Abstract
Tau is a microtubule-associated protein involved in regulation of assembly and spatial organization of microtubule in neurons. However, in pathological conditions, tau monomers assemble into amyloid filaments characterized by the cross-β structures in a number of neurodegenerative diseases known as tauopathies. In this review, we summarize recent progression on the characterization of structures of tau monomer and filament, as well as the dynamic liquid droplet assembly. Our aim is to reveal how post-translational modifications, amino acid mutations, and interacting molecules modulate the conformational ensemble of tau monomer, and how they accelerate or inhibit tau assembly into aggregates. Structure-based aggregation inhibitor design is also discussed in the context of dynamics and heterogeneity of tau structures.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
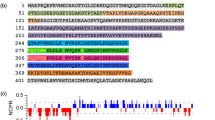
Tau protein is encoded by the MAPT gene which is located on chromosome 17. Six different tau isoforms are generated by alternative splicing, containing zero, one or two N-terminal inserts and three or four microtubule-binding repeats [1]. The constructs of 2N4R and 2N3R are illustrated in Fig. 1a. Tau protein can be divided into four distinct portions: the N-terminal domain (NTD), the proline-rich domain (PRD), the microtubule-binding domain (MTBD), and the C-terminal domain (CTD). The two aggregation-prone hexapeptide motifs, PHF6* (275VQIINK280) and PHF6 (306VQIVYK311), are located on the R2 repeat and R3 repeat of MTBD, respectively. Consequently, the 4R tau isoform contains two hexapeptide motifs, and the 3R tau isoform has only one hexapeptide motif.
Domain organization and conformation states of tau. a Illustrations of two tau isoforms, 2N4R and 2N3R. The 2N4R isoform contains two N-terminal inserts (N1 and N2), two proline-rich regions (P1 and P2), and four microtubule-binding repeats (R1, R2, R3, and R4). The repeat-like segment R’ in the C-terminus is also indicated. The microtubule-binding repeat R2 is not present in the 2N3R isoform. b Schematic illustrations of the three conformation states of tau. The disordered while compact tau monomers bind to microtubule, forming elongated conformations. Under certain conditions, tau monomers aggregate into filaments characterized by the presence of cross-β structures. The length of individual domain is not to scale
Tau is mainly expressed in central and peripheral nerve systems, where it is largely distributed in axons. As a microtubule-associated protein, tau regulates assembly and spatial organization of microtubule, thus playing a critical role in axon development and navigation (Fig. 1b) [2, 3]. Tau monomer is intrinsically disordered, with some transient secondary structure elements populated. However, tau monomers assemble into amyloid filaments characterized by the cross-β structures in a number of neurodegenerative diseases known as tauopathies, including Alzheimer’s disease (AD), Pick’s disease (PiD), chronic traumatic encephalopathy (CTE), and corticobasal degeneration (CBD) [4,5,6,7]. Consequently, tau protein is widely believed to be a major target for treatment of tauopathies [8,9,10,11]. In addition to its standard function as a microtubule regulating protein, tau has numerous binding partners and is distributed into various cell compartments. Recent studies show that tau also plays roles in signaling, cytoskeletal organization, and chromosome stability [12, 13].
In this review, we summarize recent structural characterization on tau monomer and filaments, aiming to reveal how post-translational modifications (PTMs), amino acid mutations, and interacting molecules modulate the conformational ensemble of tau monomer, and how they accelerate or inhibit tau aggregation. We pay special attention to the liquid–liquid phase separation (LLPS) of tau monomer and structure-based screening/designing of tau aggregation inhibitors.
Structure of tau monomer
Tau is a polyelectrolyte. The NTD and CTD are negatively charged, whereas the PRD and MTBD are positively charged (Fig. 2a). Disordered propensity predictions suggest that tau is highly disordered except the MTBD (Fig. 2b). Consistently, experimental characterization has shown that tau does not have well-defined three-dimensional structures [14,15,16,17,18,19,20]. Circular dichroism (CD), Fourier transform infrared spectroscopy, and nuclear magnetic resonance (NMR) have revealed that dynamic and residual secondary structures are present in tau monomer [14, 18, 21, 22]. In particular, the PHF6* and PHF6 motifs adopt β-strand conformations [20, 22, 23]. The mean radius of gyration (Rg) of tau (5.1 ± 0.5 nm from single molecule Förster resonance energy transfer (smFRET) measurement or 6.5 ± 0.3 nm from small-angle X-ray scattering (SAXS) measurement) is smaller than that of random coil of equal length [16, 17]. Therefore, tau is globally compact in solution. Further characterization shows that tau can form long-range contacts, some of which are mediated by electrostatic interactions [18, 22, 24, 25]. For example, the C-terminus folds on to the MTBD and the N-terminus is in contact with the C-terminus. Such an overall structure of tau can be described by the “paperclip-like” model [26]. Although tau is globally compact, different domains have distinct conformational characteristics and the conformation of MTBD can be altered by the flanking domains. For example, MTBD becomes more compact when it is isolated from the full-length protein [27]. Furthermore, backbone dynamics analysis revealed that the NTD is highly mobile while the MTBD shows increased rigidity [22].
Sequence and structure properties of tau monomer. a Net charge per residue (NCPR) distribution of 2N4R tau analyzed by CIDER [174]. b Disorder prediction of 2N4R tau using the PONDR VLXT [175, 176]. c Conformational ensemble of tau constructed based on cross-linking data and computational sampling [28]. Five conformations are shown by different colors. d Schematic illustration of conformation remodeling of tau
Several atomic conformational ensembles of tau monomer have been constructed based on paramagnetic relaxation enhancement, residual dipolar coupling, cross-linking data, and computational sampling, further demonstrating the globular conformations of tau with distinct topology and variable secondary-structure elements (Fig. 2c) [17, 18, 22, 25, 28]. The free energy landscape of tau monomer may have several minima separated by free energy barriers. Therefore, distinct tau conformer species can be detected and isolated. Single-molecule fluorescence anisotropy combined with an anti-Brownian electrokinetic trap has revealed that tau resides in two groups of conformations, i.e., a more compact family and a less compact family [29, 30]. In another study, after sonication treatment, two tau monomeric species were isolated from tau fibrils, i.e., the inert monomer “Mi” and the seed-competent monomer “Ms” [31]. Although CD showed no observable difference between Mi and Ms, conformation modeling using restrains from cross-linking data revealed that the PHF6* and PHF6 are buried in Mi but are relatively exposed in Ms. Furthermore, the Ms group itself is heterogeneous as Ms from AD patient brain and CBD patient brain encodes one tau prion strain and three tau prion strains, respectively [32]. Other observations also suggest that tau can reside in different conformational states, some of which are called “pathological conformation” and can be recognized by specific antibodies [33,34,35]. By monitoring the end-to-end distance distribution of PHF6 and PHF6* through pulsed double electron–electron resonance measurements, Eschmann et al. found that the extended β-strand conformational states of PHF6 and PHF6* constitute a defining signature of aggregation-prone tau [36]. Since different tau conformational states could exhibit distinct rigidity, structure propensity, and steric hindrance surrounding PHF6* and PHF6 motifs, they may possess different aggregation propensity and contribute differently to tau-related diseases.
Mutations, PTMs as well as intermolecular interactions can remodel the conformational ensemble of tau monomer (Fig. 2d), thus promoting or suppressing its aggregation. The pro-aggregant mutant ΔK280 is found to suppress long-range intermolecular contacts and stabilize the β-strand conformations [24]. The aggregation-prone PHF6 motif forms metastable compact structures with its upstream sequence and is shielded by a β-turn structure in the inert monomer state [31]. P301 mutations destabilize this local structure and trigger spontaneous aggregation [37]. Depending on the phosphorylation state, the overall conformation of tau monomer and the transient intramolecular interactions could be modified [38]. Phosphorylation mimic in the epitopes recognized by the AT8 antibody (S199E, S202E, T205E) enhances tau aggregation by reducing the electrostatic attraction between NTD and PRD or by moving NTD away from CTD [39, 40]. Phosphorylation at Ser202 and Thr205 induces formation of a turn-like structure, protecting tau against aggregation [41]. On the contrary, hyperphosphorylation by glycogen synthase kinase-3β results in global expansion of tau and increased exposure of PHF6 [38]. SmFRET characterization showed that binding of aggregation enhancers (e.g., heparin and cytoplasmic polyphosphates) eliminate long-range contacts and induce expansion of tau [17, 27, 42]. The mean Rg of tau is increased from 5.1 ± 0.5 to 6.0 ± 0.6 nm upon addition of heparin [17]. NMR titration showed that the segment at the beginning of R2 exhibits the largest chemical shift displacements [23]. Although the MTBD becomes more compact upon heparin binding [27], the PHF6 and PHF6* motifs remain extended [36]. A similar chemical shift pattern is observed when tau is bound to polyglutamic acid [23]. Binding of polyglutamic acid also compacts the MTBD and tightens the interactions between PHF6 and PHF6* [43].
In contrast to 4R tau isoform, structural investigations on 3R tau are very limited. Molecular dynamics simulation and NMR characterization have been applied to compare the structural difference between K18 (the repeat region of 4R tau) and K19 (the repeat region of 3R tau) constructs of tau [23, 25]. K18 and K19 show similar β-structure propensity at the beginning of repeats [23]. In K18, the R1 repeat interacts with R2 but not R3 or R4. In contrast, in K19, R1 interacts with both R3 and R4 [25]. Therefore, although the removal of R2 does not affect the local β-structure propensity of MTBD, it remodels the global intramolecular interactions of tau.
Liquid–liquid phase separation and structure of tau in the droplet-like assembly
When the solution conditions favor formation of massive dynamic intermolecular interactions, some intrinsically disordered proteins (IDPs) will demix into a light phase and a dense phase. Such a process has been termed as liquid–liquid phase separation, which underlies the formation of the membraneless compartments in cells [44,45,46,47,48,49]. Importantly, some IDP droplets can convert into solid states, suggesting that the liquid condensates are on pathway to fibers and LLPS can be related to the progression of some neurodegenerative diseases [50,51,52,53].
Recent studies have revealed that tau readily undergoes LLPS in the presence of macromolecular crowding agents or in cells [54,55,56,57,58]. The concentration of tau can be increased more than ten-fold upon LLPS [56, 59]. Fusion, fission, fluorescence recovery after photobleaching and electron paramagnetic resonance spectroscopy of droplets indicate that tau droplets are in liquid state [57, 60, 61].
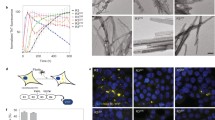
LLPS of tau is sensitive to the concentration of salt. Decreasing salt concentration promotes LLPS while increasing salt concentration suppresses droplets formation (Fig. 3a) [54, 55, 57, 62]. The sensitivity of LLPS to salt concentration suggests that electrostatic interactions are critical for the formation of tau droplets [55, 57]. The charges are clustered along the tau sequence (Fig. 2a). Truncation experiments suggest that interactions between the positively charged domains and the negatively charged domains are the main driving force of tau LLPS (Fig. 3c) [57]. Although the K18 construct of tau is also able to undergo LLPS under very high concentration, its LLPS is sensitive to 1,6-hexanediol rather than salt concentration, indicating that the main driving force of K18 LLPS is hydrophobic interactions rather than electrostatic interactions [55, 59]. Similarly, under high salt conditions where electrostatic interactions are screened, droplets formed by N-terminal truncated tau are dissolved with 1,6-hexanediol [63]. PTMs also regulate LLPS of tau (Fig. 3b). Acetylation neutralizes the positive charges on lysine residues, thereby reducing electrostatic attractions that drive intramolecular and intermolecular contacts. Consequently, acetylation suppresses LLPS of tau [64]. By introducing negatively charged groups into PRD and MTBD, phosphorylation also reduces electrostatic attractions. However, phosphorylation of tau, either introduced by MARK2 or SF9 insect cells, promotes LLPS of tau [55, 56]. Interestingly, phosphorylation reverses the sensitivity of tau LLPS to salt and 1,6-hexanediol, where LLPS of phosphorylated tau is insensitive to increasing salt concentration but sensitive to the addition of 1,6-hexanediol [56]. It seems that phosphorylation suppresses tau LLPS driven by electrostatic interactions but promotes tau LLPS driven by hydrophobic interactions. It is possible that phosphorylation modulates the conformations of PRD and MTBD and induces exposure of hydrophobic segments. Structural investigations on phosphorylated tau will be valuable for further understanding the influence of phosphorylation on tau LLPS. Taken together, these data show that tau LLPS is driven by a complex combination of electrostatic and hydrophobic interactions.
Liquid–liquid phase separation of tau. a A phase diagram illustrating the effect of salt concentration on LLPS of tau. b The mutations, acetylation sites, and phosphorylation sites that influence tau LLPS. c Intermolecular interactions that may be critical for tau LLPS, such as electrostatic interactions between the negatively charged NTD and the positively charged PRD and MTBD of tau, electrostatic interactions between tau and RNA, and metal ions mediated interactions
Tau forms droplets with various types of RNA, which may result from the complex coacervation effect [60,61,62]. Lysine/RNA and arginine/RNA interactions are critical in forming the complex coacervate phase [61]. Phosphorylation reduces the propensity of tau/RNA LLPS, likely due to electrostatic repulsion between the phosphorylated residues and the negatively charged RNA [60]. Acetylation also suppresses LLPS of tau with RNA, probably due to the removal of positive charges on lysine sidechains [61].
Metal ions play important roles in regulating the function of tau. Rane et al. showed that Al3+ and Zn2+ enhance tau LLPS [65]. Singh et al. further showed that the two cysteines within the MTBD are required for Zn2+ to induce tau droplet formation [66]. Metal ions may promote LLPS of tau through a conformational change favorable for intermolecular electrostatic attractions or mediating intermolecular contacts [65].
Structural characterization of tau droplets remains challenging. However, progress has been made. Double electron–electron resonance spectroscopy characterization indicated that the mean distance flanking the PHF6* region remains unchanged when tau undergoes LLPS with RNA [62]. However, the entire polypeptide chain of tau adopts more extended conformations in the droplet state [67]. Secondary structure characterization of tau K18 via CD and NMR indicated that the level of β-structure content and the propensity for β-hairpin conformation are increased upon LLPS [55, 59]. Although the thioflavin T fluorescence of tau droplets gradually increases over incubation, its intensity is much weaker than what is observed in the presence of heparin [55, 62]. Therefore, the β-structure content of tau in the droplets is much smaller than that in the amyloid fibrils.
Whether LLPS of tau is related to fibril formation remains controversial. Some studies suggest that LLPS of tau mediates and facilitates aggregation. Aggregation enhancing factors, including heparin, pro-aggregation mutations, and K274 acetylation, promote LLPS of tau [56, 64, 68]. Protein disulfide isomerase directly interacts with tau and suppresses the formation of tau droplets and aggregates [69]. EFhd2 is associated with aggregated tau species in AD brains. Recent results showed that EFhd2 regulates tau aggregation and LLPS in a calcium dependent manner [58]. Furthermore, droplets formed by phosphorylated tau rapidly transition from a liquid state to a gel-like state, and finally transition into large aggregates containing β-structures [56]. Therefore, tau concentrated in the droplets is postulated to be on pathway to fibril formation. On the contrary, other studies point out that LLPS and amyloid aggregation of tau are independent processes although they occur in overlapping conditions [63]. Lin et al. systematically investigated the impacts of LLPS on tau aggregation by evaluating the conformation of tau, kinetics of aggregation and fibril quantity [63]. They found that none of these properties are influenced directly by LLPS. The presence of extended β-strand conformation of PHF6 and PHF6* has been postulated as a defining signature of aggregation-prone tau [36]. However, structural investigation discussed above shows that the β-structure content of PHF6 and PHF6* in the droplets is almost indistinguishable from that in the dilute state. Though LLPS may not directly promote aggregation of tau, Kanaan et al. showed that phase separation of tau could facilitate the formation of non-filamentous pathogenic tau oligomers in vitro [70]. It is noted that all tau phase separation studies have been carried out in vitro or in cells. So far, no study shows a direct connection between LLPS of tau and neurodegenerative diseases in vivo or tau undergoing LLPS in neurons in situ in brain. Taken together, LLPS of tau is promoted by a variety of factors. Tau concentration is increased inside the droplets and tau molecules adopt conformations that are slightly different from those in dilute state. The connection between LLPS and tau aggregation remains elusive and requires further investigation.
Structures of soluble tau oligomers
Soluble tau oligomers with various molecular weights have been identified and they have been suggested as the toxic species in vivo [35, 71,72,73,74,75,76,77,78,79,80,81,82,83,84,85,86]. Once formed, tau oligomers can be released and taken up by cells [87]. The conformations of oligomeric tau could be different from those of monomeric tau as monoclonal antibodies raised against tau oligomers show no reactivity toward monomeric tau [73, 88, 89]. Since some tau oligomers can be converted to amyloid fibrils, hydrophobic segments in the MTBD may become exposed upon oligomerization [31, 75, 90]. While some studies show that tau dimers can be stabilized by intermolecular disulfide bonds [84, 85], non-disulfide linked tau dimers are also observed [91,92,93]. Via force measurement, Rosenberg et al. showed that two tau monomers can associate in an antiparallel configuration stabilized by complementary electrostatic interactions between the negatively-charged NTDs and the positively-charged PRDs [91]. Moreover, a tau dimer can be formed via bridging by a heparin molecule [72, 94, 95]. Although tau oligomers have been recognized as important players in tau pathogenesis for a long time, structural characterization on tau oligomers is very limited and requires more studies in the future.
Structures of tau filaments and aggregation mechanism
It’s challenging to characterize the structures of tau aggregates because they are partially disordered and heterogeneous. Nevertheless, accumulated evidence from studies using various constructs of recombinant tau shows that the amyloid core is dominantly formed by the MTBD, ranging from the second half of R1 to the first half of R4 [96]. Jakes et al. further revealed that the core of paired helical filament (PHF) in AD is restricted to the repeat regions of 3R and 4R tau isoforms [97]. The repeats pack against each other parallel and form β-sheets [98,99,100]. Except the amyloid core, the remainder of tau protein remains highly mobile, forming the fuzzy coat [4, 20, 101,102,103].
Since six tau isoforms are expressed in adult human brain, different neurodegenerative diseases can have different tau isoform compositions and filament structures [104, 105]. Limited proteolysis applied on aggregates extracted from patient brains shows different banding patterns in immunoblot analyses for PiD, AD, CBD, and progressive supranuclear palsy [106]. However, the atomic structures of tau fibrils in pathological conditions are not available until the last 3 years when structures of tau aggregates from various sources were determined by cryo-EM [107,108,109,110,111].
In AD, tau filaments are made of PHFs and straight filaments (SFs), where all six tau isoforms are present [107]. Early electron microscopy images indicated that PHFs and SFs have a common C-shaped morphology [112, 113]. Recently, atomic cryo-EM structures revealed that the filament core is made of amino acids V306-F378, comprising R3 and R4, as well as part of the C-terminal domain (Fig. 4a) [107]. Similar to AD, all six tau isoforms assemble into filaments in CTE, and residues K274-R379 of 3R tau and S305-R379 of 4R tau form the enlarged C-shaped filament core (Fig. 4b) [110]. Importantly, a hydrophobic cavity which is not present in the AD filament is observed within the filament core from CTE. Hydrophobic cofactors may be incorporated during tau aggregation in CTE. In PiD, two filament forms (narrow and wide) have been identified [109]. Narrow filaments are composed of a single protofilament while wide filaments are made of two narrow filaments. Different from the C-shape core of AD, the filament core of PiD is an elongated structure, which comprises amino acids K254-F378 of 3R tau (Fig. 4c). The latest tau filament structure is from CBD [111]. The CBD filaments are made of 4R tau exclusively. Two types of filaments are observed depending on the numbers of protofilaments. The CBD filament core comprises K274-E380 and adopts a four-layered fold (Fig. 4d). An additional uncharacterized density surrounded by the sidechains of K290, K294, and K370 is found within the filament core.
Structure models of tau from various filaments. a 3R/4R tau filaments from AD patients. b 3R/4R tau filaments from CTE patients. c 3R tau filaments from PiD patients. d 4R tau filaments from CBD patients. e Heparin induced 2N4R tau filaments. f Heparin induced 0N4R tau filaments. Additional uncharacterized density found within the filament core of the CTE and CBD folds are indicated the by the black dots
The structures of fibers derived from different patients are identical, indicating that they are disease- rather than patient-specific. So far, it remains unknown what factors drive tau into specific conformations in tauopathies. PTMs, mutations, and cofactors may be critical. The structural heterogeneity of tau filaments suggests that different factors or tau segments may play different roles in the aggregation process. On one hand, the cellular environment is much more complicated than what we can mimic in a test tube. It is not surprising that the structures of heparin-induced tau filaments are different from those in diseases (Fig. 4e, f) [114, 115]. On the other hand, the marked difference in the structures of tau filaments suggests the existence of distinct seeds of tau in different tauopathies [116,117,118]. Since stable seed-competent tau monomers can be isolated from patient brains [32], it is important to determine whether these seeds will serve as templates to drive tau assembling into filaments identical to those observed in patients.
In vitro, the aggregation process of tau is generally described by the classic nucleation-elongation model, with the nucleating species thought to be the assembly-competent monomer [119] or soluble oligomer [75, 85]. Recent studies have demonstrated that monomeric tau species derived from heparin-induced aggregates or isolated from patient brains are capable of seeding tau aggregation both in vitro and in cultured cells [31, 32]. Through a photochemical cross-linking technique, Patterson et al. demonstrated that dimerization is an early step in the aggregation process of tau and these dimers self-associate to form larger aggregates [73]. Tau aggregation induced by heparin further suggests that the aggregation-competent tau species is a tau dimer which may be bridged by a heparin molecule [72, 94, 95]. Exposure of hydrophobic segments in the assembly-competent monomer and dimer may be critical for the subsequent self-assembling process [31, 75].
Tau is stable under normal conditions. However, factors inducing formation of oligomers or enhancing exposure of hydrophobic segments will promote tau aggregation. Previous analysis has revealed that tau is often mutated or hyperphosphorylated in tauopathies, suggesting that mutation or phosphorylation regulates tau aggregation. Indeed, mutations found in frontotemporal dementias promote tau aggregation [19, 120,121,122,123]. Although these mutations generate marginal change on the overall structure of tau monomer, they enhance local β-structure propensity [19, 124]. On the contrary, proline mutations within the two hexapeptide motifs (I277P and I308P) disrupt β-structure and abrogate aggregation [20]. The influence of phosphorylation on tau aggregation is also complicated. The abnormally hyperphosphorylated tau from AD brain or tau phosphorylated by glycogen synthase kinase-3β has been found to self-aggregate into PHF-like structures [125, 126]. Specific phosphorylation patterns promote tau aggregation can be recognized [41, 127]. However, hyperphosphorylated tau obtained by in vitro phosphorylation with recombinant extracellular-regulated kinase or rat brain extract, or obtained from recombinant expression in Sf9 cells shows no significant increased susceptibility to in vitro aggregation than unphosphorylated tau [83, 128]. Furthermore, phosphorylation has also been found to protect tau against aggregation [129, 130]. Importantly, kinetics studies suggest that even when phosphorylated tau is aggregated, phosphorylation enhances but not triggers tau aggregation [131, 132].
Interactions with surrounding molecules can modulate the conformational ensemble of tau monomer or shift the equilibrium between tau monomer and oligomer. The tau fibril core is mainly made of the positively charged MTBD which interacts with various polyanions, including DNA, RNA, heparin, polyphosphates and polyglutamic acid. Structural and kinetic characterization suggests that polyanions enhance tau aggregation by remodeling the conformational ensemble of monomeric tau and noncovalent cross-linking of multiple tau monomers [42, 43, 94, 95, 133, 134]. Tau contains multiple metal ion binding sites. Binding to metal ions has been showed to enhance tau aggregation although the mechanism remains unclear [9, 65, 135,136,137]. Furthermore, crowded cell-like environments can significantly promote tau aggregation by accelerating the nucleation step [138,139,140].
Structure-based design of tau aggregation inhibitors
Tau aggregation can be inhibited by binding to various molecules. Molecular chaperones suppress tau aggregation efficiently. Hsp70 suppresses the formation of tau nuclei [141]. Hsp27 delays tau fibril formation by weakly interacting with early species in the aggregation process, whereas HspA8 is highly efficient at preventing tau fibril elongation, possibly by capping the ends of tau fibrils [142]. A number of chaperones bind tau at or around the PHF6* and PHF6 motifs [143,144,145]. Thus, a major mechanism of anti-aggregation activity of molecular chaperones seems to be the direct binding to tau at the aggregation-prone regions [144]. Antibodies and protein disulfide isomerase may also adopt similar mechanism to inhibit tau aggregation [69, 146].
Small molecules can inhibit tau aggregation, although the mechanism remains elusive [9, 147,148,149,150,151,152,153,154,155,156,157,158]. Based on the K18 conformational ensembles, Kiss et al. analyzed the potential hot spots and small molecule binding sites using FTMap [159]. They found that the PHF6 and PHF6* motifs have the highest probability of forming the hot spots. Chong et al. also identified nine druggable cavities from the K18 conformational ensembles [160]. Docking of methylene blue with various tau conformations revealed that methylene blue binds in close proximity of C291/C322 [159] and NMR characterization showed that the molecular tweezer CLR01 binds preferentially to Lys residues in the MTBD [161]. Recently, through molecular dynamics simulations and ensemble docking, Baggett and Nath identified novel tau aggregation inhibitors [162]. Since the structure of tau monomer is highly dynamic, small molecules may bind to tau in a fuzzy way [163, 164].
PHF6 and PHF6* are critical for tau aggregation. Tau molecules lacking these two hexapeptide motifs cannot aggregate [165]. Consequently, it is possible to block tau aggregation by shielding these two motifs. Based on the atomic structures of amyloid fibrils formed by PHF6 and PHF6*, molecules have been designed to cap the ends of tau fibrils and they are found to efficiently inhibit the aggregation of 3R and 4R tau isoforms [166,167,168,169,170].
Conclusions and perspectives
Tau is a major target for tauopathies treatment, the structures of whose monomer and fiber have been studied for decades. It turns out that the conformational ensemble of tau monomer is very dynamic and can be remodeled by a variety of factors. Up to date, atomic models of unmodified free tau monomer are available. For better understanding the conversion of tau from inert state to aggregation-prone state, it is urgent to determine the structures of tau monomer upon phosphorylation, acetylation, or binding to other molecules.
As indicated above, the cryo-EM structures of tau fibers in distinct diseases are different. It remains unclear what factors induce or determine the heterogeneity of tau fiber structures. It is noted that the PRD is absent in the cores of available fibril structures. However, PRD is subjected to extensive PTMs. One possibility is that PTMs on PRD induce formation of various aggregation-prone tau species, which act as templates in the subsequent aggregation process. Furthermore, unknown densities are present in the cryo-EM structures of tau fibers. Clarifying their identities will be also valuable for understanding the heterogeneity of tau structures in the future.
Inhibiting the aggregation of tau has been widely accepted as a therapeutic strategy for tauopathies. It is appearing that inhibitors can be designed to bind tau monomers to block their seeding or to cap tau fibrils to block their propagation. The distinct structures of filaments from different tauopathies and the difference between structures of heparin-induced tau aggregates and those of filaments isolated from diseased brains indicate the complexity of tau assembling process. Reconstruction of disease specific tau filaments will be valuable to test the efficacy of inhibitors in this regard. The LLPS of tau seems to be related to tau aggregation. Therefore, molecules designed to suppress the formation of tau droplets may be also able to inhibit tau aggregation. LMTX is a potent tau aggregation inhibitor. While LMTX failed to show effects on the primary cognitive endpoints in two phase 3 trials [171, 172], pharmacological activity has been demonstrated on brain structure and function at the 8 mg/day dose [173]. Further clinical trials in mild/moderate AD will be required to confirm efficacy at this dose. Due to the dynamics of tau monomer, heterogeneity of fibril structures, and enrichment of PTMs, a variety of tau aggregation inhibitors are expected to be designed in the future.
References
Goedert M, Spillantini MG, Potier MC, Ulrich J, Crowther RA (1989) Cloning and sequencing of the cDNA encoding an isoform of microtubule-associated protein tau containing four tandem repeats: differential expression of tau protein mRNAs in human brain. EMBO J 8:393–399
Guo T, Noble W, Hanger DP (2017) Roles of tau protein in health and disease. Acta Neuropathol 133:665–704. https://doi.org/10.1007/s00401-017-1707-9
Sayas CL, Tortosa E, Bollati F, Ramirez-Rios S, Arnal I, Avila J (2015) Tau regulates the localization and function of End-binding proteins 1 and 3 in developing neuronal cells. J Neurochem 133:653–667. https://doi.org/10.1111/jnc.13091
Berriman J, Serpell LC, Oberg KA, Fink AL, Goedert M, Crowther RA (2003) Tau filaments from human brain and from in vitro assembly of recombinant protein show cross-beta structure. Proc Natl Acad Sci USA 100:9034–9038. https://doi.org/10.1073/pnas.1530287100
Ballatore C, Lee VM, Trojanowski JQ (2007) Tau-mediated neurodegeneration in Alzheimer’s disease and related disorders. Nat Rev Neurosci 8:663–672. https://doi.org/10.1038/nrn2194
Iqbal K, Liu F, Gong CX (2016) Tau and neurodegenerative disease: the story so far. Nat Rev Neurol 12:15–27. https://doi.org/10.1038/nrneurol.2015.225
Lee G, Leugers CJ (2012) Tau and tauopathies. Prog Mol Biol Transl Sci 107:263–293. https://doi.org/10.1016/B978-0-12-385883-2.00004-7
Li C, Gotz J (2017) Tau-based therapies in neurodegeneration: opportunities and challenges. Nat Rev Drug Discov 16:863–883. https://doi.org/10.1038/nrd.2017.155
Medina M (2018) An overview on the clinical development of tau-based therapeutics. Int J Mol Sci 19:1160. https://doi.org/10.3390/ijms19041160
Long JM, Holtzman DM (2019) Alzheimer disease: an update on pathobiology and treatment strategies. Cell 179:312–339. https://doi.org/10.1016/j.cell.2019.09.001
Chong FP, Ng KY, Koh RY, Chye SM (2018) Tau proteins and tauopathies in Alzheimer’s disease. Cell Mol Neurobiol 38:965–980. https://doi.org/10.1007/s10571-017-0574-1
Morris M, Maeda S, Vossel K, Mucke L (2011) The many faces of tau. Neuron 70:410–426. https://doi.org/10.1016/j.neuron.2011.04.009
Sotiropoulos I, Galas MC, Silva JM, Skoulakis E, Wegmann S, Maina MB, Blum D, Sayas CL et al (2017) Atypical, non-standard functions of the microtubule associated Tau protein. Acta Neuropathol Commun 5:91. https://doi.org/10.1186/s40478-017-0489-6
Schweers O, Schönbrunn-Hanebeck E, Marx A, Mandelkow E (1994) Structural studies of tau protein and Alzheimer paired helical filaments show no evidence for b-structure. J Biol Chem 269:24290–24297
Smet C, Leroy A, Sillen A, Wieruszeski JM, Landrieu I, Lippens G (2004) Accepting its random coil nature allows a partial NMR assignment of the neuronal Tau protein. ChemBioChem 5:1639–1646. https://doi.org/10.1002/cbic.200400145
Mylonas E, Hascher A, Bernado P, Blackledge M, Mandelkow E, Svergun DI (2008) Domain conformation of tau protein studied by solution small-angle X-ray scattering. Biochemistry 47:10345–10353. https://doi.org/10.1021/bi800900d
Nath A, Sammalkorpi M, DeWitt DC, Trexler AJ, Elbaum-Garfinkle S, O’Hern CS, Rhoades E (2012) The conformational ensembles of alpha-synuclein and tau: combining single-molecule FRET and simulations. Biophys J 103:1940–1949. https://doi.org/10.1016/j.bpj.2012.09.032
Schwalbe M, Ozenne V, Bibow S, Jaremko M, Jaremko L, Gajda M, Jensen MR, Biernat J et al (2014) Predictive atomic resolution descriptions of intrinsically disordered hTau40 and alpha-synuclein in solution from NMR and small angle scattering. Structure 22:238–249. https://doi.org/10.1016/j.str.2013.10.020
von Bergen M, Barghorn S, Li L, Marx A, Biernat J, Mandelkow EM, Mandelkow E (2001) Mutations of tau protein in frontotemporal dementia promote aggregation of paired helical filaments by enhancing local beta-structure. J Biol Chem 276:48165–48174. https://doi.org/10.1074/jbc.M105196200
von Bergen M, Friedhoff P, Biernat J, Heberle J, Mandelkow EM, Mandelkow E (2000) Assembly of t protein into Alzheimer paired helical filaments depends on a local sequence motif (306VQIVYK311) forming b structure. Proc Natl Acad Sci USA 97:5129–5134
Mukrasch MD, Markwick P, Biernat J, Bergen M, Bernado P, Griesinger C, Mandelkow E, Zweckstetter M et al (2007) Highly populated turn conformations in natively unfolded tau protein identified from residual dipolar couplings and molecular simulation. J Am Chem Soc 129:5235–5243. https://doi.org/10.1021/ja0690159
Mukrasch MD, Bibow S, Korukottu J, Jeganathan S, Biernat J, Griesinger C, Mandelkow E, Zweckstetter M (2009) Structural polymorphism of 441-residue tau at single residue resolution. PLoS Biol 7:e34. https://doi.org/10.1371/journal.pbio.1000034
Mukrasch MD, Biernat J, von Bergen M, Griesinger C, Mandelkow E, Zweckstetter M (2005) Sites of tau important for aggregation populate {beta}-structure and bind to microtubules and polyanions. J Biol Chem 280:24978–24986. https://doi.org/10.1074/jbc.M501565200
Wegmann S, Scholer J, Bippes CA, Mandelkow E, Muller DJ (2011) Competing interactions stabilize pro- and anti-aggregant conformations of human tau. J Biol Chem 286:20512–20524. https://doi.org/10.1074/jbc.M111.237875
Luo Y, Ma BY, Nussinov R, Wei GH (2014) Structural insight into tau protein’s paradox of intrinsically disordered behavior, self-acetylation activity, and aggregation. J Phys Chem Lett 5:3026–3031. https://doi.org/10.1021/jz501457f
Jeganathan S, von Bergen M, Brutlach H, Steinhoff HJ, Mandelkow E (2006) Global hairpin folding of tau in solution. Biochemistry 45:2283–2293. https://doi.org/10.1021/bi0521543
Elbaum-Garfinkle S, Rhoades E (2012) Identification of an aggregation-prone structure of tau. J Am Chem Soc 134:16607–16613. https://doi.org/10.1021/ja305206m
Popov KI, Makepeace KAT, Petrotchenko EV, Dokholyan NV, Borchers CH (2019) Insight into the structure of the “unstructured” tau protein. Structure 27:1710–1715. https://doi.org/10.1016/j.str.2019.09.003
Manger LH, Foote AK, Wood SL, Holden MR, Heylman KD, Margittai M, Goldsmith RH (2017) Revealing conformational variants of solution-phase intrinsically disordered tau protein at the single-molecule level. Angew Chem Int Ed Engl 56:15584–15588. https://doi.org/10.1002/anie.201708242
Foote AK, Manger LH, Holden MR, Margittai M, Goldsmith RH (2019) Time-resolved multirotational dynamics of single solution-phase tau proteins reveals details of conformational variation. Phys Chem Chem Phys 21:1863–1871. https://doi.org/10.1039/c8cp06971a
Mirbaha H, Chen D, Morazova OA, Ruff KM, Sharma AM, Liu X, Goodarzi M, Pappu RV et al (2018) Inert and seed-competent tau monomers suggest structural origins of aggregation. Elife 7:e36584. https://doi.org/10.7554/eLife.36584
Sharma AM, Thomas TL, Woodard DR, Kashmer OM, Diamond MI (2018) Tau monomer encodes strains. Elife 7:e37813. https://doi.org/10.7554/eLife.37813
Carmel G, Mager EM, Binder LI, Kuret J (1996) The structural basis of monoclonal antibody Alz50’s selectivity for Alzheimer’s disease pathology. J Biol Chem 271:32789–32795. https://doi.org/10.1074/jbc.271.51.32789
Jicha GA, Bowser R, Kazam IG, Davies P (1997) Alz-50 and MC-1, a new monoclonal antibody raised to paired helical filaments, recognize conformational epitopes on recombinant tau. J Neurosci Res 48:128–132. https://doi.org/10.1002/(sici)1097-4547(19970415)48:2%3c128::aid-jnr5%3e3.0.co;2-e
Ward SM, Himmelstein DS, Lancia JK, Binder LI (2012) Tau oligomers and tau toxicity in neurodegenerative disease. Biochem Soc Trans 40:667–671. https://doi.org/10.1042/BST20120134
Eschmann NA, Georgieva ER, Ganguly P, Borbat PP, Rappaport MD, Akdogan Y, Freed JH, Shea JE et al (2017) Signature of an aggregation-prone conformation of tau. Sci Rep 7:44739. https://doi.org/10.1038/srep44739
Chen D, Drombosky KW, Hou Z, Sari L, Kashmer OM, Ryder BD, Perez VA, Woodard DR et al (2019) Tau local structure shields an amyloid-forming motif and controls aggregation propensity. Nat Commun 10:2493. https://doi.org/10.1038/s41467-019-10355-1
Zhu S, Shala A, Bezginov A, Sljoka A, Audette G, Wilson DJ (2015) Hyperphosphorylation of intrinsically disordered tau protein induces an amyloidogenic shift in its conformational ensemble. PLoS ONE 10:e0120416. https://doi.org/10.1371/journal.pone.0120416
Jeganathan S, Hascher A, Chinnathambi S, Biernat J, Mandelkow EM, Mandelkow E (2008) Proline-directed pseudo-phosphorylation at AT8 and PHF1 epitopes induces a compaction of the paperclip folding of tau and generates a pathological (MC-1) conformation. J Biol Chem 283:32066–32076. https://doi.org/10.1074/jbc.M805300200
Bibow S, Ozenne V, Biernat J, Blackledge M, Mandelkow E, Zweckstetter M (2011) Structural impact of proline-directed pseudophosphorylation at AT8, AT100, and PHF1 epitopes on 441-residue tau. J Am Chem Soc 133:15842–15845. https://doi.org/10.1021/ja205836j
Despres C, Byrne C, Qi H, Cantrelle FX, Huvent I, Chambraud B, Baulieu EE, Jacquot Y et al (2017) Identification of the tau phosphorylation pattern that drives its aggregation. Proc Natl Acad Sci USA 114:9080–9085. https://doi.org/10.1073/pnas.1708448114
Wickramasinghe SP, Lempart J, Merens HE, Murphy J, Huettemann P, Jakob U, Rhoades E (2019) Polyphosphate initiates tau aggregation through intra- and intermolecular scaffolding. Biophys J 117:717–728. https://doi.org/10.1016/j.bpj.2019.07.028
Akoury E, Mukrasch MD, Biernat J, Tepper K, Ozenne V, Mandelkow E, Blackledge M, Zweckstetter M (2016) Remodeling of the conformational ensemble of the repeat domain of tau by an aggregation enhancer. Protein Sci 25:1010–1020. https://doi.org/10.1002/pro.2911
Banani SF, Lee HO, Hyman AA, Rosen MK (2017) Biomolecular condensates: organizers of cellular biochemistry. Nat Rev Mol Cell Biol 18:285–298. https://doi.org/10.1038/nrm.2017.7
Wu H, Fuxreiter M (2016) The Structure and dynamics of higher-order assemblies: amyloids, signalosomes, and granules. Cell 165:1055–1066. https://doi.org/10.1016/j.cell.2016.05.004
Boeynaems S, Alberti S, Fawzi NL, Mittag T, Polymenidou M, Rousseau F, Schymkowitz J, Shorter J et al (2018) Protein phase separation: a new phase in cell biology. Trends Cell Biol 28:420–435. https://doi.org/10.1016/j.tcb.2018.02.004
Feng Z, Chen X, Wu X, Zhang M (2019) Formation of biological condensates via phase separation: characteristics, analytical methods, and physiological implications. J Biol Chem 294:14823–14835. https://doi.org/10.1074/jbc.REV119.007895
Cramer P (2019) Organization and regulation of gene transcription. Nature 573:45–54. https://doi.org/10.1038/s41586-019-1517-4
Rhine K, Vidaurre V, Myong S (2020) RNA droplets. Annu Rev Biophys 49:247–265. https://doi.org/10.1146/annurev-biophys-052118-115508
Aguzzi A, Altmeyer M (2016) Phase separation: linking cellular compartmentalization to disease. Trends Cell Biol 26:547–558. https://doi.org/10.1016/j.tcb.2016.03.004
Shin Y, Brangwynne CP (2017) Liquid phase condensation in cell physiology and disease. Science 357:eaaf4382. https://doi.org/10.1126/science.aaf4382
Alberti S, Dormann D (2019) Liquid-liquid phase separation in disease. Annu Rev Genet 53:171–194. https://doi.org/10.1146/annurev-genet-112618-043527
de Oliveira GAP, Cordeiro Y, Silva JL, Vieira T (2019) Liquid-liquid phase transitions and amyloid aggregation in proteins related to cancer and neurodegenerative diseases. Adv Protein Chem Struct Biol 118:289–331. https://doi.org/10.1016/bs.apcsb.2019.08.002
Hernandez-Vega A, Braun M, Scharrel L, Jahnel M, Wegmann S, Hyman BT, Alberti S, Diez S et al (2017) Local nucleation of microtubule bundles through tubulin concentration into a condensed tau phase. Cell Rep 20:2304–2312. https://doi.org/10.1016/j.celrep.2017.08.042
Ambadipudi S, Biernat J, Riedel D, Mandelkow E, Zweckstetter M (2017) Liquid-liquid phase separation of the microtubule-binding repeats of the Alzheimer-related protein tau. Nat Commun 8:275. https://doi.org/10.1038/s41467-017-00480-0
Wegmann S, Eftekharzadeh B, Tepper K, Zoltowska KM, Bennett RE, Dujardin S, Laskowski PR, MacKenzie D et al (2018) Tau protein liquid-liquid phase separation can initiate tau aggregation. EMBO J 37:e98049. https://doi.org/10.15252/embj.201798049
Boyko S, Qi X, Chen TH, Surewicz K, Surewicz WK (2019) Liquid-liquid phase separation of tau protein: the crucial role of electrostatic interactions. J Biol Chem 294:11054–11059. https://doi.org/10.1074/jbc.AC119.009198
Vega IE, Umstead A, Kanaan NM (2019) EFhd2 affects tau liquid-liquid phase separation. Front Neurosci 13:845. https://doi.org/10.3389/fnins.2019.00845
Ambadipudi S, Reddy JG, Biernat J, Mandelkow E, Zweckstetter M (2019) Residue-specific identification of phase separation hot spots of Alzheimer’s-related protein tau. Chem Sci 10:6503–6507. https://doi.org/10.1039/c9sc00531e
Lin Y, McCarty J, Rauch JN, Delaney KT, Kosik KS, Fredrickson GH, Shea JE, Han S (2019) Narrow equilibrium window for complex coacervation of tau and RNA under cellular conditions. Elife 8:e42571. https://doi.org/10.7554/eLife.42571
Ukmar-Godec T, Hutten S, Grieshop MP, Rezaei-Ghaleh N, Cima-Omori MS, Biernat J, Mandelkow E, Soding J et al (2019) Lysine/RNA-interactions drive and regulate biomolecular condensation. Nat Commun 10:2909. https://doi.org/10.1038/s41467-019-10792-y
Zhang X, Lin Y, Eschmann NA, Zhou H, Rauch JN, Hernandez I, Guzman E, Kosik KS et al (2017) RNA stores tau reversibly in complex coacervates. PLoS Biol 15:e2002183. https://doi.org/10.1371/journal.pbio.2002183
Lin Y, Fichou Y, Zeng Z, Hu NY, Han S (2020) Electrostatically driven complex coacervation and amyloid aggregation of tau are independent processes with overlapping conditions. ACS Chem Neurosci 11:615–627. https://doi.org/10.1021/acschemneuro.9b00627
Ferreon JC, Jain A, Choi KJ, Tsoi PS, MacKenzie KR, Jung SY, Ferreon AC (2018) Acetylation disfavors tau phase separation. Int J Mol Sci 19:1360. https://doi.org/10.3390/ijms19051360
Rane JS, Kumari A, Panda D (2020) The acetyl mimicking mutation, K274Q in tau, enhances the metal binding affinity of tau and reduces the ability of tau to protect DNA. ACS Chem Neurosci 11:291–303. https://doi.org/10.1021/acschemneuro.9b00455
Singh V, Xu L, Boyko S, Surewicz K, Surewicz WK (2020) Zinc promotes liquid-liquid phase separation of tau protein. J Biol Chem 295:5850–5856. https://doi.org/10.1074/jbc.AC120.013166
Majumdar A, Dogra P, Maity S, Mukhopadhyay S (2019) Liquid-liquid phase separation is driven by large-scale conformational unwinding and fluctuations of intrinsically disordered protein molecules. J Phys Chem Lett 10:3929–3936. https://doi.org/10.1021/acs.jpclett.9b01731
Rane JS, Kumari A, Panda D (2019) An acetylation mimicking mutation, K274Q, in tau imparts neurotoxicity by enhancing tau aggregation and inhibiting tubulin polymerization. Biochem J 476:1401–1417. https://doi.org/10.1042/BCJ20190042
Wang K, Liu JQ, Zhong T, Liu XL, Zeng Y, Qiao X, Xie T, Chen Y et al (2020) Phase separation and cytotoxicity of tau are modulated by protein disulfide isomerase and s-nitrosylation of this molecular chaperone. J Mol Biol 432:2141–2163. https://doi.org/10.1016/j.jmb.2020.02.013
Kanaan NM, Hamel C, Grabinski T, Combs B (2020) Liquid-liquid phase separation induces pathogenic tau conformations in vitro. Nat Commun 11:2809. https://doi.org/10.1038/s41467-020-16580-3
Wille H, Drewes G, Biernat J, Mandelkow EM, Mandelkow E (1992) Alzheimer-like paired helical filaments and antiparallel dimers formed from microtubule-associated protein tau in vitro. J Cell Biol 118:573–584. https://doi.org/10.1083/jcb.118.3.573
Sahara N, Maeda S, Murayama M, Suzuki T, Dohmae N, Yen SH, Takashima A (2007) Assembly of two distinct dimers and higher-order oligomers from full-length tau. Eur J Neurosci 25:3020–3029. https://doi.org/10.1111/j.1460-9568.2007.05555.x
Patterson KR, Remmers C, Fu Y, Brooker S, Kanaan NM, Vana L, Ward S, Reyes JF et al (2011) Characterization of prefibrillar tau oligomers in vitro and in Alzheimer disease. J Biol Chem 286:23063–23076. https://doi.org/10.1074/jbc.M111.237974
Makrides V, Shen TE, Bhatia R, Smith BL, Thimm J, Lal R, Feinstein SC (2003) Microtubule-dependent oligomerization of tau. Implications for physiological tau function and tauopathies. J Biol Chem 278:33298–33304. https://doi.org/10.1074/jbc.M305207200
Lasagna-Reeves CA, Castillo-Carranza DL, Guerrero-Muoz MJ, Jackson GR, Kayed R (2010) Preparation and characterization of neurotoxic tau oligomers. Biochemistry 49:10039–10041. https://doi.org/10.1021/bi1016233
Berger Z, Roder H, Hanna A, Carlson A, Rangachari V, Yue M, Wszolek Z, Ashe K et al (2007) Accumulation of pathological tau species and memory loss in a conditional model of tauopathy. J Neurosci 27:3650–3662. https://doi.org/10.1523/JNEUROSCI.0587-07.2007
Tian H, Davidowitz E, Lopez P, Emadi S, Moe J, Sierks M (2013) Trimeric tau is toxic to human neuronal cells at low nanomolar concentrations. Int J Cell Biol 2013:260787. https://doi.org/10.1155/2013/260787
Ren Y, Sahara N (2013) Characteristics of tau oligomers Front Neurol 4:102. https://doi.org/10.3389/fneur.2013.00102
Ait-Bouziad N, Lv G, Mahul-Mellier AL, Xiao S, Zorludemir G, Eliezer D, Walz T, Lashuel HA (2017) Discovery and characterization of stable and toxic tau/phospholipid oligomeric complexes. Nat Commun 8:1678. https://doi.org/10.1038/s41467-017-01575-4
Maeda S, Takashima A (2019) Tau oligomers. Adv Exp Med Biol 1184:373–380. https://doi.org/10.1007/978-981-32-9358-8_27
Kaniyappan S, Chandupatla RR, Mandelkow EM, Mandelkow E (2017) Extracellular low-n oligomers of tau cause selective synaptotoxicity without affecting cell viability. Alzheimers Dement 13:1270–1291. https://doi.org/10.1016/j.jalz.2017.04.002
Lasagna-Reeves CA, Castillo-Carranza DL, Sengupta U, Clos AL, Jackson GR, Kayed R (2011) Tau oligomers impair memory and induce synaptic and mitochondrial dysfunction in wild-type mice. Mol Neurodegener 6:39. https://doi.org/10.1186/1750-1326-6-39
Tepper K, Biernat J, Kumar S, Wegmann S, Timm T, Hubschmann S, Redecke L, Mandelkow EM et al (2014) Oligomer formation of tau protein hyperphosphorylated in cells. J Biol Chem 289:34389–34407. https://doi.org/10.1074/jbc.M114.611368
Kim D, Lim S, Haque MM, Ryoo N, Hong HS, Rhim H, Lee DE, Chang YT et al (2015) Identification of disulfide cross-linked tau dimer responsible for tau propagation. Sci Rep 5:15231. https://doi.org/10.1038/srep15231
Friedhoff P, von Bergen M, Mandelkow EM, Davies P, Mandelkow E (1998) A nucleated assembly mechanism of Alzheimer paired helical filaments. Proc Natl Acad Sci USA 95:15712–15717. https://doi.org/10.1073/pnas.95.26.15712
Cowan CM, Mudher A (2013) Are tau aggregates toxic or protective in tauopathies? Front Neurol 4:114. https://doi.org/10.3389/fneur.2013.00114
Wegmann S, Nicholls S, Takeda S, Fan Z, Hyman BT (2016) Formation, release, and internalization of stable tau oligomers in cells. J Neurochem 139:1163–1174. https://doi.org/10.1111/jnc.13866
Castillo-Carranza DL, Sengupta U, Guerrero-Munoz MJ, Lasagna-Reeves CA, Gerson JE, Singh G, Estes DM, Barrett AD et al (2014) Passive immunization with Tau oligomer monoclonal antibody reverses tauopathy phenotypes without affecting hyperphosphorylated neurofibrillary tangles. J Neurosci 34:4260–4272. https://doi.org/10.1523/JNEUROSCI.3192-13.2014
Lasagna-Reeves CA, Castillo-Carranza DL, Sengupta U, Sarmiento J, Troncoso J, Jackson GR, Kayed R (2012) Identification of oligomers at early stages of tau aggregation in Alzheimer’s disease. FASEB J 26:1946–1959. https://doi.org/10.1096/fj.11-199851
Maeda S, Sahara N, Saito Y, Murayama M, Yoshiike Y, Kim H, Miyasaka T, Murayama S et al (2007) Granular tau oligomers as intermediates of tau filaments. Biochemistry 46:3856–3861. https://doi.org/10.1021/bi061359o
Rosenberg KJ, Ross JL, Feinstein HE, Feinstein SC, Israelachvili J (2008) Complementary dimerization of microtubule-associated tau protein: Implications for microtubule bundling and tau-mediated pathogenesis. Proc Natl Acad Sci USA 105:7445–7450. https://doi.org/10.1073/pnas.0802036105
Watanabe A, Hong WK, Dohmae N, Takio K, Morishima-Kawashima M, Ihara Y (2004) Molecular aging of tau: disulfide-independent aggregation and non-enzymatic degradation in vitro and in vivo. J Neurochem 90:1302–1311. https://doi.org/10.1111/j.1471-4159.2004.02611.x
Feinstein HE, Benbow SJ, LaPointe NE, Patel N, Ramachandran S, Do TD, Gaylord MR, Huskey NE et al (2016) Oligomerization of the microtubule-associated protein tau is mediated by its N-terminal sequences: implications for normal and pathological tau action. J Neurochem 137:939–954. https://doi.org/10.1111/jnc.13604
Ramachandran G, Udgaonkar JB (2011) Understanding the kinetic roles of the inducer heparin and of rod-like protofibrils during amyloid fibril formation by Tau protein. J Biol Chem 286:38948–38959. https://doi.org/10.1074/jbc.M111.271874
Fichou Y, Oberholtzer ZR, Ngo H, Cheng CY, Keller TJ, Eschmann NA, Han S (2019) Tau-cofactor complexes as building blocks of tau fibrils. Front Neurosci 13:1339. https://doi.org/10.3389/fnins.2019.01339
von Bergen M, Barghorn S, Jeganathan S, Mandelkow EM, Mandelkow E (2006) Spectroscopic approaches to the conformation of tau protein in solution and in paired helical filaments. Neurodegener Dis 3:197–206. https://doi.org/10.1159/000095257
Jakes R, Novak M, Davison M, Wischik CM (1991) Identification of 3- and 4-repeat tau isoforms within the PHF in Alzheimer’s disease. EMBO J 10:2725–2729
Andronesi OC, von Bergen M, Biernat J, Seidel K, Griesinger C, Mandelkow E, Baldus M (2008) Characterization of Alzheimer’s-like paired helical filaments from the core domain of tau protein using solid-state NMR spectroscopy. J Am Chem Soc 130:5922–5928. https://doi.org/10.1021/ja7100517
Daebel V, Chinnathambi S, Biernat J, Schwalbe M, Habenstein B, Loquet A, Akoury E, Tepper K et al (2012) beta-Sheet core of tau paired helical filaments revealed by solid-state NMR. J Am Chem Soc 134:13982–13989. https://doi.org/10.1021/ja305470p
Margittai M, Langen R (2004) Template-assisted filament growth by parallel stacking of tau. Proc Natl Acad Sci USA 101:10278–10283. https://doi.org/10.1073/pnas.0401911101
Kirschner DA, Abraham C, Selkoe DJ (1986) X-ray diffraction from intraneuronal paired helical filaments and extraneuronal amyloid fibers in Alzheimer disease indicates cross-beta conformation. Proc Natl Acad Sci USA 83:503–507. https://doi.org/10.1073/pnas.83.2.503
Barghorn S, Davies P, Mandelkow E (2004) Tau paired helical filaments from Alzheimer’s disease brain and assembled in vitro are based on beta-structure in the core domain. Biochemistry 43:1694–1703. https://doi.org/10.1021/bi0357006
Bibow S, Mukrasch MD, Chinnathambi S, Biernat J, Griesinger C, Mandelkow E, Zweckstetter M (2011) The dynamic structure of filamentous tau. Angew Chem Int Ed Engl 50:11520–11524. https://doi.org/10.1002/anie.201105493
Goedert M, Eisenberg DS, Crowther RA (2017) Propagation of tau aggregates and neurodegeneration. Annu Rev Neurosci 40:189–210. https://doi.org/10.1146/annurev-neuro-072116-031153
Goedert M, Spillantini MG (2019) Ordered assembly of tau protein and neurodegeneration. Adv Exp Med Biol 1184:3–21. https://doi.org/10.1007/978-981-32-9358-8_1
Taniguchi-Watanabe S, Arai T, Kametani F, Nonaka T, Masuda-Suzukake M, Tarutani A, Murayama S, Saito Y et al (2016) Biochemical classification of tauopathies by immunoblot, protein sequence and mass spectrometric analyses of sarkosyl-insoluble and trypsin-resistant tau. Acta Neuropathol 131:267–280. https://doi.org/10.1007/s00401-015-1503-3
Fitzpatrick AWP, Falcon B, He S, Murzin AG, Murshudov G, Garringer HJ, Crowther RA, Ghetti B et al (2017) Cryo-EM structures of tau filaments from Alzheimer’s disease. Nature 547:185–190. https://doi.org/10.1038/nature23002
Falcon B, Zhang W, Schweighauser M, Murzin AG, Vidal R, Garringer HJ, Ghetti B, Scheres SHW et al (2018) Tau filaments from multiple cases of sporadic and inherited Alzheimer’s disease adopt a common fold. Acta Neuropathol 136:699–708. https://doi.org/10.1007/s00401-018-1914-z
Falcon B, Zhang W, Murzin AG, Murshudov G, Garringer HJ, Vidal R, Crowther RA, Ghetti B et al (2018) Structures of filaments from Pick’s disease reveal a novel tau protein fold. Nature 561:137–140. https://doi.org/10.1038/s41586-018-0454-y
Falcon B, Zivanov J, Zhang W, Murzin AG, Garringer HJ, Vidal R, Crowther RA, Newell KL et al (2019) Novel tau filament fold in chronic traumatic encephalopathy encloses hydrophobic molecules. Nature 568:420–423. https://doi.org/10.1038/s41586-019-1026-5
Zhang W, Tarutani A, Newell KL, Murzin AG, Matsubara T, Falcon B, Vidal R, Garringer HJ et al (2020) Novel tau filament fold in corticobasal degeneration. Nature 580:283–287. https://doi.org/10.1038/s41586-020-2043-0
Crowther RA (1991) Straight and paired helical filaments in Alzheimer disease have a common structural unit. Proc Natl Acad Sci USA 88:2288–2292. https://doi.org/10.1073/pnas.88.6.2288
Crowther RA, Wischik CM (1985) Image reconstruction of the Alzheimer paired helical filament. EMBO J 4:3661–3665
Zhang W, Falcon B, Murzin AG, Fan J, Crowther RA, Goedert M, Scheres SH (2019) Heparin-induced tau filaments are polymorphic and differ from those in Alzheimer’s and Pick’s diseases. Elife 8:e43584. https://doi.org/10.7554/eLife.43584
Dregni AJ, Mandala VS, Wu H, Elkins MR, Wang HK, Hung I, DeGrado WF, Hong M (2019) In vitro 0N4R tau fibrils contain a monomorphic beta-sheet core enclosed by dynamically heterogeneous fuzzy coat segments. Proc Natl Acad Sci USA 116:16357–16366. https://doi.org/10.1073/pnas.1906839116
Nizynski B, Dzwolak W, Nieznanski K (2017) Amyloidogenesis of tau protein. Protein Sci 26:2126–2150. https://doi.org/10.1002/pro.3275
Nizynski B, Nieznanska H, Dec R, Boyko S, Dzwolak W, Nieznanski K (2018) Amyloidogenic cross-seeding of tau protein: transient emergence of structural variants of fibrils. PLoS ONE 13:e0201182. https://doi.org/10.1371/journal.pone.0201182
Scheres SH, Zhang W, Falcon B, Goedert M (2020) Cryo-EM structures of tau filaments. Curr Opin Struct Biol 64:17–25. https://doi.org/10.1016/j.sbi.2020.05.011
Chirita CN, Congdon EE, Yin H, Kuret J (2005) Triggers of full-length tau aggregation: a role for partially folded intermediates. Biochemistry 44:5862–5872. https://doi.org/10.1021/bi0500123
Barghorn S, Zheng-Fischhofer Q, Ackmann M, Biernat J, von Bergen M, Mandelkow EM, Mandelkow E (2000) Structure, microtubule interactions, and paired helical filament aggregation by tau mutants of frontotemporal dementias. Biochemistry 39:11714–11721. https://doi.org/10.1021/bi000850r
Eckermann K, Mocanu MM, Khlistunova I, Biernat J, Nissen A, Hofmann A, Schonig K, Bujard H et al (2007) The beta-propensity of tau determines aggregation and synaptic loss in inducible mouse models of tauopathy. J Biol Chem 282:31755–31765. https://doi.org/10.1074/jbc.M705282200
Rossi G, Bastone A, Piccoli E, Mazzoleni G, Morbin M, Uggetti A, Giaccone G, Sperber S et al (2012) New mutations in MAPT gene causing frontotemporal lobar degeneration: biochemical and structural characterization. Neurobiol Aging 33:834. https://doi.org/10.1016/j.neurobiolaging.2011.08.008
Strang KH, Croft CL, Sorrentino ZA, Chakrabarty P, Golde TE, Giasson BI (2018) Distinct differences in prion-like seeding and aggregation between tau protein variants provide mechanistic insights into tauopathies. J Biol Chem 293:2408–2421. https://doi.org/10.1074/jbc.M117.815357
Margittai M, Langen R (2006) Side chain-dependent stacking modulates tau filament structure. J Biol Chem 281:37820–37827. https://doi.org/10.1074/jbc.M605336200
Alonso A, Zaidi T, Novak M, Grundke-Iqbal I, Iqbal K (2001) Hyperphosphorylation induces self-assembly of tau into tangles of paired helical filaments/straight filaments. Proc Natl Acad Sci USA 98:6923–6928. https://doi.org/10.1073/pnas.121119298
Liu F, Li B, Tung EJ, Grundke-Iqbal I, Iqbal K, Gong CX (2007) Site-specific effects of tau phosphorylation on its microtubule assembly activity and self-aggregation. Eur J Neurosci 26:3429–3436. https://doi.org/10.1111/j.1460-9568.2007.05955.x
Alonso AD, Di Clerico J, Li B, Corbo CP, Alaniz ME, Grundke-Iqbal I, Iqbal K (2010) Phosphorylation of tau at Thr212, Thr231, and Ser262 combined causes neurodegeneration. J Biol Chem 285:30851–30860. https://doi.org/10.1074/jbc.M110.110957
Qi H, Prabakaran S, Cantrelle FX, Chambraud B, Gunawardena J, Lippens G, Landrieu I (2016) Characterization of neuronal tau protein as a target of extracellular signal-regulated kinase. J Biol Chem 291:7742–7753. https://doi.org/10.1074/jbc.M115.700914
Schneider A, Biernat J, von Bergen M, Mandelkow E, Mandelkow EM (1999) Phosphorylation that detaches tau protein from microtubules (Ser262, Ser214) also protects it against aggregation into Alzheimer paired helical filaments. Biochemistry 38:3549–3558. https://doi.org/10.1021/bi981874p
Haj-Yahya M, Gopinath P, Rajasekhar K, Mirbaha H, Diamond MI, Lashuel HA (2020) Site-specific hyperphosphorylation inhibits, rather than promotes, tau fibrillization, seeding capacity, and its microtubule binding. Angew Chem Int Ed Engl 59:4059–4067. https://doi.org/10.1002/anie.201913001
Necula M, Kuret J (2004) Pseudophosphorylation and glycation of tau protein enhance but do not trigger fibrillization in vitro. J Biol Chem 279:49694–49703. https://doi.org/10.1074/jbc.M405527200
Chang E, Kim S, Schafer KN, Kuret J (2011) Pseudophosphorylation of tau protein directly modulates its aggregation kinetics. Biochim Biophys Acta 1814:388–395. https://doi.org/10.1016/j.bbapap.2010.10.005
Tetz G, Pinho M, Pritzkow S, Mendez N, Soto C, Tetz V (2020) Bacterial DNA promotes tau aggregation. Sci Rep 10:2369. https://doi.org/10.1038/s41598-020-59364-x
Maiza A, Chantepie S, Vera C, Fifre A, Huynh MB, Stettler O, Ouidja MO, Papy-Garcia D (2018) The role of heparan sulfates in protein aggregation and their potential impact on neurodegeneration. FEBS Lett 592:3806–3818. https://doi.org/10.1002/1873-3468.13082
Roman AY, Devred F, Byrne D, La Rocca R, Ninkina NN, Peyrot V, Tsvetkov PO (2019) Zinc induces temperature-dependent reversible self-assembly of tau. J Mol Biol 431:687–695. https://doi.org/10.1016/j.jmb.2018.12.008
Ahmadi S, Zhu S, Sharma R, Wu B, Soong R, Dutta Majumdar R, Wilson DJ, Simpson AJ et al (2019) Aggregation of microtubule binding repeats of tau protein is promoted by Cu(2). ACS Omega 4:5356–5366. https://doi.org/10.1021/acsomega.8b03595
Moreira GG, Cristóvão JS, Torres VM, Carapeto AP, Rodrigues MS, Landrieu I, Cordeiro C, Gomes CM (2019) Zinc binding to tau influences aggregation kinetics and oligomer distribution. Int J Mol Sci 20:5979. https://doi.org/10.3390/ijms20235979
Zhou Z, Fan JB, Zhu HL, Shewmaker F, Yan X, Chen X, Chen J, Xiao GF et al (2009) Crowded cell-like environment accelerates the nucleation step of amyloidogenic protein misfolding. J Biol Chem 284:30148–30158. https://doi.org/10.1074/jbc.M109.002832
Ma Q, Fan JB, Zhou Z, Zhou BR, Meng SR, Hu JY, Chen J, Liang Y (2012) The contrasting effect of macromolecular crowding on amyloid fibril formation. PLoS ONE 7:e36288. https://doi.org/10.1371/journal.pone.0036288
Wu Y, Teng N, Li S (2016) Effects of macromolecular crowding and osmolyte on human tau fibrillation. Int J Biol Macromol 90:27–36. https://doi.org/10.1016/j.ijbiomac.2015.11.091
Kundel F, De S, Flagmeier P, Horrocks MH, Kjaergaard M, Shammas SL, Jackson SE, Dobson CM et al (2018) Hsp70 inhibits the nucleation and elongation of tau and sequesters tau aggregates with high Affinity. ACS Chem Biol 13:636–646. https://doi.org/10.1021/acschembio.7b01039
Baughman HER, Clouser AF, Klevit RE, Nath A (2018) HspB1 and Hsc70 chaperones engage distinct tau species and have different inhibitory effects on amyloid formation. J Biol Chem 293:2687–2700. https://doi.org/10.1074/jbc.M117.803411
Thompson AD, Scaglione KM, Prensner J, Gillies AT, Chinnaiyan A, Paulson HL, Jinwal UK, Dickey CA et al (2012) Analysis of the tau-associated proteome reveals that exchange of Hsp70 for Hsp90 is involved in tau degradation. ACS Chem Biol 7:1677–1686. https://doi.org/10.1021/cb3002599
Mok SA, Condello C, Freilich R, Gillies A, Arhar T, Oroz J, Kadavath H, Julien O et al (2018) Mapping interactions with the chaperone network reveals factors that protect against tau aggregation. Nat Struct Mol Biol 25:384–393. https://doi.org/10.1038/s41594-018-0057-1
Weickert S, Wawrzyniuk M, John LH, Rudiger SGD, Drescher M (2020) The mechanism of Hsp90-induced oligomerization of tau. Sci Adv 6:eaax6999. https://doi.org/10.1126/sciadv.aax6999
McEwan WA, Falcon B, Vaysburd M, Clift D, Oblak AL, Ghetti B, Goedert M, James LC (2017) Cytosolic Fc receptor TRIM21 inhibits seeded tau aggregation. Proc Natl Acad Sci USA 114:574–579. https://doi.org/10.1073/pnas.1607215114
Cisek K, Cooper GL, Huseby CJ, Kuret J (2014) Structure and mechanism of action of tau aggregation inhibitors. Curr Alzheimer Res 11:918–927. https://doi.org/10.2174/1567205011666141107150331
Brunden KR, Ballatore C, Crowe A, Smith AB 3rd, Lee VM, Trojanowski JQ (2010) Tau-directed drug discovery for Alzheimer’s disease and related tauopathies: a focus on tau assembly inhibitors. Exp Neurol 223:304–310. https://doi.org/10.1016/j.expneurol.2009.08.031
Brunden KR, Trojanowski JQ, Lee VM (2009) Advances in tau-focused drug discovery for Alzheimer’s disease and related tauopathies. Nat Rev Drug Discov 8:783–793. https://doi.org/10.1038/nrd2959
Calcul L, Zhang B, Jinwal UK, Dickey CA, Baker BJ (2012) Natural products as a rich source of tau-targeting drugs for Alzheimer’s disease. Future Med Chem 4:1751–1761. https://doi.org/10.4155/fmc.12.124
Schafer KN, Cisek K, Huseby CJ, Chang E, Kuret J (2013) Structural determinants of tau aggregation inhibitor potency. J Biol Chem 288:32599–32611. https://doi.org/10.1074/jbc.M113.503474
Rauch JN, Olson SH, Gestwicki JE (2017) Interactions between microtubule-associated protein tau (MAPT) and small molecules. Cold Spring Harb Perspect Med 7:a024034. https://doi.org/10.1101/cshperspect.a024034
Mouchlis VD, Melagraki G, Zacharia LC, Afantitis A (2020) Computer-aided drug design of beta-secretase, gamma-secretase and anti-tau inhibitors for the discovery of novel Alzheimer’s therapeutics. Int J Mol Sci. https://doi.org/10.3390/ijms21030703
Pickhardt M, Neumann T, Schwizer D, Callaway K, Vendruscolo M, Schenk D, St George-Hyslop P, Mandelkow EM et al (2015) Identification of small molecule inhibitors of tau aggregation by targeting monomeric tau as a potential therapeutic approach for tauopathies. Curr Alzheimer Res 12:814–828. https://doi.org/10.2174/156720501209151019104951
Wobst HJ, Sharma A, Diamond MI, Wanker EE, Bieschke J (2015) The green tea polyphenol (−)-epigallocatechin gallate prevents the aggregation of tau protein into toxic oligomers at substoichiometric ratios. FEBS Lett 589:77–83. https://doi.org/10.1016/j.febslet.2014.11.026
Dubey T, Gorantla NV, Chandrashekara KT, Chinnathambi S (2019) Photoexcited toluidine blue inhibits tau aggregation in Alzheimer’s disease. ACS Omega 4:18793–18802. https://doi.org/10.1021/acsomega.9b02792
Lo CH, Lim CK, Ding Z, Wickramasinghe SP, Braun AR, Ashe KH, Rhoades E, Thomas DD et al (2019) Targeting the ensemble of heterogeneous tau oligomers in cells: a novel small molecule screening platform for tauopathies. Alzheimers Dement 15:1489–1502. https://doi.org/10.1016/j.jalz.2019.06.4954
Wischik CM, Edwards PC, Lai RY, Roth M, Harrington CR (1996) Selective inhibition of Alzheimer disease-like tau aggregation by phenothiazines. Proc Natl Acad Sci USA 93:11213–11218. https://doi.org/10.1073/pnas.93.20.11213
Kiss R, Csizmadia G, Solti K, Keresztes A, Zhu M, Pickhardt M, Mandelkow E, Toth G (2018) Structural basis of small molecule targetability of monomeric tau protein. ACS Chem Neurosci 9:2997–3006. https://doi.org/10.1021/acschemneuro.8b00182
Chong B, Li M, Li T, Yu M, Zhang Y, Liu Z (2018) Conservation of potentially druggable cavities in intrinsically disordered proteins. ACS Omega 3:15643–15652. https://doi.org/10.1021/acsomega.8b02092
Despres C, Di J, Cantrelle FX, Li Z, Huvent I, Chambraud B, Zhao J, Chen J et al (2019) Major differences between the self-assembly and seeding behavior of heparin-induced and in vitro phosphorylated tau and their modulation by potential inhibitors. ACS Chem Biol 14:1363–1379. https://doi.org/10.1021/acschembio.9b00325
Baggett DW, Nath A (2018) The rational discovery of a tau aggregation inhibitor. Biochemistry 57:6099–6107. https://doi.org/10.1021/acs.biochem.8b00581
Ruan H, Sun Q, Zhang W, Liu Y, Lai L (2019) Targeting intrinsically disordered proteins at the edge of chaos. Drug Discov Today 24:217–227. https://doi.org/10.1016/j.drudis.2018.09.017
Jin F, Yu C, Lai L, Liu Z (2013) Ligand clouds around protein clouds: a scenario of ligand binding with intrinsically disordered proteins. PLoS Comput Biol 9:e1003249. https://doi.org/10.1371/journal.pcbi.1003249
Falcon B, Cavallini A, Angers R, Glover S, Murray TK, Barnham L, Jackson S, O’Neill MJ et al (2015) Conformation determines the seeding potencies of native and recombinant Tau aggregates. J Biol Chem 290:1049–1065. https://doi.org/10.1074/jbc.M114.589309
Zheng J, Liu C, Sawaya MR, Vadla B, Khan S, Woods RJ, Eisenberg D, Goux WJ et al (2011) Macrocyclic beta-sheet peptides that inhibit the aggregation of a tau-protein-derived hexapeptide. J Am Chem Soc 133:3144–3157. https://doi.org/10.1021/ja110545h
Sievers SA, Karanicolas J, Chang HW, Zhao A, Jiang L, Zirafi O, Stevens JT, Munch J et al (2011) Structure-based design of non-natural amino-acid inhibitors of amyloid fibril formation. Nature 475:96–100. https://doi.org/10.1038/nature10154
Seidler PM, Boyer DR, Rodriguez JA, Sawaya MR, Cascio D, Murray K, Gonen T, Eisenberg DS (2018) Structure-based inhibitors of tau aggregation. Nat Chem 10:170–176. https://doi.org/10.1038/nchem.2889
Seidler PM, Boyer DR, Murray KA, Yang TP, Bentzel M, Sawaya MR, Rosenberg G, Cascio D et al (2019) Structure-based inhibitors halt prion-like seeding by Alzheimer’s disease-and tauopathy-derived brain tissue samples. J Biol Chem 294:16451–16464. https://doi.org/10.1074/jbc.RA119.009688
Berhanu WM, Masunov AE (2015) Atomistic mechanism of polyphenol amyloid aggregation inhibitors: molecular dynamics study of Curcumin, Exifone, and Myricetin interaction with the segment of tau peptide oligomer. J Biomol Struct Dyn 33:1399–1411. https://doi.org/10.1080/07391102.2014.951689
Gauthier S, Feldman HH, Schneider LS, Wilcock GK, Frisoni GB, Hardlund JH, Moebius HJ, Bentham P et al (2016) Efficacy and safety of tau-aggregation inhibitor therapy in patients with mild or moderate Alzheimer’s disease: a randomised, controlled, double-blind, parallel-arm, phase 3 trial. Lancet 388:2873–2884. https://doi.org/10.1016/S0140-6736(16)31275-2
Wilcock GK, Gauthier S, Frisoni GB, Jia J, Hardlund JH, Moebius HJ, Bentham P, Kook KA et al (2018) Potential of low dose leuco-methylthioninium bis(hydromethanesulphonate) (LMTM) monotherapy for treatment of mild Alzheimer’s disease: cohort analysis as modified primary outcome in a phase III clinical trial. J Alzheimers Dis 61:435–457. https://doi.org/10.3233/JAD-170560
Schelter BO, Shiells H, Baddeley TC, Rubino CM, Ganesan H, Hammel J, Vuksanovic V, Staff RT et al (2019) Concentration-dependent activity of hydromethylthionine on cognitive decline and brain atrophy in mild to moderate Alzheimer’s disease. J Alzheimers Dis 72:931–946. https://doi.org/10.3233/JAD-190772
Holehouse AS, Das RK, Ahad JN, Richardson MO, Pappu RV (2017) CIDER: resources to analyze sequence-ensemble relationships of intrinsically disordered proteins. Biophys J 112:16–21. https://doi.org/10.1016/j.bpj.2016.11.3200
Romero P, Obradovic Z, Dunker AK (1997) Sequence data analysis for long disordered regions prediction in the calcineurin family. Genome Inform 8:110–124
Li X, Romero P, Rani M, Dunker AK, Obradovic Z (1999) Predicting protein disorder for N-, C-, and internal regions. Genome Inform 10:30–40
Acknowledgements
This work was supported by the Natural Science Foundation of Hubei Province (2019CFB713) and funding from Hubei University of Technology (BSQD2017022).
Author information
Authors and Affiliations
Contributions
Y. H. and Z. S. had the idea for the article; Y. Z., J. Y., and B. Z. performed the literature search and data analysis; Y. Z., J. Y., and M. G. drafted the work; Y. Z., B. Z., M. G., Y. H., and Z. S. critically revised the work.
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that there are no conflicts.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zeng, Y., Yang, J., Zhang, B. et al. The structure and phase of tau: from monomer to amyloid filament. Cell. Mol. Life Sci. 78, 1873–1886 (2021). https://doi.org/10.1007/s00018-020-03681-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-020-03681-x