Abstract
In recent years, liquid biopsy has emerged as an alternative method to diagnose and monitor tumors. Compared to classical tissue biopsy procedures, liquid biopsy facilitates the repetitive collection of diverse cellular and acellular analytes from various biofluids in a non/minimally invasive manner. This strategy is of greater significance for high-grade brain malignancies such as glioblastoma as the quantity and accessibility of tumors are limited, and there are collateral risks of compromised life quality coupled with surgical interventions. Currently, blood and cerebrospinal fluid (CSF) are the most common biofluids used to collect circulating cells and biomolecules of tumor origin. These liquid biopsy analytes have created opportunities for real-time investigations of distinct genetic, epigenetic, transcriptomics, proteomics, and metabolomics alterations associated with brain tumors. This review describes different classes of liquid biopsy biomarkers present in the biofluids of brain tumor patients. Moreover, an overview of the liquid biopsy applications, challenges, recent technological advances, and clinical trials in the brain have also been provided.
Similar content being viewed by others
Introduction
Biopsy as a diagnostic approach has been routinely used for molecular testing and deciding the precise therapeutic strategies in various human diseases including cancer [1]. The gold standard, conventional tissue biopsy provides high yields of analytes to study cancer-specific alterations and the opportunities to perform the histological examination and staging. However, tissue biopsy implementations have limitations such as surgical interventions, accessibility of tumor tissue, sampling bias, localized analysis, lack of serial monitoring, and inability to study tumor heterogeneity and evolution. In contrast, liquid biopsy provides a minimally invasive alternative for systemic and real-time tumor progression monitoring and therapeutic interventions [2].
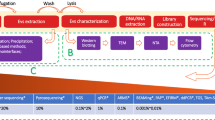
Liquid biopsy in cancer patients primarily aims at detecting, analyzing, and monitoring diverse tumor-derived analytes circulating in different biological fluids, including blood, urine, saliva, cerebrospinal fluid (CSF), pleural fluid, stool, semen, bone marrow, and ascites [3,4,5]. Tumor-derived materials in these biofluids that have shown promising future in studying diverse aspects of tumor biology involve circulating tumor cells (CTCs), cell-free DNA (cfDNA), cell-free RNA (cfRNA), tumor-specific proteins, tumor-educated platelets (TEPs), extracellular membrane-bound vesicles, and other metabolites of tumor origin [6,7,8,9,10,11]. These analytes possess immense biological information, thus empowering the researchers to perform cancer-type-specific investigations from genomics, epigenomics, transcriptomics, proteomics, and metabolomics perspectives as shown in Fig. 1, and is also frequently reviewed in the literature [12]. Consequently, in recent years, liquid biopsy has emerged as one of the diagnostic methods for monitoring and subtyping solid tumors [13]. This strategy is of additional significance to central nervous system (CNS) tumor patients because of the risks posed by the available diagnostic procedures [14].
In different types of cancers, biofluids such as blood, urine, CSF, ascites, pleural fluid, saliva, stool, semen, vitreous fluid, etc. are routinely collected to study circulating analytes which include cell-free DNA (cfDNA), cell-free coding, and non-coding RNA species (cfRNA), circulating proteins (CPs), CTCs (single or clusters), extracellular vesicles (EVs), circulating metabolites (CMs), cancer-associated (CA) fibroblast, tumor-educated platelets (TEPs), etc. Each of these circulating analytes can be investigated for cancer-specific alterations from genomics, epigenomics, transcriptomics, proteomics, and metabolomics perspectives at resolutions ranging from the single cell to the whole organism. This figure has been created using BioRender software and a publication license is obtained.
Brain malignancies include various types of neoplasms originating from either primary brain tumors or metastatic cancers. Glioblastoma is the most frequent and aggressive one in adults, with a 5-year survival rate of less than 5% [15, 16]. The similar appearances of different CNS and non-CNS tumors on neuroimaging scans, and the compromised life quality associated with the neuropathological examination techniques limit the CNS malignancies-specific diagnostic strategies successful otherwise [17]. In addition, the blood-brain barrier (BBB), the microvasculature system of CNS, significantly lowers the amounts of circulating analytes shed by the brain tumors in different biological fluids, thus posing limitations to the liquid biopsy enactments [18, 19]. Therefore, the development of specific and sensitive liquid biopsy-based protocols to diagnose brain tumors is currently of utmost need.
In light of the most recent World Health Organization (WHO) classification of tumors of the CNS edition 5 (WHO CNS5) proposed in 2021, the clinicopathologic significance of molecular profile alterations both in providing ancillary and defining diagnostic knowledge has increased [20]. WHO CNS5 advocates the adoption of integrated diagnostic approaches containing both classical histology and tissue-based tests (e.g., immunohistochemistry, ultrastructural) as well as novel molecular characteristics [21]. The liquid biopsy strategies in brain malignancies can be instrumental in capturing and defining the underlying discrete molecular changes, thus complementing the precision of diagnostic measures. This review describes brain tumor-specific biomarkers with a focus on glioblastoma that are present in various biological fluids, especially blood, and CSF. In addition, clinical applications, and technological advances toward the detection of analytes using the liquid biopsy approach in CNS tumors are explored.
Molecular biomarkers in brain tumors
Over the past few decades, international standards for brain tumor diagnosis have been improvised in a timely manner based on the new developments in the field. The latest molecular feature-centric proposal of WHO has led to a major paradigm shift in brain tumor classification, nomenclature, and grading [21]. For instance, adult-type diffuse gliomas with isocitrate dehydrogenase (IDH) mutation originally identified as glioblastoma in the previous 2016 WHO classification have now been categorized as astrocytoma [22]. Henceforth, for the remainder of this review, the term glioblastoma will refer to adult-type diffuse gliomas characterized either by IDH status or histologically defined by neuropathologists. An updated list of various brain tumor-specific molecular biomarkers has been summarized in Table 1. These genetic markers have now become an integral part of pathological reports, and their applications have transformed the brain tumor routine clinical diagnostic prototype [23, 24]. In this section, a few of the well-characterized molecular signatures of brain tumors are described.
IDH mutations have been observed both in CNS and non-CNS tumors, predominantly in secondary adult-type diffuse gliomas (astrocytoma and oligodendroglioma) [20, 25]. The IDH protein forms a dimer in the activated state and prevents intracellular oxidative injury by catalyzing the reversible oxidative decarboxylation of the substrate isocitrate to α-ketoglutarate (α-KG) in the tricarboxylic acid (TCA) cycle [26]. In brain malignancies, the most common IDH mutations identified include arginine residues (R132 for IDH1, R140, or R172 for IDH2) that are crucial for the identification of the substrate isocitrate [27]. A homodimer resulting from an IDH-mutant gene shows neomorphic activity and catalyzes the conversion of isocitrate to an oncometabolite i.e., D-2-hydroxyglutarate (D-2-HG), which has long been associated with altered cellular metabolism and oncogenesis, mostly through DNA and histone methylation [28, 29]. The inhibitors-mediated direct targeting of the IDH mutant-enzyme prevents the manifestation of this malignant transformation, but it also results in the increased resistance of the tumor cells to genotoxic therapies such as radiation and chemo agents [30, 31]. Therefore, the IDH mutations-associated prolonged median survival is considered a positive prognostic biomarker in grade 2–4 gliomas [32]. Furthermore, distinct mutations and amplifications in the epidermal growth factor receptor (EGFR) gene have also been classified as glioblastoma prognostic biomarkers [15, 21]. These genomic changes have been reported to confer a ligand-independent constitutive activated state to EGFR in cancer cells, which in turn stimulate responses and pathways required for tumorigenesis [33]. In brain tumors, the most studied EGFR variation is transcript variant III (EGFRvIII) (exons 2-7 deletion), and its expression has been reported to promote cell proliferation, invasion, and angiogenesis [34]. In recent years, the EGFRvIII variant has emerged as one of the preferred candidates for glioblastoma-targeted therapy as it is exclusively present in tumor cells [35, 36]. Moreover, the elevated levels of glial fibrillary acidic protein (GFAP) in blood have been correlated with the size of the tumor, the extent of necrosis, and the intratumor expression [37]. GFAP is the principal component of the cytoskeletal intermediate filaments produced by different cells of the central nervous system including astrocytes and is currently used as a prevalent marker for the identification of CTCs across gliomas [37,38,39,40].
Mutations altering the tumor-suppressing activity of the p53 protein predominantly occur in secondary brain malignancies (90%) compared to the primary lesions (30%) [41]. As an early event, mutations in the TP53 (gene encoding p53 protein) are coupled with concomitant loss of chromosome 17p in grade 2-3 astrocytoma, thus implying its significance in tumorigenesis [42]. In WHO CNS5, the clinicopathological significance of TP53 has been used to identify subtypes of molecularly defined medulloblastoma sonic hedgehog (SHH) activated brain tumors [21]. Overall TP53 gene mutations can attribute gain-of-functions, loss-of-functions, or dominant negative phenotypes, but the impact of these discrete mutational profiles in glioblastoma pathogenesis is still unclear [43,44,45]. The enhanced activity of the mevalonate (MVA) pathway pro-tumorigenesis enzymes MVA kinase and 3’-hydroxy-3’-methylglutaryl coenzyme A reductase is one of the suggested mechanisms of the p53-mediated glioma etiology [46]. In addition, other effector molecules of the p53 signaling pathway, such as mouse-double minute 2 homolog (MDM2), mouse-double minute 4 homolog MDM4, inhibitors of CDK4 (INK4), ADP-ribosylation factor (ARF), etc., are also known to modulate the p53 tumor suppressor activity [41, 47]. In a recent study involving glioblastoma-patients-derived brain tumor stem cells and orthotopic xenograft models, MDM2 inhibitor (BI-907828) appears as a promising new therapeutic for primary brain tumor patients having wildtype TP53 gene [48].
When compared to other adult-type diffuse gliomas carrying IDH mutations, glioblastoma patients with the wildtype IDH gene possess significantly lower levels of methylation at O6-methylguanine-DNA-methyltransferase (MGMT) gene promoter [49, 50]. In healthy cells, the MGMT gene codes for the O6 alkylguanine-DNA-alkyltransferase (AGT) protein involved in maintaining cell physiology and genomic stability by removing the alkylating lesions [51, 52]. The higher levels of AGT protein thus decrease both the risks of carcinogenesis and the chances of deleterious mutations on exposure to therapeutic methylating agents. In contrast, methylation of the MGMT promoter regions in brain malignancies prevents MGMT expression and increases genomic instability by enhancing the efficacy of alkylating agents. In glioblastoma patients, even the extent of MGMT promoter methylation has been shown to impact survival outcomes [53, 54]. Thus, while defining alkylating agent-based therapies in brain tumors, detecting the methylation status of the MGMT promoter region is highly recommended [55]. Furthermore, two specific point mutations (C228T and C250T) in the promoter region of the telomerase reverse transcriptase (pTERT) gene occur at higher frequency in glioblastoma patients (~80%) [56, 57]. These mutations are anticipated to enhance the telomerase enzyme activity through the formation of the TERT-activating GA binding protein (GABP) transcription factor complex, leading to the progressive increase in the length of telomeres essential for the uncontrolled proliferation of tumor cells [56, 58, 59]. In addition, single nucleotide polymorphism (C- allele of rs2853669) variants located in the proximity of TERT mutation hotspots are associated with increased risks of developing glioblastoma and reduced overall survival (OS) [60, 61]. Therefore, MGMT promoter methylation and pTERT mutation status can be used for glioblastoma patients’ prognosis [62].
Loss of heterozygosity (LOH) mutations, characterized by the loss of a copy of a gene or group of genes, is also prevalent in brain tumors. Almost 60-80% of primary and secondary high-grade gliomas contain 10q LOH. The most frequently deleted genetic loci on chromosome 10 include 10q23-24, 10q25-qter, and 10q14-p15. Deletion of 10q23-24 loci results in the loss of tumor suppressor gene PTEN (phosphatase activity) along with DMBT1, FGFR2, WDRI1, LGI1, and MIX1 genes that play a critical role in inhibiting P13K/AKT/mTOR pathway involved in cell proliferation. 10q25-qter deletion is associated with the progression of low-grade brain tumors to high-grade [63]. In addition, 22q LOH is also present in 80% of secondary, and 41% of primary brain tumors. Loss of 22q12.3 loci results in the absence of tissue inhibitor of metalloproteinases-3 protein (TIMP-3) protein involved in the inhibition of tumorigenesis and induction of apoptosis [64]. Other commonly occurring LOH mutations in brain malignancies are located on chromosomes 1p, 9p, 17p, and 19q [65]. Thus, identification of the abovementioned LOH through microsatellite and PCR-based assays is very helpful in classifying and designing informed therapeutic strategies for brain tumor patients.
In combination with the aforementioned molecular features and patient characteristics, Alpha Thalassemia/Mental Retardation Syndrome X-linked (ATRX), a transcriptional regulator protein, has been now routinely used to identify biological distinct glioma subclasses. Most of the studies involving glioma and ATRX are correlation-based, and the role of ATRX is not precisely defined. For example, there exists a strong correlation between IDH canonical mutations and ATRX mutation, however, the coexistence of 1p/19q codeletion and ATRX loss has been rarely observed in the clinics [66,67,68]. Moreover, ATRX mutations have also been correlated to genomic instability conferring telomerase-independent telomere lengthening mechanism termed ‘Alternate Lengthening of Telomeres’ (ALT) [69,70,71,72,73,74]. Altogether, each of these brain tumor-specific molecular biomarkers has some advantages and disadvantages, therefore, defining a combination of multiple markers may increase the diagnostic utility of these molecular changes in clinical settings.
Brain tumor-specific biomarkers in blood
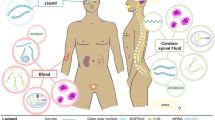
Blood is the most common source of biological samples in liquid biopsy-based oncology studies. It is of greater importance when the size and accessibility of the tumor are the constraints. Identification of blood-based biomarkers using liquid biopsy has been explored extensively in different extra-cerebral cancers such as breast cancer, non-small cell lung cancer, colorectal cancer, etc [75,76,77]. However, the exploration of brain space for searching blood-based biomarkers is still in its initial developmental stages. Various analytes present in a brain tumor patient’s blood, and their utility in defining disease phenotype are depicted in Fig. 2 and are also a very active research domain [78]. In the following sections, specific analytes in the blood used for biomarker identification in brain tumors will be described.
Analytes such as nucleic acids (DNA/RNAs), proteins, and metabolites can be either collected as circulating cell-free entities or extracted from circulating tumor cells/extracellular vesicles/tumor-educated platelets. Each of these circulating analytes can be explored for tumor-specific alterations such as different classes of mutations, epigenetic modifications, the fragmentation pattern of DNA, nucleosome patterning, chromosomal aberrations, presence/absence/change in levels of RNAs/proteins/metabolites, post-translational modifications, etc. This figure has been created using BioRender software and a publication license is obtained.
Circulating tumor cells (CTCs)
CTCs are phenotypically distinct rare subsets of the tumor cell population released by primary or metastatic lesions into the biofluids [79, 80]. The abundance of CTCs in biofluids is very low (<10 cells/mL of biofluid) even in metastatic conditions, and varies drastically between various cancer types [81, 82]. CTCs can be retrieved from a cancer patient’s body fluids either as single cells or cell aggregates. CTC clusters have also been reported to contain WBCs, and the presence of CTC-WBC clusters suggests poor prognosis in cancer patients [83, 84]. From the disease pathology perspective, CTCs represent the potential of epithelial tumor cells to metastasize. Moreover, CTCs can attain either stem cell-like or mesenchymal phenotypes upon an epithelial-to-mesenchymal transition (EMT) [85]. At present, the identification and isolation of CTCs from biofluids are mainly based on the presence/absence of specific cell-surface epithelial markers or biophysical properties (size, deformity, etc.) of the CTCs [86]. For instance, an antibody against epithelial cell adhesion molecule (EpCAM) and cytokines are currently being used to enrich and detect CTCs in the FDA-approved CellSearch System (Veridex, Warren, NJ, USA) [80, 87].
CTCs were first identified in common non-CNS cancer types such as breast cancers, prostate cancers, etc., and more recently in CNS tumors [88, 89]. Since tumor cells from high-grade gliomas including glioblastoma patients preferentially adopt mesenchymal phenotype compared to epithelial, therefore, the conventional methods of CTCs identification using the CellSearch system are not very effective for the detection and enrichment of CTCs originating from brain tumors [90]. Recent studies have shown the presence of glioma CTCs in CSF and blood, suggesting that brain tumor cells are primed to breach BBB and enter systemic circulation [91, 92]. Several noteworthy attempts have been made to detect CTCs in the blood of glioblastoma patients such as the application of antibodies to target GFAP, amplification of the EGFR gene, etc, [39]. In addition, an assay detecting the increased activity of telomerase enzyme exclusively in tumor cells has been devised to identify circulating brain tumor cells. Clinical data suggest that this adenoviral detection-based strategy is capable of distinguishing pseudoprogression from true tumor progression in brain malignancies [93]. Moreover, CTCs have also been detected in the peripheral blood of glioma patients through examination of aneuploidy of chromosome 8 (CEP8-FISH) [94]. A recently developed microfluidic device, the CTC-iChip, selectively depletes leukocytes and efficiently identifies CTCs enriched in the peripheral blood of glioblastoma patients by staining with antibody cocktail referred to as STEAM (SOX2, Tubulin beta-3, EGFR, A2B5, and c-MET) [95].
Proteoglycans are complex molecules that contain a protein core with a sugar side chain. Although yet to undergo clinical testing, proteoglycans have been used to detect CTCs in many cancers. In high-grade gliomas, rVAR2 (recombinant malaria VAR2CSA protein), a ligand for the proteoglycan chondroitin can efficiently detect CTCs in the blood by binding to tumor-specific oncofetal chondroitin sulfate [91]. Glioblastoma’s recurrence rate and progression of low-grade gliomas have been suggested to correlate with the presence of CTCs in gliomas, hence making it an attractive target for liquid biopsy-based diagnostic strategies [96]. In addition to the adults, CTCs can also be captured and identified from the blood of pediatric brain tumor patients [97]. The successful capture of viable CTCs from brain tumor patients has enabled us to develop CTCs-derived cell lines, xenograft, and 3D models, which in turn are useful for conducting functional analyses like therapy testing [82, 98,99,100,101]. Taken together, the CTCs obtained through liquid biopsy are suitable models to study the CNS tumors-specific molecular alterations, and thus helpful in monitoring tumor progression and defining targeted therapies [102].
Circulating nucleic acid biomarkers
Circulating cell-free tumor DNA
Cell-free DNA (cfDNA) is typically 180–200 bp long DNA fragments circulating in various biofluids [103,104,105]. In healthy individuals, cfDNA is mainly the outcome of underlying inflammatory or apoptotic processes [106]. Under physiological conditions, most of these released cfDNA pieces are cleared by phagocytosis, thus maintaining a low level of cfDNA in circulation [85]. By contrast, in cancer patients, there is an accumulation of cfDNA in circulation due to a relatively higher rate of their generation as compared to phagocytic removal. Such cfDNA fragments released by tumor cells are known as circulating cell-free tumor DNA (ctDNA). In advanced solid tumors, the proportion of ctDNA in the whole cfDNA has been correlated with tumor burden [107,108,109,110]. Therefore, ctDNA load can be used to efficiently determine the tumor burden in cancer patients as it requires less volume of biofluid compared to that of CTCs [111].
As a biomarker, ctDNA possesses several distinct characteristics. Its short half-life of approximately 1.5 h is advantageous while investigating the dynamic modifications in tumor homeostasis [105, 112]. The tumor-specific mutations harbored by ctDNA provide specificity for the detection of genetic alterations present exclusively in tumor cells [112]. Quantitatively, the ctDNA carrying tumor-specific variations such as copy number variations, chromosomal rearrangements, point mutations, etc., can contribute 0.1–90% fractions of the total circulating cfDNA [105, 113]. Additionally, the detection rate of ctDNA is much higher in metastatic and advanced disease stages compared to localized disease [114,115,116,117]. The comparative molecular profiling studies between matched ctDNA and tumor samples from colorectal, non-small cell lung carcinoma, and metastatic breast cancers have shown a high degree of concordance to the clinically significant known mutations [105, 113,114,115, 118,119,120,121,122,123]. Furthermore, the abundance of ctDNA has been reported to be negatively correlated with the survival outcomes in various cancer types such as melanoma, ovarian, breast, and colon cancers, etc, [105, 113, 115, 124, 125]. Therefore, ctDNA-based diagnostic assays for brain tumor patients hold a promising future in clinical settings.
In gliomas, the blood-brain barrier limits the potential of ctDNA as an efficient biomarker [112]. This structurally and functionally significant endothelial cell layer prevents the release and detection of ctDNA in circulation. Additionally, the lower frequency of ctDNA detection in CNS malignancies compared to non-CNS tumors has also been partially attributed to its extremely low concentrations in body fluids [115, 116]. For instance, in a study cohort of 419 primary brain tumor patients, ctDNA was detected only in 60% of the total 222 high-grade glioma cases [126]. Several investigations have also demonstrated the effectiveness of ctDNA in identifying brain tumor-specific molecular biomarker gene alterations, of which IDH mutations were detected at a higher frequency [127,128,129,130,131,132,133,134]. For example, in a study involving 157 adult glioma patients’ plasma ctDNA, TERT promoter mutations were identified with a specificity of 90% and sensitivity of 62.5% [135]. Currently, fewer genes with mutation frequency greater than 5% in brain tumors, and the deep sequencing cost involved in their identification are the two major constraints associated with the genetic variations-based diagnosis. In contrast, genome-wide epigenetic landscapes captured on ctDNA represent the real-time dynamic changes linked to the disease phenotype. The most commonly studied epigenetics modification on DNA concerning brain tumors is methylation. The ctDNA of glioma patients has been reported to contain significantly lower levels of Alu methylation than the control group which includes benign intracranial tumor patients and healthy individuals [136]. This is significant given that Alu is the abundant interspersed element that occupies about 10% of the genomes. Several studies implementing an antibody-based capture method to isolate cfDNA have also reported a differential methylation pattern in different cancer types [137, 138]. Moreover, in an independent study, the plasma-cfDNA methylome profile has been exhibited to detect and discriminate intracranial tumors [139]. In recent years, the diagnostic significance of another epigenetic modification i.e., DNA hydroxymethylation has also been realized [140,141,142]. While its overall abundance is ~10% compared to methylation marks, the tissue-specificity it provides is of special interest from a liquid biopsy perspective [143, 144]. Moreover, in contrast to other tumor types, brain tumors possess a higher amount of hydroxymethylation marks and have also been correlated with the survival outcomes of glioblastoma patients [145]. Thus, blood-based liquid biopsy assays using cfDNA, especially epigenetic features, are very critical in defining the diagnosis and treatments of patients with brain malignancies, and the development of more sensitive and accurate ctDNA detection methods will help to achieve the optimum potential of these approaches.
Circulating cell-free RNA
Tumor cells also release cell-free RNA (cfRNA) in circulation [146,147,148]. In patients with low tumor shedding, the signals from tumor-specific overexpressed transcripts in the blood are helpful in detecting cancer. Unlike cfDNA which is contributed to the blood by necrotic cells, the cfRNA can also be added to the systemic circulation through exosome-mediated signaling by living cells. Therefore, cfDNA and cfRNA fractions in the blood due to distinct cell populations wide open the opportunities for cancer detection through their combined assessment [149]. On the technical front, several optimized protocols are currently available to capture unstable cfRNA from blood alone or in combination with cfDNA [150]. For instance, low centrifugation protocols give a higher yield of cfRNA, and the BD Vacutainer K2EDTA (EDTA) tubes are best suited for the simultaneous enrichment of cfDNA and cfRNA.
Most of the previous cfRNA studies are focused either on highly stable and abundant circulating micro RNAs (miRNAs) in plasma or a few of the already reported cancer-associated messenger RNA (mRNAs) [149, 151, 152]. The hypothesis-driven cell-free mRNA studies can only identify genetic alterations or expression changes of previously characterized tumor-associated genes but could miss a substantial number of potential biomarkers. Also, detecting small expression changes linked to early disease states in the presence of abundant circulating transcripts of red blood cells, immune cells, and platelets is difficult [146, 148, 149, 153]. Therefore, the knowledge of circulating cfRNA baseline concentrations in disease-free conditions, and its cancer-subtype specificity is very crucial [154]. Interestingly, transcriptome-wide cfRNA characterization studies in brain tumors are yet to be reported. Furthermore, RNA methylation as a regulatory biological mechanism has also gained extensive momentum in recent years, but the utility of cfRNA methylation has not been fully explored. It is possible that methylation of cfRNA obtained through liquid biopsy could uncover novel cancer-specific alterations. Altogether, cell-free mRNA profiling represents transcriptionally active living tumor cells and proposes a promising approach for detecting, localizing, and identifying biomarkers in different cancers [149].
miRNAs are usually 18-25 nucleotides long non-coding RNAs of endogenous origin, involved in the post-transcriptional regulation of gene expression through translational repression or mRNA destabilization, and can function either as an oncogene or tumor suppressor [155]. In healthy tissues, miRNAs regulate diverse biological functions such as cell proliferation, development, metabolism, differentiation, and intercellular communication. Therefore, perturbation in miRNA expression can significantly impact cell growth, differentiation, and tumor cell apoptosis [156]. Several studies have shown the role of circulating miRNAs in the initiation and progression of different cancer types, including brain tumors [157, 158]. For instance, the levels of miR21, miR115b, miR-23a, and miR-146b species in pediatric juvenile pilocytic astrocytoma patients can predict the tumor nodule sizes and the response to therapies with 100% specificity and sensitivity of 86% [159]. A comparative study between matched tumor and blood samples of pediatric astrocytoma patients has revealed disease-specific miR-130 upregulation and downregulation of miR-145 and miR-335 [160]. In other independent studies, significantly higher levels of miR-21 and miR-15b, and lower quantities of miR-128 and miR-342-3p have been observed in grade 4 glioma patients [161, 162]. Moreover, high-grade gliomas can be distinguished from low-grade based on differential enrichment of miR-29, miR-125b, miR-497, and miR-16 in the blood [158, 163]. Also, the elevated levels of the miR-221/222 family of miRNAs in the plasma are negatively correlated with the overall survival of glioma patients [164]. Qu et al. performed a meta-analysis to gauge the diagnostic potential of miRNAs as a glioma biomarker and identified miR-21 as the most significant and reproducible miRNA [165]. In conclusion, the inconsistencies and reproducibility of miRNA repertoire in cancer patients limit its potential as a biomarker.
Through tumor biopsy studies, the role of non-coding RNAs and tRNA-derived fragments (tRFs) in various cancer types has been known for a long time [166,167,168]. For instance, higher expression of RNAs such as nlr-20-2IHZI73Z, tRF-18-HRE9XFD2, and tRF-22-WB86N7O52 are positively correlated with IDH mutant pathology (astrocytoma and oligodendroglioma) and confers improved survival of glioma patients [169]. However, the exploration of these rare RNA species as biomarkers using liquid biopsy approaches is still limited due to the lack of sensitive detection methods. Taken together, addressing the technical and analytical inaccuracies associated with the sample type, the platform used for measurements and data normalization will allow the usage of cfRNA species as a prognostic biomarker with higher confidence [170].
Circulating protein biomarkers
The correlation of tumor pathology with the expression of proteins and peptides has provided a new search space for novel biomarkers and therapeutic target identification [171]. Increased secretion of proteins in cancer patients may result in elevated levels of circulating proteins (CPs) in various biological fluids, including the blood [15]. The high degree of quantitative disparity in the concentrations of abundant proteins and CPs secreted in the blood limits the detection and clinical usefulness of CPs [15, 171]. Currently, most clinical centers routinely measure tumor protein markers such as PSA, CEA, CA15-3, CA125, CA19-9, CYFRA21-1, S100, NSE, ProGRP, sHER2, SCCA, HE-4, CA72-4, etc., for various cancer types. The proteomics profiling of cancer patients has suggested a few potential circulating tumor protein biomarkers including circulating nucleosomes, thymidine kinases, immunogenic cell death markers, soluble receptors of advanced glycation end products (sRAGE), DNAse activity and high-mobility group box 1 (HMGB1) [172, 173].
Previously, several attempts had been made to identify brain malignancies-specific circulating proteins. Kikuchi et al. were the first to report blood-based protein biomarkers in brain tumors. They showed higher levels of immunosuppressive acidic proteins (IAP) like alpha-1 antitrypsin and alpha-1 acidic glycoprotein, and endothelial cells-derived thrombomodulin and glycoprotein fibronectin in the glioma patients compared to non-glioma and healthy individuals [174]. Besides, the levels of circulating angiogenesis-related proteins such as vascular endothelial growth factor (VEGF), soluble EEGP receptor- (sVEGFR-1), primary fibroblast growth factor (FGF-2), etc., are significantly elevated in different glioma grades [175,176,177]. Other potential circulating protein biomarkers of brain tumors and metastases include tumor cells’ extracellular matrix remodeling proteins such as tissue inhibitors of metalloproteinases (TIMPs) and matrix metalloproteinases (MMPs). These circulating proteins help to classify tumors based on their staging [178]. Also, the plasma levels of interleukins 2(IL-2) and its receptor, neural cell adhesion molecule (NCAM), chitinase-3-like protein 1 (CHI3L1/YKL-40), tumor necrosis factor-alpha (TNFα), neuropeptide Y (NPY) and tumor necrosis factor beta (TNFβ) have diagnostic significance in brain malignancies [179,180,181]. Amongst these, YKL-40 appears as a promising grade 4 glioma prognostic biomarker, inversely associated with overall survival [180, 182]. Plasminogen activator inhibitor-1 (PAI-1) is another marker of interest, as its serum levels are negatively correlated to the progression-free survival (PFS) of brain tumor patients [183]. In addition to early prognosis, CPs can also be implemented to oversee the effectiveness of cancer therapies. For instance, brain tumor patients treated with irinotecan and bevacizumab have decreased plasma levels of VEGF protein (measured after 8 weeks) with an improvement in OS and PFS [184]. Overall, the proteomics studies inferring potential circulating tumor-specific protein biomarkers in blood have contributed substantially to our understanding, but their translational relevance in clinics needs further exploration.
Circulating metabolic biomarkers
Similar to proteome studies, liquid biopsy-based metabolome profiling is also used to identify and quantify different classes of compounds in the biofluids of cancer patients [185]. The circulating metabolites include amino acids, carbohydrates, nucleosides, nucleotides, lipids, vitamins, fatty acids, etc., and are mostly involved in the maintenance of cellular architecture and signal transduction through secondary messenger molecules [186, 187]. The metabolic reprogramming in brain tumors is known to compromise both catabolic and anabolic processes and ultimately signaling pathways involved in various cellular effector functions, leading to the development of tumors and resistance to treatment in the long run [188, 189]. For instance, in plasma metabolic profiling of 159 high-grade brain tumor samples, methionine, arginine, and kynurenine were found to be significantly associated with survival outcomes. While methionine and arginine are positive prognostic markers, kynurenine (intermediate of tryptophan metabolism) is associated with poor survival outcomes [190]. In another study, the levels of metabolites uridine and ornithine were found to be significantly different between low and higher-grade glioma patients [191]. Moreover, as compared to healthy controls, the serum of primary grade 4 patients was observed to contain a higher concentration of antioxidant properties compounds i.e., α- and γ-tocopherol, proposing them as the latent markers of high-grade brain tumor progression [192].
Because of the higher amounts of lipids in the brain, and their involvement in critical biological functions like lipid membrane formation, energy metabolism, and signal transduction, lipidomics has emerged as one of the specialized arms of brain tumor metabolomics studies [193]. However, unlike other neurological disorders, studies involving a liquid biopsy approach toward identifying and characterizing brain tumor-specific lipid biomarkers are very few. For example, Zhou et al. have recently identified 11 plasma lipids as candidate diagnostic biomarkers of malignant brain tumors [194]. Moreover, a landmark lipid profiling study involving 99 glioblastoma tumor tissues has identified >500 unique significantly different lipid species, thus proposing new avenues of lipid biomarker identifications [195, 196]. In conclusion, recent advancements in metabolite detection strategies and the integration of prediction models into lipid biomarker screening workflows offer unprecedented biomarker screening opportunities [197].
Extracellular vesicle (EV)
EV is a lipid-bilayer-bound organelle released into the extracellular space by both tumor and healthy cells. Under physiological conditions, the rate of EV generation of tumor cells is much higher than that of normal cells [198]. Based on the size, morphology, and method of generation, EVs are broadly classified as exosomes, microvesicles, and apoptotic bodies [199, 200]. As an analyte, EV offers several advantages over other liquid biopsy-derived substrates [14]. For instance, EV contains a variety of biomolecules such as DNA, coding and non-coding RNA, miRNA, lipids, proteins, metabolites, etc., and is present in almost every biofluid including blood, CSF, semen, urine, saliva, and ascites [201,202,203,204]. The biological stability of EV allows its storage without degradation at different temperatures [201]. The contents of EV give a more precise representation of the altered biological processes than the ctDNA arising from apoptotic bodies because their parental cells are living [205]. Moreover, EV can manifest surface markers specific to the parental cell of origin, thus helping to predict organ-specific metastases [206]. Finally, the frequency of detecting cancer mutations on DNA enclosed inside EV is much higher than on ctDNA [207, 208].
In cancer, extracellular vesicles (EVs) are suggested to facilitate tumor progression and propagation by increasing cell proliferation, extracellular matrix remodeling, promoting angiogenesis, modulating immune responses, and eventually, metastasis [209,210,211,212,213]. Quantitatively, EVs derived from tumor cells have been shown to correlate with prognosis in various cancer types [214, 215]. Tumor origin exosome’s DNA has been shown to harbor KRAS, EGFR, BRAF, and TP53 mutations in pancreatic, non-small cell lung carcinoma, melanoma, and CRC, respectively [216,217,218]. Several studies have also used EVs collected using liquid biopsy approaches from brain tumor patients as a source of the analyte. For example, specific EGFR gene mutations were detected in DNA isolated from serum-derived EVs in glioblastoma patients [210]. While the mRNA, microRNAs (miR-320 and miR-574-3p), and noncoding RNA (RNU6-1) expression patterns in glioblastoma patients were found to be distinct as compared to the normal healthy individuals, some studies have also shown a panel of surface proteins of EVs as distinguishing feature among brain tumor subtypes [219]. Glioma patients with a higher quantity of tumor-associated molecules such as podoplanin and EGFRvIII in their circulating microvesicles are more prone to chemoradiation therapy failures (radiotherapy plus temozolomide) [220]. In general, the tumor recurrence post-resection is also coupled with the increase in EV concentration in the plasma of patients with brain malignancies [221]. Furthermore, the high-grade gliomas-derived exosomes have been shown to stimulate tumor growth and neoangiogenesis, implying the potential of EVs to mediate metastasis [222]. Similarly, the proteomic profiling of serum-derived EVs from histologically defined medulloblastoma patients indicated their potential role in cancer cell proliferation and migration and suggested tumor-repressor activity of transcription factor hepatocyte nuclear factor 4 alpha (HNF4A) [223]. Thus, the EVs-based liquid biopsy approach provides an opportunity to explore a variety of biomolecules of tumor origin for biomarker identification and monitor therapeutic responses in CNS malignancies [14]. However, further investigation of EVs-based biomarkers is required in large and diverse cohorts to increase the sensitivity, specificity, and reproducibility before its routine clinical application can be considered.
Tumor-educated platelets (TEPs)
Recently, TEPs have been added as a cancer biomarker resource to the blood-based liquid biopsies [2]. An increase in platelet counts (thrombocytosis) is associated with poor survival in various malignant tumors, including mesothelioma, melanoma, breast, lung, and ovarian cancers. A retrospective examination of platelet concentrations in 122 glioma patients predominantly glioblastoma cases (88/122) at longitudinal time points (pre-operative, pre-RT, pre-adjuvant temozolomide (TMZ), and after 2 cycles TMZ) showed a correlation of changes in platelet concentrations during therapy with the survival outcomes [224]. Several previous studies have also shown the prognostic significance of platelet aggregation in brain malignancies. During tumorigenesis and metastasis, the platelets from the blood function as the localized and systemic responders, and get exposed to tumor cells leading to their altered behavior [225,226,227,228]. Besides education through platelet-cancer cell direct interaction, platelets can also uptake RNA and proteins released by tumor cells [227, 229,230,231,232,233,234]. In two independent studies, glioma and other cancer subtypes patients’ platelets RNA profiles have been detected to be altered [229, 235]. Education of platelets in different cancer types is reflected in terms of distinct differentially spliced RNA profiles as compared to healthy individuals [230]. Using these TEPs spliced RNA profiles, the origin of different types of primary cancers, which include pancreatic cancer, breast cancer, colorectal cancer, non-small cell lung carcinoma [NSCLC], and localized high-grade brain tumors, has been successfully identified with an accuracy of 71% [230]. In a recent study, spliced RNA profiles from TEPs were shown to distinguish glioblastoma patients from healthy individuals, brain metastases, and inflammatory processes like multiple sclerosis with an average precision of 85%. Moreover, the genetic arrangement in TEPs (e.g., EGFRvIII rearrangement) can act as an onco-signature for localized high-grade brain tumors [230, 236]. Also, TEPs-specific RNA signatures of glioblastoma patients were observed to be correlated with the tumor volume and its recurrence, which in turn may help in differentiating true tumor progression from false positive advancement [237]. Hence, TEPs offer alternative blood-based diagnostic and monitoring opportunities in brain tumor patients.
Brain tumor-specific biomarkers in CSF
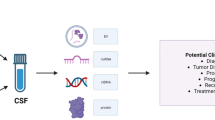
In addition to peripheral blood, CSF-based liquid biopsy is critical in the diagnosis, monitoring, and biomarker identification of CNS tumors [238]. The presence of BBB restricts the movement of CNS tumor-specific analytes in the systemic circulation, thus limiting their detection in cancer patient serum/plasma. In contrast, CSF that comes in direct contact with primary CNS tumors are enriched in tumor-originated CTCs, nucleic acids, proteins, metabolites, etc, [115, 239, 240]. A major limitation of CSF-based liquid biopsy procedures is the requirement of lumbar puncturing to collect CSF from cancer patients [14].
Recent studies have revealed the presence of CTCs in CSF of adult as well as pediatric brain tumor patients. Using antibodies against cytoplasmic GFAP protein, CTCs have been successfully isolated from CSF of various types of pediatric brain tumor patients, and the overall detection has been observed to be more efficient than in systemic circulation [91, 92, 97, 241, 242]. Single-cell RNA sequencing of CTCs sequestered from CSF of lung adenocarcinoma leptomeningeal metastases has led to the identification of metastatic-CTCs signature genes enriched for cell adhesion molecules and metabolic pathway categories [243]. Compared to the reference samples, CSF of HER2+ leptomeningeal metastasis patients possess enhanced levels of CTCs, and these changes have been observed 2-3 months before they appear on magnetic resonance imaging (MRI) or CSF cytology. Thus, the enumeration of CSF CTCs may offer a more dynamic and quantitative assessment of tumor burden, leading to successful CNS tumor diagnosis [244].
As compared to plasma, there is a 10-fold enrichment of tumor-derived DNA in CSF (CSF-tDNA), and its detection partially depends on the tumor location [239, 245]. Cancer-specific mutations including single nucleotide variants of NRAS, BRAF, EGFR, AKT1, and KRAS have been identified in CSF-derived cfDNA of primary and metastatic brain tumor patients [246]. In a cohort of 35 patients, tumor-specific mutational profiling of cfDNA from CSF successfully identified different types of CNS malignancies in 74% of cases [239]. In contrast to low-grade lesions, more mutations were observed in high-grade brain tumor lesions probably due to a higher mutational burden. Hence, the possibility of detecting cancer-related mutations in high-grade lesions is more as compared to that of low-grade lesions. In another study, matched tissue, blood, and CSF from 57 brain tumor patients were collected and analyzed. In 82.5% of these CSF-tDNA samples, at least one tumor-specific mutation was identified, which is concordant with those observed in the primary tumors. In brain-stem gliomas that reside close to the CSF reservoir, a higher percentage of alterations in CSF-tDNA has been detected [246]. The utility of CSF-tDNA has also been studied concerning response to therapies and tumor evolution [3, 247]. For instance, a decrease in the levels of CSF-tDNA after resection was observed, which corroborated with the radiographic evidence in response to chemotherapy. The CSF-tDNA level increases again with increasing radiographic progression. Moreover, sequential biopsies of CSF are very helpful in understanding tumor evolution [3]. Highly diverse mutational profiles of the initial CSF-tDNA samples were observed especially within genes coding for growth factors signaling pathways [247]. Missense and amplification mutations of EGFR genes were observed in the initial sampling, whereas the platelet-derived growth factor receptor alpha (PDFRA) gene mutations appeared in subsequent CSF sampling time points without any alteration in EGFR. These outcomes thus suggest the partial concordance of mutations detected between sequential biopsies over time [248,249,250,251,252,253]. Additionally, CSF-tDNA has also been used to identify copy number variations in glioma patients [254, 255]. Somatic copy number variations in CSF-tDNA have been observed only in a subset of glioma patients [256]. Furthermore, during tumorigenesis, CSF-tDNA from matched CSF and tumor samples have also been used to study alteration in the methylation patterns of promoter regions of MGMT, THBS1, TIMP-3, TERT, and p-16 genes [127, 129, 257,258,259].
Like CSF-tDNA, cfRNA in CSF can also be used as one of the detection modalities for cancer. Most commonly the cfRNA species are encapsulated within extracellular vesicles such as exosomes [260,261,262]. The ability to detect the fusion genes is one of the prime benefits associated with these RNA-based approaches in liquid biopsy [238]. For example, the fusion gene KIAA: BRAF was identified in pilocytic astrocytoma by profiling cfRNA in CSF [263]. In addition, miRNAs isolated from CSF also provide crucial information about various CNS malignancies. While primary diffuse B-cell lymphomas of CNS are characterized by the presence of miR-21, miR-19, and miR-92a, glioma patients showed overexpression of miR-15b and miR-21 [162, 264]. The integrated analysis of miR-15b and miR-21 expression has the potential to differentiate glioma patients from primary CNS lymphomas, leptomeningeal carcinomas, and brain metastasis [162]. In grade 4 glioma patients, miR-10b and miR-21 are present at higher levels in the CSF [265].
CNS tumors also release proteins into the CSF and can be detected by mass spectrometry and ELISA [238]. In contrast to cfDNA or cfRNA, identification of protein biomarkers specific to malignancies is difficult because the alterations in the baseline levels of proteins in CSF may be either because of disease or an outcome of the inflammation or unrelated sufferings [238]. Intracranial germ cell malignancies are the most significant example where specific CSF proteins have an impact on CNS tumor care [266]. For instance, elevated levels of beta human chorionic gonadotropin (bHCG) and alpha-fetoprotein (AFP) proteins have been associated with intracranial germ cell tumors [267]. Levels of both these proteins have been currently used to diagnose a tumor, monitor response to therapy, and predict recurrence. In independent studies, 19 upregulated proteins were identified reproducibly in different types of CNS malignancies [268, 269]. Further studies showed that elevated levels of four peptides (alpha- 1-antichymotrypsin, osteopontin, transthyretin, and the N-terminal residue of albumin) in the CSF of glioma patients, and the reduced levels of prostaglandin D2 synthase in the CSF of pediatric medulloblastoma patients can distinguish these disease pathologies from healthy individuals [270, 271]. Moreover, compared to control patients, CSF of glioma patients has also been reported to be differentially enriched with metabolites such as lactic acid, malic acid, succinate, phosphoenolpyruvate, etc., in an IDH-stratified manner [272]. Taken together, CSF is a highly significant and very promising biofluid for biomarkers identification, therapy monitoring, and tumor recurrence prediction, especially in the case of CNS tumors.
Brain tumor-specific biomarkers in other biofluids
In addition to blood and CSF, vitreous fluid and urine samples have also been used as a source of analytes to determine brain tumor-specific biomarkers [85]. For example, vitreous fluid has been used for the diagnosis of lymphoid malignancies of the brain i.e., primary central nervous system lymphoma (PCNSL). The spread of PCNSL to the retina and vitreous of the eyes is known as vitreoretinal lymphoma (VRL). For PCNSL diagnostic purposes, vitreous fluid offers easy collection and decreased invasiveness compared to other liquid biopsy approaches for brain tumors. Targeted next-generation sequencing on matched vitreous and brain samples has led to the discovery of unique sets of common and distinct genetic alterations both in PCNSL and VRL patients, which in turn suggests the common origin of both the lymphomas. Therefore, vitreous liquid biopsy can be used for the treatment of PCNSL/VRL through accurate diagnosis [273]. In addition, the presence of tumor cells and interleukins ratio (IL10/IL6) ratio of more than 1 in the vitreous fluid also indicate the VRL condition [274]. The mutational profiling of ctDNA from vitreous fluid detected specific mutations in the signaling protein MYD88 in 69% of VRL patients [275]. Besides, the targeted sequencing of 16 genes of VRL patients, revealed copy number variations in CDKN2A or PTEN, and at least one-point mutations impacting the MYD88 gene [276].
Urine has been considered an ideal biomarker resource for early disease detection, as it gathers all the systemic changes within the body. As compared to healthy individuals, significantly elevated levels of MMP-2, MMP-9, neutrophil gelatinase-associated proteins (NGAP), and vascular endothelial growth factor (VEGF) have been reported in the urine of brain tumor patients [277]. These identified urinary biomarkers were used to design a panel containing MMPs and associated proteins, capable of predicting new brain tumors in the early stages of development and monitoring brain malignancies after treatment [277]. The proteomic analysis of glioma patients has revealed a considerable change in the level of twenty-seven urinary proteins after tumor resection, and several of them have been previously correlated with glioma [278]. Functions like angiogenesis and autophagy, which have been formerly associated with glioma development were significantly enriched in these identified proteins. After the targeted proteomics validation, another glioma detection biomarker panel of six proteins, including AHSG, LEG1, CALR, TSP4, MDHM, and AACT, was designed [278]. Also, a diagnostic model has recently been developed based on the expression pattern of specific urinary microRNAs. This model can distinguish CNS tumor patients from noncancerous patients with a specificity of 97% and sensitivity of 100% [279]. On the whole, additional studies need to be conducted before urine and vitreous fluid along with other alternate biofluids can be used as the preferred resource for biomarkers identification, and monitoring disease response and therapeutic efficacy.
Clinical applications, challenges and technological advances in liquid biopsy
The liquid biopsy approach has opened the window of opportunities in brain tumor patient management. Similar to other cancer types, in clinical settings, this strategy can be implemented for brain tumor early detection, staging, identification of treatment targets, defining personalized therapeutic strategies, longitudinal monitoring of disease progression and therapy in advanced stages, and understanding resistance mechanisms [12]. However, most of these applications in brain malignancies are restricted due to the presence of BBB and the inherent limitations associated with each of the analytes [19]. Currently, liquid biopsy research in the context of brain tumors is mainly focused on increasing the availability of analytes in circulation and developing sensitive and specific analytical tools.
Zhu et al. used focused ultrasound (FUS) to transiently disrupt BBB, and enhance the release of brain tumor biomarkers in blood circulation. The presence of tumor-derived eGFP mRNA in blood suggested that FUS-induced BBB disruption could augment brain-blood trafficking. In another advanced study, spatial precision to create temporary brain openings was achieved using low-intensity MR in conjunction with FUS [280]. However, these transient openings allow bidirectional movement of biological entities, leading to the access of systemic circulation to the brain. Therefore, further explorations are required to achieve the regulated unidirectional release of biomarkers from the brain to the blood-only [18, 281].
In the last few years, the evolving field of liquid biopsy has witnessed unprecedented advances toward reducing technical and biological variabilities, leading to the development of more sensitive and specific analytical strategies. Collectively, these efforts aim to achieve an analyte of interest in the purest form and extract maximum insights about the tumor by minimizing the effect of confounding or background signals. Lately, integrated platforms for CTC enrichment, detection, and characterization have been developed to prevent sample loss. The in vivo detection or capture of CTCs using a capture wire, intravascular aphaeretic system, and acoustics have the potential to overcome the CTC’s low count problem [282,283,284]. Moreover, epithelial ImmunoSPOT (EPISPOT) and its microfluidic version i.e., EPIDROP have enabled the investigations of CTCs at a single-cell resolution, thus defining tumor cell heterogeneity from biofluids of cancer patients [285, 286]. Several targeted and untargeted approaches for the precise detection of ctDNA have also been developed namely BEAMing Safe-Sequencing System (BEAMing Safe-SeqS), Cancer Personalized Profiling by deep sequencing (CAPP-Seq), Tagged Amplicon deep Sequencing (TamSeq), and digital PCR. All these ultrasensitive technologies can provide insights into tumor-related single nucleotide variations, copy number variations, changes in methylation patterns, etc [287,288,289,290]. Furthermore, recent technological advances can also filter out ctDNA mutations contributed by factors such as age, leukocytes, cfDNA-sequencing, etc. from those of tumor origin [291,292,293].
Unlike cfDNA reproducible isolation protocols, cfRNA sequestering and recovery from biofluids are challenging. Even from the same biofluid aliquot, different RNA isolation methods result in the enrichment of distinct RNA species. With the commercial availability of spike-ins and control libraries, the technical variations due to PCR amplifications and batch effects can be minimized now [294]. In addition, the application of cross-validation and model-simplification strategies prevents the overfitting problem of cfRNA-based ML biomarker detection methods. Most of the efforts in cfRNA research are focused on extracting pure cfRNA fractions from EVs and lipoprotein complexes, enrichment of poly-A tail lacking distinct coding and non-coding cfRNA fragments, preventing library preparation biases due to cfRNA secondary structure and enzyme affinity, and studying differential epigenomic profiles of cfRNA [149, 294]. The lack of FDA-approved cfRNA-based tests in clinics defines both the experimental difficulties as well as immense possibilities within this research arena.
In a coherent approach, multimodal assays defining protein/metabolites and transcript composition have also been developed. For instance, advanced techniques like cellular indexing of transcriptomes and epitopes by sequencing (CITE-seq) and proximity extension assays (PEAs) can identify proteins and transcripts from CTCs simultaneously [295, 296]. Also, cleavage under targets and tagmentation (scCUT & Tag) and chromatin accessibility assay for transposase-accessible chromatin (ATAC) with select antigen profiling by sequencing (ASAP–seq) can provide insights about protein levels and chromatin accessibility in CTCs concurrently [297, 298]. Moreover, circulating proteins/metabolites in biofluids can be studied using various mass spectrometry-based techniques such as MALDI-TOF, SELDI-TOF, etc. Alternatively, the targeted approaches using antibodies and aptamers (short peptide sequence) can also be used to identify the level of known circulating proteins/metabolites biomarkers [299]. Cytometry by time-of-flight (CyTOF) that uses non-biological metal isotopes with precise mass spectrometry parameters can be used to quantify cell surface and intracellular components of tumor cells [300]. Single CTC proteome can also be studied using unique mass bar-code for multiple samples (18 samples at a time) by the technique known as isobaric labeling [301]. Plasma proteomics-specific advancements and their application to precision medicine have been covered in detail in a series of recent articles [302, 303].
TEP-derived RNA panels have also been frequently used for biomarker screening purposes in several cancer types. However, minimizing the contribution of potential contaminants such as erythrocytes, leukocytes and residual plasma components which include EVs, cfDNA, etc. is essential to prevent the activation of platelets during pure sample preparation, storage, and transport, and obtain TEP-specific RNA profiles [304]. The recently developed pan-cancer particle swarm optimization (PSO)-enhanced thromboSeq algorithm trained on TEP-derived RNA profiles can detect 18 different cancer types with high (99%) specificity, but for various non-cancer diseases its specificity is low (78%) [305]. Also, this algorithm and TEP RNA profiles can be used to identify the tumor site of origin. However, before bringing the thromboSeq test to clinics, independent validation studies at population strata are required.
Exosomes promote tumor development and progression by facilitating the intercellular transfer of bioactive molecules from parent cells with altered pathological states to recipient cells. Therefore, examining cancerous exosome analytes provides new opportunities for biomarker identification. While the number of exosomes in circulation is very large compared to CTCs and ctDNA, identifying and isolating cancerous exosomes and detecting different analytes they contain is challenging. In a recent review article, Yu et al. present a detailed description of the technological advances in exosome separation and analyte detection techniques in detail [306]. However, studies discussing the translational relevance of exosomes originating from brain tumors are very few and need further investigation [307, 308].
In conclusion, overcoming the limitations of the availability of brain-tumor-derived analytes in circulation and the development of ultrasensitive detection and characterization techniques will allow the successful implementation of liquid biopsy clinical applications in the context of brain malignancies.
Liquid biopsy-based brain-tumor-specific clinical trials
In contrast to other cancer types, brain tumor clinical studies are more challenging, and there is a lack of participation. Additionally, issues like brain tumor subtype, tumor size, tumor stage, the extent of BBB disruption, inherent sensitivity, specificity, and clinical utility associated with liquid biopsy markers, and biofluid used for analyte collection greatly impact the study design. Therefore, these confounding factors need to be addressed with precision. Currently, clinical trials involving liquid biopsy and brain tumors are very few, this count goes further down for glioblastoma cases. Various liquid biopsy-based clinical trials involving glioblastoma patients in the USA are listed in Table 2. Detailed description for each of these clinical trials is available at https://beta.clinicaltrials.gov (as of 22nd July 2023).
Conclusion and future perspectives
Emerging liquid biopsy strategies are currently being deployed to detect prognostic and diagnostic markers in biofluids as well as monitor disease progression, response to therapies, and treatment planning. In several disease pathologies, liquid biopsy has even replaced invasive methods like tissue biopsy and image-based methods. The significance of the liquid biopsy-based assays becomes more relevant to cancer types where the size of the tumor and its location is a major concern such as CNS tumors, especially brain tumors. Moreover, routinely used MRI scans are cost-intensive, and the resection of brain tumor tissue may lead to neurological deficits, thus compromising the quality of life of cancer patients. Therefore, liquid biopsy appears to be the promising approach toward brain tumor diagnosis and treatment management.
Liquid biopsy procedures used to explore brain malignancies are typically based on four types of biofluids (blood, CSF, vitreous, and urine), and multiple categories of biomarkers (tumor cells, nucleic acids, proteins, metabolites, extracellular vesicles, and platelets). The diagnostic and prognostic strategies, mostly aim at defining diverse genetic, epigenetic, transcriptomic, proteomic, and metabolomics alterations in brain tumor patients as compared to healthy individuals. While significant progress has been made toward clinical applications of liquid biopsies in other cancer types, these implementations have not been yet employed as a standard of care neuro-oncology exercise for brain tumor patients. Incorporation of WHO CNS5 recommended key genes/molecules/pathways involved in glioblastoma pathogenesis can also be helpful in designing high-precision diagnostic approaches. Also, addressing the technological challenges and enriching the understanding of brain tumor biology in the context of liquid biopsy will help to achieve these goals quickly.
At the technological front, increasing availability of liquid biopsy analytes in biofluids of brain tumor patients, development of standard and reproducible processes of sample collection, improved specificity and the sensitivity of tumor-associated signal detection, and the employment of analyte-specific tailored downstream analytical and bioinformatics techniques need absolute attention. Additionally, brain tumor-specific biological insights like distinct roles of CTCs as single cells and clusters (CTCs-clusters as well as CTCs-WBC clusters), estimation of genetic mutation frequency on ctDNA in context to biofluid and ctDNA size profiles, the significance of cfRNA modifications and tRNA fragments, the role of various secondary messenger molecules in altering transcriptome profiles of platelets and effector outcomes of alternate splice forms, and EVs contribution to metastasis and its utility as drug delivery vehicle in the brain will complement the efforts regarding the technology and assays development. In conclusion, all these discovery, exploration, and validation efforts will be expensive and time-consuming, but their clinical implementations are expected to be cost-effective. Besides, the liquid biopsy research fraternity intends to reach a consensus regarding analyte and biofluid with more biological information for specific cancer types.
Finally, the introduction of liquid biopsy brain tumor-specific multi-national/continental initiatives/societies/consortiums will help to orient the specialized scientific community towards common goals and increase public awareness about participation in brain tumor clinical trials. Global organizations such as IBRO (https://ibro.org) and ABTA (https://www.abta.org), etc. can act coherently to establish brain-tumor-specific liquid biopsy functional bodies similar to that of already existing initiatives such as BloodPac (https://www.bloodpac.org), Cancer-ID (https://www.imi.europa.eu/projects-results/project-factsheets/cancer-id), PANCAID (https://www.pancaid-project.eu), European Liquid Biopsy Society (ELBS) (https://www.uke.de/english/departments-institutes/institutes/tumor-biology/european-liquid-biopsy-society-elbs/index.html), Hellenic Society of Liquid Biopsy (http://en.actc-lab.chem.uoa.gr/hellenic-society-of-liquid-biopsy.html), Liquid Biopsy Consortium (https://prevention.cancer.gov/major-programs/liquid-biopsy-consortium), International Society of Liquid Biopsy (ISLB) (https://islb.info), etc. Altogether, these concerted efforts will facilitate the quick realization of the proposed milestone of ‘liquid biopsy as the standard care in brain malignancies’.
References
Corcoran RB, Chabner BA. Application of cell-free DNA analysis to cancer treatment. N Engl J Med. 2018;379:1754–65.
Poulet G, Massias J, Taly V. Liquid biopsy: general concepts. Acta Cytol. 2019;63:449–55.
De Mattos-Arruda L, Mayor R, Ng CKY, Weigelt B, Martínez-Ricarte F, Torrejon D, et al. Cerebrospinal fluid-derived circulating tumour DNA better represents the genomic alterations of brain tumours than plasma. Nat Commun. 2015;6:8839.
Wang Y, Springer S, Mulvey CL, Silliman N, Schaefer J, Sausen M, et al. Detection of somatic mutations and HPV in the saliva and plasma of patients with head and neck squamous cell carcinomas. Sci Transl Med. 2015;7:293ra104.
Di Meo A, Bartlett J, Cheng Y, Pasic MD, Yousef GM. Liquid biopsy: a step forward towards precision medicine in urologic malignancies. Mol Cancer. 2017;16:80.
Schwarzenbach H, Nishida N, Calin GA, Pantel K. Clinical relevance of circulating cell-free microRNAs in cancer. Nat Rev Clin Oncol. 2014;11:145–56.
Heitzer E, Ulz P, Geigl JB. Circulating tumor DNA as a liquid biopsy for cancer. Clin Chem. 2015;61:112–23.
Alix-Panabières C, Pantel K. Clinical applications of circulating tumor cells and circulating tumor DNA as liquid biopsy. Cancer Discov. 2016;6:479–91.
Duffy MJ, McDermott EW, Crown J. Blood-based biomarkers in breast cancer: from proteins to circulating tumor cells to circulating tumor DNA. Tumour Biol. 2018;40:1010428318776169.
Yu W, Hurley J, Roberts D, Chakrabortty SK, Enderle D, Noerholm M, et al. Exosome-based liquid biopsies in cancer: opportunities and challenges. Ann Oncol. 2021;32:466–77.
Kalluri R, LeBleu VS. The biology, function, and biomedical applications of exosomes. Science. 2020;367:eaau6977.
Alix-Panabières C, Pantel K. Liquid biopsy: from discovery to clinical application. Cancer Discov. 2021;11:858–73.
Sato Y, Matoba R, Kato K. Recent advances in liquid biopsy in precision oncology research. Biol Pharm Bull. 2019;42:337–42.
Bunda S, Zuccato JA, Voisin MR, Wang JZ, Nassiri F, Patil V, et al. Liquid biomarkers for improved diagnosis and classification of CNS tumors. Int J Mol Sci. 2021;22:4548.
Jelski W, Mroczko B. Molecular and circulating biomarkers of brain tumors. Int J Mol Sci. 2021;22:7039.
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2019. CA Cancer J Clin. 2019;69:7–34.
Yan PF, Yan L, Zhang Z, Salim A, Wang L, Hu TT, et al. Accuracy of conventional MRI for preoperative diagnosis of intracranial tumors: a retrospective cohort study of 762 cases. Int J Surg. 2016;36:109–17.
Zhu L, Cheng G, Ye D, Nazeri A, Yue Y, Liu W, et al. Focused ultrasound-enabled brain tumor liquid biopsy. Sci Rep. 2018;8:6553.
Touat M, Duran-Peña A, Alentorn A, Lacroix L, Massard C, Idbaih A. Emerging circulating biomarkers in glioblastoma: promises and challenges. Expert Rev Mol Diagn. 2015;15:1311–23.
WHO classifications of tumors editorial board. Central nervous system tumors. International agency for research on cancer; 2022;5:1–584.
Louis DN, Perry A, Wesseling P, Brat DJ, Cree IA, Figarella-Branger D, et al. The 2021 WHO Classification of Tumors of the Central Nervous System: a summary. Neuro Oncol. 2021;23:1231–51.
Whitfield BT, Huse JT. Classification of adult-type diffuse gliomas: Impact of the World Health Organization 2021 update. Brain Pathol. 2022;32:e13062.
Aquilanti E, Miller J, Santagata S, Cahill DP, Brastianos PK. Updates in prognostic markers for gliomas. Neuro Oncol. 2018;20:vii17–vii26.
Senhaji N, Squalli Houssaini A, Lamrabet S, Louati S, Bennis S. Molecular and circulating biomarkers in patients with glioblastoma. Int J Mol Sci. 2022;23:7474.
Szopa W, Burley TA, Kramer-Marek G, Kaspera W. Diagnostic and therapeutic biomarkers in glioblastoma: current status and future perspectives. Biomed Res Int. 2017;2017:8013575.
Lee SM, Koh HJ, Park DC, Song BJ, Huh TL, Park JW. Cytosolic NADP(+)-dependent isocitrate dehydrogenase status modulates oxidative damage to cells. Free Radic Biol Med. 2002;32:1185–96.
Yan H, Parsons DW, Jin G, McLendon R, Rasheed BA, Yuan W, et al. IDH1 and IDH2 mutations in gliomas. N Engl J Med. 2009;360:765–73.
Losman JA, Kaelin WG Jr. What a difference a hydroxyl makes: mutant IDH, (R)-2-hydroxyglutarate, and cancer. Genes Dev. 2013;27:836–52.
SongTao Q, Lei Y, Si G, YanQing D, HuiXia H, XueLin Z, et al. IDH mutations predict longer survival and response to temozolomide in secondary glioblastoma. Cancer Sci. 2012;103:269–73.
Johannessen TA, Mukherjee J, Viswanath P, Ohba S, Ronen SM, Bjerkvig R, et al. Rapid conversion of mutant IDH1 from driver to passenger in a model of human gliomagenesis. Mol Cancer Res. 2016;14:976–83.
Han S, Liu Y, Cai SJ, Qian M, Ding J, Larion M, et al. IDH mutation in glioma: molecular mechanisms and potential therapeutic targets. Br J Cancer. 2020;122:1580–9.
Houillier C, Wang X, Kaloshi G, Mokhtari K, Guillevin R, Laffaire J, et al. IDH1 or IDH2 mutations predict longer survival and response to temozolomide in low-grade gliomas. Neurology. 2010;75:1560–6.
Brennan CW, Verhaak RG, McKenna A, Campos B, Noushmehr H, Salama SR, et al. The somatic genomic landscape of glioblastoma. Cell. 2013;155:462–77.
An Z, Aksoy O, Zheng T, Fan QW, Weiss WA. Epidermal growth factor receptor and EGFRvIII in glioblastoma: signaling pathways and targeted therapies. Oncogene. 2018;37:1561–75.
Congdon KL, Gedeon PC, Suryadevara CM, Caruso HG, Cooper LJ, Heimberger AB, et al. Epidermal growth factor receptor and variant III targeted immunotherapy. Neuro Oncol. 2014;16:viii20–5.
Zhang LY, Ge HJ, Wang LM, Zhao LH, Liu L, Zhang DJ, et al. Prognostic implication of alterations in epidermal growth factor receptor and MGMT in glioblastoma. Zhonghua Bing Li Xue Za Zhi. 2019;48:186–91.
Tichy J, Spechtmeyer S, Mittelbronn M, Hattingen E, Rieger J, Senft C, et al. Prospective evaluation of serum glial fibrillary acidic protein (GFAP) as a diagnostic marker for glioblastoma. J Neurooncol. 2016;126:361–9.
Jung CS, Foerch C, Schänzer A, Heck A, Plate KH, Seifert V, et al. Serum GFAP is a diagnostic marker for glioblastoma multiforme. Brain. 2007;130:3336–41.
Müller C, Holtschmidt J, Auer M, Heitzer E, Lamszus K, Schulte A, et al. Hematogenous dissemination of glioblastoma multiforme. Sci Transl Med. 2014;6:247ra101.
Zhang H, Yuan F, Qi Y, Liu B, Chen Q. Circulating tumor cells for glioma. Front Oncol. 2021;11:607150.
England B, Huang T, Karsy M. Current understanding of the role and targeting of tumor suppressor p53 in glioblastoma multiforme. Tumour Biol. 2013;34:2063–74.
von Deimling A, Eibl RH, Ohgaki H, Louis DN, von Ammon K, Petersen I, et al. p53 mutations are associated with 17p allelic loss in grade II and grade III astrocytoma. Cancer Res. 1992;52:2987–90.
Guimaraes DP, Hainaut P. TP53: a key gene in human cancer. Biochimie. 2002;84:83–93.
Brázdová M, Quante T, Tögel L, Walter K, Loscher C, Tichý V, et al. Modulation of gene expression in U251 glioblastoma cells by binding of mutant p53 R273H to intronic and intergenic sequences. Nucleic Acids Res. 2009;37:1486–500.
Petitjean A, Achatz MI, Borresen-Dale AL, Hainaut P, Olivier M. TP53 mutations in human cancers: functional selection and impact on cancer prognosis and outcomes. Oncogene. 2007;26:2157–65.
Furnari FB, Fenton T, Bachoo RM, Mukasa A, Stommel JM, Stegh A, et al. Malignant astrocytic glioma: genetics, biology, and paths to treatment. Genes Dev. 2007;21:2683–710.
Cancer genome atlas research network. Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature. 2008;455:1061–8.
Hao X, Bahia RK, Cseh O, Bozek DA, Blake S, Rinnenthal J, et al. BI-907828, a novel potent MDM2 inhibitor, inhibits glioblastoma brain tumor stem cells in vitro and prolongs survival in orthotopic xenograft mouse models. Neuro Oncol. 2023;25:913–26.
Chai R, Li G, Liu Y, Zhang K, Zhao Z, Wu F, et al. Predictive value of MGMT promoter methylation on the survival of TMZ treated IDH-mutant glioblastoma. Cancer Biol Med. 2021;18:272–82.
Mulholland S, Pearson DM, Hamoudi RA, Malley DS, Smith CM, Weaver JM, et al. MGMT CpG island is invariably methylated in adult astrocytic and oligodendroglial tumors with IDH1 or IDH2 mutations. Int J Cancer. 2012;131:1104–13.
Patel M, Vogelbaum MA, Barnett GH, Jalali R, Ahluwalia MS. Molecular targeted therapy in recurrent glioblastoma: current challenges and future directions. Expert Opin Investig Drugs. 2012;21:1247–66.
Gerson SL. MGMT: its role in cancer aetiology and cancer therapeutics. Nat Rev Cancer. 2004;4:296–307.
Radke J, Koch A, Pritsch F, Schumann E, Misch M, Hempt C, et al. Predictive MGMT status in a homogeneous cohort of IDH wildtype glioblastoma patients. Acta Neuropathol Commun. 2019;7:89.
Kim M, Yoo J, Chang JH, Kim SH. Association of MGMT gene promoter methylation with clinicopathological parameters in patients with wild-type IDH glioblastoma. Anticancer Res. 2022;42:335–41.
Thon N, Kreth S, Kreth FW. Personalized treatment strategies in glioblastoma: MGMT promoter methylation status. Onco Targets Ther. 2013;6:1363–72.
Horn S, Figl A, Rachakonda PS, Fischer C, Sucker A, Gast A, et al. TERT promoter mutations in familial and sporadic melanoma. Science. 2013;339:959–61.
Killela PJ, Reitman ZJ, Jiao Y, Bettegowda C, Agrawal N, Diaz LA Jr., et al. TERT promoter mutations occur frequently in gliomas and a subset of tumors derived from cells with low rates of self-renewal. Proc Natl Acad Sci USA 2013;110:6021–6.
Huang FW, Hodis E, Xu MJ, Kryukov GV, Chin L, Garraway LA. Highly recurrent TERT promoter mutations in human melanoma. Science. 2013;339:957–9.
Amen AM, Fellmann C, Soczek KM, Ren SM, Lew RJ, Knott GJ, et al. Cancer-specific loss of TERT activation sensitizes glioblastoma to DNA damage. Proc Natl Acad Sci USA 2021;118:e2008772118.
Mosrati MA, Malmström A, Lysiak M, Krysztofiak A, Hallbeck M, Milos P, et al. TERT promoter mutations and polymorphisms as prognostic factors in primary glioblastoma. Oncotarget. 2015;6:16663–73.
Giunco S, Padovan M, Angelini C, Cavallin F, Cerretti G, Morello M, et al. Prognostic role and interaction of TERT promoter status, telomere length and MGMT promoter methylation in newly diagnosed IDH wild-type glioblastoma patients. ESMO Open. 2023;8:101570.
Corless BC, Chang GA, Cooper S, Syeda MM, Shao Y, Osman I, et al. Development of novel mutation-specific droplet digital PCR assays detecting TERT promoter mutations in tumor and plasma samples. J Mol Diagn. 2019;21:274–85.
Fujisawa H, Kurrer M, Reis RM, Yonekawa Y, Kleihues P, Ohgaki H. Acquisition of the glioblastoma phenotype during astrocytoma progression is associated with loss of heterozygosity on 10q25-qter. Am J Pathol. 1999;155:387–94.
Nakamura M, Ishida E, Shimada K, Kishi M, Nakase H, Sakaki T, et al. Frequent LOH on 22q12.3 and TIMP-3 inactivation occur in the progression to secondary glioblastomas. Lab Invest. 2005;85:165–75.
Haruna M, Ueyama A, Yamamoto Y, Hirata M, Goto K, Yoshida H, et al. The impact of CCR8+ regulatory T cells on cytotoxic T cell function in human lung cancer. Sci Rep. 2022;12:5377.
Brat DJ, Verhaak RG, Aldape KD, Yung WK, Salama SR, Cooper LA, et al. Comprehensive, integrative genomic analysis of diffuse lower-grade gliomas. N Engl J Med. 2015;372:2481–98.
Kannan K, Inagaki A, Silber J, Gorovets D, Zhang J, Kastenhuber ER, et al. Whole-exome sequencing identifies ATRX mutation as a key molecular determinant in lower-grade glioma. Oncotarget. 2012;3:1194–203.
Leeper HE, Caron AA, Decker PA, Jenkins RB, Lachance DH, Giannini C. IDH mutation, 1p19q codeletion and ATRX loss in WHO grade II gliomas. Oncotarget. 2015;6:30295–305.
Koschmann C, Lowenstein PR, Castro MG. ATRX mutations and glioblastoma: Impaired DNA damage repair, alternative lengthening of telomeres, and genetic instability. Mol Cell Oncol. 2016;3:e1167158.
Amorim JP, Santos G, Vinagre J, Soares P. The role of ATRX in the alternative lengthening of telomeres (ALT) phenotype. Genes. 2016;7:66.
Ramamoorthy M, Smith S. Loss of ATRX suppresses resolution of telomere cohesion to control recombination in ALT cancer cells. Cancer Cell. 2015;28:357–69.
Lovejoy CA, Li W, Reisenweber S, Thongthip S, Bruno J, de Lange T, et al. Loss of ATRX, genome instability, and an altered DNA damage response are hallmarks of the alternative lengthening of telomeres pathway. PLoS Genet. 2012;8:e1002772.
Heaphy CM, Subhawong AP, Hong SM, Goggins MG, Montgomery EA, Gabrielson E, et al. Prevalence of the alternative lengthening of telomeres telomere maintenance mechanism in human cancer subtypes. Am J Pathol. 2011;179:1608–15.
Nandakumar P, Mansouri A, Das S. The role of ATRX in glioma biology. Front Oncol. 2017;7:236.
Pi C, Zhang MF, Peng XX, Zhang YC, Xu CR, Zhou Q. Liquid biopsy in non-small cell lung cancer: a key role in the future of personalized medicine? Expert Rev Mol Diagn. 2017;17:1089–96.
De Mattos-Arruda L, Caldas C. Cell-free circulating tumour DNA as a liquid biopsy in breast cancer. Mol Oncol. 2016;10:464–74.
Jia S, Zhang R, Li Z, Li J. Clinical and biological significance of circulating tumor cells, circulating tumor DNA, and exosomes as biomarkers in colorectal cancer. Oncotarget. 2017;8:55632–45.
Rehman AU, Khan P, Maurya SK, Siddiqui JA, Santamaria-Barria JA, Batra SK, et al. Liquid biopsies to occult brain metastasis. Mol Cancer. 2022;21:113.
Castro-Giner F, Aceto N. Tracking cancer progression: from circulating tumor cells to metastasis. Genome Med. 2020;12:31.
Masuda T, Hayashi N, Iguchi T, Ito S, Eguchi H, Mimori K. Clinical and biological significance of circulating tumor cells in cancer. Mol Oncol. 2016;10:408–17.
Chen L, Bode AM, Dong Z. Circulating tumor cells: moving biological insights into detection. Theranostics. 2017;7:2606–19.
Siravegna G, Marsoni S, Siena S, Bardelli A. Integrating liquid biopsies into the management of cancer. Nat Rev Clin Oncol. 2017;14:531–48.
Szczerba BM, Castro-Giner F, Vetter M, Krol I, Gkountela S, Landin J, et al. Neutrophils escort circulating tumour cells to enable cell cycle progression. Nature. 2019;566:553–7.
Chen J, Luo Y, Xi X, Li H, Li S, Zheng L, et al. Circulating tumor cell associated white blood cell cluster as a biomarker for metastasis and recurrence in hepatocellular carcinoma. Front Oncol. 2022;12:931140.
Fontanilles M, Duran-Peña A, Idbaih A. Liquid biopsy in primary brain tumors: looking for stardust! Curr Neurol Neurosci Rep. 2018;18:13.
Alix-Panabières C, Pantel K. Technologies for detection of circulating tumor cells: facts and vision. Lab Chip. 2014;14:57–62.
Allard WJ, Matera J, Miller MC, Repollet M, Connelly MC, Rao C, et al. Tumor cells circulate in the peripheral blood of all major carcinomas but not in healthy subjects or patients with nonmalignant diseases. Clin Cancer Res. 2004;10:6897–904.
Ross AA, Cooper BW, Lazarus HM, Mackay W, Moss TJ, Ciobanu N, et al. Detection and viability of tumor cells in peripheral blood stem cell collections from breast cancer patients using immunocytochemical and clonogenic assay techniques. Blood. 1993;82:2605–10.
Wang ZP, Eisenberger MA, Carducci MA, Partin AW, Scher HI, Ts’o PO. Identification and characterization of circulating prostate carcinoma cells. Cancer. 2000;88:2787–95.
Müller Bark J, Kulasinghe A, Chua B, Day BW, Punyadeera C. Circulating biomarkers in patients with glioblastoma. Br J Cancer. 2020;122:295–305.
Bang-Christensen SR, Pedersen RS, Pereira MA, Clausen TM, Løppke C, Sand NT, et al. Capture and detection of circulating glioma cells using the recombinant VAR2CSA malaria protein. Cells. 2019;8:998.
Chistiakov DA, Chekhonin VP. Circulating tumor cells and their advances to promote cancer metastasis and relapse, with focus on glioblastoma multiforme. Exp Mol Pathol. 2018;105:166–74.
Macarthur KM, Kao GD, Chandrasekaran S, Alonso-Basanta M, Chapman C, Lustig RA, et al. Detection of brain tumor cells in the peripheral blood by a telomerase promoter-based assay. Cancer Res. 2014;74:2152–9.
Gao F, Cui Y, Jiang H, Sui D, Wang Y, Jiang Z, et al. Circulating tumor cell is a common property of brain glioma and promotes the monitoring system. Oncotarget. 2016;7:71330–40.
Sullivan JP, Nahed BV, Madden MW, Oliveira SM, Springer S, Bhere D, et al. Brain tumor cells in circulation are enriched for mesenchymal gene expression. Cancer Discov. 2014;4:1299–309.
Liu T, Xu H, Huang M, Ma W, Saxena D, Lustig RA, et al. Circulating glioma cells exhibit stem cell-like properties. Cancer Res. 2018;78:6632–42.
Shankar GM, Balaj L, Stott SL, Nahed B, Carter BS. Liquid biopsy for brain tumors. Expert Rev Mol Diagn. 2017;17:943–7.
Kitz J, Lowes LE, Goodale D, Allan AL. Circulating tumor cell analysis in preclinical mouse models of metastasis. Diagnostics. 2018;8:30.
Pantel K, Alix-Panabières C. Functional studies on viable circulating tumor cells. Clin Chem. 2016;62:328–34.
Pantel K, Alix-Panabières C. Cell lines from circulating tumor cells. Oncoscience. 2015;2:815–6.
Maheswaran S, Haber DA. Ex vivo culture of CTCs: an emerging resource to guide cancer therapy. Cancer Res. 2015;75:2411–5.
Adamczyk LA, Williams H, Frankow A, Ellis HP, Haynes HR, Perks C, et al. Current understanding of circulating tumor cells - potential value in malignancies of the central nervous system. Front Neurol. 2015;6:174.
Jahr S, Hentze H, Englisch S, Hardt D, Fackelmayer FO, Hesch RD, et al. DNA fragments in the blood plasma of cancer patients: quantitations and evidence for their origin from apoptotic and necrotic cells. Cancer Res. 2001;61:1659–65.
Diehl F, Li M, Dressman D, He Y, Shen D, Szabo S, et al. Detection and quantification of mutations in the plasma of patients with colorectal tumors. Proc Natl Acad Sci USA 2005;102:16368–73.
Diehl F, Schmidt K, Choti MA, Romans K, Goodman S, Li M, et al. Circulating mutant DNA to assess tumor dynamics. Nat Med. 2008;14:985–90.
Stroun M, Lyautey J, Lederrey C, Olson-Sand A, Anker P. About the possible origin and mechanism of circulating DNA apoptosis and active DNA release. Clin Chim Acta. 2001;313:139–42.
El Messaoudi S, Mouliere F, Du Manoir S, Bascoul-Mollevi C, Gillet B, Nouaille M, et al. Circulating DNA as a strong multimarker prognostic tool for metastatic colorectal cancer patient management care. Clin Cancer Res. 2016;22:3067–77.
Steffensen KD, Madsen CV, Andersen RF, Waldstrøm M, Adimi P, Jakobsen A. Prognostic importance of cell-free DNA in chemotherapy resistant ovarian cancer treated with bevacizumab. Eur J Cancer. 2014;50:2611–8.
Yanagita M, Redig AJ, Paweletz CP, Dahlberg SE, O’Connell A, Feeney N, et al. A prospective evaluation of circulating tumor cells and cell-free DNA in EGFR-mutant non-small cell lung cancer patients treated with erlotinib on a phase II trial. Clin Cancer Res. 2016;22:6010–20.
Rossi G, Mu Z, Rademaker AW, Austin LK, Strickland KS, Costa RLB, et al. Cell-Free DNA and circulating tumor cells: comprehensive liquid biopsy analysis in advanced breast cancer. Clin Cancer Res. 2018;24:560–8.
Bonner ER, Bornhorst M, Packer RJ, Nazarian J. Liquid biopsy for pediatric central nervous system tumors. NPJ Precis Oncol. 2018;2:29.
Wang J, Bettegowda C. Applications of DNA-based liquid biopsy for central nervous system neoplasms. J Mol Diagn. 2017;19:24–34.
Dawson SJ, Tsui DW, Murtaza M, Biggs H, Rueda OM, Chin SF, et al. Analysis of circulating tumor DNA to monitor metastatic breast cancer. N. Engl J Med. 2013;368:1199–209.
Newman AM, Bratman SV, To J, Wynne JF, Eclov NC, Modlin LA, et al. An ultrasensitive method for quantitating circulating tumor DNA with broad patient coverage. Nat Med. 2014;20:548–54.
Bettegowda C, Sausen M, Leary RJ, Kinde I, Wang Y, Agrawal N, et al. Detection of circulating tumor DNA in early- and late-stage human malignancies. Sci Transl Med. 2014;6:224ra24.
Beaver JA, Jelovac D, Balukrishna S, Cochran R, Croessmann S, Zabransky DJ, et al. Detection of cancer DNA in plasma of patients with early-stage breast cancer. Clin Cancer Res. 2014;20:2643–50.
Diaz LA Jr., Bardelli A. Liquid biopsies: genotyping circulating tumor DNA. J Clin Oncol. 2014;32:579–86.
Rothé F, Laes JF, Lambrechts D, Smeets D, Vincent D, Maetens M, et al. Plasma circulating tumor DNA as an alternative to metastatic biopsies for mutational analysis in breast cancer. Ann Oncol. 2014;25:1959–65.
Narayan A, Carriero NJ, Gettinger SN, Kluytenaar J, Kozak KR, Yock TI, et al. Ultrasensitive measurement of hotspot mutations in tumor DNA in blood using error-suppressed multiplexed deep sequencing. Cancer Res. 2012;72:3492–8.
Higgins MJ, Jelovac D, Barnathan E, Blair B, Slater S, Powers P, et al. Detection of tumor PIK3CA status in metastatic breast cancer using peripheral blood. Clin Cancer Res. 2012;18:3462–9.
Board RE, Wardley AM, Dixon JM, Armstrong AC, Howell S, Renshaw L, et al. Detection of PIK3CA mutations in circulating free DNA in patients with breast cancer. Breast Cancer Res Treat. 2010;120:461–7.
Thierry AR, Mouliere F, El Messaoudi S, Mollevi C, Lopez-Crapez E, Rolet F, et al. Clinical validation of the detection of KRAS and BRAF mutations from circulating tumor DNA. Nat Med. 2014;20:430–5.
Punnoose EA, Atwal S, Liu W, Raja R, Fine BM, Hughes BG, et al. Evaluation of circulating tumor cells and circulating tumor DNA in non-small cell lung cancer: association with clinical endpoints in a phase II clinical trial of pertuzumab and erlotinib. Clin Cancer Res. 2012;18:2391–401.
Forshew T, Murtaza M, Parkinson C, Gale D, Tsui DW, Kaper F, et al. Noninvasive identification and monitoring of cancer mutations by targeted deep sequencing of plasma DNA. Sci Transl Med. 2012;4:136ra68.
Shinozaki M, O’Day SJ, Kitago M, Amersi F, Kuo C, Kim J, et al. Utility of circulating B-RAF DNA mutation in serum for monitoring melanoma patients receiving biochemotherapy. Clin Cancer Res. 2007;13:2068–74.
Piccioni DE, Achrol AS, Kiedrowski LA, Banks KC, Boucher N, Barkhoudarian G, et al. Analysis of cell-free circulating tumor DNA in 419 patients with glioblastoma and other primary brain tumors. CNS Oncol. 2019;8:Cns34.
Majchrzak-Celińska A, Paluszczak J, Kleszcz R, Magiera M, Barciszewska AM, Nowak S, et al. Detection of MGMT, RASSF1A, p15INK4B, and p14ARF promoter methylation in circulating tumor-derived DNA of central nervous system cancer patients. J Appl Genet. 2013;54:335–44.
Balańa C, Carrato C, Ramírez JL, Cardona AF, Berdiel M, Sánchez JJ, et al. Tumour and serum MGMT promoter methylation and protein expression in glioblastoma patients. Clin Transl Oncol. 2011;13:677–85.
Balaña C, Ramirez JL, Taron M, Roussos Y, Ariza A, Ballester R, et al. O6-methyl-guanine-DNA methyltransferase methylation in serum and tumor DNA predicts response to 1,3-bis(2-chloroethyl)-1-nitrosourea but not to temozolamide plus cisplatin in glioblastoma multiforme. Clin Cancer Res. 2003;9:1461–8.
Weaver KD, Grossman SA, Herman JG. Methylated tumor-specific DNA as a plasma biomarker in patients with glioma. Cancer Invest. 2006;24:35–40.
Wakabayashi T, Natsume A, Hatano H, Fujii M, Shimato S, Ito M, et al. p16 promoter methylation in the serum as a basis for the molecular diagnosis of gliomas. Neurosurgery. 2009;64:455–61.
Lavon I, Refael M, Zelikovitch B, Shalom E, Siegal T. Serum DNA can define tumor-specific genetic and epigenetic markers in gliomas of various grades. Neuro Oncol. 2010;12:173–80.
Boisselier B, Gállego Pérez-Larraya J, Rossetto M, Labussière M, Ciccarino P, Marie Y, et al. Detection of IDH1 mutation in the plasma of patients with glioma. Neurology. 2012;79:1693–8.
Salkeni MA, Zarzour A, Ansay TY, McPherson CM, Warnick RE, Rixe O, et al. Detection of EGFRvIII mutant DNA in the peripheral blood of brain tumor patients. J Neurooncol. 2013;115:27–35.
Muralidharan K, Yekula A, Small JL, Rosh ZS, Kang KM, Wang L, et al. TERT promoter mutation analysis for blood-based diagnosis and monitoring of gliomas. Clin Cancer Res. 2021;27:169–78.
Chen J, Huan W, Zuo H, Zhao L, Huang C, Liu X, et al. Alu methylation serves as a biomarker for non-invasive diagnosis of glioma. Oncotarget. 2016;7:26099–106.
Shen SY, Singhania R, Fehringer G, Chakravarthy A, Roehrl MHA, Chadwick D, et al. Sensitive tumour detection and classification using plasma cell-free DNA methylomes. Nature. 2018;563:579–83.
Nuzzo PV, Berchuck JE, Korthauer K, Spisak S, Nassar AH, Abou Alaiwi S, et al. Detection of renal cell carcinoma using plasma and urine cell-free DNA methylomes. Nat Med. 2020;26:1041–3.
Nassiri F, Chakravarthy A, Feng S, Shen SY, Nejad R, Zuccato JA, et al. Detection and discrimination of intracranial tumors using plasma cell-free DNA methylomes. Nat Med. 2020;26:1044–7.
Thomson JP, Meehan RR. The application of genome-wide 5-hydroxymethylcytosine studies in cancer research. Epigenomics. 2017;9:77–91.
Wilkins OM, Johnson KC, Houseman EA, King JE, Marsit CJ, Christensen BC. Genome-wide characterization of cytosine-specific 5-hydroxymethylation in normal breast tissue. Epigenetics. 2020;15:398–418.
Walker NJ, Rashid M, Yu S, Bignell H, Lumby CK, Livi CM, et al. Hydroxymethylation profile of cell-free DNA is a biomarker for early colorectal cancer. Sci Rep. 2022;12:16566.
He B, Zhang C, Zhang X, Fan Y, Zeng H, Liu J, et al. Tissue-specific 5-hydroxymethylcytosine landscape of the human genome. Nat Commun. 2021;12:4249.
Cui XL, Nie J, Ku J, Dougherty U, West-Szymanski DC, Collin F, et al. A human tissue map of 5-hydroxymethylcytosines exhibits tissue specificity through gene and enhancer modulation. Nat Commun. 2020;11:6161.
Johnson KC, Houseman EA, King JE, von Herrmann KM, Fadul CE, Christensen BC. 5-Hydroxymethylcytosine localizes to enhancer elements and is associated with survival in glioblastoma patients. Nat Commun. 2016;7:13177.
Kopreski MS, Benko FA, Gocke CD. Circulating RNA as a tumor marker: detection of 5T4 mRNA in breast and lung cancer patient serum. Ann N. Y Acad Sci. 2001;945:172–8.
Castellanos-Rizaldos E, Zhang X, Tadigotla VR, Grimm DG, Karlovich C, Raez LE, et al. Exosome-based detection of activating and resistance EGFR mutations from plasma of non-small cell lung cancer patients. Oncotarget. 2019;10:2911–20.
Perhavec A, Cerkovnik P, Novakovic S, Zgajnar J. The hTERT mRNA in plasma samples of early breast cancer patients, non-cancer patients and healthy individuals. Neoplasma. 2008;55:549–54.
Larson MH, Pan W, Kim HJ, Mauntz RE, Stuart SM, Pimentel M, et al. A comprehensive characterization of the cell-free transcriptome reveals tissue- and subtype-specific biomarkers for cancer detection. Nat Commun. 2021;12:2357.
Sorber L, Zwaenepoel K, Jacobs J, De Winne K, Goethals S, Reclusa P, et al. Circulating cell-free DNA and RNA analysis as liquid biopsy: optimal centrifugation protocol. Cancers. 2019;11:458.
Arroyo JD, Chevillet JR, Kroh EM, Ruf IK, Pritchard CC, Gibson DF, et al. Argonaute2 complexes carry a population of circulating microRNAs independent of vesicles in human plasma. Proc Natl Acad Sci USA 2011;108:5003–8.
Godoy PM, Bhakta NR, Barczak AJ, Cakmak H, Fisher S, MacKenzie TC, et al. Large differences in small RNA composition between human biofluids. Cell Rep. 2018;25:1346–58.
Kopreski MS, Benko FA, Kwak LW, Gocke CD. Detection of tumor messenger RNA in the serum of patients with malignant melanoma. Clin Cancer Res. 1999;5:1961–5.
Raez LE, Danenberg K, Sumarriva D, Usher J, Sands J, Castrellon A, et al. Using cfRNA as a tool to evaluate clinical treatment outcomes in patients with metastatic lung cancers and other tumors. Cancer Drug Resist. 2021;4:1061–71.
Jonas S, Izaurralde E. Towards a molecular understanding of microRNA-mediated gene silencing. Nat Rev Genet. 2015;16:421–33.
Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D, et al. MicroRNA expression profiles classify human cancers. Nature. 2005;435:834–8.
Li R, Gao K, Luo H, Wang X, Shi Y, Dong Q, et al. Identification of intrinsic subtype-specific prognostic microRNAs in primary glioblastoma. J Exp Clin Cancer Res. 2014;33:9.
Regazzo G, Terrenato I, Spagnuolo M, Carosi M, Cognetti G, Cicchillitti L, et al. A restricted signature of serum miRNAs distinguishes glioblastoma from lower grade gliomas. J Exp Clin Cancer Res. 2016;35:124.
Bookland M, Tang-Schomer M, Gillan E, Kolmakova A. Circulating serum oncologic miRNA in pediatric juvenile pilocytic astrocytoma patients predicts mural nodule volume. Acta Neurochir. 2018;160:1571–81.
López-Aguilar JE, Velázquez-Flores MA, Simón-Martínez LA, Ávila-Miranda R, Rodríguez-Florido MA, Ruiz-Esparza Garrido R. Circulating microRNAs as biomarkers for pediatric astrocytomas. Arch Med Res. 2017;48:323–32.
Wang Q, Li P, Li A, Jiang W, Wang H, Wang J, et al. Plasma specific miRNAs as predictive biomarkers for diagnosis and prognosis of glioma. J Exp Clin Cancer Res. 2012;31:97.
Baraniskin A, Kuhnhenn J, Schlegel U, Maghnouj A, Zöllner H, Schmiegel W, et al. Identification of microRNAs in the cerebrospinal fluid as biomarker for the diagnosis of glioma. Neuro Oncol. 2012;14:29–33.
Wu J, Li L, Jiang C. Identification and evaluation of serum microRNA-29 family for glioma screening. Mol Neurobiol. 2015;52:1540–6.
Zhang R, Pang B, Xin T, Guo H, Xing Y, Xu S, et al. Plasma miR-221/222 family as novel descriptive and prognostic biomarkers for glioma. Mol Neurobiol. 2016;53:1452–60.
Qu S, Guan J, Liu Y. Identification of microRNAs as novel biomarkers for glioma detection: a meta-analysis based on 11 articles. J Neurol Sci. 2015;348:181–7.
Anfossi S, Babayan A, Pantel K, Calin GA. Clinical utility of circulating non-coding RNAs - an update. Nat Rev Clin Oncol. 2018;15:541–63.
Fu M, Gu J, Wang M, Zhang J, Chen Y, Jiang P, et al. Emerging roles of tRNA-derived fragments in cancer. Mol Cancer. 2023;22:30.
Toden S, Goel A. Non-coding RNAs as liquid biopsy biomarkers in cancer. Br J Cancer. 2022;126:351–60.
Xu B, Liang J, Zou H, Wang J, Xiong Y, Pei J. Identification of Novel tRNA-Leu-CAA-derived tsRNAs for the diagnosis and prognosis of diffuse gliomas. Cancer Manag Res. 2022;14:2609–23.
Chevillet JR, Lee I, Briggs HA, He Y, Wang K. Issues and prospects of microRNA-based biomarkers in blood and other body fluids. Molecules. 2014;19:6080–105.
Tang K, Gardner S, Snuderl M. The role of liquid biopsies in pediatric brain tumors. J Neuropathol Exp Neurol. 2020;79:934–40.
Molina R, Holdenrieder S, Auge JM, Schalhorn A, Hatz R, Stieber P. Diagnostic relevance of circulating biomarkers in patients with lung cancer. Cancer Biomark. 2010;6:163–78.
Wittwer C, Boeck S, Heinemann V, Haas M, Stieber P, Nagel D, et al. Circulating nucleosomes and immunogenic cell death markers HMGB1, sRAGE and DNAse in patients with advanced pancreatic cancer undergoing chemotherapy. Int J Cancer. 2013;133:2619–30.
Kikuchi K, Gotoh H, Kowada M. Immunosuppressive acidic protein in patients with brain tumours: a preliminary report. Acta Neurochir. 1987;86:42–9.
Rafat N, Beck G, Schulte J, Tuettenberg J, Vajkoczy P. Circulating endothelial progenitor cells in malignant gliomas. J Neurosurg. 2010;112:43–9.
Ilhan, Gartner A, Neziri W, Czech D, Base T, Hörl WH W, et al. Angiogenic factors in plasma of brain tumour patients. Anticancer Res. 2009;29:731–6.
Reynés G, Vila V, Martín M, Parada A, Fleitas T, Reganon E, et al. Circulating markers of angiogenesis, inflammation, and coagulation in patients with glioblastoma. J Neurooncol. 2011;102:35–41.
Lin Y, Wang JF, Gao GZ, Zhang GZ, Wang FL, Wang YJ. Plasma levels of tissue inhibitor of matrix metalloproteinase-1 correlate with diagnosis and prognosis of glioma patients. Chin Med J. 2013;126:4295–300.
Yoshida S, Morii K. Serum concentrations of soluble interleukin-2 receptor in patients with malignant brain tumors. J Surg Oncol. 2000;75:131–5.
Bernardi D, Padoan A, Ballin A, Sartori M, Manara R, Scienza R, et al. Serum YKL-40 following resection for cerebral glioblastoma. J Neurooncol. 2012;107:299–305.
Ilhan-Mutlu A, Wagner L, Widhalm G, Wöhrer A, Bartsch S, Czech T, et al. Exploratory investigation of eight circulating plasma markers in brain tumor patients. Neurosurg Rev. 2013;36:45–55.
Hormigo A, Gu B, Karimi S, Riedel E, Panageas KS, Edgar MA, et al. YKL-40 and matrix metalloproteinase-9 as potential serum biomarkers for patients with high-grade gliomas. Clin Cancer Res. 2006;12:5698–704.
Xu BJ, An QA, Srinivasa Gowda S, Yan W, Pierce LA, Abel TW, et al. Identification of blood protein biomarkers that aid in the clinical assessment of patients with malignant glioma. Int J Oncol. 2012;40:1995–2003.
Tabouret E, Boudouresque F, Barrie M, Matta M, Boucard C, Loundou A, et al. Association of matrix metalloproteinase 2 plasma level with response and survival in patients treated with bevacizumab for recurrent high-grade glioma. Neuro Oncol. 2014;16:392–9.
Gonzalez-Covarrubias V, Martínez-Martínez E, Del Bosque-Plata L. The potential of metabolomics in biomedical applications. Metabolites. 2022;12:194.
Pienkowski T, Kowalczyk T, Garcia-Romero N, Ayuso-Sacido A, Ciborowski M. Proteomics and metabolomics approach in adult and pediatric glioma diagnostics. Biochim Biophys Acta Rev Cancer. 2022;1877:188721.
Bratulic S, Limeta A, Dabestani S, Birgisson H, Enblad G, Stålberg K, et al. Noninvasive detection of any-stage cancer using free glycosaminoglycans. Proc Natl Acad Sci USA 2022;119:e2115328119.
Muller Bark J, Karpe AV, Doecke JD, Leo P, Jeffree RL, Chua B, et al. A pilot study: metabolic profiling of plasma and saliva samples from newly diagnosed glioblastoma patients. Cancer Med. 2023;12:11427–37.
Pandey R, Caflisch L, Lodi A, Brenner AJ, Tiziani S. Metabolomic signature of brain cancer. Mol Carcinog. 2017;56:2355–71.
Shen J, Song R, Hodges TR, Heimberger AB, Zhao H. Identification of metabolites in plasma for predicting survival in glioblastoma. Mol Carcinog. 2018;57:1078–84.
Zhao H, Heimberger AB, Lu Z, Wu X, Hodges TR, Song R, et al. Metabolomics profiling in plasma samples from glioma patients correlates with tumor phenotypes. Oncotarget. 2016;7:20486–95.
Björkblom B, Wibom C, Jonsson P, Mörén L, Andersson U, Johannesen TB, et al. Metabolomic screening of pre-diagnostic serum samples identifies association between α- and γ-tocopherols and glioblastoma risk. Oncotarget. 2016;7:37043–53.
Taïb B, Aboussalah AM, Moniruzzaman M, Chen S, Haughey NJ, Kim SF, et al. Lipid accumulation and oxidation in glioblastoma multiforme. Sci Rep. 2019;9:19593.
Zhou J, Ji N, Wang G, Zhang Y, Song H, Yuan Y, et al. Metabolic detection of malignant brain gliomas through plasma lipidomic analysis and support vector machine-based machine learning. EBioMedicine. 2022;81:104097.
Miska J, Chandel NS. Targeting fatty acid metabolism in glioblastoma. J Clin Invest. 2023;133:e163448.
Wang LB, Karpova A, Gritsenko MA, Kyle JE, Cao S, Li Y, et al. Proteogenomic and metabolomic characterization of human glioblastoma. Cancer Cell. 2021;39:509–28.e20.
Neil ZD, Pierzchajlo N, Boyett C, Little O, Kuo CC, Brown NJ, et al. Assessing metabolic markers in glioblastoma using machine learning: a systematic review. Metabolites. 2023;13:161.
Balaj L, Lessard R, Dai L, Cho YJ, Pomeroy SL, Breakefield XO, et al. Tumour microvesicles contain retrotransposon elements and amplified oncogene sequences. Nat Commun. 2011;2:180.
Théry C, Zitvogel L, Amigorena S. Exosomes: composition, biogenesis and function. Nat Rev Immunol. 2002;2:569–79.
Akers JC, Gonda D, Kim R, Carter BS, Chen CC. Biogenesis of extracellular vesicles (EV): exosomes, microvesicles, retrovirus-like vesicles, and apoptotic bodies. J Neurooncol. 2013;113:1–11.
Zhou B, Xu K, Zheng X, Chen T, Wang J, Song Y, et al. Application of exosomes as liquid biopsy in clinical diagnosis. Signal Transduct Target Ther. 2020;5:144.
Srivastava A, Moxley K, Ruskin R, Dhanasekaran DN, Zhao YD, Ramesh R. A non-invasive liquid biopsy screening of urine-derived exosomes for miRNAs as biomarkers in endometrial cancer patients. AAPS J. 2018;20:82.
Wang S, Yang Y, Sun L, Qiao G, Song Y, Liu B. Exosomal microRNAs as liquid biopsy biomarkers in hepatocellular carcinoma. Onco Targets Ther. 2020;13:2021–30.
Otake K, Kamiguchi H, Hirozane Y. Identification of biomarkers for amyotrophic lateral sclerosis by comprehensive analysis of exosomal mRNAs in human cerebrospinal fluid. BMC Med Genomics. 2019;12:7.
Cai X, Janku F, Zhan Q, Fan JB. Accessing genetic information with liquid biopsies. Trends Genet. 2015;31:564–75.
Hoshino A, Costa-Silva B, Shen TL, Rodrigues G, Hashimoto A, Tesic Mark M, et al. Tumour exosome integrins determine organotropic metastasis. Nature. 2015;527:329–35.
Allenson K, Castillo J, San Lucas FA, Scelo G, Kim DU, Bernard V, et al. High prevalence of mutant KRAS in circulating exosome-derived DNA from early-stage pancreatic cancer patients. Ann Oncol. 2017;28:741–7.
Möhrmann L, Huang HJ, Hong DS, Tsimberidou AM, Fu S, Piha-Paul SA, et al. Liquid biopsies using plasma exosomal nucleic acids and plasma cell-free DNA compared with clinical outcomes of patients with advanced cancers. Clin Cancer Res. 2018;24:181–8.
Kim JW, Wieckowski E, Taylor DD, Reichert TE, Watkins S, Whiteside TL. Fas ligand-positive membranous vesicles isolated from sera of patients with oral cancer induce apoptosis of activated T lymphocytes. Clin Cancer Res. 2005;11:1010–20.
Skog J, Würdinger T, van Rijn S, Meijer DH, Gainche L, Sena-Esteves M, et al. Glioblastoma microvesicles transport RNA and proteins that promote tumour growth and provide diagnostic biomarkers. Nat Cell Biol. 2008;10:1470–6.
Wieckowski EU, Visus C, Szajnik M, Szczepanski MJ, Storkus WJ, Whiteside TL. Tumor-derived microvesicles promote regulatory T cell expansion and induce apoptosis in tumor-reactive activated CD8+ T lymphocytes. J Immunol. 2009;183:3720–30.
Cai Z, Yang F, Yu L, Yu Z, Jiang L, Wang Q, et al. Activated T cell exosomes promote tumor invasion via Fas signaling pathway. J Immunol. 2012;188:5954–61.
Al-Nedawi K, Meehan B, Micallef J, Lhotak V, May L, Guha A, et al. Intercellular transfer of the oncogenic receptor EGFRvIII by microvesicles derived from tumour cells. Nat Cell Biol. 2008;10:619–24.
Baran J, Baj-Krzyworzeka M, Weglarczyk K, Szatanek R, Zembala M, Barbasz J, et al. Circulating tumour-derived microvesicles in plasma of gastric cancer patients. Cancer Immunol Immunother. 2010;59:841–50.
Taylor DD, Gercel-Taylor C. MicroRNA signatures of tumor-derived exosomes as diagnostic biomarkers of ovarian cancer. Gynecol Oncol. 2008;110:13–21.
Thakur BK, Zhang H, Becker A, Matei I, Huang Y, Costa-Silva B, et al. Double-stranded DNA in exosomes: a novel biomarker in cancer detection. Cell Res. 2014;24:766–9.
Kahlert C, Melo SA, Protopopov A, Tang J, Seth S, Koch M, et al. Identification of double-stranded genomic DNA spanning all chromosomes with mutated KRAS and p53 DNA in the serum exosomes of patients with pancreatic cancer. J Biol Chem. 2014;289:3869–75.
Melo SA, Luecke LB, Kahlert C, Fernandez AF, Gammon ST, Kaye J, et al. Glypican-1 identifies cancer exosomes and detects early pancreatic cancer. Nature. 2015;523:177–82.
Manterola L, Guruceaga E, Gállego Pérez-Larraya J, González-Huarriz M, Jauregui P, Tejada S, et al. A small noncoding RNA signature found in exosomes of GBM patient serum as a diagnostic tool. Neuro Oncol. 2014;16:520–7.
Shao H, Chung J, Balaj L, Charest A, Bigner DD, Carter BS, et al. Protein typing of circulating microvesicles allows real-time monitoring of glioblastoma therapy. Nat Med. 2012;18:1835–40.
Osti D, Del Bene M, Rappa G, Santos M, Matafora V, Richichi C, et al. Clinical significance of extracellular vesicles in plasma from glioblastoma patients. Clin Cancer Res. 2019;25:266–76.
Kucharzewska P, Christianson HC, Welch JE, Svensson KJ, Fredlund E, Ringnér M, et al. Exosomes reflect the hypoxic status of glioma cells and mediate hypoxia-dependent activation of vascular cells during tumor development. Proc Natl Acad Sci USA 2013;110:7312–7.
Epple LM, Griffiths SG, Dechkovskaia AM, Dusto NL, White J, Ouellette RJ, et al. Medulloblastoma exosome proteomics yield functional roles for extracellular vesicles. PLoS One. 2012;7:e42064.
Boonyawan K, Hess KR, Yang J, Long L, Wang Q, Ezhilarasan R, et al. A relative increase in circulating platelets following chemoradiation predicts for poor survival of patients with glioblastoma. Oncotarget. 2017;8:90488–95.
McAllister SS, Weinberg RA. The tumour-induced systemic environment as a critical regulator of cancer progression and metastasis. Nat Cell Biol. 2014;16:717–27.
Schumacher D, Strilic B, Sivaraj KK, Wettschureck N, Offermanns S. Platelet-derived nucleotides promote tumor-cell transendothelial migration and metastasis via P2Y2 receptor. Cancer Cell. 2013;24:130–7.
Kerr BA, McCabe NP, Feng W, Byzova TV. Platelets govern pre-metastatic tumor communication to bone. Oncogene. 2013;32:4319–24.
Labelle M, Begum S, Hynes RO. Direct signaling between platelets and cancer cells induces an epithelial-mesenchymal-like transition and promotes metastasis. Cancer Cell. 2011;20:576–90.
Nilsson RJ, Balaj L, Hulleman E, van Rijn S, Pegtel DM, Walraven M, et al. Blood platelets contain tumor-derived RNA biomarkers. Blood. 2011;118:3680–3.
Best MG, Sol N, Kooi I, Tannous J, Westerman BA, Rustenburg F, et al. RNA-seq of tumor-educated platelets enables blood-based pan-cancer, multiclass, and molecular pathway cancer diagnostics. Cancer Cell. 2015;28:666–76.
Klement GL, Yip TT, Cassiola F, Kikuchi L, Cervi D, Podust V, et al. Platelets actively sequester angiogenesis regulators. Blood. 2009;113:2835–42.
Nilsson RJ, Karachaliou N, Berenguer J, Gimenez-Capitan A, Schellen P, Teixido C, et al. Rearranged EML4-ALK fusion transcripts sequester in circulating blood platelets and enable blood-based crizotinib response monitoring in non-small-cell lung cancer. Oncotarget. 2016;7:1066–75.
Tjon-Kon-Fat LA, Lundholm M, Schröder M, Wurdinger T, Thellenberg-Karlsson C, Widmark A, et al. Platelets harbor prostate cancer biomarkers and the ability to predict therapeutic response to abiraterone in castration resistant patients. Prostate. 2018;78:48–53.
Luo CL, Xu ZG, Chen H, Ji J, Wang YH, Hu W, et al. LncRNAs and EGFRvIII sequestered in TEPs enable blood-based NSCLC diagnosis. Cancer Manag Res. 2018;10:1449–59.
Calverley DC, Phang TL, Choudhury QG, Gao B, Oton AB, Weyant MJ, et al. Significant downregulation of platelet gene expression in metastatic lung cancer. Clin Transl Sci. 2010;3:227–32.
Varkey J, Nicolaides T. Tumor-educated platelets: a review of current and potential applications in solid tumors. Cureus. 2021;13:e19189.
Sol N, In ‘t Veld S, Vancura A, Tjerkstra M, Leurs C, Rustenburg F, et al. Tumor-educated platelet RNA for the detection and (pseudo)progression monitoring of glioblastoma. Cell Rep Med. 2020;1:100101.
Mattox AK, Yan H, Bettegowda C. The potential of cerebrospinal fluid-based liquid biopsy approaches in CNS tumors. Neuro Oncol. 2019;21:1509–18.
Wang Y, Springer S, Zhang M, McMahon KW, Kinde I, Dobbyn L, et al. Detection of tumor-derived DNA in cerebrospinal fluid of patients with primary tumors of the brain and spinal cord. Proc Natl Acad Sci USA 2015;112:9704–9.
Chen WW, Balaj L, Liau LM, Samuels ML, Kotsopoulos SK, Maguire CA, et al. BEAMing and droplet digital PCR analysis of mutant IDH1 mRNA in glioma patient serum and cerebrospinal fluid extracellular vesicles. Mol Ther Nucleic Acids. 2013;2:e109.
Zhao Y, Jiang F, Wang Q, Wang B, Han Y, Yang J, et al. Cytoplasm protein GFAP magnetic beads construction and application as cell separation target for brain tumors. J Nanobiotechnol. 2020;18:169.
Guerreiro Stucklin AS, Ramaswamy V, Daniels C, Taylor MD. Review of molecular classification and treatment implications of pediatric brain tumors. Curr Opin Pediatr. 2018;30:3–9.
Ruan H, Zhou Y, Shen J, Zhai Y, Xu Y, Pi L, et al. Circulating tumor cell characterization of lung cancer brain metastases in the cerebrospinal fluid through single-cell transcriptome analysis. Clin Transl Med. 2020;10:e246.
Malani R, Fleisher M, Kumthekar P, Lin X, Omuro A, Groves MD, et al. Cerebrospinal fluid circulating tumor cells as a quantifiable measurement of leptomeningeal metastases in patients with HER2 positive cancer. J Neurooncol. 2020;148:599–606.
Panditharatna E, Kilburn LB, Aboian MS, Kambhampati M, Gordish-Dressman H, Magge SN, et al. Clinically relevant and minimally invasive tumor surveillance of pediatric diffuse midline gliomas using patient-derived liquid biopsy. Clin Cancer Res. 2018;24:5850–9.
Pan W, Gu W, Nagpal S, Gephart MH, Quake SR. Brain tumor mutations detected in cerebral spinal fluid. Clin Chem. 2015;61:514–22.
Miller AM, Shah RH, Pentsova EI, Pourmaleki M, Briggs S, Distefano N, et al. Tracking tumour evolution in glioma through liquid biopsies of cerebrospinal fluid. Nature. 2019;565:654–8.
Suzuki H, Aoki K, Chiba K, Sato Y, Shiozawa Y, Shiraishi Y, et al. Mutational landscape and clonal architecture in grade II and III gliomas. Nat Genet. 2015;47:458–68.
Johnson BE, Mazor T, Hong C, Barnes M, Aihara K, McLean CY, et al. Mutational analysis reveals the origin and therapy-driven evolution of recurrent glioma. Science. 2014;343:189–93.
Bai H, Harmancı AS, Erson-Omay EZ, Li J, Coşkun S, Simon M, et al. Integrated genomic characterization of IDH1-mutant glioma malignant progression. Nat Genet. 2016;48:59–66.
Kim J, Lee IH, Cho HJ, Park CK, Jung YS, Kim Y, et al. Spatiotemporal evolution of the primary glioblastoma genome. Cancer Cell. 2015;28:318–28.
Wang J, Cazzato E, Ladewig E, Frattini V, Rosenbloom DI, Zairis S, et al. Clonal evolution of glioblastoma under therapy. Nat Genet. 2016;48:768–76.
Lee JK, Wang J, Sa JK, Ladewig E, Lee HO, Lee IH, et al. Spatiotemporal genomic architecture informs precision oncology in glioblastoma. Nat Genet. 2017;49:594–9.
Leary RJ, Kinde I, Diehl F, Schmidt K, Clouser C, Duncan C, et al. Development of personalized tumor biomarkers using massively parallel sequencing. Sci Transl Med. 2010;2:20ra14.
Leary RJ, Sausen M, Kinde I, Papadopoulos N, Carpten JD, Craig D, et al. Detection of chromosomal alterations in the circulation of cancer patients with whole-genome sequencing. Sci Transl Med. 2012;4:162ra54.
Mouliere F, Mair R, Chandrananda D, Marass F, Smith CG, Su J, et al. Detection of cell-free DNA fragmentation and copy number alterations in cerebrospinal fluid from glioma patients. EMBO Mol Med. 2018;10:e9323.
Liu BL, Cheng JX, Zhang W, Zhang X, Wang R, Lin H, et al. Quantitative detection of multiple gene promoter hypermethylation in tumor tissue, serum, and cerebrospinal fluid predicts prognosis of malignant gliomas. Neuro Oncol. 2010;12:540–8.
Wang Z, Jiang W, Wang Y, Guo Y, Cong Z, Du F, et al. MGMT promoter methylation in serum and cerebrospinal fluid as a tumor-specific biomarker of glioma. Biomed Rep. 2015;3:543–8.
Juratli TA, Stasik S, Zolal A, Schuster C, Richter S, Daubner D, et al. TERT promoter mutation detection in cell-free tumor-derived DNA in patients with IDH wild-type glioblastomas: a pilot prospective study. Clin Cancer Res. 2018;24:5282–91.
Hunter MP, Ismail N, Zhang X, Aguda BD, Lee EJ, Yu L, et al. Detection of microRNA expression in human peripheral blood microvesicles. PLoS One. 2008;3:e3694.
Fernandez-Mercado M, Manterola L, Larrea E, Goicoechea I, Arestin M, Armesto M, et al. The circulating transcriptome as a source of non-invasive cancer biomarkers: concepts and controversies of non-coding and coding RNA in body fluids. J Cell Mol Med. 2015;19:2307–23.
El-Hefnawy T, Raja S, Kelly L, Bigbee WL, Kirkwood JM, Luketich JD, et al. Characterization of amplifiable, circulating RNA in plasma and its potential as a tool for cancer diagnostics. Clin Chem. 2004;50:564–73.
Behling F, Schittenhelm J. Oncogenic BRAF alterations and their role in brain tumors. Cancers. 2019;11:794.
Baraniskin A, Kuhnhenn J, Schlegel U, Chan A, Deckert M, Gold R, et al. Identification of microRNAs in the cerebrospinal fluid as marker for primary diffuse large B-cell lymphoma of the central nervous system. Blood. 2011;117:3140–6.
Teplyuk NM, Mollenhauer B, Gabriely G, Giese A, Kim E, Smolsky M, et al. MicroRNAs in cerebrospinal fluid identify glioblastoma and metastatic brain cancers and reflect disease activity. Neuro Oncol. 2012;14:689–700.
Samuel N, Remke M, Rutka JT, Raught B, Malkin D. Proteomic analyses of CSF aimed at biomarker development for pediatric brain tumors. J Neurooncol. 2014;118:225–38.
Nishizaki T, Kajiwara K, Adachi N, Tsuha M, Nakayama H, Ohshita N, et al. Detection of craniospinal dissemination of intracranial germ cell tumours based on serum and cerebrospinal fluid levels of tumour markers. J Clin Neurosci. 2001;8:27–30.
Shen F, Zhang Y, Yao Y, Hua W, Zhang HS, Wu JS, et al. Proteomic analysis of cerebrospinal fluid: toward the identification of biomarkers for gliomas. Neurosurg Rev. 2014;37:367–80.
Khwaja FW, Reed MS, Olson JJ, Schmotzer BJ, Gillespie GY, Guha A, et al. Proteomic identification of biomarkers in the cerebrospinal fluid (CSF) of astrocytoma patients. J Proteome Res. 2007;6:559–70.
Schuhmann MU, Zucht HD, Nassimi R, Heine G, Schneekloth CG, Stuerenburg HJ, et al. Peptide screening of cerebrospinal fluid in patients with glioblastoma multiforme. Eur J Surg Oncol. 2010;36:201–7.
Rajagopal MU, Hathout Y, MacDonald TJ, Kieran MW, Gururangan S, Blaney SM, et al. Proteomic profiling of cerebrospinal fluid identifies prostaglandin D2 synthase as a putative biomarker for pediatric medulloblastoma: a pediatric brain tumor consortium study. Proteomics. 2011;11:935–43.
Ballester LY, Lu G, Zorofchian S, Vantaku V, Putluri V, Yan Y, et al. Analysis of cerebrospinal fluid metabolites in patients with primary or metastatic central nervous system tumors. Acta Neuropathol Commun. 2018;6:85.
Balikov DA, Hu K, Liu CJ, Betz BL, Chinnaiyan AM, Devisetty LV, et al. Comparative molecular analysis of primary central nervous system lymphomas and matched vitreoretinal lymphomas by vitreous liquid biopsy. Int J Mol Sci. 2021;22:9992.
Pochat-Cotilloux C, Bienvenu J, Nguyen AM, Ohanessian R, Ghesquières H, Sève P, et al. Use of a threshold of interleukin-10 and IL-10/IL-6 ratio in ocular samples for the screening of vitreoretinal lymphoma. Retina. 2018;38:773–81.
Bonzheim I, Giese S, Deuter C, Süsskind D, Zierhut M, Waizel M, et al. High frequency of MYD88 mutations in vitreoretinal B-cell lymphoma: a valuable tool to improve diagnostic yield of vitreous aspirates. Blood. 2015;126:76–9.
Cani AK, Hovelson DH, Demirci H, Johnson MW, Tomlins SA, Rao RC. Next generation sequencing of vitreoretinal lymphomas from small-volume intraocular liquid biopsies: new routes to targeted therapies. Oncotarget. 2017;8:7989–98.
Smith ER, Zurakowski D, Saad A, Scott RM, Moses MA. Urinary biomarkers predict brain tumor presence and response to therapy. Clin Cancer Res. 2008;14:2378–86.
Wu J, Zhang J, Wei J, Zhao Y, Gao Y. Urinary biomarker discovery in gliomas using mass spectrometry-based clinical proteomics. Chin Neurosurg J. 2020;6:11.
Kitano Y, Aoki K, Ohka F, Yamazaki S, Motomura K, Tanahashi K, et al. Urinary MicroRNA-based diagnostic model for central nervous system tumors using nanowire scaffolds. ACS Appl Mater Interfaces. 2021;13:17316–29.
Meng Y, Pople CB, Suppiah S, Llinas M, Huang Y, Sahgal A, et al. MR-guided focused ultrasound liquid biopsy enriches circulating biomarkers in patients with brain tumors. Neuro Oncol. 2021;23:1789–97.
Rincon-Torroella J, Khela H, Bettegowda A, Bettegowda C. Biomarkers and focused ultrasound: the future of liquid biopsy for brain tumor patients. J Neurooncol. 2022;156:33–48.
Galanzha EI, Menyaev YA, Yadem AC, Sarimollaoglu M, Juratli MA, Nedosekin DA, et al. In vivo liquid biopsy using Cytophone platform for photoacoustic detection of circulating tumor cells in patients with melanoma. Sci Transl Med. 2019;11:eaat5857.
Kim TH, Wang Y, Oliver CR, Thamm DH, Cooling L, Paoletti C, et al. A temporary indwelling intravascular aphaeretic system for in vivo enrichment of circulating tumor cells. Nat Commun. 2019;10:1478.
Kuske A, Gorges TM, Tennstedt P, Tiebel AK, Pompe R, Preißer F, et al. Improved detection of circulating tumor cells in non-metastatic high-risk prostate cancer patients. Sci Rep. 2016;6:39736.
Soler A, Cayrefourcq L, Mazel M, Alix-Panabières C. EpCAM-independent enrichment and detection of viable circulating tumor cells using the EPISPOT assay. Methods Mol Biol. 2017;1634:263–76.
Eyer K, Doineau RCL, Castrillon CE, Briseño-Roa L, Menrath V, Mottet G, et al. Single-cell deep phenotyping of IgG-secreting cells for high-resolution immune monitoring. Nat Biotechnol. 2017;35:977–82.
Heitzer E, Haque IS, Roberts CES, Speicher MR. Current and future perspectives of liquid biopsies in genomics-driven oncology. Nat Rev Genet. 2019;20:71–88.
Liu MC, Oxnard GR, Klein EA, Swanton C, Seiden MV. Sensitive and specific multi-cancer detection and localization using methylation signatures in cell-free DNA. Ann Oncol. 2020;31:745–59.
Luo H, Zhao Q, Wei W, Zheng L, Yi S, Li G, et al. Circulating tumor DNA methylation profiles enable early diagnosis, prognosis prediction, and screening for colorectal cancer. Sci Transl Med. 2020;12:eaax7533.
Keller L, Belloum Y, Wikman H, Pantel K. Clinical relevance of blood-based ctDNA analysis: mutation detection and beyond. Br J Cancer. 2021;124:345–58.
Razavi P, Li BT, Brown DN, Jung B, Hubbell E, Shen R, et al. High-intensity sequencing reveals the sources of plasma circulating cell-free DNA variants. Nat Med. 2019;25:1928–37.
Genovese G, Kähler AK, Handsaker RE, Lindberg J, Rose SA, Bakhoum SF, et al. Clonal hematopoiesis and blood-cancer risk inferred from blood DNA sequence. N Engl J Med. 2014;371:2477–87.
Chan HT, Nagayama S, Chin YM, Otaki M, Hayashi R, Kiyotani K, et al. Clinical significance of clonal hematopoiesis in the interpretation of blood liquid biopsy. Mol Oncol. 2020;14:1719–30.
Cabús L, Lagarde J, Curado J, Lizano E, Pérez-Boza J. Current challenges and best practices for cell-free long RNA biomarker discovery. Biomark Res. 2022;10:62.
Stoeckius M, Hafemeister C, Stephenson W, Houck-Loomis B, Chattopadhyay PK, Swerdlow H, et al. Simultaneous epitope and transcriptome measurement in single cells. Nat Methods. 2017;14:865–8.
Wik L, Nordberg N, Broberg J, Björkesten J, Assarsson E, Henriksson S, et al. Proximity extension assay in combination with next-generation sequencing for high-throughput proteome-wide analysis. Mol Cell Proteom. 2021;20:100168.
Bartosovic M, Kabbe M, Castelo-Branco G. Single-cell CUT&Tag profiles histone modifications and transcription factors in complex tissues. Nat Biotechnol. 2021;39:825–35.
Mimitou EP, Lareau CA, Chen KY, Zorzetto-Fernandes AL, Hao Y, Takeshima Y, et al. Scalable, multimodal profiling of chromatin accessibility, gene expression and protein levels in single cells. Nat Biotechnol. 2021;39:1246–58.
Veyssière H, Bidet Y, Penault-Llorca F, Radosevic-Robin N, Durando X. Circulating proteins as predictive and prognostic biomarkers in breast cancer. Clin Proteom. 2022;19:25.
Spitzer MH, Nolan GP. Mass cytometry: single cells, many features. Cell. 2016;165:780–91.
Specht H, Emmott E, Petelski AA, Huffman RG, Perlman DH, Serra M, et al. Single-cell proteomic and transcriptomic analysis of macrophage heterogeneity using SCoPE2. Genome Biol. 2021;22:50.
Yu X, Schwenk JM, Xu P, LaBaer J. Advances in plasma proteomics: moving from technology to precision medicine. Proteom Clin Appl. 2022;16:e2200083.
Bennett HM, Stephenson W, Rose CM, Darmanis S. Single-cell proteomics enabled by next-generation sequencing or mass spectrometry. Nat Methods. 2023;20:363–74.
In ‘t Veld S, Arkani M, Post E, Antunes-Ferreira M, D’Ambrosi S, Vessies DCL, et al. Detection and localization of early- and late-stage cancers using platelet RNA. Cancer Cell. 2022;40:999–1009.e6.
Best MG, In ‘t Veld S, Sol N, Wurdinger T. RNA sequencing and swarm intelligence-enhanced classification algorithm development for blood-based disease diagnostics using spliced blood platelet RNA. Nat Protoc. 2019;14:1206–34.
Yu D, Li Y, Wang M, Gu J, Xu W, Cai H, et al. Exosomes as a new frontier of cancer liquid biopsy. Mol Cancer. 2022;21:56.
Karami Fath M, Azami J, Masoudi A, Mosaddeghi Heris R, Rahmani E, Alavi F, et al. Exosome-based strategies for diagnosis and therapy of glioma cancer. Cancer Cell Int. 2022;22:262.
Del Bene M, Osti D, Faletti S, Beznoussenko GV, DiMeco F, Pelicci G. Extracellular vesicles: the key for precision medicine in glioblastoma. Neuro Oncol. 2022;24:184–96.
Ohba S, Kuwahara K, Yamada S, Abe M, Hirose Y. Correlation between IDH, ATRX, and TERT promoter mutations in glioma. Brain Tumor Pathol. 2020;37:33–40.
Qin T, Mullan B, Ravindran R, Messinger D, Siada R, Cummings JR, et al. ATRX loss in glioma results in dysregulation of cell-cycle phase transition and ATM inhibitor radio-sensitization. Cell Rep. 2022;38:110216.
Cohen AL, Colman H. Glioma biology and molecular markers. Cancer Treat Res. 2015;163:15–30.
Cao Y, Li X, Kong S, Shang S, Qi Y. CDK4/6 inhibition suppresses tumour growth and enhances the effect of temozolomide in glioma cells. J Cell Mol Med. 2020;24:5135–45.
Baker SJ, Reddy EP. CDK4: a key player in the cell cycle, development, and cancer. Genes Cancer. 2012;3:658–69.
Kurokawa R, Kurokawa M, Baba A, Ota Y, Pinarbasi E, Camelo-Piragua S, et al. Major changes in 2021 World Health Organization classification of central nervous system tumors. Radiographics. 2022;42:1474–93.
Leske H, Blakstad H, Lund-Iversen M, Skovholt EK, Niehusmann P, Ramm-Pettersen JT, et al. Astrocytoma (CNS WHO grade 4), IDH-mutant with co-occurrence of BRAF p.V600E mutation, and homozygous loss of CDKN2A. Neuropathology. 2023. https://doi.org/10.1111/neup.12895.
Malta TM, de Souza CF, Sabedot TS, Silva TC, Mosella MS, Kalkanis SN, et al. Glioma CpG island methylator phenotype (G-CIMP): biological and clinical implications. Neuro Oncol. 2018;20:608–20.
Xu Y, Xiao H, Hu W, Shen HC, Liu W, Tan S, et al. CIMP-positive glioma is associated with better prognosis: a systematic analysis. Medicine. 2022;101:e30635.
Wen PY, Kesari S. Malignant gliomas in adults. N Engl J Med. 2008;359:492–507.
Riemenschneider MJ, Jeuken JW, Wesseling P, Reifenberger G. Molecular diagnostics of gliomas: state of the art. Acta Neuropathol. 2010;120:567–84.
Towner RA, Jensen RL, Colman H, Vaillant B, Smith N, Casteel R, et al. ELTD1, a potential new biomarker for gliomas. Neurosurgery 2013;72:77–90.
McNamara MG, Sahebjam S, Mason WP. Emerging biomarkers in glioblastoma. Cancers. 2013;5:1103–19.
Di Stefano AL, Picca A, Saragoussi E, Bielle F, Ducray F, Villa C, et al. Clinical, molecular, and radiomic profile of gliomas with FGFR3-TACC3 fusions. Neuro Oncol. 2020;22:1614–24.
Frattini V, Pagnotta SM, Tala, Fan JJ, Russo MV, Lee SB, et al. A metabolic function of FGFR3-TACC3 gene fusions in cancer. Nature 2018;553:222–7.
Jung CS, Foerch C, Schänzer A, Heck A, Plate KH, Seifert V, et al. Serum GFAP is a diagnostic marker for glioblastoma multiforme. Brain 2007;13012:3336–41.
Radu R, Petrescu GED, Gorgan RM, Brehar FM. GFAPδ: a promising biomarker and therapeutic target in glioblastoma. Front Oncol. 2022;12:859247.
Maiti S, Mondal S, Satyavarapu EM, Mandal C. mTORC2 regulates hedgehog pathway activity by promoting stability to Gli2 protein and its nuclear translocation. Cell Death Dis. 2017;8:e2926.
Huang D, Wang Y, Xu L, Chen L, Cheng M, Shi W, et al. GLI2 promotes cell proliferation and migration through transcriptional activation of ARHGEF16 in human glioma cells. J Exp Clin Cancer Res. 2018;37:247.
Castel D, Philippe C, Calmon R, Le Dret L, Truffaux N, Boddaert N, et al. Histone H3F3A and HIST1H3B K27M mutations define two subgroups of diffuse intrinsic pontine gliomas with different prognosis and phenotypes. Acta Neuropathol. 2015;130:815–27.
Karremann M, Gielen GH, Hoffmann M, Wiese M, Colditz N, Warmuth-Metz M, et al. Diffuse high-grade gliomas with H3 K27M mutations carry a dismal prognosis independent of tumor location. Neuro Oncol. 2018;20:123–31.
Domènech M, Hernández A, Plaja A, Martínez-Balibrea E, Balañà C. Hypoxia: the cornerstone of glioblastoma. Int J Mol Sci. 2021;22:12608.
Dang L, White DW, Gross S, Bennett BD, Bittinger MA, Driggers EM, et al. Cancer-associated IDH1 mutations produce 2-hydroxyglutarate. Nature. 2009;462:739–44.
Flavahan WA, Drier Y, Liau BB, Gillespie SM, Venteicher AS, Stemmer-Rachamimov AO, et al. Insulator dysfunction and oncogene activation in IDH mutant gliomas. Nature. 2016;529:110–4.
Bettegowda C, Agrawal N, Jiao Y, Sausen M, Wood LD, Hruban RH, et al. Mutations in CIC and FUBP1 contribute to human oligodendroglioma. Science. 2011;333:1453–5.
Borgenvik A, Čančer M, Hutter S, Swartling FJ. Targeting MYCN in molecularly defined malignant brain tumors. Front Oncol. 2020;10:626751.
Faria MH, Khayat AS, Burbano RR, Rabenhorst SH. c -MYC amplification and expression in astrocytic tumors. Acta Neuropathol. 2008;116:87–95.
Turner KM, Deshpande V, Beyter D, Koga T, Rusert J, Lee C, et al. Extrachromosomal oncogene amplification drives tumour evolution and genetic heterogeneity. Nature. 2017;543:122–5.
Karami Fath M, Karimfar N, Fazlollahpour Naghibi A, Shafa S, Ghasemi Shiran M, Ataei M, et al. Revisiting characteristics of oncogenic extrachromosomal DNA as mobile enhancers on neuroblastoma and glioma cancers. Cancer Cell Int. 2022;22:200.
Lobbous M, Bernstock JD, Coffee E, Friedman GK, Metrock LK, Chagoya G, et al. An update on neurofibromatosis type 1-associated gliomas. Cancers. 2020;12:114.
Esteller M, Garcia-Foncillas J, Andion E, Goodman SN, Hidalgo OF, Vanaclocha V, et al. Inactivation of the DNA-repair gene MGMT and the clinical response of gliomas to alkylating agents. N Engl J Med. 2000;343:1350–4.
Aldape K, Zadeh G, Mansouri S, Reifenberger G, von Deimling A. Glioblastoma: pathology, molecular mechanisms and markers. Acta Neuropathol. 2015;129:829–48.
Li SY, Johnson R, Smyth LC, Dragunow M. Platelet-derived growth factor signalling in neurovascular function and disease. Int J Biochem Cell Biol. 2022;145:106187.
Rascio F, Spadaccino F, Rocchetti MT, Castellano G, Stallone G, Netti GS, et al. The pathogenic role of PI3K/AKT pathway in cancer onset and drug resistance: an updated review. Cancers. 2021;13:3949.
Yan Y, Takayasu T, Hines G, Dono A, Hsu SH, Zhu JJ, et al. Landscape of genomic alterations in IDH wild-type glioblastoma identifies PI3K as a favorable prognostic factor. JCO Precis Oncol. 2020;4:575–84.
Yang JM, Schiapparelli P, Nguyen HN, Igarashi A, Zhang Q, Abbadi S, et al. Characterization of PTEN mutations in brain cancer reveals that pten mono-ubiquitination promotes protein stability and nuclear localization. Oncogene. 2017;36:3673–85.
Zhang Y, Dube C, Gibert M, Jr., Cruickshanks N, Wang B, Coughlan M, et al. The p53 pathway in glioblastoma. Cancers. 2018;10:297.
Mao H, Lebrun DG, Yang J, Zhu VF, Li M. Deregulated signaling pathways in glioblastoma multiforme: molecular mechanisms and therapeutic targets. Cancer Invest. 2012;30:48–56.
Adhikari AS, Sullivan T, Bargaje R, Lu L, O’Sullivan TN, Song Y, et al. Abrogation of Rb tumor suppression initiates GBM in differentiated astrocytes by driving a progenitor cell program. Front Oncol. 2022;12:904479.
Hung HC, Liu CC, Chuang JY, Su CL, Gean PW. Inhibition of sonic hedgehog signaling suppresses glioma stem-like cells likely through inducing autophagic cell death. Front Oncol. 2020;10:1233.
Yan GN, Yang L, Lv YF, Shi Y, Shen LL, Yao XH, et al. Endothelial cells promote stem-like phenotype of glioma cells through activating the hedgehog pathway. J Pathol. 2014;234:11–22.
Higa N, Akahane T, Yokoyama S, Yonezawa H, Uchida H, Takajo T, et al. A tailored next-generation sequencing panel identified distinct subtypes of wildtype IDH and TERT promoter glioblastomas. Cancer Sci. 2020;111:3902–11.
Liu EM, Shi ZF, Li KK, Malta TM, Chung NY, Chen H, et al. Molecular landscape of IDH-wild type, pTERT-wild type adult glioblastomas. Brain Pathol. 2022;32:e13107.
Stead LF, Verhaak RGW. Doomed from the TERT? A two-stage model of tumorigenesis in IDH wild-type glioblastoma. Cancer Cell. 2019;35:542–4.
Xu C, Wu X, Zhu J. VEGF promotes proliferation of human glioblastoma multiforme stem-like cells through VEGF receptor 2. ScientificWorldJournal 2013;2013:417413.
Chaudhry IH, O’Donovan DG, Brenchley PE, Reid H, Roberts IS. Vascular endothelial growth factor expression correlates with tumour grade and vascularity in gliomas. Histopathology. 2001;39:409–15.
Pulvirenti T, Van Der Heijden M, Droms LA, Huse JT, Tabar V, Hall A. Dishevelled 2 signaling promotes self-renewal and tumorigenicity in human gliomas. Cancer Res. 2011;71:7280–90.
Zuccarini M, Giuliani P, Ziberi S, Carluccio M, Iorio PD, Caciagli F, et al. The role of wnt signal in glioblastoma development and progression: a possible new pharmacological target for the therapy of this tumor. Genes. 2018;9:105.
Tanwar MK, Gilbert MR, Holland EC. Gene expression microarray analysis reveals YKL-40 to be a potential serum marker for malignant character in human glioma. Cancer Res. 2002;62:4364–8.
Chen WJ, Zhang X, Han H, Lv JN, Kang EM, Zhang YL, et al. The different role of YKL-40 in glioblastoma is a function of MGMT promoter methylation status. Cell Death Dis. 2020;11:668.
Acknowledgements
The authors thank The University of Texas MD Anderson Cancer Center, Houston, USA for providing the computational facilities and literature required to complete this review. The authors also gratefully acknowledge stimulating discussions with their colleagues Dr. Marcos Estecio and Dr. Chad Tang which helped in completing this review.
Funding
We would like to thank funding from MD Anderson Startup funds and a grant from the NCI 5R01 CA225963 to Krishna P Bhat for financial support.
Author information
Authors and Affiliations
Contributions
RT and KB established the review idea and finalized the framework. RT and KB were involved in the writing and editing of the review. All authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Trivedi, R., Bhat, K.P. Liquid biopsy: creating opportunities in brain space. Br J Cancer 129, 1727–1746 (2023). https://doi.org/10.1038/s41416-023-02446-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-023-02446-0
- Springer Nature Limited