Abstract
Papaya (Carica papaya L.) is a versatile tropical fruit crop with economic and nutritional benefits. Assessing genetic diversity of the germplasm is essential for effective collection, conservation management and utilization of resources for breeding purposes. Genetic variability and population structure were studied for 55 papaya accessions including landraces, cultivars, and exotic collections, using 15 simple sequence repeat (SSR) markers. SSR analysis data revealed a total of 95 alleles with an average of 6.3 alleles per marker and an average heterozygosity of 0.75. All the markers were polymorphic, with an average PIC value of 0.72. The unweighted pair group method with arithmetic mean (UPGMA) dendrogram based on shared allele distance grouped the accessions into four main clusters. Genetic structure based on the structure algorithm identified two populations, whereas discriminant analysis of principal components revealed four distinct genetic clusters. Remarkably, these clusters were not linked to the geographic regions. The analysis of molecular variance (AMOVA) indicated a variation of 75% within individuals and 22% between populations. Fst = 0.216 and Nm = 0.205 showed moderate genetic relationship within populations. These findings have implications for marker-assisted breeding, improvement of conservation strategies, detection of duplicates, and framing policies for sustainable crop utilization.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Papaya is the sole representative of the genus Carica in the family Caricaceae, and its wild relatives are now classified under Vasconcellea (Badillo 2000). Papaya is believed to have originated in the Mesoamerican center (south of Mexico and Central America) (Fuentes and Santamaría, 2014) and it was introduced to India by Spaniards in the sixteenth century (Singh 1990). Currently, it is cultivated in tropical and subtropical regions worldwide. Globally, papaya is cultivated in an area of 0.48 million ha with a production of 14.1 million metric tonnes. India contributes 40% (5.54 million tonnes) of papaya production with 30% (0.14 million ha) of the global papaya cultivated area (FAOSTAT 2022).
Papaya, with its rich nutrient profile and wide adaptability, plays a vital role in food and nutritional security (Pinnamaneni 2017) in tropical and subtropical regions. Its cultivation is profitable globally owing to its demand and efficient productivity. In addition, dried milky latex from mature papaya, called papain, has significant applications in biotechnology and industrial sectors (Elsson et al. 2019). It is particularly used in industries such as pharmaceuticals, breweries, tanneries, cosmetics, detergents (Saran and Choudhary 2013), and the processing of cheese, meat, and fish (Mamboya and Amri 2012).
Genetic resources of crops are essential for food security (Toledo and Burlingame 2006). A wide gene pool aids in understanding evolutionary relationships and breeding better traits such as disease resistance and fruit quality. A larger population increases the chances of identifying individuals with the desired traits in various environments. Morphological and agronomic traits, such as plant height, juvenile period, flower initiation, leaf shape, fruit shape, flesh color, stamen abortion, carpelloidy, and fruit yield, can vary owing to genotype and environment interaction (Campostrini and Glenn 2007; Silva et al. 2007; Kumar et al. 2015; Kaluram et al. 2018). Field observations can help to estimate genetic diversity; however, environmental factors can affect the same gene differently (Weckwerth et al. 2020), making it difficult to draw conclusions. Genotyping is the most reliable method because it is unaffected by environmental factors and can identify variations at the genome level.
Genotyping using molecular markers have been used for germplasm characterization and conservation for many years. A large extent of genetic diversity has been reported within Caricaceae and the genus Carica using molecular marker studies. Different molecular markers have been used, such as random amplified polymorphic DNA (RAPD) (Stiles et al. 1993; Jobin-Décor et al. 1997), restriction fragment length polymorphism (RFLP), and amplified fragment length polymorphism (AFLP) (Van Droogenbroeck et al. 2002; Kim et al. 2002; Ratchadaporn et al. 2007; Oliveira et al. 2011), inter-simple sequence repeats (ISSR) (Costa et al. 2011; Kanupriya et al. 2012), and simple sequence repeats (SSR) (Oliveira et al. 2010a, b; Matos et al. 2013; Sengupta et al. 2013; Pirovani et al. 2021) were used to analyze genetic diversity. Among these, SSR markers are considered robust molecular tools for the analysis of genetic diversity because of their abundance in the genome and their high reproducibility (Eustice et al. 2008). In addition, SSR markers have been used for sex identification (Parasnis et al. 1999), segregating populations (Pinto et al. 2013), DNA fingerprinting (Vitoria et al. 2004), and genetic mapping (Blas et al. 2012).
In India, studies on the genetic diversity of papaya have been conducted based on morphological traits and conventional molecular markers (Singh et al. 1997; Singh and Kumar 2010; Sudha et al. 2013; Saran et al. 2015). However, the extent of genetic diversity within the active germplasm of papaya remains unexplored. The Tamil Nadu Agricultural University (TNAU) has a long history of collecting papaya genotypes (Ram 2005) and releasing five inbred cultivars and three hybrids. Here, we maintained a diverse population of papaya genotypes consisting of landraces, cultivars, improved cultivars, and exotic collections. Despite its potential significance, evaluation of genetic diversity among the germplasm collections available in TNAU using molecular markers has not been attempted. Fifteen SSR primers were used to analyze 55 papaya accessions of the TNAU papaya germplasm. The objective of this study was to assess the genetic variation within the germplasm and determine its population structure. The results provide insights into genetic diversity and population structure, aiding conservation management, targeted breeding, and collection expansion. In addition, this study can be instrumental in framing policies related to germplasm conservation and utilization.
Material and methods
Plant material
Seeds of 55 papaya accessions were procured from the germplasm repository of the Department of Fruit Crops, Horticultural College & Research Institute, TNAU, Coimbatore. Subsequently, the seeds were sown in polybags and after 45 days, a polybag containing five to six seedlings was transplanted (spacing 1.8 m X 1.8 m) into the field at the College Orchard, TNAU, Coimbatore. The experiment conducted using randomized block design with 15 plants per accession in experimental plot. A list of the 55 papaya accessions is presented in Table 1.
Genomic DNA isolation
At the fruit maturation stage, fourth leaf from the top of tree was collected from the selected female or hermaphrodite plant. Healthy papaya leaves were collected, a Genomic DNA was extracted using the CTAB method (Doyle 1991). DNA quality was determined using 0.8% agarose gel, and the quantity and purity were recorded using a spectrophotometer (NanoDrop1000c, Thermo Scientific). The extracted DNA was diluted to 50 ng/µL and stored at − 20 °C until further analysis.
SSR analysis
A set of 16 SSR primers (Table 2) were selected from the microsatellite sequences developed by Perez et al. (2006). Reaction mixture for PCR assay:10 µL containing 1.0 µL of reaction buffer (10X with 20 mM MgCl2), 0.2 µL of 10 mM dNTP, 0.5 µL from forward and reverse primers of 10 µM, 1.0 µL of genomic DNA and 0.5 U of Taq polymerase. The amplification reaction was performed as follows: initial denaturation at 94 °C for 4 min, 35 cycles of denaturation at 94 °C for 1 min, primer annealing temperature (adjusted according to primers) for 1 min, primer extension 72 °C for 45 s, final extension at 72 °C for 4 min and finally, hold at 4 °C. The amplified PCR products were resolved by agarose gel electrophoresis (3%) and visualized using a gel documentation system (Alpha Imager, USA). The amplicon size was measured using a Takara 100 bp ladder.
SSR-based diversity analysis
The gel images of the SSR bands were scored using Gel Analyzer (Version 19.1 (www.gelanalyzer.com) by Istvan Lazar Jr., PhD and Istvan Lazar Sr., PhD, CSc) based on the molecular weight and data were recorded. From the recorded molecular data, the number of alleles, effective alleles, Shannon’s information index, observed heterozygosity, and expected and unbiased heterozygosity were computed using the GenAIex software (Version:6.0.5) (Peakall and Smouse 2012). Power marker (Liu and Muse 2005) was used to calculate the allele frequency and polymorphism information content (PIC) of the markers and generate a unweighted pair group method with arithmetic mean (UPGMA) dendrogram based on the shared allele frequency.
Population structure analysis
A Bayesian model of clustering was performed using STRUCTURE V.2.3.4 (Pritchard, 2000) to categorize the individuals into clusters (subpopulations). Without prior population information, the parameters were configured as an admixture model with correlated allelic frequencies. Ten independent runs with K values ranging from 1 to 10 were performed, with a burn-in period of 500,000 iterations and 500,000 Monte Carlo–Markov iterations. The generated output was compressed and uploaded to STRUCTURE HARVESTER V.0.9.94 (http://taylor0.biology.ucla.edu/structureHarvester/) (Earl and Holdt 2012). This software was used to determine the best K value, as outlined by Evanno et al. 2005. Individuals were assigned to clusters using a membership coefficient (q) and samples showing q < 0.8 termed as “genetic admixture” within that particular cluster.
Discriminant analysis of principal components was performed for the SSR dataset in R (version4.3.1) using the adegenet package (Jombart 2008). The SSR dataset was imported using the poppr package (Kamvar et al. 2014). The major advantage of DAPC is that it is not reliant on population genetics models, such as Hardy–Weinberg equilibrium or linkage equilibrium (Jombart et al. 2010). Data was first transformed into PCA, followed by a discriminant analysis of the retained principal components (PC). First, the clusters were identified using the find.clusters function based on the K-means algorithm, with K values varying from 1 to 10. The number of clusters was chosen based on the Bayesian information clustering (BIC) value. Next, the number of principal components (PC’s) was retained using the a-score optimization method function from the adegenet. The final cluster was generated using discriminant analysis.
Analysis of molecular variance (AMOVA)
Genetic differentiation within the population and individuals was determined using AMOVA (Excoffier et al. 1992) implemented in the GenAlex 6.503 software. To calculate the significance among the populations, pairwise Fst values and gene flow (Nm) were computed.
Results
Assessment of polymorphisms in SSR loci
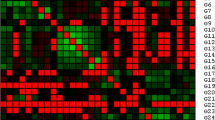
A set of 15 SSR markers was efficiently amplified in DNA fragments of 55 accessions of papaya and the results are given in Table 3. The SSR markers generated 95 alleles in all accessions, with an average of 6 alleles per marker. The number of alleles detected per primer varied from four to ten. The lowest number of alleles was recorded for the three primers S285, mcpCIR09, and mcpCIR16, whereas the highest number of alleles was present for mcpCIR28. The number of effective alleles ranged from 3.05 (mcpCIR16) to 7.70 (mcpCIR28), with an average of 4.37. The mean major allele frequency was 0.34, with a range of 0.23—0.55. The most frequent allele was recorded in S 285 and the least frequent alleles were in mcpCIR28.
The Shannon’s information index (I) was highest in mcpCIR28 (2.15) and lowest in S 285 (0.96). The observed heterozygosity ranged from 0.00 to 0.27 with an average value of 0.03. The expected heterozygosity or gene diversity detected by all SSR loci varied from 0.75 to 0.87, with an average value of 0.75. The polymorphism information content of the loci ranged from 0.48 (S 285) to 0.85 (mcpCIR28), with an average value of 0.72. The size of the alleles produced by the 15 SSR primers ranged from 67 to 780 bp.
Dendrogram
Based on the SSR marker data, a neighbor-joining tree of 55 accessions of papaya was constructed using the unweighted pair group method with an arithmetic mean (UPGMA) algorithm (Fig. 1). The dendrogram clearly grouped the 55 papaya accessions into four groups. Group 1 (G1) contained 13 papaya accessions and was divided into two subgroups: two accessions from IIHR-Bangalore, Sunrise Solo, CO.7, and Waimanalo in one group and seven exotic collections in another group. Group 2 (G2) was composed of 11 accessions with two subgroups, four local collections in one subgroup, and four exotic collections, Washington, Tainung II, and PAU selection were placed in another subgroup. Group 3 (G3) included 13 accessions, including two subgroups consisting of TNAU cultivars (CO.1 (R &Y), CO.2 (Y), CO.4(R &Y) and CO.5 (R)) in subgroup 1 and five nearby local collections grouped in subgroup 2. Group 4 (G4) was separated into two groups: 18 accessions consisting of three exotic collections (Singapore, Mexican, EC.611 100), one inter-varietal hybrid (CO.3 × Washington), one local collection (Perur), six Pusa varieties (Pusa Dwarf, Pusa Giant, Pusa Delicious, Pusa Majesty, Giant) and their derivatives (CO 5 and CO 6), and two cultivars from the Madhya Pradesh region (Barwani (R&Y)).
Population structure
To understand the population structure of the 55 accessions of papaya, Bayesian clustering analysis using Structure software and discriminant analysis of principal components were performed. The optimal K value was obtained using methods described by Pritchard et al. (2000) and Evanno et al. (2005). As shown in the figure (Fig. 2), the highest delta K value was K = 2. The bar plot of cluster K = 2 showed that out of 55 accessions, 31 accessions were grouped in one population and 24 accessions in another population, of which one accession was genetically admixed (Fig. 3). Population I consisted of most of the dioecious accessions, and population II consisted of gynodioecious accessions.
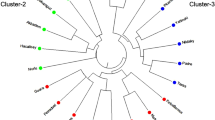
The results from the DAPC method revealed four distinct clusters (Fig. 4), which corresponded to the four BIC values obtained using the find.clusters function (Supplementary Fig. 1). Principal components were retained using an a-score optimization method (Supplementary Fig. 2). Clusters were formed by retaining the first five major principal components (cumulative variance = 50%), with three discriminant eigenvalues. A total of 12 accessions were assigned to Cluster I, 14 to Cluster II, 17 to Cluster III, and 12 to Cluster IV (Fig. 5). Varieties derived from Sunrise Solo and local accessions are grouped in cluster I. In comparison with dendrogram Group 1, the six accessions were similar in DAPC Cluster I. Cluster II grouped the varieties released from TNAU, Coimbatore and accessions collected in nearby areas. Cluster III comprises Pusa varieties and varieties derived from Pusa cultivars. Both DAPC clusters II and III were similar to dendrogram groups G3 and G4. Cluster IV grouped all the exotic collections in the germplasm, and Group 2 was in contrast to DAPC cluster IV, which contained only four similar accessions.
Genetic diversity of the identified populations by DAPC analysis
The gene diversity of the populations generated from the DAPC analysis was calculated, and the results are presented in Table 4. Among the populations, cluster III recorded the highest number of alleles (4.20), whereas the lowest number of alleles (2.60) was recorded in cluster IV. Similarly, the allele frequency and number of effective alleles were highest in cluster III (4.06 and 3.01, respectively) and lowest in cluster IV (2.60 and 2.03, respectively). Cluster III recorded the highest expected heterozygosity of 0.63, and Cluster IV recorded the lowest value (0.43).
Analysis of molecular variance within and among the population of papaya accessions based on results of DAPC analysis
The populations derived from DAPC analysis were tested for genetic differentiation using SSR genotypic data. The extent of genetic variability between populations, within individuals, and among individuals in the germplasm was analyzed using AMOVA (Table 5). The analysis showed that 75% of the variation existed between individuals, which was significantly higher than the variation obtained between the population (22%) and within individuals (3%). The Fst value was 0.216 (0.15 < Fst < 0.25) indicating a moderate level of genetic differentiation among the population. The Nm value of 0.905 indicated low gene flow among the populations. The pairwise Fst value was highest (0.29) between Clusters II and IV, and the lowest Fst value was between Clusters II and III. The gene flow (Nm) was highest (1.26) between clusters II and III (Supplementary Table 1).
Discussion
Molecular markers provide a comprehensive understanding of the genetic diversity and population structure of germplasms without any environmental influence. SSR markers are better suited for germplasm diversity analysis because they are easy to use, highly polymorphic, and reliable (Powell et al. 1996; Varshney et al. 2007). Earlier studies have reported that papaya contains abundant SSRs in its genome and is more useful for detailed genetic studies of population structure, hybrid testing, evolutionary studies, and QTL mapping (Santos et al. 2003; Perez et al. 2006; Eustice et al. 2008; Oliveira et al. 2011; Matos et al. 2013). In this study, using SSR genotypic data, the genetic diversity among 55 selected accessions of papaya was evaluated to understand the genetic variation and existing population structure between individuals.
In this study, the choice of markers was based on previous study conducted by Perez et al. 2006. The set of SSR markers used in this study provided a distinct genetic structure of the individuals in the papaya germplasm. Fifteen polymorphic simple sequence repeat (SSR) markers revealed 95 alleles across 55 papaya germplasm accessions. The alleles per locus ranged from 4 to 10, with an average of 6.3. This is lower than the 7 alleles per locus reported by Sengupta et al. (2013) for 34 accessions, including Indian and non-Indian accessions. Our results are similar to those of Ocampo Perez et al. (2006), who found an average of 6.6 alleles per locus in 72 accessions using 15 SSR markers, and Hasibuzzaman et al. (2020), who reported six alleles per locus for 34 genotypes with 10 SSR markers. In contrast, De Oliveira et al. (2010a) found 4.02 alleles per marker in 48 papaya accessions with 59 SSR markers, whereas Matos et al. (2013) reported 4.08 alleles per marker in 96 accessions with 15 microsatellite markers. The high number of alleles in the papaya germplasm may be due to the collection and conservation of accessions from all papaya-growing regions in India and exotic collections. In India, over the past 500 years since the papaya introduction, it has been naturalized and widely cultivated, leading to considerable genetic diversity. A wide range of cultivars exists in India, including primitive types, local adaptive cultivars, minor cultivars and principal cultivars released from Indian Agricultural Research Institute—Regional Station at Pusa, Tamil Nadu Agricultural University, Coimbatore and Indian Horticultural Research Institute, Bengaluru (Ram 2005).
Gene diversity and polymorphism information content
Nei’s gene diversity or expected heterozygosity and polymorphism information content is the reliable measure for assessing genetic variation in the population. The average gene diversity in this study was 0.75, similar to 31 papaya genotypes (0.74) from various countries including Bangladesh (Hasibuzzaman et al. 2020). This is higher than Caribbean populations (0.37–0.69) (Ocampo Perez et al. 2007), USDA germplasm (0.58) (Luciano-Rosario et al. 2018), and Costa Rica’s natural populations (0.62) (Brown et al. 2012).
Botstein et al. (1980) stated that PIC value > 0.5 as high locus diversity, PIC < 0.25 as limited diversity and values between 0.25 and 0.50 as intermediate diversity. Our germplasm’s average PIC value of 0.72 indicates high genetic diversity. Comparatively, Hasibuzzaman et al. (2020) reported a value of 0.70, showing a similar level of diversity in the Bangladeshi germplasm. Sengupta et al. (2013) observed a slightly higher PIC value of 0.74 in analyzing Caricaceae accessions and Asudi et al. (2013) reported the highest of 0.81 indicating diverse Kenyan germplasm. In the Embrapa papaya genebank, Oliveira et al. (2010a) found 0.52 in 30 select accessions and Matos et al. (2013) reported 0.47 in 96 accessions, indicating lower diversity in their genebank accessions than in our study.
Genetic structure of the germplasm
The population structure of germplasm facilitate effective management and utilization of resources. SSR analysis data clearly revealed the genetic similarity cluster between the accessions based on the shared allele distance computed using the UPGMA method. Fifty-five accessions in the germplasm were divided into four main groups and subgroups within it. Group 1 comprised Waimanalo, Sunrise Solo, IIHR-39, IIHR-57, CO 7, Malaysian Long, Singapore, and six other exotic collections. Some of these accessions were interlinked to the Hawaiian cultivar “Solo”, from which Waimanalo and Sun rise solo were derived (Ram 2005). Accession IIHR-39 has sunrise solo as the main parent, and IIHR-57 is derived from Arka Surya and Tainung-I. Accessions IIHR-39 (Arka Surya) and IIHR-57(Arka Prabhath) were released as cultivars from IIHR, Bengaluru, suitable for the institute region (Mitra and Dinesh 2019).
Group 2 had two subgroups: subgroup 1 included three local accessions (Sathyamangalam Dwarf, Valliyur collection, and Red flesh) and PAU selection from PAU, Ludhiana were closely related. These collections were not related to a specific region. The second subgroup included an open-pollinated accession and Tainung II from Taiwan, four exotic collections (EC.100 211, EC.100 135, EC.100 012, and EC.100 064) and Washington. “Washington” papaya has a distinct character of purple-colored petiole and it has been domesticated for a long time in the Maharashtra region of India (Ram 2005).
Group 3 comprised most accessions belonging to the Coimbatore region of Tamil Nadu. The cultivars CO.1 (red), CO.1 (yellow), CO.2 (yellow), CO.4(yellow), CO.4 (red), and CO.5 (yellow) were closely connected in subgroup 1, and the second subgroup had local collections (MD.13 (Veda Patti), M1 (OP), Local Acc (Y), and MD Telungu palayam), Carica pink petiole, and PKM-1 long from Periyakulam, Theni. Over the past five decades, TNAU, Coimbatore have made significant advancements in papaya crop improvement, resulting in the release of eight elite cultivars from CO.1 to CO.8 (Mitra and Dinesh 2019). The narrow genetic diversity among the TNAU released cultivars, as revealed in the present study, could be due to the parental materials involved in the development of these cultivars. As the local accessions were collected within 20 km of Coimbatore, these genotypes have common alleles, probably because of the exchange of seeds among farmers (Matos et al. 2013), therefore, limited genetic differentiation existed between these groups.
Lastly, group 4 was divided into two subgroups, one group dominated by Pusa varieties such as Pusa Dwarf, Pusa Giant, Pusa Majesty, Pusa Delicious, CO.6 (selection from Pusa Majesty), CO.5 (selection from Washington), Barwani (locally adapted genotype from Madhya Pradesh) and Manila Pink from the Philippines. Interestingly, this subgroup consisted of the hybrid Carica (wild) X CO-6 (CP-50), which was reported to be a PRSV-tolerant genotype by Balamohan et al. (2010). CO.5, derived from the Washington variety (Sharma and Mitra 2014), was distantly related to its parent. This can be attributed to many factors, such as the outcrossing nature of papaya increasing genetic distance (Kim et al. 2002), and a limited number of SSR markers influencing differentiation. In another subgroup, accessions such as Mexican, Singapore, and Perur (local collection from Coimbatore). Nevertheless, this subgroup was comprised of a mixture of accessions collected from various regions. Ocampo Perez et al. (2007) analyzed genotypes from various regions, including Costa Rica, Colombia, Venezuela, Guadeloupe, and the Antillean islands and found geographic based clustering regions with few exceptions. In our study, despite the small number of accessions are region specific, we did not observe a correlation between geographic region and cluster formation.
In addition to the dendrogram, we applied both structure and DAPC approaches to infer the population structure of the 55 accessions. A model-based approach by structure distinguished the germplasm accessions into two populations based on the delta value (K = 2), while Hasibuzzaman et al. (2020) reported six populations from 31 papaya accessions collected around the world. However, the DAPC method revealed a remarkably distinct clustering pattern that deviated significantly from the results obtained using structure. DAPC analysis successfully classified the 55 selected accessions of papaya into four distinct clusters, irrespective of their region of collection. The clustering pattern derived from the DAPC analysis exhibited a close alignment with the hierarchical structure depicted in the dendrogram, except for Clusters I and IV. The difference in clustering is attributed to the methodologies and principles underlying both the analytical approaches. Using the DAPC method, Matos et al. (2013) clearly identified that the papaya germplasm of 96 selected accessions was classified into six distinct clusters; however, in contrast to our study, the DAPC classified clusters were concordant with Bayesian clustering by STRUCTURE algorithm. However, Campoy et al. (2016) and Mariette et al. (2010) reported that DAPC analysis yielded a comprehensive clustering pattern within the germplasm compared with the results obtained from structure analysis.
Molecular variation in the populations
The results of molecular variation between the populations indicated 22%, whereas a variation of 75% among the individuals of the population represented the overall genetic diversity. The increased variation is possibly due to the reproductive biology of papaya with three sex forms (Matos et al. 2013), evolutionary forces such as the hybridization of the most divergent parents (Goulet et al. 2017), and the introduction of exotic collections in germplasm (Scherlosky et al. 2018). Wright (1965) stated that Fst (Fixation index) close to 0 signifies low genetic differentiation, 0 to 1 indicates moderate and close to 1 shows high genetic differentiation. Nm (Gene flow) value below 1 signify limited gene flow within the population. Fst (0.22) and Nm (0.21) values showed moderate genetic differentiation and limited gene flow.
Conclusion
In this study, 55 accessions selected from papaya germplasm collected worldwide were genotyped using 15 SSR markers. Allelic richness and extensive gene diversity indicated broad genetic variation in the germplasm. DAPC and UPGMA analyses separated the accessions into four subpopulations, irrespective of their region. These findings can potentially optimize the expansion of collection, effective management of resources, parental line selection for hybridization, and tailor breeding programs.
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author upon reasonable request.
References
Asudi GO, Ombwara F, Rimberia F, Ateka E, Nyende A, Wamocho L (2013) Evaluating diversity among Kenyan papaya germplasm using simple sequence repeat markers. Afr J Food Agric Nutr Dev 13(1):7307–7324. https://doi.org/10.18697/ajfand.56.11950
Badillo (2000) Carica L. vs. Vasconcella St. Hil. (Caricaceae)—Google Scholar. Retrieved 28 Sep, 2023
Balamohan TN, Auxcilia J, Thirugnanavel A, Manoranjitham SK (2010) CP-50: a papaya ring spot virus (PRSV) tolerant papaya genotype under field conditions. In: Acta horticulturae, international symposium on papaya 851:153–156. https://doi.org/10.17660/ActaHortic.2010.851.22
Blas AL, Yu Q, Veatch OJ, Paull RE, Moore PH, Ming R (2012) Genetic mapping of quantitative trait loci controlling fruit size and shape in papaya. Mol Breed 29(2):457–466. https://doi.org/10.1007/s11032-011-9562-1
Botstein D, White RL, Skolnick M, Davis RW (1980) Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am J Hum Genet 32(3):314–331
Brown JE, Bauman JM, Lawrie JF, Rocha OJ, Moore RC (2012) The structure of morphological and genetic diversity in natural populations of Carica papaya (Caricaceae) in Costa Rica: morphological and genetic diversity of natural papaya. Biotropica 44(2):179–188. https://doi.org/10.1111/j.1744-7429.2011.00779.x
Campostrini E, Glenn DM (2007) Ecophysiology of papaya: a review. Braz J Plant Physiol 19(4):413–424. https://doi.org/10.1590/S1677-04202007000400010
Campoy JA, Lerigoleur-Balsemin E, Christmann H, Beauvieux R, Girollet N, Quero-García J et al (2016) Genetic diversity, linkage disequilibrium, population structure and construction of a core collection of Prunus avium L. landraces and bred cultivars. BMC Plant Biol 16(1):49. https://doi.org/10.1186/s12870-016-0712-9
da Costa FR, Pereira TNS, Gabriel APC, Pereira MG (2011) ISSR markers for genetic relationships in Caricaceae and sex differentiation in papaya. Crop Breed Appl Biotechnol 11(4):352–357. https://doi.org/10.1590/S1984-70332011000400009
da Silva J, Rashid Z, Nhut DT, Sivakumar D, Gera A, Souza MT, Tennant P (2007) Papaya (Carica papaya L.) biology and biotechnology. Tree for Sci Biotechnol 1(1):47–73
De Oliveira EJ, Amorim VBO, Matos ELS, Costa JL, Da Silva CM, Pádua JG, Dantas JLL (2010a) Polymorphism of microsatellite markers in papaya (Carica papaya L.). Plant Mol Biol Rep 28(3):519–530. https://doi.org/10.1007/s11105-010-0180-6
De Oliveira EJ, Dos Santos Silva A, De Carvalho FM, Dos Santos LF, Costa JL, De Oliveira Amorim VB, Dantas JLL (2010b) Polymorphic microsatellite marker set for Carica papaya L. and its use in molecular-assisted selection. Euphytica 173(2):279–287. https://doi.org/10.1007/s10681-010-0150-y
de Oliveira EJ, Costa JL, dos Santos LF, de Carvalho FM, Silva Ados S, Dantas JLL (2011) Molecular characterization of papaya genotypes using AFLP markers. Rev Bras Frutic 33(3):849–858. https://doi.org/10.1590/S0100-29452011000300020
Doyle J (1991) DNA protocols for plants. In: Hewitt GM, Johnston AWB, Young JPW (eds) Molecular techniques in taxonomy. Springer, Berlin Heidelberg, pp 283–293
Earl DA, vonHoldt BM (2012) Structure harvester: a website and program for visualizing structure output and implementing the Evanno method. Conserv Genet Resour 4(2):359–361. https://doi.org/10.1007/s12686-011-9548-7
Eustice M, Yu Q, Lai CW, Hou S, Thimmapuram J, Liu L et al (2008) Development and application of microsatellite markers for genomic analysis of papaya. Tree Genet Genomes 4(2):333–341. https://doi.org/10.1007/s11295-007-0112-2
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software structure: a simulation study. Mol Ecol 14(8):2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x.Pubmed:15969739
Excoffier L, Smouse PE, Quattro JM (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 131(2):479–491. https://doi.org/10.1093/genetics/131.2.479
FAOSTAT (2022) FAO statistical database https://www.fao.org/faostat/en/#data/QCL
Fuentes G, Santamaría JM (2014) Papaya (Carica papaya L.): origin, domestication, and production. In: Ming R, Moore PH (eds) Genetics and genomics of papaya. Springer, New York p, pp 3–15
Goulet BE, Roda F, Hopkins R (2017) Hybridization in plants: old ideas, new techniques. Plant Physiol 173(1):65–78. https://doi.org/10.1104/pp.16.01340
Hasibuzzaman ASM, Islam AKMA, Miah MdG, Hasan M (2020) Phylogeographic diversity and population structure of Carica papaya L. revealed through nuclear microsatellites. Braz J Bot 43(1):147–154. https://doi.org/10.1007/s40415-020-00594-8
Jobin-Decor MP, Graham GC, Henry RJ, Drew RA (1997) RAPD and isozyme analysis of genetic relationships between Carica papaya and wild relatives. Genet Resour Crop Evol 44(5):471–477. https://doi.org/10.1023/A:1008644901727
Jombart T (2008) Adegenet: AR package for the multivariate analysis of genetic markers. Bioinformatics 24(11):1403–1405. https://doi.org/10.1093/bioinformatics/btn129
Jombart T, Devillard S, Balloux F (2010) Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genet 11(1):94. https://doi.org/10.1186/1471-2156-11-94
Kaluram, Prakash J, Singh SK, Goswami AK, Singh P, Hussain Z, Singh AK (2018) Genetic diversity in qualitative and quantitative traits of papaya. Indian J Hortic. https://doi.org/10.5958/0974-0112.2018.00031.2
Kamvar ZN, Tabima JF, Grünwald NJ (2014) Poppr: an R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. PeerJ 2:e281. https://doi.org/10.7717/peerj.281
Kanupriya C, Shobhana M, Vasugi C, Aswath C, Radhika V, Reddy L, Dinesh M (2012) Genetic relationship among papaya (Carica papaya) and wild papaya (Vasconcellea species) using RAPD and ISSR markers. Indian J Agric Sci 82(4):366–369
Kim MS, Moore PH, Zee F, Fitch MM, Steiger DL, Manshardt RM et al (2002) Genetic diversity of Carica papaya as revealed by AFLP markers. Genome 45(3):503–512. https://doi.org/10.1139/g02-012
Kumar M, Prasad KM, Prakash S, Kumar S (2015) Evaluation of genetic variability, genetic advance, heritability and character association for yield and its contributing traits in papaya (Carica papaya L.). Soc Plant Res. 28(2):99–102
Liu K, Muse SV (2005) PowerMarker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21(9):2128–2129. https://doi.org/10.1093/bioinformatics/bti282
Luciano-Rosario D, Cruz-Saavedra LA, Siritunga D (2018) Genetic diversity of Puerto Rican farmer-held papaya (Carica papaya) using SSR markers. Hortic Sci 53(8):1109–1114. https://doi.org/10.21273/HORTSCI12943-18
Mamboya EA, Amri E (2012) Papain, a plant enzyme of biological importance: a review. Am J Biochem Biotechnol 8(2):99–104. https://doi.org/10.3844/ajbbsp.2012.99.104
Mariette S, Tavaud M, Arunyawat U, Capdeville G, Millan M, Salin F (2010) Population structure and genetic bottleneck in sweet cherry estimated with SSRs and the gametophytic self-incompatibility locus. BMC Genet 11(1):77. https://doi.org/10.1186/1471-2156-11-77
Matos ELS, Oliveira EJ, Jesus ON, Dantas JLL (2013) Microsatellite markers of genetic diversity and population structure of Carica papaya. Ann Appl Biol 163(2):298–310. https://doi.org/10.1111/aab.12053
Mitra SK, Dinesh MR (2019) Papaya breeding in India—achievements and future thrust. Acta Hortic 1250:23–28. https://doi.org/10.17660/ActaHortic.2019.1250.4
Ocampo Pérez J, Coppens D’Eeckenbrugge G, Risterucci AM, Dambier D, Ollitrault P (2007) Papaya genetic diversity assessed with microsatellite markers in germplasm from the Caribbean region. Acta Hortic 740:93–101. https://doi.org/10.17660/ActaHortic.2007.740.9
Parasnis AS, Ramakrishna W, Chowdari KV, Gupta VS, Ranjekar PK (1999) Microsatellite (GATA)n reveals sex-specific differences in papaya. Theor Appl Genet 99(6):1047–1052. https://doi.org/10.1007/s001220051413
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in excel. Population genetic software for teaching and research—an update. Bioinformatics 28(19):2537–2539. https://doi.org/10.1093/bioinformatics/bts460
Perez JO, Dambier D, Ollitrault P, D’Eeckenbrugge GC, Brottier P, Froelicher Y, Risterucci AM (2006) Microsatellite markers in Carica papaya L.: isolation, characterization and transferability to Vasconcellea species. Mol Ecol Notes 6(1):212–217. https://doi.org/10.1111/j.1471-8286.2006.01197.x
Pinnamaneni R (2017) Nutritional and medicinal value of papaya (Carica papaya Linn.). World J Pharm Pharm Sci 6(8):2559–2578. https://doi.org/10.20959/wjpps20178-9947
Pinto FO, Pereira MG, Luz LN, Cardozo DL, Ramos HCC, Macedo CMP (2013) Use of microsatellite markers in molecular analysis of segregating populations of papaya (Carica papaya L.) derived from backcrossing. Genet Mol Res 12(3):2248–2259. https://doi.org/10.4238/2013.July.8.6
Pirovani AAV, Ramos HCC, Duarte RP, Miranda DP, Bohry D, Santana JGS et al (2021) Molecular characterization of elite lines of papaya (Carica papaya L.) via SSR markers. Funct Plant Breed J 3(1):49–58
Powell W, Morgante M, Andre C, Hanafey M, Vogel J, Tingey S, Rafalski A (1996) The comparison of RFLP, RAPD, AFLP and SSR (microsatellite) markers for germplasm analysis. Mol Breed 2(3):225–238. https://doi.org/10.1007/BF00564200
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155(2):945–959. https://doi.org/10.1093/genetics/155.2.945
Ram M (2005) Papaya. In: Introduction and production technology. Indian Council of Agricultural Research, New Delhi. pp. 2 & 89
Ratchadaporn J, Sureeporn K, Khumcha U (2007) An analysis on DNA fingerprints of thirty papaya cultivars (Carica papaya L.), grown in Thailand with the use of amplified fragment length polymorphisms technique. Pak J Biol Sci 10(18):3072–3078. https://doi.org/10.3923/pjbs.2007.3072.3078
Santos SC, Ruggiero C, Silva CL, Lemos EG (2003) A microsatellite library for Carica papaya L. cv. Sunr Solo Rev Bras Frutic 25:263–267. https://doi.org/10.17660/ActaHortic.2007.740.9
Saran PL, Choudhary R (2013) Drug bioavailability and traditional medicaments of commercially available papaya: a review. Afr J Agric Res 8(25):3216–3223
Saran PL, Choudhary R, Solanki IS, Patil P, Kumar S (2015) Genetic variability and relationship studies in new Indian papaya (Carica papaya L.) germplasm using morphological and molecular markers. Turk J Agric for 39(2):310–321. https://doi.org/10.3906/tar-1409-148
Scherlosky A, Marchioro VS, Franco A, Braccini AL, Schuster I (2018) Genetic variability of Brazilian wheat germplasm obtained by high-density SNP genotyping. Crop Breed Appl Biotechnol 18(4):399–408. https://doi.org/10.1590/1984-70332018v18n4a59
Sengupta S, Das B, Prasad M, Acharyya P, Ghose TK (2013) A comparative survey of genetic diversity among a set of Caricaceae accessions using microsatellite markers. Springerplus 2(1):345. https://doi.org/10.1186/2193-1801-2-345
Sharma SK, Mitra SK (2014) An overview of biodiversity in papaya in India with future prospects. Acta Hortic 1022(1022):149–155. https://doi.org/10.17660/ActaHortic.2014.1022.19
Singh ID (1990) Papaya. Oxford & IBH publishing company private limited, New Delhi
Singh IP, Barman D, Sharma CK (1997) Genetic variability and association of characters in papaya. Indian J Hortic 54(2):135–138
Singh K, Kumar A (2010) Genetic variability and correlation studies in papaya under Bihar conditions. In: Acta Horticulture International Symposium on Papaya 851. (851):145–50. https://doi.org/10.17660/ActaHortic.2010.851.20
Stiles JI, Lemme C, Sondur S, Morshidi MB, Manshardt R (1993) Using randomly amplified polymorphic DNA for evaluating genetic relationships among papaya cultivars. Theor Appl Genet 85(6–7):697–701. https://doi.org/10.1007/BF00225007
Sudha R, Singh DR, Sankaran M, Singh S, Damodaran V, Simachalam P (2013) Genetic diversity analysis of papaya (Carica papaya L.) genotypes in Andaman Islands using morphological and molecular markers. Afr J Agric Res 8(41):5187–5192
Toledo Á, Burlingame B (2006) Biodiversity and nutrition: a common path toward global food security and sustainable development. J Food Compos Anal 19(6–7):477–483. https://doi.org/10.1016/j.jfca.2006.05.001
Van Droogenbroeck B, Breyne P, Goetghebeur P, Romeijn-Peeters E, Kyndt T, Gheysen G (2002) AFLP analysis of genetic relationships among papaya and its wild relatives (Caricaceae) from Ecuador. Theor Appl Genet 105(2–3):289–297. https://doi.org/10.1007/s00122-002-0983-4
Varshney RK, Chabane K, Hendre PS, Aggarwal RK, Graner A (2007) Comparative assessment of EST-SSR, EST-SNP and AFLP markers for evaluation of genetic diversity and conservation of genetic resources using wild, cultivated and elite barleys. Plant Sci 173(6):638–649. https://doi.org/10.1016/j.plantsci.2007.08.010
Vitória AP, Filho GADS, Bressan-Smith RE, Pinto FO, Paiva LB, Guimarães PS et al (2004) DNA fingerprint of Carica papaya L, genotypes by RAPD markers. J New Seeds 6(1):51–65. https://doi.org/10.1300/J153v06n01_04
Weckwerth W, Ghatak A, Bellaire A, Chaturvedi P, Varshney RK (2020) Panomics meets germplasm. Plant Biotechnol J 18(7):1507–1525. https://doi.org/10.1111/pbi.13372
Wright S (1965) The interpretation of population structure by F-statistics with special regard to systems of mating. Evolution 1:395–420
Acknowledgements
The authors thank Professor and Head, Department of fruit science, TNAU, Coimbatore for providing seeds and field to conduct the experiment and Professor and Head, Department of Vegetable science for the laboratory infrastructure and equipment for the experiment.
Funding
No external funds were supported.
Author information
Authors and Affiliations
Contributions
ML executed the work, analysed the data and wrote the draft of the manuscript. CK provided resources, critically reviewed and finalized the draft. KS conceptualized the work and supervised the research activity.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
The research does not involve human or animal subjects.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Leela, M., Kavitha, C. & Soorianathasundaram, K. Genetic diversity and population structure analysis of papaya (Carica papaya L.) germplasm using simple sequence repeat (SSR) markers. Genet Resour Crop Evol (2024). https://doi.org/10.1007/s10722-024-01883-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10722-024-01883-x