Abstract
According to the 2016 World Health Organization (WHO) classification of central nervous system tumors, diffuse astrocytic and oligodendroglial tumors are differentiated by the presence of isocitrate dehydrogenase 1 or 2 (IDH1/2) mutation and the combined loss of the short arm of chromosome 1 and the long arm of chromosome 19 (1p/19q co-deletion). IDH-mutant astrocytoma often has p53 and alpha-thalassemia/mental retardation syndrome X-linked (ATRX) mutation, showing the alternative lengthening of telomeres (ALT) phenotype, while IDH-mutant and 1p/19q-co-deleted oligodendroglioma often have wild-type p53 and telomerase reverse transcriptase (TERT) promoter mutation, showing telomerase activation. This study analyzed IDH, ATRX, and TERT promoter mutations, and the correlation between them. Immortalized cells overcome the telomere-related crisis by activating telomerase or ALT. In glioma, telomerase is mainly activated by TERT promoter mutation, while ALT is usually associated with ATRX mutation. Although the mechanism of how ATRX mutation induces ALT remains unclear, ATRX loss alone is believed to be insufficient to induce ALT. Treatments targeting telomere maintenance are promising.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the 2016 World Health Organization (WHO) classification of central nervous system (CNS) tumors, diffuse astrocytic and oligodendroglial tumors are differentiated by the presence of isocitrate dehydrogenase 1 or 2 (IDH1/2) mutation. IDH1/2-mutant gliomas are further subdivided on the basis of the presence of a combined loss of the short arm of chromosome 1 and the long arm of chromosome 19 (1p/19q co-deletion). Diffuse or anaplastic astrocytoma does not have 1p/19q co-deletion, while oligodendroglioma or anaplastic oligodendroglioma shows 1p/19q co-deletion [1]. Astrocytoma often has p53 and alpha-thalassemia/mental retardation syndrome X-linked (ATRX) mutation, showing the alternative lengthening of telomeres (ALT) phenotype, while oligodendroglioma often has wild-type (WT) p53 and telomerase reverse transcriptase (TERT) promoter mutation, showing activation of telomerase [1] (Fig. 1). This study analyzed IDH, ATRX, and TERT promoter mutations and the correlation between them.
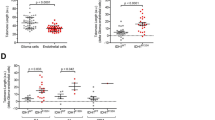
The pathological characteristics of diffuse astrocytoma, IDH-mutant, and oligodendroglioma, IDH-mutant and 1p/19q-codeleted. These figures show representative pictures of diffuse astrocytoma, IDH-mutant, and oligodendroglioma, IDH-mutant and 1p/19q-codeleted. Both types of tumors are usually immunopositive for IDH1R132H. Diffuse astrocytoma, IDH-mutant is often immunopositive for p53 and immunonegative for alpha-thalassemia/mental retardation syndrome X-linked (ATRX), while oligodendroglioma, IDH-mutant and 1p/19q codeleted is often immunonegative for p53 and immunopositive for ATRX having telomerase reverse transcriptase promoter mutation
IDH mutation
IDHs are a group of enzymes that catalyze the oxidative decarboxylation of isocitrate to alpha-ketoglutarate (α-KG). The human body contains three types of IDHs: IDH1, IDH2, and IDH3 [2]. IDH1 is located in the cytosol/peroxisomes, while IDH2 and IDH3 are present in the mitochondria [3]. IDH1 and IDH2 are nicotinamide adenine dinucleotide phosphate (NADP) dependent, while IDH3 is nicotinamide adenine dinucleotide (NAD) dependent [4]. Only IDH3 is associated with the tricarboxylic acid cycle [5]. Mutant IDH1/2 converts α-KG into 2-hydroxyglutarate (2-HG) [6].
IDH1/2 mutation has been reported in not only gliomas [7] but also other tumors, such as acute myelogenous leukemia [8] and chondrosarcoma [9]. In addition, 2-HG accumulation reportedly contributes to tumorigenesis in these cases [7].
After Parsons [10], many studies have been conducted on IDH mutations in glioma [11,12,13,14]. Nearly all IDH mutations involve a single amino acid substitution. The mutation occurs at the arginine residue at codon 132 in IDH1 and codon 140 or 172 in IDH2, although R140 mutations are not found in glioma. The commonest is the R132H mutation (c.395G > A), which accounts for ~ 90% of all IDH mutations. IDH1 and IDH2 mutations usually occur exclusively [7]. IDH1/2 mutation is reportedly detected in 80–90% of grade II and III glioma and 5% of primary glioblastoma [3]. After publication of the 2016 WHO classification of CNS tumors, a glioma needs to have an IDH mutation to be classified as an oligodendroglioma [1]. Since the IDH1R132H alteration occupies ~ 90% of IDH mutations, immunohistochemistry (IHC) evaluation with the IDH1R132H antibody can cover ~ 90% of IDH1/2 mutation cases. In addition, direct DNA sequence analysis is performed to detect the IDH mutations, depending on the facilities.
IDH mutation has been detected as a common mutation between primary and recurrent tumors [15,16,17]. It has also been detected as a common mutation among samples from multiple regions in the same patients [16]. Therefore, IDH mutation is considered to occur at the early stage of tumorigenesis and as a driver mutation in IDH-mutant gliomas [7, 15]. The mechanism underlying gliomagenesis by mutant IDH1/2 has not been clearly elucidated. However, several possible hypotheses are reported (Fig. 2). The structures of 2-HG and α-KG are similar, and 2-HG competes with α-KG and inhibits α-KG-dependent enzymes, such as the ten-eleven translocation (TET) enzyme family of five methylcytosine hydroxylases and Jumonji C domain-containing histone demethylases [15, 18]. Mutant IDH-induced 2-HG in glioma inhibits the TET family, which might lead to global DNA methylation in the glioma CpG island methylation phenotype (G-CIMP) [19]. Gliomas with G-CIMP show gene expression of the proneural type classified by Verhaak et al. [20, 21]. These epigenetic changes are believed to affect gene expression, contributing to tumorigenesis. Histone demethylase inhibition by 2-HG increases histone methylation. This epigenetic status might also affect gene expression [18]. 2-HG reportedly stabilizes hypoxia-inducible factor 1-alpha (HIF-1α) expression and subsequently results in an increase in HIF-1α target gene expression by inhibition of prolyl-hydroxylase enzymes [22]. Consequently, IDH-mutant glioma cells escape from a hypoxic environment [4]. In addition, 2-HG increases reactive oxygen species through the NAPD/NAPDH balance, contributing to tumorigenesis [23].
The hypothesis of mutant IDH1/2-induced tumorigenesis. Mutant IDH1/2-induced 2-hydroxyglutarate (2-HG) competes with α-ketoglutaric acid (α-KG) and inhibits α-KG-dependent enzymes, such as the ten-eleven translocation (TET) enzyme family, histone demethylases, and prolyl-hydroxylase. G-CIMP induced by inhibition TET family, or histone methylation leads to gene expression. 2-HG stabilizes hypoxia-inducible factor 1-alpha (HIF-1α) and expression and increases the expression of HIF-1α target gene by inhibition of prolyl-hydroxylase. 2-HG also increases reactive oxygen species through the NAPD/NAPDH balance. These factors are considered to contribute to tumorigenesis
IDH1R132H is a main mutation among IDH mutations, so IHC evaluation using IDH1R132H antibody is widely used. To check other types of mutations, direct DNA sequence analysis is also performed. IHC is reportedly 100% sensitive and specific in detecting IDH1R132H mutations [24]. The consistency rate between IHC and DNA sequence analysis is reported to be 88% in diffuse glioma patients [25]. As noninvasive methods, mutant IDH1/2-induced 2-HG is reported to be detectable by magnetic resonance spectroscopy [26].
IDH1/2 mutation is considered a driver mutation in IDH-mutant glioma, and inhibition of mutant IDH1/2 is a promising treatment for such gliomas. Several clinical trials are ongoing [4, 27]. In experimental studies, the effects of IDH inhibitors are controversial. IDH inhibitors inhibit the proliferation of IDH1-mutant glioma cells [28], while inhibition of IDH mutation is ineffective [29] or works only in a narrow window of time [30].
Telomere length in gliomas
Telomeres are DNA–protein complexes that protect chromosome ends. These protein complexes are called Shelterin, which comprise TRF1, TRF2, POT1, TIN1, TPP1, and RAP1 [31]. Telomeres in vertebrates comprise a region of 3000–20,000 TTAGGG repeats at the ends of chromosomes [32]. The length of a telomere shortens after each cell division, and cells go to the arrest stage after limitation [33]. To overcome this issue, many types of tumor cells maintain the telomere length via telomerase activation, while some types of tumors elongate the telomere length by telomere-independent manner, which is known as ALT [32]. These oncogenic changes in glioma are generally exclusive [34]. Telomerase has two principle components, the 1132-amino acid telomerase reverse transcriptase (TERT) and an associated telomerase RNA molecule (TERC) [35]. TERT promoter mutation increases TERT expression and activates telomerase activity [36, 37]. Another mechanism is ALT, with which ATRX, death-associated protein 6 (DAXX), and histone H3.3 are reportedly associated [32, 38].
IDH-mutant diffuse or anaplastic astrocytoma shows the ALT phenotype, while IDH-mutant and 1p/19q-co-deleted oligodendroglioma or IDH-WT glioblastoma shows TERT promoter mutation [36, 39, 40]. Although both tumor types have the same IDH mutation, why astrocytoma selects the ALT phenotype and oligodendroglioma selects TERT promoter mutation to resolve telomeric dysfunction and maintain telomere length remains unknown.
Telomerase
TERT promoter mutation occurs at positions 124 and 146 bp upstream of the TERT ATG start site, called C228T and C250T, respectively [34]. These mutations are exclusive. Messenger RNA (mRNA) expression of TERT increases in gliomas with TERT promoter mutation compared to gliomas without TERT promoter mutation [36, 37]. The telomere length is shorter in glioma with TERT promoter mutation compared to glioma without TERT promoter mutation [37]. TERT promoter mutations generate identical 11 bp sequences that form a de novo binding site. The E26 transformation-specific transcription factor GA-binding protein selectively binds to the site and activates TERT [41]. TERT promoter mutation is believed to be associated with tumorigenesis in two phases: by promoting immortalization and genomic instability. In the first phase, TERT promoter mutation extends the cellular life span by extending the shortest telomeres but does not prevent most telomere shortening. In the second phase, the critically short telomeres inducing genome instability and telomerase are upregulated. Consequently, cells maintain growth [42]. TERT promoter mutation is considered the main mechanism of TERT transcript upregulation; however, other mechanisms have also been reported [43, 44]. For example, mutant IDH activates the TERT promoter, increasing histone lysine methylation and c-Myc/Max binding at the TERT promoter [44].
Several studies have reported the correlation between TERT promoter mutation and prognosis in glioma [39, 45,46,47], but few studies have compared prognoses within the same diagnosis based on the 2016 WHO classification [46]. Pekmezci et al. showed that of 291 grade II or III oligodendroglioma, IDH-mutant and 1p/19q-codeleted patients, 94% had only TERT promoter mutation, 0.69% had only ATRX mutation, 1.7% had both TERT promoter and ATRX mutation, and 4% had neither TERT promoter nor ATRX mutation [46]. The group with TERT promoter mutation had significant better overall survival (OS) compared to the group with TERT promoter WT group [46]. Of 154 IDH-WT astrocytoma patients, 60% showed only TERT promoter mutation, and 2% had both TERT promoter and ATRX mutation. The group with TERT promoter mutation had significantly worse OS compared to the TERT promoter WT group [46]. Akyerli et al. classified hemispheric diffuse glioma using the IDH and TERT promoter mutation status: both IDH and TERT promoter mutation, only IDH mutation, only TERT promoter mutation, and neither IDH nor TERT promoter mutation. Every group had a distinct demographic, anatomical, clinical, and prognostic correlation [39], and the TERT promoter mutation status of tumors was unchanged over time or recurred [39]. Similarly, in a meta-analysis, Vuong et al. classified WHO grade II or III glioma patients into four groups on the basis of the IDH and TERT promoter mutation status and showed the difference in OS between the groups (both IDH and TERT promoter mutation > only IDH mutation > neither IDH nor TERT promoter mutation > only TERT promoter mutation) [47].
Telomere maintenance is expected to be a new therapeutic target. Several approaches have been followed, such as vaccines, antisense oligonucleotides, and small-molecule inhibitors, to inhibit TERT, TRRC, or TERT promoter mutation [48,49,50,51,52,53]. However, these reagents might be ineffective in tumors with the ALT phenotype.
ALT
Although telomerase activity is the most frequent mechanism for maintaining telomere length, 10%–15% of cancers also show a telomerase-independent mechanism for elongating telomeres (ALT) [54]. Although the ALT phenotype is uncommon in tumors, it is common in certain cancer subtypes, including glioma and sarcoma [32, 54]. Among gliomas, ALT is often detected in WHO grade II or III IDH-mutant astrocytoma, and IDH-mutant glioblastoma [46]. Tumor cells with the ALT phenotype depend on the activation of a homologous recombination DNA repair mechanism to maintain telomere length [55]. ALT often begins with the loss of chromatin-remodeling proteins in telomeres, resulting in DNA damage response, recombination, and abnormal protein behavior, which reportedly initiates ALT [32, 55].
Addition of ATRX suppresses the ALT phenotype, so ATRX loss is associated with the ALT phenotype [56]. However, studies have reported that ATRX loss alone is insufficient to induce the ALT phenotype [56, 57]. In contrast, IDH1R132H overexpression in glioma cell lines downregulates ATRX and induces telomere lengthening consistent with ALT [58]. The authors suggested that IDH1R132H alone is sufficient to diminish ATRX expression and induces ALT. Inconsistent with their findings, neither overexpressed IDH1R132H alone nor ATRX knockout alone induces the ALT phenotype [44, 57]. Exogenous IDH1R132H expression combined with ATRX mutation induces the ALT phenotype [57]. Mutant IDH1 downregulates a Shelterin protein, RAP1, which leads to telomere dysfunction. Mutant IDH1 also downregulates XRCC, which leads to nonhomologous end-joining (NHEJ) inhibition. This downregulation of RAP1 and XRCC reportedly contributes to ALT in glioma cells with ATRX loss [57].
The cIMPACT Update 2 report showed that diffuse astrocytic-appearing WHO grade II or III glioma with IDH mutation and ATRX nuclear expression loss and/or strong, diffuse p53 immunopositivity can be diagnosed as diffuse astrocytoma, IDH-mutant or anaplastic astrocytoma, IDH-mutant without checking the 1p/19q co-deletion status [59]. IHC analysis for ATRX, not sequence analysis for ATRX, was described in this report. Since ATRX is relatively large, it is difficult to perform the assay to detect the mutation as a routine test. Therefore, IHC analysis is usually performed as an alternative method for evaluating ATRX mutation. Ikemura et al. (2016) examined 193 patients and divided them into three groups: ATRX loss with staining loss in > 90% of tumor cells (n = 43), ATRX indeterminate with staining loss in 10%–90% of tumor cells (n = 0), and ATRX retained with staining loss in < 10% of tumor cells (n = 150) [60]. Other studies divided patients into two groups, with nuclear ATRX loss and with nuclear ATRX retention, using a cutoff value of 10% [61, 62].
ATRX mutation is associated with prognosis in glioma [63, 64]. However, in most cases, tumor diagnosis is performed on the basis of pre-2016 WHO classification. Since most IDH-mutant astrocytomas and IDH-mutant glioblastomas show ATRX loss and most IDH-mutant and 1p/19q-co-deleted oligodendrogliomas and IDH-WT glioblastomas show ATRX expression, the comparison between low-grade gliomas or between glioblastomas might be a comparison between astrocytoma and oligodendroglioma or between IDH-WT glioblastoma and IDH-mutant glioblastoma. In a comparison within the same diagnosis of 220 cases of IDH-mutant diffuse astrocytoma and 181 cases of IDH-mutant anaplastic astrocytoma, 78% had only ATRX mutation, 2% had both TERT and ATRX mutation, 5% had only TERT mutation, and 16% had neither TERT nor ATRX mutation. Neither TERT nor ATRX status was associated with survival [46]. In IDH-mutant glioblastoma patients, the rate of each mutation is similar to that in IDH-mutant WHO grade II and III astrocytoma patients. Sixty-three percentage had only ATRX mutation, 6% had both TERT and ATRX mutation, 12% had only TERT mutation, and 20% had neither TERT nor ATRX mutation. Neither TERT nor ATRX status was associated with survival [46].
Several new treatments have been recommended for targeting tumor cells with the ALT phenotype [35, 54]. CRISPR-Cas9 screening detected Wee1 as a target in ATRX-null tumor cells [65]. The ATR inhibitor reportedly disrupts ALT and triggers chromosome fragmentation and apoptosis in ALT cells [66]. Histone deacetylase inhibitors might suppress ALT through NuRD inhibition [67]. Although these treatments are expected to be effective in tumor cells with the ALT phenotype, treatments that are effective independent of activated telomerase or ALT are promising. G-quadruplexes are tetra-stranded DNA structures formed by guanine-rich sequences. Studies have reported a G-quadruplex stabilizer that activated the pathways of response to DNA damage and induce senescence in both telomerase-positive and ALT phenotype cells [68].
Conclusion
In this review, we discussed the three mutations of IDH, ATRX, and TERT promoter in glioma. It remains unclear why astrocytoma, IDH-mutant has ALT phenotype, whereas oligodendroglioma, IDH-mutant and 1p/19q-codeleted showed activated telomerase. Further studies may clarify it and new treatments targeting maintenance of telomere are promising.
References
Louis DN, Perry A, Reifenberger G, von Deimling A, Figarella-Branger D, Cavenee WK, Ohgaki H, Wiestler OD, Kleihues P, Ellison DW (2016) The 2016 World Health Organization classification of tumors of the central nervous system: a summary. Acta Neuropathol 131(6):803–820
May JL, Kouri FM, Hurley LA, Liu J, Tommasini-Ghelfi S, Ji Y, Gao P, Calvert AE, Lee A, Chandel NS, Davuluri RV, Horbinski CM, Locasale JW, Stegh AH (2019) IDH3α regulates one-carbon metabolism in glioblastoma. Sci Adv 5(1):eaat0456
Miller JJ, Shih HA, Andronesi OC, Cahill DP (2017) Isocitrate dehydrogenase-mutant glioma: evolving clinical and therapeutic implications. Cancer 123(23):4535–4546
Huang J, Yu J, Tu L, Huang N, Li H, Luo Y (2019) Isocitrate dehydrogenase mutations in glioma: from basic discovery to therapeutics development. Front Oncol 9:506
Ichimura K, Narita Y, Hawkins CE (2015) Diffusely infiltrating astrocytomas: pathology, molecular mechanisms and markers. Acta Neuropathol 129:789–808
Dang L, White DW, Gross S, Bennett BD, Bittinger MA, Driggers EM, Fantin VR, Jang HG, Jin S, Keenan MC, Marks KM, Prins RM, Ward PS, Yen KE, Liau LM, Rabinowitz JD, Cantley LC, Thompson CB, Vander Heiden MG, Su SM (2009) Cancer-associated IDH1 mutations produce 2-hydroxyglutarate. Nature 462:739–744
Ohba S, Hirose Y (2016) Biological significance of mutant isocitrate dehydrogenase 1 and 2 in gliomagenesis. Neurol Med Chir (Tokyo) 56(4):170–179
Patel JP, Gönen M, Figueroa ME, Fernandez H, Sun Z, Racevskis J, Van Vlierberghe P, Dolgalev I, Thomas S, Aminova O, Huberman K, Cheng J, Viale A, Socci ND, Heguy A, Cherry A, Vance G, Higgins RR, Ketterling RP, Gallagher RE, Litzow M, van den Brink MR, Lazarus HM, Rowe JM, Luger S, Ferrando A, Paietta E, Tallman MS, Melnick A, Abdel-Wahab O, Levine RL (2012) Prognostic relevance of integrated genetic profiling in acute myeloid leukemia. N Engl J Med 366(12):1079–1089
Amary MF, Bacsi K, Maggiani F, Damato S, Halai D, Berisha F, Pollock R, O'Donnell P, Grigoriadis A, Diss T, Eskandarpour M, Presneau N, Hogendoorn PC, Futreal A, Tirabosco R, Flanagan AM (2011) IDH1 and IDH2 mutations are frequent events in central chondrosarcoma and central and periosteal chondromas but not in other mesenchymal tumours. J Pathol 224(3):334–343
Parsons DW, Jones S, Zhang X, Lin JC, Leary RJ, Angenendt P, Mankoo P, Carter H, Siu IM, Gallia GL, Olivi A, McLendon R, Rasheed BA, Keir S, Nikolskaya T, Nikolsky Y, Busam DA, Tekleab H, Diaz LA Jr, Hartigan J, Smith DR, Strausberg RL, Marie SK, Shinjo SM, Yan H, Riggins GJ, Bigner DD, Karchin R, Papadopoulos N, Parmigiani G, Vogelstein B, Velculescu VE, Kinzler KW (2008) An integrated genomic analysis of human glioblastoma multiforme. Science 321(5897):1807–1812
Hartmann C, Meyer J, Balss J, Capper D, Mueller W, Christians A, Felsberg J, Wolter M, Mawrin C, Wick W, Weller M, Herold-Mende C, Unterberg A, Jeuken JW, Wesseling P, Reifenberger G, von Deimling A (2009) Type and frequency of IDH1 and IDH2 mutations are related to astrocytic and oligodendroglial differentiation and age: a study of 1,010 diffuse gliomas. Acta Neuropathol 118:469–474
Ichimura K, Pearson DM, Kocialkowski S, Bäcklund LM, Chan R, Jones DT, Collins VP (2009) IDH1 mutations are present in the majority of common adult gliomas but rare in primary glioblastomas. Neuro-oncology 11:341–347
Mukasa A, Takayanagi S, Saito K, Shibahara J, Tabei Y, Furuya K, Ide T, Narita Y, Nishikawa R, Ueki K, Saito N (2012) Significance of IDH mutations varies with tumor histology, grade, and genetics in Japanese glioma patients. Cancer Sci 103:587–592
Sonoda Y, Kumabe T, Nakamura T, Saito R, Kanamori M, Yamashita Y, Suzuki H, Tominaga T (2009) Analysis of IDH1 and IDH2 mutations in Japanese glioma patients. Cancer Sci 100:1996–1998
Johnson BE, Mazor T, Hong C, Barnes M, Aihara K, McLean CY, Fouse SD, Yamamoto S, Ueda H, Tatsuno K, Asthana S, Jalbert LE, Nelson SJ, Bollen AW, Gustafson WC, Charron E, Weiss WA, Smirnov IV, Song JS, Olshen AB, Cha S, Zhao Y, Moore RA, Mungall AJ, Jones SJM, Hirst M, Marra MA, Saito N, Aburatani H, Mukasa A, Berger MS, Chang SM, Taylor BS, Costello JF (2014) Mutational analysis reveals the origin and therapy-driven evolution of recurrent glioma. Science 343(6167):189–193
Suzuki H, Aoki K, Chiba K, Sato Y, Shiozawa Y, Shiraishi Y, Shimamura T, Niida A, Motomura K, Ohka F, Yamamoto T, Tanahashi K, Ranjit M, Wakabayashi T, Yoshizato T, Kataoka K, Yoshida K, Nagata Y, Sato-Otsubo A, Tanaka H, Sanada M, Kondo Y, Nakamura H, Mizoguchi M, Abe T, Muragaki Y, Watanabe R, Ito I, Miyano S, Natsume A, Ogawa S (2015) Mutational landscape and clonal architecture in grade II and III gliomas. Nat Genet 47(5):458–468
Zhang Z, Chan AK, Ding X, Li Y, Zhang R, Chen L, Liu Y, Wang Y, Xiong J, Ng HK, Yao Y, Zhou L (2017) Glioma groups classified by IDH and TERT promoter mutations remain stable among primary and recurrent gliomas. Neuro Oncol 19(7):1008–1010
Xu W, Yang H, Liu Y, Yang Y, Wang P, Kim SH, Ito S, Yang C, Wang P, Xiao MT, Liu LX, Jiang WQ, Liu J, Zhang JY, Wang B, Frye S, Zhang Y, Xu YH, Lei QY, Guan KL, Zhao SM, Xiong Y (2011) Oncometabolite 2-hydroxyglutarate is a competitive inhibitor of α-ketoglutarate-dependent dioxygenases. Cancer Cell 19(1):17–30
Malta TM, de Souza CF, Sabedot TS, Silva TC, Mosella MS, Kalkanis SN, Snyder J, Castro AVB, Noushmehr H (2018) Glioma CpG island methylator phenotype (G-CIMP): biological and clinical implications. Neuro Oncol 20(5):608–620
Noushmehr H, Weisenberger DJ, Diefes K, Phillips HS, Pujara K, Berman BP, Pan F, Pelloski CE, Sulman EP, Bhat KP, Verhaak RG, Hoadley KA, Hayes DN, Perou CM, Schmidt HK, Ding L, Wilson RK, Van Den Berg D, Shen H, Bengtsson H, Neuvial P, Cope LM, Buckley J, Herman JG, Baylin SB, Laird PW, Aldape K; Cancer Genome Atlas Research Network (2010) Identification of a CpG island methylator phenotype that defines a distinct subgroup of glioma. Cancer Cell 17(5):510–522
Verhaak RG, Hoadley KA, Purdom E, Wang V, Qi Y, Wilkerson MD, Miller CR, Ding L, Golub T, Mesirov JP, Alexe G, Lawrence M, O'Kelly M, Tamayo P, Weir BA, Gabriel S, Winckler W, Gupta S, Jakkula L, Feiler HS, Hodgson JG, James CD, Sarkaria JN, Brennan C, Kahn A, Spellman PT, Wilson RK, Speed TP, Gray JW, Meyerson M, Getz G, Perou CM, Hayes DN, Cancer Genome Atlas Research Network (2010) Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 17:98–110
Zhang C, Moore LM, Li X, Yung WK, Zhang W (2013) IDH1/2 mutations target a key hallmark of cancer by deregulating cellular metabolism in glioma. Neuro-oncology 15:1114–1126
Rinaldi M, Caffo M, Minutoli L, Marini H, Abbritti RV, Squadrito F, Trichilo V, Valenti A, Barresi V, Altavilla D, Passalacqua M, Caruso GROS, Gliomas B (2016) An Overview of Potential and Innovative Therapeutic Strategies. Int J Mol Sci 17(6):E984
Capper D, Weissert S, Balss J, Habel A, Meyer J, Jäger D, Ackermann U, Tessmer C, Korshunov A, Zentgraf H, Hartmann C, von Deimling A (2010) Characterization of R132H mutation-specific IDH1 antibody binding in brain tumors. Brain Pathol 20:245–254
Agarwal S, Sharma MC, Jha P, Pathak P, Suri V, Sarkar C, Chosdol K, Suri A, Kale SS, Mahapatra AK, Jha P (2013) Comparative study of IDH1 mutations in gliomas by immunohistochemistry and DNA sequencing. Neuro Oncol 15:718–726
Di Ieva A, Magnussen JS, McIntosh J, Mulcahy MJ, Pardey M, Choi C (2020) Magnetic resonance spectroscopic assessment of isocitrate dehydrogenase status in gliomas: the new frontiers of spectrobiopsy in neurodiagnostics. World Neurosurg 133:e421–e442
Golub D, Iyengar N, Dogra S, Wong T, Bready D, Tang K, Modrek AS, Placantonakis DG (2019) Mutant isocitrate dehydrogenase inhibitors as targeted cancer therapeutics. Front Oncol 17(9):417
Rohle D, Popovici-Muller J, Palaskas N, Turcan S, Grommes C, Campos C, Tsoi J, Clark O, Oldrini B, Komisopoulou E, Kunii K, Pedraza A, Schalm S, Silverman L, Miller A, Wang F, Yang H, Chen Y, Kernytsky A, Rosenblum MK, Liu W, Biller SA, Su SM, Brennan CW, Chan TA, Graeber TG, Yen KE, Mellinghoff IK (2013) An inhibitor of mutant IDH1 delays growth and promotes differentiation of glioma cells. Science 340(6132):626–630
Tateishi K, Wakimoto H, Iafrate AJ, Tanaka S, Loebel F, Lelic N, Wiederschain D, Bedel O, Deng G, Zhang B, He T, Shi X, Gerszten RE, Zhang Y, Yeh JJ, Curry WT, Zhao D, Sundaram S, Nigim F, Koerner MVA, Ho Q, Fisher DE, Roider EM, Kemeny LV, Samuels Y, Flaherty KT, Batchelor TT, Chi AS, Cahill DP (2015) Extreme vulnerability of IDH1 mutant cancers to NAD+ depletion. Cancer Cell 28(6):773–784
Johannessen TA, Mukherjee J, Viswanath P, Ohba S, Ronen SM, Bjerkvig R, Pieper RO (2016) Rapid conversion of mutant IDH1 from driver to passenger in a model of human gliomagenesis. Mol Cancer Res 14(10):976–983
de Lange T (2005) Shelterin: the protein complex that shapes and safeguards human telomeres. Genes Dev 19(18):2100–2110
Fan HC, Chen CM, Chi CS, Tsai JD, Chiang KL, Chang YK, Lin SZ, Harn HJ (2019) Targeting telomerase and ATRX/DAXX inducing tumor senescence and apoptosis in the malignant glioma. Int J Mol Sci 20(1):E200
Bodnar AG, Ouellette M, Frolkis M, Holt SE, Chiu CP, Morin GB, Harley CB, Shay JW, Lichtsteiner S, Wright WE (1998) Extension of life-span by introduction of telomerase into normal human cells. Science 279(5349):349–352
Killela PJ, Reitman ZJ, Jiao Y, Bettegowda C, Agrawal N, Diaz LA Jr, Friedman AH, Friedman H, Gallia GL, Giovanella BC, Grollman AP, He TC, He Y, Hruban RH, Jallo GI, Mandahl N, Meeker AK, Mertens F, Netto GJ, Rasheed BA, Riggins GJ, Rosenquist TA, Schiffman M, IeM S, Theodorescu D, Torbenson MS, Velculescu VE, Wang TL, Wentzensen N, Wood LD, Zhang M, McLendon RE, Bigner DD, Kinzler KW, Vogelstein B, Papadopoulos N, Yan H (2013) TERT promoter mutations occur frequently in gliomas and a subset of tumors derived from cells with low rates of self-renewal. Proc Natl Acad Sci USA 110(15):6021–6026
Kent T, Gracias D, Shepherd S, Clynes D (2020) Alternative lengthening of telomeres in pediatric cancer: mechanisms to therapies. Front Oncol 9:1518
Arita H, Narita Y, Fukushima S, Tateishi K, Matsushita Y, Yoshida A, Miyakita Y, Ohno M, Collins VP, Kawahara N, Shibui S, Ichimura K (2013) Upregulating mutations in the TERT promoter commonly occur in adult malignant gliomas and are strongly associated with total 1p19q loss. Acta Neuropathol 126(2):267–276
Heidenreich B, Rachakonda PS, Hosen I, Volz F, Hemminki K, Weyerbrock A, Kumar R (2015) TERT promoter mutations and telomere length in adult malignant gliomas and recurrences. Oncotarget 6(12):10617–10633
Haase S, Garcia-Fabiani MB, Carney S, Altshuler D, Núñez FJ, Méndez FM, Núñez F, Lowenstein PR, Castro MG (2018) Mutant ATRX: uncovering a new therapeutic target for glioma. Expert Opin Ther Target 22(7):599–613
Akyerli CB, Yüksel Ş, Can Ö, Erson-Omay EZ, Oktay Y, Coşgun E, Ülgen E, Erdemgil Y, Sav A, von Deimling A, Günel M, Yakıcıer MC, Pamir MN, Özduman K (2018) Use of telomerase promoter mutations to mark specific molecular subsets with reciprocal clinical behavior in IDH mutant and IDH wild-type diffuse gliomas. J Neurosurg 128(4):1102–1114
Lee Y, Koh J, Kim SI, Won JK, Park CK, Choi SH, Park SH (2017) The frequency and prognostic effect of TERT promoter mutation in diffuse gliomas. Acta Neuropathol Commun 5(1):62
Bell RJ, Rube HT, Kreig A, Mancini A, Fouse SD, Nagarajan RP, Choi S, Hong C, He D, Pekmezci M, Wiencke JK, Wrensch MR, Chang SM, Walsh KM, Myong S, Song JS, Costello JF (2015) Cancer: the transcription factor GABP selectively binds and activates the mutant TERT promoter in cancer. Science 348(6238):1036–1039
Chiba K, Lorbeer FK, Shain AH, McSwiggen DT, Schruf E, Oh A, Ryu J, Darzacq X, Bastian BC, Hockemeyer D (2017) Mutations in the promoter of the telomerase gene TERT contribute to tumorigenesis by a two-step mechanism. Science 357(6358):1416–1420
Masui K, Komori T, Kato Y, Masutomi K, Ichimura K, Ogasawara S, Kaneko MK, Oki H, Suzuki H, Nitta M, Maruyama T, Muragaki Y, Kawamata T, Sawada T, Shibata N (2018) Elevated TERT expression in TERT-wildtype adult diffuse gliomas: histological evaluation with a novel TERT-specific antibody. Biomed Res Int 2018:7945845
Ohba S, Mukherjee J, Johannessen TC, Mancini A, Chow TT, Wood M, Jones L, Mazor T, Marshall RE, Viswanath P, Walsh KM, Perry A, Bell RJ, Phillips JJ, Costello JF, Ronen SM, Pieper RO (2016) Mutant IDH1 expression drives TERT promoter reactivation as part of the cellular transformation process. Cancer Res 76(22):6680–6689
Labussière M, Di Stefano AL, Gleize V, Boisselier B, Giry M, Mangesius S, Bruno A, Paterra R, Marie Y, Rahimian A, Finocchiaro G, Houlston RS, Hoang-Xuan K, Idbaih A, Delattre JY, Mokhtari K, Sanson M (2014) TERT promoter mutations in gliomas, genetic associations and clinico-pathological correlations. Br J Cancer 111(10):2024–2032
Pekmezci M, Rice T, Molinaro AM, Walsh KM, Decker PA, Hansen H, Sicotte H, Kollmeyer TM, McCoy LS, Sarkar G, Perry A, Giannini C, Tihan T, Berger MS, Wiemels JL, Bracci PM, Eckel-Passow JE, Lachance DH, Clarke J, Taylor JW, Luks T, Wiencke JK, Jenkins RB, Wrensch MR (2017) Adult infiltrating gliomas with WHO 2016 integrated diagnosis: additional prognostic roles of ATRX and TERT. Acta Neuropathol 133(6):1001–1016
Vuong HG, Altibi AMA, Duong UNP, Ngo HTT, Pham TQ, Chan AK, Park CK, Fung KM, Hassell L (2017) TERT promoter mutation and its interaction with IDH mutations in glioma: combined TERT promoter and IDH mutations stratifies lower-grade glioma into distinct survival subgroups—a meta-analysis of aggregate data. Crit Rev Oncol Hematol 120:1–9
Li X, Qian X, Wang B, Xia Y, Zheng Y, Du L, Xu D, Xing D, DePinho RA, Lu Z (2020) Programmable base editing of mutated TERT promoter inhibits brain tumour growth. Nat Cell Biol 2020:1
Mancini A, Xavier-Magalhães A, Woods WS, Nguyen KT, Amen AM, Hayes JL, Fellmann C, Gapinske M, McKinney AM, Hong C, Jones LE, Walsh KM, Bell RJA, Doudna JA, Costa BM, Song JS, Perez-Pinera P, Costello JF (2018) Disruption of the β1L isoform of GABP reverses glioblastoma replicative immortality in a TERT promoter mutation-dependent manner. Cancer Cell 34(3):513–528.e8
Ouellette MM, Wright WE, Shay JW (2011) Targeting telomerase-expressing cancer cells. J Cell Mol Med 15(7):1433–1442
Parsch D, Brassat U, Brümmendorf TH, Fellenberg J (2008) Consequences of telomerase inhibition by BIBR1532 on proliferation and chemosensitivity of chondrosarcoma cell lines. Cancer Invest 26(6):590–596
Röth A, Harley CB, Baerlocher GM (2010) Imetelstat (GRN163L)–telomerase-based cancer therapy. Recent Results Cancer Res 184:221–234
Takahashi M, Miki S, Fujimoto K, Fukuoka K, Matsushita Y, Maida Y, Yasukawa M, Hayashi M, Shinkyo R, Kikuchi K, Mukasa A, Nishikawa R, Tamura K, Narita Y, Hamada A, Masutomi K, Ichimura K (2019) Eribulin penetrates brain tumor tissue and prolongs survival of mice harboring intracerebral glioblastoma xenografts. Cancer Sci 110(7):2247–2257
Sommer A, Royle NJ (2020) ALT: a multi-faceted phenomenon. Genes (Basel) 11(2):E133
Cesare AJ, Reddel RR (2010) Alternative lengthening of telomeres: models, mechanisms and implications. Nat Rev Genet 11(5):319–330
Napier CE, Huschtscha LI, Harvey A, Bower K, Noble JR, Hendrickson EA, Reddel RR (2015) ATRX represses alternative lengthening of telomeres. Oncotarget 6(18):16543–16558
Mukherjee J, Johannessen TC, Ohba S, Chow TT, Jones L, Pandita A, Pieper RO (2018) Mutant IDH1 cooperates with ATRX loss to drive the alternative lengthening of telomere phenotype in glioma. Cancer Res 78(11):2966–2977
Ferreira MSV, Sørensen MD, Pusch S, Beier D, Bouillon AS, Kristensen BW, Brümmendorf TH, Beier CP, Beier F (2020) Alternative lengthening of telomeres is the major telomere maintenance mechanism in astrocytoma with isocitrate dehydrogenase 1 mutation. J Neurooncol (in press)
Louis DN, Giannini C, Capper D, Paulus W, Figarella-Branger D, Lopes MB, Batchelor TT, Cairncross JG, van den Bent M, Wick W, Wesseling P (2018) cIMPACT-NOW update 2: diagnostic clarifications for diffuse midline glioma, H3 K27M-mutant and diffuse astrocytoma/anaplastic astrocytoma. IDH-Mutant Acta Neuropathol 135(4):639–642
Ikemura M, Shibahara J, Mukasa A, Takayanagi S, Aihara K, Saito N, Aburatani H, Fukayama M (2016) Utility of ATRX immunohistochemistry in diagnosis of adult diffuse gliomas. Histopathology 69(2):260–267
Ebrahimi A, Skardelly M, Bonzheim I, Ott I, Mühleisen H, Eckert F, Tabatabai G, Schittenhelm J (2016) ATRX immunostaining predicts IDH and H3F3A status in gliomas. Acta Neuropathol Commun 4(1):60
Kuwahara K, Ohba S, Nakae S, Hattori N, Pareira ES, Yamada S, Sasaki H, Abe M, Hasegawa M, Hirose Y (2019) Clinical, histopathological, and molecular analyses of IDH-wild-type WHO grade II-III gliomas to establish genetic predictors of poor prognosis. Brain Tumor Pathol 36(4):135–143
Cai J, Zhu P, Zhang C, Li Q, Wang Z, Li G, Wang G, Yang P, Li J, Han B, Jiang C, Sun Y, Jiang T (2016) Detection of ATRX and IDH1-R132H immunohistochemistry in the progression of 211 paired gliomas. Oncotarget 7(13):16384–16395
Xie Y, Tan Y, Yang C, Zhang X, Xu C, Qiao X, Xu J, Tian S, Fang C, Kang C (2019) Omics-based integrated analysis identified ATRX as a biomarker associated with glioma diagnosis and prognosis. Cancer Biol Med 16(4):784–796
Liang J, Zhao H, Diplas BH, Liu S, Liu J, Wang D, Lu Y, Zhu Q, Wu J, Wang W, Yan H, Zeng YX, Wang X, Jiao Y (2020) Genome-wide CRISPR-Cas9 screen reveals selective vulnerability of ATRX-mutant cancers to WEE1 inhibition. Cancer Res 80(3):510–523
Flynn RL, Cox KE, Jeitany M, Wakimoto H, Bryll AR, Ganem NJ, Bersani F, Pineda JR, Suvà ML, Benes CH, Haber DA, Boussin FD, Zou L (2015) Alternative lengthening of telomeres renders cancer cells hypersensitive to ATR inhibitors. Science 347(6219):273–277
Conomos D, Reddel RR, Pickett HA (2014) NuRD-ZNF827 recruitment to telomeres creates a molecular scaffold for homologous recombination. Nat Struct Mol Biol 21(9):760–770
Huang FC, Chang CC, Wang JM, Chang TC, Lin JJ (2012) Induction of senescence in cancer cells by the G-quadruplex stabilizer, BMVC4, is independent of its telomerase inhibitory activity. Br J Pharmacol 167(2):393–406
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Ohba, S., Kuwahara, K., Yamada, S. et al. Correlation between IDH, ATRX, and TERT promoter mutations in glioma. Brain Tumor Pathol 37, 33–40 (2020). https://doi.org/10.1007/s10014-020-00360-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10014-020-00360-4