Abstract
The ecological distribution of yeasts through the various stages of winemaking process have an important role on the characteristics of wines influencing both the analytical composition and the sensory profile of product. The yeast community of grapes, affected by several biotic and abiotic factors, is directly involved in grape juice fermentation taking part more or less actively in both spontaneous or inoculated fermentations. The yeast community of winery also take part in winemaking process influencing positively or negatively the final wine composition. Selected starter cultures of Saccharomycs cerevisiae are usually added to control the winemaking process and to achieve specific desired enological traits. The aim is that to dominate indigenous yeasts belonging to the vineyard environment, winery facilities and cellar equipment obtaining wines tailored to consumer requests. More recently, the revaluation of the role of non-Saccharomyces yeasts allowed to enologists to obtain some peculiar characters and more complex aroma to the wines. In this regard, the use of controlled mixed fermentation with selected non-Saccharomyces and S. cerevisiae wine yeasts represents a novelty in oenological field.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
1 Introduction
Wine fermentation is a complex biotechnological process in which yeasts play an essential role. In this context, the ecological distribution of yeasts through the production chain of wine production is a crucial factor the quality of wine. Although Saccharomyces cerevisiae is the main microorganism involved during the transformation of grape juice in wine, many other yeasts species occur in grape juice fermentation and may actively take part in the process. Nowadays, selected starter cultures of S. cerevisiae are usually added by oenologists to control the fermentative process and to achieve specific desired enological characters (inoculated fermentations). The aim is that to dominate indigenous yeasts belonging to the vineyard environment, winery facilities and cellar equipment. Indeed, it has been clearly demonstrated that the microbial population is a multi-comprehensive consortium that includes filamentous fungi, yeasts and bacteria with different physiological characteristics and different impact on the grape metabolome and final wine quality (Pinto et al. 2015; Verginer et al. 2010). The composition of grape microbiota can be influenced, in complexity and frequency, by various abiotic or biotic factors, including climatic conditions, temperature, UV exposure, rainfall, sunlight and winds, ripeness or variety of grapes and interaction within strains that co-habitat. The study and the monitoring of microbiota of grape barriers is important to recognize the evolution of yeasts and the relationship between the microorganisms, fundamental to predict the progress of fermentative process. The use of conventional and innovative molecular methods allow to analyse the microbial members of consortium from grape berries to wine. Indeed, spontaneous wine fermentation is typically carried out by a complex evolution of microorganisms extensively examined during the years. Now, it is well established that together with S. cerevisiae, non-Saccharomyces species actively participate during the alcoholic fermentation and their contribution was recently positively revaluated. Non-Saccharomyces yeasts, coming from grape berry and winery environment, if well managed, can positively impact on the analytical and sensory characteristics of wines. In this regard, growing interest on the use of controlled mixed fermentation with selected non-Saccharomyces and S. cerevisiae wine yeasts draw the applied research in oenological field.
2 Yeasts on Grapes
Grapes represent a complex ecological niche where filamentous fungi, yeasts and bacteria cohabit. The microbial community colonizing this ecological niche includes microbial species whose concentration depending on multiple factors; the most important are related to grape ripening and nutrients availability. Actually, the microbial ecology of grape berry is a wide concept including closed relations between the ecosystems and their microbial interactions, microbial vectors and sources of microorganisms. Herman Phaff, the pioneer of yeast ecology, described the concept of ecology as “where microbes live and why they live in one habitat and how yeasts interact with other microorganisms” (Lachance 2003). This comprehensive approach implies that microbial communities may be affected by many other variables in grapes, such as viticultural practices, pedoclimatic factors, diseases and pests that could modify grape integrity. In general, the yeast populations of mature grapes are comprised of between 103 and 105 cells/g (Fleet et al. 2002), but approximately one log higher values have often been found on damaged berries in presence of higher availability of sugar and nutrients (Barata et al. 2008). Over the last century many researchers have described the occurrence and association of yeasts with grape surface and the results were reviewed by Amerine and Kunkee 1968; Kunkee and Goswell 1977; Kunkee and Bisson 1993. More recently, the yeast ecology of wine grapes was reviewed by Fleet et al. 2002, Barata et al. 2012 and Jolly et al. 2014 evaluating the factors that affect their occurrence and quantitative presence.
2.1 Occurrence and Diversity of Yeasts
The composition, in terms of occurrence and amount, of indigenous microbiota naturally present on grape berry surfaces is crucial during winemaking process, as it can positively or negatively affect the quality of final wine. The presence and fitness of yeasts are essential in alcoholic fermentation, as promoters of transformation of grape sugars into principal products of fermentations: ethanol, carbon dioxide and hundreds of other metabolites responsible for aroma and flavours (Romano et al. 2003; Fleet 2003).
Kurtzman et al. (2011) already several years ago, ascribed overall yeasts potentially associated with grape/wine ecosystem in 15 different yeast genera, such as Dekkera/Brettanomyces, Candida, Cryptococcus , Debaryomyces, Hanseniaspora/Kloeckera, Kluyveromyces, Metschnikowia, Pichia, Rhodotorula, Saccharomyces, Saccharomycodes, Schizosaccharomyces and Zygosaccharomyces. On the other hand, the dynamic yeast taxonomy poses challenge on the nomenclature of wine microbiology (Bisson et al. 2017). The yeast Hanseniaspora and its anamorph counterpart Kloeckera are the numerically predominant genera present on the surface of grape, with more than 50% of the total yeast population (Fleet and Heard 1993). To a lesser extent, species belonging to Candida, Starmerella, Cryptococcus , Pichia, Metschnikowia and Kluyveromyces (Lachancea) genera are detected (Heard and Fleet 1988; Mills et al. 2002; Rosini et al. 1982). However, the variability may be reduced to few groups of similar physiological characteristics. For instance, the ubiquitous Candida spp. and Pichia spp. are highly heterogeneous, and new species are likely to be found in each new survey because the accuracy of molecular identifications is constantly increasing. A division of yeast biota of grape berries into three main groups with similar characteristics are proposed: (i) oxidative yeasts as basidiomycetous Rhodotorula and Cryptococcus along with the yeast-like fungus Aerobasidium pullulans and some Candida species; (ii) oxidative- fermentative ascomycetes Hanseniaspora spp., Pichia spp., and Metschnikowia spp. together with some Candida species; (iii) strongly fermentative yeasts with higher alcohol producing Saccharomyces spp., Starmerella spp. Torulaspora spp., Zygosaccharomyces spp., and Lachancea spp. In Table 1.1 are summarized the main yeast species colonizing wine making environment.
Oxidative Yeasts
Relatively to oxidative yeasts, these are present on the surface of the berries still before this reach maturity when there is a high sugar content and can be found in the early stages of fermentation. In the middle and last phase of grape ripeness, the oxidative yeasts decrease in concurrently to the detriment of nutrient availability due to the competition with other yeast species, but they are still widely present at harvest time depending on the agronomical practices (Fleet et al. 2002; Hernández et al. 2018).
Oxidative-Fermenting Yeasts
Hanseniaspora/Kloeckera species are the most abundant ascomycetes yeasts colonizing the grape surface of grape berries at harvest time. Regardless of the geographic distribution of winemaking areas, the presence and colonization of the yeasts Hanseniaspora / Kloeckera on grape surface is everywhere dominant over the other yeast species. Within the apiculate yeasts the species Hanseniaspora uvarum (Kloeckera apiculata) are the most frequent but other species such as Hanseniaspora hosmophila or Hanseniaspora guilliermondii can be found at lower concentration (Giorello et al. 2018). Other ascomycetes widely found at harvest time on grape surfaces are species belonging to Pichia, Candida and Metschnikowia genera. In this regard, several species have been described. Among the species described within Pichia genera, Pichia membranifaciens , Pichia fermentans , Pichia kluyvery and Pichia kudriadzvii (synonimum Issatchenkia orientalis ) are the most widely isolated (del Monaco et al. 2014). In the Candida genus several fermenting and non-fermenting species were isolated from grapes. The most diffused fermenting species is Candida stellata that it was successively reclassified as Candida zemplinina and more recently enclosed in clade of Starmerella as Starmerella bacillaris (Duarte et al. 2012)
Within Metschnikowia genera, new species Metschnikowia viticola was recently isolated, studied and characterized from a Hungarian vineyard. From a genetic point of view M. viticola is well disconnected species within the genus Metschnikowia. However, very little is known about the ecological distribution of M. viticola and their frequency on grape berries (Peter et al. 2005; Brysch-Herzberg and Seidel 2015). Other many new species have recently been described in the Metschnikowia genus, including Metschnikowia chrysoperlae (Suh et al. 2004), Metschnikowia fructicola (Kurtzman and Droby 2001) and Metschnikowia andauensis (Molnar and Prillinger 2005). In these cases, there was a real difficult in the delimitation among new species and the well characterized Metschnikowia pulcherrima The experimental results obtained by Sipiczki et al. (2013) explain that the type strains of M. andauensis and M. fructicola possess divergent copies of the rDNA gene will lead to further investigations of the species concept in the clade. This support the importance of unambiguous yeast identification in any study of the yeast diversity in grape habitat.
Strongly Fermentative Yeasts
Regarding to the fermentative, higher alcohol tolerant yeasts, their colonization is related to the high nutrient availability resulting from grape damage that possess, besides much higher cell counts, wider species diversity than sound grapes (Barata et al. 2012). S. bacillaris may be present in higher numbers but its relative proportion also decreases in favour of higher fermentative yeasts such as Zygosaccharomyces spp., Lachancea spp. and Torulaspora spp., which, as mentioned above, may occasionally dominate the overall microbiota.
2.2 Factors Affecting Yeast Community
The composition and complexity of microbiota of grape berries depend on the interactions between individuals. The resulting consortium is generally stable over time and depending on several biotic and abiotic factors (Fig. 1.1). Relative to abiotic factors, the climatic and microclimatic conditions, including the effect of temperature, UV exposure, rainfall, sunlight and winds, can influence microbial populations.
Among biotic factors, microbial vectors, such as bees and wasps, can actively transfer yeasts on the grape surfaces (Francesca et al. 2012; Goddard et al. 2010; Stefanini et al. 2012). The microbial habitat associated with birds represents the object of several ecological surveys not only in applied food microbiology. Indeed, associations between wild birds and microorganisms have been studied mainly focusing on bacteria, whereas limited studies on yeasts are available (Cafarchia et al. 2006). The monitoring of bird movements allows the investigation about behavioural and demographical responses to a given environment (Riffell et al. 2006). The migration of birds includes a round trip to the resting areas and a return to the territories of nesting, which occur in autumn and spring respectively. These movements follow the seasonality of food resources. Since birds act as microorganism vectors, the analysis of the microflora they host may be important to evaluate the microbial diversity of the sites visited. From an applicative perspective, yeasts carried out by birds have not been deeply investigated. Nowadays, there is a growing interest of wine producers to perform winemaking employing ‘autochthonous’ strains which may ensure typical terroir characteristics (Capozzi et al., 2015). At this regard, it was recently reported the dissemination of oenological yeasts by vineyard inhabiting birds, mainly Black birds (Turdus merula), although no yeast with technological relevant traits was found in the few migratory birds analysed (Francesca et al. 2012). Those authors evidenced an issue related to the autochthonous status of yeasts, since they may not be indigenous in each environment. Yeasts may be moved at different distances depending on the vector type. Some studies provided evidences that insects such as honey bees disseminate S. cerevisiae strains until approximately 10 Km (Goddard et al. 2010), so that the investigation of migratory birds need clarifications for the associated movements of yeasts with the support of technological relevance. During migration, several sites are visited by birds because they represent important stop-over points. For example, during migration from Africa to Europe and vice versa, Lampedusa and Ustica islands are visited in spring when the direction is from sub-Saharan areas to North Europe, while Linosa is visited in autumn during the opposite fly.
It is well known that climate change is partly responsible for the elevated sugar concentration and lower acidity in grape berry and then in must (Godden et al. 2015; Neumann and Matzarakis 2014; Petrie and Sadras 2008; Teslić et al. 2018). This is strictly related to the microbial composition in grape berries. Vine sensitivity to weather properties (Holland and Smit 2014), narrow spatial surfaces suitable for producing high-quality grapes as wine industry raw material, and possibility of perennial plant exploitation (Lereboullet et al. 2014) are indicative of the need for a climate change assessment associated with winemaking. Despite the importance of the global climate change trend, from the vine grower/winemaker perspective, it is more essential to understand regional atmospheric conditions (Orlandini et al. 2009) and local microclimatic environment as well. Generally, increasing average global temperature over the last few decades is more than evident, as is the increasing temperature trend, although is not homogenous in every vine-growing region (Pielke et al. 2002; Van Leeuwen et al. 2013). For example, a significant growing season temperature trend for the majority of northern hemisphere wine-producing regions between 1950 and 1999, with an average increase of 1.26 °C, was demonstrated. However, there was also an insignificant trend in the majority of southern hemisphere wine regions, which emphasizes the necessity to focus study on smaller study areas.
Since climate modifications are vastly complex, examinations of simple temperature and precipitation values are insufficient to explain climate change trends. Therefore, several bioclimatic indices, such as Huglin index (Huglin 1978), Cool night index (Tonietto 1999), Winkler or growing degree day index (Winkler 1962), number of days with maximum temperatures higher than 30 °C (ND > 30 °C) (Ramos et al. 2008), number of days with precipitations <1 mm (Dry spell index, DSI) (Moisselin and Dubuisson 2006), etc. are commonly used in viticulture to provide an improved insight into climate change tendencies. However, the selected bioclimatic indices were mainly based only on-air temperature, as it has the strongest influence on overall growth, productivity, and berry ripening of the grapevine (Jones-Vaid et al. 2012).
Another important parameter influencing the grape microbiota is related to the water intended as rainfall. Indeed, moderate water stress may positively affect berry sugar accumulation during grape-growing season (Coombe et al. 1989), while increasing temperature advances phenological stages and speeds up sugar accumulation in grape berries (Jones-Vaid et al. 2012; Bonnefoy et al. 2013). The association of water stress together with increasing temperature later lead to the production of wines with higher alcohol content and other microbiological, technological, sensorial, and financial implications (Mira de Orduña et al. 2000). As direct consequence, increase of grape sugar content at harvest may cause slow or stuck alcoholic fermentations during hot years as well as alter sensory features due to the ethanol’s tendency to increase bitterness perception (Sokolowsky and Fischer 2012), suppress the perception of sourness, and reduce astringency perception.
Concerning the total yeast counts, Combina et al. (2005) found that rainy years increased yeast presence. This climatic condition probably increases the berry volume and permits the release of juice in joint areas, such as the part between the pedicel and the berry, and higher exosmosis leads to nutrients on the grape surface. With careful and sound berry sampling, Čadež et al. (2010) found that colder harvests with higher rainfall lead to increased yeast counts. In contrast, Comitini and Ciani (2006) found ten-fold less total counts in years with high rainfall. In addition, the geographic location, grape variety and vineyard age and size can influence the composition and occurrence of microflora that are present on the surface of grape berries.
Another important aspect is related to vineyard chemical treatments. A lot of studies showed that agronomical practices, such as organic or biodynamic management can modify the microbiota of grape and must (Cordero-Bueso et al. 2011; Milanovic et al. 2013; Pretorius 2000; Mezzasalma et al. 2017). Some authors suggested that the occurrence of specific bacteria in must and wine influences wine characteristics and typicity (Belda et al. 2017a; Liu et al. 2017).
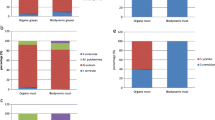
The main vineyard treatment studied is related with the use of pesticide treatments, mainly those against fungi (downy mildew, powdery mildew and grey rot). The studies are either based on analysing grapes after vineyard treatment, which do not exclude the influence of other factors, or from auto-enrichment fermentations which cannot be correctly extrapolated to evaluate the variations on berry microbiota. Conventional pesticides can produce a decrement in the yeast population and diversity in fermenting musts. Ganga and Martínez (2004) detected less diversity of non-Saccharomyces species, which was explained using fungicides against Botritys cinerea. Differently, there are discordant results on the effect of chemical treatments on S. cerevisiae presence on grapes. Ganga and Martínez (2004) did not find reduced S. cerevisiae occurrence after fungicide use while other investigations recovered lower numbers of this species (Regueiro et al. 1993; Van der Westhuizen et al. 2000). It is quite evident that the influence of chemical pesticides on microbiota of grape berry is related to other factors, such as climatic conditions or grape variety, which cannot be correctly extrapolated to evaluate the single effect on berry microbiota. About this concern, Ganga and Martínez (2004) detected less diversity of non-Saccharomyces species, which was explained using fungicides against B. cinerea, while Regueiro et al. (1993) and recovered lower numbers of these species. Milanovic et al. (2013) found that Candida zemplinina (synonimus S. bacillaris) and Hanseniaspora species colonised surface of grapes treated with both organic and conventional treatment, while M. pulcherrima was widely found in conventional samples and only occasionally in organic grapes.
A specific influence of grape varieties on indigenous yeast community of grape berries was found. Clavijo and collegues (2010) carried out an ecological survey of wine yeasts present on grapes growing in two vineyards located in the southern Spain (Serranía de Ronda region). They found that, although Kluyveromyces (Lachancea) thermotolerans , H. guilliermondii , H. uvarum and Issatchenkia orientalis (Pichia kudriavzevii ) are the most frequent species, a specific distribution of strains was found in the three grape varieties studied. The influence of grape varieties on the indigenous yeast community of grape berries was also evaluated by Raspor et al. (2006) The frequency of occurrence of yeast species showed their preferences for certain grape varieties. The white grape variety mostly attracted pigmented Basidiomycetous yeasts belonging to the genera Rhodotorula, Sporobolomyces and Cryptococcus that dominated on all sampling locations. Differently, yeast populations isolated from the red grape surfaces belonged both to Ascomycetous and Basidiomycetous yeasts in the ratio of 1: 1.
In the last 10 years, due to the advances in metagenomics, it has become clearer and clearer that in general, plants host a wide array of bacteria and yeasts most of which are not cultivable and therefore are almost unknown at the taxonomic and metabolic levels. Such microorganisms interact with the plant organs and can influence plant nutrition, development, productivity, and stress responses (Bacon and White 2016).
Another important question regards the influence of grapevine cultivar on the grape microbiota. Recently, it was shown that some epiphytic bacteria were shared by aerial plant portions and the soil (Martins et al. 2013). This finding led them to propose that the physical proximity between soil and the plant might facilitate microbial migration through rain splash, winds, pollinators and other foragers, and parasites.
Moreover, any grapevine cultivar shows peculiar secondary metabolites, and most of these are concentrated in the fruit. Some of these metabolites have antimicrobial properties (Chong et al. 2009; Katalinić et al. 2010) and could influence the composition of grape microbiota both quantitatively and qualitatively. Based on these assumptions, it was hypothesized that each cultivar could have an active and specific role in the interaction with and selection of its microbial community (Mezzasalma et al. 2017).
2.3 Recent Methodologies for Detecting the Presence of Yeasts on the Grape Berry
To know the microbial composition in grape barriers and to further monitor their evolution during wine fermentation understanding the relationship between the microorganisms is of relevant importance in applied studies (Bokulich et al. 2014; Piao et al. 2015; Stefanini and Cavalieri 2018). The use of conventional methods including culture-dependent techniques, allow to analyze culturable fungi, yeasts, acetic acid- and lactic acid-bacteria associated with grape berries and wine. As well as in many other natural habitats, there are several viable but non-culturable wine microorganisms, that could not be studied under conventional laboratory microbial conditions, leaving an incomplete knowledge about the occurrence and dynamics of the microbial community involved in winemaking (Cocolin et al. 2013; Piao et al. 2015). Recent advances in sequencing technologies based on culture-independent techniques allow to capture a large proportion of cells (culturable, non-culturable and slight represented species) finding a complete microbial ecology picture (Bokulich et al. 2013; De Filippis et al. 2013; Abdelfattah et al. 2016). The beginning of massively parallel, high-throughput sequencing approach (sometimes referred to as next-generation sequencing) represent a revolution in applied microbial ecology research. Several platforms and chemistries exist, such as Illumina, 454/pyrosequencing, ion semiconductor, and nanopore sequencing, also if all employ nanotechnology to tether individual strands of DNA and detect the incorporation of individual nucleotides into each strand during polymerization events. Each system has its strengths and weaknesses, including different sequence read lengths, number of strands sequenced, and error rates – but each has been a stepping stone in advancing the ability to investigate the inner workings of the microbial community. This approach is appropriate for the food sciences, bringing manifold improvements over earlier mixed-microbial detection techniques (Bokulich and Mills 2012).
These new sequencing strategies rely on the analysis of a single core molecule DNA (and by transcription RNA) yet possess many applications for microbial ecology analysis. The first is amplicon sequencing whereby marker-genes are amplified from mixed genomic DNA by PCR, sequenced directly, and aligned against a reference dataset to identify the taxonomic composition of whole microbial communities. This same process can also be applied to RNA, by reverse-transcribed to cDNA, to profile the actively transcribing community within a sample. The taxonomic information provided by amplicon sequencing is frequently lower-resolution than that delivered by metagenome sequencing (which enables reconstruction of full-length marker genes) but is substantially higher throughput, facilitating exploration of massive numbers of unique microbial communities. With the availability of new metagenomic approaches the monitoring and composition of microbial populations can be better and faster described. In this regard, the relation between complexity of microbial community and geographical wine producing area represent a very interesting current topic. Using the metagenomic approaches, several studies showed the variation of the microbial community of grapes in function of regional distribution, (Gilbert et al. 2014b; Taylor et al. 2014; Morrison-Whittle and Goddard 2015; Pinto et al. 2015; Belda et al. 2017b). Moreover, the correlation among microbial complexity and organoleptic characteristics of wine was studied (Knight et al. 2015; Bokulich et al. 2016). An unambiguously explanation to the diversity of microbial communities among geographic locations is not currently known. In addition, recent studies showed that microbial populations found in musts may originate also from the environment surrounding the vineyard (Morrison-Whittle and Goddard 2018). Because of the observation of a putative microbial “terroir”, the role and persistence of environmental microbial species in the wine fermentative process gained a renewed interest.
In this regard, from the application point of view, studies on indigenous yeasts strongly adapted to specific grape musts are growing, both to study the biodiversity associated to different geographic area and to select new indigenous strains associated with “terroir” (Capozzi et al. 2016; Zarraonaindia et al. 2015). These new concepts of microbial colonization and effectiveness showed that microbiomes associated with grapes and with the earlier stage of fermentation are biogeographically defined, illustrating that different regional wine profiles are related with specific microbial communities.
3 Yeast in Winery
In addition to natural habitats such as woods and agricultural areas near the vineyard, vineyard soil, vines and grapes, a relevant and consistent yeast community have found niches in man-made environments such as wine cellars. During the vinification process, grape juice and wine encounter a large area of equipment surfaces within wineries which may serve as important reservoirs of microorganisms that influence and contribute the final composition of the wines. For these reasons the surfaces of winery equipments become locations for the developments of so-called residential or winery microflora (Peynaud and Domercq 1959; Pretorius et al. 1999; Rosini 1984).
3.1 Diversity of Yeasts in Winery Environment
The role of winery environments in shaping the microbiota of wine fermentations and vectoring wine spoilage organisms is poorly understood at the systems level. Indeed, although the presence and importance of winery yeasts have been known or surmised for a long-time year (Peynaud and Domercq 1959) their actual contribution to must fermentations has been poor investigated and somewhat ignored (Pretorius et al. 1999). However there are several factors that potentially determine a stable colonization of yeasts in this anthropized environment (Fig. 1.2).
Winery equipment, including crush/press equipment, valves, collectors and barrels, for its difficult to clean, become useful for microbial adhesion and biofilm production and consequent potential sources of contamination. In this regard, one of the most important features that characterize the winery microbiota is the survival and the modality of colonization over the course of harvest campaign (before, during and after grape harvest). Therefore, to track the occurrence of equipment microbiota and evaluate the fluctuation of yeast population, samplings before, during, and after grape harvest are an important aspect to be investigated.
The pre-harvest yeast communities represent the resting state of the winery and play an important role since these is the first population encountered by fresh grape juice prior to fermentation. The composition of this microbial community may play an important role and can impact on wine fermentation qualities downstream.
Most of the studies on the occurrence and yeasts colonization of cellar were carried out on S. cerevisiae the main agent involved in alcoholic fermentation. Indeed, colonization of winery surfaces by Saccharomyces has been widely reported and it is probably an important source of this yeast in wine fermentations, particularly in non-inoculated wines (Bokulich et al. 2013). Studies, investigating on the yeast biota of winery, showed the constant presence of S. cerevisiae and had identified this species as dominant on winery surfaces at pre-harvest time (Ciani and Rosini 1986; Ocón et al. 2010). Indeed, the winery environment is colonized by many cells of S. cerevisiae, which go through innumerable generations during fermentation for each vintage (Rosini 1984; Ciani and Rosini 1990; Ocón et al. 2010). Here may exist a significant selective pressure on the S. cerevisiae population of the winery by factors such as ethanol and SO2 (Cocolin et al. 2004). However, even if S. cerevisiae is the most abundant species in winery environments may account only 30–40% of the total yeast population, other non-Saccharomyces yeast species may colonize the winery surfaces and equipments depending on the spatial variation in the winery surface. Indeed, some crush equipment (hopper, elevator, crusher, and press samples) that entering almost exclusively in contact with the grapes are colonized by yeast-like (e.g., Aureobasidium pullulans) and yeast genera that colonize the grape surfaces such as Hanseniaspora, Candida or Metschnikowia (Bokulich et al. 2013). The occurrence and persistence of non-Saccharomyces in the cellar environment was well documented (Ciani and Rosini 1986; Ocón et al. 2010). A more recently work, using identification methods at the strain level, found a large number of isolates belong to S. bacillaris , H. guilliermondii and H. uvarum demonstrating the persistence of non-Saccharomyces yeast strains from year to year in the cellar. Indeed, some strains of these three non-Saccharomyces species were found in the must for two consecutive years and found also in cellar environment before the second harvest indicating the persistence of these yeasts in this environment (Grangeteau et al. 2016). In fermentation equipment samples (fermentors, hoses, filters, and pumps) that all deal strictly with fermenting and fermented wine, S. cerevisiae is largely present, together with other fermenting yeasts such as of Pichia kudiavzevii (synonimum Issatchenkia orientalis ), Torulopsis bacillaris (synonymus of Candida zemplinina ) and Pichia spp. These non-Saccharomyces yeasts can explain for 60–70% of total yeast biota which colonizes the winery surfaces and their role in this context has been little investigated (Ciani and Rosini 1986; Ocón et al. 2010; Sabate et al. 2002; Bokulich et al. 2013). Some oenological practices such as cold maceration prior to fermentation may affect the yeast ecology during wine fermentation in favour of non-Saccharomyces species. Hierro et al. (2005) found that cold maceration favoured the presence of H. osmophila, Candida tropicalis and Zygosaccharomyces bisporus , the only species isolated from the unripe and overripe fermentations after cold maceration.
Another important feature of winery biota that involves ecological and technological aspect is the flux of specific S. cerevisiae strain from winery surface to must fermentation and vice versa.
A series of studies have found an effective flow of strains of S. cerevisiae from the cellar equipments and surfaces to fermentation musts. Rosini in 1984 in a new pilot scale winery demonstrated the occurrence of a flow of S. cerevisiae cells from the winery surfaces to freshly pressed grape musts, and vice versa. These results were then confirmed by Costantì et al. (1997) that found a competition of resident winery yeasts and pure S. cerevisiae starter cultures. A 6 year follow-up study carried out in a new built winery showed that indigenous winery resident S. cerevisiae strains competed with commercial strains inoculated in other fermentation tanks of the cellar (Beltran et al. 2002). The contribution of winery-resident S. cerevisiae strains to spontaneous grape must fermentation was shown under real vinification conditions. The S. cerevisiae strains colonizing the winery surfaces were the ones that conducted the natural must fermentation (Ciani et al. 2004). Other investigations found that specific Saccharomyces strains become established on winery surfaces, resulted in repeatable detection over multiple years in uninoculated wines (Santamarıa et al. 2008; Blanco et al. 2011; Ciani et al. 2004).These results support the role of winery as a man-made niche of S. cerevisiae and a possible reproducibility, as well as regionality, of wine sensory characteristics produced at a given winery. In this regard the selectivity of the winery environment (winery effect) may have a selective pressure towards some enological characters as maximum ethanol production (ethanol resistance) fermentation rate and SO2 resistance of S. cerevisiae population (Cocolin et al. 2004). On the other hand, the role of S. cerevisiae of winery environment on the specificity of wine sensory profile at regional level remains unclear.
3.2 Factors Affecting Yeast Community in Winery
The extended of the development of a residential microflora will depends by several factors such as nature of the surfaces (irregular, unpolished surfaces, cracks and welds) and cleaning and sanitization procedures and possible biocontrol procedures (Fig. 1.2). The nature of the surfaces may strongly influence the colonization of yeast species and determines their persistence from one harvest to another.
On the other hand, the modalities of cleaning and sanitization procedures also influence the quantitative presence and relative abundance of the different yeasts species. However, several works reported that also in well cleaned wineries, the widely presence of microorganisms and specifically of yeast biota, was found. However, under normal correct procedures of cleaning and sanitization the presence of spoilage-related microorganisms (e.g. Brettanomyces spp.) was undetected or detected at very low levels (Bokulich et al., 2013; Ocòn et al., 2013).
Classical studies on spoilage yeasts by Van der Walt and van der Kerken (1961) on Brettanomyces spp., Rankine and Pilone (1973) and on Zygosaccharomyces bailii , Peynaud and Domercq (1955) on Saccharomycodes ludwigii have demonstrated that they may be winery contaminants, even if most results from literature suggest that their prevalence is low. Chatonnet et al. (1992) were the first authors to identify oak barrels as an ecological niche for Dekkera/Brettanomyces spp., which become more dangerous with repeated use. This suggests that barrel sanitation and sulfite utilization (sulfur burning in empty barrels) is not enough to eliminate Dekkera/ Brettanomyces spp., which develop during the lifetime of the barrel. Connell et al. (2002) also recovered Dekkera bruxellensis from air samples of crush, tank, barrel, and bottling line areas using BSM medium (Millipore) followed by a filter-based chemiluminescent in situ hybridization technique. However, the primary source of these yeasts remains obscure. A recent study investigating on the occurrence of Brettanomyces bruxellensis found a flux of isolates form grapes to winery (Comitini et al. 2019). Currently, some of the procedures that applied to limit the risks of Brettanomyces/Dekkera colonization in wineries and wines are not particularly appropriate for use during wine ageing. This has led to increased interest in the exploration of yeasts that can counteract the activities of these undesired microorganisms in wine (Comitini et al. 2004). Investigations on biocontrol topic, relative to killer yeasts as producers of mycocins that can neutralize the activities of undesired microorganisms in wines represent a valid strategy for the control of these undesired yeasts (Druvefors et al. 2005).
4 Alcoholic Fermentation
Wine fermentation is typically carried out by a complex evolution of microorganisms involving both yeasts and bacteria. During the years, a lot of studies extensively examined the succession of yeasts that occurs during the spontaneous fermentation in must as non-sterile source. Now, it is well established that together with S. cerevisiae, non-Saccharomyces species actively participate during the alcoholic fermentation. In the past these non-Saccharomyces wine yeasts were negatively considered because of reduced fermentation power, high production of undesired products that affects the aromatic profile of wines. For these reasons, the use of selected S. cerevisiae as starter culture was a common and widely diffused winemaking practice to control the fermentation process and give the desired characteristics to the wines. More recently, several studies have been revaluated the role of non-Saccharomyces yeasts during alcoholic fermentation and their metabolic impact on the analytical sensory characteristics of white and red wines (Benito et al., 2014). In this regard, there are a growing interest on the use of controlled mixed fermentation with selected non-Saccharomyces wine yeasts tin co-culture or sequential inoculation.
4.1 Spontaneous Fermentation
Grape bunches, the primary substrate of winemaking process, are perhaps the most obvious potential source of microbial diversity of spontaneous grape juice fermentation. However, the winery, as previously indicated as a man-made ecological niche, may play also an important role in spontaneous grape juice fermentation particularly regarding to the fermenting yeasts. Indeed, a serial of ecological surveys of the yeast flora associated with spontaneous fermentation of grape juice in almost all the geographical winemaking areas revealed a sequential occupation of the substrate: initially apiculate yeasts (Hanseniaspora, Kloeckera) take over, after 3–4 days they are replaced by S. cerevisiae (Martini 1993; Pretorius 2000). While the first one yeasts are abundant on the grape surfaces at harvesting time, S. cerevisiae (Saccharomyces uvarum ) species colonize the winery surface where resulted the most widely diffused species during the different stages of winemaking (before during and after the fermentation). With ethanol increasing S. cerevisiae the higher resistant to alcohol is the first explanation to this substitution but other contributing factors may be involved.
In this sequential occupation of the grape juice by apiculate-elliptical yeasts, during the various stages of fermentation it is possible to isolate other yeast genera, such as Starmerella, Candida, Pichia, Zygosaccharomyces, Schizosaccharomyces, Torulaspora, Lachancea (Kluyveromyces) and Metschnikowia (Fleet et al. 1984; Pardo and Serrano 1989; Belda et al. 2015, del Carmen Portillo and Mas 2016; Garofalo et al. 2016). The indigenous non-Saccharomyces yeasts, are present in the grape juice in high numbers in active growth state, which gives them a competitive edge (Cray et al. 2013).
The growth of non-Saccharomyces species belonging to the genera Kloeckera/Hanseniaspora, Starmerella and Metschnikowia is generally limited to the first few days of fermentation, because of their weak ethanol tolerance and less efficient fermentation. Other more ethanol tolerant fermenting yeasts such as Torulaspora delbrueckii , Lachancea theromotolerans and Zygosaccharomyces spp. are generally present less frequently but occasionally they were found at higher concentration. (Clavijo et al. 2010; Zott et al. 2008; Jolly et al. 2003; Garofalo et al. 2016).
Therefore, spontaneous fermentation contains a mixture of yeast species but, as the fermentation progresses, the environment becomes more selective and the dominance of S. cerevisiae is expected and desired. However, S. cerevisiae populations showed a genetic variability with the presence of more than one strain. Indeed, several studies carried out in various winemaking areas showed that different strains of S. cerevisiae are involved during the fermentation process. In this regard, some S. cerevisiae strains are present at high relative amount and were able to dominate the alcoholic fermentation and are denominated “dominant” or “predominant” strains while other strains occur at lower relative amounts and are defined “secondary” strains. The association of the dominant S. cerevisiae strains in spontaneous fermentation with the environment is not well defined and it was linked both winery (Frezier and Dubourdieu 1992; Guillamón et al. 1996; Ganucci et al. 2018) or grape variety (Blanco et al. 2006; Schuller et al. 2012).
As reported above, in a spontaneous fermentation several yeast species and strains coexist interacting with each other and environment factors. The specific fitness of yeast species and the evolution occurring in grape must during alcoholic fermentation toward more selective conditions determined the sequential occupation of the substrate and the progressive dominant presence of S. cerevisiae. Similarly, within S. cerevisiae strains the dominant strains were selected on the bases of its fitness toward the specific environmental factors. Currently, there are two lines of research that investigate the origin and occurrence of S. cerevisiae dominating spontaneous fermentation and that characterize the analytical sensory characteristics of the wine of a given territory or winery.
As previously reported, several recent investigations (Bokulich et al. 2014; Gilbert et al. 2014a; Taylor et al. 2014; Morrison-Whittle and Goddard 2015; Pinto et al. 2015; Belda et al. 2017a, b), showed evidences for a relation between microbial community of grapes and geographic distribution and organoleptic characteristics of fermenting musts (Knight et al. 2015; Bokulich et al. 2016). This adaptation, however, could be due to the selective pressure performed during the winemaking process in the winery. Some factors such as ethanol, temperature, SO2 and others can play a fundamental role in the yeast species and strains selection during spontaneous fermentation and determining the dominant S. cerevisiae strains that for these reasons, colonize the man-made winery environment.
4.2 Factors Affecting the Occurrence and Succession of Yeast During Spontaneous Fermentation
There are two principal factors determining the evolution of yeast community during the spontaneous fermentation: (i) the quantitative occupation of the substrate by yeast species; (ii) the progressive increase of ethanol concentration. These features play a key role on the yeast species succession in spontaneous grape juice fermentation. However, the dynamics among the yeast species present during fermentation are more complex and strongly influenced by their interactions and other environmental factors (Ciani and Comitini 2015).
Temperature of grape juice fermentation is one of the most influencing factors on yeast species dynamics. Indeed, the presence and permanence of non-Saccharomyces yeast species during fermentation is affected by temperature. Indeed low temperatures (10–15 °C). increased tolerance to ethanol of K. apiculata and C. stellata (formerly S. bacillaris ) (Erten 2002; Gao and Fleet 1988: Ciani et al. 2006; Mendoza et al. 2007; Ciani et al. 2010). Such increase in ethanol tolerance of non-Saccharomyces yeasts at low temperatures appear to be the major factor that affects their stronger contribution in this condition.
Another factor that regulates the presence and occurrence of yeast species during spontaneous fermentation is the availability of oxygen. Reduced oxygen availability during fermentation may have a key role in yeast-yeast interactions. Indeed, the limited availability of oxygen could explain in part the reduced competitiveness showed by K. thermotolerans and T. delbrueckii toward S. cerevisiae (Hansen et al. 2001).
In the yeast species interactions during wine fermentation cell-to cell contact appears to be also involved. Indeed, in the presence of high concentrations of viable cells of S. cerevisiae, the growth of T. delbrueckii and K. thermotolerans is inhibited (Nissen and Arneborg 2003; Nissen et al. 2003; Arneborg et al. 2005).
The quorum-sensing-like phenomena could also be involved in some yeast interactions during spontaneous wine fermentation and the identification of active molecules and their influence on gene expression of yeast co-culture deserves to be investigated. In this regard, recent investigations on the putative quorum-sensing molecules as 2-phenylethanol, tryptophol, and tyrosol have begun to elucidate the mechanisms and role of quorum sensing in yeast under winemaking condition (high cell density or under low nutrient conditions) (Zupan et al. 2013; Williams et al. 2015). In addition to ethanol, acetic acid, medium chain fatty acids, acetaldehyde and the synergistic action of their combinations, could play an important role on the inhibitory mechanism that can occur in wine fermentation (Bisson 1999; Fleet 2003).
The production of toxic compounds from S. cerevisiae has also been hypothesized as a cause of the early death of H. guilliermondi in mixed fermentations (Pérez-Nevado et al. 2006). Indeed, several compounds produced by yeasts during must fermentation may become inhibitory to other yeast species or strains. Between them, secretion of killer toxins by specific yeasts represents an efficient tool to eliminate competitors without direct cell-to-cell contact. Yeasts with killer phenotype secrete protein or glycoprotein that generally kill sensitive cells in a two-step receptor-mediated manner: First, there is a specific bind to primary cell wall receptors; secondary the killer toxin translocate to the plasma membrane where they interact with secondary receptors or enter susceptible cells, thus exerting their cytocidal effect through different mode of actions. In addition to killer proteins other proteinacenous compounds have been found in the yeast–yeast and yeast–bacteria interactions in wine fermentations.
Indeed, it was found that certain S. cerevisiae strains produce proteinaceous compounds that are active against malolactic bacteria (Comitini et al. 2005; Osborne and Edwards 2007), peptides with molecular mass less than 10 kDa, that inhibit the growth of some non-Saccharomyces such as H. guilliermondii , T. delbrueckii , K. marxianus and K. thermotolerans (Albergaria et al. 2010; Peña and Ganga 2018). However, also if the identity of these antimicrobial peptides remained elusive and need some deepness, the possibility of using them as natural biopreservatives in alcoholic fermentations could be an interesting alternative for the microbial control of winemaking process.
4.3 S. cerevisiae Inoculated Fermentation
From the ecology surveys carried out in different winemaking environments S. cerevisiae is a minority species and it is difficult to isolate from vineyard soil or the surface of ripe grapes, while it is the dominant yeast species of winery and its equipment (Martini 1993). Indeed, S. cerevisiae is generally found in association with the production of alcoholic beverages and for this reason it is defined a “domesticated” species, strongly specialized for fermenting high sugars substrates. For their fermenting features and oenological aptitude, S. cerevisiae is the species that conduct and determine the rightness of the fermentation process characterizing the chemical and sensory profile of wine. However, for the long time the fermentation of grape juice was carried out without yeast starter strain inoculation and spontaneous must fermentation occurred. After 1960 scientific and technological improvements allowed the diffusion of active dry yeasts commercial preparation belonging to S. cerevisiae. The diffusion of commercial starters in active dry form was one of the most significant technological advances in winemaking. As direct result, the quality and quantity of wine production were highly improved, as the winemaking process was controlled and safe (Heard and Fleet 1985; Henick-Kling et al. 1998). The introduction in winemaking process of selected and efficient strains announced the concept of innovations revolutionizing the wine industry and market. Nowadays, the large-scale wine production, where rapid and reliable fermentations are essential for wine flavour and predictable quality, the practice of the inoculation of selected pure starter strains of known ability is a common practice. However, the current challenge of applied research in biotechnology is the producing new yeast strains with even more reliable performance, reducing processing inputs, and facilitating the production of peculiar and high-quality wines (Pretorius 2000). The forces of market and technology continue to challenge the tension between tradition and innovation. On the one hand, it is evident the tendency to use commercial strains that guarantee controlled processes, from another hand it is still recognized the potentially of native yeasts to obtain distinctive features. Indeed, despite the immense wealth of natural yeast diversity, the extremely selective and specific conditions of industrial fermentations sometimes require a combination of phenotypic traits that might not be commonly encountered in nature. In this picture, a lot studies focused on the isolation, manipulation and develop of novel S. cerevisiae strains tailored for a specific wine product that bring greater complexity to wine than strains currently available to the industry (Bellon et al. 2013). The most intuitive way to generate artificial diversity in yeasts is based on genetic manipulation, to artificially increase the already existing yeast diversity and generate variants that may perform better in industrial settings than the strains that are selected in natural environments. A specific approach is the genetic engineering reshuffle in selected strains, applied to modify single genes by introducing, disrupting or modulating enzymatic key-steps of metabolic networks (Santos and Stephanopoulos 2008). However, wine yeasts show complex and continuous variation for most industrially relevant traits, such as stress-related response, fermentative performance and profile of secondary metabolites and this approach is ineffective in modifying quantitative traits. Furthermore, genetically engineered strains are opposed by regulation No 1829/2003 of the European Parliament and of the Council, which prohibits GMOs in foodstuffs. In this picture, genome hybridization techniques, understood as natural and random rearrangement between strains exploiting the natural phenotypic variation within wild yeast populations, is a valid biotechnological tool to create genetically non-modified organisms (non-GMO) with improved phenotypes (Steensels et al. 2014). The hybridization (both sexual and asexual) produce random gene arrangements that are then tested and selected through screening procedures and technological simulations. Saccharomyces sensu stricto interspecific hybrids have been found in different fermentation processes: in addition to Saccharomyces. pastorianus , present in lager brewing, other hybrid strains have also been described from wine and cider (Masneuf et al. 1998; Groth et al. 1999; Naumova et al. 2005). For example, the type strain of Saccharomyces bayanus, originally isolated from beer, has recently been suggested to be a hybrid between S. cerevisiae and S. bayanus based on the presence of subtelomeric repeated sequences and genes (Nguyen et al. 2000; de Barros Lopes et al. 2002; Nguyen and Gaillardin 2005).
In general, the strength of this approach is the production of new fermenting strains that acquire physiological properties from both parents. The principal improved phenotype traits concern the ethanol and acetic acid tolerance, the copper resistance, the glycerol production, the high osmotic stress resistance, and the utilization of xylose (Brown and Oliver 1982; Aarnio et al. 1991; Adamo et al. 2012; Kutyna et al. 2012; Ekberg et al. 2013; Shen et al. 2012).
The prospects to obtain superior industrial wine yeasts are extremely bright. Yeasts offer unique advantages for strain improvement: they combine sexual and asexual life cycles, they can be easily cultivated in high numbers, and genetic transformation is often easy. Moreover, most strain improved by hybridization could be profitable involved in the rectification of fermentation disorders in spontaneous fermentations has been recently described in the literature (König and Claus 2018). Recent investigations have provided convincing evidence that fermentation problems can be overcome when must fermentations are successively performed with S. bayanus and the triple hybrid S. cerevisiae × Saccharomyces kudriavzevii × S. bayanus. The triple hybrid uses amino acids as a nitrogen source in the absence of ammonium and it also exhibits a fructophilic character with an enhanced uptake of fructose in comparison to glucose. This applicative example revealed that hybrid strains could be a promising tool for winemakers not always for the creation of novel wine types with desired sensory characteristics under more challenging conditions, but also ex post to solve fermentation problems during spontaneous fermentation or especially when the composition of the must components is not optimal because of critical climatic or soil conditions.
Several yeast hybrid strains are already commercially available, such as the strain “Oenoferm® X-treme” that is a GMO-free hybrid yeast obtained from the protoplast fusion of two different S. cerevisiae strains; the strain “Cross Evolution” a natural cross hybrid between S. cerevisiae yeasts; strain NT 202 is a product of the yeast hybridization program; the strain S6U a hybrid of S. cerevisiae × S. bayanus and the strain VIN7 an allotriploid interspecific hybrid of a heterozygous diploid complement of S. cerevisiae chromosomes and a haploid S. kudriavzevii genomic contribution (Borneman et al. 2012, 2016; Hart et al. 2016; Pérez-Torrado et al. 2018).
4.4 Controlled Mixed Fermentation
Although most research on wine microbiology has focused on Saccharomyces yeasts, particularly S. cerevisiae, there is a growing interest in studying and characterising non-Saccharomyces yeasts for development of starter cultures.
As already reported above, in the past wine was produced through spontaneous non-controlled fermentation by microflora residing on grapes, vineyard and in winery. In this way, many yeast species, not always identified, contribute to wine fermentation to obtain not reproducible and determining sometime failed results. Afterwards, the use of pure S. cerevisiae starter cultures, which establishes a dominant yeast population from the beginning of fermentation, has enabled modern wineries to produce predictable and reliable wines with established quality standards.
The sensory profile of wines produced by monoculture-inoculated fermentations differ substantially from those that are spontaneously fermented, principally for the biochemical composition of un-inoculated wines, which are distinctly different from wine obtained by pure fermentations (Varela et al. 2009). Certainly, spontaneous fermentations imply a higher risk of sluggish and/or incomplete fermentation and spoilage trend if compared to pure processes characterized by many default desirable characteristics but less complex flavour profiles (Jolly et al. 2014; Ugliano and Henschke 2009).
On the basis of this view, in the last years wine researchers have explored the controlled use of non-Saccharomyces starter cultures in addition to commercial and conventional S. cerevisiae starters. It is certainly known that non-Saccharomyces yeasts are generally unable to complete alcoholic fermentation on their own, for this they are used in pairs with S. cerevisiae wine strain. This can be achieved by inoculating first with the non-Saccharomyces yeast followed by a wine strain of S. cerevisiae to finish the fermentation. This is known as sequential inoculation, as opposed to simultaneous inoculation, in which two or more yeasts are added at the same time (Ciani et al. 2010; Jolly et al. 2003). In this regard, The use of non-Saccharomyces yeast in winemaking has grown enormously in the last years and several investigations have been carried out to better understand the impact of non-Saccharomyces strains on the chemical and sensorial properties of wine (Ciani and Maccarelli 1998; Comitini et al. 2011; Renault et al. 2015; Swiegers et al. 2005). In this regard, it is well established a wide intraspecific variability of oenological characters, peculiar positive oenological traits and, above all, a different behaviour in co-culture due to interactions with S. cerevisiae. All these aspects have highlighted a significant role of these non-conventional yeasts in determining the analytical and sensory profile and the aromatic complexity of wine.
4.5 Non-Saccharomyces Yeasts as Biotechnological Tool
The controlled multistarter fermentation with S. cerevisiae is the most profitable modality to use of these selected non-conventional wine yeasts. Several objectives can be pursued with the use of controlled mixed cultures with non-conventional yeasts: (i) enhancement of flavour and aroma complexity; (ii) distinctive features; (iii) ethanol reduction; (iv) control of spoilage microflora.
4.5.1 Aroma Enhancement
The contribution of selected non-Saccharomyces yeasts during wine production will be provided focusing the attention on the principal features such as aromatic profile, the color stability and polysaccharides production, the modulation of acidity, the ethanol reduction and concerning about antimicrobial activity toward undesired strains.
Certainly, the aromatic profile is one of most important traits that contribute to the quality of wine. As in many foods, wine aroma is composed by 100 s of different compounds with concentrations that can vary between 10−1 and 10−10 mg/mL. The balance and interaction of all of them determine the wine aromatic quality (Padilla et al. 2016). In literature, several works investigated on the production of volatile aromas, such as esters by different non-Saccharomyces yeast species that positively contribute to enhance the aroma profile of wines. (Moreira et al. 2008; Rojas et al. 2003; Viana et al. 2008). Between them, ethyl acetate and isoamyl acetate is often produced by yeast strains in natural grape juice during fermentation. For example, Kloeckera apiculata exhibited the highest ability for acetate formation; Hansenula subpelliculosa , Kluyveromyces marxianus , T. delbrueckii and S. cerevisiae produced intermediate levels and P. membranaefaciens and C. guilliermondii very low levels of the two esters. In general, the high production of esters did not always negatively influence the aromatic profile of wines Moreira et al. (2008).
Several applied studies focused the attention on T. delbrueckii, a species low frequently isolated on grape surface but one of the most studied species to increase flavour and aroma complexity in alcoholic beverages. Indeed, T. delbrueckii possesses several positive features that could be profitable used. Several investigations agree that T. delbrueckii impact on aromatic composition and sensory attributes of wines in both simultaneous and sequential fermentation through an increase of acetate ester (Cordero-Bueso 2013), thiols (3-sulfanylhexan-1-ol and 3-sulfanylhexyl acetate (Renault et al. 2015; Zott et al. 2008), terpenes (α terpineol and linalool) (Čuš and Jenko 2013), 2 phenyl-ethanol (Comitini et al. 2011).
Another non-Saccharomyces yeast is M. pulcherrima , species, frequently present on the grape surface and often recovered during the initial stages of alcoholic fermentation. M. pulcherrima is a high producer of β-glucosidase (Rodriguez et al. 2010), and its presence in mixed cultures can provide significant enhancements in the wine of higher alcohols, esters and terpenoids. Its aromatic profile in mixed fermentation was characterized by “citrus/grape fruits” some smoky and flowery attributes in Risling and Macabeo grape varieties respectively González-Royo et al. 2015). Also W. anomalus (formerly P. anomala ) resulted in positive contribution to aroma profile of wines in mixed fermentation determining an enhancement of isoamyl acetate and ethyl esters (Kurita 2008). Finally, an interesting non-Saccharomyces wine yeast to enhance complexity and overall aroma profile is Zygotorulaspora florentina , a yeast responsible of increased fruity and floral notes as well as lower perception of astringency (Lencioni et al. 2018). A wide and deepened information about the aroma enhancement of non-Saccharomyces yeasts in winemaking are dealt in the Chap. 2.
4.5.2 Distinctive Features
It has long been known the ability of some yeast species to metabolize malic acid. Schizosaccharomyces yeasts (Schizosaccharomyces pombe, Schyzosaccharomyces japonicus ) are characterized by malo-alcoholic fermentation and they are capable to completely metabolize the malic acid present in grape must and wine (Magyar and Panyik 1989; Ciani 1995) and could be profitable used in winemaking. In addition, more recently works showed that these yeasts species in mixed fermentation determined and increase in the production of pigments and large amounts of polysaccharides (Domizio et al. 2017; Escott et al. 2018). On the other hand, biological acidification is a desired feature in grape juices deficient in acidity generally coming from wines of warm climates. In addition in the last years, there is an increasing interest due to a progressive reduction of the total acidity of wines caused by global climate change and variations in viticulture and oenology practices. In this context Lachancea thermotolerans showed a peculiar ability to produce large amounts of lactic acid, together with glycerol and 2-phenyl ethanol during fermentation of grape musts (Kapsopoulou et al. 2007; Comitini et al. 2011; Gobbi et al. 2013). Glycerol production is another relevant feature of non - Saccharomyces yeasts. S. bacillaris (synonym Candida zemplinina ) (Rantsiou et al. 2012; Duarte et al. 2012) and Starmerella bombicola (formerly C. stellata ) exhibit strong fructophilic character and shows the ability to produce high amounts of glycerol (Ciani and Ferraro 1996). In mixed fermentation with S. cerevisiae these yeast species exhibited positive interactions in the production and degradation of metabolites (Ciani and Ferraro 1998). In addition to large glycerol production M. pulcherrima exhibited some positive features such as polysaccharides and glycosidase activity (Comitini et al. 2011). Various enzymatic activities important for enzymatic release of aromatic compounds in winemaking, were found in several other non Saccharomyces yeasts such as Hanseniaspora, Pichia and Candida genera (Rodríguez et al. 2007) and well as large production of polysaccharides (Domizio et al. 2011).
Another positive trait desired and pursued by non-Saccharomyces yeast is the low production of volatile acidity. This feature is one of the fundamental character to select strain for the oenological use. Some non-Saccharomyces species such as T. delbrueckii and C. stellata (now reclassified as Starmerella bombicola ) exhibited a very low production of volatile acidity (Ciani and Maccarelli 1998). In mixed fermentation with S. cerevisiae both T. delbrueckii and C. stellata showed a consistent reduction of volatile acidity (Ciani and Ferraro 1998). Similarly a reduction of acetic acid production was obtained in sweet wine fermentations in mixed fermentations using C. zemplinina (now reclassified as S. bacillaris) (Rantsiou et al. 2012).
4.5.3 Ethanol Reduction in Wine
Nowadays, the progressive increase in alcohol levels in wine, is a growing problem affecting the winemaking industry. Indeed, over the last two decades, there has been a progressive increase in the ethanol content in wines of c.a. two degrees over the viticulture areas (Alston et al. 2011; Gonzalez et al. 2013). This increase is mainly due to two main concerns: global climate change and the new wine styles often associated with increased grape maturity. For example, in wine the harvest the grapes at complete phenolic maturation may determine a overripe grapes and consequently the production of wines with high ethanol content. On the other hand, global climate change has deeply influenced the vine phenology and the grape composition, resulting in grapes with lower acidity, phenolic maturation and tannin content modifying other wine sensory attributes.
In order to overcome these issues, the market focus is directed to wines with a moderate alcohol content. In addition, lowering ethanol content has an economic interest due to the high taxes imposed in some countries. In this context, there are a rising interest in ethanol reduction in wine. Microbiological approach for decreasing ethanol concentrations appears a promising way and there is a growing interest to evaluate the use of non-Saccharomyces wine yeasts. There are several features possessed by non-Saccharomyces wine yeast that are a potential tool for the reduction of alcohol content in wine: a wide variability in ethanol yield (Contreras et al. 2014; 2015; Gobbi et al. 2014; Magyar and Tóth 2011) and the differences in regulatory respiro-fermentative metabolism with S. cerevisiae (Gonzalez et al. 2013). Indeed, among non-Saccharomyces wine yeasts some strains/species showed and sugar consumption by respiration (Crabtree negative). Therefore, both of these features of non-Saccharomyces yeasts have indicated a promising way to limit ethanol production. The approach used to use non-Saccharomyces wine yeasts to limit the production of ethanol is the mixed culture (simultaneous or sequential) since the inability of these yeasts of completing alcoholic fermentation (Ciani et al. 2016).
4.5.4 Control of Spoilage Microflora
Another possible applicative use of non-Saccharomyces yeasts in winemaking regards the control of spoilage microorganisms. During different stages of fermentation, a punctual and timely control of potential spoilage microorganisms is needed. In particular, during fermentation and aging stages of wine, the most spoilage yeast is B. bruxellensis responsible of undesired odors and considered the current major concern for winemakers, since an effective method to control their growth has not yet been developed.
Dekkera/Brettanomyces are described in the literature as part of the microbiota of many fermented beverages including cider, some type of beer, kombucha and kefyr, etc. (Morrissey et al. 2004). Dekkera/Brettanomyces can grow during the wine aging and even after their bottling; on the contrary, these yeasts are rarely found during the alcoholic fermentation of grape must. A few studies have reported their presence on grapes due to the difficult cultivation while in winery, in particular in vats, pumps or equipments difficult to sanitizes, Brettanomyces yeasts are more easily found (Fugelsang and Zoecklein 2003; Pretorius 2000; Renouf and Lonvaud-Funel 2007).
Different strains of Brettanomyces can show great differences in their production of volatile phenols. The variety of grape used also affects the sensorial perception of ethylphenols. Phister and Mills (2004) indicate detection thresholds to be high in monovarietal Cabernet Sauvignon wines, and lower in Tempranillo wines. The treatments to reduce the negative effect caused by Dekkera/Brettanomyces are based on both preventive and curative actions. Certain additives can inhibit the growth of Brettanomyces, including sulphur dioxide (SO2). The recommended molecular dose of SO2 is highly variable, from 0.3 to 0.8 mg/L. But these doses do not consider differences of strain resistance to sulfites or yeast population levels. Moreover, SO2 is known as a chemical stressor inducing a viable but nonculturable (VBNC) state of B. bruxellensis that are non-detectable by plate counting, can lead to new contamination when the amount of sulfite decreases over time (Capozzi et al. 2016). Moreover, the SO2 preservative agent has been largely demonstrated to have negative effects in wine consumers, including allergic reactions, asthma and headaches. This led to the establishment of strict regulations governing its use in the wine industry (Guerrero and Cantos-Villar 2015) with a direct consequence that the industry is interested to new ways to reduce sulphur dioxide levels, without changing the sensory quality of the wine. On the basis of this, a valid and natural alternative could be represented by bioactive compound produced by yeasts (Muccilli and Restuccia 2015). Biopreservation or biocontrol refers to the use of natural or controlled microorganisms, or their antimicrobial products, to extend the shelf life and to enhance the safety of food and beverages. This can be achieved by the addition of antimicrobial metabolites, such as killer toxins, or the direct application of pro-technological killer strain. A number of microorganisms and other biological agents have been regarded to be crucial in the biopreservation of food and beverages. In this context, a large group of non-Saccharomyces killer yeasts, able to produce killer toxins, can counteract Dekkera/Brettanomyces spoilage yeasts in wine.
Yeast killer toxins, also named mycocins or zymocins were initially defined as extracellular proteins, glycoproteins or glycolipids that disrupt the cell membrane function in susceptible yeast bearing receptors for the compound, whose activity is directed primarily against yeast closely related to the producer strain, which has a protective factor. The first mycocins were identified in association with S. cerevisiae in the brewing industry, but several others have since been isolated, frequently where yeast populations occur in high density and in highly competitive conditions, as for example fermented olive brine and fermenting grape must. Biological control could have an important application during the maturation and wine ageing of wines. In this regard, killer toxins secreted by W. anomalus (Pikt) and K. wickerhamii (Kwkt) were tested to control Dekkera/Brettanomyces spoilage yeasts. The stability in wine and the fungicidal effect of these two zymocins were demonstrated (Comitini et al. 2004). Thus, a potential application for the two toxins as antimicrobial agents active on Dekkera/Brettanomyces during wine ageing and storage can be hypothesized. Also, another killer toxin produced by Ustilago maydis it was seen to have efficacy to control B. bruxellensis , in mixed cultures under winemaking conditions Santos et al. (2011).
Recently, two new killer toxins produced by Candida pyralidae with an antimicrobial effect against B. bruxellensis , was tested in wine (Mehlomakulu et al. 2014). The killer toxins were stable under winemaking conditions and the activity was not affected by the ethanol and sugar concentrations typically found in grape juice and wine. Another new killer toxin from T. delbrueckii was identified and partial characterized. This zymocin, showed also a potential biocontrol effect on B. bruxellensis and other spoilage non-Saccharomyces yeasts such as Pichia spp.
However, other biological methods besides killer yeasts, were evaluated to control B. bruxellensis using non-Saccharomyces specific strains. For example, M. pulcherrima secretes pulcherriminic acid that exhibits an effective inhibitory effect to the growth of B. bruxellensis. In this case, the antimicrobial activity of M. pulcherrima does not seem due to proteinaceous compounds but to the precursor of pulcherrimin pigment that depletes iron present in the medium, making it not available to the other yeasts. Moreover, cell-to-cell contact and quorum sensing have been investigated as mechanisms involved in non-Saccharomyces-mixed fermentation (Oro et la. 2014). Quorum sensing was recently examined in H. uvarum , Torulaspora pretoriensis , Zygosaccharomyces bailii , C. zemplinina (S. bacillaris ), and B. bruxellensis . Results indicated species-specific kinetics for the production of 2-phenylethanol, tryptophol, and tyrosol, considered the main molecules involved in the quorum sensing mechanism (Zupan et al. 2013; Avbelj et al. 2016).
5 Conclusions
Grape must is a complex matrix where grapes, microbes and technological process determine the final composition of wine. Yeasts associated with grapes and winery environment may influence both the analytical composition and sensorial profile of final wine. Indeed, the ecological distribution of microbial community along the whole of the wine production chain plays an important role in the composition of the final wine. For these reasons investigations on yeast microflora of the different wine regions, interactions among them and with other biotic and abiotic factors are of significant importance in wine production. The use of new metagenomic techniques such as new generation sequencing (NGS) strategies will allow to acquire additional knowledge to have a more complete picture on wine yeast ecology. Investigations on physiology features of selected wine yeasts (Saccharomyces and non-Saccharomyces yeasts) may positively contribute to achieve some goals as: enhanced aroma profile and complexity, ethanol reduction and biocontrol strategy. In this way, applied studies on fermentative yeasts could provide new opportunities in the oenological field, such as the introduction on the market of products with distinctive analytical and sensory characteristics due to recognized yeast strains.
References
Aarnio, T., Suikho, M., & Kauppinen, V. (1991). Isolation of acetic acid tolerant baker’s yeast variants in a turbidostat. Applied Biochemistry and Biotechnology, 27, 55–63.
Abdelfattah, A., Wisniewski, M., Droby, S., & Schena, L. (2016). Spatial and compositional variation in the fungal communities of organic and conventionally grown apple fruit at the consumer point-of-purchase. Horticulture Research, 3, 16047.
Adamo, G. M., Brocca, S., Passolunghi, S., Salvato, B., & Lotti, M. (2012). Laboratory evolution of copper tolerant yeast strains. Microbial Cell Factories, 11(1), 1.
Albergaria, H., Francisco, D., Gori, K., Arneborg, N., & Gírio, F. (2010). Saccharomyces cerevisiae CCMI 885 secretes peptides that inhibit the growth of some non-Saccharomyces wine-related strains. Applied Microbiology and Biotechnology, 86, 965–972.
Alston, J. M., Fuller, K. B., Lapsley, J. T., & Soleas, G. (2011). Too much of a good thing? Causes and consequences of increases in sugar content of California wine grapes. Journal of Wine Economics, 6, 135–159.
Arneborg, N., Siegumfeldt, H., Andersen, G. H., Nissen, P., Daria, V. R., Rodrigo, P. G., & Glückstad, J. (2005). Interactive optical trapping shows that confinement is a determinant of growth in a mixed yeast culture. FEMS Microbiology Letters, 245, 155–159.
Avbelj, M., Zupan, J., Kranjc, L., & Raspor, P. (2016). Quorum-sensing kinetics in Saccharomyces cerevisiae: A symphony of ARO genes and aromatic alcohols. Journal of Agricultural and Food Chemistry, 63, 8544–8550.
Bacon, C. W., & White, J. F. (2016). Functions, mechanisms and regulation of endophytic and epiphytic microbial communities of plants. Symbiosis, 68, 87–98.
Barata, A., González, S., Malfeito-Ferreira, M., Querol, A., & Loureiro, V. (2008). Sour rot-damaged grapes are sources of wine spoilage yeasts. FEMS Yeast Research, 8, 1008–1017.
Barata, A., Malfeito-Ferreira, M., & Loureiro, V. (2012). The microbial ecology of wine grape berries. International Journal of Food Microbiology, 153, 243–259.
Belda, I., Navascués, E., Marquina, D., Santos, A., Calderon, F., & Benito, S. (2015). Dynamic analysis of physiological properties of Torulaspora delbrueckii in wine fermentations and its incidence on wine quality. Applied Microbiology and Biotechnology, 9, 1911–1922.
Belda, I., Ruiz, J., Esteban-Fernández, A., Navascués, E., Marquina, D., Santos, A., & Moreno-Arribas, M. (2017a). Microbial contribution to wine aroma and its intended use for wine quality improvement. Molecules, 22, 189.
Belda, I., Zarraonaindia, I., Perisin, M., Palacios, A., & Acedo, A. (2017b). From vineyard soil to wine fermentation: Microbiome approximations to explain the “terroir” concept. Frontiers in Microbiology, 8, 821.
Bellon, J. R., Schmid, F., Capone, D. L., Dunn, B. L., & Chambers, P. J. (2013). Introducing a new breed of wine yeast: Interspecific hybridisation between a commercial Saccharomyces cerevisiae wine yeast and Saccharomyces mikatae. PLoS One, 8, e62053.
Beltran, G., Torija, M. J., Novo, M., Ferrer, N., Poblet, M., Guillamón, J. M., & Mas, A. (2002). Analysis of yeast populations during alcoholic fermentation: A six year follow-up study. Systematic and Applied Microbiology, 25, 287–293.
Benito, S., Palomero, F., Calderón, F., Palmero, D., & Suárez-Lepe, J. A. (2014). Selection of appropriate Schizosaccharomyces strains for winemaking. Food Microbiology, 42, 218–224.
Bisson, L. F. (1999). Stuck and sluggish fermentations. American Journal of Enology and Viticulture, 50, 107–119.
Bisson, L. F., Joseph, C. L., & Domizio, P. (2017). Yeasts. In Biology of microorganisms on grapes, in must and in wine (pp. 65–101). Cham: Springer.
Blanco, P., Ramilo, A., Cerdeira, M., & Orriols, I. (2006). Genetic diversity of wine Saccharomyces cerevisiae strains in an experimental winery from Galicia (NW Spain). Antonie Van Leeuwenhoek, 89, 351–357.
Blanco, P., Orriols, I., & Losada, A. (2011). Survival of commercial yeasts in the winery environment and their prevalence during spontaneous fermentations. Journal of Industrial Microbiology & Biotechnology, 38, 235–239.
Bokulich, N. A., & Mills, D. A. (2012). Next-generation approaches to the microbial ecology of food fermentations. BMB Reports, 45, 377–389.
Bokulich, N. A., Ohta, M., Richardson, P. M., & Mills, D. A. (2013). Monitoring seasonal changes in winery-resident microbiota. PLoS One, 8, e66437.
Bokulich, N. A., Thorngate, J. H., Richardson, P. M., & Mills, D. A. (2014). Microbial biogeography of wine grapes is conditioned by cultivar, vintage, and climate. Proceedings of the National Academy of Sciences of the United States of America, 111, E139–E148.
Bokulich, N. A., Collins, T. S., Masarweh, C., Allen, G., Heymann, H., Ebeler, S. E., & Mills, D. A. (2016). Associations among wine grape microbiome, metabolome, and fermentation behavior suggest microbial contribution to regional wine characteristics. MBio, 7, e00631–e00616.
Bonnefoy, C., Quénol, H., Bonnardot, V., Barbeau, G., Madelin, M., Planchon, O., & Neethling, E. (2013). Temporal and spatial analyses of temperature in a French wine-producing area: The Loire Valley. International Journal of Climatology, 33, 1849–1862.
Borneman, A. R., Desany, B. A., Riches, D., Affourtit, J. P., Forgan, A. H., Pretorius, I. S., Egholm, M., & Chambers, P. J. (2012). The genome sequence of the wine yeast VIN7 reveals an allotriploid hybrid genome with Saccharomyces cerevisiae and Saccharomyces kudriavzevii origins. FEMS Yeast Research, 12, 88–96.
Borneman, A. R., Forgan, A. H., Kolouchova, R., Fraser, J. A., & Schmidt, S. A. (2016). Whole genome comparison reveals high levels of inbreeding and strain redundancy across the spectrum of commercial wine strains of Saccharomyces cerevisiae. G3: Genes, Genomes, Genetics, 6(4), 115.
Brown, S. W., & Oliver, S. G. (1982). Isolation of ethanol-tolerant mutants of yeast by continuous selection. European Journal of Applied Microbiology and Biotechnology, 16, 119–122.
Brysch-Herzberg, M., & Seidel, M. (2015). Yeast diversity on grapes in two German wine growing regions. International Journal of Food Microbiology, 214, 137–144.
Čadež, N., Zupan, J., & Raspor, P. (2010). The effect of fungicides on yeast communities associated with grape berries. FEMS Yeast Research, 10, 619–630.
Cafarchia, C., Camarda, A., Romito, D., Campolo, M., Quaglia, N. C., Tullio, D., & Otranto, D. (2006). Occurrence of yeasts in cloacae of migratory birds. Mycopathologia, 16, 229–234.
Capozzi, V., Garofalo, C., Chiriatti, M. A., Grieco, F., & Spano, G. (2015). Microbial terroir and food innovation: The case of yeast biodiversity in wine. Microbiological Research, 181, 75–83.
Capozzi, V., Di Toro, M. R., Grieco, F., Michelotti, V., Salma, M., Lamontanara, A., et al. (2016). Viable But Not Culturable (VBNC) state of Brettanomyces bruxellensis in wine: New insights on molecular basis of VBNC behaviour using a transcriptomic approach. Food Microbiology, 59, 196–204.
Chatonnet, P., Dubourdie, D., Boidron, J. N., & Pons, M. (1992). The origin of ethylphenols in wines. Journal of the Science of Food and Agriculture, 60, 165–178.
Chong, K. P., Rossall, S., & Atong, M. (2009). In vitro antimicrobial activity and fungitoxicity of syringic acid, caffeic acid and 4-hydroxybenzoic acid against Ganoderma boninense. Journal of Agricultural Science, 1, 15.
Ciani, M. (1995). Continuous deacidification of wine by immobilized Schizosaccharomyces pombe cells: Evaluation of malic acid degradation rate and analytical profiles. Journal of Applied Microbiology, 79, 631–634.
Ciani, M., & Comitini, F. (2015). Yeast interactions in multi-starter wine fermentation. Current Opinion in Food Science, 1, 1–6.
Ciani, M., & Ferraro, L. (1996). Enhanced glycerol content in wines made with immobilized Candida stellata cells. Applied and Environmental Microbiology, 62, 128–132.
Ciani, M., & Ferraro, L. (1998). Combined use of immobilized Candida stellata cells and Saccharomyces cerevisiae to improve the quality of wines. Journal of Applied Microbiology, 85, 247–254.
Ciani, M., & Maccarelli, F. (1998). Oenological properties of non-Saccharomyces yeasts associated with wine-making. World Journal of Microbiology and Biotechnology, 14, 199–203.
Ciani M., & Rosini G. (1986). Sulla microflora blastomicetica del Sagrantino D.O.C.: Microflora dei vini e dei locali di vinificazione. Annals Fac. Agr. Univ. Perugia 40: 103–112.
Ciani, M., & Rosini, G. (1990). Selection of strains Saccharomyces cerevisiae of “Sagrantino D.O.C.” for their wine – making properties: Preliminary results. Italian Journal of Food Science, 3, 151–158.
Ciani, M., Mannazzu, I., Marinangeli, P., Clementi, F., & Martini, A. (2004). Contribution of winery-resident Saccharomyces cerevisiae strains to spontaneous grape must fermentations. Antonie Van Leeuwenhoek, 85, 159–164.
Ciani, M., Beco, L., & Comitini, F. (2006). Fermentation behaviour and metabolic interactions of multistarter wine yeast fermentations. International Journal of Food Microbiology, 108, 239–245.
Ciani, M., Comitini, F., Mannazzu, I., & Domizio, P. (2010). Controlled mixed culture fermentation: A new perspective on the use of non-Saccharomyces yeasts in winemaking. FEMS Yeast Research, 10, 123–133.
Ciani, M., Morales, P., Comitini, F., Tronchoni, J., Canonico, L., Curiel, J. A., & Gonzalez, R. (2016). Non-conventional yeast species for lowering ethanol content of wines. Frontiers in Microbiology, 7, 642.
Clavijo, A., Calderón, I. L., & Paneque, P. (2010). Diversity of Saccharomyces and non-Saccharomyces yeasts in three red grape varieties cultured in the Serrania de Ronda (Spain) vine-growing region. International Journal of Food Microbiology, 143, 241–245.
Cocolin, L., Rantsiou, K., Iacumin, L., Urso, R., Cantoni, C., & Comi, G. (2004). Study of the ecology of fresh sausages and characterization of populations of lactic acid bacteria by molecular methods. Applied and Environmental Microbiology, 70, 1883–1894.
Cocolin, L., Alessandria, V., Dolci, P., Gorra, R., & Rantsiou, K. (2013). Culture independent methods to assess the diversity and dynamics of microbiota during food fermentation. International Journal of Food Microbiology, 167, 29–43.
Combina, M., Elía, A., Mercado, L., Catania, C., Ganga, A., & Martinez, C. (2005). Dynamics of indigenous yeast populations during spontaneous fermentation of wines from Mendoza, Argentina. International Journal of Food Microbiology, 99, 237–243.
Comitini, F., & Ciani, M. (2006). Survival of inoculated Saccharomyces cerevisiae strain on wine grapes during two vintages. Letters in Applied Microbiology, 42, 248–253.
Comitini, F., Ingeniis De, J., Pepe, L., Mannazzu, I., & Ciani, M. (2004). Pichia anomala and Kluyveromyces wickerhamii killer toxins as new tools against Dekkera/Brettanomyces spoilage yeasts. FEMS Microbiology Letters, 238, 235–240.
Comitini, F., Ferretti, R., Clementi, F., Mannazzu, I., & Ciani, M. (2005). Interactions between Saccharomyces cerevisiae and malolactic bacteria: preliminary characterization of a yeast proteinaceous compound (s) active against Oenococcus oeni. Journal of Applied Microbiology, 99, 105–111.
Comitini, F., Gobbi, M., Domizio, P., Romani, C., Lencioni, L., Mannazzu, I., & Ciani, M. (2011). Selected non-Saccharomyces wine yeasts in controlled multistarter fermentations with Saccharomyces cerevisiae. Food Microbiology, 28, 873–882.
Comitini, F., Oro, L., Canonico, L., Marinelli, V., & Ciani, M. (2019). Occurrence of Brettanomyces bruxellensis on grape berries and in related winemaking cellar. Frontiers in Microbiology, 10, 415.
Connell, L., Stender, H., & Edwards, C. G. (2002). Rapid detection and identification of Brettanomyces from winery air samples based on peptide nucleic acid analysis. American Journal of Enology and Viticulture, 53, 322–324.
Contreras, A., Hidalgo, C., Henschke, P. A., Chambers, P. J., Curtin, C., & Varela, C. (2014). Evaluation of non-Saccharomyces yeasts for the reduction of alcohol content in wine. Applied and Environmental Microbiology, 80, 1670–1678.
Contreras, A., Curtin, C., & Varela, C. (2015). Yeast population dynamics reveal a potential ‘collaboration’ between Metschnikowia pulcherrima and Saccharomyces uvarum for the production of reduced alcohol wines during Shiraz fermentation. Applied Microbiology and Biotechnology, 99, 1885–1895.
Coombe, P. E., Srinivasan, M. V., & Guy, R. G. (1989). Are the large monopolar cells of the insect lamina on the optomotor pathway. Journal of Comparative Physiology. A, Neuroethology, Sensory, Neural, and Behavioral Physiology, 166, 23–35.
Cordero-Bueso, G. A. (2013). Aplicación del Análisis Sensorial de los Alimentos en la Cocina y en la Industria Alimentaria. Sede de Carmona de la Universidad Pablo de Olavide, XI, 13–96.
Cordero-Bueso, G., Arroyo, T., Serrano, A., Tello, J., Aporta, I., Vélez, M. D., & Valero, E. (2011). Influence of the farming system and vine variety on yeast communities associated with grape berries. International Journal of Food Microbiology, 145, 132–139.
Costantì, M., Poblet, M., Arola, L., Mas, A., & Guillarmon, J. M. (1997). Analysis of yeast population during alcoholic fermentation in a newly established winery. American Journal of Enology and Viticulture, 48, 339–344.
Cray, J. A., Bel, l., Bhaganna, P., Mswaka, A. Y., Timson, D. J., & Hallsworth, J. (2013). The biology of habitat dominance; can microbes behave as weeds? Microbial Biotechnology, 6, 453–492.
Čuš, F., & Jenko, M. (2013). The influence of yeast strains on the composition and sensory quality of Gewürztraminer wine. Food Technology and Biotechnology, 51, 547–553.
De Barros, L. M., Bellon, J. R., Shirley, N. J., & Ganter, P. F. (2002). Evidence for multiple interspecific hybridization in Saccharomyces sensu stricto species. FEMS Yeast Research, 1, 323–331.
De Filippis, F., La Storia, A., Villani, F., & Ercolini, D. (2013). Exploring the sources of bacterial spoilers in beefsteaks by culture-independent high-throughput sequencing. PLoS One, 8, e70222.
del Carmen Portillo, M., & Mas, A. (2016). Analysis of microbial diversity and dynamics during wine fermentation of Grenache grape variety by high-throughput barcoding sequencing. LWT-Food Science and Technology, 72, 317–321.
del Monaco, S. M., Barda, N., Rubio, N., & Caballero, A. (2014). Selection and characterization of a Patagonian Pichia kudriavzevii for wine deacidification. Journal of Applied Microbiology, 117, 451–464.
Domizio, P., Romani, C., Lencioni, L., Comitini, F., Gobbi, M., Mannazzu, I., & Ciani, M. (2011). Outlining a future for non-Saccharomyces yeasts: Selection of putative spoilage wine strains to be used in association with Saccharomyces cerevisiae for grape juice fermentation. International Journal of Food Microbiology, 147, 170–180.
Domizio, P., Liu, Y., Bisson, L. F., & Barile, D. (2017). Cell wall polysaccharides released during the alcoholic fermentation by Schizosaccharomyces pombe and Schizosaccharomyces japonicus: Quantification and characterization. Food Microbiology, 61, 136–149.
Druvefors, U. A., Passoth, V., & Schnürer, J. (2005). Nutrient effects on biocontrol of Penicillium roqueforti by Pichia anomala J121 during airtight storage of wheat. Applied and Environmental Microbiology, 71, 1865–1869.
Duarte, F. L., Pimentel, N. H., Teixeira, A., & Fonseca, A. (2012). Saccharomyces bacillaris is not a synonym of Candida stellata: Reinstatement as Starmerella bacillaris comb. nov. Antonie Van Leeuwenhoek, 102(4), 653–658.
Ekberg, J., Rautio, J., Mattinen, L., Vidgren, V., Londesborough, J., & Gibson, B. R. (2013). Adaptive evolution of the lager brewing yeast Saccharomyces pastorianus for improved growth under hyperosmotic conditions and its influence on fermentation performance. FEMS Yeast Research, 13, 335–349.
Erten, H. (2002). Relations between elevated temperatures and fermentation behaviour of Kloeckera apiculata and Saccharomyces cerevisiae associated with winemaking in mixed cultures. World Journal of Microbiology and Biotechnology, 18, 373–378.
Escott, C., Del Fresno, J. M., Loira, I., Morata, A., Tesfaye, W., del Carmen González, M., et al. (2018). Formation of polymeric pigments in red wines through sequential fermentation of flavanol-enriched musts with non-Saccharomyces yeasts. Food Chemistry, 239, 975–983.
Fleet, G. H. (2003). Yeast interactions and wine flavour. International Journal of Food Microbiology, 86, 11–22.
Fleet, G. H., & Heard, G. M. (1993). Yeasts-growth during fermentation. In G. H. Fleet (Ed.), Wine microbiology and biotechnology (pp. 27–54). Chur: Harwood Academic Publishers.
Fleet, G. H., Lafon-Lafourcade, S., & Ribéreau-Gayon, P. (1984). Evolution of yeasts and lactic acid bacteria during fermentation and storage of Bordeaux wines. Applied and Environmental Microbiology, 48, 1034–1038.
Fleet, G. H., Prakitchaiwattana, C., Beh, A. L., & Heard, G. (2002). The yeast ecology of wine grapes. In Biodiversity and biotechnology of wine yeasts (Vol. 95, pp. 1–17). Kerala: Research Signpost.
Francesca, N., Canale, D. E., Settanni, L., & Moschetti, G. (2012). Dissemination of wine-related yeasts by migratory birds. Environmental Microbiology Reports, 4, 105–112.
Frezier, V., & Dubourdieu, D. (1992). Ecology of yeast strain Saccharomyces cerevisiae during spontaneous fermentation in a Bordeaux winery. American Journal of Enology and Viticulture, 53, 375–380.
Fugelsang, K. C., & Zoecklein, B. W. (2003). Population dynamics and effects of Brettanomyces bruxellensis strains on Pinot noir (Vitis vinifera L.) wines. American Journal of Enology and Viticulture, 54, 294–300.
Ganga, M. A., & Martinez, C. (2004). Effect of wine yeast monoculture practice on the biodiversity of non-Saccharomyces yeasts. Journal of Applied Microbiology, 96, 76–83.
Ganucci, D., Guerrini, S., Mangani, S., Vincenzini, M., & Granchi, L. (2018). Quantifying the effects of ethanol and temperature on the fitness advantage of predominant Saccharomyces cerevisiae strains occurring in spontaneous wine fermentations. Frontiers in Microbiology, 9.
Gao, C., & Fleet, G. H. (1988). The effects of temperature and pH on the ethanol tolerance of the wine yeasts Saccharomyces cerevisiae, Candida stellata and Kloeckera apiculata. The Journal of Applied Bacteriology, 65, 405–410.
Garofalo, C., Tristezza, M., Grieco, F., Spano, G., & Capozzi, V. (2016). From grape berries to wine: Population dynamics of cultivable yeasts associated to “Nero di Troia” autochthonous grape cultivar. World Journal of Microbiology and Biotechnology, 32, 59.
Gilbert, J. A., Jansson, J. K., & Knight, R. (2014a). The earth microbiome project: Successes and aspirations. BMC Biology, 12, 69.
Gilbert, J. A., van der Lelie, D., & Zarraonaindia, I. (2014b). Microbial terroir for wine grapes. Proceedings of the National Academy of Sciences of the United States of America, 111, 5–6. https://doi.org/10.1073/pnas.1320471110.
Giorello, F., Valera, M. J., Martin, V., Parada, A., Salzman, V., Camesasca, L., & Berna, L. (2018). Genomic and phenomic analysis of Hanseniaspora vineae provides insights for understanding yeast fermentation flavours that contribute to wine quality. Applied and Environmental Microbiology, 85(1), e01959–e01918.
Gobbi, M., Comitini, F., Domizio, P., Romani, C., Lencioni, L., Mannazzu, I., & Ciani, M. (2013). Lachamcea thermotolerans and Saccharomyces cerevisiae in simultaneous and sequential co-fermentation: A strategy to enhance acidity and improve the overall quality of wine. Food Microbiology, 33, 271–281.
Gobbi, M., De Vero, L., Solieri, L., Comitini, F., Oro, L., Giudici, P., & Ciani, M. (2014). Fermentative aptitude of non-Saccharomyces wine yeast for reduction in the ethanol content in wine. European Food Research and Technology, 239, 41–48.
Goddard, M. R., Anfang, N., Tang, R., Gardner, R. C., & Jun, C. (2010). A distinct population of Saccharomyces cerevisiae in New Zealand: Evidence for local dispersal by insects and human-aided global dispersal in oak barrels. Environmental Microbiology, 12(1), 63–73.
Godden, P., Wilkes, E., & Johnson, D. (2015). Trends in the composition of Australian wine 1984–2014. Australian Journal of Grape and Wine Research, 21, 741–753.
Gonzalez, R., Quirós, M., & Morales, P. (2013). Yeast respiration of sugars by non-Saccharomyces yeast species: A promising and barely explored approach to lowering alcohol content of wines. Trends in Food Science and Technology, 29, 55–61.
González-Royo, E., Pascual, O., Kontoudakis, N., Esteruelas, M., Esteve-Zarzoso, B., Mas, A., & Zamora, F. (2015). Oenological consequences of sequential inoculation with non-Saccharomyces yeasts (Torulaspora delbrueckii or Metschnikowia pulcherrima) and Saccharomyces cerevisiae in base wine for sparkling wine production. European Food Research and Technology, 240, 999–1012.
Grangeteau, C., Gerhards, D., von Wallbrunn, C., Alexandre, H., Rousseaux, S., & Guilloux-Benatier, M. (2016). Persistence of two non-Saccharomyces yeasts (Hanseniaspora and Starmerella) in the cellar. Frontiers in Microbiology, 7, 268.
Groth, G., Hansen, J., & Piškur, J. (1999). A natural chimeric yeast containing genetic material from three species. International Journal of Systematic Bacteriology, 49, 1933–1938.
Guerrero, R. F., & Cantos-Villar, E. (2015). Demonstrating the efficiency of sulphur dioxide replacements in wine: A parameter review. Trends in Food Science and Technology, 42, 27–43.
Guillamón, J. M., Barrio, E., & Querol, A. (1996). Characterization of wine yeast strains of the Saccharomyces genus on the basis of molecular markers: Relationships between genetic distance and geographic or ecological origin. Systematic and Applied Microbiology, 19, 122–132.
Hansen, E. H., Nissen, P., Sommer, P., Nielsen, J. C., & Arneborg, N. (2001). The effect of oxygen on the survival of non-Saccharomyces yeasts during mixed culture fermentations of grape juice with Saccharomyces cerevisiae. Journal of Applied Microbiology, 91, 541–547.
Hart, R. S., Jolly, N. P., Mohamed, G., Booyse, M., & Ndimba, B. K. (2016). Characterisation of Saccharomyces cerevisiae hybrids selected for low volatile acidity formation and the production of aromatic Sauvignon blanc wine. African Journal of Biotechnology, 15, 2068–2081.
Heard, G. M., & Fleet, G. H. (1985). Growth of natural yeast flora during the fermentation of inoculated wines. Applied and Environmental Microbiology, 50, 727–728.
Heard, G. M., & Fleet, G. H. (1988). The effects of temperature and pH on the growth of yeast species during the fermentation of grape juice. The Journal of Applied Bacteriology, 65, 23–28.
Henick-Kling, T., Edinger, W., Daniel, P., & Monk, P. (1998). Selective effects of sulfur dioxide and yeast starter culture addition on indigenous yeast populations and sensory characteristics of wine. Journal of Applied Microbiology, 84, 865–876.
Hernández, A., Pérez-Nevado, F., Ruiz-Moyano, S., Serradilla, M. J., Villalobos, M. C., Martín, A., & Córdoba, M. G. (2018). Spoilage yeasts: What are the sources of contamination of foods and beverages? International Journal of Food Microbiology, 286, 98–110.
Hierro, J. L., Maron, J. L., & Callaway, R. M. (2005). A biogeographical approach to plant invasions: The importance of studying exotics in their introduced and native range. Journal of Ecology, 93, 5–15.
Holland, T., & Smit, B. (2014). Recent climate change in the Prince Edward County winegrowing region, Ontario, Canada: Implications for adaptation in a fledgling wine industry. Regional Environmental Change, 14, 1109–1121.
Huglin, P. (1978). Nouveau mode d’évaluation des possibilités héliothermiques d’un milieu viticole. Comptes Rendus de l’Académie de l’Agriculture de France, 64, 1117–1126.
Jolly, N. P., Augustyn, O. P. H., & Pretorius, I. S. (2003). The occurrence of non-Saccharomyces cerevisiae yeast species over three vintages in four vineyards and grape musts from four production regions of the Western Cape, South Africa. South African Journal of Enology and Viticulture, 24, 35–42.
Jolly, N. P., Varela, C., & Pretorius, I. S. (2014). Not your ordinary yeast: Non-Saccharomyces yeasts in wine production uncovered. FEMS Yeast Research, 14, 215–237.
Jones-Vaid, M., Prasad, R., Singh, T., Jones, V., & Katiyar, S. K. (2012). Grape seed proanthocyanidins reactivate silenced tumor suppressor genes in human skin cancer cells by targeting epigenetic regulators. Toxicology and Applied Pharmacology, 263, 122–130.
Kapsopoulou, K., Mourtzini, A., Anthoulas, M., & Nerantzis, E. (2007). Biological acidification during grape must fermentation using mixed cultures of Kluyveromyces thermotolerans and Saccharomyces cerevisiae. World Journal of Microbiology and Biotechnology, 23, 735–739.
Katalinić, V., Možina, S. S., Skroza, D., Generalić, I., Abramovič, H., Miloš, M., & Boban, M. (2010). Polyphenolic profile, antioxidant properties and antimicrobial activity of grape skin extracts of 14 Vitis vinifera varieties grown in Dalmatia (Croatia). Food Chemistry, 119, 715–723.
Knight, S., Klaere, S., Fedrizzi, B., & Goddard, M. R. (2015). Regional microbial signatures positively correlate with differential wine phenotypes: Evidence for a microbial aspect to terroir. Scientific Reports, 5, 14233.
König, H., & Claus, H. (2018). A future place for Saccharomyces mixtures and hybrids in winemaking. Fermentation, 4, 67.
Kurita, O. (2008). Increase of acetate ester-hydrolysing esterase activity in mixed cultures of Saccharomyces cerevisiae and Pichia anomala. Journal of Applied Microbiology, 104, 1051–1058.
Kurtzman, C. P., & Droby, S. (2001). Metschnikowia fructicola, a new ascosporic yeast with potential for biocontrol of postharvest fruit rots. Systematic and Applied Microbiology, 24, 395–399.
Kurtzman, C., Fell, J. W., & Boekhout, T. (Eds.). (2011). The yeasts: a taxonomic study. Elsevier, Amsterdam.
Kutyna, D. R., Varela, C., Stanley, G. A., Borneman, A. R., Henschke, P. A., & Chambers, P. J. (2012). Adaptive evolution of Saccharomyces cerevisiae to generate strains with enhanced glycerol production. Applied Microbiology and Biotechnology, 93, 1175–1184.
Lachance, M. A. (2003). The Phaff school of yeast ecology. International Microbiology, 63, 163–167.
Lencioni, L., Taccari, M., Ciani, M., & Domizio, P. (2018). Zygotorulaspora florentina and Starmerella bacillaris in multistarter fermentation with Saccharomyces cerevisiae to reduce volatile acidity of high sugar musts. Australian Journal of Grape and Wine Research, 24, 368.
Lereboullet, A. L., Beltrando, G., Bardsley, D. K., & Rouvellac, E. (2014). The viticultural system and climate change: coping with long-term trends in temperature and rainfall in Roussillon, France. Regional Environmental Change, 14, 1951–1966.
Liu, X., Hoque, M., Larochelle, M., Lemay, J. F., Yurko, N., Manley, J. L., & Tian, B. (2017). Comparative analysis of alternative polyadenylation in S. cerevisiae and S. pombe. Genome Research, 27(10), 1685–1695.
Magyar, I., & Panyik, I. (1989). Biological deacidification of wine with Schizosaccharomyces pombe entrapped in Ca-alginate gel. American Journal of Enology and Viticulture, 40, 233–240.
Magyar, I., & Tóth, T. (2011). Comparative evaluation of some oenological properties in wine strains of Candida stellata, Candida zemplinina, Saccharomyces uvarum and Saccharomyces cerevisiae. Food Microbiology, 28, 94–100.
Martini, A. (1993). Origin and domestication of the wine yeast Saccharomyces cerevisiae. Journal of Wine Research, 4, 165–176.
Martins, G., Lauga, B., Miot-Sertier, C., Mercier, A., Lonvaud, A., Soulas, M. L., Soulas, G., & Masneuf-Pomarède, I. (2013). Characterization of epiphytic bacterial communities from grapes, leaves, bark and soil of grapevine plants grown, and their relations. PLoS One, 8, e73013.
Masneuf, I., Hansen, J., Groth, C., Piškur, J., & Dubourdieu, D. (1998). New hybrids between Saccharomyces sensu stricto yeast species found among wine and cider production strains. Applied and Environmental Microbiology, 64, 3887–3892.
Mehlomakulu, N. N., Setati, B., & Divol, G. (2014). Characterization of novel killer toxins secreted by wine- related non-Saccharomyces yeasts and their action on Brettanomyces spp. International Journal of Food Microbiology, 188, 83–91.
Mendoza, L. M., Manca de Nadra, M. C., & Farías, M. E. (2007). Kinetics and metabolic behaviour of a composite culture of Kloeckera apiculata and Saccharomyces cerevisiae wine related strains. Biotechnology Letters, 29, 1057–1063.
Mezzasalma, V., Sandionigi, A., Bruni, I., Bruno, A., Lovicu, G., Casiraghi, M., & Labra, M. (2017). Grape microbiome as a reliable and persistent signature of field origin and environmental conditions in Cannonau wine production. PLoS One, 12, e0184615.
Milanovic, V., Comitini, F., & Ciani, M. (2013). Grape berry yeast communities: Influence of fungicide treatments. International Journal of Food Microbiology, 161, 240–246.
Mills, D. A., Johannsen, E. A., & Cocolin, L. (2002). Yeast diversity and persistence in botrytis-affected wine fermentations. Applied and Environmental Microbiology, 68, 4884–4893.
Mira de Orduña, R., Liu, S.-Q., Patchett, M. L., & Pilone, G. J. (2000). Kinetics of the arginine metabolism of malolactic wine lactic acid bacteria Lactobacillus buchneri CUC-3 and Oenococcus oeni Lo111. Journal of Applied Microbiology, 89, 547–552.
Moisselin, J. M., & Dubuisson, B. (2006). Évolution des valeurs extrêmes de température et de précipitations au cours du XXe siècle en France.
Molnar, O., & Prillinger, H. (2005). Analysis of yeast isolates related to Metschnikowia pulcherrima using the partial sequences of the large subunit rDNA and the actin gene; description of Metschnikowia andauensis sp. Nov. Systematic and Applied Microbiology, 28, 717–726.
Moreira, N., Mendes, F., de Pinho, P. G., Hogg, T., & Vasconcelos, I. (2008). Heavy sulphur compounds, higher alcohols and esters production profile of Hanseniaspora uvarum and Hanseniaspora guilliermondii grown as pure and mixed cultures in grape must. International Journal of Food Microbiology, 124, 231–238.
Morrison-Whittle, P., & Goddard, M. R. (2015). Quantifying the relative roles of selective and neutral processes in defining eukaryotic microbial communities. ISME Journal, 9, 2003–2011.
Morrison-Whittle, P., & Goddard, M. R. (2018). From vineyard to winery: A source map of microbial diversity driving wine fermentation. Environmental Microbiology, 20, 75–84.
Morrissey, J. P., Dow, J. M., Mark, G. L., & O’Gara, F. (2004). Are microbes at the root of a solution to world food production?: Rational exploitation of interactions between microbes and plants can help to transform agriculture. EMBO Reports, 5, 922–926.
Muccilli, S., & Restuccia, C. (2015). Bioprotective role of yeasts. Microorganisms, 3, 588–611.
Naumova, E. S., Naumov, G. I., Masneuf-Pomarède, I., Aigle, M., & Dubourdieu, D. (2005). Molecular genetic study of introgression between Saccharomyces bayanus and S. cerevisiae. Yeast, 22, 1099–1115.
Neumann, P. A., & Matzarakis, A. (2014). Potential climate change impacts on winegrape must density and titratable acidity in southwest Germany. Climate Research, 59, 161–172.
Nguyen, H. V., & Gaillardin, C. (2005). Evolutionary relationships between the former species Saccharomyces uvarum and the hybrids Saccharomyces bayanus and Saccharomyces pastorianus; reinstatement of Saccharomyces uvarum (Beijerinck) as a distinct species. FEMS Yeast Research, 5, 471–483.
Nguyen, H. V., Lepingle, A., & Gaillardin, C. (2000). Molecular typing demonstrates homogeneity of Saccharomyces uvarum strains and reveals the existence of hybrids between S. uvarum and S. cerevisiae including the S. bayanus type strain CBS 380. Systematic and Applied Microbiology, 23, 71–85.
Nissen, P., & Arneborg, N. (2003). Characterization of early deaths of non-Saccharomyces yeasts in mixed cultures with Saccharomyces cerevisiae. Archives of Microbiology, 180, 257–263.
Nissen, P., Nielsen, D., & Arneborg, N. (2003). Viable Saccharomyces cerevisiae cells at high concentrations cause early growth arrest of non-Saccharomyces yeasts in mixed cultures by a cell-cell-contact-mediated mechanism. Yeast, 20, 331–341.
Ocón, E., Gutierrez, A. R., Garijo, P., Lope, Z. R., & Santamaria, P. (2010). Presence of non- Saccharomyces yeasts in cellar equipment and grape juice during harvest time. Food Microbiology, 27, 1023–1027.
Ocón, E., Garijo, P., Sanz, S., Olarte, C., Lo’pez, R., Santamaria, P., & Gutierrez, A. R. (2013). Analysis of airborne yeast in one winery over a period of one year. Food Control, 30, 585–589.
Orlandini, S., Di Stefano, V., Lucchesini, P., Puglisi, A., & Bartolini, G. (2009). Current trends of agroclimatic indices applied to grapevine in Tuscany (Central Italy). Idojaras, 113, 69–78.
Oro, L., Ciani, M., & Comitini, F. (2014). Antimicrobial activity of Metschnikowia pulcherrima on wine yeasts. Journal of Applied Microbiology, 11, 1209–1217.
Osborne, J. P., & Edwards, C. G. (2007). Inhibition of malolactic fermentation by a peptide produced by Saccharomyces cerevisiae during alcoholic fermentation. International Journal of Food Microbiology, 118, 27–34.
Padilla, B., Gil, J. V., & Manzanares, P. (2016). Past and future of non-Saccharomyces yeasts: From spoilage microorganisms to biotechnological tools for improving wine aroma complexity. Frontiers in Microbiology, 7, 411.
Pardo, J. M., & Serrano, R. (1989). Structure of a plasma membrane H+-ATPase gene from the plant Arabidopsis thaliana. The Journal of Biological Chemistry, 264, 8557–8562.
Peña, R., & Ganga, M. A. (2018). Novel antimicrobial peptides produced by Candida intermedia LAMAP1790 active against the wine-spoilage yeast Brettanomyces bruxellensis. Antonie Van Leeuwenhoek, 9, 1–8.
Pérez-Nevado, F., Albergaria, H., Hogg, T., & Girio, F. (2006). Cellular death of two non-Saccharomyces wine-related yeasts during mixed fermentations with Saccharomyces cerevisiae. International Journal of Food Microbiology, 108, 336–345.
Pérez-Torrado, R., Barrio, E., & Querol, A. (2018). Alternative yeasts for winemaking: Saccharomyces non-cerevisiae and its hybrids. Critical Reviews in Food Science and Nutrition, 58, 1780–1790.
Peter, G., Tornai-Lehoczki, J., Suzuki, M., & Dlauchy, D. (2005). Metschnikowia viticola sp. nov., a new yeast species from grape. Antonie Van Leeuwenhoek, 87, 155–160.
Petrie, P. R., & Sadras, V. O. (2008). Advancement of grapevine maturity in Australia between 1993 and 2006: putative causes, magnitude of trends and viticultural consequences. Australian Journal of Grape and Wine Research, 14, 33–45.
Peynaud, E., & Domercq, S. (1959). Possibilité de provoquer la fermentation malolactique à l’aide de bactéries cultivées. Comptes Rendus de l’Academie d’Agriculture de France, 45, 355–358.
Piao, H., Hawley, E., Kopf, S., DeScenzo, R., Sealock, S., Henick-Kling, T., & Hess, M. (2015). Insights into the bacterial community and its temporal succession during the fermentation of wine grapes. Frontiers in Microbiology, 6, 809.
Pielke, S. R., Stohlgren, T., Schell, L., Parton, W., Doesken, N., Redmond, K., & Kittel, T. G. F. (2002). Problems in evaluating regional and local trends in temperature: An example from eastern Colorado, USA. International Journal of Climatology, 22, 421–434.
Pinto, C., Pinho, D., Cardoso, R., Custódio, V., Fernandes, J., & Susana. (2015). Wine fermentation microbiome: A landscape from different Portuguese wine appellations. Frontiers in Microbiology, 6, 905.
Pretorius, I. S. (2000). Tailoring wine yeast for the new millennium: Novel approaches to the ancient art of winemaking. Yeast, 16, 675–729.
Pretorius, I. S., Van der Westhuizen, T. J., & Augustyn, O. P. H. (1999). Yeast biodiversity in vineyards and wineries and its importance to the South African wine industry. A review. South African Journal of Enology and Viticulture, 20, 61–70.
Ramos, M. C., Jones, G. V., & Martínez-Casasnovas, J. A. (2008). Structure and trends in climate parameters affecting winegrape production in Northeast Spain. Climate Research, 38(1), 1–15.
Rankine, B. C., & Pilone, D. A. (1973). Saccharomyces bailii, a resistant yeast causing serious spoilage of bottled table wine. American Journal of Enology and Viticulture, 24, 55–58.
Rantsiou, K., Dolci, P., Giacosa, S., Torchio, F., Tofalo, R., Torriani, S., Suzzi, G., Rolle, L., & Cocolin, L. (2012). Candida zemplinina can reduce acetic acid produced by Saccharomyces cerevisiae in sweet wine fermentations. Applied and Environmental Microbiology, 78(6), 1987–1994.
Raspor, P., Milek, D. M., Polanc, J., Možina, S. S., & Čadež, N. (2006). Yeasts isolated from three varieties of grapes cultivated in different locations of the Dolenjska vine-growing region, Slovenia. International Journal of Food Microbiology, 109, 97–102.
Regueiro, L. A., Costas, C. L., & Rubio, J. E. L. (1993). Influence of viticultural and enological practices on the development of yeast populations during winemaking. American Journal of Enology and Viticulture, 44, 405–408.
Renault, P., Coulon, J., de Revel, G., Barbe, J. C., & Bely, M. (2015). Increase of fruity aroma during mixed T. delbrueckii/S. cerevisiae wine fermentation is linked to specific esters enhancement. International Journal of Food Microbiology, 207, 40–48.
Renouf, V., & Lonvaud-Funel, A. (2007). Development of an enrichment medium to detect Dekkera/Brettanomyces bruxellensis, a spoilage wine yeast, on the surface of grape berries. Microbiological Research, 162, 154–167.
Riffell, S., Burton, T., & Murphy, M. (2006). Birds in depressional forested wetlands: Area and habitat requirements and model uncertainty. Wetlands, 26, 107–118.
Rodríguez, M. E., Lopes, C. A., Valles, S., Giraudo, M. R., & Caballero, A. (2007). Selection and preliminary characterization of β-glycosidases producer Patagonian wild yeasts. Enzyme and Microbial Technology, 41, 812–820.
Rodríguez-Gómez, F., Arroyo-López, F. N., López-López, A., Bautista-Gallego, J., & Garrido-Fernández, A. (2010). Lipolytic activity of the yeast species associated with the fermentation/storage phase of ripe olive processing. Food Microbiology, 27, 604–612.
Rojas, V., Gil, J. V., Piñaga, F., & Manzanares, P. (2003). Acetate ester formation in wine by mixed cultures in laboratory fermentations. International Journal of Food Microbiology, 86, 181–188.
Romano, P., Fiore, C., & Paraggio, M. (2003). Function of yeast species and strains in wine flavour. International Journal of Food Microbiology, 86, 169–180.
Rosini, G. (1984). Assessment of dominance of added yeast in wine fermentation and origin of Saccharomyces cererevisiae in wine-making. The Journal of General and Applied Microbiology, 30, 249–256.
Rosini, G., Federici, F., & Martini, A. (1982). Yeast flora of grape berries during ripening. Microbial Ecology, 8, 83–89.
Sabate, J., Cano, J., Esteve-Zarzoso, B., & Guillamon, J. M. (2002). Isolation and identification of yeasts associated with vineyard and winery by RFLP analysis of ribosomal genes and mitochondrial DNA. Microbiological Research, 157, 267–274.
Santamarıa, P., Lopez, R., Lopez, E., Garijo, P., & Gutierrez, A. R. (2008). Permanence of yeast inocula in the winery ecosystem and presence in spontaneous fermentations. European Food Research and Technology, 227, 1563–1567.
Santos, C. N. S., & Stephanopoulos, G. (2008). Combinatorial engineering of microbes for optimizing cellular phenotype. Current Opinion in Chemical Biology, 12, 168–176.
Santos, C., Lima, N., Sampaio, P., & Pais, C. (2011). Matrix-assisted laser desorption/ionization time-of-flight intact cell mass spectrometry to detect emerging pathogenic Candida species. Diagnostic Microbiology and Infectious Disease, 71, 304–308.
Schuller, D., Cardoso, F., Sousa, S., Gomes, P., Gomes, A. C., & Santos, M. A. S. (2012). Genetic diversity and population structure of Saccharomyces cerevisiae strains isolated from different grape varieties and winemaking regions. PLoS One, 7, e32507.
Shen, Y., Chen, X., Peng, B., Chen, L., Hou, J., & Bao, X. (2012). An efficient xylose-fermenting recombinant Saccharomyces cerevisiae strain obtained through adapted evolution and its global transcription profile. Applied Microbiology and Biotechnology, 96, 1079–1091.
Sipiczki, M., Pfliegler, W. P., & Holb, I. J. (2013). Species share a pool of diverse rRNA genes differing in regions that determine hairpin-loop structures and evolve by reticulation. PLoS One, 8, e67384.
Sokolowsky, M., & Fischer, U. (2012). Evaluation of bitterness in white wine applying descriptive analysis, time-intensity analysis, and temporal dominance of sensations analysis. Analytica Chimica Acta, 732, 46–52.
Steensels, J., Snoek, T., Meersman, E., Nicolino, M. P., Voordeckers, K., & Verstrepen, K. J. (2014). Improving industrial yeast strains: Exploiting natural and artificial diversity. FEMS Microbiology Reviews, 38, 947–995.
Stefanini, I., & Cavalieri, D. (2018). Metagenomic approaches to investigate the contribution of the vineyard environment to the quality of wine fermentation: Potentials and difficulties. Frontiers in Microbiology, 9.
Stefanini, I., Dapporto, L., Legras, J. L., Calabretta, A., Di Paola, M., & De Filippo, C. (2012). Role of social wasps in Saccharomyces cerevisiae ecology and evolution. Proceedings of the National Academy of Sciences, 109, 13398–13403.
Suh, S. O., Gibson, C. M., & Blackwell, M. (2004). Metschnikowia chrysoperlae sp. nov., Candida picachoensis sp. nov. and Candida pimensis sp. nov., isolated from the green lacewings Chrysoperla comanche and Chrysoperla carnea (Neuroptera: Chrysopidae). International Journal of Systematic and Evolutionary Microbiology, 54, 1883–1890.
Swiegers, J. H., Bartowsky, E. J., Henschke, P. A., & Pretorius, I. (2005). Yeast and bacterial modulation of wine aroma and flavour. Australian Journal of Grape and Wine Research, 11, 139–173.
Taylor, M. W., Tsai, P., Anfang, N., Ross, H. A., & Goddard, M. R. (2014). Pyrosequencing reveals regional differences in fruit-associated fungal communities. Environmental Microbiology, 16, 2848–2858. https://doi.org/10.1111/1462-2920.12456.
Teslić, N., Zinzani, G., Parpinello, G. P., & Versari, A. (2018). Climate change trends, grape production, and potential alcohol concentration in wine from the “Romagna Sangiovese” appellation area (Italy). Theoretical and Applied Climatology, 131, 793–803.
Tonietto, J. (1999). Les macroclimats viticoles mondiaux et l’influence du mesoclimat sur la typicite de la Syrah et du Muscat de Hamboug dans le sud de la France: Methodologie de caracterisation. Embrapa Uva e Vinho-Outras publicações científicas. (ALICE).
Ugliano, M., & Henschke, P. A. (2009). Yeasts and wine flavour. In Wine chemistry and biochemistry (pp. 313–392). New York: Springer.
Van der Walt, J. P., & van Kerken, A. E. (1961). The wine yeast sf the cape. Antonie Van Leeuwenhoek, 27, 81–90.
Van der Westhuizen, T. J., Augustyn, O. P. H., Khan, W., & Pretorius, I. S. (2000). Seasonal variation of indigenous Saccharomyces cerevisiae strains isolated from vineyards of the Western Cape in South Africa. South African Journal of Enology & Viticulture, 21(1), 10–16.
Van Leeuwen, C., Schultz, H. R., de Cortazar-Atauri, I. G., Duchêne, E., Ollat, N., Pieri, P., & Malheiro, A. C. (2013). Why climate change will not dramatically decrease viticultural suitability in main wine-producing areas by 2050. Proceedings of the National Academy of Sciences of the United States of America, 110, E3051–E3052.
Varela, C., Siebert, T., Cozzolino, D., Rose, L., McLean, H., & Henschke, P. A. (2009). Discovering a chemical basis for differentiating wines made by fermentation with ‘wild’ indigenous and inoculated yeasts: Role of yeast volatile compounds. Australian Journal of Grape and Wine Research, 15, 238–248.
Verginer, M., Leitner, E., & Berg, G. (2010). Production of volatile metabolites by grape associated microorganisms. Journal of Agricultural and Food Chemistry, 58, 8344–8350.
Viana, M., Kuhlbusch, T. A. J., Querol, X., Alastuey, A., Harrison, R. M., Hopke, P. K., & Hueglin, C. (2008). Source apportionment of particulate matter in Europe: A review of methods and results. Journal of Aerosol Science, 39, 827–849.
Williams, T. C., Averesch, N. J. H., Winter, G., Plan, M. R., Vickers, C. E., Nielsen, L. K., & Krömer, J. O. (2015). Quorum-sensing linked RNA interference for dynamic metabolic pathway control in Saccharomyces cerevisiae. Metabolic Engineering, 29, 124–134.
Winkler, A. J. (1962). General viticulture. Berkeley: University of California Press.
Zarraonaindia, I., Owens, S. M., Weisenhorn, P., West, K., Hampton-Marcell, J., Lax, S., & van der Lelie, D. (2015). The soil microbiome influences grapevine-associated microbiota. MBio, 6, e02527–e02514.
Zott, K., Miot-Sertier, C., Claisse, O., Lonvaud-Funel, A., & Masneuf-Pomarede, I. (2008). Dynamics and diversity of non-Saccharomyces yeasts during the early stages in winemaking. International Journal of Food Microbiology, 125, 197–203.
Zupan, J., Avbelj, M., Butinar, B., Kosel, J., Šergan, M., & Raspor, P. (2013). Monitoring of quorum-sensing molecules during minifermentation studies in wine yeast. Journal of Agricultural and Food Chemistry, 61, 2496–2505.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Science+Business Media, LLC, part of Springer Nature
About this chapter
Cite this chapter
Ciani, M., Comitini, F. (2019). Yeast Ecology of Wine Production. In: Romano, P., Ciani, M., Fleet, G. (eds) Yeasts in the Production of Wine. Springer, New York, NY. https://doi.org/10.1007/978-1-4939-9782-4_1
Download citation
DOI: https://doi.org/10.1007/978-1-4939-9782-4_1
Published:
Publisher Name: Springer, New York, NY
Print ISBN: 978-1-4939-9780-0
Online ISBN: 978-1-4939-9782-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)