Abstract
Agricultural production system is extremely vulnerable to climate change, and this change will heavily affect the grain yields, thereby threating the food security worldwide. People from developing countries are at greatest risk of experiencing food insecurity, and today, millions of people are going to bed hungry. Wheat is serving as a staple food for millions of people around the world. Development of high-yielding wheat varieties during the Green Revolution is considered an important event in agricultural history. However, these plant breeding activities also resulted in genetic erosion in wheat. Moreover, it is also believed that after domestication process, selection process also resulted in the loss of genetic diversity of wheat. Therefore, commercial wheat cultivars are prone to various biotic and abiotic stresses. To combat with climate changes and to serve enough quantity of food with quality, there is a need to harness wheat landraces. Landraces are considered as repository of gene pool that enhance the biodiversity and maintain and stabilize the ecosystem in a sustainable way to make it functional. Wheat landraces are traditional crop populations developed by the farmers through natural and human selection under their years of cultivations and have adaptation to local environment and management practices. Wheat landraces have more genetic diversity compared to their cultivated ones, and breeding community has utilized their potential in development of climate-resilient wheat cultivars. Here, we are exploring the role of landraces in wheat breeding and hoping that provided information will catch the attention of breeding community to collect, conserve, and perform breeding activities using wheat landraces.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Crop wild relatives
- Germplasm characterization
- Genetic diversity
- Adaptive traits breeding
- Stress breeding
11.1 Introduction of Landraces
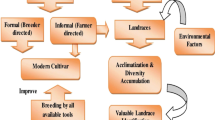
Agriculture is one of the oldest livelihood sources of mankind. Humans remained actively involved in the selection of favorable traits which resulted in significant changes in the phenotype and genotype of wild plants. In addition to man’s selection activities, environmental factors also played a significant contribution in the selection of various favorable traits suitable for man-made land and gardens. Combination of these activities resulted in the development of distinctive populations called “landraces” (Zeven 1998). Landraces are dynamic populations of cultivated plants having a historical background, genetically diverse and distinct identity, and good adaptation to local environment and that are associated with traditional farming systems (Villa et al. 2005). Dwivedi et al. (2016) stated that landraces are heterogeneous populations of domesticated species having great adaptation to local environment and can serve as a source of genetic variations that can be very helpful to combat the current and new challenges for farming in changing environments. Landraces are found phenotypically diverse and less productive compared to their cultivated types (Mir et al. 2020). However, regarding their quality attributes, landraces have been found highly nutritious compared to their cultivated ones (Azeez et al. 2018). Landraces played a major role in plant breeding by providing novel genes for various agronomic, quality, mineral, biotic, and abiotic traits (Azeez et al. 2018; Lopes et al. 2015). An impressive increase in yields per hectare was the result of the “Green Revolution” due the inclusion of high-yielding varieties (HYVs) having better response to inputs (Mir et al. 2020). After the inclusion of these high-yielding varieties, it was supposed that landraces will inevitably disappear with time (Frankel and Bennett 1970; Zeven 1998). However, these breeding activities led to genetic erosion and emergence of various modern cultivars that are prone to various biotic and abiotic stresses. It is estimated that approximately 75% loss of genetic diversity is observed in the last 100 years (Hammer et al. 1996). Globally, loss of genetic diversity is very alarming because it can be used to combat food scarcity problems in the long term. Therefore, it is very important to pay attention to collect, preserve, and grow these landraces as they guarantee the existence of variations that can be used for breeding of crops for the production of more quantity of food with high quality. Besides the inclusion of HYVs, landraces maintained their position by playing a key role in agricultural production, specifically in those environments where commercial cultivars failed their competitive advantage (Casañas et al. 2017).
11.2 Origin of Wheat Landraces
Wheat is one of the domesticated food crops cultivated in mild temperature and consumed as a staple crop by millions of people (Lodhi et al. 2020). Domestication of wheat is considered a key reason behind increased human population, thereby participating in the emergence of the human civilization (Jaradat 2011). Domestication of wild emmer (Triticum dicoccoides), which is the progenitor of all polyploid cultivated wheats, is considered an important event in the emergence of agriculture in Southwest Asia. Domestication of wild emmer occurs in the Fertile Crescent, and it acted as a prerequisite for the evolution of tetraploid durum and hexaploid bread wheat (Jaradat 2011). However, the domestication and the subsequent breeding activities drastically reduced wheat genetic diversity (Dvorak et al. 1998).
Ancient farmers planted diverse assemblages of wheat genotypes (i.e., landraces) aiming to decrease the risk of crop failure and to improve food security because they had limited capacity to control the spatially heterogeneous and temporally unpredictable environment (Jaradat 2006; Peng et al., 2011a, b). This exercise resulted in the development of wheat landrace meta-populations and the evolution of farmers’ seed systems through which they accessed and exchanged diverse genetic material. A meta-population structure can be defined as a group of subpopulations that is interconnected through gene flow and seed exchange and favors the evolution of diversity (Jaradat 2011). It is believed that natural interference, human skills, and years of continuous cultivation resulted in great diversity in wheat genotypes (Lodhi et al. 2020). Zeven (2000) stated that previously many farmers used wheat crops to develop new cultivars. Archaeological evidence are present revealing the cultivation of wheat in Iberian Peninsula, since the fifth millennium BC, and the development of wild wheats, traditional wheat varieties, and other crops happened in the Fertile Crescent (Diamond 2002).
Wheat landraces were developed from their older ones having the ability to grow in such conditions which are not feasible for the growth of the regular wheat (Witcombe et al. 1996). Zeven (1999) stated that wheat landraces are crop varieties developed by farmers through human and natural selection and reflect adaptation to local management practices and environmental conditions. Combination of both human and natural selection resulted in changes in the architecture of genotypes having better attributes like drought, salt, cold, or heat tolerance, quality traits, time to heading and maturity, and seed filling duration (Masood et al. 2005). Due to genetically distinct plant populations, wheat landraces are conserved, and some specific names were given by the traditional farmers in order to meet their environmental, cultural, social, and economic needs. Therefore, landraces are also known as farmer’s varieties or folk varieties (Belay et al. 1995).
11.3 How Landraces Contributed in Wheat Breeding
Landraces played a significant role in wheat breeding by gaining focus from breeding community. Wheat landraces served as genetic resource for the development of climate-resilient cultivars with high yield (Abu-Zaitoun et al. 2018). An increasing interest has been observed for the usage of landraces as source of nutritional traits and flavor repertoire and landrace cultivation for niche markets (Roselló et al. 2018). Wheat landraces contain higher genetic diversity compared to most modern wheat landraces, and this diversity includes their adaptation to environmental conditions according to the place of origin. Some countries used this characteristic in the development of first improved cultivars through the selection of local landraces. For example, “Aragon 03” was the leading variety in Spain during the period 1960–1976. It was developed from indigenous landrace population “Catalan de Monte” (Gadea 1958) and showed high ability to drought resistance (Royo and Briceño-Félix 2011a; b). Similarly, “Turkey” (syn. “Turkey Red”), a hard red winter wheat having better adaption for cold regions, showed marvelous impact on wheat cultivation in the United States at the turn of the last century due to decreased winterkill, among other traits (Olmstead and Rhode 2002). A Japanese landrace “Akakomugi,” containing Rht8c and Ppd-D1, was used by Italian breeder Nazareno Strampelli to improve Italian wheat gene pool (Salvi et al. 2013). The sensational varieties “Ardito and Mentana” developed from the crosses of Strampelli, including Akakomugi, became the basis of most of the new varieties developed in Mediterranean countries, South American countries, and several distant countries such as Russia and China. In Argentina, “Ardito” was used as parent to develop the variety Klein-33, which became the backbone of the former USSR breeding program, generating the variety Bezostaja-1 (Borojevic and Borojevic 2005). Contribution of landraces in wheat breeding for various traits is discussed comprehensively.
11.3.1 Role of Landraces in Adaptive Traits
Adaptive traits suited to target the environment have acted a decisive role during domestication and the spread out of domesticated wheat. Fitting flowering time to the current conditions in the target environments is presumably one of the main important factors during dispersal (Peng et al. 2011a; b; Royo et al. 2020). The first domesticated cereals/old landraces had most probably response to day length and cold temperatures like their wild relatives/progenitors. Motzo and Giunta (2007) hypothesized that old cultivars/landraces had the greatest day-length sensitivity and vernalization in comparison to intermediate and modern ones. However, novel adaptive traits for each target environment were naturally or artificially selected during the domestication and spreading process from the Fertile Crescent to new agricultural areas (Kilian et al. 2009). Especially other yield-related traits such as plant height, waxiness, number of spikes, and weight of spikes and grains were also co-selected by ancient farmers, and many botanical variants have been developed in this process (Peng et al. 2011a; b). Wheat landraces arising from the migration from the Fertile Crescent to the other regions of the world had been grown extensively until the Green Revolution in the early 1970s (Harlan 1975). As a result of the Green Revolution, more productive semidwarf wheat cultivar shaving better response to inputs replaced the landraces/local populations which are generally identified as tall, tended to lodge, sensitive to the foliar diseases, and low yielded (Reynolds and Borlaug 2006a; b; Lopes et al. 2015). Nevertheless, their cultivation has continued in marginal environments and they currently support subsistence farming in many regions of the world (Newton et al. 2010).
The wide range adaptability of wheat is mainly based on three genetic groups such as vernalization (Vrn) genes, photoperiod (Ppd) genes, and genes controlling “narrow-sense earliness” or “earliness per se” (Eps). Vernalization, which is inducting flowering by exposure to cold, basically determines plant growth habit types as winter (strong vernalization requirement) and spring (no vernalization requirement). Vernalization in wheat has very allelic complex and previous studies have presented that Vrn allele combinations or frequencies with an adaptive value in target growing areas are varied geographically (Stelmakh 1990; Damania et al. 1996; Iwaki et al. 2001; Zhang et al. 2008; Sun et al. 2009). Kato and Yokoyama (1992) observed the main adaptive traits in 158 bread wheat landraces collected from various climatic regions including Asian and European countries, and they claimed that nearly half of the variation for observed traits was accounted by geographical differences of their origin centers. Kato et al. (1997) also studied geographical variation of wild emmer (Triticum dicoccoides) accessions for vernalization response and earliness in comparison to other tetraploid relatives such as cultivated emmer (T. dicoccum), durum wheat (T. durum), and T. turgidum. They concluded that spring growth habit in T. dicoccoides could have evolved from a winter type especially in temperate conditions.
Many studies presented that the vernalization requirement in wheat is considered to be genetically controlled by at least three loci, Vrn-A1 (Vrn-1), Vrn-B1 (Vrn-2), and Vrn-D1 (Vrn-3), located in chromosomes 5A, 5B, and 5D, respectively (Pugsley 1971, 1972; Law et al. 1976; Galiba et al. 1995; Dubcovsky et al. 1998; Yan et al. 2003). While Vrn-A1 has the major impact on transition from vegetative to generative phase, recessive mutants of Vrn-B1 trigger flowering. While a dominant allele of any Vrn genes causes spring growth habit, wheats classified as winter type must have recessive alleles at all Vrn loci (Turner et al. 2013). On the other hand, photoperiodic response in wheat is primarily controlled by three major genes, Ppd-D1 (Ppd1), Ppd-B1 (Ppd2), and Ppd-A1 (Ppd3), located in 2DS, 2BS, and 2AS chromosomes, respectively. It is known that Ppd-D1 plays an important role in regulation of photoperiodic response. In addition, “earliness per se” or “narrow-sense earliness” is the difference in flowering times of genotypes whose vernalization and day-length requirements have been completed (Kato et al. 2001). Earliness per se genes can also affect flowering time independently, but these genes have not been studied in detail because of major effects of vernalization and photoperiod genes on flowering time. Moreover, this trait is highly heritable and can be effectively used in breeding programs (Kato and Wada 1999). Many QTLs have been identified for earliness per se in all three genomes with previous studies (Bullrich et al. 2002; Hanocq et al. 2004; Kamran et al. 2013).
Previous studies with a marker-assisted selection approach have clarified that landraces/accessions have a huge genetic diversity and very allelic complex for vernalization and photoperiod genes. Jiang et al. (2010) found that the frequencies of the dominant Vrn genes in 153 Chinese wheat landraces were 60.78% (Vrn-D1), 5.88% (Vrn-A1a), 5.23% (Vrn-B1), and 0 (Vrn-B3), respectively. Andeden et al. (2011) determined that Turkish wheat germplasm has mostly the dominant Vrn-B1 allele followed by Vrn-D1 and Vrn-A1. Derakhshan et al. (2013) reported that the frequencies of dominant Vrn-D1 and Vrn-B1 alleles in 395 Iranian wheat landraces were 67.35% and 38.48%, respectively. Manickavelu et al. (2014) characterized 400 wheat landraces genetically collected from different agroecological zones of Afghanistan for adaptive and other yield-related traits, and they reported that 53% of all landraces were winter types, 43% had one or more dominant Vrn alleles, and 4% were either unknown or had Vrn-A1c – a rare spring allele. Guo et al. (2015) also studied distribution of the Vrn-D1b allele in Chinese wheat accessions and determined that the frequencies of Vrn-D1a, Vrn-D1b, and Vrn-D1 alleles were 27.3, 20.6, and 52.1%, respectively, of 689 accessions. They also claimed that Vrn-D1b allele originated from Chinese landraces as a result of pedigree analysis. Goncharov (1998) claimed that there is a high rate of the Vrn-D1 allele in countries near the equator in addition to Pakistan, Afghanistan, and China.
Other important genetic factors like dwarfing genes (Rht) are critical against environmental stresses to guarantee both adaptability and grain yield in addition to vernalization, photoperiod, and earliness. It is known that over 30 height-reducing genes have been identified so far (McIntosh et al. 2013). The major dwarfing genes “Rht-B1” and “Rht-D1” known as the Reduced height (Rht) loci were introduced during the “Green Revolution” that achieved to improve harvest index by reducing plant height. These genes are known as gibberellic acid (GA)-insensitive dwarfing genes and located on chromosomes 4BS and 4DS, respectively. Another important height-reducing gene is Rht8 classified as GA-sensitive. Rht8 is located in chromosome 2D close to Ppd-D1 and previous studies clarified that Rht8 and Ppd-D1a alleles are often derived together (Worland et al. 1998), but Ppd-D1a has pleiotropic effects independently on plant height, grain yield, and yield-related traits (Börner et al. 2002; Chebotar et al. 2013; Zhang et al. 2019a, b). Zhang et al. (2006) determined that Rht-B1b, Rht-D1b, and Rht8 are common in autumn-sown Chinese wheat germplasm, while the frequencies of alleles vary from between regions. Kolev et al. (2011) also reported the most frequent alleles as Ppd-D1b, vrn-A1, vrn-B1, vrn-D1, and Rht-B1a in Bulgarian germplasm including old cultivars and landraces. Rasheed et al. (2016) studied the allelic variation of economically important traits such as Vrn, Ppd and Rht, in 107 wheat landraces collected from different geographic zones of Pakistan. They determined that less than half of the landraces has Ppd-D1a, Rht-B1b, Rht-D1b, and spring-type alleles of Vrn-A1 and Vrn-D1. The studies explained above highlight how these genes from landraces have geographically evolved in the target areas.
11.3.1.1 Success Stories of Wheat Landraces for Adaptive and Yield-Related Traits
There are many successful reports in the development of new wheat varieties with the use of landraces containing different dwarfing genes. A Japanese landrace “Akakomugi,” containing Rht8c and Ppd-D1, was used by Italian breeder Nazareno Strampelli to improve Italian wheat gene pool (Salvi et al. 2013). The crosses between Italian genotypes and Akakomugi resulted into the introgression of new alleles such as Ppd-D1 and Rht8c. The sensational varieties “Ardito and Mentana” developed from the crosses of Strampelli, including Akakomugi, became the basis of most of the new varieties developed in Mediterranean countries, South American countries, and several distant countries such as Russia and China. In Argentina, “Ardito” was used as parent to develop the variety Klein-33, which became the backbone of the former USSR breeding program, generating the variety Bezostaja-1 (Borojevic and Borojevic 2005). Another variety, Frontana, derived from a cross with Mentana, was part of the pedigree of the varieties Penjamo 62, Yaqui 48, Lerma 50, Escobar, and Supremo. Similarly, many genotypes derived from Mentana were developed in breeding programs of Canada and Australia (Salvi et al. 2013; Tadesse et al. 2016).
A similar success story from the Nobel laureate Norman Borlaug in the mid-20th century was recorded with the Norin 10/Brevor cross containing Rht-B1 and Rht-D1. The lineage of Norin-10, developed by a Japanese breeder G. Inazuka, is tracked back to a Japanese short-straw landrace “Shiro Daruma” containing Rht-B1 and Rht-D1 crossed with the American high-yielding varieties Fultz and later Turkey Red (Reitz and Salmon 1968). Norin 10-Brevor 14 cross was sent to N. Borlaug at the International Maize and Wheat Improvement Center (CIMMYT) in Mexico, and this cross and new crosses with Norin/Brevor 14 variants were tested for adaptation in tropical and subtropical climates in the center (Hedden 2003). Wheat varieties developed from semidwarf wheats developed by N. Borlaug and his colleagues in CIMMYT are grown in millions of hectares in many regions of the world.
The story of the Turkey Red brought to America is also very interesting. This bread wheat landrace, which was firstly grown in the USA around Kansas in the 1870s, was introduced to this region by German Mennonites who migrated from Crimea to the USA (Quisenberry and Reitz 1974; Smale 1996). The landrace has thin stem, high plant height, tended to lodging, narrow and dark green leaves, resistance to harsh climate conditions, white grain, high biomass, resistance to rust diseases, and tolerance to other foliar diseases (Quisenberry and Reitz 1974; Lopes et al. 2015). In addition, the landrace “Crimean,” introduced at the same time as Turkey Red, was directly included into the Nebraska gene pool. The effects of these two landraces on wheat improvement were indirectly reported with previous studies (Ali et al. 2011; Mengistu et al. 2012). Previous reports reported the investigation of major quantitative trait locus (QTL) related to grain yield on chromosome 3A originated from the cultivar “Wichita” which was obtained from these landraces.
Another important landrace “Chinese Spring” (CS) has also affected the wheat improvement and genetics in depth. This variety is known to be a Sichuan landrace, and Yen et al. (1988) claimed that CS is similar to a Sichuan white landrace “Cheng-du-guang-tou” (CDGT) in terms of morphology, physiology, and cytogenetics-based comparison. The similarity of these two landraces was also presented with RFLP profiling by Ward et al. (1998). CDGT has still been used widely in Sichuan breeding programs because of its high tillering potential, high number of spikelets, and high level of floret fertility (Liu et al. 2018). In addition, the landrace has been widely used to develop wheat-rye translocation lines because of its ready crossability, and therefore, many cultivars and pre-/breeding lines have been developed using CS as parent both in China and many different regions of the world. However, the main important impact of CS is on genetics and molecular breeding of wheat in which CS (IWGSC RefSeq v2.0) was sequenced at genome and single chromosome level and released the genomic data for public access (http://www.wheatgenome.org/News2/IWGSC-RefSeq-v2.0-now-available-at-URGI).
In addition to important wheat landraces mentioned above, several landraces with important adaptive and yield-related traits used by plant breeders in the early twentieth century have intensely been used in pedigrees of modern wheats such as Zeeuwse Witte in the Netherlands, Blount’s Lambrigg and Purple Straw in Australia, Marquis and Red Fife in Canada, Kunduru in Turkey, Saragolla in Italy, and Turkey Red in USA, which actually originated from Turkey (Gökgöl 1935; Quisenberry and Reitz 1974; Ozberk et al. 2016; Alsaleh et al. 2016) and became a cornerstone of the early European and indirectly world breeding programs (Smale 1996; Braun et al. 2001). In addition to these examples, wild progenitors/relatives and transition forms of wheat have formed the evolution and distribution of modern wheat landraces and (indirectly) cultivars. Especially, many unique alleles that provide resistance to different diseases and pests, including rust diseases, powdery mildew, Septoria tritici blotch, Septoria nodorum blotch, tan spot, cyst nematode, root knot nematode, Hessian fly, greenbug, Russian wheat aphid, wheat curl mite, and soilborne cereal mosaic virus, have been introgressed to modern wheat cultivars (Kishii 2019).
Introgression of new alleles from the locally adapted landraces to modern wheat cultivars should be one of the main breeding targets. Unfortunately, most of landraces have not still been identified both genetically and agronomically. However, the efficient use of landraces in breeding programs requires understanding their genetic diversity and population structure. Baloch et al. (2017) evaluated the genetic diversity of 92 durum wheat landraces from the Central Fertile Crescent including Turkey and Syria with 39,568 DArT-seq and 20,661 SNP markers. As a result of the study, Turkish and Syrian landraces complexly clustered into three groups, and the results illustrated that farmer-mediated selection and lack of the commercial varieties might have concluded in the exchange of genetic materials between two neighboring regions. Soriano et al. (2016) classified 172 durum wheat landraces, using molecular markers, into four genetic populations in relation to their geographic origin: eastern Mediterranean (EM), eastern Balkans and Turkey, western Balkans and Egypt, and western Mediterranean (WM). They determined that the genetic diversity among landraces increased during migration to West Mediterranean basin due to lower genetic diversity in the eastern Mediterranean population. Soriano et al. (2018) also support the theory with an association mapping study that 23 marker alleles in relation to important agronomic traits with different frequencies from east and west regions of Mediterranean basin were identified. With a similar approach, Liu et al. (2017a; b; c) reported a genome-wide association study with 52,303 DArT-seq markers that 723 wheat landraces collected from ten different agroecological zones of China were investigated for 23 agronomic traits in six environments. As a result of the study, all landraces were classified into five clusters based on phenotypic data, and 25 candidate genes associated with significant markers were characterized.
Unveiling the genetic basis of yield-related traits in wheat landraces is vital to ensure global food security because of their higher genetic diversity, large number of alleles, and potency of unique variants of alleles compared to modern wheat varieties. The advent of new technologies about sequencing, mapping, and other related technologies has been facilitating high-quality sequences of wheat and its relatives. The sequences will likely stimulate many new studies on evolution, genetics, and genomics of wheat, and accelerate characterization of novel genes controlling important adaptive and yield-related traits from landraces and wild relatives of wheat.
11.3.2 Role of Landraces in Abiotic Stress
Resistance to abiotic and biotic stresses, productivity, seed quality, seed mineral content, and many other traits will be future breeding aims to meet the world’s rapidly increasing food demand. Availability of higher natural genetic diversity to increase selection efficiency is one of the most critical and significant objectives of breeding programs. The abiotic stress factors (salinity, heat, drought, etc.) adversely affect crop production and yield (Jaleel et al. 2009; Thakur et al. 2010; Mantri et al. 2012). Traditional plant breeding is a long-term process that has been used effectively for many years, and molecular tools can be employed to overcome complications and to ensure the improvement of speed breeding strategies (Nadeem et al. 2018; Baloch et al. 2016). In this part, we discussed the role of landraces in different abiotic stress conditions such as salinity, heat, and drought to provide a significant resource for wheat breeders.
11.3.2.1 Wheat Landraces’ Role in Salinity Tolerance
Salinity is a major feature that reduces crop production and affects nearly 1 billion hectares of land worldwide (Fageria et al. 2012). Therefore, developing crops providing a satisfactory amount of product in salty soils or different climatic conditions is important to meet the growing food demand. Screening of wheat germplasm for salt tolerance has been conducted by various researchers (Kumar et al. 2017; Arabbeigi et al., 2018). For example, Shahzad et al. (2012) evaluated wheat landrace genotypes using morphological and molecular markers for salinity tolerance at the vegetative stage. The authors proposed that accessions 10793 (Pakistan), 10790 (Pakistan), 10821 (Pakistan), and 11526 (Pakistan) are found salt-tolerant at 200 mM NaCl stress. At 250 mM NaCl stress, accession 11299 (Pakistan) was the most salt-tolerant followed by accessions 11335 (Pakistan), 11370 (Italy), and 11214 (Pakistan). Additionally, accessions 10790 (Pakistan), 10828 (Pakistan), 10823 (Pakistan), and 4098805 (4098805) performed better at both 200 and 250 mM NaCl stresses. In another study, Chaparzadeh et al. (2014) determined the effects of NaCl (control, 75, and 150 mM) on the plant leaves of 18 bread wheat (Triticum aestivum L.) landraces from the west area of the Urmia Saline Lake. While accessions 12194 (from Piranshahr), 11199 (from Urmia), and 11488 (from Salmas) were found as the most tolerant with combined salt tolerance indexes for all biochemical and physiological parameters, accessions 11479 (from Mahabad) and 11492 (from Urmia) were determined as the least tolerant. It was suggested that these parameters could be used together as powerful biomarkers to screen for salt-tolerant landraces using the cluster analysis method. Al-maskri et al. (2014) investigated specific stem and leaf structural traits for water conservation. Based on the results of the study, cultivars/landraces were rated according to their degree of drought and salt tolerance as S-24 (from Pakistan) > J-305 (from Oman) > Sarraya (from Northern Asia, Africa, Middle East, Asia Minor) > Senain (from Oman) > Cooley (from Chile and Mongolia) > MH-97 (from Pakistan) > Missani (from the Mediterranean, Middle East Asia, and North Africa) > Hamira (from Oman) > Shwairaa (from Oman). Two of them (S-24 and J-305) are rated as highly tolerant, five moderately tolerant (Sarraya, Senain, Cooley, MH-97, and Missani), and two sensitive (Hamira and Shwairaa). The recent advances in genomic information and technology have opened new horizons and foundations for genetic breeding of salt tolerance. Various QTL mapping studies for salt tolerance in wheat were conducted by Quarrie et al. (2005), Ma et al. (2007), Genc et al. (2010), Hussain et al. (2017), Shamaya et al. (2017), Ren et al. (2018), Devi et al. (2019), and Ilyas et al. (2020). On the other hand, Yu et al. (2020) analyzed in a GWAS using 307 wheat accessions including local landraces and exotic cultivars. Researchers found that some Chinese landraces such as Baihuamai, Youzimai, Beijing 10, Jimai 1, and Zaosui 30 displayed superior salt tolerance. According to kinship analysis, Chinese landraces revealed a source of rare favorable genetic variation. Moreover, many of these landraces have already adapted to the different environments in China (Liu et al. 2017c; Zhou et al. 2018). In addition to these examples, wild relatives of wheat are also potential sources of important genetic materials such as salinity tolerance for wheat breeding. The use of wild relatives of Triticum species is one of the main breeding targets and may offer an opportunity to improve salinity tolerance by presenting availability to more variable germplasm (Shavrukov et al. 2009). For this content, researchers investigated the salinity tolerance of various accessions of Aegilops tauschii, and determined that the accessions studied are found similar to bread wheat. On the other hand, it was presented that accessions of Aegilops tauschii had a much lower Na+ ratio but higher K+/Na+ ratios in their leaves than did durum wheat (Gorham et al. 1987, 1990). Another important wild relative of wheat is jointed goatgrass, Aegilops cylindrica Host. (2n = 4x = 28; CCDD) species, which was formed through amphidiploidization of a hybrid or hybrids between Ae. tauschii Coss. (2n = 2x = 14; DD) and Ae. markgrafii (Greuter) Hammer (2n = 2x = 14; CC). Farooq et al. (1989) screened Ae. cylindrica accessions obtained from inland Pakistan and oversea, and determined that some of salinity-tolerant accessions survived at 300 mM NaCl and 400 mM NaCl in treatments using Hoagland solution. Another researcher reviewed the use of wild relatives of wheat for salinity tolerance (Colmer et al. 2006). Arabbeigi et al. (2014) evaluated the physiological response of the highly salinity-tolerant Ae. cylindrica genotypes and the SSR and EST-SSR markers linked to the salinity tolerance. As a result of the study, ten most salinity-tolerant genotypes of Ae. cylindrical were identified. In addition, Xgwm312, Xwmc170, Xgwm291, and Xgwm410 microsatellite markers produced a distinguished banding pattern in the ten most salinity-tolerant genotypes in the study. These markers can play important role in wheat breeding programs. Very recently, Ahmadi et al. (2020) investigated the domesticated and ancestral wheat genotypes, including Ae. triuncialis, Ae. neglecta, Ae. umbellulata, Ae. caudata, Ae. speltoides, Ae. tauschii, T. boeoticum, T. durum, T. urartu, and T. aestivum, under control and salinity stress to evaluate the mechanisms involved in salinity tolerance. It was found that two neglected (Ae. triuncialis) and ancestral (Ae. tauschii) wheat genotypes responded better to salinity tolerance than other genotypes. The studies explained above revealed that variation among the wild relatives and landraces of wheat is available for salinity tolerance, and they can be used to develop modern wheat cultivars in breeding studies.
11.3.2.2 Drought and Heat Stress Tolerance in Wheat
Drought and heat stress are important climatic factors that occur in almost all climatic areas of wheat-growing areas and cause a significant crop loss of up to 40% and 60% by drought and heat stresses in fields, respectively (Zampieri et al. 2017; Thirumalaikumar et al. 2018). These factors affect crops at the physiological, morphological, and biochemical levels (Guo et al. 2020); reduce photosynthesis (McKay et al. 2003), cell turgor (Taiz and Zeiger 2006), and chlorophyll fluorescence with a critical reduction of the Fv/Fm ratio (Mohammed and Tarpley, 2009; Izanloo et al. 2008), and impair cell division and elongation (Bal et al. 2010) in sensitive wheat lines compared with tolerant lines. Wheat yield is particularly sensitive to drought and heat stress factors that reduce spikelet productivity, individual grain weight, grain number, and grain filling time during the breeding season (Mahrookashani et al. 2017). The lack of water is not invincible (Ballesta et al. 2019). The adverse effects of drought and heat factors can be overcome by using drought- and heat-resistant cultivars (Van Oosten et al. 2016). The global scenario consists of having a genetic balance of major/minor genes suitable key for these stress factors and developing stress-resistant varieties (Mujeeb-Kazi et al. 2009). Success in plant development commonly depends on the size of genetic variability and the extent to which the beneficial traits are inherited (Kahrizi et al. 2010). Information from the germplasm evaluation will be of great importance for drought- and heat-tolerant genotype selection (Okechukwu et al. 2016). Breeding wheat varieties that tolerate these stressors is currently a major challenge for wheat breeders (Mwadzingeni et al. 2016). Exotic wheat landraces have been shown to be an excellent source of various genes and to function better under stressful conditions (Reynolds et al. 2007). Various studies were conducted to evaluate genetic resources in terms of drought and heat resistance (Hede et al. 1999; Sareen et al. 2014; Pinto et al. 2017; Al Khateeb et al. 2017; Ullah et al. 2018; Korkut et al. 2019). Hede et al. (1999) used a group of 2255 accessions from a Mexican landrace collection in which three landrace accessions (CWI 60155, CWI 59788, and CWI 60391) were determined as having superior and stable leaf chlorophyll content in both environments in 1997. In a study conducted by Sareen et al. (2014), six wheat genotypes (IC 28661, IC 57586, IC 78856, IC 28938B, IC 36761A, and IC 78869A) were identified as tolerant to drought and heat stresses. Al-maskri et al. (2014) rated cultivars/landraces according to their degree of drought and salt tolerance as S-24 (from Pakistan) > J-305 (from Oman) > Sarraya (from Northern Asia, Africa, Middle East, Asia Minor) > Senain (from Oman) > Cooley (from Chile, Mongolia) > MH-97 (from Pakistan) > Missani (from Mediterranean, Middle East Asia, North Africa) > Hamira (from Oman) > Shwairaa (from Oman). Aktaş (2016) determined the most tolerant genotypes (SEN-DER genotypes G7, G10, landrace group genotype G11 (Sorık)) to be used to improve drought-tolerant varieties. Al Khateeb et al. (2017) used four wheat landraces collected from Jordan and indicated that Karak landrace may be selected as the most tolerant wheat capable of adapting to drought-prone environments. Chaichi et al. (2019) screened 123 Iranian wheat (Triticum aestivum L.) landraces (spring and winter genotype) for drought tolerance using morphological and physiological features. They determined L-82 and Marvdasht genotypes as drought-tolerant and sensitive genotypes, respectively. Korkut et al. (2019) determined that some genotypes (Nota, Dropia, CIMMYT-HTN 2014/15-6, CIMMYT-HTN 2014/15-2, CIMMYT HTN 2014/15-10) could be evaluated as genitor(s)/progenitor(s) in the wheat breeding programs for heat tolerance.
Landraces, wild relatives, and traditional varieties are potential reservoirs of novel alleles for improving abiotic stress tolerance (Karan and Subudhi 2012). In this context, a deeper understanding of the genetic mechanisms of drought and heat resistance is important to maintain and further develop the efficiency of wheat breeding programs (Arriagada et al. 2017). The initial genetic investigations of wheat under both drought and heat stress in controlled conditions were conducted in durum wheat and bread wheat by Aprile et al. (2013) and Qaseem et al. (2018), respectively. Merchuk-Ovnat et al. (2016) revealed that introgression of QTLs on chromosomes 1B and 2B of T. turgidum into T. aestivum can improve drought tolerance in domesticated wheat. B genome has been identified carrying loci controlling water utilization efficiency, associated traits, and grain yield under water stress conditions (Mohammadi et al. 2012; Poersch-Bortolon et al. 2016). In another study conducted by Touzy et al. (2019), a panel of 210 elite European wheat varieties in 35 field trials was evaluated, and GWAS (genome-wide association study) was done with six characters in four different environment types to confirm 590 QTLs, some of which were specific to the different water stress patterns. Schmidt et al. (2020) used 315 spring bread wheat accessions to evaluate in pots with semi-controlled environmental conditions that combined drought and heat stress in 2016 and 2017. Australian and Mexican varieties were rated as having great productivity potential under both stresses, which have been selected for their yield performance and made up about 70% of the spring wheat panels. Nearly one-fifth of the tolerant wheat came from varieties of various origins such as the Middle East, the USA, Central Africa, India, and Canada. In the study, QTLs were determined on all chromosomes, most of which were on chromosomes 3B, 5A, 5B, and 6B. Drought and heat stress factors, which together can lead to significant yield losses, have restricted wheat yields in various wheat-growing areas worldwide, and their combined impact could result in critical yield losses (Toreti et al. 2019). Information about QTLs can help breeders to improve new cultivars tolerant to drought and heat stress in marginal environments in future global margins.
11.3.3 Role of Wheat Landraces in Quality Traits
11.3.3.1 Landraces for Biofortification
“Biofortification ” or “biological fortification” is the process of improving the nutritional status of staple crops such as minerals, vitamins, and proteins through traditional breeding, modern biotechnological methods, and agronomic approaches (Garg et al. 2018; Yeken et al. 2018; Saini et al. 2020). It is a long-term and sustainable approach, and a cost-effective way to overcome hidden hunger, which is a progressively severe universal challenge for humanity around the world (De Valença et al. 2017). In low-income countries, micronutrient deficiencies have largely increased in the last decades. Zn and Fe deficiencies in particular are a serious public health problem that negatively affects people’s lifespan, health, and productivity (WHO 2009; Khan et al. 2008). People need cereals for their dietary requirements; hence, biofortification of cereals is important worldwide (Saini et al. 2020). Wheat is one of the world’s most important crops for global food grain production, which was adversely affected by several biotic and abiotic stresses (Ozer et al. 2020). Annual wheat production is expected to increase in the coming years depending on increases of population (Iizumi et al. 2017). Biofortification can be divided into two categories as agronomic biofortification and genetic biofortification (Saini et al. 2020). The first step of biofortification in food crops for plant breeders is to understand the current genetic diversity in germplasm collections (Baloch et al. 2014). Wheat has a large number of wild relatives that can lead to its genetic development (Dempewolf et al. 2017; Ahmadi et al. 2018; Saini et al. 2020). The most frequently required mineral elements in the human diet can be obtained from genetic variations, which improve the levels of nutrients in crops (White and Broadley 2005; Bouis and Saltzman 2017). Agronomical biofortification techniques include fertilizing crops with different fertilizers containing elements such as zinc, iron, and selenium, while genetic biofortification includes traditional and molecular breeding approaches. These techniques have the potential to increase the levels of these minerals in grains (Saini et al. 2020). Monasterio and Graham (2000) claimed that iron and zinc concentrations especially in some bread wheat genotypes were negatively correlated with Rht genes. They also reported that the high-yielding wheat cultivars developed after Green Revolution contained less iron and zinc compared to old cultivars/landraces. Heidari et al. (2016) reported that landraces had higher Fe and Zn concentrations compared with commercial cultivars. Ram and Govindan (2020) clarified that genetic diversity in wheat landraces and wild relatives provides novel alleles for genetic enhancement of Zn and Fe. Lyons et al. (2005) examined 665 wheats (ancestral and wild relatives, landrace accessions, and registered cultivars) in Australia and Mexico for Se concentration in grain. They found that Se concentrations of grains changed between 5 and 720 microgr/kg. Khokhar et al. (2020) studied 245 bread wheat genotypes derived from crosses with landraces and the modern wheat cultivar Paragon to detect grain Zn concentration, and they reached promising results for high level of grain Zn where Zn concentration in wholegrain was positively correlated with Fe concentration and grain protein content. They claimed that landraces have a huge potential to increase the concentration of Zn in whole grain and flour of modern high-yielding bread wheat cultivars.
11.3.3.2 Landraces for Some Important Quality Traits
It is generally known that old landraces or cultivars have a huge diversity for some quality traits such as grain protein content, grain texture (hardness), and gluten strength and quality (glutenin and gliadin subunits) than modern wheat cultivars (Aguiriano et al. 2006; Moragues et al. 2006; Ruiz et al. 2012). The grain protein content (GPC) is a crucial trait in determining the quality of wheat (Veraverbeke and Delcour 2002), and modern wheat grains include inherently low protein levels. Hence, breeding for an increase in the protein levels of grain wheat is required to alleviate hunger and nutrient deficiencies. However, the grain protein content was negatively related to grain yield (Blanco et al. 2006; Iqbal et al. 2007; Klindworth et al. 2009). Avivi (1978) claimed that wild emmer wheat (T. turgidum ssp. dicoccoides) can be a potential gene source to improve grain protein content in modern wheat. Joppa and Cantrell (1990) also studied this hypothesis that they crossed wild emmer wheat and durum wheat, and obtained substitution lines with high GPC. Joppa et al. (1997) reported that a QTL explained 66% of total variation in these substitution lines for GPC. The QTL was named as Gpc-B1 (Distelfeld et al. 2004), and Uauy et al. (2006a) also positionally cloned the locus and renamed as NAM-B1. Hagenblad et al. (2012) studied 367 bread wheat germplasm with worldwide origin and determined that five accessions had wild-type NAM-B1 allele where it confers high levels of protein and microelements. They also indicated that several accessions with wild-type NAM-B1 were traced back to Fennoscandian origin. In addition to landraces, cultivated transitional forms of wheat such as einkorn (T. monococcum ssp. monococcum), emmer (T. turgidum ssp. dicoccum), and spelt (T. aestivum ssp. spelta) and wild relatives have the possibility to contain the wild-type NAM-B1 allele. Uauy et al. (Uauy et al. 2006a; b) reported that wild emmer accessions and most of cultivated emmer accessions studied had wild-type NAM-B1 allele. Asplund et al. (2010) also determined that only two spelts had a wild-type NAM-B1 allele among 62 wheat germplasm displayed at the International Exhibition in London in 1862. It’s likely that unique variants for grain protein content can be uncovered due to higher genetic diversity of landraces.
As another important trait, endosperm texture is mainly controlled by the Hardness (Ha) locus located in 5DS, and it’s simply inherited despite the fact that softness is the dominant trait. The lipid binding proteins, puroindoline genes (Pina-D1 and Pinb-D1), which are tightly linked to Ha locus, have been used to determine the differences between hard- and soft-textured wheats, and landraces that originated from different geographic regions had different Puroindoline allele combinations. As an example of this situation, Ayala et al. (2013) studied 102 lines selected from 15 Mexican landraces and determined that while 16 lines had hard texture, 86 lines were soft-textured. Ten out of 16 lines had presence of both Pina-D1 and Pinb-D1 alleles. They concluded that the Mexican old landraces are potential sources for important quality traits to develop new wheat varieties with hard grain texture. Li et al. (2019) also studied 107 Chinese wheat cultivars and landraces in terms of diversity of Puroindoline genes and their association with kernel hardness. The most frequent combinations were Pina-D1a/Pinb-D1a and PinaD1a/Pinb-D1b with 39.3% and 34.6% ratios, respectively. They indicated that Chinese landraces had more allelic than do cultivars and are a valuable source of genetic variability in Puroindoline genes. Gluten strength and quality are other important quality traits of wheat. Many studies were conducted to determine the genetic variability of old durum wheat cultivars or landraces for glutenin and gliadin profiles, which affected viscoelastic properties of dough, especially in Mediterranean basin (Melnikova et al. 2010; Xynias et al. 2011; Ribeiro et al. 2011; Ruiz et al. 2012; Janni et al. 2018). Nazco et al. (2012) studied the variability of some quality traits such as protein content, SDS sedimentation, and yellow color index and gluten strength in 154 durum wheat landraces from 20 Mediterranean countries with 18 modern wheat cultivars. They determined that the largest variability for quality traits was observed in landraces from eastern Mediterranean basin followed by landraces from western Mediterranean basin, and identified landraces could be used to improve quality traits especially for gluten strength and grain weight in durum wheat breeding programs. While Glu-A1c was the most frequent allele in almost all genetic materials studied for Glu-A1 locus, but Glu-A1a was found at low frequency in Mediterranean basin (Mir Ali et al. 1999; Moragues et al. 2006; Naghavi et al. 2009). In addition to Glu-A1a, Glu-A1b, and Glu-A1VI, encoding the subunits 2* and 2*** were determined at very low frequency. However, Henkrar et al. (2017) reported that in Moroccan genotypes, Glu-A1a and Glu-A1b were the predominant alleles. On the other hand, at the Glu-B1 locus, there were more genetic variation between genotypes with Glu-B1b, Glu-B1d, and Glu-B1e alleles encoding the subunits 7+8, 6+8, and 20, respectively. Moreover, the variation varied geographically that while Glu-B1d allele was predominant in Algerian, Syrian, and Spanish germplasm (Mir Ali et al. 1999; Moragues et al. 2006; Hamdi et al. 2010), the allele was not present in Iranian landraces that they had more Glu-B1a, Glu-B1e, and Glu-B1i alleles (Naghavi et al. 2009). Similar genetic variation was determined for low molecular weight glutenin subunits (LMW-GS). Li et al. (2009) studied 615 Chinese wheat germplasm including 390 landraces and 225 varieties, for HMW-GS, LMW-GS, Zeleny sedimentation, volume, dough development time, stability time, and strength, and reported that genetic materials with good gluten strength and quality were identified in landraces that did not contain wheat-rye translocation. Wheat-rye (the 1BL/1RS) translocation has been used widely in breeding programs because of its disease resistance genes especially for foliar diseases and increased grain yield in some environments, but it negatively affects bread-making quality of wheat at the same time (Zhao et al. 2012; Oak and Tamhankar 2017).
On the other hand, the new technologies such as sequencing, mapping, and other related technologies have been recently used to reveal genetic diversity and novel variants/alleles among landraces related to quality traits of wheat. For instance, Giraldo et al. (2016) performed an association mapping study with 183 Spanish wheat landraces using 749 DArT markers for 18 agromorphological and grain quality traits including protein content, gluten strength, vitreousness, yellow color index, thousand kernel weight, and test weight. They identified 85 stable MTAs (marker-trait associations) with more than 10% explained phenotypic variation, and claimed that novel MTAs were identified and can provide new information to understand genetic control of complex traits. Roselló et al. (2018) also performed an association mapping study with 165 durum wheat landraces from 21 Mediterranean countries using 1149 DArT markers. Landraces had generally higher GPC than modern ones in this study but lower gluten strength. In addition to this, while eastern landraces showed the highest yellow color index, Balkan landraces had the lowest test weight. They also identified 15 meta-QTL (MQTL) for grain quality traits of wheat.
Various studies about landraces conducted in different countries have been briefly summarized and discussed above. We hope that improving the grain quality via agronomic/genetic biofortification and quality breeding studies and producing wheat genotypes with better quality will be beneficial to prevent hidden hunger and to live healthy. In this regard, collaboration among various specialists from public and private research institutes and universities can accelerate the improvement of wheat varieties with high bread- and pasta-making quality. This section will be helpful for wheat breeders, providing knowledge of the advancement made so far in wheat biofortification and quality.
11.4 Role of Landraces in Biotic Stress
There are many studies conducted to discover resistance properties of wheat landraces for different biotic stresses. Since in the wild the host and the pathogen have co-lived in mutual habitats for long periods of time, they co-evolved together. Thus, the sources of resistance can be found most often at these centers of origin, among the wild relatives and landraces of wheat (McIntosh et al. 1995). Pinpointing the resistance factors and genes in the genome and development of molecular markers to test their presence are of great importance.
11.4.1 Role of Wheat Landraces in Disease Resistance
11.4.1.1 Role of Wheat Landraces in Rust Diseases
11.4.1.1.1 Yellow Rust or Stripe Rust
Rust diseases of wheat are among most important and economically devastating diseases of wheat. Rust diseases of wheat consist of yellow (stripe) rust (YR) caused by Puccinia striiformis f. sp. tritici, leaf rust (LR) caused by Puccinia triticina, and stem rust (SR) caused by Puccinia graminis f. sp. tritici (Reynolds and Borlaug 2006a; b). Genes that confer resistance to the rust diseases are generally designated as Yr, Lr, and Sr for the effectiveness against yellow rust, leaf rust, and stem rust, respectively. Resistances against rust diseases are the most studied resistance properties in wheat landraces. To date, some genes against rust diseases have been identified from landraces and wild relatives of wheat. Among them, Sr2 gene, which provides resistance against stem rust, has been incorporated from an emmer wheat landrace (McIntosh et al. 1995). Race-nonspecific resistance genes Yr52, Yr56, Yr57, and Yr62, which provide adult plant resistance (APR) against yellow rust, have been also incorporated from landraces (Mondal et al. 2016).
Yellow rust or stripe rust is one of the most prevalent and devastating wheat foliar diseases worldwide (Kumar et al. 2016). It is observed mostly on cool and moist regions and causes lower kernel quality and massive yield losses (Chen et al. 2013). Recently, there are many studies on YR done by genome wide association studies (GWAS) using bread and durum wheat landraces (Tehseen et al. 2020; Long et al. 2019; Liu et al. 2017a; b; c; Manickavelu et al. 2016). Wu et al. (2016) used simple sequence repeats (SSR), sequence-related amplified polymorphism (SRAP), and resistance gene analog polymorphism (RGAP) markers, Ma et al. (2015) used SSR and SRAP markers, while Wang et al. (2010) used SSR markers to find the source of resistance in a known resistant wheat landrace. Kandel et al. (2017) used microsatelite markers to pinpoint the resistance in the genome of known resistant wheat landrace. Wu et al. (2015) used molecular markers to screen wheat landraces to find a suppressor gene of the known resistance gene Yr18. Li et al. (2015) used DArT-seq genotyping-by-sequencing (GBS) on 8416 Mexican Creole landrace wheats and found seven accessions from them with less than 20% disease severity after YR inoculation. Gessese et al. (2019) screened resistant landrace Aus27430 with 90K wheat SNP chip array by selective genotyping to locate a new resistance gene “Yr81.” Yuan et al. (2018), Wang et al. (2019), and Liu et al. (2020) used also wheat SNP chip to locate resistance characteristics of wheat landraces. Bux et al. (2012) evaluated Pakistani wheat landraces phenotypically against the disease; on the other hand Akar et al. (2009) used durum wheat landraces from Turkey to evaluate their resistance against the yellow rust disease. Rola et al. (2019) have found two Lebanese wheat landraces that are resistant to different yellow rust pathogen races, including the devastating Warrior pathotype. Wamalwa et al. (2020) found that Kenyan Kenya Tai landrace shows resistance against many YR races. Mohammadi et al. (2015) screened 380 durum wheat landraces and found 46 accessions to be resistant against YR.
11.4.1.1.2 Leaf Rust
Leaf rust is one of the main wheat diseases seen worldwide, which can affect kernel weight and wheat biomass, causing major yield losses (Herrera-Foessel et al. 2006). There are many studies done on leaf rust resistance. Qureshi et al. (2018) identified a novel resistance gene “Lr79,” from genotyping analysis of resistant durum wheat by using DArT-seq and 90K chip array, and also developed a Kompetitive Allele Specific Polymerase (KASP) marker to locate the gene. Kolmer et al. (2018) used DArT-seq technology to genotype Uruguayan wheat landrace Americano 44. Qureshi et al. (2017) used DArT-seq markers to locate disease resistance in the genome of two wheat landraces from Portugal. Zhang et al. (2019a), b) screened 46 Chinese wheat landraces for resistance against LR and used molecular markers to find out the presence of known resistance genes in those accessions. Akcura et al. (2017) used Turkish wheat landraces, while Riaz et al. (2017) used 136 wheat landraces from Vavilov Institute of Plant Genetic Resources in Russia to test against YR phenotypically. Andenow et al. (1997) used ten Ethiopian tetraploid wheat (Triticum turgidum L.) landraces and found some degree of resistance toward the YR disease.
11.4.1.1.3 Stem Rust
Stem rust is one of the major diseases of wheat which hinders with the nutrient flow to developing ears and result in shriveling of the grain and the breakage of the stem that can cause total yield loss (Roelfs et al. 1992; Leonard and Szabo 2005). Studies on SR have been conducted by Babiker et al. (2015) and Zurn et al. (2014), which used quantitative trait loci (QTL) and linkage map, respectively, to locate the resistance region in the known resistant landrace against stem rust pathogen. Haile et al. (2013) used molecular markers for genotyping the Ethiopian durum wheat landraces. Newcomb et al. (2013) and Toor et al. (2013) have screened the landrace collection phenotypically against the SR disease and genotyped using molecular markers. Denbel and Badebo (2012) screened Ethiopian durum wheat landraces against SR race Ug99. On the other hand, Endresen et al. (2011) used ecogeographic data of landrace accessions to predict the resistance against SR according to climatic factors of their location of origin, while Bonman et al. (2007) studied the geographic origin or the resistant accessions. There are also studies conducted to find multiple rust resistance in wheat landraces. Studies which include resistance against all three rust diseases were conducted by DArT and molecular markers (Rahmatov et al. 2019; Bansal et al. 2013) by GWAS and resistance gene prediction (Kankwatsa et al. 2017; Pasam et al. 2017; Jordan et al. 2015; Daetwyler et al. 2014). Kertho et al. (2015) studied YR and SR resistance traits with GWAS technique, Sthapit et al. (2014) used simple sequence repeat (SSR) markers to study YR and SR resistance, and Aoun et al. (2019) used QTL in durum wheat to locate the resistance region against LR and SR in the known resistant durum wheat landrace.
11.4.1.2 Role of Wheat Landraces in Powdery Mildew (PM)
Powdery mildew (PM) is a foliar fungal disease caused by Blumeria graminis f. sp. tritici, an obligate biotrophic fungus that causes yield and quality loss in wheat grains (Newton et al. 2011). Chinese wheat landraces known for their PM resistance were screened by microsatelite markers (Xue et al. 2009, Huang et al. 2000), SSR markers (Qie et al. 2019; Sun et al. 2018; Fu et al. 2017; Wang et al. 2015; Xu et al. 2015; Fu et al. 2013; Xue et al. 2012), and RNA-seq SNP markers (Li et al. 2020, Xu et al. 2018) to locate genes in the plants’ genome, responsible for the resistance trait. Li et al. (2018a; b) used SSR marker to pinpoint resistance in an Afghan wheat landrace. Tan et al. (2019) and Tan et al. (2018) used single Iranian and Afghan PM-resistant wheat landrace to define new resistance genes “Pm63” and “Pm59,” respectively, using SSR markers. Identification of germplasm strategy (FIGS) was used on wheat landraces in a study conducted by Wang et al. (2015), Bhullar et al. (2010), and Bhullar et al. (2009) to discover new alleles of powdery mildew resistance gene Pm3. Huang (1997) also used APR against powdery mildew found in the landrace accession k-15560, and monosomic and hybridological analyses were used to locate the gene (Peusha et al. 2002). Amplified fragment length polymorphism (AFLP) markers and microsatellite markers were used to locate Pm24 resistance gene in a Chinese spring wheat landrace. Li et al. (2012) used SSR markers to test the diversity of the single wheat landrace and its relation to the PM resistance. In their study, Li et al. (2016a; b) used 1,297 landraces from 57 countries to screen for the PM resistance, and molecular markers were used to check the presence of known resistance genes. Hysing et al. (2007) screened 155 Nordic wheat landraces phenotypically and with molecular markers for resistance to PM.
11.4.1.3 Role of Wheat Landraces in Fusarium Head Blight (FHB)
Fusarium head blight (FHB) is caused by the fungal pathogen Fusarium graminearum Schwabe and has destructive effects on cereals and especially on wheat production all over the world. Moreover, the diseased plants become contaminated with mycotoxins which are poisonous to mammals (Cetin and Bullerman 2005; Goswami and Kistler 2004). Cai et al. (2019) used meta-analysis of previous QTL studies (MQTL) of five wheat landraces to construct a consensus map, and they also developed 22 KASP markers to ease the MAS in breeding programs. Xiao et al. (2011) located a chromosomal region responsible for FHB resistance by fast-neutron induced chromosome fragment deletion, causing the resistant wheat landrace to lose its resistance and become susceptible. Li et al. (2016a, b) used SSR and sequence-tagged site (STS) markers in 195 wheat accessions to find the presence of known resistance genes, whereas Wei et al. (2005) used microsatellite markers to compare the difference between 20 resistant wheat landraces and 4 susceptible wheat lines. Xiao et al. (2013) used RNA sequencing to determine expression of a resistant wheat landrace during FHB infection. There are also studies where wheat landraces known for their resistance against Fusarium head blight have been screened with SSR markers to pinpoint the resistance source in the genome (Cai et al. 2016; Zhang et al. 2012; Li et al. 2011). Talas et al. (2011) screened 68 Syrian durum wheat landraces and Yu et al. (2008) screened 94 wheat accessions to find new sources of resistance to FHB.
11.4.1.4 Role of Wheat Landraces in Septoria Tritici Blotch (STB)
Septoria tritici blotch (STB) is major foliar wheat disease caused by the fungal pathogen Zymoseptoria tritici previously known as Mycosphaerella graminicola . It is a major threat to wheat production globally, and it is the most damaging pathogen of wheat in Europe causing loss in chlorophyll, premature death of leaves, and reduction of grain production (O’Driscoll et al. 2014; Ziv and Eyal 1977). Many European and Chinese landraces have been found to contain Stb6 gene which provides resistance against STB (Chartrain et al. 2005a; b). Kidane et al. (2019) used 318 Ethiopian wheat landraces for GWAS analysis and found four putative loci for STB resistance. Ouaja et al. (2020) screened 304 Tunisian wheat landraces, and Ghaneie et al. (2012) screened 45 tetraploid Iranian wheat landraces to test against STB disease phenotypically and found some promising accessions.
11.4.1.5 Role of Wheat Landraces in Tan Spot
Tan spot is caused by Pyrenophora tritici-repentis and is an important foliar wheat disease causing severe loss in the grain yield. The disease causes large-scale chlorosis and tan necrosis on leaves and grain shriveling (Maraite et al., 1997, de Wolf et al. 1998). In their study, Gurung et al. (2011) assessed the resistance of 567 wheat landraces against P. tritici-repentis races 1 and 5 using DArT markers and developed association mapping.
11.4.1.6 Role of Wheat Landraces in Eyespot
Eyespot is caused by soilborne necrotrophic funguses Oculimacula acuformis and Oculimacula yallundae. The disease is seen in temperate areas and affects the stem base of the cereals including wheat, causing premature grain ripening and heavy crop losses (Crous et al. 2003, Fitt et al. 1990, Scott and Hollins 1974). Burt et al. (2014) screened all 1056 hexaploid wheat landraces of Watkins collection against both funguses and found two promising accessions with high level of resistance. They also genotyped the accessions that showed resistance to one or both funguses by SSR, STS, and QTL-linked markers.
11.4.1.7 Role of Wheat Landraces in Stagonospora Nodorum Blotch (SNB)
Stagonospora nodorum blotch (SNB) is caused by Phaeosphaeria nodorum and constitutes a serious disease of wheat worldwide (Eyal 1987). SNB disease infects both leaves and glumes, subsequently causing decreased grain quality and yield losses (King et al. 1983). Adhikari et al. (2011a, b) evaluated 567 spring wheat landraces of different origin for resistance to SNB and used DArT markers to genotype and develop association map of the resistance traits.
11.4.1.8 Role of Wheat Landraces in Bacterial Leaf Streak (BLS)
Bacterial leaf streak (BLS) is caused by Xanthomonas translucens pv. undulosa, the most important wheat bacterial pathogen which can cause major outbreaks in the wheat fields under favorable conditions (Adhikari et al. 2011b, Bragard et al. 1997). Adhikari et al. (2012) screened 566 spring wheat landraces for resistance against BLS and used DArT markers to generate association mapping of the resistance regions. They found five genomic regions which are associated with resistance to the BLS disease.
11.4.1.9 Role of Wheat Landraces in Spot Blotch (SB)
Spot blotch (SB) is caused by Cochliobolus sativus which is a fungal disease of wheat and barley, observed globally which results in severe yield losses (Kumar et al. 2002). Adhikari et al. (2012) screened 566 spring wheat landraces also for resistance against SB and used DArT markers to create association mapping of the resistance regions. They found four genomic regions which are associated with resistance to the SB disease.
11.4.1.10 Role of Wheat Landraces in Common Bunt (CB)
Common bunt (CB) is caused by the fungal pathogen Tilletia tritici that causes significant yield losses in spring and winter wheat production worldwide (Goates and Peterson 1999). Bonman et al. (2006) investigated 10,759 wheat accessions for resistance against the common bunt disease. Accessions from Bakhtaran province in Iran showed the most resistance.
11.4.1.11 Role of Wheat Landraces in Dwarf Bunt (DB)
Dwarf bunt (DB) is caused by the fungus Tilletia controversa in winter wheat in regions where snow is persistent (Goates and Peterson 1999). Bonman et al. (2006) studied 8167 wheat accessions against dwarf bunt resistance. Accessions from Hakkari province in Turkey showed the highest resistance against DB.
11.4.1.12 Role of Wheat Landraces in Wheat Blast (WB)
Wheat blast (WB) is a relatively new emerging disease (mid-1980s) caused by Triticum pathotype of Pyricularia oryzae fungus. It has immense impacts on wheat production (Inoue et al. 2017). Wang et al. (2018a, b) evaluated 520 landraces of common wheat from different regions of the world for the resistance to Br48 isolate of the fungus and found a unique accession resistant to WB. The resistance was due to combination effect of two genes “Rmg8” and newly found “RmgGR119” gene.
11.4.2 Role of Wheat Landraces in Pest Resistance
11.4.2.1 Role of Wheat Landraces in Root Lesion Nematodes
Root lesion nematodes Pratylenchus thornei and Pratylenchus neglectus are the most common root lesion parasites that grow and develop in wheat roots, causing damage and substantial losses in wheat production (Nicol et al. 2002). Thompson and Seymour (2011) analyzed the modes of inheritance of resistance to P. thornei in seven wheat accessions that showed resistance against the nematode. Schmidt et al. (2005) studied two resistant Middle Eastern wheat landraces with AFLP and microsatellite markers for QTL analysis of resistance to P. thornei. Thompson et al. (2009) screened 207 bread wheat and 102 durum wheat accessions from West Asia and North Africa for resistance against P. thornei. Among them, 13 bread wheat and 10 durum wheat showed significant resistance. Thompson et al. (2016) screened 78 Iranian wheat accessions for resistance against P. thornei and P. neglectus. Among them, 32 showed some degree of resistance to both nematodes.
11.4.2.2 Role of Wheat Landraces in Russian Wheat Aphid (RWA)
Russian wheat aphid (RWA) (Diuraphis noxia) is an important wheat pest indigenous to southern Russia and Mediterranean countries which have spread to all continents causing substantial damage to wheat fields (DuToit and Walters 1984; Hewitt et al. 1984). Valdez et al. (2012) have evaluated a resistant Iranian wheat landrace using SSR markers to identify the location of resistance trait. It was found that the trait was due to dominant gene. Similarly, Li et al. (2018a) used an Iranian wheat landrace known for its resistance to RWA to locate the trait in the genome using SSR markers.
11.4.2.3 Role of Wheat Landraces in Wheat Stem Sawfly (WSS)
Wheat stem sawfly (WSS), Cephus cinctus Norton, is a major pest insect of wheat observed in North America, with devastating consequences in wheat production (Michael et al. 1992). Mohammadi et al. (2015) evaluated the collection of 380 durum wheat landraces against WSS and found that 33 accessions showed resistance to the pest. Varella et al. (2017) screened 1409 accessions of wheat landraces collected from different regions to WSS. They found 204 accessions that have resistance to the disease. The resistant accessions were screened with KASP markers for QTL analysis. Varella et al. (2019) used four resistant wheat accessions and generated six recombinant inbred lines (RIL) with them and genotyped with 90K iSelect assay to find novel QTL related to WSS resistance.
11.4.2.4 Role of Wheat Landraces in Cereal Cyst Nematodes (CCN)
Cereal cyst nematodes (CCN) (Heterodera spp.) are a group of 12 known species with H. avenae, H. filipjevi, and H. latipons being the most important ones. The pest is observed in many regions of the world and causes major yield losses in cereals (Nicol et al. 2003). Yavuzaslanoglu et al. (2016) studied the response of 31 Iranian wheat landraces against H. filipjevi and found one resistant and five moderately resistant accessions.
11.4.2.5 Role of Wheat Landraces in Cereal Aphids
Cereal aphids cause important yield losses in wheat. There are 14 species of aphids that were observed causing damage to wheat. Sitobion avenae, Rhopalosiphum maidis, R. padi, and Metopolophium dirhodum are the most common of these (Popov et al. 1988). Amin et al. (2019) observed 114 wheat landraces for their resistance level against the disease and population dynamics of R. padi. They found promising accessions which can be used for breeding of resistant cultivars.
11.5 Landraces and the Future of Wheat Diversity
The world is confronting food scarcity problem due to rapid increase in population and climate change. Previous report showed 6–13% reduction in wheat yield for each °C rise in temperature. Continuously changing climate, extreme weather events, new pathogen strains, and pests further jeopardize linear productivity growth into the future (Mondal et al. 2016). It is believed that the world’s population will cross the nine billion mark in 2050. By considering this factor, it is very important to increase wheat production by a rate of 1.6% (Lodhi et al. 2020). To feed the rapidly increasing world’s population under changing climatic conditions, more pressure is put on agriculture to produce enough quantity of food. Therefore, it is very important to increase the wheat production to serve enough quantity of food. By considering these factors, it is very important to develop wheat cultivars having higher production and better adaptation to biotic and abiotic stresses (Khan et al. 2013). These targets can be achieved by harnessing wheat genetic diversity. Previous studies explored the existence of higher genetic diversity in wheat landraces compared to its commercial cultivars (Lodhi et al. 2020; Jaradat 2011; Jaradat 2013).
Genetic diversity present in wheat landraces has been successfully utilized for breeding perspectives. Wheat landraces possess a sufficient amount of diversity, including useful genes to adapt to stressful environments such as salinity, heat, and drought (Karagöz and Zencirci 2005; Özkan et al. 2011). The evaluation of genetic diversity in wheat landraces is important for the selection of the suitable landraces as donors of traits in breeding studies (Gurcan et al. 2017; Abbasov et al. 2018). Landraces represent significantly broader genetic diversity than modern varieties (Azeez et al. 2018). For this reason, they can help to increase the genetic source of modern cultivars. However, for their utilization in breeding programs, it is very important that breeders should make crosses among elite lines having the highest likelihood of developing new varieties (Baenziger and DePauw 2009). There is scarcity of information about the successful release of cultivars using wheat landraces. Gerek 79 which is a Turkish variety is developed through crosses with landraces (Smale and McBride 1996). One of the best examples of landraces serving as a source of novel genes is the identification of Rht dwarfing gene that was available through the Japanese variety “Norin 10” originating from a Japanese landrace Shiro Daruma (Reitz and Salmon 1968; Dreisigacker et al. 2005). Dr. Norman E. Borlaug utilized these genes to develop the high-yielding semidwarf wheat varieties that resulted in Green Revolution. Similarly, various wheat landraces served as a foundation in the wheat germplasm pool impotent like: “Cheyenne,” a selection from landrace Crimea, founded the Nebraska wheat gene pool. Moreover, “Turkey Red” has been successfully used in winter wheat breeding in the US Great Plains (Lopes et al. 2015). Similarly, previous studies confirmed landrace diversity as a potential source for the breeding of grain yield and climate resilience, for example, the drought-tolerant variety “Aragon 03” was developed from a selection of a landrace population “Catalan de Monte” (Royo and Briceño-Félix 2011a; b). Vikram et al. (2016a; b) stated that a group of Creole wheat landraces (the landraces introduced to Mexico from Europe) has better adaptation to various abiotic stresses including drought because of the presence of rare but beneficial alleles. Further, wheat landraces reflected genetic diversity for various traits like 1000-kernel weight, biomass, and photosynthesis that can be used for cultivar development (Lopes et al. 2015). Various studies have been conducted using wheat landraces as germplasm through molecular markers and explored their potential as a source of novel variations (Sansaloni et al. 2020; Alipour et al. 2017; Lopes et al. 2015; Sofalian et al. 2008; Alsaleh et al. 2015; Jorgensen et al. 2017; Arystanbekkyzy et al. 2019; Dababat et al. 2020; Ozer et al. 2020). As is obvious from the above-provided information, there is a need to utilize wheat landrace diversity to develop climate-resilient cultivars having high yield. Similarly, some nonbreeding efforts that should be used to promote on-farm dynamic conservation and sustainable utilization of wheat landraces include the following:
-
1.
Awareness should be raised in the farming community about their potential in changing climate.
-
2.
Availability of wheat landrace seeds to the farmers.
-
3.
Development of niche market for landrace products.
-
4.
Involvement of wheat breeders, seed producers, farmers, and end-users, as stakeholders in wheat breeding activities to develop new cultivars (Newton et al. 2011).
References
Abbasov M, Akparov Z, Gross T, Babayeva S, Izzatullayeva V, Hajiyev E, Rustamov K, Gross P, Tekin M, Akar T, Chao S (2018) Genetic relationship of diploid wheat (Triticum spp.) species assessed by SSR markers. Genet Resour Crop Evol 65(5):1441–1453
Abu-Zaitoun SY, Chandrasekhar K, Assili S, Shtaya MJ, Jamous RM, Mallah OB, Nashef K, Sela H, Distelfeld A, Alhajaj N, Ali-Shtayeh MS (2018) Unlocking the genetic diversity within a Middle-East panel of durum wheat landraces for adaptation to semi-arid climate. Agronomy 8(10):233
Adhikari TB, Gurung S, Hansen JM, Jackson EW, Bonman JM (2012) Association mapping of quantitative trait loci in spring wheat landraces conferring resistance to bacterial leaf streak and spot blotch. Plant Genome 5(1):1–16
Adhikari TB, Jackson EW, Gurung S, Hansen JM, Bonman JM (2011a) Association mapping of quantitative resistance to Phaeosphaeria nodorum in spring wheat landraces from the USDA National Small Grains Collection. Phytopathology 101(11):1301–1310
Adhikari TB, Hansen JM, Gurung S, Bonman JM (2011b) Identification of new sources of resistance in winter wheat to multiple strains of Xanthomonas translucens pv. undulosa. Plant disease 95(5):582–588
Aguiriano E, Ruiz M, Fite’ R, Carrillo JM (2006) Analysis of genetic variability in a sample of the durum wheat (Triticum durum Desf.) Spanish collection based on gliadin markers. Genet Resour Crop Evol 53(8):1543–1552
Ahmad M, Shahzad A, Iqbal M, Asif M, Hirani AH (2013) Morphological and molecular genetic variation in wheat for salinity tolerance at germination and early seedling stage. Aust J Crop Sci 7(1):66
Ahmadi J, Pour-Aboughadareh A, Ourang SF, Mehrabi AA, Siddique KHM (2018) Wild relatives of wheat: Aegilops–Triticum accessions disclose differential antioxidative and physiological responses to water stress. Acta Physiol Plant 40:1–14
Ahmadi J, Pour-Aboughadareh A, Ourang SF, Khalili P, Poczai P (2020) Unraveling salinity stress responses in ancestral and neglected wheat species at early growth stage: A baseline for utilization in future wheat improvement programs. Physiol Mol Biol Plants 1–13.
Akar T, Mert Z, Yazar S, Sanal T, Avci M (2009) Sustainable use of winter Durum wheat landraces under Mediterranean conditions. Afr J Biotechnol 8(17)
Akcura M, Kadir A, Hocaoglu O (2017) Biplot analysis of leaf rust resistance in pure lines selected from eastern Anatolian bread wheat landraces of turkey. Turkish J Field Crops 22(2):227–234
Aktaş H (2016) Drought tolerance indices of selected landraces and bread wheat (Triticum aestivum L.) genotypes derived from synthetic wheats. Appl Ecol Environ Res 14(4):177–189
Al Khateeb W, Schroeder D, Musallam I (2017) Phenotypic and molecular variation in drought tolerance of Jordanian durum wheat (Triticum durum Desf.) landraces. Physiol Mol Biol Plants 23(2):311–319
Ali ML, Baenziger PS, Ajlouni ZA, Campbell BT, Gill KS, Eskridge KM, Mujeeb-Kazi A, Dweikat I (2011) Mapping QTLs for yield and agronomic traits on wheat chromosome 3A and a comparison of recombinant inbred chromosome line populations. Crop Sci 51:553–566
Alipour H, Bihamta MR, Mohammadi V, Peyghambari SA, Bai G, Zhang G (2017) Genotyping-by-sequencing (GBS) revealed molecular genetic diversity of Iranian wheat landraces and cultivars. Front Plant Sci 8:1293
Al-maskri A, Hameed M, Ashraf M, Khan MM, Fatima S, Nawaz T, Batool R (2014) Structural features of some wheat (Triticum spp.) landraces/cultivars under drought and salt stress. Arid Land Res Manag 28(3):355–370
Al-Naggar AMM, Abd El-Shafi MAE, El-Shal MH, Anany AH (2020) Evaluation of Egyptian wheat landraces (Triticum aestivum L.) for drought tolerance, agronomic, grain yield and quality traits. Plant Archives 20(Supplement 1):3487–3504
Alsaleh A, Baloch FS, Derya M, Azrak M, Kilian B, Özkan H, Nachit M (2015) Genetic linkage map of Anatolian durum wheat derived from a cross of Kunduru-1149× Cham1. Plant Mol Biol Rep 33:209–220
Alsaleh A, Baloch FS, Nachit M, Ozkan H (2016) Phenotypic and genotypic intra-diversity among Anatolian durum wheat “Kunduru” landraces. Biochem Syst Ecol 65:9–16
Amin M, Mahmood K, Nazir N, Kassi AK, Ahmed S (2019) Population dynamics of wheat aphid on different landraces of wheat under field conditions. Plant Protect 3(2):59–66
Andeden EE, Yediay FE, Baloch FS, Shaaf S, Kilian B, Nachit M, Ozkan H (2011) Distribution of vernalization and photoperiod genes (Vrn-A1, Vrn-B1, Vrn-D1, Vrn-B3, Ppd-D1) in Turkish bread wheat cultivars and landraces. Cereal Res Commun 39:352–364
Andenow Y, Hullukal M, Belay G (1997) Resistance and tolerance to leaf rust in Ethiopian tetraploid wheat landraces. Plant breeding 116(6):533–536
Aoun M, Kolmer JA, Rouse MN, Elias EM, Breiland M, Bulbula WD, Chao S, Acevedo M (2019) Mapping of novel leaf rust and stem rust resistance genes in the Portuguese durum wheat landrace PI 192051. G3: Genes, Genomes. Genetics 9(8):2535–2547
Aprile A, Havlickova L, Panna R, Mare C, Borrelli GM, Marone D, Perrotta C, Rampino P, De Bellis L, Curn V, Mastrangelo AM, Rizza F, Cattivelli L (2013) Different stress responsive strategies to drought and heat in two durum wheat cultivars with contrasting water use efficiency. BMC Genomics 14:821–838
Arabbeigi M, Arzani A, Majidi MM, Kiani R, Tabatabaei BES, Habibi F (2014) Salinity tolerance of Aegilops cylindrica genotypes collected from hyper-saline shores of Uremia Salt Lake using physiological traits and SSR markers. Acta Physiol Plant 36(8):2243–2251
Arabbeigi M, Arzani A, Majidi MM, Sayed-Tabatabaei BE, Saha P (2018) Expression pattern of salt tolerance-related genes in Aegilops cylindrica. Physiol Mol Biol Plants 24(1):61–73
Arriagada O, Mora F, Quitral Y, Del Pozo A (2017) Identification of QTL underlying agronomic, morphological and physiological traits in barley under rainfed conditions using SNP markers. Acta Sci Agron 39:321–329
Arystanbekkyzy M, Nadeem MA, Aktas H, Yeken MZ, Zencirci N, Nawaz MA, Ali F, Haider MS, Tunç K, Chung G, Baloch FS (2019) Phylogenetic and taxonomic relationship of Turkish wild and cultivated emmer (Triticum turgidum ssp. dicoccoides) revealed by iPBSretrotransposons markers. Int J Agric Biol 21(1):155–163
Asplund L, Hagenblad J, Leino MW (2010) Re-evaluating the history of the wheat domestication gene NAM-B1 using historical plant material. J Archaeol Sci 37:2303–2307
Avivi L (1978) High protein content in wild tetraploid Triticum dicoccoides Korn. In Ramanujam S (ed). In: Proceedings of the 5th international wheat genetics symposium, New Delhi, 23–28 Feb 1978. Indian Soc Genet Plant Breed, Indian Agric Res Inst, New Delhi, pp 372–380
Ayala M, Guzmán AJB, Peña RJ (2013) Characterization of genetic diversity of puroindoline genes in Mexican wheat landraces. Euphytica 190(1):53–63
Azeez MA, Adubi AO, Durodola FA (2018) Landraces and crop genetic improvement. In Rediscovery of landraces as a resource for the future. IntechOpen.
Babiker EM, Gordon TC, Chao S, Newcomb M, Rouse MN, Jin Y, Wanyera B, Acevedo M, Brown-Guedira G, Williamson S, Bonman JM (2015) Mapping resistance to the Ug99 race group of the stem rust pathogen in a spring wheat landrace. Theor Appl Genet 128(4):605–612
Baenziger PS, Depauw RM (2009) Wheat breeding: Procedures and strategies. In: Wheat science and trade. Wiley-Blackwell, Ames, pp 273–308
Bal W, Kozlowski H, Robbins R, Pettit LD (2010) Improving drought tolerance by exogenous application of glycinebetaine and salicylic acid in sunflower. J Agron Crop Sci 194:193–199
Blanco A, Simeone R, Gadaleta A (2006) Detection of QTLs for grain protein content in durum wheat. Theor Appl Genet 112:1195–1204
Ballesta P, Mora F, Del Pozo A (2019) Association mapping of drought tolerance indices in wheat: QTL-rich regions on chromosome 4A. Sci Agri 77(2)
Baloch FS, Alsaleh A, Andeden EE, Hatipoğlu R, Nachit M, Özkan H (2016) High levels of segregation distortion in the molecular linkage map of bread wheat representing the West Asia and North Africa region. Turk J Agric For 40(3):352–364
Baloch FS, Alsaleh A, Shahid MQ, Çiftçi V, Aasim M, Nadeem MA, Aktaş H, Özkan H, Hatipoğlu R (2017) A whole genome DArTseq and SNP analysis for genetic diversity assessment in durum wheat from central fertile crescent. PLoS One 12(1):e0167821
Baloch FS, Karaköy T, Demirbaş A, Toklu F, Özkan H, Hatipoğlu R (2014) Variation of some seed mineral contents in open pollinated faba bean (Vicia faba L.) landraces from Turkey. Turk J Agric For 38(5):591–602
Bansal UK, Arief VN, DeLacy IH, Bariana HS (2013) Exploring wheat landraces for rust resistance using a single marker scan. Euphytica 194(2):219–233
Belay G, Tesemma T, Bechere E, Mitiku D (1995) Natural and human selection for purple-grain tetraploid wheats in the Ethiopian highlands. Genet Resour Crop Evol 42:387–391
Bhullar NK, Street K, Mackay M, Yahiaoui N, Keller B (2009) Unlocking wheat genetic resources for the molecular identification of previously undescribed functional alleles at the Pm3 resistance locus. Proc Natl Acad Sci U S A 106(23):9519–9524
Bhullar NK, Zhang Z, Wicker T, Keller B (2010) Wheat gene bank accessions as a source of new alleles of the powdery mildew resistance gene Pm3: a large scale allele mining project. BMC Plant Biol 10(1):88
Bonman JM, Bockelman HE, Goates BJ, Obert DE, McGuire PE, Qualset CO, Hijmans RJ (2006) Geographic distribution of common and dwarf bunt resistance in landraces of Triticum aestivum subsp. aestivum. Crop Sci 46(4):1622–1629
Bonman JM, Bockelman HE, Jin Y, Hijmans RJ, Gironella AIN (2007) Geographic distribution of stem rust resistance in wheat landraces. Crop Sci 47(5):1955–1963
Börner A, Schumann E, Fürste A, Cöster H, Leithold B, Röder M, Weber W (2002) Mapping of quantitative trait loci determining agronomic important characters in hexaploid wheat (Triticum aestivum L.). Theor Appl Genet 105(6-7):921–936
Börner A, Worland AJ, Plaschke J, Schumann E, Law CN (1993) Pleiotropic effects of genes for reduced height (Rht) and day-length insensitivity (Ppd) on yield and its components for wheat grown in middle Europe. Plant Breed 111:204–216
Borojevic K, Borojevic K (2005) Historic role of the wheat variety Akakomugi in Southern and Central European wheat breeding programs. Breed Sci 55:253–256
Bouffier B (2014) Genetic and ecophysiological dissection of tolerance to drought and heat stress in bread wheat: from environmental characterization to QTL detection (Doctoral dissertation).
Bouis HE, Saltzman A (2017) Improving nutrition through biofortification: a review of evidence from HarvestPlus, 2003 through 2016. Glob Food Sec 12:49–58
Bragard C, Singer E, Alizadeh A, Vauterin L, Maraite H, Swings J (1997) Xanthomonas translucens from small grains: diversity and phytopathological relevance. Phytopathology 87(11):1111–1117
Braun HJ, Zencirci N, Altay F, Atli A, Avci M, Eser V, Kambertay M, Payne TS (2001) Turkish wheat pool. In: Bonjean AP, Agnus WJ (eds) The world wheat book: A history of wheat breeding. Lavosier, Paris, pp 851–879
Budak H, Shearman RC, Parmaksiz I, Gaussoin RE, Riordan TP, Dweikat I (2004) Molecular characterization of Buffalograss germplasm using sequence-related amplified polymorphism markers. Theor Appl Genet 108:328–334
Bullrich L, Appendino ML, Tranquilli G, Lewis S, Dubcovsky J (2002) Mapping of a thermo-sensitive earliness per se gene on Triticum monococcum chromosome 1Am. Theor Appl Genet 105:585–593
Burt C, Griffe LL, Ridolfini AP, Orford S, Griffiths S, Nicholson P (2014) Mining the Watkins collection of wheat landraces for novel sources of eyespot resistance. Plant Pathol 63(6):1241–1250
Bux H, Ashraf M, Chen X (2012) Expression of high-temperature adult-plant (HTAP) resistance against stripe rust (Puccinia striiformis f. sp. tritici) in Pakistan wheat landraces. Can J Plant Pathol 34(1):68–74
Cai J, Wang S, Li T, Zhang G, Bai G (2016) Multiple minor QTLs are responsible for Fusarium head blight resistance in Chinese wheat landrace Haiyanzhong. PLoS One 11(9):e0163292
Cai J, Wang S, Su Z, Li T, Zhang X, Bai G (2019) Meta-analysis of QTL for Fusarium head blight resistance in Chinese wheat landraces. Crop J 7(6):784–798
Casañas F, Simó J, Casals J, Prohens J (2017) Toward an evolved concept of landrace. Front Plant Sci 8:145
Chaichi M, Sanjarian F, Razavi K, Gonzalez-Hernandez JL (2019) Phenotypic diversity among Iranian bread wheat landraces, as a screening tool for drought tolerance. Acta Physiol Plant 41(6):1–15
Cetin Y, Bullerman LB (2005) Cytotoxicity of Fusarium mycotoxins to mammalian cell cultures as determined by the MTT bioassay. Food Chem Toxicol 43(5):755–764
Chaparzadeh N, Aftabi Y, Dolati M, Mehrnejad F, Pessarakli M (2014) Salinity tolerance ranking of various wheat landraces from the west of the Urmia saline lake in Iran by using physiological parameters. J Plant Nutr 37(7):1025–1039
Chartrain L, Berry ST, Brown JKM (2005a) Resistance of wheat line Kavkaz-K4500 L. 6. A. 4 to Septoria tritici blotch controlled by isolate-specific resistance genes. Phytopathology 95(6):664–671
Chartrain L, Brading PA, Brown JKM (2005b) Presence of the Stb6 gene for resistance to Septoria tritici blotch (Mycosphaerella graminicola) in cultivars used in wheat-breeding programmes worldwide. Plant Pathol 54(2):134–143
Chebotar GO, Chebotar SV, Motsnyy II, Sivolap YM (2013) Clarification of the Rht8–PpdD1 gene linkage on the 2D chromosome of winter bread wheat. Cytol Genet 47:70–74
Chen W, Wellings C, Chen X, Kang Z, Liu T (2014) Wheat stripe (yellow) rust caused by Puccinia striiformis f. sp. tritici. Mol Plant Pathol 15(5):433–446
Chen P, You C, Hu Y, Chen S, Zhou B, Cao A, Wang X (2013) Radiation-induced translocations with reduced Haynaldia villosa chromatin at the Pm21 locus for powdery mildew resistance in wheat. Mol Breed 31(2):477–484
Colmer TD, Flowers TJ, Munns R (2006) Use of wild relatives to improve salt tolerance in wheat. J Exp Bot 57:1059–1078
Crous PW, Groenewald JE, Gams W (2003) Eyespot of cereals revisited: ITS phylogeny reveals new species relationships. Eur J Plant Pathol 109(8):841–850
Dababat A, İmren M, Pridannikov M, Özer G, Zhapayev R, Mokrini F, Otemissova A, Yerimbetova A, Morgounov A (2020. Plant-parasitic nematodes on cereals in northern Kazakhstan. J Plant Dis Prot 1–9
Daetwyler HD, Bansal UK, Bariana HS, Hayden MJ, Hayes BJ (2014) Genomic prediction for rust resistance in diverse wheat landraces. Theor Appl Genet 127(8):1795–1803
Damania AB, Pecetti L, Qualset CO, Humeid BO (1996) Diversity and geographic distribution of adaptive traits in Triticum turgidum L. (durum group) wheat landraces from Turkey. Genet Resour Crop Evol 43:409–422
De Valença AW, Bake A, Brouwer ID, Giller KE (2017) Agronomic biofortification of crops to fight hidden hunger in sub-Saharan Africa. Glob Food Sec 12:8–14
De Wolf ED, Effertz RJ, Ali S, Francl LJ (1998) Vistas of tan spot research. Canadian journal of plant pathology= Revue Canadienne de phytopathologie
Dempewolf H, Baute G, Anderson J, Kilian B, Smith C, Guarino L (2017) Past and future use of wild relatives in crop breeding. Crop Sci 57(3):1070–1082
Denbel W, Badebo A (2012) Valuable sources of resistance in the Ethiopian durum wheat landraces to Ug33 and other stem rust races. Int. J. Agron. Plant Prod 3:191–195
Derakhshan B, Mohammadi SA, Moghaddam M, Jalal Kamali MR (2013) Molecular characterization of vernalization genes in Iranian wheat landraces. Crop Breed J 3:11–14
Devi R, Ram S, Rana V, Malik VK, Pande V, Singh GP (2019) QTL mapping for salt tolerance associated traits in wheat (Triticum aestivum L.). Euphytica 215(12):210
Diamond J (2002) Evolution, consequences and future of plant and animal domestication. Nature 418:700–707
Distelfeld A, Uauy C, Olmos S, Schlatter AR, Dubcovsky J, Fahima T (2004) Microcolinearity between a 2-cM region encompassing the grain protein content locus Gpc-6B1 on wheat chromosome 6B and a 350-kb region on rice chromosome 2. Funct Integr Genomics 4:59–66
Dreisigacker S, Zhang P, Warburton ML, Skovmand B, Hoisington D, Melchinger AE (2005) Genetic diversity among and within CIMMYT wheat landrace accessions investigated with SSRs and implications for plant genetic resources management. Crop Sci 45:653–661
Dubcovsky J, Lijavetzky D, Appendino L, Tranquilli G (1998) Comparative RFLP mapping of Triticum monococcum genes controlling vernalization requirement. Theor Appl Genet 97:968–975
DuToit F, Walters MC (1984) Damage assessment and economic threshold values for the chemical control of the Russian wheat aphid, Diuraphis noxia (Mordvilko) on winter wheat. Technical communication-South Africa, Department of Agriculture
Dvorak J, Luo MC, Yang ZL, Zhang HB (1998) The structure of the Aegilops tauschii genepool and the evolution of hexaploid wheat. Theor Appl Genet 97:657–670
Dvorak J, Noaman MM, Goyal S, Gorham J (1994) Enhancement of the salt tolerance of Triticum turgidum L by the Kna1 locus transferred from Triticum aestivum L. chromosome 4D by homoeologous recombination. Theor Appl Genet 87:872–877
Dwivedi SL, Ceccarelli S, Blair MW, Upadhyaya HD, Are AK, Ortiz R (2016) Landrace germplasm for improving yield and abiotic stress adaptation. Trends Plant Sci 21:31–42
ELshafei AA, Afiah SA, Amer MA, El-enany MAM (2019) Validation of molecular markers linked with salinity tolerance in wheat (Triticum aestivum L.) grown on saline soil. Biosci Res 16(2):963–978
Endresen DTF, Street K, Mackay M, Bari A, DePauw E (2011) Predictive association between biotic stress traits and eco-geographic data for wheat and barley landraces. Crop Sci 51(5):2036–2055
Epstein E, Bloom AJ (2005) Mineral nutrition of plants: principles and perspectives. Sinauer, Sunderland
Eyal Z (1987) The Septoria diseases of wheat: concepts and methods of disease management. CIMMYT
Fageria NK, Stone LF, dos Santos AB (2012) Breeding for salinity tolerance. In plant breeding for abiotic stress tolerance. Springer, Berlin, pp 103–122
Farooq S, Niazi M, Iqbal N, Shah TM (1989) Salt tolerance potential of wild resources of the tribe Triticeae II. Screening of species of the genus Aegilops. Plant and Soil 119:255–260
Fei X, Wen-Wen Z, Xia-Yu D, Yi-Lin Z, Wan-Quan J (2009) Microsatellite mapping of a powdery mildew resistance gene in wheat landrace Xiaobaidong. Acta Agronomica Sinica 35(10):1806–1811
Fitt BD, Goulds A, Hollins TW, Jones DR (1990) Strategies for control of eyespot (Pseudocercosporella herpotrichoides) in UK winter wheat and winter barley. Ann Appl Biol 117(2):473–486
Flowers TJ (2004) Improving crop salt tolerance. J Exp Bot 55:307–319
Frankel OH, Bennett E (1970) Genetic resources in plants-their exploration and conservation. In: Genetic resources in plants-their exploration and conservation. Distributed by Blackwell Scientific, Oxford
Fu B, Chen Y, Li N, Ma H, Kong Z, Zhang L, Jia H, Ma Z (2013) pmX: a recessive powdery mildew resistance gene at the Pm4 locus identified in wheat landrace Xiaohongpi. Theor Appl Genet 126(4):913–921
Fu B, Zhang Z, Zhang Q, Wu X, Wu J, Cai S (2017) Identification and mapping of a new powdery mildew resistance allele in the Chinese wheat landrace Hongyoumai. Molecular Breeding 37(11):133
Gadea M (1958) Trigos cultivados en España y nuevas variedades recomendadas. Ministerio de Agricultura, Madrid
Galiba G, Quarrie SA, Sutka J, Morgounov A, Snape JW (1995) RFLP mapping of the vernalization (Vrnl) and frost resistance (Frl) genes on chromosome 5A of wheat. Theor Appl Genet 90:1174–1179
Garg AK, Kim JK, Owens TG, Ranwala AP, Do Choi Y, Kochian LV, Wu RJ (2002) Trehalose accumulation in rice plants confers high tolerance levels to different abiotic stresses. Proc Natl Acad Sci U S A 99:15898–15903
Garg M, Sharma N, Sharma S, Kapoor P, Kumar A, Chunduri V, Arora P (2018) Biofortified crops generated by breeding, agronomy, and transgenic approaches are improving lives of millions of people around the world. Front Nutr 5:12
Genc Y, Oldach K, Verbyla AP, Lott G, Hassan M, Tester M, Wallwork H, McDonald GK (2010) Sodium exclusion QTL associated with improved seedling growth in bread wheat under salinity stress. Theor Appl Genet 121(5):877–894
Genc Y, Verbyla AP, Torun AA, Cakmak I, Willsmore K, Wallwork H, McDonald GK (2009) Quantitative trait loci analysis of zinc efficiency and grain zinc concentration in wheat using whole genome average interval mapping. Plant and Soil 314:49
Gessese M, Bariana H, Wong D, Hayden M, Bansal U (2019) Molecular mapping of stripe rust resistance gene Yr81 in a common wheat landrace Aus27430. Plant disease 103(6):1166–1171
Ghaneie A, Mehrabi R, Safaie N, Abrinbana M, Saidi A, Aghaee M (2012) Genetic variation for resistance to septoria tritici blotch in Iranian tetraploid wheat landraces. Eur J Plant Pathol 132(2):191–202
Giraldo P, Royo C, Gonzalez M, Carrillo JM, Ruiz M (2016) Genetic diversity and association mapping for agromorphological and grain quality traits of a structured collection of durum wheat landraces including subsp. durum, turgidum and diccocon. PLoS One 11(11):e0166577
Goates BJ, Peterson GL (1999) Relationship between soilborne and seedborne inoculum density and the incidence of dwarf bunt of wheat. Plant Dis 83(9):819–824
Gökgöl M (1935) Turkish wheats, vol I. Ministry of Agriculture, Yesilkoy Seed Breeding Institute Publications. No: 7, Devlet Press, Istanbul. (In Turkish), 436 pp.
Goncharov NP (1998) Genetic resources of wheat related species: The Vrn genes controlling growth habit (spring vs. winter). Euphytica 100(1):371–376
Gonzalez-Hernandez JL, Elias EM, Kianian SF (2004) Mapping genes for grain protein concentration and grain yield on chromosome 5B of Triticum turgidum (L.) var. dicoccoides. Euphytica 139:217–225
Gorham J, Bristol A, Young EM, Wyn Jones RG, Kashour G (1990) Salt tolerance in the Triticeae: K/Na discrimination in barley. J Exp Bot 41:1095–1101
Gorham J (1994) Salt tolerance in the Triticeae: K/Na discrimination in some perennial wheatgrasses and their amphiploids with wheat. J Exp Bot 45:441–447
Gorham J, Bridges J, Dubcovsky J, Dvoák J, Hollington PA, Luo MC, Khan JA (1997) Genetic analysis and physiology of a trait for enhanced K + /Na + discrimination in wheat. New Phytol 137:109–116
Gorham J, Hardy C, WynJones RG, Joppa LR, Law CN (1987) Chromosomal location of a K/Na discrimination character in the D genome of wheat. Theor Appl Genet 74:584–588
Goswami RS, Kistler HC (2004) Heading for disaster: Fusarium graminearum on cereal crops. Mol Plant Pathol 5(6):515–525
Graham RD, Welch RM, Bouis HE (2001) Addressing micronutrient malnutrition through enhancing the nutritional quality of staple foods: principles, perspectives and knowledge gaps. Adv Agron 70:77–142
Guo X, Wang Y, Meng L, Liu H, Yang L, Zhou Y, Zhang H (2015) Distribution of the Vrn-D1b allele associated with facultative growth habit in Chinese wheat accessions. Euphytica 206:1–10
Guo X, Xin Z, Yang T, Ma X, Zhang Y, Wang Z, Ren Y, Lin T (2020) Metabolomics Response for Drought Stress Tolerance in Chinese Wheat Genotypes (Triticum aestivum). Plan Theory 9(4):520
Gurcan K, Demirel F, Tekin M, Demirel S, Akar T (2017) Molecular and agro-morphological characterization of ancient wheat landraces of Turkey. BMC Plant Biol 17(1):171
Gurung S, Mamidi S, Bonman JM, Jackson EW, DelRio LE, Acevedo M, Mergoum M, Adhikari TB (2011) Identification of novel genomic regions associated with resistance to Pyrenophora tritici-repentis races 1 and 5 in spring wheat landraces using association analysis. Theor Appl Genet 123(6):1029
Haile JK, Hammer K, Badebo A, Singh RP, Röder MS (2013) Haplotype analysis of molecular markers linked to stem rust resistance genes in Ethiopian improved durum wheat varieties and tetraploid wheat landraces. Genetic resources and crop evolution 60(3):853–864
Hammer K, Knüpffer H, Xhuveli L, Perrino P (1996) Estimating genetic erosion in landraces—two case studies. Genet Resour Crop Evol 43:329–336
Hanocq E, Niarquin M, Heumez E, Rousset M, Le Gouis J (2004) Detection and mapping of QTL for earliness components in a bread wheat recombinant inbred lines population. Theor Appl Genet 110:106–115
Hao Y, Velu G, Peña RJ, Singh S, Singh RP (2014) Genetic loci associated with high grain zinc concentration and pleiotropic effect on kernel weight in wheat (Triticum aestivum L.). Mol Breed 34:1893–1902
Hagenblad J, Asplund L, Balfourier F, Ravel C, Leino MW (2012) Strong presence of the high grain protein content allele of NAM-B1 in Fennoscandian wheat. Theor Appl Genet 125:1677–1686
Hamdi O, Bellil I, Branlard G, Khelii D (2010) Genetic variation and geographical diversity for seed storage proteins of seventeen durum wheat populations collected in Algeria. Not Bot Horti Agrobo 38(2):22–32
Harlan JR (1975) Our vanishing genetic resources. Science 188(4188):618–621
Hedden P (2003) The genes of the Green Revolution. Trends Genet 19:5–9
Hede AR, Skovmand B, Reynolds MP, Crossa J, Vilhelmsen AL, Stølen O (1999) Evaluating genetic diversity for heat tolerance traits in Mexican wheat landraces. Genet Resour Crop Evol 46(1):37–45
Heidari B, Padash S, Dadkhodaie A (2016) Variations in micronutrients, bread quality and agronomic traits of wheat landrace varieties and commercial cultivars. Aust J Crop Sci 10:377–384
Henkrar F, El-Haddoury J, Iraqi D, Bendaou N, Udupa SM (2017) Allelic variation at high-molecular weight and low-molecular weight glutenin subunit genes in Moroccan bread wheat and durum wheat cultivars.3. Biotech 7:287
Herrera-Foessel S, Singh R, Huerta-Espino J, Crossa J, Yuen J, Djurle A (2006) Effect of leaf rust on grain yield and yield traits of durum wheats with race-specific and slow-rusting resistance to leaf rust. Plant Dis 90:1065–1072
Hewitt PH, Van Niekerk, GJJ, Walters MC, Kriel CF, Fouche A (1984) Aspects of the ecology of the Russian wheat aphid, Diuraphis noxia, in the Bloemfontein district. I. The colonization and infestation of sown wheat, identification of summer hosts and cause of infestation symptoms. Technical Communication, Department of Agriculture, South Africa, (191), p. 3–13
Huang XQ, Hsam SLK, Zeller FJ (1997) Identification of powdery mildew resistance genes in common wheat (Triticum aestivum L. em Thell.). IX. Cultivars, land races and breeding lines grown in China. Plant Breed 116:233–238
Huang XQ, Hsam SLK, Zeller FJ, Wenzel G, Mohler V (2000) Molecular mapping of the wheat powdery mildew resistance gene Pm24 and marker validation for molecular breeding. Theor Appl Genet 101(3):407–414
Hussain B, Lucas SJ, Ozturk L, Budak H (2017) Mapping QTLs conferring salt tolerance and micronutrient concentrations at seedling stage in wheat. Sci Rep 7(1):1–14
Hysing SC, Merker A, Liljeroth E, Koebner RM, Zeller FJ, Hsam SL (2007) Powdery mildew resistance in 155 Nordic bread wheat cultivars and landraces. Hereditas 144(3):102–119
Iizumi T, Furuya J, Shen Z, Kim W, Okada M, Fujimori S, Hasegawa T, Nishimori M (2017) Responses of crop yield growth to global temperature and socioeconomic changes. Sci Rep 7(1):1–10
Ilyas N, Amjid MW, Saleem MA, Khan W, Wattoo FM, Rana RM, Rana HM, Zahid A, Shah GA, Anwar A, Ahmad MQ, Shaheen M, Riaz H, Ansari MJ (2020) Quantitative trait loci (QTL) mapping for physiological and biochemical attributes in a Pasban90/Frontana recombinant inbred lines (RILs) population of wheat (Triticum aestivum) under salt stress condition. Saudi J Biol Sci 27(1):341–351
Inoue Y, Vy TT, Yoshida K, Asano H, Mitsuoka C, Asuke S, Anh VL, Cumagun CJR, Chuma I, Terauchi R, Kato K, Mitchell T, Valent B, Farman M, Yukio TY (2017) Evolution of the wheat blast fungus through functional losses in a host specificity determinant. Science 357(6346):80–83
Iqbal M, Navabi A, Salmon DF, Yang RC, Spaner D (2007) Simultaneous selection for early maturity, increased grain yield and elevated grain protein content in spring wheat. Plant Breed 126:244–250
Iwaki K, Haruna S, Niwa T, Kato K (2001) Adaptation and ecological differentiation in wheat with special reference to geographical variation of growth habit and Vrn genotype. Plant Breed 120:107–114
Izanloo A, Condon AG, Langridge P, Tester M, Schnurbusch T (2008) Different mechanisms of adaptation to cyclic water stress in two South Australian bread wheat cultivars. J Exp Bot 59(12):3327–3346
Jalal A, Shah S, Filho MCMT, Khan A, Shah T, Ilyas M, Rosa PAL (2020) Agro-Biofortification of Zinc and Iron in Wheat Grains. Gesunde Pflanzen 72:227–236
Jaleel CA, Manivannan P, Wahid A, Farooq M, Somasundaram R, Panneerselvam R (2009) Drought stress in plants: a review on morphological characteristics and pigments composition. Int J Agric Biol 11:100–105
Jan SU, Jamil M, Alipour H, Bhatti MF, Gul A (2017) Analysis of salinity tolerance potential in synthetic hexaploid wheat. Pak J Bot 49(4):1269–1278
Janni M, Cadonici S, Bonas U, Grasso A, Dahab AAD, Visioli G, Pignone D, Ceriotti A, Marmiroli N (2018) Gene-ecology of durum wheat HMW glutenin reflects their diffusion from the center of origin. Sci Rep 8:16929
Jaradat A (2006) Phenotypic divergence in the meta-population of the Hourani wheat landrace. J Food Agric Env 4:186–191
Jaradat AA (2011) Wheat landraces: genetic resources for sustenance and sustainability. usda-ars, pp 1–20. http://www.usmarc.usda.gov/SP2UserFiles/Place/36450000/products-wheat/AAJ-wheatlandraces.pdf
Jaradat AA (2013) Wheat landraces: a mini review. Emir J Food Agric 25:20–29
Jiang Y, Huang L, Hu Y (2010) Distribution of vernalization genes in Chinese wheat landraces and their relationship with winter hardness. Sci Agric Sin 43:2619–2632
Joppa LR, Cantrell RG (1990) Chromosomal location of genes for grain protein content of wild tetraploid wheat. Crop Sci 30:1059–1064
Joppa LR, Du C, Hart GE, Hareland GA (1997) Mapping gene(s) for grain protein in tetraploid wheat (Triticum turgidum L.) using a population of recombinant inbred chromosome lines. Crop Sci 37:1586–1589
Jordan KW, Wang S, Lun Y, Gardiner LJ, MacLachlan R, Hucl P, Wiebe K, Wong D, Forrest KL, Sharpe AG, Sidebottom CHD, Hall N, Toomajian C, Close T, Dubcovsky J, Akhunova A, Talbert L, Bansal UK, Bariana HS, Hayden MJ, Pozniak C, Jeddeloh JA, Anthony Hall A, Akhunov E (2015) A haplotype map of allohexaploid wheat reveals distinct patterns of selection on homoeologous genomes. Genome Biol 16(1):1–18
Jorgensen C, Luo MC, Ramasamy R, Dawson M, Gill BS, Korol AB, Distelfeld A, Dvorak J (2017) A high-density genetic map of wild emmer wheat from the Karaca Dağ region provides new evidence on the structure and evolution of wheat chromosomes. Front Plant Sci 8:1798
Kahrizi D, Cheghamirza K, Kakaei M, Mohammadi R, Ebadi A (2010) Heritability and genetic gain of some morphophysiological variables of durum wheat (Triticum turgidum var. durum). Afr J Biotechnol 9(30):4687–4691
Kamal NM, Gorafi YSA, Mega R, Tsujimoto H (2018) Physiological response of wheat to chemical desiccants used to simulate post-anthesis drought stress. Agronomy 8(4):44
Kamran A, Iqbal M, Navabi A, Randhawa HS, Pozniak C, Spaner D (2013) Earliness per QTLs and their interaction with photoperiod insensitive allele Ppd-D1a in Cutler x AC Barrie spring wheat population. Theor Appl Genet 126:1965–1976
Kandel JS, Krishnan V, Jiwan D, Chen X, Skinner DZ, See DR (2017) Mapping genes for resistance to stripe rust in spring wheat landrace PI 480035. PloS one 12(5):e0177898
Kankwatsa P, Singh D, Thomson PC, Babiker EM, Bonman JM, Newcomb M, Park RF (2017) Characterization and genome-wide association mapping of resistance to leaf rust, stem rust and stripe rust in a geographically diverse collection of spring wheat landraces. Mol Breed 37(9):113
Karagöz A, Zencirci N (2005) Variation in wheat (Triticum spp.) landraces from different altitudes of three regions of Turkey. Genet Resour Crop Ev 52(6):775–785
Karan R, Subudhi PK (2012) Approaches to increasing salt tolerance in crop plants. In: Abiotic stress responses in plants. Springer, New York, pp 63–88
Kato K, Mori Y, Beiles A, Nevo E (1997) Geographical variation in heading traits in wild emmer wheat, Triticum dicoccoides. I. Variation in vernalization response and ecological differentiation. Theor Appl Genet 95:546–552
Kato K, Taketa S, Ban T, Iriki N, Miura K (2001) The influence of a spring habit gene, Vrn-D1, on heading time in wheat. Plant Breed 120:115–120
Kato K, Wada T (1999) Genetic analysis and selection experiment for narrow-sense earliness in wheat by using segregating hybrid progenies. Breed Sci 49:233–238
Kato K, Yokoyama H (1992) Geographical variation in heading characters among wheat landraces, Triticum aestivum L., and its implication for their adaptability. Theor Appl Genet 84:259–265
Kertho A, Mamidi S, Bonman JM, McClean PE, Acevedo M (2015) Genome-wide association mapping for resistance to leaf and stripe rust in winter-habit hexaploid wheat landraces. PLoS One 10(6):e0129580
Khan MA, Fuller MP, Baloch FS (2008) Effect of soil applied zinc sulphate on wheat (Triticum aestivum L.) grown on a calcareous soil in Pakistan. Cereal Res Commun 36:571–582
Khan MH, Bukhari A, Dar ZA, Rizvi SM (2013) Status and strategies in breeding for rust resistance in wheat. Agr Sci 4:292
Khokhar JS, King J, King IP, Young SD, Foulkes MJ, De Silva J et al (2020) Novel sources of variation in grain Zinc (Zn) concentration in bread wheat germplasm derived from Watkins landraces. PLoS One 15(2):e0229107
Kidane YG, Gesesse CA, Hailemariam BN, Desta EA, Mengistu DK, Fadda C, Pe ME, Dell’Acqua M (2019) A large nested association mapping population for breeding and quantitative trait locus mapping in Ethiopian durum wheat. Plant Biotechnol J 17(7):1380–1393
Kidane YG, Hailemariam BN, Mengistu DK, Fadda C, Pè ME, Dell’Acqua M (2017) Genome-wide association study of Septoria tritici blotch resistance in Ethiopian durum wheat landraces. Front Plant Sci 8:1586
Kilian B, Ozkan H, Pozzi C, Salamini F (2009) Domestication of the Triticeae in the Fertile Crescent. In: Feuillet C, Muehlbauer GJ (eds) Genetics and genomics of the Triticeae. USA pp, Springer-Verlag, New York, pp 81–119
King JE, Cook RJ, Melville SC (1983) A review of Septoria diseases of wheat and barley. Ann Appl Biol 103(2):345–373
Kishii M (2019) An update of recent use of Aegilops species in wheat breeding. Front Plant Sci 10:585
Klindworth DL, Hareland GA, Elias EM, Faris JD, Chao S, Xu SS (2009) Agronomic and quality characteristics of two new sets of Langdon durum-wild emmer wheat chromosome substitution lines. J Cereal Sci 50:29–35
Kolev S, Vassilev D, Kostov K, Todorovska E (2011) Allele variation in loci for adaptive response in Bulgarian wheat cultivars and landraces and its effect on heading date. Plant Genet Res 9:251–255
Kolmer JA, Garvin DF, Hayden M, Spielmeyer W (2018) Adult plant leaf rust resistance derived from the wheat landrace cultivar Americano 44d is conditioned by interaction of three QTL. Euphytica 214(3):59
Korkut ZK, Balkan A, Başer İ, Bilgin O (2019) Grain Yield and Some Physiological Traits Associated with Heat Tolerance in Bread Wheat (Triticum aestivum L.) Genotypes. J Agric Sci 25(3):391–400
Kumar J, Jaiswal V, Kumar A, Kumar N, Mir RR, Kumar S, Dhariwala R, Tyagia S, Khandelwale M, Prabhub KV, Prasade R, Balyana HS, Guptaa PK (2011) Introgression of a major gene for high grain protein content in some Indian bread wheat cultivars. Field Crop Res 123:226–233
Kumar J, Schäfer P, Hückelhoven R, Langen G, Baltruschat H, Stein E, Nagarajan S, Kogel KH (2002) Bipolaris sorokiniana, a cereal pathogen of global concern: cytological and molecular approaches towards better control. Mol Plant Pathol 3(4):185–195
Kumar S, Archak S, Tyagi RK, Kumar J, Vikas VK, Jacob SR, Srinivasan K, Radhamani J, Parimalan R, Sivaswamy M, Tyagi S, Yadav M, Kumari J, Deepali SS, Bhagat I, Meeta M, Bains NS, Chowdhury AK, Saha BC, Bhattacharya PM, Kumari J, Singh MC, Gangwar OP, Prasad P, Bharadwaj SC, Gogoi R, Sharma JB, Kumar GMS, Saharan MS, Bag M, Roy A, Prasad TV, Sharma RK, Dutta M, Sharma I, Bansal KC (2016) Evaluation of 19,460 wheat accessions conserved in the Indian national genebank to identify new sources of resistance to rust and spot blotch diseases. PLoS One 12:e0175610
Kumar S, Beena AS, Awana M, Singh A (2017) Physiological, biochemical, epigenetic and molecular analyses of wheat (Triticum aestivum) genotypes with contrasting salt tolerance. Front Plant Sci 8:1151
Law CN, Worland AJ, Giorgi B (1976) The genetic control of ear-emergence time by chromosomes 5A and 5D of wheat. Heredity 36:49–58
Leonard K, Szabo L (2005) Stem rust of small grains and grasses caused by Puccinia graminis. Mol Plant Pathol 6:99–111
Li G, Carver BF, Cowger C, Bai G, Xu X (2018a) Pm223899, a new recessive powdery mildew resistance gene identified in Afghanistan landrace PI 223899. Theor Appl Genet 131(12):2775–2783
Li G, Xu X, Bai G, Carver BF, Hunger R, Bonman JM (2016a) Identification of novel powdery mildew resistance sources in wheat. Crop Sci 56(4):1817–1830
Li G, Xu X, Carver BF, Guo P, Puterka G (2018b) Dn10, a new gene conferring resistance to Russian wheat aphid biotype 2 in Iranian wheat landrace PI 682675. Crop Sci 58(3):1219–1225
Li H, Vikram P, Singh RP, Kilian A, Carling J, Song J, Burgueno-Ferreira JA, Bhavani S, Huerta-Espino J, Payne T, Sehgal D, Wenzl P, Singh S (2015) A high density GBS map of bread wheat and its application for dissecting complex disease resistance traits. BMC Genomics 16(1):1–15.
Li T, Bai G, Wu S, Gu S (2011) Quantitative trait loci for resistance to fusarium head blight in a Chinese wheat landrace Haiyanzhong. Theor Appl Genet 122(8):1497–1502
Li T, Zhang D, Zhou X, Bai G, Li L, Gu S (2016b) Fusarium head blight resistance loci in a stratified population of wheat landraces and varieties. Euphytica 207(3):551–561
Li XJ, Xu X, Yang XM, Li XQ, Liu WH, Gao AN, Li LH (2012) Genetic diversity of the wheat landrace Youzimai from different geographic regions investigated with morphological traits, seedling resistance to powdery mildew, gliadin and microsatellite markers. Cereal Res Commun 40(1):95–106
Li X, Li Y, Zhang M, Yu X, Hu R, Chang J, Yang G, Wang Y, He G (2019) Diversity of Puroindoline genes and their association with kernel hardness in Chinese wheat cultivars and landraces. Mol Breed 39(4):1–13
Li Y, Huang C, Sui X, Fan Q, Li G, Chu X (2009) Genetic variation of wheat glutenin subunits between landraces and varieties and their contributions to wheat quality improvement in China. Euphytica 169:159–168
Li Y, Shi X, Hu J, Wu P, Qiu D, Qu Y, Xie J, Wu Q, Zhang H, Yang L, Liu H, Zhou Y, Liu Z, Li H (2020) Identification of a recessive gene PmQ conferring resistance to powdery mildew in wheat landrace Qingxinmai using BSR-Seq analysis. Plant Disease 104(3):743–751
Liu D, Liu Y, Zhang W, Chen X, Zou C (2017a) Agronomic approach of zinc biofortification can increase zinc bioavailability in wheat flour and thereby reduce zinc deficiency in humans. Nutrients 9(5):465
Liu D, Zhang L, Hao M, Ning S, Yuan Z, Dai S, Huang L, Wu B, Yan Z, Lan X, Zheng Y (2018) Wheat breeding in the hometown of Chinese Spring. Crop J 6:82–90
Liu J, Huang L, Wang C, Liu Y, Yan Z, Wang Z, Xiang L, Zhong X, Gong F, Zheng Y, Liu D, Wu B (2019) Genome-Wide Association Study Reveals Novel Genomic Regions Associated With High Grain Protein Content in Wheat Lines Derived From Wild Emmer Wheat. Front Plant Sci 10:464
Liu W, Maccaferri M, Rynearson S, Letta T, Zegeye H, Tuberosa R, Chen X, Pumphrey M (2017b) Novel sources of stripe rust resistance identified by genome-wide association mapping in Ethiopian durum wheat (Triticum turgidum ssp. durum). Front Plant Sci 8:774.1
Liu Y, Lin Y, Gao S, Li Z, Deng M, Chen G, Wei Y, Zheng Y (2017c) A genome-wide association study of 23 agronomic traits in Chinese wheat landraces. Plant J 91:861–873
Liu Y, Qie Y, Li X, Wang M, Chen X (2020) Genome-Wide Mapping of Quantitative Trait Loci Conferring All-Stage and High-Temperature Adult-Plant Resistance to Stripe Rust in Spring Wheat Landrace PI 181410. Int J Mol Sci 21(2):478
Lodhi SS, Maryam S, Rafique K, Shafique A, Yousaf ZA, Talha AM, Gul A, Amir R (2020) Overview of the prospective strategies for conservation of genomic diversity in wheat landraces. In: Climate change and food security with emphasis on wheat. Academic Press, London, pp 293–309
Long L, Yao F, Yu C, Ye X, Cheng Y, Wang Y, Wu Y, Li J, Wang J, Jiang Q, Li W, Ma J, Liu Y, Deng M, Wei Y, Zheng Y, Chen G (2019) Genome-Wide association study for adult-plant resistance to stripe rust in Chinese wheat landraces (Triticum aestivum L.) from the yellow and huai river valleys. Front Plant Sci 10:596
Lopes MS, El-Basyoni I, Baenziger PS, Singh S, Royo C, Ozbek K, Aktas H, Ozer E, Ozdemir F, Manickavelu A, Ban T, Vikram P (2015) Exploiting genetic diversity from landraces in wheat breeding for adaptation to climate change. J Exp Bot 66:3477–3486
Lyons G, Ortiz-Monasterio I, Stangoulis J, Graham R (2005) Selenium concentration in wheat grain: Is there sufficient genotypic variation to use in breeding? Plant and Soil 269:369–380
Ma D, Li Q, Tang M, Chao K, Li J, Wang B, Jing J (2015) Mapping of gene conferring adult-plant resistance to stripe rust in Chinese wheat landrace Baidatou. Mol Breed 35(8):157
Ma L, Zhou E, Huo N, Zhou R, Wang G, Jia J (2007) Genetic analysis of salt tolerance in a recombinant inbred population of wheat (Triticum aestivum L.). Euphytica 153(1-2):109–117
Mahrookashani A, Siebert S, Hüging H, Ewert F (2017) Independent and combined effects of high temperature and drought stress around anthesis on wheat. J Agron Crop Sci 203(6):453–463
Manickavelu A, Joukhadar R, Jighly A, Lan C, Huerta-Espino J, Stanikzai AS, Kilian A, Singh RP, Ban T (2016) Genome wide association mapping of stripe rust resistance in Afghan wheat landraces. Plant Sci 252:222–229
Manickavelu A, Niwa S, Ayumi K, Komatsu K, Naruoka Y, Ban T (2014) Molecular evaluation of Afghan wheat landraces. Plant Genet Resour-C 12:S31–S35
Mantri N, Patade V, Penna S, Ford R, Pang E (2012) In: Ahmad P, Prasad MNV (eds) Abiotic stress responses in plants: metabolism, productivity and sustainability. Springer Science+Business Media, LLC, New York
Maraite H, Di Zinno T, Longree H, Daumerie V, Duveiller E (1997) Fungi associated with foliar blight of wheat in warm areas. In Proceedings of the international workshop on helminthosporium diseases of wheat: Spot Blotch and Tan Spot, El Batán (pp. 293–300)
Masood MS, Javaid A, Rabbani MA, Anwar R (2005) Phenotypic diversity and trait association in bread wheat (Triticum aestivum L.) landraces from Baluchistan, Pakistan. Pak J Bot 37:949
McIntosh RA, Wellings CR, Park RF (1995) Wheat rusts: an atlas of resistance genes. CSIRO Publications, East Melbourne (Australia)
McIntosh RA, Yamazaki Y, Dubcovsky J, Rogers J, Morris C, Appels R, Xia XC (2013) Catalogue of gene symbols for wheat. https://shigen.nig.ac.jp/wheat/komugi/genes/download.jsp
McKay JK, Richards JH, Mitchell-Olds T (2003) Genetics of drought adaptation in Arabidopsis thaliana: I. Pleiotropy contributes to genetic correlations among ecological traits. Mol Ecol 12:1137–1151
Melnikova NV, Ganeva GD, Popova ZG, Landjeva SP, Kudryavtsev AM (2010) Gliadins of Bulgarian durum wheat Triticum durum Desf. landraces: genetic diversity and geographical distribution. Genet Resour Crop Evol 57:587–595
Mengistu N, Baenziger PS, Eskridge KM, Dweikat I, Wegulo SN, Gill KS, Mujeeb-Kazi A (2012) Validation of QTL for grain yield-related traits on wheat chromosome 3a using recombinant inbred chromosome lines. Crop Sci 52:1622–1632
Merchuk-Ovnat L, BarakV FT, Ordon F, Lidzbarsky GA, Krugman T, Saranga Y (2016) Ancestral QTL alleles from wild emmer wheat improve drought resistance and productivity in modern wheat cultivars. Front Plant Sci 7:452
Mir RA, Sharma A, Mahajan R (2020) Crop landraces: present threats and opportunities for conservation. In: Rediscovery of genetic and genomic resources for future food security 2020. Springer, Singapore, pp 335–349
Mir Ali N, Arabi MIE, Al-Safadi B (1999) Frequencies of high and low molecular weight glutenin subunits in durum wheat grown in Syria. Cereal Res Commun 27:301–305
Mishra VK, Gupta PK, Arun B, Vasistha NK, Vishwakarma MK, SinghYadav P, Kumar H, Joshiac AK (2015) Introgression of a gene for high grain protein content (Gpc-B1) into two leading cultivars of wheat in Eastern Gangetic Plains of India through marker assisted backcross breeding. J Plant Breed Crop Sci 7:292–300
Mitrofanova OP, Khakimova AG (2017) New genetic resources in wheat breeding for increased grain protein content. Russ J Genet Appl Res 7(4):477–487
Mohammadi R, Armion M, Kahrizi D, Amri A (2012) Efficiency of screening techniques for evaluating durum wheat genotypes under mild drought conditions. Int J Plant Prod 4(1):11–24
Mohammadi R, Sadeghzadeh B, Ahmadi H, Bahrami N, Amri A (2015) Field evaluation of durum wheat landraces for prevailing abiotic and biotic stresses in highland rainfed regions of Iran. The Crop Journal 3(5):423–433
Monasterio I, Graham RD (2000) Breeding for trace minerals in wheat. Food Nutr Bull 21:392–396
Mondal S, Rutkoski JE, Velu G, Singh PK, Crespo-Herrera LA, Guzman C, Bhavani S, Lan C, He X, Singh RP (2016) Harnessing diversity in wheat to enhance grain yield, climate resilience, disease and insect pest resistance and nutrition through conventional and modern breeding approaches. Front Plant Sci 7:991
Mora F, Castillo D, Lado B, Matus I, Poland J, Belzile F, Zitzewitz JV, del Pozo A (2015) Genome-wide association mapping of agronomic traits and carbon isotope discrimination in a worldwide germplasm collection of spring wheat using SNP markers. Mol Breed 35(2):69
Moragues M, Zarco-Hernández J, Moralejo MA, Royo C (2006) Genetic diversity of glutenin protein subunits composition in durum wheat landraces [Triticum turgidum ssp. turgidum convar. durum (Desf.) MacKey] from the Mediterranean basin. Genet Resour Crop Evol 53(5):993–1002
Motzo R, Giunta F (2007) The effect of breeding on the phenology of Italian durum wheats: From landraces to modern cultivars. Eur J Agron 26:462–470
Mohammed AR, Tarpley L (2009) High nighttime temperatures affect rice productivity through altered pollen germination and spikelet fertility. Agric For Meteorol 149(6–7):999–1008
Mujeeb-Kazi A, Gul A, Ahmad I, Farooq M, Rauf Y, Riaz H (2009) Genetic resources for some wheat abiotic stress tolerances. In: Salinity and water stress. Springer, Dordrecht, pp 149–163
Muqaddasi QH, Reif JC, Li Z, Basnet BR, Dreisigacker S, Röder MS (2017) Genome-wide association mapping and genome-wide prediction of anther extrusion in CIMMYT spring wheat. Euphytica 213(3):73
Mwadzingeni L, Shimelis H, Tesfay S, Tsilo TJ (2016) Screening of bread wheat genotypes for drought tolerance using phenotypic and proline analyses. Front Plant Sci 7:1276
Nadeem MA, Nawaz MA, Shahid MQ, Doğan Y, Comertpay G, Yıldız M, Hatipoğlu R, Ahmad F, Alsaleh A, Labhane N, Özkan H (2018) DNA molecular markers in plant breeding: current status and recent advancements in genomic selection and genome editing. Biotechnol Biotechnol Equip 32:261–285
Naghavi MR, Monfared SR, Ahkami AH, Ombidbakhsh MA (2009) Genetic variation of durum wheat landraces and cultivars using morphological and protein markers. Proceedings of world academy of science. Eng Technol 37:73–75
Nazco R, Villegas D, Ammar K, Pena RJ, Moragues M, Royo C (2012) Can Mediterranean durum wheat landraces contribute to improved grain quality attributes in modern cultivars? Euphytica 185(1):1–17
Newcomb M, Acevedo M, Bockelman HE, Brown-Guedira G, Goates BJ, Jackson EW, Jin Y, Njau P, Rouse MN, Singh RD, Wanyera R, Bonman JM (2013) Field resistance to the Ug99 race group of the stem rust pathogen in spring wheat landraces. Plant disease 97(7):882–890
Newton AC, Akar T, Baresel JP, Bebeli PJ, Bettencourt E, Bladenopoulos KV, Czembor JH, Fasoula DA, Katsiotis A, Koutis K, Koutsika-Sotiriou M, Kovacs G, Larsson H, Pinheiro de Carvalho MAA, Rubiales D, Russell J, Dos Santos TMM, Vaz Patto MC (2010) Cereal landraces for sustainable agriculture. Sustain Agric 2:147–186
Newton AC, Johnson SN, Gregory PJ (2011) Implications of climate change for diseases, crop yields and food security. Euphytica 179:3–18
Nicol J, Rivoal R, Taylor S, Zaharieva M (2003) Global importance of cyst (Heterodera spp.) and lesion nematodes (Pratylenchus spp.) on cereals: distribution, yield loss, use of host resistance and integration of molecular tools. Nematol Monogr Perspect 2:1–19
Nicol JM, Rivoal R, Bolat N, Aktas H, Braun HJ, Mergoum M, Yildrim AF, Bagci A, Eleckcioglu IH, Yahyaoui A (2002) The frequency and diversity of the cyst and lesion nematode on wheat in the Turkish Central Anatolian Plateau. Nematology 4(2):272
Norman A, Taylor J, Tanaka E, Telfer P, Edwards J, Martinant JP, Kuchel H (2017) Increased genomic prediction accuracy in wheat breeding using a large Australian panel. Theor Appl Genet 130(12):2543–2555
O’Driscoll A, Kildea S, Doohan F, Spink J, Mullins E (2014) The wheat–Septoria conflict: a new front opening up? Trends Plant Sci 19(9):602–610
Okechukwu EC, Agbo CU, Uguru MI, Ogbonnaya FC (2016) Germplasm evaluation of heat tolerance in bread wheat in Tel Hadya, Syria. Chil J Agric Res 76(1):9–17
Olmstead AL, Rhode PW (2002) The red queen and the hard reds: productivity grown in American wheat 1800–1940. J Econ Hist 62:929–966
Ouaja M, Aouini L, Bahri B, Ferjaoui S, Medini M, Marcel TC, Hamza S (2020) Identification of valuable sources of resistance to Zymoseptoria tritici in the Tunisian durum wheat landraces. Eur J Plant Pathol 156(2):647–661
Oak MD, Tamhankar SA (2017) 1BL/1RS translocation in durum wheat and its effect on end use quality traits. J Plant Biochem Biot 26(1):91–96
Ozberk I, Atay S, Altay F, Cabi E, Ozkan H, Atli A (2016) The Wheat Atlas of Turkey. (World Wildlife Fund), Istanbul (in Turkish)
Ozer G, Paulitz TC, Imren M, Alkan M, Muminjanov H, Dababat AA (2020) Identity and Pathogenicity of Fungi Associated with Crown and Root Rot of Dryland Winter Wheat in Azerbaijan. Plant Disease 104:2149–2157
Özkan H, Brandolini A, Schäfer-Pregl R, Salamini F (2002) AFLP analysis of a collection of tetraploid wheats indicates the origin of emmer and hard wheat domestication in southeast Turkey. Mol Biol Evol 19(10):1797–1801
Özkan H, Willcox G, Graner A, Salamini F, Kilian B (2011) Geographic distribution and domestication of wild emmer wheat (Triticum dicoccoides). Genet Resour Crop Evol 58(1):11–53
Pasam RK, Bansal U, Daetwyler HD, Forrest KL, Wong D, Petkowski J, Willey N, Randhawa M, Chhetri M, Miah H, Tibbits J, Bariana H, Hayden MJ (2017) Detection and validation of genomic regions associated with resistance to rust diseases in a worldwide hexaploid wheat landrace collection using BayesR and mixed linear model approaches. Theor Appl Genet 130(4):777–793
Peng JH, Sun D, Nevo E (2011a) Domestication evolution, genetics and genomics in wheat. Mol Breed 28:281–301
Peng ZS, Li X, Yang ZJ, Liao ML (2011b) A new reduced height gene found in the tetraploid semi-dwarf wheat landrace Aiganfanmai. Genet Mol Res 10:2349–2357
Peusha H, Lebedeva T, Prilinn O, Enno T (2002) Genetic analysis of durable powdery mildew resistance in a common wheat line. Hereditas 136:201–206
Pinto RS, Molero G, Reynolds MP (2017) Identification of heat tolerant wheat lines showing genetic variation in leaf respiration and other physiological traits. Euphytica 213(3):76
Poersch-Bortolon LB, Pereira JF, Nhani Junior A, Gonzáles HHS, Torres GAM, Consoli L, Arenhart RA, Bodanese-Zanettini MH, Margis-Pinheiro M (2016) Gene expression analysis reveals important pathways for drought response in leaves and roots of a wheat cultivar adapted to rainfed cropping in the Cerrado biome. Genet Mol Biol 39(4):629–645
Popov C, Hondru N, Bărbulescu A, Vonica I, Mărgărit G (1988) Species of aphids attacking wheat and barley crops. Analele Institutului de Cercetări pentru Cereale și Plante Tehnice, Fundulea 56:379–384
Pu Z, Pei Y, Yang J, Ma J, Li W, Liu D, Wang J, Wei Y, Zheng Y (2018) A QTL located on chromosome 3D enhances the selenium concentration of wheat grain by improving phytoavailability and root structure. Plant and Soil 425(1-2):287–296
Pugsley AT (1971) A genetic analysis of the spring-winter habit of growth in wheat. Aust J Agr Res 22:21–23
Pugsley AT (1972) Additional genes inhibiting winter habit in wheat. Euphytica 21:547–552
Qaseem MF, Qureshi R, Muqaddasi QH, Shaheen H, Kousar R, Röder MS (2018) Genome-wide association mapping in bread wheat subjected to independent and combined high temperature and drought stress. PLoS One 13:6
Qiang LI, Wang ZR, Ding LI, Wei JW, Qiao WC, Meng XH, Sun SI, Li HM, Zhao MH, Chen XM, Zhao FW (2018) Evaluation of a new method for quantification of heat tolerance in different wheat cultivars. J Integr Agric 17(4):786–795
Qie Y, Sheng Y, Xu H, Jin Y, Ma F, Li L, Li X, An D (2019) Identification of a new powdery mildew resistance gene pmDHT at or closely linked to the Pm5 locus in the Chinese wheat landrace Dahongtou. Plant disease 103(10):2645–2651
Quarrie SA, Steed A, Calestani C, Semikhodskii A, Lebreton C, Chinoy C, Steele N, Pljevljakusić D, Waterman E, Weyen J, Schondelmaier J, Habash DZ, Farmer P, Saker L, Clarkson DT, Abugalieva A, Yessimbekova M, Turuspekov Y, Abugalieva S, Tuberosa R, Sanguineti MC, Hollington PA, Aragués R, Royo A, Dodig D (2005) A high-density genetic map of hexaploid wheat (Triticum aestivum L.) from the cross Chinese Spring× SQ1 and its use to compare QTLs for grain yield across a range of environments. Theor Appl Genet 110(5):865–880
Quisenberry KS, Reitz LP (1974) Turkey wheat: The cornerstone of an empire. Agric Hist 48:98–110
Qureshi N, Bariana H, Kolmer JA, Miah H, Bansal U (2017) Genetic and molecular characterization of leaf rust resistance in two durum wheat landraces. Phytopathology 107(11):1381–1387
Qureshi N, Bariana H, Kumran VV, Muruga S, Forrest KL, Hayden MJ, Bansal U (2018) A new leaf rust resistance gene Lr79 mapped in chromosome 3BL from the durum wheat landrace Aus26582. Theor Appl Genet 131(5):1091–1098
Rahmatov M, Otambekova M, Muminjanov H, Rouse MN, Hovmøller MS, Nazari K, Steffenson BJ, Johansson E (2019) Characterization of stem, stripe and leaf rust resistance in Tajik bread wheat accessions. Euphytica 215(3):1–22
Ram S, Govindan V (2020) Improving wheat nutritional quality through biofortification. In: Igrejas G, Ikeda TM, Guzman C (eds) Wheat quality for improving processing and human health. Springer, Switzerland, pp 205–224
Rasheed A, Xia X, Mahmood T, Quraishi UM, Bux AAH, Mahmood Z, Mirza JI, Mujeeb-Kazi A, He Z (2016) Comparison of economically important loci in landraces and improved wheat cultivars from Pakistan. Crop Sci 56:287–301
Rawson HM, Richards RA, Munns R (1988) An examination of selection criteria for salt tolerance in wheat, barley and triticale genotypes. Aust J Agr Res 39:759–772
Reitz LP, Salmon SC (1968) Origin, history, and use of Norin 10 wheat. Crop Sci 8:686–689
Ren Y, Xu Y, Teng W, Li B, Lin T (2018) QTLs for seedling traits under salinity stress in hexaploid wheat. Cienc Rural 48(3):e20170446
Reynolds M, Dreccer F, Trethowan R (2007) Drought-adaptive traits derived from wheat wild relatives and landraces. J Exp Bot 58(2):177–186
Reynolds MP, Borlaug NE (2006a) Applying innovations and new technologies for international collaborative wheat improvement. J Agr Sci-Cambridge 144:95
Reynolds MP, Borlaug NE (2006b) Impacts of breeding on international collaborative wheat improvement. J Agr Sci-Cambridge 144:3–17
Riaz A, Athiyannan N, Periyannan S, Afanasenko O, Mitrofanova O, Aitken EA, Lagudah E, Hickey LT (2017) Mining Vavilov’s treasure chest of wheat diversity for adult plant resistance to Puccinia triticina. Plant disease 101(2):317–323
Ribaut JM, Hoisington D (1998) Marker-assisted selection: new tools and strategies. Trends Plant Sci 3(6):236–239
Ribeiro M, Carvalho C, Carnide V, Guedes-Pinto H, Igrejas G (2011) Towards allelic diversity in the storage proteins of old and currently growing tetraploid and hexaploid wheats in Portugal. Genet Resour Crop Evol 58:1051–1073
Roelfs A, Huerto-Espino J, Marshall D (1992) Rust diseases of wheat: concepts and methods of disease management. CIMMYT, Mexico
Rola EL, De Vallavieille-Pope C, Leconte M, Nazari K (2019) Diversity of genes for resistance to stripe rust in wheat elite lines, commercial varieties and landraces from Lebanon and Syria. Phytopathologia Mediterranea 58(3):607–627
Roselló M, Royo C, Álvaro F, Villegas D, Nazco R, Soriano JM (2018) Pasta-making quality QTLome from Mediterranean durum wheat landraces. Front Plant Sci 9:1512
Roshanzamir H, Kordenaeej A, Bostani A (2013) Mapping QTLs related to Zn and Fe concentrations in bread wheat (Triticum aestivum) grain using microsatellite markers. Iran J Genet Plant Breed 2:10–17
Royo C, Briceño-Félix GA (2011a) Spanish wheat pool. In: Bojean AP, Angus WJ, van Ginkel M (eds) The world wheat book. A history of wheat breeding, vol 2. Lavoisier, Paris, pp 121–154
Royo C, Dreisigacker S, Soriano JM, Lopes MS, Ammar K, Villegas D (2020) Allelic variation at the vernalization response (Vrn-1) and photoperiod sensitivity (Ppd-1) genes and their association with the development of durum wheat landraces and modern cultivars. Front Plant Sci 11:838
Royo C, Briceño-Félix GA (2011b) Spanish wheat pool. In: Bojean AP, Angus WJ, van Ginkel M (eds) The world wheat book. A history of wheat breeding. Lavoisier Publishing, Paris, pp 121–154
Ruiz M, Giraldo P, Royo C, Villegas D, Jose Aranzana M, Carrillo JM (2012) Diversity and genetic structure of a collection of spanish durum wheat landraces. Crop Sci 52(5):2262–2275
Saini DK, Devi P, Kaushik P (2020) Advances in genomic interventions for wheat biofortification: a review. Agronomy 10(1):62
Salvi S, Porfiri O, Ceccarelli S (2013) Nazareno Strampelli, the ‘Prophet’ of the green revolution. J Agr Sci-Cambridge 151:1–5
Sansaloni C, Franco J, Santos B, Percival-Alwyn L, Singh S, Petroli C, Campos J, Dreher K, Payne T, Marshall D, Kilian B (2020) Diversity analysis of 80,000 wheat accessions reveals consequences and opportunities of selection footprints. Nat Commun 11:1–2
Sareen S, Tyagi BS, Sarial AK, Tiwari V, Sharma I (2014) Trait analysis, diversity, and genotype x environment interaction in some wheat landraces evaluated under drought and heat stress conditions. Chil J Agric Res 74(2):135–142
Schmidt AL, McIntyre CL, Thompson J, Seymour NP, Liu CJ (2005) Quantitative trait loci for root lesion nematode (Pratylenchus thornei) resistance in Middle-Eastern landraces and their potential for introgression into Australian bread wheat. Aust J Agr Res 56(10):1059–1068
Schmidt J, Tricker PJ, Eckermann P, Kalambettu P, Garcia M, Fleury DL (2020) Novel alleles for combined drought and heat stress tolerance in wheat. Front Plant Sci 10:1800
Scott PR, Hollins TW (1974) Effects of eyespot on the yield of winter wheat. Ann Appl Biol 78(3):269–279
Shahzad A, Ahmad M, Iqbal M, Ahmed I, Ali GM (2012) Evaluation of wheat landrace genotypes for salinity tolerance at vegetative stage by using morphological and molecular markers. Genet Mol Res 11(1):679–692
Shamaya NJ, Shavrukov Y, Langridge P, Roy SJ, Tester M (2017) Genetics of Na+ exclusion and salinity tolerance in Afghani durum wheat landraces. BMC Plant Biol 17(1):209
Sharma DK, Torp AM, Rosenqvist E, Ottosen CO, Andersen SB (2017) QTLs and potential candidate genes for heat stress tolerance identified from the mapping populations specifically segregating for Fv/Fm in wheat. Front Plant Sci 8:1668
Shavrukov Y, Langridge P, Tester M (2009) Salinity tolerance and sodium exclusion in genus Triticum. Breed Sci 59(5):671–678
Shi R, Li H, Tong Y, Jing R, Zhang F, Zou C (2008) Identification of quantitative trait locus of zinc and phosphorus density in wheat (Triticum aestivum L.) grain. Plant and Soil 306:95–104
Singh R, Govindan V, Andersson MS (2017) Zinc-Biofortified Wheat: Harnessing Genetic Diversity for Improved Nutritional Quality. Sci Br Biofortif Ser 1:1–4
Smale M (1996) Understanding global trends in the use of wheat diversity and international flows of wheat genetic resources. CIMMYT, Mexico
Smale, M. and McBride, T., 1996. Understanding global trends in the use of wheat diversity and international flows of wheat genetic resources: part 1. CIMMYT 1995/96 World Wheat Facts and Trends: Understanding Global Trends in the Use of Wheat Diversity and International Flows of Wheat Genetic Resources (No. Look under series title. CIMMYT.). Centro Internacional de Mejoramiento de Maiz y Trigo (CIMMYT), Mexico
Sofalian O, Chaparzadeh N, Javanmard A, Hejazi MS (2008) Study the genetic diversity of wheat landraces from northwest of Iran based on ISSR molecular markers. Int J Agric Biol 10:466–468
Soriano JM, Villegas D, Aranzana MJ, García del Moral LF, Royo C (2016) Genetic structure of modern durum wheat cultivars and Mediterranean landraces matches with their agronomic performance. PLoS One 11:e0160983
Soriano JM, Villegas D, Sorrells MR, Royo C (2018) Durum wheat landraces from east and west regions of the Mediterranean basin are genetically distinct for yield components and phenology. Front Plant Sci 9:80
Stelmakh AF (1990) Geographic distribution of Vrn-genes in landraces and improved varieties of spring bread wheat. Euphytica 45:113–118
Sthapit J, Newcomb M, Bonman JM, Chen X, See DR (2014) Genetic diversity for stripe rust resistance in wheat landraces and identification of accessions with resistance to stem rust and stripe rust. Crop Sci 54(5):2131–2139
Sun H, Hu J, Song W, Qiu D, Cui L, Wu P, Zhang H, Liu H, Li L, Qu Y, Li Y, Li T, Cheng W, Zhou Y, Liu Z, Li J, Li H (2018) Pm61: A recessive gene for resistance to powdery mildew in wheat landrace Xuxusanyuehuang identified by comparative genomics analysis. Theor Appl Genet 131(10):2085–2097
Sun QM, Zhou RH, Gao LF, Zhao GY, Jia JZ (2009) The characterization and geographical distribution of the genes responsible for vernalization requirement in Chinese bread wheat. J Integr Plant Biol 51:423–432
Tadesse W, Amri A, Ogbonnaya FC, Sanchez-Garcia M, Sohail Q, Baum M (2016) Wheat. In: Mohar S, Upadhyaya H (eds) Genetic and genomic resources for grain cereals improvement. Academic Press, Oxford, UK, pp 81–124
Taiz L, Zeiger E (2006) Plant Physiology, 4th edn. Sinauer Associates Inc Publishers, Sunderland
Talas F, Longin F, Miedaner T (2011) Sources of resistance to Fusarium head blight within Syrian durum wheat landraces. Plant breeding 130(3):398–400
Tan C, Li G, Cowger C, Carver BF, Xu X (2018) Characterization of Pm59, a novel powdery mildew resistance gene in Afghanistan wheat landrace PI 181356. Theor Appl Genet 131(5):1145–1152
Tan C, Li G, Cowger C, Carver BF, Xu X (2019) Characterization of Pm63, a powdery mildew resistance gene in Iranian landrace PI 628024. Theor Appl Genet 132(4):1137–1144
Tehseen MM, Tonk FA, Tosun M, Amri A, Sansaloni CP, Kurtulus E, Yazbek M, Al-Sham’aa K, Ozseven I, Safdar LB, Shehadeh A, Nazari K (2020) Genome Wide Association Study of Resistance to PstS2 and Warrior Races of Stripe (Yellow) Rust in Bread Wheat Landraces. bioRxiv
Thakur P, Kumar S, Malik JA, Berger JD, Nayyar H (2010) Cold stress effects on reproductive development in grain crops: an overview. Environ Exp Bot 67(3):429–443
Thirumalaikumar VP, Devkar V, Mehterov N, Ali S, Ozgur R, Turkan I, Mueller-Roeber B, Balazadeh S (2018) NAC transcription factor JUNGBRUNNEN 1 enhances drought tolerance in tomato. Plant Biotechnol J 16(2):354–366
Thompson AL, Smiley RW, Paulitz TC, Garland-Campbell K (2016) Identification of resistance to Pratylenchus neglectus and Pratylenchus thornei in Iranian landrace accessions of wheat. Crop Sci 56(2):654–672
Thompson JP, O’reilly MM, Clewett TG (2009) Resistance to the root-lesion nematode Pratylenchus thornei in wheat landraces and cultivars from the West Asia and North Africa (WANA) region. Crop Pasture Sci 60(12):1209–1217
Thompson JP, Seymour NP (2011) Inheritance of resistance to root-lesion nematode (Pratylenchus thornei) in wheat landraces and cultivars from the West Asia and North Africa (WANA) region. Crop Pasture Sci 62(1):82–93
Toor AK, Bansal UK, Bhardwaj S, Badebo A, Bariana HS (2013) Characterization of stem rust resistance in old tetraploid wheat landraces from the Watkins collection. Genet Resour Crop Evol 60(7):2081–2089
Toreti A, Cronie O, Zampieri M (2019) Concurrent climate extremes in the key wheat producing regions of the world. Sci Rep 9:5493
Touzy G, Rincent R, Bogard M, Lafarge S, Dubreuil P, Mini A, Deswarte JC, Beauchene K, Le Gouis J, Praud S (2019) Using environmental clustering to identify specific drought tolerance QTLs in bread wheat (T. aestivum L.). Theor Appl Genet 132(10):2859–2880
Turner AS, Faure S, Zhang Y, Laurie DA (2013) The effect of day-neutral mutations in barley and wheat on the interaction between photoperiod and vernalization. Theor Appl Genet 126:2267–2277
Uauy C, Brevis JC, Dubcovsky J (2006a) The high grain protein content gene Gpc-B1 accelerates senescence and has pleiotropic effects on protein content in wheat. J Exp Bot 57:2785–2794
Uauy C, Distelfeld A, Fahima T, Blechl A, Dubcovsky J (2006b) A NAC gene regulating senescence improves grain protein, zinc, and iron content in wheat. Science 314:1298–1301
Ullah S, Bramley H, Daetwyler H, He S, Mahmood T, Thistlethwaite R, Trethowan R (2018) Genetic contribution of emmer wheat (Triticum dicoccon Schrank) to heat tolerance of bread wheat. Front Plant Sci 9:1529
Valdez VA, Byrne PF, Lapitan NL, Peairs FB, Bernardo A, Bai G, Haley SD (2012) Inheritance and genetic mapping of Russian wheat aphid resistance in Iranian wheat landrace accession PI 626580. Crop Sci 52(2):676–682
Valluru R, Reynolds MP, Davies WJ, Sukumaran S (2017) Phenotypic and genome-wide association analysis of spike ethylene in diverse wheat genotypes under heat stress. New Phytol 214(1):271–283
Van Oosten MJ, Costa A, Punzo P, Landi S, Ruggiero A, Batelli G, Grillo S (2016) Genetics of drought stress tolerance in crop plants. In Drought Stress Tolerance in Plants, vol 2. Springer, Cham, pp 39–70
Varella AC, Weaver DK, Blake NK, Hofland ML, Heo HY, Cook JP, Lamb PF, Jordan KW, Akhunov E, Chao S, Talbert LE (2019) Analysis of recombinant inbred line populations derived from wheat landraces to identify new genes for wheat stem sawfly resistance. Theor Appl Genet 132(8):2195–2207
Varella AC, Weaver DK, Cook JP, Blake NK, Hofland ML, Lamb PF, Talbert LE (2017) Characterization of resistance to the wheat stem sawfly in spring wheat landrace accessions from targeted geographic regions of the world. Euphytica 213(7):153
Vats P, Banerjee UC (2004) Production studies and catalytic properties of phytases (myo-inositol hexakisphosphate phosphohydrolases): an overview. Enzyme Microb Technol 35(1):3–14
Veraverbeke WS, Delcour JA (2002) Wheat protein composition and properties of wheat glutenin in relation to breadmaking functionality. Crit Rev Food Sci Nutr 42:179–208
Vikram P, Franco J, Burgueño-Ferreira J, Li H, Sehgal D, Saint Pierre C, Ortiz C, Sneller C, Tattaris M, Guzman C, Sansaloni CP (2016a) Unlocking the genetic diversity of Creole wheats. Sci Rep 6:23092
Vikram P, Franco J, Burgueño-Ferreira J, Li H, Sehgal D, Saint Pierre C, Cynthia Ortiz C, Clay Sneller C, Maria Tattaris M, Carlos Guzman C, Carolina Paola Sansaloni CP, Ellis M, Fuentes-Davila G, Reynolds M, Sonder K, Singh P, Payne T, Wenzl P, Sharma A, Bains NS, Singh GP, Crossa J, Singh S (2016b) Unlocking the genetic diversity of Creole wheats. Sci Rep 6:23092
Villa TC, Maxted N, Scholten M, Ford-Lloyd B (2005) Defining and identifying crop landraces. Plant Genet Res 3(3):373–384
Vishwakarma MK, Arun B, Mishra VK, Yadav PS, Kumar H, Joshi AK (2016) Marker-assisted improvement of grain protein content and grain weight in Indian bread wheat. Euphytica 208:313–321
Von Rünker K (1908) Die Systematischeeinteilung und Benen-ung der Getreidesortenfu¨rpr aktische Zwecke. Jahrbuch der Deutschenlandwirtschafts-Gesellschaft 23:137–167
Wamalwa MN, Owuoche J, Ogendo J, Wanyera R (2019) Multi-Pathotype Testing of Selected Kenyan Wheat Germplasm and Watkin Landraces for Resistance to Wheat Stripe Rust (Puccinia striiformis f. sp tritici). Races Agronomy 9(11):770
Wamalwa M, Tadesse Z, Muthui L, Yao N, Zegeye H, Randhawa M, Wanyera R, Uauy C, Shorinola O (2020) Allelic diversity study of functional genes in East Africa bread wheat highlights opportunities for genetic improvement. Mol Breedi 40(11):1–14
Wang S, Asuke S, Vy TTP, Inoue Y, Chuma I, Win J, Kato K, Tosa Y (2018a) A new resistance gene in combination with Rmg8 confers strong resistance against Triticum isolates of Pyricularia oryzae in a common wheat landrace. Phytopathology 108(11):1299–1306
Wang Y, Peng H, Liu G, Xie C, Ni Z, Yang T, Liu Z, Sun Q (2010) Identification and molecular mapping of a leaf rust resistance gene in spelt wheat landrace Altgold. Euphytica 174(3):371–375
Wang Z, Huang L, Wu B, Hu J, Jiang Z, Qi P, Zheng Y, Liu D (2018b) Characterization of an integrated active Glu-1Ay allele in common wheat from wild emmer and its potential role in flour improvement. Int J Mol Sci 19:E923
Wang Z, Li H, Zhang D, Guo L, Chen J, Chen Y, Wu Q, Xie J, Zhang Y, Sun Q, Dvorak J, Luo M, Liu Z (2015) Genetic and physical mapping of powdery mildew resistance gene MlHLT in Chinese wheat landrace Hulutou. Theor Appl Genet 128(2):365–373
Wang Z, Ren J, Du Z, Che M, Zhang Y, Quan W, Jiang X, Ma Y, Zhao Y, Zhang Z (2019) Identification of a major QTL on chromosome arm 2AL for reducing yellow rust severity from a Chinese wheat landrace with evidence for durable resistance. Theor Appl Genet 132(2):457–471
Ward RW, Yang ZL, Kim HS, Yen C (1998) Comparative analyses of RFLP diversity in landraces of Triticum aestivum and collections of T. tauschii from China and Southwest Asia. Theor Appl Genet 96:312–318
Wei YM, Hou YC, Yan ZH, Wu W, Zhang ZQ, Liu DC, Zheng YL (2005) Microsatellite DNA polymorphism divergence in Chinese wheat (Triticum aestivum L.) landraces highly resistant to Fusarium head blight. J Appl Genet 46(1):3–9
White PJ, Broadley MR (2005) Biofortifying crops with essential mineral elements. Trends Plant Sci 10(12):586–593
WHO (2009) Global health risks, mortality and burden of disease attributable to selected major risks. Geneva, Switzerland, WHO
Witcombe JR, Joshi A, Joshi KD, Sthapit BR (1996) Farmer Participatory Crop Improvement. I. Varietal Selection and Breeding Methods and Their Impact on Biodiversity. Exp Agric 32(04):445–460
Worland AJ, Korzun V, Röder MS, Ganal MW, Law CN (1998) Genetic analysis of the dwarfing gene Rht8 in wheat. Part II. The distribution and adaptive significance of allelic variants at the Rht8 locus of wheat as revealed by microsatellite screening. Theor Appl Genet 96:1110–1120
Wu L, Xia X, Rosewarne GM, Zhu H, Li S, Zhang Z, He Z (2015) Stripe rust resistance gene Yr18 and its suppressor gene in Chinese wheat landraces. Plant Breed 134(6):634–640
Wu XL, Wang JW, Cheng YK, Ye XL, Li W, Pu ZE, Jiang QT, Wei YM, Deng M, Zheng YL, Chen GY (2016) Inheritance and molecular mapping of an all-stage stripe rust resistance gene derived from the Chinese common wheat landrace “Yilongtuomai”. J Hered 107(5):463–470
Xiao J, Jia X, Wang H, Zhao R, Fang Y, Gao R, Wu Z, Cao A, Wang J, Xue Z, Zhao W, Kang J, Chen Q, Chen P, Wang X (2011) A fast-neutron induced chromosome fragment deletion of 3BS in wheat landrace Wangshuibai increased its susceptibility to Fusarium head blight. Chromosome Res 19(2):225–234
Xiao J, Jin X, Jia X, Wang H, Cao A, Zhao W, Pei H, Xue Z, He L, Chen Q, Wang X (2013) Transcriptome-based discovery of pathways and genes related to resistance against Fusarium head blight in wheat landrace Wangshuibai. BMC Genomics 14(1):1–19
Xu H, Yi Y, Ma P, Qie Y, Fu X, Xu Y, Zhang X, An D (2015) Molecular tagging of a new broad-spectrum powdery mildew resistance allele Pm2c in Chinese wheat landrace Niaomai. Theor Appl Genet 128(10):2077–2084
Xu X, Li Q, Ma Z, Fan J, Zhou Y (2018) Molecular mapping of powdery mildew resistance gene PmSGD in Chinese wheat landrace Shangeda using RNA-seq with bulk segregant analysis. Mol Breed 38(3):23
Xu X, Liu W, Liu Z, Fan JR, Zhou Y (2020) Mapping powdery mildew resistance gene pmYBL on chromosome 7B of Chinese Wheat (Triticum aestivum L.) Landrace Youbailan. Plant Dis 104(9):2411–2417
Xu Y, An D, Liu D, Zhang A, Xu H, Li B (2012) Molecular mapping of QTLs for grain zinc, iron and protein concentration of wheat across two environments. Field Crop Res 138:57–62
Xue F, Wang C, Li C, Duan X, Zhou Y, Zhao N, Wang Y, Ji W (2012) Molecular mapping of a powdery mildew resistance gene in common wheat landrace Baihulu and its allelism with Pm24. Theor Appl Genet 125(7):1425–1432
Xue F, Zhai WW, Duan XY, Zhou YL, Ji WQ (2009) Microsatellite mapping of powdery mildew resistance gene in wheat landrace Xiaobaidong. Acta Agron Sin 34:1193–1198
Xynias IN, Kozub NA, Sozinov IA (2011) Analysis of hellenic durum wheat Triticum turgidum L. var. durum germplasm using gliadin and high-molecular-weight glutenin subunit loci. Cereal Res Commun 39:415–425
Yan L, Loukoianov A, Tranquilli G, Helguera M, Fahima T, Dubcovsky J (2003) Positional cloning of the wheat vernalization gene VRN1. Proc Natl Acad Sci U S A 100:6263–6268
Yeken MZ, Akpolat H, Karaköy T, Çiftçi V (2018) Assessment of Mineral Content Variations for Biofortification of the Bean Seed. Int J Agri Wild Sci 4(2):261–269
Yen C, Luo MC, Yang JL (1988) The origin of the Tibetan weedrace of hexaploid wheat, Chinese Spring, Chengdu-guang-tou and other landraces of the white wheat complex from China. In: Miller TE, Koebner RMD (eds.) Proceedings of the 7th International Wheat Genetics Symposium, Cambridge, pp 175–179
Yu S, Wu J, Wang M, Shi W, Xia G, Jia J, Kang Z, Han D (2020) Haplotype variations in QTL for salt tolerance in Chinese wheat accessions identified by marker-based and pedigree-based kinship analyses. Crop J. https://doi.org/10.1016/j.cj.2020.03.007
Yuan FP, Zeng QD, Wu JH, Wang QL, Yang ZJ, Liang BP, Kang ZS, Chen XH, Han DJ (2018) QTL mapping and validation of adult plant resistance to stripe rust in Chinese wheat landrace Humai 15. Front Plant Sci 9:968
Zampieri M, Ceglar A, Dentener F, Toreti A (2017) Wheat yield loss attributable to heat waves, drought and water excess at the global, national and subnational scales. Environ Res Lett 12(6):064008
Zeven AC (1998) Landraces: a review of definitions and classifications. Euphytica 104:127–139
Zeven AC (1999) The traditional inexplicable replacement of seed and seed ware of landraces and cultivars: a review. Euphytica 110:181–191
Zeven AC (2000) Traditional maintenance breeding of landraces: 1. Data by crop. Euphytica 116:65–85
Zhang K, Wang J, Qin H, Wei Z, Hang L, Zhang P, Reynolds M, Wang D (2019a) Assessment of the individual and combined effects of Rht8 and Ppd-D1a on plant height, time to heading and yield traits in common wheat. The Crop Journal 7:845–856
Zhang P, Gebrewahid TW, Zhou Y, Ll Q, LI Z, LIu D (2019b) Seedling and adult plant resistance to leaf rust in 46 Chinese bread wheat landraces and 39 wheat lines with known Lr genes. J Integr Agric 18(5):1014–1023
Zhang X, Pan H, Bai G (2012) Quantitative trait loci responsible for Fusarium head blight resistance in Chinese landrace Baishanyuehuang. Theor Appl Genet 125(3):495–502
Zhang X, Yang S, Zhou Y, He Z, Xia X (2006) Distribution of the Rht-B1b, Rht-D1b and Rht8 reduced height genes in autumn-sown Chinese wheats detected by molecular markers. Euphytica 152:109–116
Zhang XK, Xiao YG, Zhang Y, Xia XC, Dubcovsky J, He ZH (2008) Allelic variation at the vernalization genes Vrn-A1, Vrn-B1, Vrn-D1, and Vrn-B3 in Chinese wheat cultivars and their association with growth habit. Crop Sci 48:458–470
Zhao C, Cui F, Wang X, Shan S, Li X, Bao Y, Wang H (2012) Effects of 1BL/1RS translocation in wheat on agronomic performance and quality characteristics. Field Crops Res 127:79–84
Zhao FJ, Su YH, Dunham SJ, Rakszegi M, Bedo Z, McGrath SP, Shewry PH (2009) Variation in mineral micronutrient concentrations in grain of wheat lines of diverse origin. J Cereal Sci 49:290–295
Zhou Y, Chen Z, Cheng M, Chen J, Zhu T, Wang R, Liu Y, Qi P, Chen G, Jiang Q, Wei Y, Luo MC, Nevo E, Allaby RG, Liu D, Wang J, Dvorak J, Zheng Y (2018) Uncovering the dispersion history, adaptive evolution and selection of wheat in China. Plant Biotechnol J 16(1):280–291
Ziv O, Eyal Z (1977) Assessment of yield component losses caused in plants of spring wheat cultivars by selected isolates of septoria tritici. Phytopathology 68:791–796
Zurn JD, Newcomb M, Rouse MN, Jin Y, Chao S, Sthapit J, See DR, Wanyera R, Njau P, Bonman JM, Brueggeman R, Acevedo M (2014) High-density mapping of a resistance gene to Ug99 from the Iranian landrace PI 626573. Molecular breeding 34(3):1
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Nadeem, M.A. et al. (2021). Contribution of Landraces in Wheat Breeding. In: Zencirci, N., Baloch, F.S., Habyarimana, E., Chung, G. (eds) Wheat Landraces. Springer, Cham. https://doi.org/10.1007/978-3-030-77388-5_11
Download citation
DOI: https://doi.org/10.1007/978-3-030-77388-5_11
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-77387-8
Online ISBN: 978-3-030-77388-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)