Abstract
Branched chain amino acids (BCAA) are catabolized into various acyl-CoA compounds, which are key precursors used in polyketide productions. Because of that, BCAA catabolism needs fine tuning of flux balances for enhancing the production of polyketide antibiotics. To enhance BCAA catabolism for pikromycin production in Streptomyces venezuelae ATCC 15439, three key enzymes of BCAA catabolism, 3-ketoacyl acyl carrier protein synthase III, acyl-CoA dehydrogenase, and branched chain α-keto acid dehydrogenase (BCDH) were manipulated. BCDH overexpression in the wild type strain resulted in 1.3 fold increase in pikromycin production compared to that of WT, resulting in total 25 mg/L of pikromycin. To further increase pikromycin production, methylmalonyl-CoA mutase linked to succinyl-CoA production was overexpressed along with BCDH. Overexpression of the two enzymes resulted in the highest titer of total macrolide production of 43 mg/L, which was about 2.2 fold increase compared to that of the WT. However, it accumulated and produced dehydroxylated forms of pikromycin and methymycin, including their derivatives as well. It indicated that activities of pikC, P450 monooxygenase, newly became a bottleneck in pikromycin synthesis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Streptomyces species are soil-dwelling bacteria producing vast numbers of polyketide secondary metabolites such as antibiotics and anti-cancer drugs, but such secondary metabolites are often produced in very small quantities [18]. There are many efforts and studies to increase the productions of such secondary metabolites [11, 33] in academia as well as fermentation industry. Among them, flux balance between fatty acids (FA) and polyketide (PK) biosynthesis is one of general strategies to overproduce molecules of interests, because FAs and PKs share common precursor pools [1, 10]. The commonality of using the same acyl-CoA substrates between fatty acid synthase (FAS) genes and polyketide synthase (PKS) genes would result a reciprocal relationship in their production yields at their high level productions, due to similarities in their gene sequences and protein structures [4]. To increase PK antibiotics productions, the precursors in FA biosynthesis should be redirected towards PK biosynthesis.
Dynamic controlling fluxes of precursor pools towards FA biosynthesis and antibiotics production is very challenging, since FAs are essential components of lipid, consisting of 7–10% of dry cell weight, as well as energy source for cell growth. One method to repress FA synthesis and increase PK productions is “redirection”, a strategy to reduce fluxes toward acyl-CoA consuming pathways. In a recent publication, addition of other antibiotics such as triclosan was newly suggested in Streptomyces sp. to achieve successful repression of fatty acids synthesis [33]. And, enhancements of actinorhodin (ACT) and salinomycin productions from Streptomyces coelicolor and Streptomyces albus, respectively [32], were also suggested by the addition of triclosan, which repressed activities of enoyl-acyl carrier protein reductase of FAS gene. However, this strategy has some drawbacks. First, additions of antibiotics such as triclosan are impractical and unreasonably expensive in real application [19]. In addition, their clearance during purification generates another problem. As a result, controlling FAS gene expression may be a better strategy to reduce FA synthesis. Another general, the so-called “pull” method to redirect acyl-CoA fluxes from FAS into PKS, is to overexpress an entire PKS gene cluster [20], or by manipulations of transcriptional regulators [34]. But, PKS gene clusters are very bulky, which is hard to overexpress.
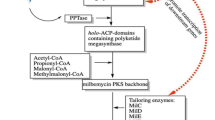
Enhancements of PK antibiotics productions can also be achieved with increasing entire precursor pools, so-called “push” method, by enhancing the fluxes toward catabolism of key nutrients such as glucose [27, 41]. Glycolysis is a major source of acyl-CoA precursor productions [2, 3, 22]. However, as there are limits to how much antibiotics could be produced from one nutrient source, [12, 24] metabolisms of other nutrients such as amino acids should also be considered. In case of PK, branched chain amino acid (BCAA) degradation was known to provide sufficient amounts of acyl-CoA precursors (Fig. 1a). Branched chain α-keto acid dehydrogenase (BCDH) is a key common enzyme complex, participating in the catabolism of BCAA. Direct products of BCDH from valine, isoleucine, and leucine catabolism, are isobutyryl-CoA, α-methylbutyryl-CoA, and isovaleryl-CoA, respectively. The three direct products of BCDH are further metabolized by acyl-CoA dehydrogenase (ACAD) [25, 31], or are used in synthesis of branched chain fatty acids (BCFA) [35]. It was previously reported that 50% of acetyl-CoA for ACT biosynthesis was supplied from the BCAA degradation pathway [31], and about 52 fold increase in ACT production was observed with BCDH overexpression mutant [10]. Along with BCDH, ACAD also plays a key role in BCAA catabolism. ACAD pulls the fluxes of the three products of BCDH reactions away from fatty acid synthesis, and direct them to synthesis of the short chain acyl-CoA compounds, which are acetyl-CoA, propionyl-CoA, and methylmalonyl-CoA [25, 42].
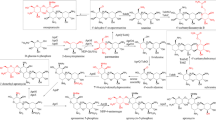
a TCA cycle and valine catabolism pathway leading to methylmalonyl-CoA. Overexpression targets are indicated by bold arrow, and repression target by dashed arrow. b Pikromycin (PKM) and methymycin (MTM) biosynthesis pathway. PKM and MTM are produced from type I PKS genes, composed of 6 modules, from PikAI through PikAV, and a thioesterase (TE). Two different TEs, TEI and TEII, determine the production of PKM or MTM. Post-production modification occurs after the synthesis of PK backbone. Glycosylation by DesVII and hydroxylation by PikC result various macrolides productions in S. venezuelae
Despite of such importance of the two enzymes in the secondary metabolism, effects of BCDH and ACAD overexpression for antibiotics productions were not thoroughly investigated in S. coelicolor or S. lividans model strains, since ACT overproduced by the two strains do not use propionyl-CoA and methylmalonyl-CoA, but only use acetyl-CoA and malonyl-CoA as their precursors [27].
We were interested in how BCAA catabolism would affect pikromycin (PKM) production in S. venezuelae, as PKM is biosynthesized using six methylmalonyl-CoAs and one malonyl-CoA [23, 36] by PKS genes (Fig. 1b), and many previous reports on PKM overproduction focused on the manipulations of transcriptional regulators such as wblA [37], afsR, and metK [17] rather than actual metabolic enzymes participating in the biosynthetic or degradation pathways. A simple solution to investigate the effects of BCAA catabolism on the PKM production is to make net fluxes of precursor pools positive by the overexpression of the key enzymes enhancing its biosynthesis.
In this study, we applied and combined different methylmalonyl-CoA synthetic routes for PKM biosynthesis in S. venezuelae ATCC 15439. As overexpression of PKS gene cluster may result in the limitations of acyl-CoA precursor pools, we wanted to set up a guideline for selection of optimal metabolic pathways to overproduce PK antibiotics (for example, PDH for ACT productions in S. coelicolor). Using BCAA catabolic pathway, we applied three different strategies for PKM overproduction: (1) “redirection” of acyl-CoA fluxes by repression of β-ketoacyl acyl carrier protein synthase III (KASIII), competing pathway, (2) “push” by overexpression of BCDH, and (3) “pull” by overexpression of ACAD to increase BCAA catabolism. Methylmalonyl-CoA mutase (MCM) was overexpressed along with BCDH to combine methylmalonyl-CoA synthesis from both BCAA catabolism and tricarboxylic acid (TCA) cycle.
Materials and methods
Bacterial strains, plasmids, and culture conditions
All strain information and plasmids used in this study are listed in Table 1. E. coli strains were cultured in Luria–Bertani (LB) broth (Becton–Dickinson, USA) with 0.1 mg/mL ampicillin or 50 μg/mL apramycin when needed.
R2YE complex medium was made following standard protocols [9], and used for cultures of S. venezuelae strains. R2YE, Difco Nutrient Agar (DNA) media (Becton–Dickinson, USA), and Mannitol Soya flour Agar (MSA) media made by standard protocols were used for conjugation of cloned plasmids such as p1220FabH and pSET152PikC, and S. venezuelae spore preparations. For selection of S. venezuelae mutants and conjugates, 8 μg/mL thiostrepton and 50 μg/mL apramycin were used. Filter sterilized 100 mM valine was added to S. venezuelae cultures at the time of inoculation to study the effects of valine on pikromycin production. Cell growth was also observed by UV absorbance at 600 nm using Multiskan spectroscopy (Thermo Fisher Scientific, USA), and cell pellets after pikromycin extractions were dried in an oven at 60 °C for dry cell weight (DCW) measurements.
Escherichia coli DH5α was cultured at 37 °C. E. coli ET12567 and Streptomyces strains were cultured at 30 °C, both with 200 rpm shaking in Lab Companion SK-71 Bench top shaker (Jeio Tech, South Korea). 200 mg of wet weighted cells from 50 mL seed cultures grown overnight were inoculated into 50 mL R2YE media in 250 mL flasks.
Strain construction
All primers and their sequences are listed in Table S1. PCR amplifications were performed with LA-taq DNA polymerase with GC buffer (TaKaRa, Japan) using Thermal Cycler Dice (TaKaRa, Japan). FastDigest (Thermo Fisher Scientific, USA) restriction enzymes were used for digestions of PCR amplified DNA fragments.
Promoter and 5′ UTR for repression of KASIII were selected from S. coelicolor, which was described in the previous study [40]. Three promoters were tested including PSCO4505, PSCO4808 and PSCO1214. Promoter of SCO1214 and 5′ UTR of SCO2078 were selected to repress KASIII transcription. A cosmid of KASIII carrying promoter of SCO1214 and 5′ UTR of SCO2078 was constructed using SuperCos-1 with Gibson Assembly (New England BioLabs, USA) following a procedure of the manufacturer. Overexpression plasmids of KASIII (SCO2388), BCDH (SCO3815-3817), ACAD (SCO2279), and MCM (SCO4869), were constructed by conventional cloning methods with restriction enzymes. pikC overexpression plasmid was constructed in the same manner. For the integration of ermE*P pikC into the host chromosome, pikC was first inserted into pIBR25 plasmid, resulting in pIBR25PikC. ermE*P and pikC were PCR amplified together using pIBR25PikC as a template, and then inserted into pSET152.
Plasmids were conjugated to S. venezuelae using E. coli ET12567, and all other overexpression plasmids were transformed into S. venezuelae using protoplast transformation methods [9].
Quantification of KASIII transcription level
mRNA of S. venezuelae mutants were extracted with RNeasy Mini Kit (Qiagen, Germany). cDNAs were synthesized from 1.5 µg of the extracted mRNAs using M-MLV reverse transcriptase (Promega, USA). mRNA extraction and cDNA synthesis were performed according to standard protocols provided by the manufacturers. 16S rRNA was used as an internal standard for normalization of real-time PCR (qPCR) data, and calculation of relative KASIII expression levels. LightCycler 480 (Roche, Switzerland) was used for qPCR analysis. TOPreal qPCR 2X PreMIX containing SYBR Green with low ROX (Enzynomics, Korea), and standard qPCR settings from the SYBR manufacturer were used for the analysis.
Pikromycin extraction and quantification
On day 3 of S. venezuelae cultures, 40 mL culture broths were harvested and supernatants were collected by centrifugation at 2840×g for 10 min. PKM was extracted from the supernatants using two volumes (80 mL) of ethyl acetate. After the extracts were dried with a rotary vacuum evaporator, the remaining part was dissolved with 1.8 mL methanol. Leftover cell pellets from the harvested 40 mL cultures were used for calculations of DCW.
PKM, methymycin (MTM) and their derivatives were quantified using 20 µL of cell extracts by high-performance liquid chromatography (HPLC) YL-9100 (Younglin, Korea). A C18 reverse phase column (Waters, USA) with a linear gradient system at flow rate of 1 mL/min was used as described in our previous study [39]. PKM, MTM, and their derivatives corresponding peaks were analyzed and confirmed by TSQ Quantum Access Max Triple Quadrupole Mass Spectrometer, (TSQ-MS, Thermo Fisher Scientific, USA) using 5% ACN with 0.28% ammonium hydroxide and 5 mM ammonium acetate.
Results and discussion
Impact of valine supplementation on pikromycin production
Catabolism of BCAA results in various low molecular weight acyl-CoA (Fig. 1a). Valine can possibly produce three CoA compounds, acetyl-CoA, propionyl-CoA, and methylmalonyl-CoA. Thus, we focused on the effects of valine supplementation on PKM production. To examine an optimal level of valine feeding in batch flask cultures of S. venezuelae, R2YE complex media containing five different final valine concentrations (i.e., 2, 4, 6, 8, and 10 mM) were compared for cell growths and PKM production preliminarily. Valine supplementation tends to decrease cell mass (Fig. 2a). The maximum PKM titer, 22.6 mg/L, was observed at 2 mM valine, which was 1.2 folds more than that without valine supplementation (Fig. 2b), and 1.7 fold increase in production per cell mass (Fig. S1).
A reduction of PKM production with increasing valine concentrations may be explained by roles of valine. Valine supplementation was previously reported to activate expressions of pyruvate dehydrogenase, but repress glycolysis, inhibiting pyruvate formation in Bacillus subtilis [38]. In Streptomyces ambofaciens, excess valine resulted in ammonium formation and accumulation, reducing cell mass and spiramycin production yields [16]. Reduced cell mass in S. venezuelae at higher valine concentrations may have reduced pikromycin productions as well. In addition, valine is an activator of a global regulator codY. codY is activated by access valine, and represses formation of acetyl-CoA from acetate, isocitrate from citrate, and glutamate from 2-oxoglutarate [5, 8, 29]. Additionally, biosynthesis of BCAAs are subjected to many other regulations such as amino acid metabolisms (i.e., glutamate and arginine), and Lrp family transcriptional regulators [29], so that an optimal valine feeding concentration is required. Since 2 mM valine supplementation resulted in some increase in PKM production, it appeared that there were still rooms to utilize BCAA degradation pathways by modulating metabolic enzymes, and/or regulators to increase PKM production.
BCAA catabolism and pikromycin productions
When BCAAs are catabolized into acyl-CoA compounds via BCDH, they are used for either synthesis of BCFA or PK secondary metabolites. KASIII for BCFA [7] and ACAD for short chain acyl-CoAs [42] are responsible for the two pathways, respectively. Since FAS and PKS compete for the same substrates, and reduction of FA synthesis was effective to improve PK productions [33], reduction of KASIII expression was first performed for PKM. We aimed to reduce acyl-CoA flux into FA synthesis by controlling KASIII gene expression, replacing a promoter and 5′ UTR of KASIII with the ones developed from our previous study [40].
Promoter of SCO1214 (PSCO1214) was included on top of the two previously developed promoter and 5′ UTR sets from S. coelicolor. Total three sets of promoters and 5′ UTRs were inserted and replaced the WT promoter and RBS of KASIII in the chromosome. The three sets are PSCO4505RSCO0641, PSCO4808RSCO2078, and PSCO1214RSCO2078, and they were chosen for low expression levels of β-d-glucuronidase. Nonfunctional S. coelicolor promoters and 5′ UTRs in S. venezuelae were easily screened out by live and death assay, because without KASIII, cells could not grow [15]. PSCO1214RSCO2078 was selected to repress KASIII transcription. All the names of the promoters and 5′ UTRs with their sequences are listed in Table S2. According to data of qPCR measurement, successful 24 folds repression of KASIII transcription was achieved using PSCO1214 and RSCO2078 (SVEN1220KASIII) compared to that of the wild type (Fig. S2a).
The growth of SVEN1220KASIII slightly decreased compared to that of the wild type, measured by cell optical density (OD) (Fig. S2b), which was also confirmed by DCW measurement [i.e., DCW1220FabH = 6 g/L, which is about 83% of the control (Fig. S2c)]. In contrast, PKM titer of SVEN1220KASIII decreased more dramatically, by almost a half (Fig. S2d). Previous reports indicated that cell mass at the time of secondary metabolite production is very important, and severe decrease in cell mass may also decrease the production of secondary metabolites [33]. Decrease in PKM production from SVEN1220KASIII can be such case. Therefore, we realized that KASIII repression is not a good strategy for enhancing PKM production at this moment. It would require more thorough understanding of regulating KASIII expression.
To increase metabolic flux towards methylmalonyl-CoA synthesis, we used push and pull strategy by overexpressing BCDH (SVENBCDH) and ACAD (SVENACAD). DCW among SVEN1220KASIII, SVENBCDH, and SVENACAD were almost identical (i.e., about 7 g/L DCW) (Fig. 3a). The maximum concentration of PKM was about 25 mg/L, which was 1.3 fold increase compared to the WT (Fig. 3b). Increase in PKM production due to BCDH or ACAD overexpression were almost identical. Overexpression of the two well-known positive regulators metK and afsR produced about 1.5 and 2 mg/L of PKM, respectively, which were 1.6 fold and 2.6 fold increase of the WT [17]. Deletion of antibiotic biosynthesis down-regulator, wblA, increased PKM production from 0.2 to 0.6 mg/L, which was 3.5 fold increase [37]. It would be difficult to compare our data with the previous reports due to the differences in culture and production scale. Although our result was a rather small increase in fold changes compared to the previous studies from other groups, such mutations in transcriptional regulators do not generally affect the activity of other metabolic enzymes, so that they can be independently and easily combined with our mutants (data not shown).
One possible explanation of smaller increase could be a low possibility of methylmalonyl-CoA synthesis from the BCAA catabolism. Theoretically, only acetyl-CoAs could be produced from leucine. Either one acetyl-CoA or one propionyl-CoA could be produced from isoleucine, and either one acetyl-CoA, one propionyl-CoA, or one methylmalonyl-CoA could be produced from valine [28, 43]. Therefore, there is a lower probability of methylmalonyl-CoA synthesis for PKM, compared to acetyl-CoA synthesis from BCAA catabolism for ACT. But, since acetyl-CoA can be converted into methylmalonyl-CoA via succinyl-CoA through TCA cycle, perhaps the rate-determining step (RDS) of the synthesis of PKM may be in a different step or in some gene regulation of the pathway.
Double overexpression mutant of BCDH and MCM
A double overexpression mutant of BCDH and MCM (SVENBCMC) was constructed to examine if there were any additive effects of the two genes on PKM production. From central carbon metabolism, methylmalonyl-CoA is produced from succinyl-CoA of TCA cycle by MCM. It is a well-known overexpression target, which increased methylmalonyl-CoA pool. Overexpression of MCM overproduced erythromycin by about 1.4–1.64 folds from Saccharopolyspora erythraea [26]. MCM was a good overexpression candidate, along with BCDH, to maximize PKM production in S. venezuelae for that reason. All the individual enzymes were overexpressed with ermE promoter in pIBR25 plasmids. All the strains except SVENMCM yielded almost the same DCW, about 7 g/L (Fig. S3a). The cell mass of SVENMCM was much higher than that of others, 9.5 g/L, suggesting that the contribution of the supply of methylmalonyl-CoA via TCA cycle possibly be more significant than that via BCAA degradation. The most PKM was produced from SVENMCM, 33.9 mg/L, which was 1.74 fold increase compared to that of the WT.
Surprisingly, PKM titer of SVENBCMC decreased down to 25.0 mg/L (Fig. S3b), which was almost the same level as that of SVENBCDH. To further examine a reason for such a decrease, we performed more detailed HPLC and LC–MS analysis. Three new compound peaks were detected from chromatograms of HPLC (Fig. 4), and four from TSQ-MS analysis (Fig. 5). The three newly detected compounds from HPLC were collected and further analyzed by MS/MS.
According to MS/MS analysis of the three unidentified compounds from SVENBCMC culture extracts, the three peaks were 486.45 m/z, 470.40 m/z, and 510.53 m/z, identified as [novamethymycin + H]+ (Fig. S4a), [neomethymycin + H]+ (Fig. S4b), and [narbomycin + H]+ (Fig. S5a), respectively. Novapikromycin was also detected from crude extract of PK antibiotics from SVENBCMC (Fig. S5b). Nova-forms are C12 hydroxylated forms of MTM and PKM. In SVENBCMC, fair amounts of deoxy forms of PK antibiotics were accumulated, which may also explain the decrease in the PKM production. Based on the TSQ-MS data, macrolides were quantified to calculate total production.
Despite the reduction of PKM production in SVENBCMC compared to that of SVENMCM, total macrolide productions increased on the other hand (Fig. 6a). Total 43 mg/L of macrolides were produced from SVENBCMC, which was about 2.2 fold increase compared to that of the WT. Considering error ranges, it was about the same as that of SVENMCM. PKM and MTM derivatives, including their deoxy forms, increased dramatically, taking a huge portion in macrolides production (Fig. 6b).
Total macrolides production per biomass of SVENBCMC was increased by 2.3 fold of the WT, which was about 7.41 mg/L/gcell (Fig. S6a). Interestingly, PKM production per biomass was about the same among the five strains, considering their error ranges (Fig. S6b). Such results indicated that supplementation of acyl-CoA precursors from metabolism was no longer a bottleneck within SVENBCMC. But, activities of pikC, cytochrome P450 monooxygenase, became a new bottleneck.
pikC is responsible for production of various PKM and MTM derivatives (Fig. S7). For PKM family, pikC recognizes desosamine groups, and adds hydroxyl groups at C12 and C14 positions [13, 30]. The enzyme appears to have somewhat broad substrate specificity, and it alone produces neo and nova form derivatives of PKM and MTM. As a result, the increase in PKM production may also mean the increase in its substrate production for pikC as well. It applies the same for MTM. But, pikC expression throughout the study was not manipulated according to increasing PKM production. In other words, pikC expressions in SVENBCDH, SVENACAD, SVENMCM, and SVENBCMC were the same. It was reasonable to assume that there were no sufficient pikC available for the hydroxylation of PKM and MTM deoxy forms. Such explanation is not enough, and optimizations in the productions of PKM and MTM would require further detailed studies, but to convert the accumulated deoxy form antibiotics into PKM and MTM, overexpression of pikC in SVENBCMC was attempted.
To make a stable expression of pikC, the gene was integrated into the chromosome (SVENpikC). Interestingly enough, transformation of pIBR25BM into SVENpikC was not possible, because the transformed cells died (data not shown). There are some reports mentioning that P450 overexpression in E. coli reduces cell growth fitness [6], but there were no vivid differences in growth rates of the WT and SVENpikC. However, whenever pikC was overexpressed with other enzymes, via either integrated into chromosome or plasmid amplification, S. venezuelae cells died, which we do not understand well yet. As a result, we are suggesting that there needs to be another method to hydroxylate narbomycin and 10-deoxymethymycin.
In result, conversion of narbomycin into PKM was not possible with pikC overexpression, but a possible strategy to solve such problem would be modifications of pikC substrate specificity. As long as desosamine was attached to PK compounds, pikC was shown to hydroxylate many of them in the previous reports, resulting in numerous hydroxylated aminoglycoside products [14]. Taking advantages of its large substrate specificity, pikC was mutated to accept various substrates with different amino sugars or types of aglycones [21]. If pikC could be mutated to accept narbomycin and nothing else, it would be possible to further maximize the PKM production.
Conclusion
Streptomyces venezuelae, producing mainly PKM and MTM, received spotlights as a good heterologous expression host due to its fast growing nature compared to other Streptomyces sp. [23]. In this study, synthesis of methylmalonyl-CoA from BCAA degradation was applied in S. venezuelae by manipulating expressions of three key enzymes involved. In S. coelicolor, it was previously reported that BCAA catabolism showed similar contributions towards acetyl-CoA for ACT production with carbon metabolism, and its importance in PK antibiotics was emphasized [10, 31]. Unfortunately, however, methylmalonyl-CoA coming from succinyl-CoA of TCA cycle had greater influence than that from BCAA degradation towards PKM productions in S. venezuelae. The highest titer of PKM was achieved by MCM overexpression, resulting in 1.7 fold increase, 33.9 mg/L, compared to that of the WT. Overexpression of BCDH and MCM together resulted in the highest macrolide production titer of 43 mg/L, which were about 2.2 fold increase compared to that of the WT. Intermediates and other forms of PKM and MTM began to be accumulated, suggesting enzymatic activities or control nods of pikC (i.e., gene regulation) were altered. It would be necessary to manipulate activities or substrate specificities of pikC to accept only narbomycin as a substrate in BCDH and MCM double overexpression mutant to further maximize PKM productions.
References
Brakhage AA (2004) Molecular biotechnology of fungal beta-lactam antibiotics and related peptide synthetases. Springer Sciences and Business Media, Germany, pp 160–177
Chan YA, Podevels AM, Kevany BM, Thomas MG (2009) Biosynthesis of polyketide synthase extender units. Nat Prod Rep 26:90–114
Dekleva ML, Strohl WR (1988) Biosynthesis of epsilon-rhodomycinone from glucose by Streptomyces C5 and comparison with intermediary metabolism of other polyketide-producing streptomycetes. Can J Microbiol 34:1235–1240
Gajewski J, Buelens F, Serdjukow S, Janben M, Cortina N, Grumuller H, Grininger M (2017) Engineering fatty acid synthases for directed polyketide production. Nat Chem Biol 13:363–365
Gopalani M, Dhiman A, Rahi A, Bhatnagar R (2016) Overexpression of the pleiotropic regulator CodY decreases sporulation, attachment and pellicle formation in Bacillus anthracis. Biochem Biophys Res Commun 469:672–678
Julsing MK, Fichera MA, Malz F, Ebbelaar M, Bos R, Woerdenbag HJ, Quax WJ, Kayser O (2008) Bioconversion of mono- and sesquiterpenoids by recombinant human cytochrome P450 monooxygenases. Pharm Biol 46:710–718
Kaneda T (1991) Iso- and anteiso-fatty acids in bacteria: biosynthesis, function, and taxonomic significance. Microbiol Rev 55:288–302
Katz E, Brown D (1989) A possible rold of d-valine and related d-amino acids in repression of enzyme and actinomycin synthesis. Appl Microbiol Biotechnol 30:67–70
Kieser T, Bibb MJ, Buttner MJ, Chater KF, Hopwood DA (2000) Practical Streptomyces genetics. John Innes Foundation Norwich, England, pp 229–422
Kim M, Yi JS, Kim J, Kim JN, Kim MW, Kim BG (2014) Reconstruction of a high-quality metabolic model enables the identification of gene overexpression targets for enhanced antibiotic production in Streptomyces coelicolor A3(2). Biotechnol J 9:1185–1194
Komatsu M, Uchiyama T, Omura S, Cane DE, Ikeda H (2010) Genome-minimized Streptomyces host for the heterologous expression of secondary metabolism. Proc Natl Acad Sci USA 107:2646–2651
Laakel M, Lebrihi A, Khaoua S, Schneider F, Lefebvre G, Germain P (1994) Relationship between valine, fatty acids, and spiramycin biosynthesis in Streptomyces ambofaciens. Can J Microbiol 40:672–676
Lee SK, Park JW, Kim JW, Jung WS, Park SR, Choi CY, Kim ES, Kim BS, Ahn JS, Sherman DH, Yoon YJ (2006) Neopikromycin and novapikromycin from the pikromycin biosynthetic pathway of Streptomyces venezuelae. J Nat Prod 69:847–849
Li S, Chaulagain MR, Knauff AR, Podust LM, Montgomery J, Sherman DH (2009) Selective oxidation of carbolide C–H bonds by an engineered macrolide P450 mono-oxygenase. Proc Natl Acad Sci USA 106:18463–18468
Li Y, Florova G, Reynolds KA (2005) Alteration of the fatty acid profile of Streptomyces coelicolor by replacement of the initiation enzyme 3-ketoacyl acyl carrier protein synthase III (FabH). J Bacteriol 187:3795–3799
Lounes A, Lebrihi A, Benslimane C, Lefebvre G, Germain P (1995) Regulation of valine catabolism by ammonium in Streptomyces ambofaciens, producer of spiramycin. Can J Microbiol 41:800–808
Maharjan S, Oh TJ, Lee HC, Sohng JK (2008) Heterologous expression of metK1-sp and afsR-sp in Streptomyces venezuelae for the production of pikromycin. Biotechnol Lett 30:1621–1626
McKenzie NL, Thaker M, Koteva K, Hughes DW, Wright GD, Nodwell JR (2010) Induction of antimicrobial activities in heterologous streptomycetes using alleles of the Streptomyces coelicolor gene absA1. J Antibiot (Tokyo) 63:177–182
Morvan C, Halpern D, Kenanian G, Hays C, Anba-Mondoloni J, Brinster S, Kennedy S, Trieu-Cuot P, Poyart C, Lamberet G, Gloux K, Gruss A (2016) Environmental fatty acids enable emergence of infectious Staphylococcus aureus resistant to FASII-targeted antimicrobials. Nat Commun 7:12944
Nah HJ, Pyeon HR, Kang SH, Choi SS, Kim ES (2017) Cloning and heterologous expression of a large-sized natural product biosynthetic gene cluster in Streptomyces species. Front Microbiol 8:394–404
Negretti S, Narayan AR, Chiou KC, Kells PM, Stachowski JL, Hansen DA, Podust LM, Montgomery J, Sherman DH (2014) Directing group-controlled regioselectivity in an enzymatic C–H bond oxygenation. J Am Chem Soc 136:4901–4904
Panagiotou G, Andersen MR, Grotkjaer T, Regueira TB, Nielsen J, Olsson L (2009) Studies of the production of fungal polyketides in Aspergillus nidulans by using systems biology tools. Appl Environ Microbiol 75:2212–2220
Phelan RM, Sachs D, Petkiewicz SJ, Barajas JF, Blake-Hedges JM, Thompson MG, Reider Apel A, Rasor BJ, Katz L, Keasling JD (2017) Development of next generation synthetic biology tools for use in Streptomyces Venezuelae. ACS Synth Biol 6:159–166
Podojil M, Steinerova N, Cudlin J (1989) Relationship between the fatty acid composition and the type of antibiotics produced by Streptomyces lasaliensis. J Basic Microbiol 29:605–609
Pulsawat N, Kitani S, Kinoshita H, Lee CK, Nihira T (2007) Identification of the BkdAB gene cluster, a plausible source of the starter-unit for virginiamycin M production in Streptomyces virginiae. Arch Microbiol 187:459–466
Reeves AR, Brikun IA, Cernota WH, Leach BI, Gonzalez MC, Weber JM (2007) Engineering of the methylmalonyl-CoA metabolite node of Saccharopolyspora erythraea for increased erythromycin production. Metab Eng 9:293–303
Ryu YG, Butler MJ, Chater KF, Lee KJ (2006) Engineering of primary carbohydrate metabolism for increased production of actinorhodin in Streptomyces coelicolor. Appl Environ Microbiol 72:7132–7139
She P, Olson KC, Kadota Y, Inukai A, Shimomura Y, Hoppel CL, Adams SH, Kawamata Y, Matsumoto H, Sakai R, Lang CH, Lynch CJ (2013) Leucine and protein metabolism in obese Zucker rats. PLoS ONE 8:e59443
Sonenshein AL (2007) Control of key metabolic intersections in Bacillus subtilis. Nat Rev Microbiol 5:917–927
Srinivasan A, Bach H, Sherman DH, Dordick JS (2004) Bacterial P450-catalyzed polyketide hydroxylation on a microfluidic platform. Biotechnol Bioeng 88:528–535
Stirrett K, Denoya C, Westpheling J (2009) Branched-chain amino acid catabolism provides precursors for the type II polyketide antibiotic, actinorhodin, via pathways that are nutrient dependent. J Ind Microbiol Biotechnol 36:129–137
Swiatek MA, Gubbens J, Bucca G, Song E, Yang YH, Laing E, Kim BG, Smith CP, van Wezel GP (2013) The ROK family regulator Rok7B7 pleiotropically affects xylose utilization, carbon catabolite repression, and antibiotic production in Streptomyces coelicolor. J Bacteriol 195:1236–1248
Tanaka Y, Izawa M, Hiraga Y, Misaki Y, Watanabe T, Ochi K (2017) Metabolic perturbation to enhance polyketide and nonribosomal peptide antibiotic production using triclosan and ribosome-targeting drugs. Appl Microbiol Biotechnol 101:4417–4431
Tang ZK, Li XM, Pang AP, Lin CY, Zhang Y, Zhang J, Qiao J, Zhao GR (2017) Characterization of three pathway-specific regulators for high production of monensin in Streptomyces cinnamonensis. Appl Microbiol Biotechnol 101:6083–6097
Wallace KK, Zhao B, McArthur HA, Reynolds KA (1995) In vivo analysis of straight-chain and branched-chain fatty acid biosynthesis in three actinomycetes. FEMS Microbiol Lett 131:227–234
Wilson DJ, Xue Y, Reynolds KA, Sherman DH (2001) Characterization and analysis of the PikD regulatory factor in the pikromycin biosynthetic pathway of Streptomyces venezuelae. J Bacteriol 183:3468–3475
Woo MW, Nah HJ, Choi SS, Kim ES (2014) Pikromycin production stibulation through antibiotic down-regulatory gene disruption in Streptomyces venezuelae. Biotechnol Bioprocess Eng 19:973–977
Ye BC, Zhang Y, Yu H, Yu WB, Liu BH, Yin BC, Yin CY, Li YY, Chu J, Zhang SL (2009) Time-resolved transcriptome analysis of Bacillus subtilis responding to valine, glutamate, and glutamine. PLoS ONE 4:e7073
Yi JS, Kim MS, Kim SJ, Kim BG (2015) Effects of sucrose, phosphate, and calcium carbonate on the production of pikromycin from Streptomyces venezuelae. J Microbiol Biotechnol 24:496–502
Yi JS, Kim MW, Kim M, Jeong Y, Kim EJ, Cho BK, Kim BG (2017) A novel approach for gene expression optimization through native promoter and 5′ UTR combinations based on RNA-seq, Bibo-seq, and TSS-seq of Streptomyces coelicolor. ACS Synth Biol 6:555–565
Zabala D, Brana AF, Florez AB, Salas JA, Mendez C (2013) Engineering precursor metabolite pools for increasing production of antitumor mithramycins in Streptomyces argillaceus. Metab Eng 20:187–197
Zhang YX, Denoya CD, Skinner DD, Fedechko RW, McArthur HA, Morgenstern MR, Davies RA, Lobo S, Reynolds KA, Hutchinson CR (1999) Genes encoding acyl-CoA dehydrogenase (AcdH) homologues from Streptomyces coelicolor and Streptomyces avermitilis provide insights into the metabolism of small branched-chain fatty acids and macrolide antibiotic production. Microbiology 145:2323–2334
Zhang YX, Tang L, Hutchinson CR (1996) Cloning and characterization of a gene (msdA) encoding methylmalonic acid semialdehyde dehydrogenase from Streptomyces coelicolor. J Bacteriol 178:490–495
Acknowledgements
This research was supported by the National Research Foundation of Korea (NRF) funded by the Ministry of Science, ICT & Future Planning (2016953757). The Institute of Engineering Research at Seoul National University provided research facilities for this work.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yi, J.S., Kim, M., Kim, EJ. et al. Production of pikromycin using branched chain amino acid catabolism in Streptomyces venezuelae ATCC 15439. J Ind Microbiol Biotechnol 45, 293–303 (2018). https://doi.org/10.1007/s10295-018-2024-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-018-2024-6