 2018.
Furukori H, Otani K, Yasui N, Kondo T, Kaneko S, Shimoyama R, et al. Effect of carbamazepine on the single oral dose pharmacokinetics of alprazolam. Neuropsychopharmacology. 1998;18(5):364–9.
Yasui N, Otani K, Kaneko S, Ohkubo T, Osanai T, Sugawara K, et al. A kinetic and dynamic study of oral alprazolam with and without erythromycin in humans: in vivo evidence for the involvement of CYP3A4 in alprazolam metabolism. Clin Pharmacol Ther. 1996;59(5):514–9.
Fleishaker JC, Hulst LK. A pharmacokinetic and pharmacodynamic evaluation of the combined administration of alprazolam and fluvoxamine. Eur J Clin Pharmacol. 1994;46(1):35–9.
Yasui N, Kondo T, Otani K, Furukori H, Kaneko S, Ohkubo T, et al. Effect of itraconazole on the single oral dose pharmacokinetics and pharmacodynamics of alprazolam. Psychopharmacology. 1998;139(3):269–73.
Greenblatt DJ, Wright CE, von Moltke LL, Harmatz JS, Ehrenberg BL, Harrel LM, et al. Ketoconazole inhibition of triazolam and alprazolam clearance: differential kinetic and dynamic consequences. Clin Pharmacol Ther. 1998;64(3):237–47.
Schmider J, Brockmoller J, Arold G, Bauer S, Roots I. Simultaneous assessment of CYP3A4 and CYP1A2 activity in vivo with alprazolam and caffeine. Pharmacogenetics. 1999;9(6):725–34.
Markowitz JS, DeVane CL, Boulton DW, Carson SW, Nahas Z, Risch SC. Effect of St. John’s wort (Hypericum perforatum) on cytochrome P-450 2D6 and 3A4 activity in healthy volunteers. Life Sci. 2000;66(9):PL133–9.
Jones TE, Morris RG, Mathew TH. Diltiazem–cyclosporin pharmacokinetic interaction—dose–response relationship. Br J Clin Pharmacol. 1997;44(5):499–504.
Gupta SK, Bakran A, Johnson RW, Rowland M. Cyclosporin–erythromycin interaction in renal transplant patients. Br J Clin Pharmacol. 1989;27(4):475–81.
Bauer S, Stormer E, Johne A, Kruger H, Budde K, Neumayer HH, et al. Alterations in cyclosporin A pharmacokinetics and metabolism during treatment with St John’s wort in renal transplant patients. Br J Clin Pharmacol. 2003;55(2):203–11.
Robinson K, Johnston A, Walker S, Mulrow JP, McKenna WJ, Holt DW. The digoxin–amiodarone interaction. Cardiovasc Drugs Ther. 1989;3(1):25–8.
Dorian P, Strauss M, Cardella C, David T, East S, Ogilvie R. Digoxin–cyclosporine interaction: severe digitalis toxicity after cyclosporine treatment. Clin Invest Med. 1988;11(2):108–12.
Summary of product characteristics: AFINITOR (everolimus). http://www.ema.europa.eu/docs/en_GB/document_library/EPAR_-_Product_Information/human/001038/WC500022814.pdf. Accessed 24 Aug 2018.
Kovarik JM, Hartmann S, Figueiredo J, Rouilly M, Port A, Rordorf C. Effect of rifampin on apparent clearance of everolimus. Ann Pharmacother. 2002;36(6):981–5.
Tayrouz Y, Ganssmann B, Ding R, Klingmann A, Aderjan R, Burhenne J, et al. Ritonavir increases loperamide plasma concentrations without evidence for P-glycoprotein involvement. Clin Pharmacol Ther. 2001;70(5):405–14.
Bui K, Zhou D, Sostek M, She F, Al-Huniti N. Effects of CYP3A modulators on the pharmacokinetics of naloxegol. J Clin Pharmacol. 2016;56(8):1019–27.
Tortorici MA, Matschke K, Korth-Bradley JM, DiLea C, Lasseter KC. The effect of rifampin on the pharmacokinetics of sirolimus in healthy volunteers. Clin Pharmacol Drug Dev. 2014;3(1):51–6.
O’Leary JG, McKenna GJ, Klintmalm GB, Davis GL. Effect of telaprevir on the pharmacokinetics of sirolimus in liver transplant recipients. Liver Transpl. 2013;19(4):463–5.
Mai I, Stormer E, Bauer S, Kruger H, Budde K, Roots I. Impact of St John’s wort treatment on the pharmacokinetics of tacrolimus and mycophenolic acid in renal transplant patients. Nephrol Dial Transpl. 2003;18(4):819–22.
von Richter O, Burk O, Fromm MF, Thon KP, Eichelbaum M, Kivisto KT. Cytochrome P450 3A4 and P-glycoprotein expression in human small intestinal enterocytes and hepatocytes: a comparative analysis in paired tissue specimens. Clin Pharmacol Ther. 2004;75(3):172–83.
Yang J, Tucker GT, Rostami-Hodjegan A. Cytochrome P450 3A expression and activity in the human small intestine. Clin Pharmacol Ther. 2004;76(4):391.
Tse FL, Ballard F, Skinn J. Estimating the fraction reabsorbed in drugs undergoing enterohepatic circulation. J Pharmacokinet Biopharm. 1982;10(4):455–61.
Kawai R, Mathew D, Tanaka C, Rowland M. Physiologically based pharmacokinetics of cyclosporine A: extension to tissue distribution kinetics in rats and scale-up to human. J Pharmacol Exp Ther. 1998;287(2):457–68.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
No external funding was used in the conduct of this study.
Conflict of Interest
Michel Tod, Nathalie Bleyzac, Sylvain Goutelle, and Laurent Bourguignon do not have any potential conflicts of interest that might be relevant to the content of this article.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Appendices
Appendix A: Derivation of the Model Equations
We need to derive the expression of (1) the concentration of the victim drug in blood; (2) its value when the interactor is co-administered; and (3) the metrics of the interaction: the AUC ratio and the ratio of renal clearances.
1.1 Model with No Interaction (Substrate Alone)
The full model (Fig. 4) is the starting point of our approach. It is not the model ultimately used, but it allows understanding of the derivation of the final model. Because we are only interested by the impact of drug interaction on substrate clearance, the non-eliminating organs need not be considered in the model. In the gut and the liver compartment, the upper compartment represents blood, while the lower compartment represents the intracellular fluid. In compartment A (the gut lumen), the drug may be passively absorbed (with a permeability surface produt PSAB) or eliminated by lumen fluid flow in the feces, with a flow rate CLA. The drug is absorbed in the intracellular space of the gut (compartment B), where it may diffuse passively in the blood (PSBC = PSCB) or undergo metabolism by CYP3A4 (with an intrinsic clearance CLig-cyp) or efflux in the lumen (CLig-pgp). The drug in the gut blood (compartment C) flows to the liver blood (compartment D) where it may diffuse in the intracellular space of hepatocytes (compartment E, PSh). In this study, PSh is a passive process and the permeability of the drug is assumed to be high. Hence, the unbound drug concentrations in compartments D and E are considered to be equal, and compartment E is ignored in the equations.
Full model. CLA clearance in compartment A, CLiEH intrinsic clearance of the enterohepatic cycle, CLig-cyp intrinsic clearance of the drug in the gut due to cytochromes, CLig-other intrinsic clearance of the gut due to other metabolism, CLig-pgp intrinsic clearance of the gut due to P-glycoprotein metabolism, CLih-cyp intrinsic clearance of the drug in the liver by cytochromes, CLih-other intrinsic clearance of the drug in the liver by other metabolism, CLr renal clearance, fub unbound drug concentration in blood, Qg blood flow rate in gut blood, Qh blood flow rate in liver blood, PSAB permeability surface product in compartments A and B, PSBC permeability surface product in compartments B and C, PSh permeability surface product in the intercellular space of hepatocytes, R dosing rate
The drug in the liver may undergo metabolism by CYP3A4 (CLih-cyp) and an enterohepatic cycle with an intrinsic clearance CLiEH that is the sum of CLih-pgp (the intrinsic clearance for efflux in bile mediated by P-gp) and CLih-mr (the intrinsic clearance for metabolites that may be recycled, e.g., glucuronides, plus efflux by other transporters).
The total intrinsic clearance in the liver is as follows:
CLih = CLih-pgp + CLih-mr + CLih-cyp + CLih-other.
The drug that escapes the liver reaches the systemic circulation (compartment F) where it may be eliminated by renal route (with renal clearance CLr).
This model is not useful in practice for our purpose because it contains too many parameters, some of which cannot be easily measured or calculated. A more useful model is obtained by splitting the full model into two submodels: the absorption model and the clearance model.
The absorption model represents the lumen and the gut. Because we are not interested in the pharmacokinetics of the drug, but rather only in the steady-state concentrations, the concentration in the lumen and in the intracellular space of the gut need not be distinguished. The enterohepatic cycle will be considered in the clearance model and does not appear in the absorption model. Therefore, compartment A and B are lumped together and the absorption model is represented by Fig. 1a.
From compartment 1, the drug may disappear with an apparent clearance CLg, which is due to CYP3A4 metabolism (CLg-cyp), elimination by the lumen flow of the drug that has been effluxed and not reabsorbed (CLg-pgp), and elimination by the lumen flow before absorption (CLg-other). The passive diffusion between compartment 1 and 2 (PSg) is actually a tuning parameter that controls the residence time of the drug in compartment 1. A slow PSg results in a long residence time in compartment 1, allowing more drug to be eliminated before reaching the liver, for a given CLg value. The output rate of the drug from compartment 2 is Qg.C2, which is by definition Fa.Fg.R, the rate of drug input to the liver that may be measured in a subject in whom the enterohepatic cycle has been interrupted by mechanical or pharmacological means. Note that the systemic blood returning to the gut (from compartment F to compartment C in the full model) that would lead to a clearance from the gut is neglected, because the amount of CYP3A4 and P-gp in the gut is 90- and 41-fold lower, respectively, than in the liver [147, 148].
The clearance model represents the liver and the systemic blood compartment (Fig. 1b).
The fraction of drug that undergoes an enterohepatic cycle goes into the bile compartment, where it returns eventually in the liver after reabsorption in the gut. The corresponding rate of input is Fa.Fg.CLiEH.C5. If the recycling is not fully efficient (e.g., because the glucuronides are not completely hydrolyzed in the lumen), an additional parameter must be introduced, but its value has to be fixed.
The model corresponding to Fig. 1a, b is described as a set of five differential equations [Eqs. (5)–(9)]. Since the system is considered at steady state, each equation is equal to zero:
Equation (9) implies that, at steady state, C5 = fub.C3. Hence, the bile compartment may be omitted in the system provided that Eq. (7) is rewritten as follows:
The system of equations may be further simplified by deriving an expression for FaFg.
Rearranging Eq. (6) yields:
Let’s define the apparent PSg as follows:
Using this definition, the concentration C2 is expressed as follows:
The sum of Eqs. (5) and (6) yields:
Replacing C1 with its expression and rearranging, we get the following:
The output rate of the drug escaping from the system is Qg.C2. By definition the product FaFg is as follows:
Substituting R with Eq. (14) yields:
With this expression, Eqs. (5) and (6) are no longer needed to compute C3 and C4.
The system of equations reduces to its final form:
This system is solved in matrix form by matrix inversion to obtain (C3,C4).
In Eqs. (18)–(20), the drug is assumed to be administered at a constant rate R and C4 is the steady-state concentration. In case of repeated administration by oral route at intervals (τ), C4 would be the time-average concentration, the ratio of AUC from time zero to τ on τ. Because the pharmacokinetics of the victim drug are assumed to be linear, AUCτ is equivalent to the AUC from time zero to infinity (AUC∞) after the first dose. Hence, Eqs. (18)–(20) may be used to characterize the AUC after single or repeated dosing.
1.2 Model with Interaction (Substrate + Interactor)
The DDI is assumed to alter the efflux by P-gp (in the gut, liver, and kidney) and the metabolism by cytochromes (in the gut and liver).
To describe the DDI, we define the CR of each pathway that may be impacted by the interaction as follows:
where each CR may range from 0 to 1. In addition, the sum of CRs in the gut and the sum of CRs in the liver must both be ≤ 1.
The characteristic parameters of the interactor are defined as follows:
IXg-cyp and IXh-cyp are the inhibition or induction potency of the interactor on CYP metabolism in the gut and the liver, respectively.
IXg-pgp, IXh-pgp, and IXr-pgp are the inhibition or induction potency of the interactor on the efflux transporter in the gut, liver, and kidney, respectively.
For inhibitors, each IX may range from 0 (no inhibition) to – 1 (complete inhibition). For inducers, each IX may range from 0 (no induction) to ∞ in theory. In practice, IX estimates of the most potent inducers have been ≤ 10.
Using these definitions, the expression of the parameters impacted by the interaction may be derived. The superscript * denotes the parameters whose value is changed by the interaction.
In case of interaction, the value of CLg becomes CLg*, which is expressed as follows:
which becomes the following after simplification:
And:
which becomes the following after simplification:
The value of CLih becomes CLih*, which is expressed as follows:
while
with
The value of CLr becomes CLr*:
The value of the substrate concentration in blood in case of interaction, C4*, is calculated by solving the system of Eqs. (18)–(20) with CLg*, CLih*, CLEH*, and CLr* replacing CLg, CLih, CLEH, and CLr, respectively. The predicted AUC ratio is therefore as follows:
But, as explained in the following section, the AUC ratio is not obtained exactly in this way.
1.3 Calculation of the Substrate Parameters and the Area Under the Plasma Concentration–Time Curve (AUC) Ratio
The goal is to estimate the CRs (other than CRh-mr) and the IXs by regression, given the data (the AUC ratios).
For a given substrate, the values of oral clearance from plasma in the control group (CLoralp), bioavailability (F), CLr, fub, blood-to-plasma concentration ratio (BPR), Qh, and CRh-mr are common to all interactions with this substrate. These values are extracted or imputed from the literature.
When there is an enterohepatic cycle, it is necessary to distinguish the ‘effective’ fraction of dose absorbed are oral administration, which will be denoted F+, from the fraction of dose absorbed when the enterohepatic cycle is suppressed by pharmacological or mechanical means, FnoEHC. Following Tse et al. [149], the relationship between F+ and FnoEHC may be derived as follows:
where Eb, the coefficient of extraction of the drug to the bile, is approximated by (CRh-pgp + CRh-mr). For our purpose, the values of the pharmacokinetic parameters without an enterohepatic cycle need to be derived in order to calculate the impact of the interaction properly.
The rest of the parameters are therefore calculated as follows:
where CLr is renal clearance from blood.
where Qh is set to 89 L/h [150].
Using the current estimate of CRh-pgp:
Using the current estimates of CRs and IXs, FaFg*, CLih*, CLEH*, and CLr* are calculated according to Eqs. (27)–(30), allowing C4* to be calculated by solving Eqs. (18)–(20). In the computations, the dosing rate R is fixed to an arbitrary value (e.g., R = 100). The output of the model is expressed as CLoral = R/C4 and CLoral* = R/C4*. The AUC ratio is CLoral/CLoral*. Assuming that the BPR of the victim drug concentration is unchanged by the interactor, the ratio of blood clearances is equal to the ratio of clearances from plasma, because the BPR cancels each of these clearances out in the ratio.
1.4 Calculation of the Inhibition or Induction Potencies (IXs) of Interactors with Midazolam and Alprazolam as Substrates
Midazolam and alprazolam are not effluxed by P-gp. Hence, all parameters pertaining to the efflux may be removed from the model. After rearrangement, the following expression for the AUC ratio is obtained:
where the CRh-cyp values are known from our previous work [35] and CRg-cyp values are fixed to 0.9 because no other metabolic pathways are known in the intestine for these substrates. If the AUC ratios of the interaction of midazolam and alprazolam are known for an interactor, then we have two equations with two unknowns (IXg-cyp and IXh-cyp). The solution may be obtained by root finding or regression. The values of IXg-cyp and IXh-cyp were obtained in this way for carbamazepine, ketoconazole, itraconazole, erythromycin, fluvoxamine, rifampicin, ritonavir, and St John’s Wort. These values are weakly sensitive to the value assumed for CRg-cyp (i.e., 0.9).
1.5 Calculation of the IXs of Interactors with Aliskiren as Substrate
The interactions with aliskiren are assumed to arise solely by P-gp inhibition in the gut. Therefore:
where CRg-pgp = 0.87. Hence, the IXg-pgp values of amlodipine, atorvastatin, cyclosporine, ketoconazole, and verapamil were fixed by solving Eq. (41).
Appendix B: Objective Function
The objective function to be minimized with respect to the parameters is finally as follows:
where \(\hat{P}\) are the current estimates of CRs and IXs, Rpred and Robs are the predicted and observed AUC and CLr ratios (i.e., AUC*/AUC and CLr*/CLr), ns is the number of observations (number of AUC ratio and number of CLr ratio), np is the number of parameters to be estimated, and pen is the value of the penalty if some constraints are violated. The expected standard error of Robs is calculated according to our previous work [25].
The constraints and the penalties associated with violation of the constraints are as follows:For each substrate:
For each inhibitor:
For each inducer:
The expected standard error of Robs is calculated as follows.When the AUC ratio is > 1:
When the AUC ratio is < 1:
where nsuj is the number of subjects in the clinical study from which the geometric mean ratio \({\bar{\text{R}}\text{obs}}\) was calculated. These equations reflect the larger uncertainty associated with \({\bar{\text{R}}\text{obs}}\) values far from unity.
A confidence interval for the parameter estimates was obtained by a parametric bootstrap of the observations. Based on these equations for SE(Ln Robs), a synthetic dataset is generated by sampling Ln Robs values in a normal distribution with mean Ln Robs and standard deviation SE(Ln Robs). The parameters are estimated with this synthetic dataset. This procedure is repeated 30 times. The final point estimate of each parameter and its standard error are taken as the mean and standard deviation of the 30 estimates. A higher number of replicates did not result in significant variations of the standard deviation.
Rights and permissions
About this article
Cite this article
Tod, M., Goutelle, S., Bleyzac, N. et al. A Generic Model for Quantitative Prediction of Interactions Mediated by Efflux Transporters and Cytochromes: Application to P-Glycoprotein and Cytochrome 3A4. Clin Pharmacokinet 58, 503–523 (2019). https://doi.org/10.1007/s40262-018-0711-0
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40262-018-0711-0