Abstract
A set of 81 new microsatellite markers for Carica papaya L. previously identified by data mining using freely available sequence information from Genbank were tested for polymorphism using 30 germplasm accessions from the Papaya Germplasm Bank (PGM) at Embrapa Mandioca e Fruticultura Tropical (CNPMF) and 18 landraces. The data were used to estimate pairwise genetic distances between the genotypes. A neighbor-joining based dendrogram was used to define clusters and infer possible genetic structuring of the collection. Most microsatellites were polymorphic (73%), with an observed number of alleles per locus ranging from one to eleven. The levels of observed and expected heterozygosity for 51 polymorphic loci varied from 0.00 to 0.85 and from 0.08 to 0.82, averaging 0.19 and 0.59, respectively. Forty-four percent of microsatellites showed polymorphism information content (PIC) higher than 0.50. The compound microsatellites seem to be more informative than dinucleotide and trinucleotide repeats in average alleles per locus and PIC. Among dinucleotides, AG/TC or GA/CT repeat motifs exhibited more informativeness than TA/AT, GT/CA and TG/AC repeat motifs. The neighbor-joining analysis based on shared allele distance could differentiate all the papaya accessions and landraces as well as differences in their genetic structure. This set of markers will be useful for examining parentage, inbreeding and population structure in papaya.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genus Carica is monotypic and only includes papaya (Carica papaya L.). Papaya belongs to the small family Caricaceae which consists of six genera and 35 species (Badillo 2000). Most of them are diploid (2n = 18) and a relatively small genome of 372 Mb was found in C. papaya (Arumuganathan and Earle 1991; Parasnis et al. 1999; Kim et al. 2002; Liu et al. 2004). There are three distinct types of C. papaya plants: (1) dioecious papayas have male and female flowers on separate trees, (2) gynodioecious papayas bare female flowers on some trees and bisexual (hermaphrodite) flowers on others, and (3) trioiceous papayas have male, female, and hermaphrodite flowers in different plants.
Many landraces and cultivars present hermaphrodite plants, bearing perfect flowers and producing fruits shaped from long-cylindrical to ellipsoidal, which are preferred for commercial production. The economic importance of papaya resides largely in the fruit production, and Brazil is the world's major producer (FAOSTAT 2007).
Papaya exhibits considerable phenotypic variation for many morphological and horticultural traits, including fruit size, fruit shape, flesh color, flavor and sweetness, length of juvenile period, plant stature, stamen carpellody, and carpel abortion (Kim et al. 2002; Ocampo Pérez et al. 2006). Nevertheless, low levels of genetic variation for resistance to major fungal and virus diseases were observed in Carica genus (Nishijima 1994). On the other hand, resistance to several diseases which affect papaya are identified in the Vasconcellea genepool, including resistance to PRSV-P (V. cundinamarcensis, V. candicans, V. stipulata, V. cauliflora e V. quercifolia; Horovitz and Jimenez 1967; Alvizo and Rojkind 1987), Asperisporium Caricae (V. cundinamarcensis; Drew et al. 1998); Fusarium and Meloidogyne (V. weberbaueri; Scheldeman et al. 2003), phytoplasma (V. parviflora; Drew et al. 1998), and Phytophthora (V. goudotiana; Drew et al. 1998).
In recent years, molecular markers have been widely used in practical plant breeding to access the genetic variability available in germplasm banks, manage and develop core collections, target crosses, classify germplasm into interest groups and identify duplicate accessions (Rafalski and Tingey 1993; Manifesto et al. 2001; Martins-Lopes et al. 2007). Additionally, available molecular markers linked to important genes have proved to be useful for early selection of different desirable traits (Deputy et al. 2002; Dillon et al. 2005). These molecular markers have also been used to develop genetic maps, and for analysis of qualitative and quantitative inheritance (Luo et al. 2001; Jansen 2005; Milczarski et al. 2007). Other applications, such as marker-assisted backcrossing can reduce the number of generations needed to obtain a genotype with 98% or 99% genetic similarity to the recurrent parent for a fixed sample size (Liang et al. 2004; Oliveira et al. 2005). The first study of genetic diversity in papaya was made by Tan and Weinheimer (1976) using isozyme. Genetic relationships between papaya and related wild species have been investigated using isozyme (Morshidi 1998), Random Amplification of Polymorphic DNA (RAPD) (Sondur et al. 1996; Stiles et al. 1993; Jobin-Décor et al. 1997), Restriction Fragment Length Polymorphism (RFLP) (Aradhya et al. 1999), Amplified Fragment Length Polymorphism (AFLP) markers (Kim et al. 2002; Van Droogenbroeck et al. 2002), PCR-RFLP (Van Droogenbroeck et al. 2004), SSR (Kyndt et al. 2005; Kyndt et al. 2006) as well as Inter-Simple Sequence Repeats (ISSR) (Carrasco et al. 2009).
A large number of different molecular techniques are at present available and each of them differs in its informational content. Although in principle all types of markers would be suitable for our purpose, microsatellites (Single Simple Repeat (SSR)) are especially useful for diversity studies (Baumung et al. 2004).
Multi-locus approaches, such as RAPD, AFLP, and ISSR, may be convenient but have some technical and analytical drawbacks, such as dominance. Multi-locus data are typically analyzed as pairwise comparison of complex patterns that only have meaning relative to others in the same study, thus results are to a limited extent comparable among studies. By contrast, single-locus markers, such as SSR, are usually characterized by co-dominance and thus are more flexible and supply more robust and comparable data (Brondani et al. 1998; Rallo et al. 2000; Karp 2002).
Despite the present availability of molecular marker techniques, papaya have just recently received molecular attention, especially by development of a large number of SSR markers with different motifs, by data mining analysis from DNA databases (Oliveira et al. 2008a) and by screening of sequence data from bacterial artificial chromosomes (BAC) ends and complementary DNA (Eustice et al. 2008). The utility of SSR as genetic markers to investigate relationships among plants has been clearly established (Zhou et al. 2003). Microsatellites have also been extensively exploited for fingerprinting, phylogenetic studies, genetic and QTL mapping for a wide range of species. The genetic analysis based on SSR made it possible to investigate the occurrence and variability of simple sequence repeats at the whole genome level in germplasm accessions of papaya. The objective of this investigation was to test the suitability of SSR for genomic analysis in C. papaya.
Material and Methods
Plant Material
Thirty papaya accessions and eighteen landraces collected from Muritiba, Bahia-Brazil, were used to screen for SSR polymorphisms (Table 1). These papaya plants were maintained in the Papaya Germplasm Bank (PGM) at Embrapa Mandioca e Fruticultura Tropical (CNPMF), in Cruz das Almas, BA, Brazil. Among the 30 papaya accessions, two were cultivars, 12 were improved (but not released) breeding lines, and 16 were unimproved germplasm. All landraces were unimproved germplasm (Table 1).
DNA Extraction
Young papaya leaves were harvested and stored at −80°C for long-term storage. Genomic DNA was extracted based on the procedure described by Doyle and Doyle (1990). DNA quantification was carried out in an agarose gel (1.0% w/v) by comparing the fluorescent intensity of the sample stained with ethidium bromide (1.0 mg/mL), relative to a dilution series of Lambda DNA (Invitrogen, Carlsbad, CA) as standard of known concentration.
PCR Amplification
A set of 100 SSR primers developed by Oliveira et al. (2008a) was tested for amplification and polymorphism. Each PCR reaction was prepared as follows: 20 ng DNA template, 20 mM Tris-HCl (pH 8.4), 50 mM KCl, 0.3 mM of each primer, 1.5 mM MgCl2, 0.2 mM dNTPs, and 0.5 U Taq DNA Polymerase (Invitrogen, Carlsbad, CA) in a total volume of 20 µL.
PCR cycling consisted of 94°C for 4 min, followed by 35 cycles of 94°C for 40 s, (annealing temperature of 55°C, 60°C, or 62°C according to each SSR primer (Table 2)) for 40 s, and 72°C for 1 min, with a final extension at 72°C for 2 min, on a PTC-100 thermal cycler (MJ Research, Inc., Watertown, MA). After cycling, fragments with size difference shorter than 10 base pairs (bp) were electrophoresed on a 6% (w/v) denaturing polyacrylamide gel in a Hoefer SQ3 DNA sequencer gel electrophoresis unit (Pharmacia Biotech Inc., San Francisco, CA) at 70 W for 2.5 h. The gels were stained with silver nitrate, according to Creste et al. (2001). Previous analysis in polyacrylamide gel showed some primers that produces fragments longer than 10 bp in size difference. These loci were electrophoresed on a 3% agarose 1,000 gel (Invitrogen, Carlsbad, CA) at 130 V for 3.5 h. The 50-bp ladder (New England Biolabs, Inc., Beverly, MA) was used as a molecular weight standard to estimate the size of microsatellite alleles.
Data Analysis
Genetic variability was measured as allelic richness determined by the total number of the detected alleles and the number of alleles per locus (N A ), observed heterozygosity (H O ), expected heterozygosity (H E ) and polymorphism information content (PIC). We defined rare alleles as those whose individual frequency is lower than 1% in the investigated materials. Common alleles are those that occur with a frequency between 1% to 20%, while those whose frequency is higher than 20% are classified as most frequent alleles. Genetic distances between individuals were estimated by shared allele distance (SAD). All these analysis were carried out using the software POWERMARKER version 3.25 (Liu and Muse 2005). The matrix of genetic distance was used to construct the neighbor-joining tree, using the MEGA 4.1 package (Tamura et al. 2007). To assess confidence in the nodes of a tree, bootstrap values were obtained from 1,000 replicates by re-sampling microsatellite loci.
Results
Polymorphism of Microsatellites
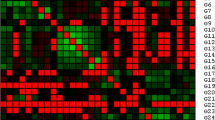
All 100 SSR loci were screened for patterns of amplification using two individuals of the PGM-CNPMF. Eighty-one primer pairs were selected according to strength, clarity of banding patterns, and successfully amplified PCR product of high quality, besides the polymorphic amplification product in the expected size. Characteristics of the 81 primer pairs and optimal conditions for their amplification are given in Table 2. Subsequently, all primer pairs that amplified a specific band were used for the genotyping of 30 germplasm accessions and 18 landraces. In total, 59 primer pairs amplified a polymorphic and easily scorable PCR product, while 22 pairs amplified a monomorphic one.
Considering the 59 SSR loci analyzed in the present study and a total of 48 genotyped individuals obtained from partial outcrossing and selfing germplasm, the SSR markers detected a total of 237 alleles. The least and the most variable loci displayed 2 (CP06, CP09, CP11, CP20, CP22, CP23, CP24, CP27, CP28, CP29, CP33, CP36 and CP38) and 11 (CP16) alleles, respectively (Table 3). The average allele number per locus was 4.02 (monomorphic loci excluded). Table 3 summarizes the locus specific descriptive statistics for the 59 SSR markers.
Expected heterozygosity was nominally larger than the observed heterozygosity under HWE for all loci, except on CP31 and CP48. Only loci CP21, CP33, and CP38 were found to be in HWE. The observed heterozygosity ranged from 0.00 (CP09, CP14, CP22, CP27, CP47, CP52, CP55, CP57, CP59, CP63, CP68, CP69, CP72, CP73, CP80, CP83, CP89, CP94, CP95, CP97, and CP100) to 0.85 (CP31), and gene diversity (H E ) from 0.08 (CP22) to 0.82 (CP16). The CP16 locus showed the highest polymorphism information content (PIC = 0.81) and the CP22 locus the lowest (PIC = 0.08), average PIC being 0.53.
When looking at SSR classes and motifs, the compound SSRs showed higher allele numbers (average 4.27 per locus) and PIC values (average 0.55 per marker) followed by dinucleotide (average alleles, 4.08 per locus; PIC value average 0.52 per marker) and trinucleotide SSRs (average alleles, 3.2 per locus; PIC value, average 0.47 per marker). Among dinucleotide SSRs, AG/TC or GA/CT repeat motifs exhibited more informativeness (average alleles, 4.56 per locus and PIC value, average 0.59 per marker) as compared with TA/AT repeat motifs (average alleles, 4.0 per locus and PIC value, average 0.50 per marker) and to GT/CA or TG/AC repeat motifs (average alleles, 3.5 per locus and PIC value, average 0.52 per marker) (Table 3).
Allelic Composition
The allelic composition revealed that rare alleles were represented by < 1.3% of the total number of alleles detected. Of the 237 alleles detected, three were rare, 115 common, and 119 most frequent alleles. Rare alleles were detected from CP16, CP30, and CP53. Common alleles were detected at 46 SSR loci, with an average of 1.95 alleles. In contrast, all SSR loci detected 1 to 3 most frequent alleles in the individuals (Table 3).
This study detected unique alleles within groups (Table 3). The germplasm accessions contained the largest number of unique alleles (46), while 14 unique alleles differentiated landraces from the papaya germplasm. Of the total alleles detected, papaya germplasm and landraces shared 74.7% of alleles (177).
Genetic Diversity
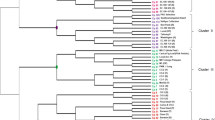
Based on the unique DNA fingerprint profiles of each genotype obtained by the polymorphic markers, a dendrogram was constructed to understand the relationships among the germplasm and landraces surveyed. The neighbor-joining cluster analysis based on shared-allele distance could successfully differentiate all the papaya accessions (group A) and landraces (group B) included in this study (Fig. 1) with the high bootstrap value of 95%. Clearly, four subgroups could be seen in germplasm accessions while landraces presented just two.
Neighbor-joining tree based on the shared allele distance of 59 SSR markers using 30 germplasm accessions at PGM-CNPMF (code CMF) and 18 landraces (code M). A1, A2, A3, and A4 are subgroups of the A group clustered by germplasm accessions; B1 and B2 are subgroups of B group that contained only papaya landraces. Distance bootstrap values are given together with subgroups in parenthesis
Subgroup A1 comprised all gynodioecious accessions, being one cultivar (CM128), eight improved germplasm (CMF068, CMF082, CMF108, CMF115, CMF125, CMF138, CMF142 and CMF143) and five unimproved germplasm (CMF054, CMF058, CMF101, CMF102 and CMF129). Subgroup A2 had two improved (CMF008 and CMF038) and three unimproved germplasm (CMF011, CMF017 and CMF188), whereas subgroup A3 was formed by one improved (CMF123) and six unimproved germplasm (CMF134, CMF135, CMF157, CMF165, CMF189 and CMF191). The other cultivars analyzed (CMF024) were clustered with three germplasm accessions in subgroup A4. Subgroups A1, A2, A3 and A4 were clustered with a relatively low bootstrap support value (36%, 25%, 28%, and 30%, respectively), but subgroups B1 and B2, had a moderate bootstrap support value (48% and 38%, respectively).
The genotypes belonging to landraces collected in the State of Bahia (Brazil) were grouped in clusters (B1and B2), with 14 and four individuals, respectively. No specific grouping was observed for the improved and unimproved cultivars, which dispersed in all subgroups.
Discussion
Microsatellite Informativeness
SSR markers have been extensively used for DNA fingerprinting and elucidating genetic relationships within plant species (Paniego et al. 2002; Stajner et al. 2005). The ability to distinguish between closely related genotypes is a function of the high heterozygosity values of SSR markers. The set of SSR markers characterized in this study proved to be useful for a broad genetic analysis of C. papaya.
Previous studies have shown that papaya contains abundant SSRs (Santos et al. 2003; Ocampo Pérez et al. 2006; Eustice et al. 2008; Oliveira et al. 2008a), but the characterization of SSR markers in C. papaya germplasm is not yet well studied. Here, we observed relatively high levels of multiallelism at all 59 SSR loci analyzed. Mean number of alleles per locus (4.02, for 59 loci) and expected heterozygosity (mean of 0.59) were higher than those reported by Ocampo Pérez et al. (2006) who used 26 polymorphic markers, and observed 3.8 alleles per locus and H O and H E values of 0.42 and 0.57, respectively, but intermediate for samples collected in Guadeloupe, Venezuela, Colombia, Barbados and Costa Rica, that showed 6.6 alleles per locus for 15 loci, and H E values between 0.37 and 0.69 (Ocampo Pérez et al. 2007).
The number of alleles per locus reported in our study is most likely a minimum value. The lower number of alleles can be due to the relatively few samples analyzed and to the different mating systems of these genotypes. Half of the accessions were gynodioecious, which is almost exclusively inbreeding and tends to decrease the allele number per locus. The number of alleles should increase when all germplasm accessions of the PGM of CNPMF were sampled.
Except CP11 and CP40, all SSR described here are derived from a genomic sequence and showed considerable polymorphism. This is in agreement with Eustice et al. (2008), who found a relatively high level of polymorphism in the genomic rather than the genic region, using seven papaya accessions (SunUp, Kapoho, 2H94, UH918, Kaek Dum, UH928, and AU9) to screen SSR polymorphisms. As in other crops, the selective pressures from breeding can significantly reduce genetic diversity in the target genes or genic regions, while levels of genetic diversity in the genomic region remain high.
The broad range of observed heterozygosity (0.00 to 0.85) and expected heterozygosity (0.08 to 0.82) result from the broad variation in number of alleles per locus and allele frequency distribution within genotypes. The obtained zero values of H O at twenty-one loci could be explained by the low number of alleles at these loci (2 to 6 alleles) as well as their combination in homozygous state. Loci with smaller numbers of alleles or with a skewed frequency distribution such as CP06, CP09, CP11, CP20, CP22, CP23, CP24, CP27, CP28, CP29, CP33, CP36, and CP38, tend to have lower heterozygosity values and, consequently, lower probability of paternity exclusion when studying natural population.
Ninety-five percent of the SSR markers surveyed for heterozygosity (56 of 59) deviated significantly from Hardy–Weinberg equilibrium. In all cases, the deviation was in the direction of reduced heterozygosity due to inbreeding, presence of null alleles, natural or artificial selection favoring homozygosity for particular loci and population bottlenecks. Such result has been commonly observed in surveys of other species (Carrasco et al. 2009). A possible reason for high levels of inbreeding in C. papaya is the elimination of the male plants in trioic accessions (male, female, and hermaphrodite plants) and self-compatibility that stimulates crossing among related individuals and increases the degree of selfing.
In relation to polymorphism, microsatellites of papaya showed enrichment for di and trinucleotide SSR repeat motifs (Eustice et al. 2008; Oliveira et al. 2008a). Overall, AT/TA is the predominant dinucleotide motif, and AAT/TTA is the predominant trinucleotide motif. The previous surveys carried out on microsatellite abundance in plant genomes have shown AT as the most frequently occurring dinucleotide repeat motif followed by AG/TC and GT/CA (Condit and Hubbel 1991; Powell et al. 1996; Yonemaru et al. 2009). Compound SSRs, mainly from different dinucleotide repeats, and dinucleotide repeats were more informative (more average alleles per locus and PIC). Among dinucleotides, although AT/TA-rich motif is prominent, AG/TC or GA/CT repeat motifs exhibited more informativeness. The same pattern was observed in other species (Ferguson et al. 2004; Moretzsohn et al. 2005).
Diversity Structuring
Our study based on SSR analysis clearly revealed genetic diversity in C. papaya germplasm samples and some landraces cultivated by farmers in the State of Bahia. Neighbor-joining tree broadly separated germplasm accessions from landraces with high bootstrap support (95%). However, clusters from each subgroup were poorly supported by bootstrap values (Fig. 1). A low bootstrap value means that a grouping is sensitive to the combinations of genotypes that are evaluated, implying that more data may alter the grouping.
The main cluster (group A) is composed of germplasm accessions of C. papaya subdivided in four smaller clusters. The smaller one (group B) is clustered only by landraces, which have two subgroups. The grouping of genotypes representing landraces in different clusters is in agreement with their origin, and is important as a first indicative of the genetic background of most of the germplasm of PGM at CNPMF.
The genetic variability in landraces grown by small farmers in the region of Recôncavo of Bahia, where the landraces were obtained, was maintained over generations. This variability is quite different from the one in the PGM, because papaya accessions from the Recôncavo of Bahia has not been deposited in the germplasm bank. The additional genetic variability will be used in breeding programmes to enhance the diversity of breeding populations for selection gains in the future.
Ocampo Pérez et al. (2007) analysed genotypes from Costa Rica, Colombia, Venezuela, Guadeloupe and Antillean islands using SSR markers and PCO analysis, and found that, with few exceptions, they were clustered according to their geographic origin. In our study, although there are few accessions belonging to different countries, no high correlation between the clustering pattern and the geographical location was observed. The Brazilian accessions were clustered in all subgroups. The accessions from South Africa and Hawaii grouped into subgroup A1, Thailand into subgroup A3; whereas accessions from Costa Rica and Malaysia grouped into A2 and A4 subgroup; and Taiwan into A1 and A2 subgroup (Fig. 1).
The 30 germplasm accessions and 18 landraces used for characterization of SSR polymorphism allowed the detection of considerable genetic variability, as showed on six diversity subgroups (Fig. 1). The average similarity based on shared allele distance between accessions was 0.44, which is very similar to the value of 0.48 obtained by Ocampo Pérez et al. (2007), using Dice distance. Previous reports using dominant markers, as RAPD (Stiles et al. 1993) and AFLP (Kim et al. 2002), showed values of 0.78 (Jaccard distance) and 0.88 (Dice distance), respectively. So, the SSRs markers revealed a more important polymorphism than the other nuclear DNA markers used so far in papaya (Ocampo Pérez et al. 2007).
Most of the observed variation occurred among accessions. All genotypes present in A1 and A4 subgroup are gynodioecious, whereas B1 and B2 are dioecious. In addition, the A2 and A3 subgroups are composed of gynodioecious and dioecious genotypes. The gynodioecious tends to show lower genetic variability within accessions than the dioecious, because self-compatibility increases the inbreeding.
Many aspects, such as the breeding system, seed and pollen dispersal, plant longevity and agricultural practices influence the genetic diversity, including the proportion of variation distributed within and between populations (Hamrick and Godt 1996). Kim et al. (2002) analysed the genetic relationships among C. papaya cultivars, breeding lines, unimproved germplasm, and related species using AFLP markers and suggested limited genetic variation in papaya, with smaller genetic diversity within the same gene pools, such as gynodioecious and dioecious cultivars. According to Carrasco et al. (2009), the genetic diversity of Vasconcellea pubescens, a species of the Caricaceae family, was remarkably low using ISSR (Inter Single Simple Repeat). The major genetic diversity was found within groups (65%) when South and the North samples from Chile were analyzed together. In addition, samples from South and the North analyzed separately showed that 82% and 60% of genetic diversity were within groups, respectively.
Efforts are made to maintain the genetic diversity of papaya breeding through the stocks in germplasm banks to study natural variation. Although artificial by their nature and submitted to several forces, including genetic drift and man driven artificial selection, germplasm collections play an important role in the way the banks are structured. In this study, we found a great depth of allelic diversity among landraces and germplasm, because this diversity is not distributed randomly among the genotypes, but rather structured into two groups. Thus, many landraces will be incorporated to PGM at CNPMF, to increase the stored variability for future use in conservation and breeding programs.
SSR in Genetic Studies of C. papaya
SSRs are powerful molecular biology tools that have been used for a wide variety of applications. Although good estimators of population genetic parameters have been presented for dominant markers (Lynch and Milligan 1994; Zhivotovsky 1999), the codominant nature of SSR allows more precise calculations of heterozygosity, Hardy-Weinberg equilibrium, differentiation and gene flow (Lowe et al. 2004). Polymorphisms based on SSR are more powerful to estimate genetic parameters of populations and understand detailed patterns of gene flow and parentage composition (Dow and Ashley 1996; Collevatti et al. 1999; Ren et al. 2008). Besides, SSRs markers are more useful for genetic mapping (Oliveira et al. 2008b; Wang et al. 2009), and ancestry studies (Guilford et al. 1997; Gianfranceschi et al. 1998).
The papaya microsatellite loci published here provide an abundant set of genetic markers for detailed studies of population genetic structure, hybridization among populations, paternity, migration, phylogeography of the genus Carica, creation of linkage and physical maps, and location of genes of interest, particularly those functioning as QTL, encoding important agricultural traits such as disease resistance, yield, fruit type, and fruit size. Specifically, we are now using these markers to understand the genetic variability of 160 papaya germplasm accessions collected from different countries, and that are part of the PGM at CNPMF.
References
Alvizo VHF, Rojkind MC (1987) Resistencia al virus mancha anular del papaya en Carica cauliflora. Rev Mex Fitopat 5:61–62
Aradhya MK, Manshardt RM, Zee F, Morden CW (1999) A phylogenetic analysis of the genus Carica L (Caricaceae) based on restriction fragment length variation in a cpDNA intergenic spacer region. Genet Resour Crop Evol 46:579–586
Arumuganathan K, Earle ED (1991) Nuclear DNA content of some important plant species. Plant Mol Biol Rep 9:208–218
Badillo VM (2000) Carica L vs Vasconcella St Hil (Caricaceae): con la rehabilitación de este úlltimo. Ernstia 10:74–79
Baumung R, Simianer H, Hoffmann I (2004) Genetic diversity studies in farm animals - a survey. J Anim Breed Genet 121:361–373
Brondani RPV, Brondani C, Tarchini R, Grattapaglia D (1998) Development characterization and mapping of microsattelite markers in Eucalyptus grandis and E. urophylla. Theor Appl Genet 97:816–827
Carrasco B, Avila P, Perez-Diaz J, Muñoz P, Garcia R, Lavandero B, Zurita-Silva A, Retamales JB, Caligari PDS (2009) Genetic structure of highland papayas (Vasconcellea pubescens (Lenné et C Koch) Badillo) cultivated along a geographic gradient in Chile as revealed by Inter Simple Sequence Repeats (ISSR). Genet Resour Crop Evol 56:331–337
Collevatti RG, Brondani RV, Grattapaglia D (1999) Development and characterisation of microsatellite markers for genetic analysis of a Brazilian endangered tree species Caryocar brasiliense. Heredity 83:748–756
Condit R, Hubbel SP (1991) Abundance and DNA sequence of two-base repeat regions in tropical tree genomes. Genome 34:66–71
Creste S, Tulmann Neto A, Figueira A (2001) Detection of single sequence repeat polymorfisms in denaturing polyacrylamide sequencing gels by silver staining. Plant Mol Biol Rep 19:299–306
Deputy JC, Ming R, Ma H, Liu Z, Fitch MMM, Wang M, Manshardt R, Stiles JI (2002) Molecular markers for sex determination in papaya (Carica papaya L.). Theor Appl Genet 106:107–111
Dillon S, Ramage C, Drew R, Ashmore S (2005) Genetic mapping of a PRSV-P resistance gene in “highland papaya” based on inheritance of RAF markers. Euphytica 145:11–23
Dow BD, Ashley MV (1996) Microsatellite analysis of seed dispersal and sapling parentage in bur oak Quercus macrocarpa. Mol Ecol 5:615–627
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Drew RA, O’Brien CM, Magdalita PM (1998) Development of interspecific Carica hybrids. Acta Hortic 461:285–292
Eustice M, Yu Q, Lai CW, Hou S, Thimmapuram J, Liu L, Alam M, Moore PH, Presting GG, Ming R (2008) Development and application of microsatellite markers for genomic analysis of papaya. Tree Genet Genomes 4:333–341
Faostat: The United Nations: Food and Agriculture Organization of The United Nations 2007 Available via: http://faostatfaoorg/site/567/defaultaspx#ancor Accessed 04 June 2009
Ferguson ME, Burow MD, Schultz SR, Bramel PJ, Paterson AH, Kresovich S, Mitchell S (2004) Microsatellite identification and characterization in peanut (Arachis hypogaea L). Theor Appl Genet 108:1064–1070
Gianfranceschi L, Seglias N, Tarchini R, Komjanc M, Gessler C (1998) Simple sequence repeats for the genetic analysis of apple. Theor Appl Genet 96:1069–1076
Guilford P, Prakash S, Zhu JM, Rikkerink E, Gardiner S, Bassett H, Forster R (1997) Microsatellites in Malus × domestica (apple): abundance polymorphism and cultivar identification. Theor Appl Genet 94:249–254
Hamrick JL, Godt MJ (1996) Conservation genetics of endemic plant species. In: Avise JC, Hamrick JL (eds) Conservation genetics case histories from nature. Chapman and Hall, New York USA, pp 281–304
Horovitz S, Jiménez H (1967) Cruzamientos interespecificos e intergenericos en Caricaceas y sus implicaciones fitotecnicas. Agron Trop 17:323–344
Jansen J (2005) Construction of linkage maps in full-sib families of diploid outbreeding species by minimizing the number of recombinations in hidden inheritance vectors. Genetics 170:2013–2025
Jobin-Decor MP, Graham GC, Henry RJ, Drew RA (1997) RAPD and isozyme analysis of genetic relationships between Carica papaya and wild relatives. Genet Resour Crop Evol 44:471–477
Karp A (2002) The new genetic Era: will it help us in managing genetic diversity? In: Engels JMM, Ramanatha Rao V, Brown AHD, Jackson MT (eds) Managing plant genetic diversity. IPGRI, Rome, pp 43–56
Kim M, Moore P, Zee F, Fitch MMM, Steiger D, Manshardt R, Paull R, Drew R, Sekioka T, Ming R (2002) Genetic diversity of Carica papaya as revealed by AFLP markers. Genome 45:503–512
Kyndt T, Haegeman A, Van Glabeke S, Maertens I, Van Droogenbroeck B, Roldán-Ruiz I, Gheysen G (2005) Isolation and characterization of microsatellite loci in the highland papaya Vasconcelle × heilbornii V. Badillo (Caricaceae). Mol Ecol Notes 5:590–592
Kyndt T, Van Droogenbroeck B, Haegeman A, Roldán-Ruiz I, Gheysen G (2006) Cross-species microsatellite amplification in Vasconcellea and related genera and their use in germplasm classification. Genome 49:786–798
Liang F, Deng Q, Wang Y, Xiong Y, Jin D, Li J, Wang B (2004) Molecular marker-assisted selection for yield-enhancing genes in the progeny of “9311 × O rufipogon” using SSR. Euphytica 139:159–165
Liu K, Muse SV (2005) PowerMarker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21:2128–2129
Liu Z, Moore PH, Ma H, Ackerman CM, Makandar R, Yu Q, Pear HM, Kim MS, Charlton JW, Stiles JI, Zee FT, Paterson AH, Ming R (2004) A primitive Y chromosome in papaya marks incipient sex chromosome evolution. Nature 427:348–352
Lowe A, Harris S, Ashton P (2004) Ecological genetics: design analysis and application. Blackwell Publishing, Oxford
Luo ZW, Hackett CA, Bradshaw JE, Mcnicol JW, Milbourne D (2001) Construction of a genetic linkage map in tetraploid species using molecular markers. Genetics 157:1369–1385
Lynch M, Milligan BG (1994) Analysis of population genetic structure with RAPD markers. Mol Ecol 3:91–99
Manifesto M, Schlatter AR, Hopp HE, Suárez EY, Dubcovsky J (2001) Quantitative evaluation of genetic diversity in wheat germplasm using molecular markers. Crop Sci 41:682–690
Martins-Lopes P, Lima-Brito J, Gomes S, Meirinhos J, Santos L, Guedes-Pinto H (2007) RAPD and ISSR molecular markers in Olea europaea L: genetic variability and molecular cultivar identification. Genet Resour Crop Evol 54:117–128
Milczarski P, Banek-Tabor A, Lebiecka K, Stojaowski S, Mysków B, Masojc P (2007) New genetic map of rye composed of PCR-based molecular markers and its alignment with the reference map of the DS2 × RXL10 intercross. J Appl Genet 48:11–24
Moretzsohn MC, Leoi L, Proite K, Guimaraes PM, Leal-Bertioli SCM, Gimanes MA, Martin WS, Valls JFM, Grattapaglia D, Bertioli D (2005) A microsatellite-based gene-rich linkage map for the AA genome of Arachis (Fabaceae). Theor Appl Genet 111:1060–1071
Morshidi M (1998) Genetic control of isozymes in Carica papaya L. Theor Appl Genet 103:89–94
Nishijima W (1994) Papaya. In: Ploetz RC, Zentmyer GA, Nishijima WT, Rohrbach KG, Ohr HD (eds) Compendium of tropical fruit disease. American Phytopath Soc Press, St. Paul, Minnesota, pp 54–70
Ocampo Pérez J, Dambier D, Ollitrault P, d’Eeckenbrugge GC, Brottier P, Froelicher Y, Risterucci AM (2006) Microsatellite markers in Carica papaya L: isolation characterization and transferability to Vasconcellea species. Mol Ecol Notes 6:212–217
Ocampo Pérez J, d'Eeckenbrugge GC, Risterucci AM, Dambier D, Ollitrault P (2007) Papaya genetic diversity assessed with microsatellite markers in germplasm from the Caribbean Region. Acta Hortic 740:93–101
Oliveira EJ, Marin ALA, Borém A, Fagundes SA, Barros EG, Moreira MA (2005) Molecular-marker assisted selection for development of common bean lines resistant to angular leaf spot. Plant Breed 124:572–575
Oliveira EJ, Dantas JLL, Castellen MS, Machado MD (2008a) Identificação de microssatélites para o mamoeiro por meio da exploração do banco de dados de DNA. Rev Bras Frutic 30:841–845
Oliveira EJ, Vieira MLC, Garcia AAF, Munhoz CF, Margarido GRA, Consoli L, Matta FP, Moraes MC (2008b) An integrated molecular map of yellow passion fruit based on simultaneous maximum-likelihood estimation of linkage and linkage phases. J Am Soc Hortic Sci 133:35–41
Paniego N, Echaide M, Marianne M, Ferrandez L, Tolales S, Faccio P, Fuxan I, Carrera M, Zandomeni R, Suarez E, Hopp HE (2002) Microsatellite isolation and characterization in sunflower (Helianthus annuus L). Genome 45:34–43
Parasnis AS, Rama Krishna W, Chowdari KY, Gupta VS, Ranjekar PK (1999) Microsatellite (GATA)4 reveals sex-specic differences in papaya. Theor Appl Genet 99:1047–1052
Powell W, Machray GC, Provan J (1996) Polymorphism revealed by simple sequence repeats. Trends Plant Sci 1:215–222
Rafalski JA, Tingey SV (1993) Genetic diagnostics in plant breeding: RAPDs microsatellites and machines. Trends Genet 9:275–279
Rallo P, Dorado G, Martín A (2000) Development of simple sequence repeats (SSRs) in olive tree (Olea europaea L). Theor Appl Genet 101:984–989
Ren J, Lu L, Liu X, Tao Z, Zhang C, Wang D, Shen J, Liu W, Tian Y, Zhu Z (2008) Paternity assessment: application on estimation of breeding value in body-weight at first egg trait of egg-laying duck (Anas platyrhynchos). Mol Biol Rep. doi:10.1007/s11033-008-9432-z
Santos SC, Ruggiero C, Silva CLSP, Lemos EGM (2003) A microsatellite library for Carica papaya L cv Sunrise Solo. Rev Bras Frutic 25:263–267
Scheldeman X, Van Damme P, Urena Alvarez JV, Romero M (2003) Horticultural potential of Andean fruit crops exploring their centre of origin. Acta Hortic 598:97–102
Sondur SN, Manshardt RM, Stiles JI (1996) A genetic linkage map of papaya based on randomly amplified polymorphic DNA markers. Theor Appl Genet 93:547–553
Stajner N, Jakse J, Kozjak P, Javornik B (2005) The isolation and characterisation of microsatellites in hop (Humulus lupulus L). Plant Sci 168:213–221
Stiles JI, Lemme C, Sondur S, Morshidi MB, Manshardt R (1993) Using randomly amplified polymorphic DNA for evaluating genetic relationships among papaya cultivars. Theor Appl Genet 85:697–701
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) Software Version 4. Mol Biol Evol 24:1596–1599
Tan SC, Weinheimer EA (1976) The isoenzyme patterns of developing fruit and leaf of papaya (Carica papaya L). Sains Malays 5:7–14
Van Droogenbroeck B, Breyne P, Gotghebeur P, Romeijn-Peeters E, Kyndt T, Gheysen G (2002) AFLP analysis of genetic relationships among papaya and its wild relatives (Caricaceae) from Ecuador. Theor Appl Genet 105:289–297
Van Droogenbroeck B, Kyndt T, Maertens I, Romeijn-Peeters E, Scheldeman X, Romero-Motochi J, Van Damme P, Goetghebeur P, Gheysen G (2004) Phylogenetic analysis of the highland papayas (Vasconcellea) and allied genera (Caricaceae) using PCR-RFLP. Theor Appl Genet 108:1473–1486
Wang X, Wadl PA, Rinehart TA, ScheZer BE, Windham MT, Spiers JM, Johnson DH, Trigiano RN (2009) A linkage map for Xowering dogwood (Cornus Xorida L) based on microsatellite markers. Euphytica 165:165–175
Yonemaru J-I, Ando T, Mizubayashi T, Kasuga S, Matsumoto T, Yano M (2009) Development of genome-wide simple sequence repeat markers using whole-genome shotgun sequences of sorghum (Sorghum bicolor (L) Moench). DNA Res 16:187–193
Zhivotovsky LA (1999) Estimating population structure in diploids with multilocus dominant DNA markers. Mol Ecol 8:907–913
Zhou H-F, Xie Z-W, Ge S (2003) Microsatellite analysis of genetic diversity and population genetic structure of a wild rice (Oryza rufipogon Griff) in China. Theor Appl Genet 107:332–339
Acknowledgements
This work was supported by Embrapa Macroprograma 2, CNPq-Conselho Nacional de Desenvolvimento Científico e Tecnológico (National Counsel of Technological and Scientific Development) and FAPESB–Fundação de Amparo à Pesquisa do Estado da Bahia (Bahia State Research Funding Institution).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
de Oliveira, E.J., Amorim, V.B.O., Matos, E.L.S. et al. Polymorphism of Microsatellite Markers in Papaya (Carica papaya L.). Plant Mol Biol Rep 28, 519–530 (2010). https://doi.org/10.1007/s11105-010-0180-6
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-010-0180-6