Abstract
In Chile Vasconcellea pubescens is cropped to produce canned fruit, juice, jam and processed sweets. Additionally this species produces latex with a high level of papain, an important and valuable proteolytic enzyme with industrial applications. In this investigation seven ISSR primers were used to study the level and organization of genetic diversity in 333 samples of V. pubescens. Out of the 114 bands recorded, 63 proved to be polymorphic (P = 55.3%). At the species level, the genetic diversity was rather low (h = 0.01 ± 6,80188E-05, Shannon’s Index I = 0.16 ± 0,000148). The major portion of the genetic diversity was found within groups (65%). The genetic differentiation between the different groups was significant, as the AMOVA analysis suggested (Φpt = 0.35). When analysing the Northern area alone, the differentiation increased to Φpt = 0.40. When only the Southern area was analysed, Φpt decreased to 0.18, indicating greater genetic similarity among the samples. The results generated from Structure and Bayesian Analysis of Population Structure distinguished 8 genetically different groups, five of them located in the north and three in the south. The results are discussed in the light of the growers’ practices.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The papaya tree cultivated in Chile belongs to the Caricaceae, a small family of dicotyledons that contains herbaceous and shrubby plants. It has been reclassified as Vasconcellea pubescens (Vasconcellea cundinamarcensis (Linden) Badillo, Carica pubescens Lenné et C. Koch) (Badillo 2000). Commonly the Vasconcellea species are called “highland papayas” because of the botanical similarity with Carica papaya L., and also because they are native to the high regions of the Ecuadorian Andes (National Research Council 1989; Badillo 2000; Van Droogenbroeck et al. 2002; Morales et al. 2004).
Although documented information about the introduction of V. pubescens into Chile is not abundant, its presence seems to be ancient. Campos (1998), points out that this species would have been introduced in the North of Chile before the Spanish arrived there—in 1535—by migrations of pre-Columbian peoples. In a similar way, information about how this species arrived to the Southern region of Chile is also unknown, but it is believed that the seeds came from the Northern Chilean plantations.
In Chile, V. pubescens fruits are mainly used to produce preserved fruit and small quantities of juice, jam and processed sweets. However, an additional feature of this species is its ability to produce latex with a high level of papain, an important and valuable proteolytic enzyme with industrial, pharmaceutical and research uses (Kiger 1986a, b).
Currently, only 225 hectares of V. pubescens are cultivated in Chile, mainly concentrated between 30° and 33° latitude south (94%). Most orchards in this area are relatively large (10–40 ha) and maintained under standard fruit management (terracing, fertigation, pest management and pruning, etc.), supplying fruit both for the local and the international market. This species is also cultivated in the south of Chile (35°–38° South Latitude) but in smaller orchards, mostly maintained as home gardens (ODEPA-CIREN 2005).
V. pubescens is an open pollinated species in which it is normally possible to distinguish male, female and hermaphrodite plants (Storey 1976). It has been a traditional practice that growers produce their own planting material from their own orchards (Kiger 1986a). During this process the growers choose the best plants to collect fruits and recover seeds to produce seedlings. The particular circumstances under which this species has evolved in Chile must have affected its genetic diversity and genetic structure.
The Inter Simple Sequence Repeats (ISSR) markers are a very useful tool for studies of genetic diversity (Wolfe et al. 1998). They are based on the use of the microsatellite sequences as primers to generate multiple band patterns. The necessity of previous knowledge of the genome is obviated in the design of primers, which makes this technique an attractive, quick and economic possibility (Zietkiewicz et al. 1994). Additionally, given the abundance of microsatellites sequences it is possible to analyze a large number of loci, giving high possibilities of finding polymorphisms, even in highly related genotypes (Arnau et al. 2003; Carrasco et al. 2007).
In this study we evaluate the level and organization of the genetic diversity in V. pubescens cultivated in Chile, in order to establish a base line to assist future conservation and breeding programmes of the Vasconcellea genus.
Material and methods
Plant material
Young leaves were collected from trees growing in all the cultivated areas of Chile, from 29°58′30.1″ to 38°06′47.9″ (Table 1), comprising 333 adult plants. The collected samples were frozen in liquid nitrogen and stored at −20°C until DNA extraction.
Sample collection
Sample size varied from 10 to 20 individuals depending on the number and size of each orchard found in each study area (see Table 1). Orchards, and samples within orchards, were collected at random in relation to morphological characteristics and position within the area. Samples collected from growers that exchange planting material were considered as the same geographic sub-group for molecular analysis. In that way, 26 orchards and home gardens sampled in this study, were grouped into 5 groups for the Southern area and 5 groups for the Northern area.
Samples collected from Lipimavida, Iloca, Curanipe, Cobquecura and Contulmo- Lleu-Lleu (between 34°50′48.9″ S and 38°06′47.9″ S, Fig. 1; Table 1) were considered as belonging to the Southern area. In these locations all orchards were small home-gardens (30–100 plants per orchards). Samples collected from La Serena, Valle del Elqui, Ovalle, Huentelauquen and La Ligua-Quillota (between 29°58′30.1″ S and 32°23′35.5″S, Fig. 1; Table 1) were considered as belonging to the Northern area. In this last case the plant material was collected from commercial orchards (10–40 ha).
Geographical locations (latitude and longitude) of the 10 V. pubescens orchards sampled in this study and genetic structure. Orchards from La Ligua to La Serena were considered as the Northern area, while orchards from Lipimavida to Lleu-Lleu were considered as the Southern area. Coloured circles is a graphic representation that show all the genetic groups obtained (8) by Structure and Bayesian Analysis of Population Structure (BAPs). Population with the same colour represents similar genetic groups (Cobquecura and Lleu-Lleu; Curanipe and Lipimavida)
DNA extraction and PCR amplification
DNA extraction and PCR amplification were carried out according to Carrasco et al. (2007). A set of 36 primers (set ISSR 100/8, Biotechnology Laboratory from University of British Columbia, Vancouver) was tested. The basic characteristics of the primers that gave positive results are shown in Table 2. All PCR products were checked by electrophoresis on 2% agarose gels, run in TAE 1X buffer and visualized by ultraviolet fluorescence after staining with ethydium bromide (0.25 μl·ml−1 staining solution).
ISSR data analysis
From the agarose gels, each generated band was considered as a locus and was designated as present (1) or absent (0) for each individual. For all statistical analyses, the samples were recorded according to geographical origin (based on GPS positions). The POPGENE version 1.31 software (Yeh et al. 1999) was used to calculate the percentage of polymorphic ISSR loci (P), Nei's gene diversity (h), (Nei 1973) and Shannon's index (I, Shannon 1948). Nei's gene diversity is a simple measure of genetic variability and is defined as: h = 1 − ∑ x i 2, where (in the case of dominant molecular markers) x i is the population frequency of each allele (1 and 0) at locus i (Nei 1978). Shannon's index (I) is defined as: I = −∑ p i ln (p i ), where p i is the proportion of the ith allele in the population (Lewontin 1972). For each locus Nei's index produces values from 0–0.5 and Shannon's index produces values ranging between 0 and 0.73 on a natural logarithm scale (Lowe et al. 2004).
The hierarchical ISSR frequency distribution was analyzed using AMOVA incorporated using software GENALEX v. 6 (Peakall and Smouse 2006). To estimate the population genetic differentiation when binary data were analysed, this software calculates a parameter Φpt, an analogue to the currently used F st. Φpt is calculated from a matrix of Euclidean distances metric, unlike F st, which is a genetic frequency based measure (Peakall and Smouse 2006). The statistical significances of Φpt were evaluated using null distributions generated by random permutation of individuals (1,000 bootstrap permutations).
Population structure
The population structure determined by methods such as AMOVA, have the disadvantage that populations must be defined “a priori”, therefore the significance of values may change depending on the “a priori” grouping. In contrast, Bayesian methods have the advantage of inferring populations based on the frequencies of the alleles. The individuals in the sample are assigned probabilistically to populations, or jointly to two or more populations if their genotypes indicate that they are genetically mixed (admixture) (Pritchard et al. 2000; Pritchard and Wen 2003). In the first method, population structure within V. pubescens samples was inferred using a Bayesian model-clustering algorithm implemented in the computer programme, Structure (version 2.1) (Pritchard and Wen 2003). To run the software, a number K of genetic clusters characterized by the matrices of allele frequencies at each locus is first assumed. Then, for each individual, the proportion of its genome derived from each genetic cluster (proportion of ancestry) is estimated. Ten independent runs of the algorithm, assuming values of K from 1 to 10 with 600,000 Markov chain Monte Carlo (MCMC) repetitions and a burning period of 60,000 were performed, assuming population admixture. The posterior probability (probability of K given the data) was then calculated for each mean value of K using the mean estimated log-likelihood of K to choose the optimal K. The proportion of ancestry in a given cluster was calculated as an assignment rate (q), in general a level of 0.8 is accepted as a correct assignment to a single cluster. The algorithm used by Structure may be poorly suited for inferring the number of genetic clusters in a data set that has a relationship of identity by descent (Pritchard and Wen 2003). Considering the latter, the second method was a Bayesian clustering method described in Corander et al. (2003), implemented in software Baps (version 4.14). In contrast with Structure, it uses stochastic optimization to infer the genetic structure and it can use a spatial model that takes into account individual geo-referenced multilocus genotypes to assign a biologically relevant structure, thereby increasing the power to detect correctly the underlying population structure. Ten independent repetitions for each K from 1 to 10 were carried out.
Results
For V. pubescens cultivated in Chile, the seven ISSR primers studied in 333 samples collected along a geographic gradient, yielded a total of 114 bands (Table 2). Out of the 114 bands recorded, 63 proved to be polymorphic (P = 55.3%). At the species level, the genetic diversity was rather low (h = 0.01 ± 6.80188E-05, Shannon’s Index I = 0.16 ± 0.000148).
Similar results were observed at the geographic group level (Table 1) where the genetic diversity was low for every parameter measured, independent of the number of trees sampled per location.
Interestingly, despite the scarcity of the genetic diversity found in the cultivated V. pubescens in Chile, it showed a clear structure (Fig. 2). The major portion of the genetic diversity was found within groups (65% Fig. 2a) when all the samples were analysed together. The genetic differentiation between the different groups was extremely high and significant, as the AMOVA analysis suggests (Φpt = 0.35, P < 0.001, Fig. 2a). This was especially remarkable when analysing the Northern area alone, as the differentiation was even higher (Φpt = 0.40, P < 0.001, Fig. 2c). Whereas when only the Southern area was analysed, Φpt was reduced to 0.18 (P < 0.001, Fig. 2b), indicating greater genetic similarity.
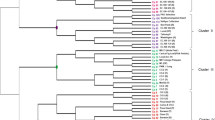
The results generated from Structure and BAPs for all the populations showed a value of K = 8 as being the best average assignment rate (q > 0.8). This means it is possible to distinguish 8 genetically different groups, five of them located in the North and three in the South (Fig. 1). Similarly with the AMOVA results, the individuals within the Southern area appear to be more similar to each other than those in the North. For example, the individuals from Buchipureo and LleuLleu, were genetically similar therefore they were shown as clustering together (Fig. 1). A similar situation was also observed between Curanipe and Lipimavida, although these locations are 120 km apart (Fig. 1). Iloca proved to be different from the other two Southern groups. On the other hand, the individuals from the Northern area were genetically dissimilar and clustered into five different groups.
Discussion
This is the first report of investigations into the organization of the genetic diversity in a cultivated species of the Vasconcellea genus. Previously Kim et al. (2002); Saxena et al. (2005) studied C. papaya cultivars and breeding lines, in order to establish the genetic relationships among specific samples, and provide information for germplasm management, improvement and cultivar protection.
The genetic diversity of V. pubescens was remarkably low (h = 0.01, I = 0.16) compared with the average diversity reported for plants when using dominant markers (h = 0.19–0.23, Nybom 2004). While at the same time, the percentage of polymorphic loci (62.3%) showed similar levels to other plant studies (P = 51–91%, Nybom 2004). This reflects an interesting situation in which a high polymorphism does not necessarily generate high gene diversity.
Many aspects, such as the breeding-system, seed and pollen dispersal, plant longevity and agricultural practices, influence the genetic diversity, including the proportion of variation distributed within and between populations (Hamrick and Godt 1989, 1996a, b). Open-pollinated, or outcrossing species show a higher proportion of genetic diversity within populations with lower levels of genetic differentiation between populations (Hamrick and Godt 1996b).
The genetic differentiation between groups are in accordance with those observed in plants using dominant markers (F st = 0.34–0.35, Nybom 2004), as well as with clonally propagated and autogamous plants species (F st = 0.43, Morjan and Rieseberg 2004).
The higher genetic similarity among the Southern groups could be due to the active exchange of plant material among growers, since in these latitudes growers usually buy and share fruits with their neighbours. Iloca is a special case because it is the largest area cultivated in the south and from discussions with the growers it is clear that they frequently include seeds from the Northern area.
Compared to the Southern, the orchards from the Northern area were genetically more distinct from each other, regardless of their geographic distance. Growers in this area have larger orchards and they are much more competitive. From discussions with them they appear to never or rarely, exchange plant material.
The level of genetic diversity and genetic structure observed for V. pubescens cultivated in Chile also may be explained by founder events and extended selection. This species is an introduced crop into Chile and a reduction of genetic diversity by founder events as compared to the center of origen in the high land of Ecuador Andes could be expected. For the future it will be interesting to carry out an extensive study including samples from center of origin of the species in order to verify its genetic diversity patterns. A second founder event can explain the lower genetic diversity in the Southern population as compared with the Northern populations, given the introduction/migration of a few seed from the Northern genetic stock and the small extension of the plantations that could reduce the chances of crossing and hybridization.
Growers tend to select their own best trees to collect fruit and produce plants. The effects caused by selection on cultivated plants have been documented extensively (Tang and Knapp 2003; Hollingsworth et al. 2005; Hyten et al. 2006). Selection is similar to having a severe bottleneck removing most of the genetic variation from the target loci and other associated loci, if not from all the genome (Wright et al. 2005.).
In addition, the agricultural practices applied by growers generate high levels of inbreeding. Similarly to Carica papaya, V. pubescens is a trioic (it is possible to find male, female and hermaphrodite plants), self-compatible and open-pollinated species (Kiger 1986a; Yagnam 1993). Currently plantation systems used by farmers establish three plants per planting-hole in the field, to thereafter eliminate male plants. Such practices stimulate crossing among related individuals, and increase the degree of selfing. For instance, it is clearly possible to view the current genetics status of papaya resources present in Chile, as reflecting the result of many years of cultivation that have reduced the genetic diversity but generated a high level of genetic structure.
From a breeding programmes perspective, it would be advisable to exploit efficiently the genetic resources by incorporating all the available genetic diversity into one gene pool, and thus start a national breeding programme by crossing material from the different cultivated areas.
References
Arnau G, Lallemand J, Bourgoin M (2003) Fast and reliable strawberry cultivar identification using inter simple sequence repeat (ISSR) amplification. Euphytica 129:69–79
Badillo V (2000) Carica L. vs. Vasconcella St. Hil. (Caricaceae) con la rehabilitación de este último. Ernstia 10:74–79
Campos J (1998). Productos forestales no Madereros en Chile. Serie Forestal No 10. Biblioteca Virtual FAO. (en línea) disponible en: http://www.fao.org/documents/show_cdr.asp?url_file=//docrep/t2368s/T2368s00.htm. Accessed March, 2007
Carrasco B, Garcés M, Rojas P, Saud G, Herrera R, Retamales JB et al (2007) The Chilean strawberry (Fragaria chiloensis (L.) Duch.): genetic diversity and structure. J Am Soc Hortic Sci 132:501–506
Corander J, Waldmann P, Sillanpää MJ (2003) Bayesian analysis of genetic differentiation between populations. Genetics 163:367–374
Hamrick JL, Godt MJ (1989) Allozyme diversity in plant species. In: Brown A, Clegg M, Khaler A, Weir B (eds) Plant population genetics breeding and genetic resources. Sinauer Press, Massachussets, USA, pp 43–63
Hamrick JL, Godt MJ (1996a) Conservation genetics of endemic plant species. In: Avise JC, Hamrick JL (eds) Conservation genetics case histories from nature. Chapman and Hall, New York, USA, pp 281–304
Hamrick JL, Godt MJ (1996b) Effects of life history traits on genetic diversity in plant species. Philos Trans R Soc Lond 351:1291–1298
Hollingsworth PM, Dawson IK, Goodall-Copestake WP, Richardson J, Weber J, Sotelo-Montes C et al (2005) Do farmers reduce genetic diversity when they domesticate tropical trees? A case study from Amazonia. Mol Ecol 14:497–501
Hyten D, Song Q, Zhu Y, Choi I, Nelson R, Costa J et al (2006) Impacts of genetic bottlenecks on soybean genome diversity. Proc Natl Acad Sci USA 103:16666–16671
Kiger M (1986a) La papaya y su aprovechamiento agroindustrial. Proxima Decada 49:4–7
Kiger M (1986b) Alternativas no tradicionales de industrialización de la papaya. Proxima Decada 50:21–24
Kim MS, Moore PH, Zee F, Fitch MMM, Steiger DL, Manshardt RM et al (2002) Genetic diversity of Carica papaya as revealed by AFLP markers. Genome 45:503–512
Lewontin R (1972) The apportionment of human diversity. Evol Biol 6:381–398
Lowe A, Harris S, Ashton P (2004) Ecological genetics: design, analysis and application. Blackwell Publishing, Oxford
Morales A, Medina D, Yaguachi B (2004) Genetic diversity, phylogeny and geographic distribution of genus Vasconcellea in Southern Ecuador. Lyonia 7:15–27
Morjan CL, Rieseberg LH (2004) How species evolve collectively: implications of gene flow and selection for the spread of advantageous alleles. Mol Ecol 13:1341–1356
National Research Council (1989) Highland papayas. In: Ruskin FR (ed) Lost crops of the Incas: little-known plants of the Andes with promise for worldwide cultivation. National Academy Press, Washington DC, pp 252–261
Nei M (1973) Analysis of gene diversity in subdivided populations. Proc Natl Acad Sci USA 70:3321–3323
Nei M (1978) Estimation of average heterozygosity and genetic distance from small number of individuals. Genetics 89:583–590
Nybom H (2004) Comparison of different nuclear DNA markers for estimating intraspecific genetic diversity in plants. Mol Ecol 13:1143–1155
ODEPA-CIREN (2005). Catastro Frutícola IV Región, Principales Resultados. En línea http://www.odepa.gob.cl/servicios-informacion/Catastrosfruticolas/catastro-IVRegion-2005.pdf. Accessed March 2007
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in Excel Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Pritchard J, Wen W (2003) Documentation for structure software: version 2. Department of Human genetics University of Chicago, USA
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Saxena S, Chandra R, Srivastava A, Mishra M, Pathak RK, Ranade SA (2005) Analysis of genetic diversity among papaya cultivars using Single Primer Amplification Reaction (SPAR) methods. J Hortic Sci Biotechnol 80:291–296
Shannon CE (1948) A mathematical theory of communication. Bell Syst Tech J 27:379–423
Storey WB (1976) Papaya, Carica papaya. In: Simmonds (ed) Evolution of crops plants. Longman, London, pp 21–24
Tang S, Knapp SJ (2003) Microsatellites uncover extraordinary diversity in native American land races and wild populations of cultivated sunflowers. Theor Appl Genet 106:990–1003
Van Droogenbroeck B, Breyne P, Goetghebeur P, Romeijn-Peeters E, Kyndt T, Heysen G (2002) AFLP analysis of genetic relationships among papaya and its wild relatives (Caricaceae) from Ecuador. Theor Appl Genet 195:289–297
Wolfe A, Xiang QY, Kephart S (1998) Assessing hybridization in natural populations of Penstemon (Scrophulariaceae) using hypervariable inter-simple sequence repeat (ISSR) bands. Mol Ecol 7:1107–1125
Wright SI, Bi IV, Schroeder SG, Yamasaki M, Doebley JF, McMullen MD et al (2005) The effects of artificial selection on the maize genome. Science 308:1310–1314
Yagnam F (1993) El cultivo del Papayo, Extracción de Papaína y Latex. Chil Hortofruticola 2:40–47
Yeh F, Boyle T, Ye Z, Xiyan JM (1999) POPGENE Version 131 Microsoft Window-based freeware for population genetic analysis. University of Alberta and Center for International Forestry Research, Edmonton
Zietkiewicz E, Rafalski A, Labuda D (1994) Genome fingerprinting by simple sequence repeat (SSR)-anchored polymorphism chain reaction amplification. Genomics 20:176–183
Acknowledgements
We acknowledge financial support of PBCT-CONICYT: Inserción de Investigadores Postdoctorales No 10 and the Universidad de Talca, Chile. The authors wish to thank for their support for technical assintance in GIS to David López. We also thank Dr. Lyndsey Nicholson, and anonymous reviewers for critically reviewing this manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Carrasco, B., Avila, P., Perez-Diaz, J. et al. Genetic structure of highland papayas (Vasconcellea pubescens (Lenné et C. Koch) Badillo) cultivated along a geographic gradient in Chile as revealed by Inter Simple Sequence Repeats (ISSR). Genet Resour Crop Evol 56, 331–337 (2009). https://doi.org/10.1007/s10722-008-9367-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-008-9367-1