Abstract
The tumors classified as gliomas include a wide variety of histologies including the more common (astrocytoma, glioblastoma), as well as the less common histologies (oligodendroglioma, mixed oligoastrocytoma, pilocytic astrocytoma). Recent efforts at comprehensive genetic characterization of various primary brain tumor types have identified a number of common alterations and pathways common to multiple tumor types. Common pathways in glioma biology include growth factor receptor tyrosine kinases and their downstream signaling via the MAP kinase cascade or PI3K signaling, loss of apoptosis through p53, cell cycle regulation, angiogenesis via VEGF signaling, and invasion. However, in addition to these common general pathway alterations, a number of specific alterations have been identified in particular tumor types, and a number of these have direct therapeutic implications. These include mutations or fusions in the BRAF gene seen in pilocytic astrocytomas (and gangliogliomas). In oligodendrogliomas, mutations in IDH1 and codeletion of chromosomes 1p and 19q are associated with improved survival with upfront use of combined chemotherapy and radiation, and these tumors also have unique mutations of CIC and FUBP1 genes. Low grade gliomas are increasingly seen to be divided into two groups based on IDH mutation status, with astrocytomas developing through IDH mutation followed by p53 mutation, while poor prognosis low grade gliomas and primary glioblastomas (GBMs) are characterized by EGFR amplification, loss of PTEN, and loss of cyclin-dependent kinase inhibitors. GBMs can be further characterized based on gene expression and gene methylation patterns into three or four distinct subgroups. Prognostic markers in diffuse gliomas include IDH mutation, 1p/19q codeletion, and MGMT methylation, and MGMT is also a predictive marker in elderly patients with glioblastoma treated with temozolomide monotherapy.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Astrocytoma
- Glioblastoma
- Oligodendroglioma
- Pilocytic astrocytoma
- IDH1
- EGFR
- BRAF
- 1p/19q codeletion
- MGMT methylation
1 Introduction
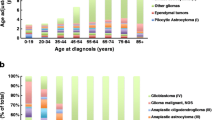
Gliomas are the most common primary brain tumor in adults, affecting about 20,000 people in the US each year [14]. Pathologically, gliomas are divided into astrocytomas, oligodendrogliomas, and mixed oligoastrocytomas based on histopathologic appearance. Most gliomas are astrocytic (82 %), followed by unspecified gliomas (7 %), oligodendroglial tumors (6 %), and mixed oligoastrocytomas (3 %) [14]. However, it is now understood that these histologies include a heterogeneous group of tumors with distinct molecular ontology and biology. Molecular classification has allowed for the identification of prognostic and predictive markers for some types of gliomas, and molecular subclasses are becoming increasingly important in the clinical classification and treatment of gliomas.
2 Histology
The 2007 World Health Organization classification recognizes three types and four grades of gliomas based on their microscopic appearance [45]. Grade 1, or pilocytic, astrocytomas are noninvasive tumors found primarily in children. Diffuse gliomas include astrocytomas, oligodendrogliomas. Grade 4 astrocytomas are called glioblastomas (GBMs). While some gliomas are mixed and have areas consistent with more than one type of histology, some investigators hypothesize that the different tumor types arise from different cells of origin and specific molecular alterations. Oligodendrogliomas probably arise from oligodendroglial precursor cells [44, 54, 72]. Radial glial cells have been proposed to be the cell of origin of ependymomas [74]. The cell of origin of astrocytomas has been proposed to be reactive astrocytes of the subventricular zone (for NF1 and PDGF-driven astrocytomas), neural progenitor cells of the frontal lobe (for IDH-mutant astrocytomas), and neural stem cells/progenitor cells in the subventricular zone [1, 22, 41].
3 Important Pathways in Glioma Biology
3.1 RTK/RAS/PI(3K), P53, Rb
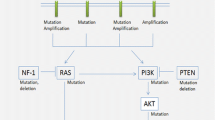
Publications from the Cancer Genome Atlas (TCGA) effort showed that more than three quarters of GBMs have alterations, including deletions, amplifications, or mutations, in three pathways: growth factor downstream signaling, particularly through the phosphatidyl inosital-3-kinase (PI3K) pathway, apoptosis regulation via p53 signaling, and cell cycle regulation via cyclin-dependant kinases and retinoblastoma 1 signaling [75]. Alterations in growth factor receptor/PI3K/MAPK signaling can include amplification of the epidermal growth factor receptor (EGFR/erbB1), platelet derived growth factor receptor (PDGFR), or MET genes or mutation in EGFR or the HER2/erbB2 receptor gene. Mutations in EGFR, particularly the EGFRvIII variant in which most of the extracellular domain is deleted, occur in 25–30 % of GBM [29, 76]. Downstream signaling from growth factor receptors can be activated by loss or mutation in the neurofibromatosis 1 gene (NF1), mutations in KRAS, mutations in PIK3CA (the gene for PI3K), and deletion or loss of heterozygosity of PTEN, the inhibitor of PI3K [75]. Although about one-third of GBM can have mutation or deletion of TP53, loss of p53 function can also be obtained through amplification of MDM2 or MDM4 [8]. Lastly, cell cycle regulation is disrupted through mutation or deletion of the cell cycle inhibitors CDKN2A, CDKN2B, and CDKN2C as well as less frequent amplifications in cyclins and cyclin-dependent kinases or actual loss of the RB1 gene.Translocations and fusion proteins may play an important role in activation of growth factor signaling. About 4 % of GBMs have fusions of EGFR with SEPT14, which activates STAT3 signaling, and another 2 % have fusions of EGFR with PSPH, which regulates neural stem cell proliferation [17, 49]. In addition, approximately 3 % of GBM have translocations causing a fusion between the fibroblast growth factor receptor genes, FGFR1 or FGFR3, and the transforming acidic coiled-coil containing protein genes, TACC1 orTACC3 [67]. There is in vitro and in vivo evidence that tumors with these fusions can be inhibited with appropriate tyrosine kinase inhibitors, but whether they are prognostic or predictive in people is not yet known.
3.2 IDH Mutation
In 2008, a multigroup collaboration using whole exome sequencing identified a common point mutation in the metabolic gene IDH1 in 12 % of glioblastoma samples [53]. Further studies found that this mutation is present in ~80 % of grade 2–3 gliomas and secondary GBM [2, 7, 26, 37, 62, 84, 88]. Mutations in IDH2 have also been identified in gliomas, although they are much less common and are mutually exclusive with mutations in IDH1 [26, 69, 88]. All mutations identified to date have been a single amino acid missense mutation in IDH1 at arginine 132 (R132) or the analogous residue in IDH2 (R172).
Although initial reports suggested that the mutant IDH functions in a dominate-negative fashion by heterodimerizing to wild-type IDH1 and impairing its activity [89, 92, 93], more recent in vitro studies have shown that the mutated IDH1 protein acquires the ability to convert α-ketoglutarate (α-KG) to R(-)-2-hydroxyglutarate (2-HG) [13, 32]. These findings led to the hypothesis that mutant IDH is an oncogene and 2-HG is an “oncometabolite” [18].
New evidence suggests that by antagonizing α-KG, 2-HG competitively inhibits the activity of many α-KG-dependent dioxygenases, including but not limited to histone demethylases (e.g., collagen prolyl-4-hydroxylase, prolyl hydroxylases, and the ten-eleven translocation (TET) family of DNA hydroxylases) [16, 63, 87]. Profiling of GBM from the TCGA demonstrated an association between IDH mutation and increased promoter methylation (G-CIMP) [50], which generally results in transcriptional silencing of the associated genes [35]. Two recent independent studies demonstrated that the G-CIMP phenotype was not correlated with IDH mutation but that IDH mutation alone is actually the cause of the G-CIMP hypermethylation phenotype in diffuse gliomas [11, 40].
3.3 Hypoxia, Pseudohypoxia, and Angiogenesis
Gliomas often exist in hypoxic or pseudohypoxic conditions. The extreme example of hypoxia is pseudopallisading necrosis in GBM, when the tumor outgrows its own blood supply. Hypoxia causes the upregulation of hypoxia inducible factor (HIF1a), and HIF1a can induce new vessel formation and stem cell survival processes. HIF1a is overexpressed in gliomas, particularly high grade gliomas [43]. HIF1a degradation via the ubiquitin-proteosome system is also inhibited by alterations in prolyl hydroxylases driven by 2-hydroxyglutarate produced by mutant IDH. Thus, IDH mutation can result in a situation in which HIF1a protein levels are high (consistent with hypoxia) while oxygen tension is normal. This situation has been called “pseudohypoxia” and may be a major driver of biology in these tumors along with other consequences of altered proline hydroxylation.
Blood vessel formation in gliomas often takes the form of angiogenesis, driven by vascular endothelial growth factor (VEGF). VEGF is overexpressed in brain tumors, with increasing expression corresponding to increased grade [10, 56, 57]. Angiogenesis can also be supported by CXCR4 signaling [90]. Alternatively, gliomas can coopt normal vessels, a process often mediated by angiopoietin signaling, or new vessels can be formed from bone marrow derived endothelial cells, in a process termed vasculogenesis [46, 60]. Glioma cells may even be able to transform into malignant endothelial cells [66]. It has been proposed that blockade of angiogenesis, particularly inhibition of VEGF signaling, may lead to increased glioma invasiveness, although this hypothesis remains controversial [38].
Invasion into normal brain is one of the hallmarks of diffuse gliomas and is one of the features making them incurable by surgery alone. Gliomas migrate along the secondary structures of Scherer, such as white matter tracks, neuronal tracks, vasculature, or subpial spaces [64]. Glioma migrations is facilitated by secretion of a variety of proteases, including matrix metalloproteases (MMPs), membrane type matrix metalloproteases (MT-MMPs), and adamalysins (ADAMS) [48]. Expression and secretion of these proteases can be regulated by TGF-beta and NF-κB.
4 Oligodendrogliomas
Oligodendrogliomas can be grade 2 (low grade) or grade 3 (anaplastic) [45]. There are no grade 4 oligodendrogliomas. One of the earliest steps in the development of oligodendrogliomas is mutation in isocitrate dehydrogenase 1 (IDH1) or 2 (IDH2) [84]. Mutation in IDH1 or IDH2 is followed by an unbalanced translocation resulting in loss of the p-arm of chromosome 1 and the q-arm of chromosome 19 [39, 59]. This translocation inactivates one copy of the Capicua transcriptional repressor (CIC) gene and the FUSE binding protein 1(FUBP1) gene [5]. Oligodendrogliomas then can develop a mutation in the other copy of these genes. Such mutations occur in 52 % of grade 2 oligodendrogliomas and 84 % of anaplastic (grade 3) oligodendrogliomas [31]. Whether these alterations are sufficient for oligodendroglioma development has not been established. The events involved in the transformation of grade 2–3 oligodendrogliomas have also not been established.
Codeletion of 1p and 19q is also a prognostic marker among all gliomas and among oligodendrogliomas, regardless of grade [91]. Indeed, mixed oligoastrocytomas with 1p/19q deletion have prognosis similar to oligodendrogliomas, while those without 1p/19q deletion have worse prognosis, similar to astrocytomas [30]. Moreover, 1p/19q codeletion is also a predictive marker among anaplastic oligodendrogliomas. Two randomized trials showed improved survival with the addition of chemotherapy to radiation for people with newly diagnosed anaplastic oligodendrogliomas whose tumors had 1p/19q codeletion but not for people whose tumors did not have this codeletion [9, 77]. Therefore, codeletion of 1p/19q, which can be detected by fluorescent in situ hybridization, is a clinically useful biomarker for oligodendrogliomas.
5 Astrocytoma Histopathologic and Molecular Classification
Grade 1, or pilocytic, astrocytomas are non-infiltrating neoplasms that occur mostly in children and adolescents but can occur in adults as well [45]. Pilocytic astrocytomas lack many of the mutations found in the diffuse gliomas. However, nearly all pilocytic astrocytomas have alterations in the BRAF oncogene leading to its activation, primarily by fusion with the KIAA1549 gene [33]. Other mechanisms of RAF pathway activation include fusion of SRGAP3 and RAF1, BRAFV600E mutation, in-frame insertion in the BRAF gene, or fusion of BRAF with FAM131B [12, 33, 34]. Activation of BRAF in neural progenitor cells is sufficient to cause pilocytic astrocytomas in mice and to transform human neural stem cells [21, 58]. How to target these alterations remains an area of study.
Diffuse gliomas can be categorized according to grade: low grade (grade II), anaplastic (grade III), and glioblastoma (GBM, grade IV). Traditionally, GBMs have been classified as primary or secondary on the basis of clinical presentation [65]. Secondary GBMs display evidence of progression from a lower grade tumor, whereas primary GBMs present as grade 4 at diagnosis. Secondary GBMs are predominantly found in younger patients (median age of ~45 years compared with median age of ~60 years for primary GBM) and tend to occur less frequently than primary GBMs, making up ~5 % of total GBMs [51]. Secondary GBMs are more likely to have mutation in IDH1 or IDH2. Despite the differences in their ontology, these high grade tumors are histopathologically indistinguishable [51].
In the 1990s, this clinical classification began to be correlated to molecular biology with the observation that amplification of the epidermal growth factor receptor (EGFR) gene and mutation or loss of heterozygosity of the TP53 gene was mutually exclusive, with the former being seen in primary glioblastoma and the latter being seen in secondary glioblastoma and lower grade gliomas [80, 83]. More recently, large-scale efforts have been made to identify the major genetic and epigenetic alterations and to define important molecular subtypes in GBM and lower grade gliomas [55, 79].
Gene expression patterns have been used by multiple groups to classify adult high grade gliomas into 3–4 groups [55, 71, 79]. The Cancer Genome Atlas (TCGA) identified four subgroups of GBM based on gene expression patterns, which were called Proneural, Neural, Mesenchymal, and Classical [79]. Individual gene expression subtypes were associated with specific genetic and epigenetic alterations. For example, proneural GBMs are enriched for the G-CIMP phenotype, IDH mutations, PDGFRA amplifications, and CDK4 amplifications. Classical GBMs are enriched for EGFR mutations, particularly the EGFRvIII variant. Mesenchymal GBMs are more likely to have MET amplification and NF1 mutation or loss and display increased angiogenesis, hypoxia, inflammatory infiltrates and inflammatory signaling pathways, including NF-κB, STAT3, TGF-β [3, 6, 55, 79] Secondary GBMs are virtually always Proneural, while primary GBM can be any of the subtypes. Most primary GBMs have EGFR amplification and deletion of the PTEN gene [8].
6 Prognostic and Predictive Markers in Astrocytomas
The two strongest prognostic factors in astrocytomas are IDH mutation status and grade [25, 88]. Indeed, prognosis for IDH-mutant GBMs is better in some series than that for IDH-wild-type grade 3 astrocytomas. The prognostic importance of IDH mutation is independent of other known prognostic factors, including age, and MGMT methylation status [62]. However, it remains to be determined whether IDH mutation is a prognostic factor only or whether it is predictive of outcome to specific treatments or mechanistically related to treatment response. Small retrospective series have suggested that the response rate to alkylating chemotherapy is also higher in IDH-mutated grade 2 tumors than in wild-type tumors and that progression-free survival after radiation or alkylating chemotherapy is higher for people with IDH-mutated tumors than for people with wild-type tumors [28, 36, 68]. In the German Glioma Group retrospective study, IDH mutation influenced survival only in those patients who received radiation or chemotherapy immediately after surgery [24]. IDH mutation does not predict progression-free survival for temozolomide treatment in low grade astrocytomas that had previously received radiation [73]. IDH mutation may also predict the benefit of complete resection in high grade gliomas [4]. However, their retrospective nature and lack of control groups limits the conclusions that can be drawn from these studies.
MGMT, the gene encoding the DNA repair enzyme O6-methylguanine-DNA methyltransferase, is methylated in 30–40 % of GBMs and 80 % of IDH-mutated low grade gliomas [19, 27, 42]. The presence of methylated MGMT is prognostic in all grades of glioma, including both oligodendroglial and astrocytic histologies [42, 61, 70, 78]. Given the strong association between G-CIMP phenotype and MGMT methylation, the prognostic significance of each of these independently cannot be assessed. It is also not clear whether the observation that MGMT methylation is more significant than expression level measured by RNA or protein is due to biology or to technical artifacts of the assays used.
MGMT was originally studied because of its role in repairing damage from alkylating agents such as the nitrosoureas or temozolomide. However, it remains controversial whether MGMT methylation is predictive of benefit from temozolomide. In the long-term follow-up of EORTC 26981-22981/NCIC CE3, MGMT methylation was prognostic in both the radiation alone and radiation plus temozolomide groups. Moreover, there was a statistically significant improvement in survival with the addition of temozolomide in both the MGMT methylated and unmethylated groups with similar hazard ratio, although the absolute benefit was much smaller in the unmethylated group [70]. In the elderly population, two trials (NOA-08 and the Nordic trial) have suggested that people with GBM with MGMT methylation have improved survival with temozolomide monotherapy compared to radiation, while the opposite is true for elderly people with GBM with unmethylated MGMT [47, 85]. The role of MGMT or other potential biomarkers other than IDH in grade 2 or 3 astrocytomas remains uncertain.
7 Ependymomas
Ependymomas have been subtyped based on both location (supratentorial, infratentorial, or spinal cord) or by gene expression pattern. Common genetic alterations in ependymomas include NF2 mutation or loss, HER2/erbB2 and erbB4 amplification, and RASSF1A and HIC1 methylation [15, 20, 23, 81]. Two groups have used gene expression to identify two types of ependymoma, one characterized by growth factor receptor signaling and the other by large chromosomal rearrangements along with altered metabolism and cytoskeleton pathways [82, 86]. Most recently, a fusion gene involving c11orf95 and relA, the active component of NF-κB, was found to be a driver alteration in supratentorial ependymoma [52]. The high frequency observed of this fusion in the supratentorial ependymomas suggest that this is both a key driver of biology in this tumor subtype and may also be a potential target for specific drug development. Prognostic markers for ependymoma are not well established.
8 Conclusion
Gliomas include multiple distinct histopathologic types of tumors under the current WHO classification. Recent findings regarding some of the key driving alterations in some tumor types elucidate both the distinct biologies that define different histopathologies, and identify distinct molecular subtypes within single histopathologies. There appear to be at least three key molecular ontology pathways to the development of diffuse glioma. One pathway starts with IDH mutation followed by TP53 mutation and results in astrocytic lineage tumors. These gliomas start as grade 2 astrocytomas, have the G-CIMP phenotype, and presumably then acquire other genetic alterations that result in progression to higher grade tumors. Another pathway starts with IDH mutation followed by loss of 1p/19q, which is associated with mutation in the CIC gene or the FUBP1 gene. These alterations result in development of grade 2 oligodendrogliomas, which can then acquire other genetic alterations to become anaplastic oligodendrogliomas. The third pathway includes those gliomas that are wild-type for IDH. These gliomas appear to rapidly acquire multiple complex genetic alterations, including amplification or mutation of EGFR, and loss of the PTEN gene, and become GBMs very early in their development. IDH, 1p/19q codeletion, and MGMT are validated prognostic and probably predictive markers as well.
References
Alcantara Llaguno S, Chen J, Kwon CH, Jackson EL, Li Y, Burns DK, Alvarez-Buylla A, Parada LF (2009) Malignant astrocytomas originate from neural stem/progenitor cells in a somatic tumor suppressor mouse model. Cancer Cell 15(1):45–56. doi:10.1016/j.ccr.2008.12.006
Balss J, Meyer J, Mueller W, Korshunov A, Hartmann C, von Deimling A (2008) Analysis of the IDH1 codon 132 mutation in brain tumors. Acta Neuropathol 116(6):597–602. doi:10.1007/s00401-008-0455-2
Beier CP, Kumar P, Meyer K, Leukel P, Bruttel V, Aschenbrenner I, Riemenschneider MJ, Fragoulis A, Rummele P, Lamszus K, Schulz JB, Weis J, Bogdahn U, Wischhusen J, Hau P, Spang R, Beier D (2012) The cancer stem cell subtype determines immune infiltration of glioblastoma. Stem Cells Dev 21(15):2753–2761. doi:10.1089/scd.2011.0660
Beiko J, Suki D, Hess KR, Fox BD, Cheung V, Cabral M, Shonka N, Gilbert MR, Sawaya R, Prabhu SS, Weinberg J, Lang FF, Aldape KD, Sulman EP, Rao G, McCutcheon IE, Cahill DP (2014) IDH1 mutant malignant astrocytomas are more amenable to surgical resection and have a survival benefit associated with maximal surgical resection. Neuro-Oncology 16(1):81–91. doi:10.1093/neuonc/not159
Bettegowda C, Agrawal N, Jiao Y, Sausen M, Wood LD, Hruban RH, Rodriguez FJ, Cahill DP, McLendon R, Riggins G, Velculescu VE, Oba-Shinjo SM, Marie SK, Vogelstein B, Bigner D, Yan H, Papadopoulos N, Kinzler KW (2011) Mutations in CIC and FUBP1 contribute to human oligodendroglioma. Science 333(6048):1453–1455. doi:10.1126/science.1210557
Bhat KP, Balasubramaniyan V, Vaillant B, Ezhilarasan R, Hummelink K, Hollingsworth F, Wani K, Heathcock L, James JD, Goodman LD, Conroy S, Long L, Lelic N, Wang S, Gumin J, Raj D, Kodama Y, Raghunathan A, Olar A, Joshi K, Pelloski CE, Heimberger A, Kim SH, Cahill DP, Rao G, Den Dunnen WF, Boddeke HW, Phillips HS, Nakano I, Lang FF, Colman H, Sulman EP, Aldape K (2013) Mesenchymal differentiation mediated by NF-kappaB promotes radiation resistance in glioblastoma. Cancer Cell 24(3):331–346. doi:10.1016/j.ccr.2013.08.001
Bleeker FE, Lamba S, Leenstra S, Troost D, Hulsebos T, Vandertop WP, Frattini M, Molinari F, Knowles M, Cerrato A, Rodolfo M, Scarpa A, Felicioni L, Buttitta F, Malatesta S, Marchetti A, Bardelli A (2009) IDH1 mutations at residue p.R132 (IDH1(R132)) occur frequently in high-grade gliomas but not in other solid tumors. Hum Mutat 30(1):7–11. doi:10.1002/humu.20937
Brennan CW, Verhaak RG, McKenna A, Campos B, Noushmehr H, Salama SR, Zheng S, Chakravarty D, Sanborn JZ, Berman SH, Beroukhim R, Bernard B, Wu CJ, Genovese G, Shmulevich I, Barnholtz-Sloan J, Zou L, Vegesna R, Shukla SA, Ciriello G, Yung WK, Zhang W, Sougnez C, Mikkelsen T, Aldape K, Bigner DD, Van Meir EG, Prados M, Sloan A, Black KL, Eschbacher J, Finocchiaro G, Friedman W, Andrews DW, Guha A, Iacocca M, O’Neill BP, Foltz G, Myers J, Weisenberger DJ, Penny R, Kucherlapati R, Perou CM, Hayes DN, Gibbs R, Marra M, Mills GB, Lander E, Spellman P, Wilson R, Sander C, Weinstein J, Meyerson M, Gabriel S, Laird PW, Haussler D, Getz G, Chin L (2013) The somatic genomic landscape of glioblastoma. Cell 155(2):462–477. doi:10.1016/j.cell.2013.09.034
Cairncross G, Wang M, Shaw E, Jenkins R, Brachman D, Buckner J, Fink K, Souhami L, Laperriere N, Curran W, Mehta M (2013) Phase III trial of chemoradiotherapy for anaplastic oligodendroglioma: long-term results of RTOG 9402. J Clin Oncol: Official J Am Soc Clin Oncol 31(3):337–343. doi:10.1200/JCO.2012.43.2674
Chaudhry IH, O’Donovan DG, Brenchley PE, Reid H, Roberts IS (2001) Vascular endothelial growth factor expression correlates with tumour grade and vascularity in gliomas. Histopathol 39(4):409–415
Christensen BC, Smith AA, Zheng S, Koestler DC, Houseman EA, Marsit CJ, Wiemels JL, Nelson HH, Karagas MR, Wrensch MR, Kelsey KT, Wiencke JK (2011) DNA methylation, isocitrate dehydrogenase mutation, and survival in glioma. J Natl Cancer Inst 103(2):143–153. doi:10.1093/jnci/djq497
Cin H, Meyer C, Herr R, Janzarik WG, Lambert S, Jones DT, Jacob K, Benner A, Witt H, Remke M, Bender S, Falkenstein F, Van Anh TN, Olbrich H, von Deimling A, Pekrun A, Kulozik AE, Gnekow A, Scheurlen W, Witt O, Omran H, Jabado N, Collins VP, Brummer T, Marschalek R, Lichter P, Korshunov A, Pfister SM (2011) Oncogenic FAM131B-BRAF fusion resulting from 7q34 deletion comprises an alternative mechanism of MAPK pathway activation in pilocytic astrocytoma. Acta Neuropathol 121(6):763–774. doi:10.1007/s00401-011-0817-z
Dang L, White DW, Gross S, Bennett BD, Bittinger MA, Driggers EM, Fantin VR, Jang HG, Jin S, Keenan MC, Marks KM, Prins RM, Ward PS, Yen KE, Liau LM, Rabinowitz JD, Cantley LC, Thompson CB, Vander Heiden MG, Su SM (2009) Cancer-associated IDH1 mutations produce 2-hydroxyglutarate. Nature 462(7274):739–744. doi:10.1038/nature08617
Dolecek T, Propp J, Stroup N, Kruchko C (2012) CBTRUS statistical report: primary brain and central nervous system tumors diagnosed in the United States in 2005–2009. Neuro-Oncology 15(suppl 5):1–49
Ebert C, von Haken M, Meyer-Puttlitz B, Wiestler OD, Reifenberger G, Pietsch T, von Deimling A (1999) Molecular genetic analysis of ependymal tumors. NF2 mutations and chromosome 22q loss occur preferentially in intramedullary spinal ependymomas. Am J Pathol 155(2):627–632. doi:10.1016/S0002-9440(10)65158-9
Figueroa ME, Abdel-Wahab O, Lu C, Ward PS, Patel J, Shih A, Li Y, Bhagwat N, Vasanthakumar A, Fernandez HF, Tallman MS, Sun Z, Wolniak K, Peeters JK, Liu W, Choe SE, Fantin VR, Paietta E, Lowenberg B, Licht JD, Godley LA, Delwel R, Valk PJ, Thompson CB, Levine RL, Melnick A (2010) Leukemic IDH1 and IDH2 mutations result in a hypermethylation phenotype, disrupt TET2 function, and impair hematopoietic differentiation. Cancer Cell 18(6):553–567. doi:10.1016/j.ccr.2010.11.015
Frattini V, Trifonov V, Chan JM, Castano A, Lia M, Abate F, Keir ST, Ji AX, Zoppoli P, Niola F, Danussi C, Dolgalev I, Porrati P, Pellegatta S, Heguy A, Gupta G, Pisapia DJ, Canoll P, Bruce JN, McLendon RE, Yan H, Aldape K, Finocchiaro G, Mikkelsen T, Prive GG, Bigner DD, Lasorella A, Rabadan R, Iavarone A (2013) The integrated landscape of driver genomic alterations in glioblastoma. Nat Genet 45(10):1141–1149. doi:10.1038/ng.2734
Garber K (2010) Oncometabolite? IDH1 discoveries raise possibility of new metabolism targets in brain cancers and leukemia. J Natl Cancer Inst 102(13):926–928. doi:10.1093/jnci/djq262
Gilbert MR, Wang M, Aldape KD, Stupp R, Hegi ME, Jaeckle KA, Armstrong TS, Wefel JS, Won M, Blumenthal DT, Mahajan A, Schultz CJ, Erridge S, Baumert B, Hopkins KI, Tzuk-Shina T, Brown PD, Chakravarti A, Curran WJ Jr, Mehta MP (2013) Dose-dense temozolomide for newly diagnosed glioblastoma: a randomized phase III clinical trial. J Clin Oncol: Official J American Society Clin Oncol 31(32):4085–4091. doi:10.1200/JCO.2013.49.6968
Gilbertson RJ, Bentley L, Hernan R, Junttila TT, Frank AJ, Haapasalo H, Connelly M, Wetmore C, Curran T, Elenius K, Ellison DW (2002) ERBB receptor signaling promotes ependymoma cell proliferation and represents a potential novel therapeutic target for this disease. Clin Cancer Res: An Official J American Assoc Cancer Res 8(10):3054–3064
Gronych J, Korshunov A, Bageritz J, Milde T, Jugold M, Hambardzumyan D, Remke M, Hartmann C, Witt H, Jones DT, Witt O, Heiland S, Bendszus M, Holland EC, Pfister S, Lichter P (2011) An activated mutant BRAF kinase domain is sufficient to induce pilocytic astrocytoma in mice. J Clin Investig 121(4):1344–1348. doi:10.1172/JCI44656
Hambardzumyan D, Cheng YK, Haeno H, Holland EC, Michor F (2011) The probable cell of origin of NF1- and PDGF-driven glioblastomas. PLoS ONE 6(9):24454. doi:10.1371/journal.pone.0024454
Hamilton DW, Lusher ME, Lindsey JC, Ellison DW, Clifford SC (2005) Epigenetic inactivation of the RASSF1A tumour suppressor gene in ependymoma. Cancer Lett 227(1):75–81. doi:10.1016/j.canlet.2004.11.044
Hartmann C, Hentschel B, Tatagiba M, Schramm J, Schnell O, Seidel C, Stein R, Reifenberger G, Pietsch T, von Deimling A, Loeffler M, Weller M (2011) Molecular markers in low-grade gliomas: predictive or prognostic? Clin Cancer Res: An Official J American Assoc Cancer Res 17(13):4588–4599. doi:10.1158/1078-0432.CCR-10-3194
Hartmann C, Hentschel B, Wick W, Capper D, Felsberg J, Simon M, Westphal M, Schackert G, Meyermann R, Pietsch T, Reifenberger G, Weller M, Loeffler M, von Deimling A (2010) Patients with IDH1 wild type anaplastic astrocytomas exhibit worse prognosis than IDH1-mutated glioblastomas, and IDH1 mutation status accounts for the unfavorable prognostic effect of higher age: implications for classification of gliomas. Acta Neuropathol 120(6):707–718. doi:10.1007/s00401-010-0781-z
Hartmann C, Meyer J, Balss J, Capper D, Mueller W, Christians A, Felsberg J, Wolter M, Mawrin C, Wick W, Weller M, Herold-Mende C, Unterberg A, Jeuken JW, Wesseling P, Reifenberger G, von Deimling A (2009) Type and frequency of IDH1 and IDH2 mutations are related to astrocytic and oligodendroglial differentiation and age: a study of 1,010 diffuse gliomas. Acta Neuropathol 118(4):469–474. doi:10.1007/s00401-009-0561-9
Hegi ME, Diserens AC, Gorlia T, Hamou MF, de Tribolet N, Weller M, Kros JM, Hainfellner JA, Mason W, Mariani L, Bromberg JE, Hau P, Mirimanoff RO, Cairncross JG, Janzer RC, Stupp R (2005) MGMT gene silencing and benefit from temozolomide in glioblastoma. The New England J Med 352(10):997–1003. doi:10.1056/NEJMoa043331
Houillier C, Wang X, Kaloshi G, Mokhtari K, Guillevin R, Laffaire J, Paris S, Boisselier B, Idbaih A, Laigle-Donadey F, Hoang-Xuan K, Sanson M, Delattre JY (2010) IDH1 or IDH2 mutations predict longer survival and response to temozolomide in low-grade gliomas. Neurology 75(17):1560–1566. doi:10.1212/WNL.0b013e3181f96282
Humphrey PA, Wong AJ, Vogelstein B, Zalutsky MR, Fuller GN, Archer GE, Friedman HS, Kwatra MM, Bigner SH, Bigner DD (1990) Anti-synthetic peptide antibody reacting at the fusion junction of deletion-mutant epidermal growth factor receptors in human glioblastoma. Proc Natl Acad Sci USA 87(11):4207–4211
Jiang H, Ren X, Cui X, Wang J, Jia W, Zhou Z, Lin S (2013) 1p/19q codeletion and IDH1/2 mutation identified a subtype of anaplastic oligoastrocytomas with prognosis as favorable as anaplastic oligodendrogliomas. Neuro-Oncology 15(6):775–782. doi:10.1093/neuonc/not027
Jiao Y, Killela PJ, Reitman ZJ, Rasheed AB, Heaphy CM, de Wilde RF, Rodriguez FJ, Rosemberg S, Oba-Shinjo SM, Nagahashi Marie SK, Bettegowda C, Agrawal N, Lipp E, Pirozzi C, Lopez G, He Y, Friedman H, Friedman AH, Riggins GJ, Holdhoff M, Burger P, McLendon R, Bigner DD, Vogelstein B, Meeker AK, Kinzler KW, Papadopoulos N, Diaz LA, Yan H (2012) Frequent ATRX, CIC, FUBP1 and IDH1 mutations refine the classification of malignant gliomas. Oncotarget 3(7):709–722
Jin G, Reitman ZJ, Spasojevic I, Batinic-Haberle I, Yang J, Schmidt-Kittler O, Bigner DD, Yan H (2011) 2-hydroxyglutarate production, but not dominant negative function, is conferred by glioma-derived NADP-dependent isocitrate dehydrogenase mutations. PLoS ONE 6(2):e16812. doi:10.1371/journal.pone.0016812
Jones DT, Kocialkowski S, Liu L, Pearson DM, Backlund LM, Ichimura K, Collins VP (2008) Tandem duplication producing a novel oncogenic BRAF fusion gene defines the majority of pilocytic astrocytomas. Cancer Res 68(21):8673–8677. doi:10.1158/0008-5472.CAN-08-2097
Jones DT, Kocialkowski S, Liu L, Pearson DM, Ichimura K, Collins VP (2009) Oncogenic RAF1 rearrangement and a novel BRAF mutation as alternatives to KIAA1549:BRAF fusion in activating the MAPK pathway in pilocytic astrocytoma. Oncogene 28(20):2119–2123. doi:10.1038/onc.2009.73
Jones PA, Baylin SB (2007) The epigenomics of cancer. Cell 128(4):683–692. doi:10.1016/j.cell.2007.01.029
Juratli TA, Kirsch M, Robel K, Soucek S, Geiger K, von Kummer R, Schackert G, Krex D (2012) IDH mutations as an early and consistent marker in low-grade astrocytomas WHO grade II and their consecutive secondary high-grade gliomas. J Neuro Oncol. doi:10.1007/s11060-012-0844-1
Kang MR, Kim MS, Oh JE, Kim YR, Song SY, Seo SI, Lee JY, Yoo NJ, Lee SH (2009) Mutational analysis of IDH1 codon 132 in glioblastomas and other common cancers. Int J cancer 125(2):353–355. doi:10.1002/ijc.24379
Keunen O, Johansson M, Oudin A, Sanzey M, Rahim SA, Fack F, Thorsen F, Taxt T, Bartos M, Jirik R, Miletic H, Wang J, Stieber D, Stuhr L, Moen I, Rygh CB, Bjerkvig R, Niclou SP (2011) Anti-VEGF treatment reduces blood supply and increases tumor cell invasion in glioblastoma. Proc Natl Acad Sci USA 108(9):3749–3754. doi:10.1073/pnas.1014480108
Kraus JA, Koopmann J, Kaskel P, Maintz D, Brandner S, Schramm J, Louis DN, Wiestler OD, von Deimling A (1995) Shared allelic losses on chromosomes 1p and 19q suggest a common origin of oligodendroglioma and oligoastrocytoma. J Neuropathol Exp Neurol 54(1):91–95
Laffaire J, Everhard S, Idbaih A, Criniere E, Marie Y, de Reynies A, Schiappa R, Mokhtari K, Hoang-Xuan K, Sanson M, Delattre JY, Thillet J, Ducray F (2011) Methylation profiling identifies 2 groups of gliomas according to their tumorigenesis. Neuro-Oncology 13(1):84–98. doi:10.1093/neuonc/noq110
Lai A, Kharbanda S, Pope WB, Tran A, Solis OE, Peale F, Forrest WF, Pujara K, Carrillo JA, Pandita A, Ellingson BM, Bowers CW, Soriano RH, Schmidt NO, Mohan S, Yong WH, Seshagiri S, Modrusan Z, Jiang Z, Aldape KD, Mischel PS, Liau LM, Escovedo CJ, Chen W, Nghiemphu PL, James CD, Prados MD, Westphal M, Lamszus K, Cloughesy T, Phillips HS (2011) Evidence for Sequenced Molecular Evolution of IDH1 Mutant Glioblastoma From a Distinct Cell of Origin. J Clin Oncol: Official J Am Soc Clin Oncol 29(34):4482–4490. doi:10.1200/JCO.2010.33.8715
Leu S, von Felten S, Frank S, Vassella E, Vajtai I, Taylor E, Schulz M, Hutter G, Hench J, Schucht P, Boulay JL, Mariani L (2013) IDH/MGMT-driven molecular classification of low-grade glioma is a strong predictor for long-term survival. Neuro-oncology 15(4):469–479. doi:10.1093/neuonc/nos317
Li Z, Bao S, Wu Q, Wang H, Eyler C, Sathornsumetee S, Shi Q, Cao Y, Lathia J, McLendon RE, Hjelmeland AB, Rich JN (2009) Hypoxia-inducible factors regulate tumorigenic capacity of glioma stem cells. Cancer Cell 15(6):501–513. doi:10.1016/j.ccr.2009.03.018
Liu C, Sage JC, Miller MR, Verhaak RG, Hippenmeyer S, Vogel H, Foreman O, Bronson RT, Nishiyama A, Luo L, Zong H (2011) Mosaic analysis with double markers reveals tumor cell of origin in glioma. Cell 146(2):209–221. doi:10.1016/j.cell.2011.06.014
Louis DN, Ohgaki H, Wiestler OD, Cavenee WK, Burger PC, Jouvet A, Scheithauer BW, Kleihues P (2007) The 2007 WHO classification of tumours of the central nervous system. Acta Neuropathol 114(2):97–109. doi:10.1007/s00401-007-0243-4
Lyden D, Hattori K, Dias S, Costa C, Blaikie P, Butros L, Chadburn A, Heissig B, Marks W, Witte L, Wu Y, Hicklin D, Zhu Z, Hackett NR, Crystal RG, Moore MA, Hajjar KA, Manova K, Benezra R, Rafii S (2001) Impaired recruitment of bone-marrow-derived endothelial and hematopoietic precursor cells blocks tumor angiogenesis and growth. Nat Med 7(11):1194–1201. doi:10.1038/nm1101-1194
Malmstrom A, Gronberg BH, Marosi C, Stupp R, Frappaz D, Schultz H, Abacioglu U, Tavelin B, Lhermitte B, Hegi ME, Rosell J, Henriksson R (2012) Temozolomide versus standard 6-week radiotherapy versus hypofractionated radiotherapy in patients older than 60 years with glioblastoma: the Nordic randomised, phase 3 trial. Lancet Oncol 13(9):916–926. doi:10.1016/S1470-2045(12)70265-6
Mentlein R, Hattermann K, Held-Feindt J (2012) Lost in disruption: role of proteases in glioma invasion and progression. Biochim Biophys Acta 1825(2):178–185. doi:10.1016/j.bbcan.2011.12.001
Nakano I, Dougherty JD, Kim K, Klement I, Geschwind DH, Kornblum HI (2007) Phosphoserine phosphatase is expressed in the neural stem cell niche and regulates neural stem and progenitor cell proliferation. Stem Cells 25(8):1975–1984. doi:10.1634/stemcells.2007-0046
Noushmehr H, Weisenberger DJ, Diefes K, Phillips HS, Pujara K, Berman BP, Pan F, Pelloski CE, Sulman EP, Bhat KP, Verhaak RG, Hoadley KA, Hayes DN, Perou CM, Schmidt HK, Ding L, Wilson RK, Van Den Berg D, Shen H, Bengtsson H, Neuvial P, Cope LM, Buckley J, Herman JG, Baylin SB, Laird PW, Aldape K (2010) Identification of a CpG island methylator phenotype that defines a distinct subgroup of glioma. Cancer Cell 17(5):510–522. doi:10.1016/j.ccr.2010.03.017
Ohgaki H, Dessen P, Jourde B, Horstmann S, Nishikawa T, Di Patre PL, Burkhard C, Schuler D, Probst-Hensch NM, Maiorka PC, Baeza N, Pisani P, Yonekawa Y, Yasargil MG, Lutolf UM, Kleihues P (2004) Genetic pathways to glioblastoma: a population-based study. Cancer Res 64(19):6892–6899. doi:10.1158/0008-5472.CAN-04-1337
Parker M, Mohankumar KM, Punchihewa C, Weinlich R, Dalton JD, Li Y, Lee R, Tatevossian RG, Phoenix TN, Thiruvenkatam R, White E, Tang B, Orisme W, Gupta K, Rusch M, Chen X, Nagahawhatte P, Hedlund E, Finkelstein D, Wu G, Shurtleff S, Easton J, Boggs K, Yergeau D, Vadodaria B, Mulder HL, Becksford J, Gupta P, Huether R, Ma J, Song G, Gajjar A, Merchant T, Boop F, Smith AA, Ding L, Lu C, Ochoa K, Zhao D, Fulton RS, Fulton LL, Mardis ER, Wilson RK, Downing JR, Green DR, Zhang J, Ellison DW, Gilbertson RJ (2014) C11orf95-RELA fusions drive oncogenic NF-kappaB signalling in ependymoma. Nature 506(7489):451–455. doi:10.1038/nature13109
Parsons DW, Jones S, Zhang X, Lin JC, Leary RJ, Angenendt P, Mankoo P, Carter H, Siu IM, Gallia GL, Olivi A, McLendon R, Rasheed BA, Keir S, Nikolskaya T, Nikolsky Y, Busam DA, Tekleab H, Diaz LA Jr, Hartigan J, Smith DR, Strausberg RL, Marie SK, Shinjo SM, Yan H, Riggins GJ, Bigner DD, Karchin R, Papadopoulos N, Parmigiani G, Vogelstein B, Velculescu VE, Kinzler KW (2008) An integrated genomic analysis of human glioblastoma multiforme. Science 321(5897):1807–1812. doi:10.1126/science.1164382
Persson AI, Petritsch C, Swartling FJ, Itsara M, Sim FJ, Auvergne R, Goldenberg DD, Vandenberg SR, Nguyen KN, Yakovenko S, Ayers-Ringler J, Nishiyama A, Stallcup WB, Berger MS, Bergers G, McKnight TR, Goldman SA, Weiss WA (2010) Non-stem cell origin for oligodendroglioma. Cancer Cell 18(6):669–682. doi:10.1016/j.ccr.2010.10.033
Phillips HS, Kharbanda S, Chen R, Forrest WF, Soriano RH, Wu TD, Misra A, Nigro JM, Colman H, Soroceanu L, Williams PM, Modrusan Z, Feuerstein BG, Aldape K (2006) Molecular subclasses of high-grade glioma predict prognosis, delineate a pattern of disease progression, and resemble stages in neurogenesis. Cancer Cell 9(3):157–173. doi:10.1016/j.ccr.2006.02.019
Pietsch T, Valter MM, Wolf HK, von Deimling A, Huang HJ, Cavenee WK, Wiestler OD (1997) Expression and distribution of vascular endothelial growth factor protein in human brain tumors. Acta Neuropathol 93(2):109–117
Plate KH, Breier G, Weich HA, Risau W (1992) Vascular endothelial growth factor is a potential tumour angiogenesis factor in human gliomas in vivo. Nature 359(6398):845–848. doi:10.1038/359845a0
Raabe EH, Lim KS, Kim JM, Meeker A, Mao XG, Nikkhah G, Maciaczyk J, Kahlert U, Jain D, Bar E, Cohen KJ, Eberhart CG (2011) BRAF activation induces transformation and then senescence in human neural stem cells: a pilocytic astrocytoma model. Clin Cancer Res: An Official J American Assoc Cancer Res 17(11):3590–3599. doi:10.1158/1078-0432.CCR-10-3349
Reifenberger J, Reifenberger G, Liu L, James CD, Wechsler W, Collins VP (1994) Molecular genetic analysis of oligodendroglial tumors shows preferential allelic deletions on 19q and 1p. Am J Pathol 145(5):1175–1190
Reiss Y, Machein MR, Plate KH (2005) The role of angiopoietins during angiogenesis in gliomas. Brain Pathol 15(4):311–317
Sadones J, Michotte A, Veld P, Chaskis C, Sciot R, Menten J, Joossens EJ, Strauven T, D’Hondt LA, Sartenaer D, Califice SF, Bierau K, Svensson C, De Greve J, Neyns B (2009) MGMT promoter hypermethylation correlates with a survival benefit from temozolomide in patients with recurrent anaplastic astrocytoma but not glioblastoma. Eur J Cancer 45(1):146–153. doi:10.1016/j.ejca.2008.09.002
Sanson M, Marie Y, Paris S, Idbaih A, Laffaire J, Ducray F, El Hallani S, Boisselier B, Mokhtari K, Hoang-Xuan K, Delattre JY (2009) Isocitrate dehydrogenase 1 codon 132 mutation is an important prognostic biomarker in gliomas. J clin Oncol: Official J Am Soc Clin Oncol 27(25):4150–4154 JCO.2009.21.9832[pii]/JCO.2009.21.9832
Sasaki M, Knobbe CB, Munger JC, Lind EF, Brenner D, Brustle A, Harris IS, Holmes R, Wakeham A, Haight J, You-Ten A, Li WY, Schalm S, Su SM, Virtanen C, Reifenberger G, Ohashi PS, Barber DL, Figueroa ME, Melnick A, Zuniga-Pflucker JC, Mak TW (2012) IDH1(R132H) mutation increases murine haematopoietic progenitors and alters epigenetics. Nature. doi:10.1038/nature11323
Scherer H (1938) Structural development in gliomas. Am J Cancer 34:333–351
Scherer HJ (1940) A critical review: the pathology of cerebral gliomas. J Neurol Psychiatry 3(2):147–177
Shaifer CA, Huang J, Lin PC (2010) Glioblastoma cells incorporate into tumor vasculature and contribute to vascular radioresistance. Int J Cancer 127(9):2063–2075. doi:10.1002/ijc.25249
Singh D, Chan JM, Zoppoli P, Niola F, Sullivan R, Castano A, Liu EM, Reichel J, Porrati P, Pellegatta S, Qiu K, Gao Z, Ceccarelli M, Riccardi R, Brat DJ, Guha A, Aldape K, Golfinos JG, Zagzag D, Mikkelsen T, Finocchiaro G, Lasorella A, Rabadan R, Iavarone A (2012) Transforming fusions of FGFR and TACC genes in human glioblastoma. Science 337(6099):1231–1235. doi:10.1126/science.1220834
SongTao Q, Lei Y, Si G, YanQing D, HuiXia H, XueLin Z, LanXiao W, Fei Y (2012) IDH mutations predict longer survival and response to temozolomide in secondary glioblastoma. Cancer Sci 103(2):269–273. doi:10.1111/j.1349-7006.2011.02134.x
Sonoda Y, Kumabe T, Nakamura T, Saito R, Kanamori M, Yamashita Y, Suzuki H, Tominaga T (2009) Analysis of IDH1 and IDH2 mutations in Japanese glioma patients. Cancer Sci 100(10):1996–1998 CAS1270 [pii] /j.1349-7006.2009.01270.x
Stupp R, Hegi ME, Mason WP, van den Bent MJ, Taphoorn MJ, Janzer RC, Ludwin SK, Allgeier A, Fisher B, Belanger K, Hau P, Brandes AA, Gijtenbeek J, Marosi C, Vecht CJ, Mokhtari K, Wesseling P, Villa S, Eisenhauer E, Gorlia T, Weller M, Lacombe D, Cairncross JG, Mirimanoff RO (2009) Effects of radiotherapy with concomitant and adjuvant temozolomide versus radiotherapy alone on survival in glioblastoma in a randomised phase III study: 5-year analysis of the EORTC-NCIC trial. Lancet Oncol 10(5):459–466. doi:10.1016/S1470-2045(09)70025-7
Sturm D, Witt H, Hovestadt V, Khuong-Quang DA, Jones DT, Konermann C, Pfaff E, Tonjes M, Sill M, Bender S, Kool M, Zapatka M, Becker N, Zucknick M, Hielscher T, Liu XY, Fontebasso AM, Ryzhova M, Albrecht S, Jacob K, Wolter M, Ebinger M, Schuhmann MU, van Meter T, Fruhwald MC, Hauch H, Pekrun A, Radlwimmer B, Niehues T, von Komorowski G, Durken M, Kulozik AE, Madden J, Donson A, Foreman NK, Drissi R, Fouladi M, Scheurlen W, von Deimling A, Monoranu C, Roggendorf W, Herold-Mende C, Unterberg A, Kramm CM, Felsberg J, Hartmann C, Wiestler B, Wick W, Milde T, Witt O, Lindroth AM, Schwartzentruber J, Faury D, Fleming A, Zakrzewska M, Liberski PP, Zakrzewski K, Hauser P, Garami M, Klekner A, Bognar L, Morrissy S, Cavalli F, Taylor MD, van Sluis P, Koster J, Versteeg R, Volckmann R, Mikkelsen T, Aldape K, Reifenberger G, Collins VP, Majewski J, Korshunov A, Lichter P, Plass C, Jabado N, Pfister SM (2012) Hotspot mutations in H3F3A and IDH1 define distinct epigenetic and biological subgroups of glioblastoma. Cancer Cell 22(4):425–437. doi:10.1016/j.ccr.2012.08.024
Sugiarto S, Persson AI, Munoz EG, Waldhuber M, Lamagna C, Andor N, Hanecker P, Ayers-Ringler J, Phillips J, Siu J, Lim DA, Vandenberg S, Stallcup W, Berger MS, Bergers G, Weiss WA, Petritsch C (2011) Asymmetry-defective oligodendrocyte progenitors are glioma precursors. Cancer Cell 20(3):328–340. doi:10.1016/j.ccr.2011.08.011
Taal W, Dubbink HJ, Zonnenberg CB, Zonnenberg BA, Postma TJ, Gijtenbeek JM, Boogerd W, Groenendijk FH, Kros JM, Kouwenhoven MC, van Marion R, van Heuvel I, van der Holt B, Bromberg JE, Sillevis Smitt PA, Dinjens WN, van den Bent MJ (2011) First-line temozolomide chemotherapy in progressive low-grade astrocytomas after radiotherapy: molecular characteristics in relation to response. Neuro-oncology 13(2):235–241. doi:10.1093/neuonc/noq177
Taylor MD, Poppleton H, Fuller C, Su X, Liu Y, Jensen P, Magdaleno S, Dalton J, Calabrese C, Board J, Macdonald T, Rutka J, Guha A, Gajjar A, Curran T, Gilbertson RJ (2005) Radial glia cells are candidate stem cells of ependymoma. Cancer Cell 8(4):323–335. doi:10.1016/j.ccr.2005.09.001
TCGA (2008) Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 455(7216):1061–1068. doi:10.1038/nature07385
van den Bent MJ, Brandes AA, Rampling R, Kouwenhoven MC, Kros JM, Carpentier AF, Clement PM, Frenay M, Campone M, Baurain JF, Armand JP, Taphoorn MJ, Tosoni A, Kletzl H, Klughammer B, Lacombe D, Gorlia T (2009) Randomized phase II trial of erlotinib versus temozolomide or carmustine in recurrent glioblastoma: EORTC brain tumor group study 26034. J Clin Oncol: Official J Am Soc Clin Oncol 27(8):1268–1274. doi:10.1200/JCO.2008.17.5984
van den Bent MJ, Brandes AA, Taphoorn MJ, Kros JM, Kouwenhoven MC, Delattre JY, Bernsen HJ, Frenay M, Tijssen CC, Grisold W, Sipos L, Enting RH, French PJ, Dinjens WN, Vecht CJ, Allgeier A, Lacombe D, Gorlia T, Hoang-Xuan K (2013) Adjuvant procarbazine, lomustine, and vincristine chemotherapy in newly diagnosed anaplastic oligodendroglioma: long-term follow-up of EORTC brain tumor group study 26951. J Clin Oncol: Official J Am Soc Clin Oncol 31(3):344–350. doi:10.1200/JCO.2012.43.2229
van den Bent MJ, Dubbink HJ, Sanson M, van der Lee-Haarloo CR, Hegi M, Jeuken JW, Ibdaih A, Brandes AA, Taphoorn MJ, Frenay M, Lacombe D, Gorlia T, Dinjens WN, Kros JM (2009) MGMT promoter methylation is prognostic but not predictive for outcome to adjuvant PCV chemotherapy in anaplastic oligodendroglial tumors: a report from EORTC Brain Tumor Group Study 26951. J Clin Oncol: Official J Am Soc Clin Oncol 27(35):5881–5886. doi:10.1200/JCO.2009.24.1034
Verhaak RG, Hoadley KA, Purdom E, Wang V, Qi Y, Wilkerson MD, Miller CR, Ding L, Golub T, Mesirov JP, Alexe G, Lawrence M, O’Kelly M, Tamayo P, Weir BA, Gabriel S, Winckler W, Gupta S, Jakkula L, Feiler HS, Hodgson JG, James CD, Sarkaria JN, Brennan C, Kahn A, Spellman PT, Wilson RK, Speed TP, Gray JW, Meyerson M, Getz G, Perou CM, Hayes DN (2010) Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 17(1):98–110. doi:10.1016/j.ccr.2009.12.020
von Deimling A, von Ammon K, Schoenfeld D, Wiestler OD, Seizinger BR, Louis DN (1993) Subsets of glioblastoma multiforme defined by molecular genetic analysis. Brain Pathol 3(1):19–26
Waha A, Koch A, Hartmann W, Mack H, Schramm J, Sorensen N, Berthold F, Wiestler OD, Pietsch T (2004) Analysis of HIC-1 methylation and transcription in human ependymomas. Int J Cancer 110(4):542–549. doi:10.1002/ijc.20165
Wani K, Armstrong TS, Vera-Bolanos E, Raghunathan A, Ellison D, Gilbertson R, Vaillant B, Goldman S, Packer RJ, Fouladi M, Pollack I, Mikkelsen T, Prados M, Omuro A, Soffietti R, Ledoux A, Wilson C, Long L, Gilbert MR, Aldape K (2012) A prognostic gene expression signature in infratentorial ependymoma. Acta Neuropathol 123(5):727–738. doi:10.1007/s00401-012-0941-4
Watanabe K, Tachibana O, Sata K, Yonekawa Y, Kleihues P, Ohgaki H (1996) Overexpression of the EGF receptor and p53 mutations are mutually exclusive in the evolution of primary and secondary glioblastomas. Brain Pathol 6(3):217–223 discussion 223-214
Watanabe T, Nobusawa S, Kleihues P, Ohgaki H (2009) IDH1 mutations are early events in the development of astrocytomas and oligodendrogliomas. Am J Pathol 174(4):1149–1153. doi:10.2353/ajpath.2009.080958
Wick W, Platten M, Meisner C, Felsberg J, Tabatabai G, Simon M, Nikkhah G, Papsdorf K, Steinbach JP, Sabel M, Combs SE, Vesper J, Braun C, Meixensberger J, Ketter R, Mayer-Steinacker R, Reifenberger G, Weller M (2012) Temozolomide chemotherapy alone versus radiotherapy alone for malignant astrocytoma in the elderly: the NOA-08 randomised, phase 3 trial. Lancet Oncol 13(7):707–715. doi:10.1016/S1470-2045(12)70164-X
Witt H, Mack SC, Ryzhova M, Bender S, Sill M, Isserlin R, Benner A, Hielscher T, Milde T, Remke M, Jones DT, Northcott PA, Garzia L, Bertrand KC, Wittmann A, Yao Y, Roberts SS, Massimi L, Van Meter T, Weiss WA, Gupta N, Grajkowska W, Lach B, Cho YJ, von Deimling A, Kulozik AE, Witt O, Bader GD, Hawkins CE, Tabori U, Guha A, Rutka JT, Lichter P, Korshunov A, Taylor MD, Pfister SM (2011) Delineation of two clinically and molecularly distinct subgroups of posterior fossa ependymoma. Cancer Cell 20(2):143–157. doi:10.1016/j.ccr.2011.07.007
Xu W, Yang H, Liu Y, Yang Y, Wang P, Kim SH, Ito S, Yang C, Xiao MT, Liu LX, Jiang WQ, Liu J, Zhang JY, Wang B, Frye S, Zhang Y, Xu YH, Lei QY, Guan KL, Zhao SM, Xiong Y (2011) Oncometabolite 2-hydroxyglutarate is a competitive inhibitor of alpha-ketoglutarate-dependent dioxygenases. Cancer Cell 19(1):17–30. doi:10.1016/j.ccr.2010.12.014
Yan H, Parsons DW, Jin G, McLendon R, Rasheed BA, Yuan W, Kos I, Batinic-Haberle I, Jones S, Riggins GJ, Friedman H, Friedman A, Reardon D, Herndon J, Kinzler KW, Velculescu VE, Vogelstein B, Bigner DD (2009) IDH1 and IDH2 mutations in gliomas. The New England journal of medicine 360(8):765–773. doi:10.1056/NEJMoa0808710
Yang B, Zhong C, Peng Y, Lai Z, Ding J (2010) Molecular mechanisms of “off-on switch” of activities of human IDH1 by tumor-associated mutation R132H. Cell Res 20(11):1188–1200. doi:10.1038/cr.2010.145
Zagzag D, Lukyanov Y, Lan L, Ali MA, Esencay M, Mendez O, Yee H, Voura EB, Newcomb EW (2006) Hypoxia-inducible factor 1 and VEGF upregulate CXCR4 in glioblastoma: implications for angiogenesis and glioma cell invasion. Lab Invest; J Tech Methods Pathol 86(12):1221–1232. doi:10.1038/labinvest.3700482
Zhao J, Ma W, Zhao H (2014) Loss of heterozygosity 1p/19q and survival in glioma: a meta-analysis. Neuro-oncology 16(1):103–112. doi:10.1093/neuonc/not145
Zhao S, Guan KL (2010) IDH1 mutant structures reveal a mechanism of dominant inhibition. Cell Res 20(12):1279–1281. doi:10.1038/cr.2010.160
Zhao S, Lin Y, Xu W, Jiang W, Zha Z, Wang P, Yu W, Li Z, Gong L, Peng Y, Ding J, Lei Q, Guan KL, Xiong Y (2009) Glioma-derived mutations in IDH1 dominantly inhibit IDH1 catalytic activity and induce HIF-1alpha. Science 324(5924):261–265. doi:10.1126/science.1170944
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Cohen, A.L., Colman, H. (2015). Glioma Biology and Molecular Markers. In: Raizer, J., Parsa, A. (eds) Current Understanding and Treatment of Gliomas. Cancer Treatment and Research, vol 163. Springer, Cham. https://doi.org/10.1007/978-3-319-12048-5_2
Download citation
DOI: https://doi.org/10.1007/978-3-319-12048-5_2
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-12047-8
Online ISBN: 978-3-319-12048-5
eBook Packages: MedicineMedicine (R0)