Abstract
A genetic linkage map of flowering dogwood (Cornus florida L.) was constructed using 94 individuals derived from a cross of two F1 trees designated 97-6 and 97-7, which were originally from a cross between ‘Appalachian Spring’ and ‘Cherokee Brave’. Out of approximately 800 SSR loci examined, 271 were polymorphic between ‘Appalachian Spring’ and ‘Cherokee Brave’, but were monomorphic between 97-6 and 97-7. These 271 segregating markers were used to build a linkage map for flowering dogwood. Eleven linkage groups were obtained with a log-of-odds (LOD) value of 6.0 using JoinMap® 4.0 software, which matches the chromosome number of flowering dogwood haploid genome. This linkage map consisted of 255 SSR loci, spanned a total of 1,175 centimorgans (cM) with an average internal distance of 4.6 cM. Several larger gaps and slight clustering of markers were present on this linkage map. This is the first linkage map of flowering dogwood and will be a fundamental tool for new gene identification and marker-assisted selection in our flowering dogwood breeding program.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Flowering dogwood (Cornus florida L.), native to Eastern North America, is an ornamental tree that plays an important role in the U.S. nursery industry (Ament et al. 2000; Witte et al. 2000). Over 100 cultivars have been released in the U.S. in the last decade (Witte et al. 2000). ‘Appalachian Spring’ and ‘Cherokee Brave’ are popular cultivars exhibiting different phenotypes. ‘Appalachian Spring’ has large white-bracts in the spring and deep green leaves that turn scarlet in fall (Windham et al. 1998). In contrast, ‘Cherokee Brave’ has red or pink bracts in the spring and green leaves that turn fiery red in the fall.
Flowering dogwood is predominantly cross-pollinating and highly self-incompatible (Ament et al. 2000; Cappiello and Shadow 2005; Witte et al. 2000; Gunatilleke and Gunatilleke 1984; Orton 1985; Reed 2004; Sork et al. 2005). Most flowering dogwood cultivars are propagated from axillary buds grafted onto rootstock grown from wild seedlings, or more rarely, from rooted cuttings of the cultivar (Dirr 1998). With a long juvenility period, flowering dogwood trees can require five to seven years to bloom. In the past, studies of flowering dogwood were focused on breeding or selecting new varieties with disease or pest resistance (Windham et al. 2003). Molecular studies on flowering dogwood, which were initiated <10 years ago, have largely been concerned with cultivar identification and included work with simple sequence repeats (SSRs or microsatellites; Cabe and Liles 2002; Wang et al. 2007, 2008); amplified fragment length polymorphisms (AFLPs; Nealed and William 1999; Smith et al. 2007) and DNA amplification fingerprinting (DAF, Ament et al. 2000). Combinations of traditional breeding methods and molecular techniques will facilitate breeding progress for C. florida. However, molecular breeding must be established on some fundamental knowledge of plant genetics. Although molecular markers were applied in flowering dogwood (Smith et al. 2007; Wang et al. 2008), a framework of linkage relationships among these markers is necessary for the identification and localization of genes controlling important horticultural traits, which will subsequently permit the application of marker-assisted selection (Baldoni et al. 1999) in a dogwood breeding program.
Although woody perennial trees, like flowering dogwood, have many advantages for genetic studies (clonally propagated, small genome size and high genetic diversity), most of them have many disadvantages such as long generation time, self incompatibility, and inbreeding depression (Weeden et al. 1994). Due to high heterozygousity in out-breeding species, an F1 population pseudo-testcross strategy has been widely used for linkage analysis (Grattapaglia and Sederoff 1994; Arcade et al. 2000; Porceddu et al. 2002; Hanley et al. 2002; Barcaccia et al. 2003; La Rosa et al. 2003; Doucleff et al. 2004; Dirlewanger et al. 2004; Graham et al. 2004; Beedanagari et al. 2005; Kenis and Keulemans 2005; Verde et al. 2005; Lanteri et al. 2006; Lowe and Walker 2006; Mandl et al. 2006). Other mapping populations such as F1 full-sib progeny (Maliepaard et al. 1997; Venkateswarlu et al. 2006) and BC1 mapping populations (Lalli et al. 2008) have also been used for linkage map construction. In a pseudo-testcross, a single full-sib population is generated by crossing two parents in which one parent is heterozygous at one locus, and the other parent is homozygous at other loci (Mandl et al. 2006). Although the pseudo-testcross strategy has been widely used in the mapping of many out-breeding species, it is difficult to determine if one of the parents is homozygous for any specific locus or loci.
A “pseudo-F2” mapping strategy was described for the construction of a linkage map of a non-inbred Solanum tubersosum (potato) cultivar (Rouppe van der Voort et al. 1999). A selfed family of three F1 Manihot esculenta (cassava) plants derived from a single progeny of a full-sib population was considered a “pseudo F2” population and used for QTL analysis (Okogbenin et al. 2008). The generation of a classical F2 population for flowering dogwood is impractical because homozygous lines are not available and flowering dogwood is self-incompatible (Reed 2004). Furthermore, the long juvenility phase prior to bloom has thwarted geneticists from creating an F2 mapping population. Weeden et al. (1994) suggested that seeds derived from a single tree represent a similar set of haploid genotypes, although complexes with another haploid genotype from the pollinator exist. The seeds from the single tree could be considered equivalent to an F1, and be used as mapping population by means of the pseudo-testcross strategy. However, factors such as unknown pollen donor or possible multiple pollinators affect the linkage analysis.
Although it is impractical to make an F2 from one F1 flowering dogwood tree, it is easy to make a “pseudo F2” generation using two F1 trees. If the F1 trees show genetic identity at a given locus, the progeny could be considered as F2, which could be assigned as a “pseudo F2”. Hence, two F1 breeding lines, 97-6 and 97-7, from a cross between ‘Appalachian Spring’ and ‘Cherokee Brave’ were crossed to create a “pseudo F2” population.
We screened the DNA of four parents with 825 microsatellite (SSR) markers developed previously (Wang et al. 2008) in order to find those loci most polymorphic between ‘Appalachian Spring’ and ‘Cherokee Brave’, but with identical allele segregation between two F1 parents, 97-6 and 97-7 at a given locus. The concept of the F2 generation in self incompatible woody plants can be compared to that of F2 or BC1 of an annual selfing crop (Tulsieram et al. 1992). Such a mapping population was used for genetic linkage map construction of flowering dogwood following the out-breeding population module of the JoinMap® 4.0 software program (Van Ooijen 2006).
The aim of this work was to build a microsatellite marker-based genetic linkage map of flowering dogwood. This initial linkage map will facilitate the location of genes of interest and subsequently permit marker-assisted selection (Strauss et al. 1992) in our breeding program for flowering dogwood. The release of the first linkage map in flowering dogwood will provide the basis for identification and cloning of important genes and quantitative trait loci (QTLs) that are beneficial to ornamental industry (Dirlewanger et al. 1999).
Materials and methods
Plant material
Two flowering dogwood (C. florida L.) F1 trees, 97-6 and 97-7, derived originally at the University of Tennessee ten years ago from the intra-specific cross ‘Cherokee Brave’ × ‘Appalachian Spring’ (Fig. 1), were used for creating a mapping population. In early spring 2005 at the Agricultural Experimental Station of the University of Tennessee, Crossville, TN, two trees, 97-6 and 97-7, were placed into a fiberglass mesh screened cage (2.5 m × 2.5 m × 2.5 m) and reciprocal honeybee-mediated crosses were performed.
In the fall of 2005, about 400 mature fruits from 97-6 and 97-7 crosses were collected, depulped, the “seeds” removed, and then dried for 24–48 h. After stratification at 4°C for about 4 months, the germinated “pseudo F2” seeds were planted in the greenhouse.
DNA isolation
Young, not fully expanded leaves were collected during April–May from seedlings and DNA extracted using DNeasy Plant Mini Kit (QIAGEN, Valencia, CA, USA) following the manufacturer’s instructions. Due to inbreeding depression, most of the F2 plants appeared dwarfed and unhealthy and grew very slowly compared to robust growth exhibited by F1 plants. As a result, some samples were collected more than once to obtain useable DNA. Genomic DNA concentration was quantified using NanoDrop® ND-1000 UV-Vis Spectrophotometer (NanoDrop ND-1000, Delaware, USA). One nanogram (ng) of DNA was used as template for PCR with four informative SSR primers (data not shown) to check for successful amplification. The PCR protocols described previously by Wang et al. (2008) were used throughout this study.
Mapping population size and determination of SSR makers
A set of 94 “pseudo F2” samples was randomly chosen from successfully amplified samples with four SSR primers. To find polymorphisms between the DNA of the parents, but identity between the DNA of the two F1 parents, 825 SSR primers previously developed (Wang et al. 2008) were screened with the DNA of the four parents (data not shown). PCR conditions and allele size determination followed the methods of previous work (Wang et al. 2008).
Mapping and linkage analysis
Markers were recorded as the following three types: (1) maternal markers, segregating only within female parent ‘Cherokee Brave’, and null in male parent ‘Appalachian Spring’ (expected segregation ratio 1:1). These markers were coded with the letter “B” followed by the marker name; (2) paternal markers, segregating only within male parent ‘Appalachian Spring’, and null in female parent ‘Cherokee Brave’ (expected segregation ratio 1:1). These markers were coded with the letter “A” followed by marker name; and (3) intercross markers, segregating within both parents, expected segregation ratio either 3:1 (dominant), 1:2:1 (co-dominant) or 1:1:1:1 (presents three or four alleles). The dominant markers with 3:1 ratio were not be used for mapping because of homozygosity in both parents. The markers with 1:2:1 and 1:1:1:1 ratios were designated with the letter “C” followed by the marker name.
Chi-square (χ 2) tests were performed to test goodness-of-fit between observed and expected segregation ratios. Markers segregating in Mendelian ratios (including some slightly skewed markers, P ≤ 0.01) were used for map construction. Heavily distorted segregation of markers (P > 0.01) were omitted from the analysis. Data were analyzed using the cross-breeding population type option of JoinMap® 4.0 mapping program (Van Ooijen 2006). Different log-of-odds (LOD) scores were produced to determine the linkage groups that match the number of chromosomes of the C. florida genome, which is eleven (Radford et al. 1979). Finally, linkage groups were determined with a LOD threshold of 6.0 and recombination frequency ≤0.40. The relative marker order within each linkage group was determined based on the following parameters: Rec = 0.40, LOD = 1.0, goodness-of-fit jump threshold = 5.0, Map distances were computed using Kosambi’s regression mapping function (Kosambi 1944). Linkage maps were drawn using MapChart 2.1 software (Voorrips 2002).
The linkage coverage (percentage) can be calculated from the total length of the linkage divided by the total genome length (G) of the genome in centimorgans (cM). Total genome length can be estimated by the methods of both Hulbert et al. (1988) and Chakravarti et al. (1991), and the formula for determining G is as follows: G = N (N − 1)X/K at an LOD threshold of T, where N is the total number of markers analyzed, X is the average distance between adjacent markers at a certain LOD value of T, and K is the observed number of pairs of markers with an LOD ≥ T.
Genome size estimates by flow cytometry
Flow cytometric measurements of nuclear DNA quantity were made from fresh C. florida leaf tissue samples using a two-step procedure originally described by Otto (1990) and modified by Dolezel and Göhde (1995). Approximately 0.5 cm2 of growing leaf tissue of sample and 1.0 ml of cold Otto I Buffer (0.1 M citric acid monohydrate and 0.5% (v/v) Tween) were chopped for 30–60 s in a plastic petri dish on ice. The resulting extract was passed through a 30 μm nylon mesh filter into a 3.5 ml plastic tube, which was centrifuged at 150g for 5 min. Supernatant was removed leaving approximately 100 μl solution and pellet, to which an additional 100 μl of cold Otto I Buffer was added. Pellet was gently resuspended and incubated on ice for 30 min. The fluorochrome 4, 6-diamidino-2-phenylidole (DAPI) was added to 1.0 ml of ice-cold Otto II Buffer (0.4 M Na2HPO4 · 12H2O) at a concentration of 4 μg/ml and mixed with the sample on ice. The relative fluorescence of the total DNA was measured for each nucleus using a Partec PA-1 ploidy analyzer (Partec GMBH). At least 5,000 nuclei were analyzed, revealing a single peak with a coefficient of variation (CV) less than 5.0%. Genome sizes were calculated as nuclear DNA content for unreduced tissue (2C) as: 2C DNA content of tissue = (mean fluorescence value of sample ÷ mean fluorescence value of standard) × 2C DNA content of standard. Lycopersicon esculentum (Stupicke polni) and Zea mays (CE-177 inbred line) have established 2C genome sizes of 1.96 and 5.43 picograms, respectively (Lysák and Doležel 1998; Dolezel et al. 1992) and were used as internal standards.
Results
Primer screening
The DNA of ‘Appalachian Spring’, ‘Cherokee Brave’, and two F1 (97-6 and 97-7) were amplified with 825 SSR loci. Markers that were heterozygous within grandparents and homozygous within parents were selected for mapping population segregation. As a result, 469 of the 825 SSR markers (56.8%) showed polymorphism between ‘Appalachian Spring’ and ‘Cherokee Brave’. Of the 469 SSR markers, 274 (58.4%) presented no difference between 97-6 and 97-7, which implies that 97-6 and 97-7 are genetically identical for these alleles. From the 274 SSR markers, 219 SSR markers presented sufficient separation among the 94 individuals in “pseudo-F2” populations that the alleles were easily scored and used for linkage analysis. Of the 219 markers recorded, 167 exhibited single locus segregation (either male parent or female parent) and 52 showed two loci segregation (co-dominant plus male or female parent). Three markers (1%) showed highly significant levels of distortion (0.01 < P < 0.001) and were excluded from the linkage analysis (Table 1). A total of 271 SSR loci were used for linkage analysis.
In all, 271 polymorphic loci were available for map construction, including 76 maternal, 48 paternal and 147 common loci. A total of 129 (47.6%) out of 271 markers showed significant deviation between 0.05 ≤ P ≤ 0.01 and were included for map construction. Alleles in the male parent showed significant segregation distortion when compared to those in the female parent (4 loci vs. 27 loci). Most co-dominant markers (66.7%) showed distortion at 0.05 ≤ P ≤ 0.01 (Table 1).
Linkage map construction
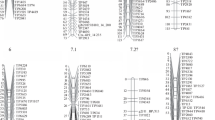
In all, 271 markers were available for map construction, including 76 maternal loci, 48 paternal loci and 147 common loci. All markers were assigned to 11 major linkage groups (LGs) with a minimum of 11 markers at a LOD value of 6.0 (Table 2). The number of linkage groups detected in this mapping population corresponds exactly with the haploid number of chromosomes in C. florida. Four maternal, seven paternal and five co-dominant loci remained unlinked (5.9% in total). A total of 255 (94.1%) markers were ordered onto 11 LGs. The length of the individual LGs varied from 69.4 to 136.5 cM with a mean of 106.8 cM. The number of loci on LGs ranged from 11 to 37 with an average of 23.2 loci. The distance for the 11 LGs spanned a total of 1,175.0 cM with an average internal distance of 4.6 cM. The majority of the marker intervals were less than 10 cM. However, some large gaps remain in each LG, and the largest gap of 26.6 cM was found on LG3 (Table 2; Fig. 2). Except for slight clustering of markers on LGs 5, 7 and 11, all markers were evenly distributed across the 11 LGs (Fig. 2).
Genetic map of an intra-specific population between C. florida ‘Appalachian Spring’ (AS) and ‘Cherokee Brave’ (CB). The letter “A” indicates markers associated with the male parent AS and the letter “B” indicates markers associated with the female parent CB. The two letter prefix “CF” for each marker is the initial letters of C. florida and have been registered at GenBank (Wang et al. 2008)
Of the 255 mapped markers, 129 distorted markers were distributed onto each LG (Table 3). LG 1, 5, 6 and 8 contained more distorted dominant maternal markers, LG 1, 5, and 11 had more distorted co-dominant markers. There were no distorted dominant markers on LG 3 and 4. Four distorted parental markers were located on LG 1, 2 and 11 (Table 3).
There were 268 marker pairs at the LOD value of 6.0. Given the formula of Hulbert et al. (1988) and the “method 3” of Chakravarti et al. (1991), the genome length of this species (C. florida L.) was estimated as 1255.9 cM. Therefore, the 11 LGs covered 93.6% of the flowering dogwood haploid nuclear genome. Given one picogram (pg) equals 978 Mb (Doležel et al. 2003), the genome size of flowering dogwood was estimated to be 1.28 pg per haploid cell (1C), which corresponds with the estimated genome size (2C) of 2.19–2.77 pg/nucleus that we found for flowering dogwood using flow cytometry.
Discussion
Unlike inbred species, out-crossing species are highly heterozygous and generally not easy to obtain a pedigree from crosses between inbred lines (Venkateswarlu et al. 2006). When attempting to build a genetic linkage map, the mapping population is the main issue for the crossing parents (Maliepaard et al. 1997). Most difficulties in linkage analysis for out-crossing species involve more than one segregating allele per locus of each parent, and the unknown linkage phase of the loci. Unlike a segregating population derived from two inbreeding species, the segregating loci present only two alleles and the alleles have the same phase in the F1 (Blenda et al. 2007). On the contrary, a segregating population between two non-identical plants of two out-crossing parents may segregate up to four alleles per locus, and the linkage phase of the marker pairs usually are unknown (Maliepaard et al. 1997; La Rosa et al. 2003; Lanteri et al. 2006).
In out-breeding trees, most researchers have used an F1 mapping population with a two-way pseudo-testcross strategy to construct an individual genetic linkage map for each parent (Grattapaglia and Sederoff 1994; Arcade et al. 2000; Porceddu et al. 2002; Hanley et al. 2002; Barcaccia et al. 2003; La Rosa et al. 2003; Doucleff et al. 2004; Dirlewanger et al. 2004; Graham et al. 2004; Beedanagari et al. 2005; Kenis and Keulemans 2005; Verde et al. 2005; Lanteri et al. 2006; Lowe and Walker 2006). F1 full-sib progeny (Maliepaard et al. 1997; Venkateswarlu et al. 2006) and BC1 population (Lalli et al. 2008) were also used as mapping population in the linkage analysis of out-crossing trees. Individual linkage maps produced by these methods may lead to inaccuracy in the estimation of recombination frequencies (Venkateswarlu et al. 2006).
Because flowering dogwood is a highly heterozygous and self-incompatible ornamental tree, we created an F2 population using two F1 breeding lines. This F2 population can be considered equivalent to the F2 of annual self-pollinated plants when the parents (two F1) are identical genetically at a given locus (Fregene et al. 1997; Okogbenin et al. 2008). All SSR markers selected for linkage analysis in this study presented identical segregation within two F1 parents. Therefore, this linkage map of flowering dogwood was built with a “pseudo-F2” mapping population. To demonstrate the marker order within each linkage group, we also used the pseudo-testcross strategy to construct an individual linkage map for both F1 parents. Eleven LGs were obtained for 97-6 (female) and twelve for 97-7 (male) at LOD value of 6.0 (data not shown). The alignment showed that eleven LGs of the female parent matched those LGs of flowering dogwood with minor distortion; while LG 5 split into two short LGs and generated 12 LGs in male parent’s map. This result indicates a necessity to develop more male parental markers to fill in 11 LGs. The alignment of common markers (co-dominant markers) with the parents’ LG showed the same or minor reversed order with present linkage map (data not shown). These results demonstrated that the genetic linkage map for flowering dogwood constructed with a “pseudo-F2” mapping population in this study is reliable. Compared to the pseudo-testcross strategy, pseudo-F2 mapping strategy has its own advantages including the following: does not require that one parent be heterozygous and other to be homozygous (Mandl et al. 2006), and is not necessary to know the allele segregation phase (Maliepaard et al. 1997; Venkateswarlu et al. 2006). However, “pseudo-F2” mapping strategy needs more molecular markers to generate fine linkage map and will lose some useful loci.
The “pseudo-F2” population of seedlings created with two F1 breeding lines showed inbreeding depression after emergence of the second pair of true leaves. About one-half of marker segregation (50.6%) showed distortion at 0.05 ≤ P ≤ 0.01. Although there is bias in the estimation of linkage analysis (Tavoletti et al. 1996; La Rosa et al. 2003; Lanteri et al. 2006), these markers were still included in the mapping data. Three markers were significantly skewed at P > 0.01 and initially excluded in the mapping data. The segregation distortion observed in this study was significantly higher than other studies (Conner et al. 1997; Casasoli et al. 2001; Scalfi et al. 2004; Pekkinen et al. 2005; Lantri et al. 2006) because both 97-6 and 97-7 are highly heterozygous at most loci.
Many biological factors, such as unequal crossover during meiosis, chromosome loss, non-random union of gametes, zygotic embryo abortion (Faris et al. 1998), changes in genetic load (Bradshaw and Stettler 1994), or lethal alleles (Pilien et al. 1993; Pekkinen et al. 2005) may cause allele segregation distortion. Forest trees often have less self-fertility and are believed to have many recessive lethal genes in the heterozygous condition (Sneizko and Zobel 1988). Due to selfing, some of the recessive lethal genes present in the heterozygous condition might have become homozygous and expressed in the progeny (Venkateswarlu et al. 2006). However, it may lead to the loss of a large part of a linkage group if those markers are significantly distorted at 0.05 ≤ P ≤ 0.01 and are excluded in the linkage analysis (Cervera et al. 2001; Doucleff et al. 2004). Some reports demonstrated that the distorted markers at 0.05 ≤ P ≤ 0.01 included in the linkage analysis were beneficial to linkage map construction (Kuang et al. 1999; Fishman et al. 2001; La Rosa et al. 2003; Lanteri et al. 2006). The use of an intraspecific cross might have contributed to the reduced level of segregation distortion observed (Becker et al. 1995; Sondur et al. 1996). In this study, we used progenies from an intraspecific cross of two cultivars. Although the distorted markers were included in the linkage analysis, these markers did not affect the linkage arrangement, as they were distributed on all linkage groups. Distorted markers that are distributed onto different linkage groups could be due to the heterogeneous transmission of chromosome fragments to progenies (Quillet et al. 1995). All distorted markers in this study demonstrated whole genome distribution. This is suggestive of a biological mechanism underlying segregation distortion (Lanteri et al. 2006), as opposed to random bias caused by scoring errors or chance (Fishman et al. 2001).
Significant clustering of markers was not observed in the LGs. However, slight clustering of markers was observed on some LGs, such LG 5, 7 and 11. Clustering is very common, especially for AFLP and RAPD marker-based linkage maps (Tanksley et al. 1992; Vallejos et al. 1992; Lantri et al. 2006; Blenda et al. 2007), whereas SSR markers were reported to randomly distribute across LGs (Lanteri et al. 2006). The slight clustering that occurred on some of the LGs in this study may have resulted from the inbreeding depression exhibited in the F2 mapping population. The small mapping population size (<100 individuals) and SSR loci mainly from a certain area of genome may result in clustering as well.
To our knowledge this is the first linkage map available for flowering dogwood. Our linkage map will benefit the following areas of genetic research for this species: 1. provide molecular fundamentals of genetic structure; 2. identify genes of interest on chromosomal regions, allowing for targeted selection of these genes in markers-assisted breeding in the future; and 3. serve as a reference map for high density saturation of future maps. By applying more SSR markers and developing other types of molecular markers, such as AFLPs, and increasing the mapping population size, a more comprehensive genetic map can be obtained.
References
Ament MH, Windham MT, Trigiano RN (2000) Determination of parentage of flowering dogwood (Cornus florida) seedlings using DNA amplification fingerprinting. J Arbor 26:206–212
Arcade A, Anselin F, Faivre Rampant P, Lesage MC, Paques LE, Prat D (2000) Application of AFLP, RAPD and ISSR markers to genetic mapping of European and Japanese larch. Theor Appl Genet 100:299–307. doi:10.1007/s001220050039
Baldoni L, Angiolillo A, Pellegrini M, Mencuccini M, Metzidakis IT, Voyiatzis DG (1999) A linkage genome map for olive as an important tool for marker-assisted selection. Acta Hortic 474:111–115
Barcaccia G, Meneghetti S, Albertini E, Triest L, Lucchin M (2003) Linkage mapping in tetraploid willows: segregation of molecular markers and estimation of linkage phases support an allotetraploid structure for Salix alba × Salix fragilis interspecific hybrids. Heredity 90:169–180. doi:10.1038/sj.hdy.6800213
Becker J, Vos P, Kuiper M, Salamini F, Heun M (1995) Combined mapping of AFLP and RFLP in barley. Mol Gen Genet 249:65–73
Beedanagari SR, Dove SK, Wood BW, Conner PJ (2005) A first linkage map of pecan cultivars based on RAPD and AFLP markers. Theor Appl Genet 110:1127–1137. doi:10.1007/s00122-005-1944-5
Blenda AV, Verde I, Georgi LL, Reighard GL, Forrest SD, Muñoz-Torres M et al (2007) Construction of a genetic linkage map and identification of molecular markers in peach rootstocks for response to peach tree short life syndrome. Tree Genet Genomes 3:341–350. doi:10.1007/s11295-006-0074-9
Bradshaw HD, Stettler RF (1994) Molecular genetics of growth and development in Populus II segregation distortion due to genetic load. Theor Appl Genet 139:963–973
Cabe PR, Liles JS (2002) Dinucleotide microsatellite loci isolated from flowering dogwood (Cornus florida L.). Mol Ecol Notes 2:150–152. doi:10.1046/j.1471-8286.2002.00169.x
Cappiello P, Shadow D (2005) Dogwoods. Timber, Portland
Casasoli M, Mattioni C, Cherubini M, Villani F (2001) A genetic linkage map of European chestnut (Castanea sativa Mill) based on RAPD, ISSR, and isozyme markers. Theor Appl Genet 102:1190–1199. doi:10.1007/s00122-001-0553-1
Cervera MT, Storme V, Ivens B, Gusmao J, Liu BH, Hostyn V et al (2001) Dense genetic linkage maps of three populus species (Populus deltoides, P nigra and Ptrichocarpa) based on AFLP and microsatellite markers. Genetics 158:787–809
Chakravarti A, Lasher LA, Reefer JE (1991) A maximum likeliwood method for estimating genome length using genetic linkage data. Genetics 128:175–182
Conner P, Brown S, Weeden N (1997) Randomly ampliWed polymorphic DNA-based genetic linkage maps of three apple cultivars. J Am Soc Hortic Sci 122:350–359
Dirlewanger E, Moing A, Rothan C, Svanella L, Pronier V, Guye A et al (1999) Mapping QTLs controlling fruit quality in peach [Prunus persica (L.) Batsch]. Theor Appl Genet 98:18–31
Dirlewanger E, Cosson P, Howad W, Capdeville G, Bosselut N, Claverie M et al (2004) Microsatellite genetic linkage maps of myrobalan plum and an almond-peach hybrid-location of root-knot nematode resistance genes. Theor Appl Genet 109:827–838. doi:10.1007/s00122-004-1694-9
Dirr MA (1998) Manual of woody landscape plants: their identification, ornamental characteristics, culture, propagation and uses, 5th edn. Stipes Publications, Champaign
Dolezel J, Göhde W (1995) Sex determination in dioecious plants Melandrium album and M. rubrum using high-resolution flow cytometry. Cytometry 19:103–106. doi:10.1002/cyto.990190203
Dolezel J, Sgorbati S, Lucretti S (1992) Comparison of three DNA fluorochromes for flow cytometric estimation of nuclear content in plants. Physiol Plant 85:625–631. doi:10.1111/j.1399-3054.1992.tb04764.x
Doležel J, Bartoš J, Voglmayr H, Greilhuber J (2003) Nuclear DNA content and genome size of trout and human. Cytometry 51A:127–128. doi:10.1002/cyto.a.10013
Doucleff M, Jin Y, Gao F, Riaz S, Krivanek AF, Walker MA (2004) A genetic linkage map of grape, utilizing Vitis rupestris and Vitis arizonica. Theor Appl Genet 109:1178–1187. doi:10.1007/s00122-004-1728-3
Faris JD, Laddomada B, Gill BS (1998) Molecular mapping of segregation distortion loci in Aegilops tauschii. Genetics 149:319–327
Fishman L, Kelly AJ, Morgan E, Willis JH (2001) A genetic map in the Mimulus guttatus species complex reveals transmission ratio distortion due to heterospecific interactions. Genetics 159:1701–1716
Fregene M, Angel F, Gomez R, Rodriguez F, Chavarriaga P, Roca W, Thome J, Bonierbale M (1997) A molecular genetic map of cassava (Manihot esculenta Crantz). Theor Appl Genet 95:431–441
Graham J, Smith K, Mackenzie K, Jorgenson L, Hackett C, Powell W (2004) The construction of a genetic linkage map of red raspberry (Rubus idaeus idaeus) based on AFLPs, genomic-SSR and EST-SSR markers. Theor Appl Genet 109:740–749. doi:10.1007/s00122-004-1687-8
Grattapaglia D, Sederoff R (1994) Genetic linkage maps of Eucalyptus grandis and Eucalyptus urophylla using a pseudo testcross mapping strategy and RAPD markers. Genetics 137:1121–1137
Gunatilleke CVS, Gunatilleke IAUN (1984) Some observations on the reproductive biology of three species of Cornus (Cornaceae). J Arnold Arbor 65:419–427
Hanley S, Barker JHA, Van Ooijen JW, Aldam C, Harris SL, Åhman I et al (2002) A genetic linkage map of willow (Salix viminalis) based on AFLP and microsatellite markers. Theor Appl Genet 105:1087–1096. doi:10.1007/s00122-002-0979-0
Hulbert S, Ilott T, Legg EJ, Lincoln S, Lander E, Michelmore R (1988) Genetic analysis of the fungus, Bremia lactucae, using restriction length polymorphism. Genetics 120:947–958
Kenis K, Keulemans J (2005) Genetic linkage maps of two apple cultivars (Malus × domestica Borkh) based on AFLP and microsatellite markers. Mol Breed 15:205–219. doi:10.1007/s11032-004-5592-2
Kosambi DD (1944) The estimation of map distance from recombination values. Ann Eugen 12:172–175
Kuang H, Richardson T, Carson S, Wilcox P, Bongarten B (1999) Genetic analysis of inbreeding depression in plus tree 85055 of Pinus radiata D Don I Genetic map with distorted markers. Theor Appl Genet 98:697–703. doi:10.1007/s001220051123
La Rosa R, Angiolillio A, Guerrero C, Pellegrini M, Rallo L, Besnard G et al (2003) A first linkage map of olive (Olea europea L.) cultivars using RAPD, AFLP, RFLP and SSR markers. Theor Appl Genet 106:1273–1282
Lalli DA, Abbott AG, Zhebentyayeva TN, Badenes ML, Damsteegt V, Polák J, Krška B, Salava J (2008) A genetic linkage map for an apricot (Prunus armeniaca L.) BC1 population mapping plum pox virus resistance. Tree Genet Genomes. doi:101007/s11295-007-0125-x
Lanteri S, Acquadro A, Comino C, Mauro R, Mauromicale G, Portis E (2006) A first linkage map of globe artichoke (Cynara cardunculus var scolymus L.) based on AFLP, S-SAP, M-AFLP and microsatellite markers. Theor Appl Genet 112:1532–1542. doi:10.1007/s00122-006-0256-8
Lowe KM, Walker MA (2006) Genetic linkage map of the interspecific grape rootstock cross Ramsey (Vitis champinii) × Riparia Gloire (Vitis riparia). Theor Appl Genet 112:1582–1592. doi:10.1007/s00122-006-0264-8
Lysák MA, Doležel J (1998) Estimation of nuclear DNA content in Sesleria (Poaceae). Caryologia 52:123–132
Maliepaard C, Jansen J, van Ooijen JW (1997) Linkage analysis in a full-sib family of an outbreeding plant species: overview and consequences for application. Genet Res 70:237–250. doi:10.1017/S0016672397003005
Mandl K, Santiago JL, Hack R, Fardossi A, Regner F (2006) A genetic map of Welschriesling × Sirius for the identification of magnesium-deficiency by QTL analysis. Euphytica 149:133–144. doi:10.1007/s10681-005-9061-8
Nealed B, William CG (1999) Restriction fragment length polymorphism mapping in conifers and applications to forest genetics and tree improvement. Can J Res 21:545–554. doi:10.1139/x91-076
Okogbenin E, Marin J, Fregene M (2008) QTL analysis for early yield in a pseudo F2 population of cassava. Afr J Biotechnol 7:131–138
Orton ER Jr (1985) Interspecific hybridization among Cornus florida, C. kousa, and C. nutallii. Proc Int Plant Prop Soc 35:655–661
Otto F (1990) DAPI staining of fixed cells for high-resolution flow cytometry of nuclear DNA. In: Crissman HA, Darzynkiewicz Z (eds) Methods in cell biology, vol 33. Academic, New York, pp 105–110
Pekkinen M, Vario S, Kuliu KKM, Käkkänen H, Smolander S, Viherä-Aarnio A et al (2005) Linkage map of birch, Betulla pendula Roth, based on microsatellites and amplified fragment length polymorphism. Genome 48:619–625. doi:10.1139/g05-031
Pilien K, Steinb_cken G, Hermann RG, Jung C (1993) An extended linkage map of sugar beet (Beta vulgaris L.) including nine putative lethal genes and the restorer gene X. Plant Breed 111:265–272. doi:10.1111/j.1439-0523.1993.tb00641.x
Porceddu A, Albertini E, Barcaccia G, Falistocco E, Falcinelli M (2002) Linkage mapping in apomictic and sexual Kentucky bluegrass (Poa pratensis L.) genotypes using a two way pseudo-testcross strategy based on AFLP and SAMPL markers. Theor Appl Genet 104:273–280. doi:10.1007/s001220100659
Quillet MC, Madjidian N, Griveau Y, Serieys H, Tersac Scaps M, Lorieux M et al (1995) Mapping genetic factors controlling pollen viability in an interspecific cross in Helianthus sect Helianthus. Theor Appl Genet 91:1195–1202. doi:10.1007/BF00220929
Radford AE, Ahles HE, Bell CR (1979) Manual of the vascular flora of the Carolinas. The University of North Carolina Press, Chapel Hill
Reed SM (2004) Self-incompatibility in Cornus florida. Hortic Sci 39:335–338
Rouppe van der Voort JNAM, van Eck HJ, van Zandvoort PM, Overmars H, Helder J, Bakker J (1999) Linkage analysis by genotyping of sibling populations: a genetic map for the potato cyst nematode constructed using a “pseudo-F2” mapping strategy. Mol Gen Genet 261:1021–1031. doi:10.1007/s004380051051
Scalfi WM, Troggio M, Piovani P, Leonardi S, Magnaschi G, Vendramin G et al (2004) A RAPD, AFLP and SSR linkage map, and QTL analysis in European beech (Fagus sylvatica L.). Theor Appl Genet 108:433–441. doi:10.1007/s00122-003-1461-3
Smith NR, Trigiano RN, Windham MT, Lamour KH, Finley LS, Wang X et al (2007) AFLP markers identify Cornus florida cultivars and lines. J Am Soc Hortic Sci 132:90–96
Sneizko RA, Zobel BJ (1988) Seedling height and diameter variation of various degrees of inbred and outcrossed progenies of loblolly pine. Silvae Genet 37:50–60
Sondur SN, Manshardt RM, Stiles JI (1996) A genetic linkage map of papaya based on random amplified polymorphic DNA markers. Theor Appl Genet 93:547–553
Sork VL, Smouse PE, Apsit VJ, Dyer RJ, Westfall RD (2005) A two-generation analysis of pollen pool genetic structure in flowering dogwood, Cornus florida (Cornaceae), in the Missouri Ozarks. Am J Bot 92:262–271. doi:10.3732/ajb.92.2.262
Strauss H, Lande R, Namkoong G (1992) Limitations of molecular-marker aided selection in forest tree breeding. Can J Res 22:1050–1061. doi:10.1139/x92-140
Tanksley SD, Ganal MW, Prince JP, Vicente de MC, Bonierbale MW, Broun P, Fulton TM, Giovannoni JJ, Grandillo S, Martin GB, Messeguer R, Miller JC, Miller L, Paterson AH, Pineda O, Roder MS, Wing RA, Wu W, Young ND (1992) High density molecular linkage maps of the tomato and potato genomes. Genetics 132:1141–1160
Tavoletti S, Veronesi F, Osborn TC (1996) RFLP linkage map of an alfalfa meiotic mutant based on an F1 population. J Hered 87:167–170
Tulsieram LK, Glaubitz JC, Kiss G, Carlson JE (1992) Single tree genetic linkage mapping in conifers using haploid DNA from megagametophyte. Biotechnology (N Y) 10:686–690. doi:10.1038/nbt0692-686
Vallejos CE, Sakiyama NS, Chase CD (1992) A molecular markerbased linkage map of Phaseolus vulgaris L. Genetics 131:733–740
Van Ooijen JW (2006) JoinMap® 4, software for the calculation of genetic linkage maps in experimental populations. Kyazma BV, Wageningen
Venkateswarlu M, Raje Urs S, Surendra Nath B, Shashidhar HE, Maheswaran M, Veeraiah TM, Sabitha MG (2006) A first genetic linkage map of mulberry (Morus spp.) using RAPD, ISSR, and SSR markers and pseudotestcross mapping strategy. Tree Genet Genomes 3:15–24. doi:10.1007/s11295-006-0048-y
Verde I, Lauria M, Dettori MT, Vendramin E, Balconi C, Micali S et al (2005) Microsatellite and AFLP markers in the Prunus persica [L. (Batsch)] × P. feganensis BC1 linkage map: saturation and coverage improvement. Theor Appl Genet 111:1013–1021. doi:10.1007/s00122-005-0006-3
Voorrips RE (2002) MapChart: software for the graphical presentation of linkage maps and QTLs. J Hered 93:77–78. doi:10.1093/jhered/93.1.77
Wang XW, Trigiano RN, Windham MT, DeVries RE, Scheffler BE, Rinehart TA et al (2007) A simple PCR procedure for discovering microsatellites from small insert libraries. Mol Ecol Notes 7:558–561. doi:10.1111/j.1471-8286.2006.01655.x
Wang XW, Trigiano RN, Windham MT, Scheffler BE, Rinehart TA, Spiers JM (2008) Development and characterization of simple sequence repeats for flowering dogwood (Cornus florida L.). Tree Genet Genomes. doi:101007/s11295-007-0123-z
Weeden NF, Hemmat M, Lawson DM, Loghi M, Manganaris AG, Reish BI et al (1994) Development and application of molecular marker linkage maps in woody fruit crops. Euphytica 77:71–75. doi:10.1007/BF02551464
Windham MT, Graham ET, Witte WT, Knighten JL, Trigiano RN (1998) Cornus florida ‘Appalachian Spring’: a white flowering dogwood resistant to dogwood anthracnose. HortScience 33:1265–1267
Windham MT, Witte WT, Trigiano RN (2003) Three white-bracted cultivars of Cornus florida resistant to powdery mildew. HortScience 38:1253–1255
Witte WT, Windham MT, Windham AS, Hale FA, Fare DC, Clatterbuck WK (2000) Dogwoods for American Gardens. The University of Tennessee Agriculture Extension Service, Knoxville
Acknowledgments
This work was supported by USDA Agreement # 58-6404-2-0057. Any information requirements can be addressed to Dr. Robert N. Trigiano at rtrigian@utk.edu. Mention of trade names or commercial products in this article is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the U.S. Department of Agriculture nor by The University of Tennessee.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wang, X., Wadl, P.A., Rinehart, T.A. et al. A linkage map for flowering dogwood (Cornus florida L.) based on microsatellite markers. Euphytica 165, 165–175 (2009). https://doi.org/10.1007/s10681-008-9802-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-008-9802-6