Abstract
Rabies is a fatal encephalitis caused by the Rabies lyssavirus (RABV). The presence of minimal neuropathological changes observed in rabies indicates that neuronal dysfunction, rather than neuronal death contributes to the fatal outcome. The role of mitochondrial changes has been suggested as a possible mechanism for neuronal dysfunction in rabies. However, these findings are mostly based on studies that have employed experimental models and laboratory-adapted virus. Studies on brain tissues from naturally infected human and animal hosts are lacking. The current study investigated the role of mitochondrial changes in rabies by morphological, biochemical and proteomic analysis of RABV-infected human and canine brains. Morphological analysis showed minimal inflammation with preserved neuronal and disrupted mitochondrial structure in both human and canine brains. Proteomic analysis revealed involvement of mitochondrial processes (oxidative phosphorylation, cristae formation, homeostasis and transport), synaptic proteins and autophagic pathways, with over-expression of subunits of mitochondrial respiratory complexes. Consistent with these findings, human and canine brains displayed elevated activities of complexes I (p < 0.05), IV (p < 0.05) and V (p < 0.05). However, this did not result in elevated ATP production (p < 0.0001), probably due to lowered mitochondrial membrane potential as noted in RABV-infected cells in culture. These could lead to mitochondrial dysfunction and mitophagy as indicated by expression of FKBP8 (p < 0.05) and PINK1 (p < 0.001)/PARKIN (p > 0.05) and ensuing autophagy, as shown by the status of LCIII (p < 0.05), LAMP1 (p < 0.001) and pertinent ultrastructural markers. We propose that altered mitochondrial bioenergetics and cristae architecture probably induce mitophagy, leading to autophagy and consequent neuronal dysfunction in rabies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Rabies is a fatal viral encephalomyelitis caused by the rabies lyssavirus (RABV), transmitted most commonly through bites of rabid dogs. About 60,000 people die of rabies worldwide every year, mostly in Asia and Africa, with India contributing to more than 30% of the global burden. Globally, canine-mediated rabies is associated with the loss of 3.7 million disability-adjusted life years (DALYs), mostly due to premature deaths and 8.6 billion USD in economic losses each year [1, 2]. The risk of acquiring rabies following exposure to a rabid animal is about 15%, but varies (from 0.1 to 60%) depending on several factors [2]. The major stumbling block in the development of effective therapy for rabies is the lack of understanding of the basic mechanisms involved in pathogenesis. Histopathological changes seen post-mortem in the central nervous system of rabies-infected patients are relatively mild and fail to explain the catastrophic outcome [3]. This led to the hypothesis that RABV infection induces neuronal dysfunction rather than structural damage or neuronal death [4]. Several plausible mechanisms that may lead to neuronal dysfunction have been proposed, including defective neurotransmission, electrophysiological alterations, nitric oxide neurotoxicity, excitotoxicity and ion channel dysfunction; however, none has provided a reasonable elucidation for the postulated neuronal dysfunction [5]. Studies using advanced laboratory techniques have demonstrated degenerative structural changes in neuronal processes, which were not obvious on routine histopathological examination [6,7,8] and suggested the role of oxidative stress in such neurodegenerative changes [9]. Reactive oxygen species (ROS), which play a significant role in oxidative stress, are generated mainly in mitochondria. Oxidative stress and mitochondrial dysfunction reportedly contribute to neurodegenerative diseases [10] and other viral infections [11]. Researchers have demonstrated mitochondrial dysfunction and oxidative stress in RABV-infected neurons in culture [5, 12, 13]. They provided further evidence to suggest the specific role of RABV phosphoprotein in mitochondrial dysfunction [14, 15]. A previous proteomics study from our institute linked altered expression of mitochondrial proteins with human RABV encephalitis [16]. However, most of the published studies reporting altered redox dynamics and mitochondrial function have been performed in experimental models that employ Challenge Virus Standard (CVS), a laboratory-adapted fixed strain of RABV [6, 12,13,14]. It is well known that the pathogenic mechanisms triggered by CVS in experimental models may not reproduce the natural disease process caused by the wild-type strain (street virus) of RABV in humans and canines. However, most researchers utilize fixed laboratory-adapted strains of RABV such as CVS for laboratory experiments because they have predictable properties (such as incubation period, clinical and pathological effects), and are relatively inexpensive and easy to handle, unlike street viruses isolated from naturally infected animals, which have a variable incubation period and unpredictable effects [17]. Studies on brain tissues from RABV-infected humans and canines, which are the most ideal samples to study the pathogenesis of this neurotropic virus are lacking. To address these lacunae, we employed morphological, biochemical and proteomic approaches to delineate the role of mitochondrial dysfunction in RABV-infected human and canine brains.
Materials and Methods
All chemicals and reagents were of analytical grade. Bulk chemicals and solvents were obtained from Sisco Research Laboratories (Mumbai, Maharashtra, India) and Merck (Whitehouse Station, NJ, USA). Fine chemicals (JC-1 dye-Cat. no. T3168), cell culture media, fetal bovine serum and anti-β-actin antibody (RRID: AB_2305186) were obtained from Sigma-Aldrich (St. Louis, MO, USA). RNA extraction kit (Cat. no. 52906) was procured from Qiagen (Hilden, Germany) and quantitative PCR kit (Cat. no. 9111082) was obtained from Geno Sen’s® (New Delhi, India). Electrophoresis and Western blot reagents (Nitrocellulose membrane, Bio-Rad Cat. no.1620112; Advansta WesternBright ECL kit, Advansta Cat. no. K-12045-D20) were from Bio-Rad Laboratories (Hercules, CA, USA), Bangalore Genie (Bangalore, Karnataka, India), and Advansta (San Jose, CA, USA). ADP/ATP ratio assay kit (Cat. no. ab65313) and ATP synthase enzyme activity kit (Cat. no. ab109714) was procured from Abcam (Cambridge, MA, USA). Anti-Rabies Light Diagnostics DFA Reagent Antibody was procured from Merck, Millipore (RRID: AB_917374). All primary antibodies [Anti-FKBP8 (dilution 1: 1000)-RRID:AB_1848627; Anti-OXPHOS (dilution 1: 100)-RRID: AB_2629281; Anti-VDAC1 (dilution 1:1000)-RRID:AB_778788; Anti-β-actin (dilution 1: 1000)- RRID-AB_2305186); Anti-LC3 (dilution 1:500)- RRID: AB_881433; Anti-LAMP1 (dilution 1:1000)-RRID: AB_775978; Anti-TOMM20 (dilution 1.500)- RRID: AB_945896; Anti-PINK1(dilution 1.500)- RRID: AB_447627; Anti-PARKIN (dilution 1.500)- RRID: AB_1566559] used for immunoblotting were obtained from Abcam and Sigma-Aldrich. HRP-conjugated secondary antibodies (Goat Anti- rabbit IgG (dilution 1: 2000)-RRID: AB_916366; Goat Anti- mouse IgG (dilution 1: 2000)-RRID: AB_11211441) were obtained from Bangalore Genei. Chemicals for electron microscopy were procured from TAAB laboratories and equipment Ltd (Aldermaston, Berkshire, England). Tandem Mass Tag (TMT) isobaric labeling kit (Cat. no. 90111) was obtained from Thermofisher Scientific (Waltham, MA, USA). Proteomics grade Trypsin/Lys-C Mix (Cat. no. 38220090) was obtained from Promega (Madison, WI, USA).
Study Design
Human brains from laboratory confirmed cases of rabies viral encephalitis (n = 10) and age-matched controls (n = 10), and RABV-infected canine brains (n = 10) and age-matched controls (n = 10) collected post-mortem were selected. The effect size was estimated as 2.53 with the sample size of 10 in each group and the estimated power was 0.99. Different anatomical brain regions were subjected to histological and ultrastructural analysis. Brain mitochondria extracted from brain tissues of cases and controls were subjected to proteomic analysis, and the protein profiles were analyzed and compared to understand the molecular basis of mitochondrial dysfunction in rabies. This was followed by validation experiments (n = 3 trials/experiment). Canine brains [(cases and controls (n = 10, each)] were utilized for validation experiments (Fig. 1).
Study design. RABV-infected human and canine brain tissues and corresponding controls were selected for the study. Different anatomical brain regions were subjected to histological (HE haematoxylin–eosin, LFB Luxol Fast Blue), ultrastructural analysis (EM electron microscopy) and estimation of viral load (qPCR Quantitative PCR). Crude brain mitochondria from human rabies cases and controls were subjected to proteomic analysis, which included tryptic digestion, Tandem-mass tag (TMT) labelling and mass spectrometry (LC–MS/MS), followed by data analysis and validation experiments
Ethical Approval and Consent to Participate
The study was approved by the NIMHANS Institutional Ethics Committee (IEC) (Approval No. NIMH/DO/ETHICS SUB-COMMITTEE MEETING/2017 dated June 19, 2017). The human brain samples were collected following written, informed consent from the next of kin of the deceased. The Human Brain Tissue Repository (HBTR), Department of Neuropathology, NIMHANS, Bangalore, India has IEC clearance to collect, store and distribute tissues for research and publication [Approval no. NIMHANS/DO/98th IEC/2015, dated July 2, 2015].
The canine tissues were collected from dogs which died of suspected rabies, through the routine rabies surveillance programme of Mission Rabies and the Department of Animal Husbandry and Veterinary Services, Government of Goa, India. These samples were sent to the NIMHANS Rabies referral laboratory, Department of Neurovirology for routine diagnostic confirmation.
Tissue Samples
Human Brain Tissues
Post-mortem brain tissues of histologically confirmed RABV-infected cases (n = 10) and age and sex-matched controls (n = 10) were obtained from HBTR. These were collected post-mortem from patients who succumbed to rabies at NIMHANS, following written informed consent of close relatives of the deceased. Brain tissues from people who succumbed to road traffic accidents were utilized as controls. Brain tissues (both disease and controls) were collected at autopsy, one half of the brain was stored in 10% neutral buffered formalin and other half was frozen and stored at − 80 °C till use [18, 19]. In the current study, frontal cortex and cerebellum of RABV-infected brains and controls were utilized.
For controls, the brain areas chosen for the current study were anatomically farthest from the site of injury without distinct oedema or grossly apparent pathology. These tissues were subjected to histopathological assessment and the samples that maintained tissue integrity were utilized. Briefly, formalin-fixed tissues from different brains were evaluated for neuronal and glial alterations, anoxic changes, vascular alterations (if any), blinded to post-mortem interval (PMI), agonal state and gender, for myelin pallor.
The protocol of autopsy, tissue handling and other procedures were uniform for all the samples. Such tissues sourced from HBTR, have been earlier utilized as controls in human brain aging, Parkinson’s disease and post-mortem studies [20,21,22,23,24,25,26,27,28,29,30,31,32,33,34].
The sample size for both RABV-infected and control brains was considered based on tissue availability in HBTR. For both infected and control brains, samples from both genders with age 11–80 years were included, while cases with a clinical history of alcoholism, diabetes, neurological, psychiatric or other infectious diseases were excluded.
Canine Brain Tissues
Canine brains used in this study, collected at necropsy from dogs that died of suspected rabies were received from Mission Rabies, an international non-government organization engaged in canine rabies elimination in Goa state, India. From these, cases (n = 10) and controls (n = 10) were included following laboratory confirmation. Canines which died due to causes other than rabies (non-neurological or non-infectious causes such as critical spinal injury following road traffic accident etc.) and confirmed to be RABV-negative were included as controls. Care was taken to ensure that whole brains were harvested from canines within PMI of < 12 h. The current study focused on frontal cortex and cerebellum regions of both RABV-infected and control canine brains.
The brain was divided in the mid-sagittal plane into left and right hemispheres. One half was transported to NIMHANS in 10% neutral buffered formalin for morphological studies. The other half was sectioned coronally into 10 to 12 slices at 0.5 cm thickness and each slice was stored in numbered zip locked bags and transported to NIMHANS on dry ice. These samples were immediately stored at − 80 °C. In addition, a small portion of the tissue was stored in 3% Glutaraldehyde for ultra-structural studies by electron microscopy (EM).
Our previous study [28] noted that PMI up to 26 h did not affect the activity of mitochondrial complex I. Assessment of the activities of mitochondrial complexes I-IV and ATP, ADP, ADP/ATP and AMP/ATP ratios with increasing PMI (0–12 h) did not show any statistically significant effect of PMI on these mitochondrial parameters (Supplementary Fig. S1).
Laboratory Confirmation of RABV in Brain Samples
This was carried out by Fluorescent antibody technique (FAT), the gold standard recommended by WHO [35]. Smears of fresh brain tissues (human and canine) on glass slides were fixed in chilled acetone and incubated with anti-RABV monoclonal antibodies conjugated with fluorescein isothiocyanate (FITC), for 45 min (Merck, Millipore). The slides were washed and viewed under a fluorescence microscope (Primostar with LED fluorescent attachment, Carl Zeiss, Weimar, Germany).
Cell Culture Experiments
Details of cell lines and their maintenance, isolation and infection with virus strains, cell viability and determination of mitochondrial membrane potential are included in Supplementary Information.
RNA Extraction and Viral Quantification by qPCR
The detection and quantitation of RABV from both human and canine brains was carried out using commercially available Rabies quantitative real time PCR (qPCR) kit (Geno Sen’s®, New Delhi, India). Brain tissue (~ 300–500 mg) was minced and homogenized in isolation buffer [320 mM sucrose, 5 mM TES (Tris (hydroxymethyl) Methyl amino ethane Sulfonic acid, 1 mM EGTA, pH 7.2] and centrifuged (1000×g for 5 min at 4 °C). One aliquot of the supernatant (140 µl) was subjected to RNA extraction using QIAamp Viral RNA Mini Kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions and the other aliquot was used for mitochondrial isolation. The RNA was subjected to qPCR using RABV nucleoprotein (N) gene-specific primers and probe. Amplification, data acquisition, and analysis were carried out using an ABI 7500 real-time PCR Instrument and software (Foster city, CA, USA) and the viral load was expressed as viral copies/ml (Lowest detection limit = 3575 copies/ml; Highest = 35,750,000 copies/ml).
Histopathology
Different anatomical regions of fresh human and canine brains were fixed in 10% neutral buffered formalin, sliced coronally and processed for paraffin embedding and sectioning. Five-micron thick sections were subjected to routine histological examination including Haematoxylin–Eosin (HE), Luxol Fast Blue (LFB) and Nissl staining as described [36].
Electron Microscopy
RABV-infected canine brain (n = 3) and human brain (n = 1) stored in 3% glutaraldehyde were processed for electron microscopy. Ultrathin sections (70 nm) were then collected on copper grids, stained with 1% Uranyl acetate for 1–2 h followed by Lead citrate for 5–7 min as described [37, 38], viewed under Transmission Electron Microscope (FEI, TECNAI G2 Spirit BioTwin, Netherlands). Selected images were captured on MegaView-III CCD camera.
Mitochondrial Isolation and Enzyme Assays
Brain mitochondria were isolated from fresh frozen human and canine brain tissues as described [39]. The tissue extract prepared as indicated in the section on viral load analysis was centrifuged (1000×g for 5 min at 4 °C). The supernatant was collected and the pellet was resuspended in isolation buffer followed by a second round of homogenization and centrifugation as described above. The supernatants from both steps were pooled and centrifuged (8500×g for 10 min at 4 °C) and the crude mitochondrial fraction in the pellet was resuspended in isolation buffer (~ 150 µl), overlaid on 6% (w/v) Ficoll solution, and centrifuged at 75,000×g for 30 min at 4 °C. The pellet was resuspended in reconstitution buffer (250 mM sucrose, 10 mM TES, pH 7.2) and stored at − 80 °C.
Mitochondria were subjected to enzyme assays of mitochondrial complex I (CI), complex II (CII), complex III (CIII) and complex IV (CIV) as described and normalized to the respective Citrate Synthase activity [39]. Complex V was assayed using a commercial kit (Abcam; Cat. no. ab109714) according to the manufacturer’s instructions.
Estimation of ADP/ATP Ratio
Estimation of ADP/ATP ratio was carried out in freshly prepared total brain extracts based on bioluminescence, using a commercial kit (Abcam; Cat. no. ab65313) according to the manufacturer’s instructions. AMP/ATP ratio was determined as described [40].
Mitochondrial Proteomics
Sample Preparation
Crude mitochondrial preparations (enriched with mitochondria, synaptosomes and microsomes) were isolated from RABV-infected human brains (frontal cortex, n = 10 and cerebellum, n = 10) and controls (frontal cortex, n = 10 and cerebellum, n = 10). The mitochondria were sonicated (1X10 s at 37% amplitude) in 1X PBS containing 0.05% SDS. Following protein estimation [41], 20 μg protein from each brain sample within a group was pooled. At each step, the samples were normalized for protein concentration by protein estimation and 10% SDS PAGE.
Tryptic Digestion, Tandem Mass Tag (TMT) Labelling and Strong Cation Exchange (SCX) Chromatography
Equal protein (200 μg) from each group was reduced and alkylated using 10 mM of dithiothreitol and iodoacetamide, respectively. Alkylated proteins were digested with sequencing grade modified trypsin (Promega) at 37 °C for 12 h. Tryptic digests were dried and labelled with TMT reagent (Thermo Fisher Scientific, Waltham, MA, USA) as per the manufacturer’s protocol. The labelling efficiency for each sample was checked by analysing 2 μg peptides on the mass spectrometer followed by pooling of equal amount of labelled peptides from control and rabies samples. The pooled sample was dried and fractionated using Strong Cation Exchange Chromatography (SCX). Each fraction was then desalted using C18 stage-tips and subjected to liquid chromatography tandem mass spectrometry (LC–MS/MS) [42].
LC–MS/MS
Each fraction was analysed in triplicate on Thermo Orbitrap Fusion tribrid mass spectrometer interfaced with Easy-nLC 1000 system (Thermo Scientific, Bremen, Germany) [43]. The peptides were loaded on 2 cm Acclaim PepMapC18 pre-column (Thermo Fisher Scientific) and further resolved on 25 cm long C18 column (3 μm, 100 Å) using a linear gradient of 5% to 30% of solvent B (0.1% formic acid in 95% Acetonitrile) over 110 min and flow rate of 300 nl/min.
Data Analysis
Raw data from the 6 fractions (in triplicates) were searched against the reference human protein database in NCBI (Refseq version 75) using MASCOT and SEQUEST search engines (version 2.4.1) through Proteome Discoverer 2.1 suite (Thermo Fisher Scientific, Bremen, Germany).
For data-sets, search parameters included carbamidylation of Cys and TMT at the peptide N-terminus as static modifications and oxidation of Met and N-terminal protein acetylation as dynamic modifications. Trypsin was considered as the proteolytic enzyme and one missed cleavage was allowed during the search. Percolator node provided in the Proteome Discoverer software was used to compute false discovery rate (FDR) and 1% peptide spectrum matches (PSM). The abundance value of the TMT-reporter ions obtained by MS analysis was used to identify the fold change of individual proteins quantified in this study. Proteins with relative fold change of ≥ 1.30 was considered as up-regulated and fold change of ≤ 0.66 as down-regulated, compared to the respective controls, as described earlier [44, 45]. Gene ontology (GO) analysis was done on Cytoscape (Version 3.6) and ClueGO analysis was used for prediction of protein–protein interactions using significance or p-value (< 0.05) [46]. Protein identification and analysis were done using MitoCarta (Version 2.0) [47, 48] for mitochondrial proteins, SynDB [49] for synaptic proteins and HADb (http://autophagy.lu/index.html) for autophagy related proteins.
SDS-PAGE and Western Blotting
Brain tissues were minced, homogenized, sonicated in 1X PBS with 1X protease inhibitor cocktail (Sigma-Aldrich) and centrifuged (14,000×g for 10 min at 4 °C). Total protein in the supernatant was subjected to protein estimation (Bradford 1976), followed by SDS-PAGE and western blot [50]. Alternatively, crude mitochondria, were solubilized in 2X Laemmli’s buffer and subjected to SDS-PAGE. Western blot was carried out with primary antibody [Anti-FKBP8 (dilution 1:1000); Anti-OXPHOS (1:100); Anti-VDAC1 (1:1000); Anti-LC3 (1:500); Anti-LAMP1 (1:1000); Anti-TOMM20 (1:500); Anti-PINK1 (1:500); Anti-PARKIN (1:500)], followed by HRP-conjugated secondary antibody [Goat Anti-rabbit IgG (1:2000); Goat Anti-mouse IgG (1:2000)]. The western signal was visualised using Advansta WesternBright ECL chemiluminescence kit (K-12045-D20). β-actin and Tom20 served as loading controls. The western signal was recorded in a gel documentation system (Bio-Rad Laboratories Inc.), quantified using ImageJ software and normalized to the respective anti-β-actin signal (dilution 1:1000) signal [51].
Statistical Analysis
Quantitative data shown as bar graphs are represented as cumulative data from n ≥ 3 independent experiments (excluding proteomics, which was in technical triplicates) and expressed as mean ± SD. For multiple comparisons, ANOVA or student’s t test using GraphPad Prism, Version 8.00 for Windows (GraphPad Software, La Jolla, California USA, www.graphpad.com) was used and p < 0.05 was considered significant. Raw data from the MS experiment (list of proteins and fold-changes across the replicates compared with controls) was uploaded on Perseus software for calculation of p value and generation of heat maps [52] and proteins identified with statistical significance of p < 0.05, were listed. Data obtained in the current study were not assessed for normality prior to statistical analysis. No test for outliers was conducted. No blinding was performed during the study, except for the histopathological assessment of brain tissues and MS experiment.
Results
Characterization of the RABV-Infected Human and Canine Brains
Two anatomical regions each from RABV-infected post-mortem human and canine brain samples (n = 10), and age-matched non-infected controls (n = 10) were selected for the study (Tables 1, 2). RABV infection in the brains was confirmed by Fluorescent Antibody technique (FAT) (Fig. 2A–D). RT-PCR data showed high viral load in all the anatomical regions in both RABV-infected human and canine brains (Table 3).
Morphological analysis of RABV-infected human and canine brains. A–D Fluorescent antibody test (FAT) to confirm the presence of RABV infection. Smears of fresh brain tissues (human and canine; n = 10, each) were fixed and stained with fluorescently labelled anti-RABV monoclonal antibodies and viewed under a fluorescence microscope. Apple green fluorescence indicates the presence of RABV antigen. A and C are controls (human and canine), while B and D are from RABV-infected human frontal cortex and canine frontal cortex respectively (A–C, 40×). E–L Neuropathological changes in RABV-infected human brains. Fresh human brain regions (n = 5) were fixed, embedded in paraffin and sectioned, followed by routine histological examination. Purkinje cells of human cerebellum (E) and pyramidal neurons of human frontal cortex (F) with characteristic eosinophilic intracytoplasmic Negri bodies (dotted circles and arrows) are shown. Neuronophagia and perivascular inflammation in frontal cortex are shown in G and H (dotted circle in both). Diffuse microglial nodules and inflammation were noted in cerebellum (I, dotted circle). The neuronal morphology of infected neurons in cerebellum was well-preserved without evidence of neuronal degeneration along with prominent nucleoli (J, dotted circles and arrows). Frontal cortex (K, star) and cerebellum (L, star) showed well preserved myelin as highlighted by Luxol Fast Blue (LFB) staining [E–J: H&E, K, L: LFB; In all the images, magnification = scalebar]. Similarly, canine RABV-infected brains (n = 5) showed characteristic eosinophilic intracytoplasmic Negri bodies in frontal cortex (M, dotted circles and arrows) and cerebellum (N, dotted circles and arrows) along with neuronophagia (O, dotted circle) and meningeal inflammation (P, dotted circle). The neuronal morphology of infected neurons was well-preserved along with prominent nucleoli (Q, R, dotted circles and arrows). Representative image of canine brain showing chromatolysis with vacuolation (S, dotted circles and arrows). Myelin was well preserved in all the tested regions (T, star). [M–R: H&E, S: Nissl, T: LFB; In all the images, magnification = scalebar]
Histopathological Changes in the Brain Following RABV Infection
RABV-infected human and canine brains (n = 5 each) displayed the presence of Negri bodies, a pathological hallmark of RABV infection, in different anatomical regions (Fig. 2E, F and M, N). The brains also revealed varying degree of perivascular and meningeal inflammation, microglial response and neuronophagia (Fig. 2G–I, O–P). Inflammation was predominant in frontal cortex (data not shown) in canine brains, whereas only one human brain showed prominent inflammation and neuronophagia. The density of Negri bodies varied across the anatomical regions. Round to oval discrete eosinophilic Negri bodies were identified in the large reticular neurons and Purkinje cells of the cerebellum (Fig. 2E). While few infected neurons exhibited chromatolysis and vacuolation (Fig. 2S), the overall neuronal morphology in all the regions tested was unchanged (Fig. 2J, Q, R) with intact myelin (Fig. 2K, L, T). Taken together, histopathology in RABV-infected human brains was similar to canine. However, the inflammatory responses in human brains were relatively lesser than canine (Relative quantitation not shown).
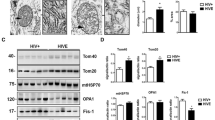
Ultrastructural analysis by TEM showed the presence of RABV as electron dense intracellular Negri bodies predominantly in canine brains (Fig. 3A–C) and as electron dense intra-cytoplasmic particles in human brains (Fig. 3J, K). Organellar ultrastructure in RABV-infected human brain (Fig. 3J) and canine brains (Fig. 3G–I) showed swollen mitochondria with altered/disintegrated cristae and dilated endoplasmic reticulum compared to controls. Compared to human, canine samples showed a greater number of abnormal mitochondria with disintegrated cristae (Relative quantification not shown). These structural changes indicated mitochondrial dysfunction, although such structural alterations were noted in many but not all the fields scanned. This prompted us to assess the proteomic changes in brain mitochondria. Considering the pathological implications for humans and the potential technical difficulty in analysis with similar robustness in canine mitochondria, proteomic analysis was carried out only in human brain samples.
Ultrastructural analysis of RABV-infected canine and human brains. Brain samples (canine brain, n = 3 and human brain, n = 1) stored in Glutaraldehyde were post-fixed with Osmium tetroxide, dehydrated, cleared and infiltrated with Araldite and Propylene oxide. Ultrathin sections on copper grids were stained with Uranyl acetate and Lead citrate and viewed under TEM. Canine: A–I, Human: J, K. A TEM image showing the presence of intracytoplasmic Negri bodies in canine brains as dense grey regions (arrows). B and C RABV particles inside Negri bodies in canine brains (arrow). The preserved structures of mitochondria (D, arrow), Golgi apparatus (E, arrow) and endoplasmic reticulum (F, arrow) in uninfected canine brain (control) are shown. RABV infected canine brains showed abnormal swollen mitochondria with vacuolation (G, arrow) dilated or disintegrated cristae architecture (H, arrow) and dilated rough endoplasmic reticulum (I, arrows). RABV particles in human brains are shown in J (white arrows) and K (white arrows). Human brains showed the presence of mitochondria with preserved (J, white arrow head) and abnormal structure (J, black filled arrows). (Scale bar corresponds to 1 μm in A and F, 0.5 μm in B–E and H–J and 0.2 μm in G and K)
Mitochondrial Proteomic Analysis of RABV-Infected Human Brains
Crude mitochondrial preparations (enriched for mitochondria, synaptosomes and microsomes) prepared from RABV-infected human brains and controls (pooled into Group 1 to 4) were subjected to proteomic analysis (Fig. 4A).
Total proteomics of RABV-infected human brain mitochondria vs. controls. A Workflow of the proteomics experiment. Total mitochondria were isolated from individual human control (C) and RABV-infected human brains (anatomical regions and number of samples are indicated) and pooled into 4 groups (1–4). Protein extracts from 4 groups were subjected to tryptic digestion, TMT labelling, strong cation exchange (SCX) fractionation, mass spectrometry (MS) and data analysis. The SDS PAGE profile of all 10 groups is shown in B. Heat map in C shows the number of upregulated (≥ 1.3 fold) and downregulated (≤ 0.66 fold) proteins in RABV-infected human brains compared to controls. The up-regulated and down-regulated proteins are indicated by red and green colours respectively (unchanged = white). D Number of differentially expressed proteins in frontal cortex and cerebellum of human brains. The number of up-regulated and down-regulated proteins common/unique to frontal cortex and cerebellum is shown in E and F respectively. Functional classification of all the up-and down-regulated proteins in RABV-infected brains highlighting the three major groups: mitochondrial proteins, synaptic compartment and autophagic pathway is shown in G
Analysis of the MS data identified 3583 proteins (1% FDR; p < 0.05) across all the groups. Among these, differentially regulated proteins included 664 up-regulated (439 in frontal cortex, 416 in cerebellum and 191 common proteins) and 245 down-regulated proteins (200 in frontal cortex, 77 in cerebellum and 32 common proteins) (Fig. 4C–F). Gene Ontology (GO) analysis and functional grouping revealed that the differentially regulated proteins were predominantly mitochondrial (n = 163 proteins), synaptic (n = 187 proteins) and autophagic (n = 83 proteins) (Fig. 4G, Table 4 and Supplementary Tables S1 and S2). Other pathways included cytoskeletal and apoptotic proteins (Supplementary Table S3).
Considering the EM data (Fig. 3), we focused on differentially expressed mitochondrial proteins, which were mainly associated with oxidative phosphorylation (OXPHOS) complexes, cristae formation, mitochondrial homeostasis and transport. Networking analysis revealed significant interlinking and strong functional interaction of these pathways (Fig. 5 and Table 4).
Among the OXPHOS complexes, 16 of CI subunits, 8 of CIII, 5 of CIV and 20 of CV were upregulated (Table 4). Western blot confirmed slight over-expression of the CI subunit NDUFB8 and the CIII subunit UQCRC2 (Fig. 6A). To assess the effect of these changes, we carried out enzyme assays of complexes CI-CV in RABV-infected human brain mitochondria vs. controls. Activities of CI, CIV and CV were significantly higher compared to control in all four regions, while the activity of CIII showed an increasing but statistically insignificant trend in frontal cortex (Fig. 6B–F). Interestingly, correlation between viral load (copy number) and activities of CI-CV was not statistically significant either in frontal cortex or cerebellum (Supplementary Table S4). Unlike OXPHOS enzymes, expression of TCA cycle enzymes such as Malate Dehydrogenase (MDH) was unchanged and the same was validated by enzyme assay (Supplementary Fig. S2).
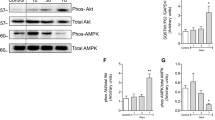
Validation of the proteomics data from RABV-infected human brains. The MS data from the mitochondrial proteomics experiment obtained using human RABV brain samples were validated by western blot and enzyme assays. A Brain mitochondria pooled from control (n = 10) and RABV-infected brains (n = 10) were subjected to western blot using mitochondrial antibody cocktail containing antibodies against one subunit each of CI (NDUFB8), CII (SDHB), CIII (UQCRC2) and CV (ATP5A). Tom20 served as loading control. Number on each band corresponds to its intensity (quantification indicated in arbitrary OD units normalized to the respective Tom20 signal) following densitometric analysis in a gel documental system (Tecan GmBH). B–F Enzyme assay of mitochondrial complexes, CI (B), CII (C), CIII (D), CIV (E) and CV (F), normalized to the respective citrate synthase (CS) activity in frontal cortex (n = 10) and cerebellum (n = 10) of both human control and RABV-infected human brains. G–K Enzyme assay of mitochondrial complexes, CI (G), CII (H), CIII (I), CIV (J) and CV (K), normalized to the respective citrate synthase (CS) activity in frontal cortex (n = 10) and cerebellum (n = 10) of canine control and RABV-infected human brains. Data represented as mean ± SD (n = 3 trials per experiment for all assays; *p < 0.05, **p < 0.01 and ***p < 0.001 compared to the respective controls; ns not significant)
To compare these data in canine brains, OXPHOS assays were carried out in canine brain mitochondria. CI and CIV activities were elevated in frontal cortex and cerebellum compared to control (Fig. 6G–K). On the other hand, CII activity was elevated only in frontal cortex (Fig. 6H), while CV activity was relatively unchanged in both frontal cortex and cerebellum (Fig. 6K). Similar to the human data, correlation between viral load of RABV-infected canine brains and complex activities was not statistically significant (Supplementary Table S4).
Analysis of the Mechanisms Underlying Mitochondrial Dysfunction in Rabies Brains
First, we assessed whether elevated OXPHOS activities influenced ATP production. While the total ATP and ADP in frontal cortex and cerebellum of human brains were significantly lower than control, the ADP/ATP ratio was not significantly different between the two (Fig. 7A–C and Supplementary Fig. S3A-C). AMP/ATP ratio revealed increasing but statistically insignificant trend in Rabies samples (vs. control) (Fig. 7D and Supplementary Fig. S3D). Although similar trend was noted in canine brains, the decrease in ATP and ADP content compared to controls was not as drastic as noted in the human brains (Fig. 7E–H and Supplementary Fig. S3E-H).
Assessment of ADP/ATP ratio in human and canine brains. The ADP and ATP content in the frontal cortex of human and canine control and RABV-infected brains were quantitated, followed by calculation of ADP/ATP and AMP/ATP ratios. A–D correspond to the data from human frontal cortex control (n = 10) and infected samples (n = 10), while E–H correspond to the data from canine frontal cortex control (n = 10) and infected samples (n = 10). Data represented as mean ± SD (n = 3 trials per experiment for all assays; ***p < 0.001, ****p < 0.0001 compared to the respective controls; ns not significant)
Since ATP levels were lower despite elevated OXPHOS activities, we surmised that altered membrane potential could contribute to mitochondrial dysfunction. Interestingly, VDAC1, a component of the mitochondrial membrane permeability transition pore was over expressed in human RABV brains (Table 4 and Fig. 8A, B). To assess the role of altered membrane potential, we assessed cultured Neuro2a cells (neuronal cell line) and BHK21 cells (non-neuronal cell line) infected by CVS-11 and SV strains of RABV for different time points (0–72 h) (Fig. 8C, D). RABV infection at 0.3 MOI caused time-dependent loss of cell viability (Fig. 8E, F) and lowered membrane potential (Fig. 8G, H). Taken together, we surmised that lowered mitochondrial membrane potential could contribute partly to lowered ATP production in RABV-infected brains, leading to mitochondrial dysfunction. Another potential mechanism is altered cristae structure, as indicated by EM (Fig. 3G, H and J) and over-expression of the cristae protein MICOS13 and transmembrane proteins TMEM11and TMEM65 (Table 4).
Assessment of the role of mitochondrial membrane potential and mitophagy following RABV infection. RABV-infected human brain tissues were assessed for components of mitochondrial permeability transition pore (mPTP), followed by validation of the role of altered mitochondrial membrane potential in cell models of RABV infection. Total mitochondrial extracts from cerebellum of control (n = 3) and RABV-infected (n = 3) human brains were subjected to anti-VDAC1 western blot with Tom20 as loading control (A) followed by quantitation of the western signal (B). Data represented as mean ± SD (n = 3 trials per experiment for all assays; **p < 0.01, compared to control). Neuronal (N2a) and non-neuronal cell lines (BHK-21) were infected with two strains of RABV, i.e., CVS-11 and SV as confirmed by Fluorescent Antibody Test (FAT). Representative images of FAT following CVS-11 and SV infection are indicated in C and D respectively. RABV-infection dependent loss of cell viability in N2a and BHK-21 cell lines at different hours post infection (hpi) vs. untreated control, as assessed by MTT assay is shown in E and F respectively. Data represented as mean ± SD (n = 3 trials per experiment for all assays; **p < 0.01, ***p < 0.001, ****p < 0.0001, ns not significant, compared to respective control). For analysis of mitochondrial membrane potential (ΔΨ), fluorescence measurement following JC-1 treatment of RABV-infected N2a and BHK-21 cell lines at different time points post infection (hpi) was carried out. Representative images of control and RABV-infected BHK-21 cells at different time points post-infection are shown in G. The ratio of red: green fluorescence (Δ JC-1 fluorescence) at different time points post infection in both cell lines is depicted in H. To understand the role of mitophagy, total mitochondrial extracts from cerebellum of control (n = 3) and RABV-infected (n = 3) human brains were subjected to anti-FKBP8 western blot with Tom20 as loading control (I) followed by quantitation of the western signal (J). Data in all graphs are represented as mean ± SD (n ≥ 3 trials per experiment for all assays; *p < 0.05, **p < 0.01, ****p < 0.0001, compared to respective control). Control experiments carried out to assess mitochondrial depolarization and hyperpolarization in N2a and BHK-21 cell lines are depicted in Supplementary Fig. S4A-B
Western blot data validated the over-expression of Peptidyl-prolyl cis–trans isomerase (FKBP8), a mitophagy protein (1.5-fold vs. control; Supplementary Table S1), indicating that RABV-mediated mitochondrial dysfunction probably culminates in mitophagy in the brain (Fig. 8I, J). Since mitophagy could ultimately lead to autophagy, we assessed the morphological and biochemical markers of autophagy. Both RABV-infected human and canine brain tissues showed evidence of autophagy in EM. Structures representing initiation and elongation of phagophore or isolation membrane and typical double membraned autophagosome like vesicles containing organelles and debris were noted in RABV-infected canine brain (Fig. 9A–C), while autolysosome was noted along with autophagosome in RABV infected human brain (Fig. 9D–F). To further investigate whether autophagy is caused by RABV infection, protein expression of LC3, an autophagy marker that binds to autophagosome membrane, was examined. Conversion of LC3I to LC3II was significantly higher in RABV-infected human brains compared vs. controls (Fig. 9G, H). LAMP1 was also overexpressed in RABV-infected human brains indicating the formation of autolysosome (Fig. 9G and I). Western blot also revealed elevated PINK1 (protein linked to mitophagy) expression in RABV-infected human brains and that mitophagy is independent of PARKIN (protein linked with PINK1 and mitophagy) (Fig. 9J–L) since PARKIN expression remained unchanged between RABV-infected and control brains.
Ultrastructural and biochemical evidences of autophagy in RABV-infected human and canine brains. Canine and human brain samples stored in Glutaraldehyde were processed for electron microscopy as described in Fig. 3. Ultrathin sections on copper grids were stained with Uranyl acetate and Lead citrate and viewed under TEM. Formation and elongation of phagophore (A and B, dotted circle) and double membraned autophagosome like vesicles (C, dotted circle) observed in RABV infected canine brains. Presence of autophagosome (D and E, dotted circles) and autolysosome like vesicle (F, dotted circle) in RABV infected human brains. Western blot analysis revealed that LC3 I to LC3 II conversion, a marker of autophagy was significantly higher in RABV infected human brains (G, H) vs. control. LAMP1, a marker of autolysosome formation showed significant increase in RABV infected brains (G, I) compared to control. PINK1 showed significantly increased expression in RABV infected brains compared to control (J, K), while the expression of PARKIN remained unchanged (L, K), (n = 3 per experiment, *p < 0.05, **p < 0.01, ns non significant, C control, R rabies)
Discussion
Most studies on RABV employ CVS, which does not mimic natural infection by wild strains and could be both strain and cell-specific [53]. Lack of mechanistic studies on natural RABV infection in humans and canines is a critical lacuna in understanding rabies pathogenesis. Hence, assessment of natural RABV infection from site of entry to the brain is physiologically pertinent to human pathology.
From the bite site, RABV is transported to the neurons via receptors or Neuronal cell adhesion molecules (NCAMs) [54]. We noted upregulation of NCAM1 (1.3-fold vs. control) (Supplementary Table S2), in the cerebellum, a predominant location for RABV antigen [55]. The virus is then internalized by endocytosis [56] for subsequent transport, which depends on actin and microtubules [57, 58]. In RABV-infected samples, Actin binding LAM protein 1 (ABLIM1), was upregulated (1.32-fold in cerebellum vs. control) (Supplementary Table S2). Most of the related proteins including BIN1 (frontal cortex-0.59 fold, cerebellum-0.66 fold, vs. control) [59], WASP-WAVE family of proteins including WASF1 (FC- 0.59 vs. control) and dynamin (DNM1- 0.49 fold vs. control) [60] were downregulated in RABV-infected human brains. We speculate that, this trend has got more to do with recruitment and promoting polymerization rather than protein expression itself. Proteins involved in endocytosis were relatively unchanged in RABV-infected brains (Data not shown).
Most viruses shift from actin to microtubule mediated retrograde transport for long range movement [58]. Microtubule associated proteins 2 and T (MAP2 and MAPT) [61] were down-regulated in RABV-infected human brains, although this did not correspond to loss of neuronal architecture (Fig. 2J, Q, R) [8]. RABV utilizes the dyenins and kinesins for transport [58], although their expression was unchanged in RABV- infected brains (Data not shown).
We hypothesize that RABV induces mitochondrial dysfunction by structural or functional mechanisms. Although the number of differentially regulated synaptic proteins was significantly higher, we focused on mitochondrial proteins, considering that previous studies from our group [16] and others [12,13,14,15] indicated mitochondrial damage following RABV infection. Further, mitochondrial dysfunction could in turn contribute to synaptic and neuronal dysfunction. RABV infection altered mitochondrial structure (Fig. 3G, H), which could probably be due to altered expression of cristae proteins, which could alter the organization of cristae, outer membrane (OM), inner membrane (IM) proteins and OXPHOS complexes [62]. Expression of mitochondrial contact site cristae junction organizing system (MICOS) proteins [63] such as DNAJC11 [63, 64], TMEM11, an OM chaperone and other OM proteins were upregulated (Table 4), probably contributing to altered organization of cristae and respiratory complexes [62]. MIC13/C19orf70, a cristae protein [63] governed by MICOS proteins [63, 65,66,67], was upregulated in RABV-infected human brain (Table 4). Apart from MICOS, subunits of F0-F1 ATP synthase, which contribute to cristae organization [68,69,70] were upregulated (Table 4).
Altered OXPHOS complexes contribute to mitochondrial dysfunction [71, 72]. Consistent with Alandijany et al. [12], CI and CIV activities were elevated and CIII activity was relatively unchanged both in canine and human brains. While CII activity was exclusively increased in canine, CV was elevated in human brains (Fig. 6B–K). Kammouni et al. [14] showed that RABV phosphoprotein could elevate CI activity, which could be over and above the upregulation of CI subunits. Unlike CI and CIV, overexpression of the CII subunit SDHB as indicated by western blot (Fig. 6A) and not in the MS data (Table 4; The expression of other CII subunits remained unchanged), did not increase the activity of CII indicating that the overexpression of this subunit alone might not be sufficient to significantly alter CII organization and enzyme activity. Similarly, overexpression of CIII subunits in human cerebellum (8 subunits) and frontal cortex (1 subunit) did not result in elevated activity (Fig. 6D). It is possible that elevated CIII activity potentially requires (i) all the three catalytic subunits to be over expressed, while CytB was the only catalytic subunit over-expressed in RABV-infected human brain vs. controls (Table 4) and (ii) increased assembly of active complex which might not have been complete. This requires further experimentation either in RABV-infected mouse or cell culture models for a meaningful inference. Interestingly, elevated OXPHOS activities did not correlate with viral load (Supplementary Table S4) indicating that the effects of viral load could be in the linear phase of disease progression. Since post-mortem samples represent end-stage disease, this could be effectively assessed only in animal models. Increased OXPHOS activities did not lead to increased ATP production (Fig. 7 and Supplementary Fig. S3), which could probably be due to lowered membrane potential as demonstrated in cell culture experiments (Fig. 8G, H). MS data revealed that protein components of the mitochondrial permeability transition pore (mPTP) such as voltage-dependent anion-selective channel proteins 1 and 2 (VDAC 1 and 2) were over-expressed (Table 4 and Fig. 8A, B). Whether this could alter mPTP opening and membrane potential contributing to mitochondrial dysfunction is only speculative at this stage and requires further experimentation.
Previous studies have demonstrated autophagy in models of RABV infection [53, 73] and others including Newcastle Disease virus and Dengue virus 2 and 3 [74, 75]. Such studies speculated that autophagy could be neuroprotective or lead to increased viral replication and exacerbated infection. Interestingly these studies did not highlight the mitophagy and autophagy markers, which make it a novel finding in human and canine brains (Table 4 and Figs. 8I, J, 9). Ultrastructural images revealed different stages of autophagosome formation both in human and canine brains (Fig. 9A–F). Biochemical evidences confirmed activation of autophagy, based on western blot of different markers including LC3, LAMP1, PINK1 and PARKIN (Fig. 9G–L).
Human and canine rabies share similar pathological and clinical features [76,77,78]. Both showed high viral load in all the regions (Table 3). While Negri bodies were distinct in canine, they were not evident in human brains (Fig. 3). The histopathology in human and canine brains was similar, although the inflammatory response in human was lesser than in canine (Fig. 2). The pathological and biochemical profiles including OXPHOS activities are comparable between RABV-infected human and canine brains. Further experimentation is required to delineate the molecular differences underlying rabies pathology between the two species.
For most part of the study, we focused only on the natural RABV infected human and canine brains, which is novel and pathologically pertinent. Although cell models used in earlier studies provide mechanistic information [12,13,14,15], they do not represent the pathological milieu relevant to humans. Secondly, the path of viral transport from the site of bite to the brain cannot be recapitulated in vitro. Thirdly, cells in culture do not have the tissue architecture applicable to the adult human brain. However, for validating the proteomic data, a cell model is ideal. We employed neuronal cells to assess RABV infection relevant to brain and a non-neuronal cell line to determine if the changes are neuron-specific (Fig. 8G, H).
Although the current study is one of the first studies done on RABV-infected human and canine brains, it has certain limitations. Since post mortem brain samples represent terminal stage of the disease early events and disease progression cannot be investigated. Such studies require an animal model of RABV infection which needs to be pursued.
Conclusion
This is one of the first comprehensive studies on human and canine brains that investigated the mechanisms underlying RABV pathogenesis, at the molecular level. Our data based on morphological, proteomic and cell culture studies revealed disrupted mitochondrial ultrastructure, altered proteome and altered enzyme activity of respiratory complexes in RABV infected brains. This along with impaired mitochondrial membrane potential, probably leads to mitochondrial dysfunction. Our data linked mitochondrial dysfunction with mitophagy and ensuing autophagy which could ultimately lead to neuronal dysfunction in rabies. These data not only delineate the events in rabies pathobiology but may help in evolving novel therapeutic strategies.
Data Availability
The MS data presented in the current manuscript have been deposited in the ProteomeXchange Consortium via the PRIDE partner repository [79] with the dataset identifier PXD020789 (http://www.ebi.ac.uk/pride).
Abbreviations
- CI:
-
Complex I
- CII:
-
Complex II
- CIII:
-
Complex III
- CIV:
-
Complex IV
- CV:
-
Complex V
- CVS:
-
Challenge Virus Standard
- EM:
-
Electron microscopy
- HE:
-
Haematoxylin and eosin
- MS:
-
Mass spectrometry
- RABV:
-
Rabies lyssavirus
- ROS:
-
Reactive oxygen species
- RRID:
-
Research resource identifier
- SV:
-
Street virus
- TMT:
-
Tandem mass tag
References
Hampson K, Coudeville L, Lembo T et al (2015) Estimating the global burden of endemic canine rabies. PLoS Negl Trop Dis 9:e0003709. https://doi.org/10.1371/Journal.PNTD.0003709
WHO expert consultation on rabies (2018): third report. World Health Organization
Murphy FA (1977) Rabies pathogenesis brief review. Arch Virol 54:279–297
Tsiang H (1982) Neuronal function impairment in rabies-infected rat brain. J Gen Virol 61:277–281. https://doi.org/10.1099/0022-1317-61-2-277
Rossiter JP, Jackson AC (2013) Chapter 9—pathology. In: Jackson AC (ed) Rabies, 3rd edn. Academic Press, Boston, pp 351–386
Scott CA, Rossiter JP, Andrew RD, Jackson AC (2008) Structural abnormalities in neurons are sufficient to explain the clinical disease and fatal outcome of experimental rabies in yellow fluorescent protein-expressing transgenic mice. J Virol 82:513–521. https://doi.org/10.1128/jvi.01677-07
Song Y, Hou J, Qiao B et al (2013) Street rabies virus causes dendritic injury and F-actin depolymerization in the hippocampus. J Gen Virol 94:276–283. https://doi.org/10.1099/vir.0.047480-0
Li X-Q, Sarmento L, Fu ZF (2005) Degeneration of neuronal processes after infection with pathogenic, but not attenuated, rabies viruses. J Virol 79:10063–10068. https://doi.org/10.1128/jvi.79.15.10063-10068.2005
Jackson AC, Kammouni W, Zherebitskaya E, Fernyhough P (2010) Role of oxidative stress in rabies virus infection of adult mouse dorsal root ganglion neurons. J Virol 84:4697–4705. https://doi.org/10.1128/jvi.02654-09
Lin MT, Beal MF (2006) Mitochondrial dysfunction and oxidative stress in neurodegenerative diseases. Nature 443:787–795. https://doi.org/10.1038/nature05292
Claus C, Liebert UG (2014) A renewed focus on the interplay between viruses and mitochondrial metabolism. Arch Virol 159:1267–1277
Alandijany T, Kammouni W, Roy Chowdhury SK et al (2013) Mitochondrial dysfunction in rabies virus infection of neurons. J Neurovirol 19:537–549. https://doi.org/10.1007/s13365-013-0214-6
Jackson AC, Kammouni W, Fernyhough P (2011) Role of oxidative stress in rabies virus infection. Adv Virus Res 79:127–138. https://doi.org/10.1016/B978-0-12-387040-7.00008-1
Kammouni W, Wood H, Jackson AC (2017) Lyssavirus phosphoproteins increase mitochondrial complex I activity and levels of reactive oxygen species. J Neurovirol 23:756–762. https://doi.org/10.1007/s13365-017-0550-z
Kammouni W, Wood H, Saleh A et al (2015) Rabies virus phosphoprotein interacts with mitochondrial Complex I and induces mitochondrial dysfunction and oxidative stress. J Neurovirol 21:370–382. https://doi.org/10.1007/s13365-015-0320-8
Venugopal AK, Ghantasala SSK, Selvan LDN et al (2013) Quantitative proteomics for identifying biomarkers for rabies. Clin Proteomics 10:1. https://doi.org/10.1186/1559-0275-10-3
Woldehiwet Z (2002) Rabies: recent developments. Res Vet Sci 73:17–25. https://doi.org/10.1016/S0034-5288(02)00046-2
Harish G, Mahadevan A, Pruthi N et al (2015) Characterization of traumatic brain injury in human brains reveals distinct cellular and molecular changes in contusion and pericontusion. J Neurochem 134:156–172. https://doi.org/10.1111/jnc.13082
Harish G, Venkateshappa C, Mahadevan A et al (2012) Effect of premortem and postmortem factors on the distribution and preservation of antioxidant activities in the cytosol and synaptosomes of human brains. Biopreserv Biobank 10:253–265. https://doi.org/10.1089/bio.2012.0001
Karunakaran S, Saeed U, Ramakrishnan S et al (2007) Constitutive expression and functional characterization of mitochondrial glutaredoxin (Grx2) in mouse and human brain. Brain Res 1185:8–17. https://doi.org/10.1016/j.brainres.2007.09.019
Karunakaran S, Saeed U, Mishra M et al (2008) Selective activation of p38 mitogen-activated protein kinase in dopaminergic neurons of substantia nigra leads to nuclear translocation of p53 in 1-methyl-4-phenyl-1,2,3,6-tetrahydropyridine-treated mice. J Neurosci 28:12500–12509. https://doi.org/10.1523/JNEUROSCI.4511-08.2008
Harish G, Venkateshappa C, Mahadevan A et al (2013) Mitochondrial function in human brains is affected by pre- and post mortem factors. Neuropathol Appl Neurobiol 39:298–315. https://doi.org/10.1111/j.1365-2990.2012.01285.x
Harish G, Mahadevan A, Srinivas Bharath MM, Shankar SK (2013) Alteration in glutathione content and associated enzyme activities in the synaptic terminals but not in the non-synaptic mitochondria from the frontal cortex of Parkinson’s disease brains. Neurochem Res 38:186–200. https://doi.org/10.1007/s11064-012-0907-x
Venkateshappa C, Harish G, Mahadevan A et al (2012) Elevated oxidative stress and decreased antioxidant function in the human hippocampus and frontal cortex with increasing age: implications for neurodegeneration in Alzheimer’s disease. Neurochem Res 37:1601–1614. https://doi.org/10.1007/s11064-012-0755-8
Venkateshappa C, Harish G, Mythri RB et al (2012) Increased oxidative damage and decreased antioxidant function in aging human substantia nigra compared to striatum: implications for Parkinson’s disease. Neurochem Res 37:358–369. https://doi.org/10.1007/s11064-011-0619-7
Mythri RB, Venkateshappa C, Harish G et al (2011) Evaluation of Markers of oxidative stress, antioxidant function and astrocytic proliferation in the striatum and frontal cortex of Parkinson’s disease brains. Neurochem Res 36:1452–1463. https://doi.org/10.1007/s11064-011-0471-9
Alladi PA, Mahadevan A, Yasha TC et al (2009) Absence of age-related changes in nigral dopaminergic neurons of Asian Indians: relevance to lower incidence of Parkinson’s disease. Neuroscience 159:236–245. https://doi.org/10.1016/j.neuroscience.2008.11.051
Chandana R, Mythri RB, Mahadevan A et al (2009) Biochemical analysis of protein stability in human brain collected at different post-mortem intervals. Indian J Med Res 129:189–199
Harish G, Venkateshappa C, Mahadevan A et al (2011) Effect of storage time, postmortem interval, agonal state, and genderon the postmortem preservation of glial fibrillary acidic protein and oxidatively damaged proteins in human brains. Biopreserv Biobank 9:379–387. https://doi.org/10.1089/bio.2011.0033
Harish G, Venkateshappa C, Mahadevan A et al (2011) Glutathione metabolism is modulated by postmortem interval, gender difference and agonal state in postmortem human brains. Neurochem Int 59:1029–1042. https://doi.org/10.1016/j.neuint.2011.08.024
Chinta SJ, Kommaddi RP, Turman CM et al (2005) Constitutive expression and localization of cytochrome P-450 1A1 in rat and human brain: presence of a splice variant form in human brain. J Neurochem 93:724–736. https://doi.org/10.1111/j.1471-4159.2005.03061.x
Agarwal V, Kommaddi RP, Valli K et al (2008) Drug metabolism in human brain: high levels of cytochrome P4503A43 in brain and metabolism of anti-anxiety drug alprazolam to its active metabolite. PLoS ONE 3:e0002337. https://doi.org/10.1371/journal.pone.0002337
Alladi PA, Mahadevan A, Shankar SK et al (2010) Expression of GDNF receptors GFRα1 and RET is preserved in substantia nigra pars compacta of aging Asian Indians. J Chem Neuroanat 40:43–52. https://doi.org/10.1016/j.jchemneu.2010.03.007
Alladi PA, Mahadevan A, Vijayalakshmi K et al (2010) Ageing enhances α-synuclein, ubiquitin and endoplasmic reticular stress protein expression in the nigral neurons of Asian Indians. Neurochem Int 57:530–539. https://doi.org/10.1016/j.neuint.2010.06.018
Dean DJ (1966) Laboratory techniques in rabies. The fluorescent antibody test. Monogr Ser World Health Organ 23:59–68
Sheehan D (1987) Theory and practice of histotechnology, 2nd edn. Battelle Press, Columbus
Frasca JM, Parks VR (1965) A routine technique for double-staining ultrathin sections using uranyl and lead salts. J Cell Biol 25:157–161. https://doi.org/10.1083/jcb.25.1.157
Hayat MA (2000) Principles and techniques of electron microscopy: biological applications, 4th edn. Cambridge University Press, Cambridge, p 543
Trounce IA, Kim YL, Jun AS, Wallace DC (1996) Assessment of mitochondrial oxidative phosphorylation in patient muscle biopsies, lymphoblasts, and transmitochondrial cell lines. Methods Enzymol 264:484–509. https://doi.org/10.1016/s0076-6879(96)64044-0
Wang W, Yang X, De Silanes IL et al (2003) Increased AMP:ATP ratio and AMP-activated protein kinase activity during cellular senescence linked to reduced HuR function. J Biol Chem 278:27016–27023. https://doi.org/10.1074/jbc.M300318200
Bradford M (1976) A Rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254. https://doi.org/10.1006/abio.1976.9999
Sharma T, Datta KK, Kumar M et al (2020) Intracranial aneurysm biomarker candidates identified by a proteome-wide study. Omi A J Integr Biol 24:483–492. https://doi.org/10.1089/omi.2020.0057
Kumar M, Varun CN, Dey G et al (2018) Identification of host-response in cerebral malaria patients using quantitative proteomic analysis. Proteomics 12:1600187. https://doi.org/10.1002/prca.201600187
Farahtaj F, Zandi F, Khalaj V et al (2013) Proteomics analysis of human brain tissue infected by street rabies virus. Mol Biol Rep 40:6443–6450. https://doi.org/10.1007/s11033-013-2759-0
Li L, Jin H, Wang H et al (2017) Autophagy is highly targeted among host comparative proteomes during infection with different virulent RABV strains. Oncotarget 8:21336–21350. https://doi.org/10.18632/oncotarget.15184
Bindea G, Mlecnik B, Hackl H, Charoentong P, Tosolini M, Kirilovsky A, Fridman WH, Pagès F, Trajanoski Z, Galon J (2009) ClueGO: a Cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics 25:1091–1093
Calvo SE, Clauser KR, Mootha VK (2016) MitoCarta: an inventory of mammalian mitochondrial genes|Broad Institute. Nucleic acids Res 44:D1251–D1257
Pagliarini DJ, Calvo SE, Chang B et al (2008) A mitochondrial protein compendium elucidates complex I disease Biology. Cell 134:112–123. https://doi.org/10.1016/j.cell.2008.06.016
Zhang W, Zhang Y, Zheng H, Zhang C, Xiong W, OlyarchukWalker JGM (2007) SynDB: a synapse protein DataBase based on synapse ontology. Nucleic Acids Res 35:D737–D741
Renjini R, Gayathri N, Nalini A, Srinivas Bharath MM (2012) Analysis of calpain-3 protein in muscle biopsies of different muscular dystrophies from India. Indian J Med Res 135:878–886
Schneider CA, Rasband WS, Eliceiri KW (2012) NIH Image to ImageJ: 25 years of image analysis. Nat Methods 9:671–675
Tyanova S, Temu T, Sinitcyn P et al (2016) The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat Methods 13:731–740
Peng J, Zhu S, Hu L et al (2016) Wild-type rabies virus induces autophagy in human and mouse neuroblastoma cell lines. Autophagy 12:1704–1720. https://doi.org/10.1080/15548627.2016.1196315
Fooks AR, Cliquet F, Finke S et al (2017) Rabies. Nat Rev Dis Prim 3:17091. https://doi.org/10.1038/nrdp.2017.91
Mahadevan A, Suja MS, Mani RS, Shankar SK (2016) Perspectives in diagnosis and treatment of rabies viral encephalitis: insights from pathogenesis. Neurotherapeutics 13:477–492
Piccinotti S, Whelan SPJ (2016) Rabies internalizes into primary peripheral neurons via clathrin coated pits and requires fusion at the cell body. PLoS Pathog 12:1–25. https://doi.org/10.1371/journal.ppat.1005753
Piccinotti S, Kirchhausen T, Whelan SPJ (2013) Uptake of rabies virus into epithelial cells by clathrin-mediated endocytosis depends upon actin. J Virol 87:11637–11647. https://doi.org/10.1128/jvi.01648-13
Walsh D, Naghavi MH (2019) Exploitation of cytoskeletal networks during early viral infection. Trends Microbiol 27:39–50
Dräger NM, Nachman E, Winterhoff M et al (2017) Bin1 directly remodels actin dynamics through its BAR domain. EMBO Rep 18:2051–2066. https://doi.org/10.15252/embr.201744137
Takenawa T, Suetsugu S (2007) The WASP-WAVE protein network: connecting the membrane to the cytoskeleton. Nat Rev Mol Cell Biol 8:37–48. https://doi.org/10.1038/nrm2069
Correas I, Padilla R, Avila J (1990) The tubulin-binding sequence of brain microtubule-associated proteins, tau and MAP-2, is also involved in actin binding. Biochem J 269:61–64. https://doi.org/10.1042/bj2690061
Cogliati S, Enriquez JA, Scorrano L (2016) Mitochondrial cristae: where beauty meets functionality. Trends Biochem Sci 41:261–273
Guarani V, McNeill EM, Paulo JA et al (2015) QIL1 is a novel mitochondrial protein required for MICOS complex stability and cristae morphology. Elife 4:1–23. https://doi.org/10.7554/eLife.06265
Xie J, Marusich MF, Souda P et al (2007) The mitochondrial inner membrane protein Mitofilin exists as a complex with SAM50, metaxins 1 and 2, coiled-coil-helix coiled-coil-helix domain-containing protein 3 and 6 and DnaJC11. FEBS Lett 581:3545–3549. https://doi.org/10.1016/j.febslet.2007.06.052
Harner M, Körner C, Walther D et al (2011) The mitochondrial contact site complex, a determinant of mitochondrial architecture. EMBO J 30:4356–4370. https://doi.org/10.1038/emboj.2011.379
Head BP, Zulaika M, Ryazantsev S, Van Der Bliek AM (2011) A novel mitochondrial outer membrane protein, MOMA-1, that affects cristae morphology in Caenorhabditis elegans. Mol Biol Cell 22:831–841. https://doi.org/10.1091/mbc.E10-07-0600
Weber TA, Koob S, Heide H et al (2013) APOOL is a cardiolipin-binding constituent of the mitofilin/MINOS protein complex determining cristae morphology in mammalian mitochondria. PLoS ONE 8:e63683. https://doi.org/10.1371/journal.pone.0063683
Davies KM, Anselmi C, Wittig I et al (2012) Structure of the yeast F 1F o-ATP synthase dimer and its role in shaping the mitochondrial cristae. Proc Natl Acad Sci USA 109:13602–13607. https://doi.org/10.1073/pnas.1204593109
Eydt K, Davies KM, Behrendt C et al (2017) Cristae architecture is determined by an interplay of the MICOS complex and the F1Fo ATP synthase via Mic27 and Mic10. Microb Cell 4:259–272. https://doi.org/10.15698/mic2017.08.585
Strauss M, Hofhaus G, Schröder RR, Kühlbrandt W (2008) Dimer ribbons of ATP synthase shape the inner mitochondrial membrane. EMBO J 27:1154–1160. https://doi.org/10.1038/emboj.2008.35
Ramadasan-Nair R, Gayathri N, Mishra S et al (2014) Mitochondrial alterations and oxidative stress in an acute transient mouse model of muscle degeneration: implications for muscular dystrophy and related muscle pathologies. J Biol Chem 289:485–509. https://doi.org/10.1074/jbc.M113.493270
Sunitha B, Gayathri N, Kumar M et al (2016) Muscle biopsies from human muscle diseases with myopathic pathology reveal common alterations in mitochondrial function. J Neurochem 138:174–191. https://doi.org/10.1111/jnc.13626
Liu J, Wang H, Gu J et al (2017) BECN1-dependent CASP2 incomplete autophagy induction by binding to rabies virus phosphoprotein. Autophagy 13:739–753. https://doi.org/10.1080/15548627.2017.1280220
Sun Y, Yu S, Ding N et al (2014) Autophagy benefits the replication of newcastle disease virus in chicken cells and tissues. J Virol 88:525–537. https://doi.org/10.1128/jvi.01849-13
Heaton NS, Randall G (2011) Dengue virus and autophagy. Viruses 3:1332–1341
Jackson AC (2000) Rabies. Can J Neurol Sci 27(4):278–282
Beck S, Gunawardena P, Horton DL et al (2017) Pathobiological investigation of naturally infected canine rabies cases from Sri Lanka. BMC Vet Res 13:1–9. https://doi.org/10.1186/s12917-017-1024-5
Farahtaj F, Alizadeh L, Gholami A et al (2019) Natural infection with rabies virus: a histopathological and immunohistochemical study of human brains. Osong Public Heal Res Perspect 10:6–11. https://doi.org/10.24171/j.phrp.2019.10.1.03
Perez-Riverol Y, Csordas A, Bai J et al (2019) The PRIDE database and related tools and resources in 2019: improving support for quantification data. Nucleic Acids Res 47:D442–D450. https://doi.org/10.1093/nar/gky1106
Acknowledgements
The technical help provided by Mr. Ramesh, Ms. Rashmi S and Mr. Satheesh from the Electron Microscopy Facility, Department of Neuropathology, NIMHANS is gratefully acknowledged. The assistance of Mr. Shivaji Rao and Mrs. Rajasakti from the Human Brain Tissue Repository, NIMHANS with the histopathology experiments is gratefully acknowledged. The authors acknowledge the help of Dr. Gajanan Sathe, Institute of Bioinformatics, Bangalore for his help with analysis of MS data.
Funding
PKH is supported through the project titled “Testing of blood samples for rabies virus neutralizing antibodies by RFFIT” (Grant no. OTHERS/001/107/2014/00713), at the department of Neurovirology, NIMHANS, Bangalore, India, funded by Cadila Pharmaceuticals, India (Principal Investigator-RSM). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
PKH: Methodology; data curation; validation; formal analysis; writing—original draft preparation, SR: conceptualization; data curation; writing—original draft preparation, GY: resources, KKM: methodology; data curation, GD: methodology; data curation, AY; methodology, BKCS: analysis, AM: conceptualization; data analysis; writing-review and editing; resources; supervision, MMSB: conceptualization; writing-review and editing, supervision; project administration, RSM: conceptualization; writing-review and editing, supervision; project administration; resources.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest with the contents of this article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Harsha, P.K., Ranganayaki, S., Yale, G. et al. Mitochondrial Dysfunction in Rabies Virus-Infected Human and Canine Brains. Neurochem Res 47, 1610–1636 (2022). https://doi.org/10.1007/s11064-022-03556-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11064-022-03556-6