Abstract
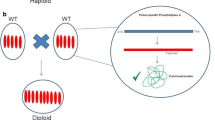
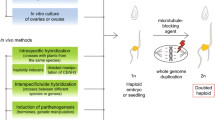
Manifold and diverse applications of doubled haploid (DH) plants have emerged in academy and in the plant breeding industry since the first discovery of a haploid mutant in the Jimson Weed (Datura stramonium), followed by the first reports about anther culture in the same species, maternal haploids by wide crosses in tobacco (Nicotiana tabacum L.) and barley (Hordeum vulgare L.), interspecific hybridization, ovary culture (gynogenesis), isolated microspore culture, and more recently the CENH3 approach in thale cress (Arabidopsis thaliana L.) and other species. Research and development efforts were and are still significant in both user groups. Luckily, often academic and industrial partners cooperate in challenging and sometimes voluminous projects worldwide. Not only to develop innovative DH protocols and technologies per se, but also to exploit the advantages of DH plants in a huge variety of research and development experiments. This review concentrates not on the DH technologies per se, but on the application of DHs in plant-related research and development projects.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Blakeslee AF, Belling J, Farnham ME, Bergner AD (1922) A haploid mutant in the Jimson Weed, “Datura stramonium”. Science 55(1433):646–647. https://doi.org/10.1126/science.55.1433.646
Guha S, Maheshwari SC (1964) In vitro production of embryos from anthers of datura. Nature 204(4957):497–497. https://doi.org/10.1038/204497a0
Burk LG, Gerstel DU, Wernsman EA (1979) Maternal haploids of Nicotiana tabacum L. from seed. Science 206(4418):585. https://doi.org/10.1126/science.206.4418.585
Kasha KJ, Kao KN (1970) High frequency haploid production in barley (Hordeum vulgare L.). Nature 225(5235):874–876. https://doi.org/10.1038/225874a0
Kalinowska K, Chamas S, Unkel K, Demidov D, Lermontova I, Dresselhaus T, Kumlehn J, Dunemann F, Houben A (2019) State-of-the-art and novel developments of in vivo haploid technologies. Theor Appl Genet 132(3):593–605. https://doi.org/10.1007/s00122-018-3261-9
Van Geyt J, Speckmann GJ Jr, D’Halluin K, Jacobs M (1987) In vitro induction of haploid plants from unpollinated ovules and ovaries of the sugarbeet (Beta vulgaris L.). Theor Appl Genet 73(6):920–925. https://doi.org/10.1007/bf00289399
Pescitelli SM, Johnson CD, Petolino JF (1990) Isolated microspore culture of maize: effects of isolation technique, reduced temperature, and sucrose level. Plant Cell Rep 8(10):628–631. https://doi.org/10.1007/bf00270070
Lichter R (1982) Induction of haploid plants from isolated pollen of Brassica napus. Z Pflanzenphysiol 105(5):427–434. https://doi.org/10.1016/S0044-328X(82)80040-8
Ravi M, Chan SW (2010) Haploid plants produced by centromere-mediated genome elimination. Nature 464(7288):615–618. https://doi.org/10.1038/nature08842
Germanà MA (2011) Gametic embryogenesis and haploid technology as valuable support to plant breeding. Plant Cell Rep 30(5):839–857. https://doi.org/10.1007/s00299-011-1061-7
L-q Y, Fu S-h, Yang J, Li Y, J-s W, M-l W (2016) Generation, identification, formation mechanism and application of plant haploids. Yi Chuan 38(11):979–991. https://doi.org/10.16288/j.yczz.16-121
Forster BP, WTB T (2005) Doubled haploids in genetics and plant breeding. Plant Breed Rev 25:57–88. https://doi.org/10.1002/9780470650301.ch3
Chang M-T, Coe EH (2009) Doubled haploids. In: Kriz AL, Larkins BA (eds) Molecular genetic approaches to maize improvement. Springer, Berlin, pp 127–142. https://doi.org/10.1007/978-3-540-68922-5_10
Choo TM (1981) Doubled haploids for studying the inheritance of quantitative characters. Genetics 99(3–4):525–540
Choo TM, Reinbergs E, Park SJ (1982) Comparison of frequency distributions of doubled haploid and single seed descent lines in barley. Theor Appl Genet 61(3):215–218. https://doi.org/10.1007/BF00273777
Powell W, Caligari PD, Swanston JS, Jinks JL (1985) Genetical investigations into β-glucan content in barley. Theor Appl Genet 71(3):461–466. https://doi.org/10.1007/BF00251188
COST
Bordes J, Charmet G, de Vaulx RD, Pollacsek M, Beckert M, Gallais A (2006) Doubled haploid versus S1 family recurrent selection for testcross performance in a maize population. Theor Appl Genet 112(6):1063–1072. https://doi.org/10.1007/s00122-006-0208-3
Sleper JA, Bernardo R (2016) Recombination and genetic variance among maize doubled haploids induced from F(1) and F(2) plants. Theor Appl Genet 129(12):2429–2436. https://doi.org/10.1007/s00122-016-2781-4
Couto EGO, Cury MN, Bandeira e Souza M, Granato ÍSC, Vidotti MS, Domingos Garbuglio D, Crossa J, Burgueño J, Fritsche-Neto R (2019) Effect of F1 and F2 generations on genetic variability and working steps of doubled haploid production in maize. PLoS One 14(11):e0224631. https://doi.org/10.1371/journal.pone.0224631
Mwathi MW, Schiessl SV, Batley J, Mason AS (2019) “Doubled-haploid” allohexaploid Brassica lines lose fertility and viability and accumulate genetic variation due to genomic instability. Chromosoma 128(4):521–532. https://doi.org/10.1007/s00412-019-00720-w
Dwivedi SL, Britt AB, Tripathi L, Sharma S, Upadhyaya HD, Ortiz R (2015) Haploids: constraints and opportunities in plant breeding. Biotechnol Adv 33(6 Pt 1):812–829. https://doi.org/10.1016/j.biotechadv.2015.07.001
Filiault DL, Seymour DK, Maruthachalam R, Maloof JN (2017) The generation of doubled haploid lines for QTL mapping. Methods Mol Biol 1610:39–57. https://doi.org/10.1007/978-1-4939-7003-2_4
Tyrka M, Oleszczuk S, Rabiza-Swider J, Wos H, Wedzony M, Zimny J, Ponitka A, Ślusarkiewicz-Jarzina A, Metzger RJ, Baenziger PS, Lukaszewski AJ (2018) Populations of doubled haploids for genetic mapping in hexaploid winter triticale. Mol Breed 38(4):46–46. https://doi.org/10.1007/s11032-018-0804-3
Werner K, Friedt W, Ordon F (2005) Strategies for pyramiding resistance genes against the barley yellow mosaic virus complex (BaMMV, BaYMV, BaYMV-2). Mol Breed 16(1):45–55. https://doi.org/10.1007/s11032-005-3445-2
Kiviharju E, Moisander S, Tanhuanpää P (2017) Oat anther culture and use of DH-lines for genetic mapping. Methods Mol Biol 1536:71–93. https://doi.org/10.1007/978-1-4939-6682-0_6
Sanchez DL, Liu S, Ibrahim R, Blanco M, Lübberstedt T (2018) Genome-wide association studies of doubled haploid exotic introgression lines for root system architecture traits in maize (Zea mays L.). Plant Sci 268:30–38. https://doi.org/10.1016/j.plantsci.2017.12.004
Lotfi M, Alan AR, Henning MJ, Jahn MM, Earle ED (2003) Production of haploid and doubled haploid plants of melon (Cucumis melo L) for use in breeding for multiple virus resistance. Plant Cell Rep 21(11):1121–1128. https://doi.org/10.1007/s00299-003-0636-3
Cegielska-Taras T, Nogala-Kałucka M, Szala L, Siger A (2016) Study of variation of tocochromanol and phytosterol contents in black and yellow seeds of Brassica napus L. doubled haploid populations. Acta Sci Pol Technol Aliment 15(3):321–332. https://doi.org/10.17306/J.AFS.2016.3.31
Siger A, Michalak M, Lembicz J, Nogala-Kałucka M, Cegielska-Taras T, Szała L (2018) Genotype × environment interaction on tocochromanol and plastochromanol-8 content in seeds of doubled haploids obtained from F1 hybrid black × yellow seeds of winter oilseed rape (Brassica napus L.). J Sci Food Agric 98(9):3263–3270. https://doi.org/10.1002/jsfa.8829
Trojak-Goluch A, Laskowska D, Kursa K (2016) Morphological and chemical characteristics of doubled haploids of flue-cured tobacco combining resistance to Thielaviopsis basicola and TSWV. Breed Sci 66(2):293–299. https://doi.org/10.1270/jsbbs.66.293
Ma H, Li G, Würschum T, Zhang Y, Zheng D, Yang X, Li J, Liu W, Yan J, Chen S (2018) Genome-wide association study of haploid male fertility in maize (Zea Mays L.). Front Plant Sci 9:974–974. https://doi.org/10.3389/fpls.2018.00974
Chaikam V, Gowda M, Nair SK, Melchinger AE, Boddupalli PM (2019) Genome-wide association study to identify genomic regions influencing spontaneous fertility in maize haploids. Euphytica 215(8):138–138. https://doi.org/10.1007/s10681-019-2459-5
Gallais A (1990) Quantitative genetics of doubled haploid populations and application to the theory of line development. Genetics 124(1):199–206
Singh S, Dey SS, Bhatia R, Kumar R, Sharma K, Behera TK (2019) Heterosis and combining ability in cytoplasmic male sterile and doubled haploid based Brassica oleracea progenies and prediction of heterosis using microsatellites. PLoS One 14(8):e0210772. https://doi.org/10.1371/journal.pone.0210772
Daetwyler HD, Hayden MJ, Spangenberg GC, Hayes BJ (2015) Selection on optimal haploid value increases genetic gain and preserves more genetic diversity relative to genomic selection. Genetics 200(4):1341. https://doi.org/10.1534/genetics.115.178038
Strigens A, Schipprack W, Reif JC, Melchinger AE (2013) Unlocking the genetic diversity of maize landraces with doubled haploids opens new avenues for breeding. PLoS One 8(2):e57234–e57234. https://doi.org/10.1371/journal.pone.0057234
Brauner PC, Müller D, Schopp P, Böhm J, Bauer E, Schön C-C, Melchinger AE (2018) Genomic prediction within and among doubled-haploid libraries from maize landraces. Genetics 210(4):1185. https://doi.org/10.1534/genetics.118.301286
Longin CFH, Utz HF, Reif JC, Schipprack W, Melchinger AE (2006) Hybrid maize breeding with doubled haploids: I. One-stage versus two-stage selection for testcross performance. Theor Appl Genet 112(5):903–912. https://doi.org/10.1007/s00122-005-0192-z
Longin CFH, Utz HF, Melchinger AE, Reif JC (2007) Hybrid maize breeding with doubled haploids: II. Optimum type and number of testers in two-stage selection for general combining ability. Theor Appl Genet 114(3):393–402. https://doi.org/10.1007/s00122-006-0422-z
Longin CFH, Utz HF, Reif JC, Wegenast T, Schipprack W, Melchinger AE (2007) Hybrid maize breeding with doubled haploids: III. Efficiency of early testing prior to doubled haploid production in two-stage selection for testcross performance. Theor Appl Genet 115(4):519–527. https://doi.org/10.1007/s00122-007-0585-2
Wegenast T, Longin CFH, Utz HF, Melchinger AE, Maurer HP, Reif JC (2008) Hybrid maize breeding with doubled haploids. IV. Number versus size of crosses and importance of parental selection in two-stage selection for testcross performance. Theor Appl Genet 117(2):251–260. https://doi.org/10.1007/s00122-008-0770-y
Wegenast T, Utz HF, Longin CFH, Maurer HP, Dhillon BS, Melchinger AE (2010) Hybrid maize breeding with doubled haploids: V. Selection strategies for testcross performance with variable sizes of crosses and S(1) families. Theor Appl Genet 120(4):699–708. https://doi.org/10.1007/s00122-009-1187-y
Marulanda JJ, Mi X, Melchinger AE, Xu J-L, Würschum T, Longin CFH (2016) Optimum breeding strategies using genomic selection for hybrid breeding in wheat, maize, rye, barley, rice and triticale. Theor Appl Genet 129(10):1901–1913. https://doi.org/10.1007/s00122-016-2748-5
Melchinger AE, Technow F, Dhillon BS (2011) Gene stacking strategies with doubled haploids derived from biparental crosses: theory and simulations assuming a finite number of loci. Theor Appl Genet 123(8):1269–1279. https://doi.org/10.1007/s00122-011-1665-x
Mi X, Wegenast T, Utz HF, Dhillon BS, Melchinger AE (2011) Best linear unbiased prediction and optimum allocation of test resources in maize breeding with doubled haploids. Theor Appl Genet 123(1):1–10. https://doi.org/10.1007/s00122-011-1561-4
Shim Y-S, Pauls KP, Kasha KJ (2009) Transformation of isolated barley (Hordeum vulgare L.) microspores: I. the influence of pretreatments and osmotic treatment on the time of DNA synthesis. Genome 52(2):166–174. https://doi.org/10.1139/g08-112
Shim Y-S, Pauls KP, Kasha KJ (2009) Transformation of isolated barley (Hordeum vulgare L.) microspores: II. Timing of pretreatment and temperatures relative to results of bombardment. Genome 52(2):175–190. https://doi.org/10.1139/g08-113
Chauhan H, Khurana P (2011) Use of doubled haploid technology for development of stable drought tolerant bread wheat (Triticum aestivum L.) transgenics. Plant Biotechnol J 9(3):408–417. https://doi.org/10.1111/j.1467-7652.2010.00561.x
Otto I, Müller A, Kumlehn J (2015) Barley (Hordeum vulgare L.) transformation using embryogenic pollen cultures. Methods Mol Biol 1223:85–99. https://doi.org/10.1007/978-1-4939-1695-5_7
Rustgi S, Ankrah NO, Brew-Appiah RAT, Sun Y, Liu W, von Wettstein D (2017) Doubled haploid transgenic wheat lines by microspore transformation. Methods Mol Biol 1679:213–234. https://doi.org/10.1007/978-1-4939-7337-8_13
Lee SY, Cheong JI, Kim TS (2003) Production of doubled haploids through anther culture of M1 rice plants derived from mutagenized fertilized egg cells. Plant Cell Rep 22(3):218–223. https://doi.org/10.1007/s00299-003-0663-0
Lu Y, Dai S, Gu A, Liu M, Wang Y, Luo S, Zhao Y, Wang S, Xuan S, Chen X, Li X, Bonnema G, Zhao J, Shen S (2016) Microspore induced doubled haploids production from ethyl methanesulfonate (EMS) soaked flower buds is an efficient strategy for mutagenesis in Chinese cabbage. Front Plant Sci 7:1780–1780. https://doi.org/10.3389/fpls.2016.01780
Ferrie AMR, Bhowmik P, Rajagopalan N, Kagale S (2020) CRISPR/Cas9-mediated targeted mutagenesis in wheat doubled haploids. Methods Mol Biol 2072:183–198. https://doi.org/10.1007/978-1-4939-9865-4_15
Kelliher T, Starr D, Su X, Tang G, Chen Z, Carter J, Wittich PE, Dong S, Green J, Burch E, McCuiston J, Gu W, Sun Y, Strebe T, Roberts J, Bate NJ, Que Q (2019) One-step genome editing of elite crop germplasm during haploid induction. Nat Biotechnol 37(3):287–292. https://doi.org/10.1038/s41587-019-0038-x
Wang B, Zhu L, Zhao B, Zhao Y, Xie Y, Zheng Z, Li Y, Sun J, Wang H (2019) Development of a haploid-inducer mediated genome editing system for accelerating maize breeding. Mol Plant 12(4):597–602. https://doi.org/10.1016/j.molp.2019.03.006
Machczyńska J, Zimny J, Bednarek PT (2015) Tissue culture-induced genetic and epigenetic variation in triticale (× Triticosecale spp. Wittmack ex A. Camus 1927) regenerants. Plant Mol Biol 89(3):279–292. https://doi.org/10.1007/s11103-015-0368-0
Wang H, Dong B, Jiang J, Fang W, Guan Z, Liao Y, Chen S, Chen F (2014) Characterization of in vitro haploid and doubled haploid Chrysanthemum morifolium plants via unfertilized ovule culture for phenotypical traits and DNA methylation pattern. Front Plant Sci 5:738–738. https://doi.org/10.3389/fpls.2014.00738
Li H, Soriano M, Cordewener J et al (2014) The histone deacetylase inhibitor trichostatin a promotes totipotency in the male gametophyte. Plant Cell 26(1):195–209. https://doi.org/10.1105/tpc.113.116491
Dirks R, van Dun K, de Snoo CB, van den Berg M, Lelivelt CLC, Voermans W, Woudenberg L, de Wit JPC, Reinink K, Schut JW, van der Zeeuw E, Vogelaar A, Freymark G, Gutteling EW, Keppel MN, van Drongelen P, Kieny M, Ellul P, Touraev A, Ma H, de Jong H, Wijnker E (2009) Reverse breeding: a novel breeding approach based on engineered meiosis. Plant Biotechnol J 7(9):837–845. https://doi.org/10.1111/j.1467-7652.2009.00450.x
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Weyen, J. (2021). Applications of Doubled Haploids in Plant Breeding and Applied Research. In: Segui-Simarro, J.M. (eds) Doubled Haploid Technology. Methods in Molecular Biology, vol 2287. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-1315-3_2
Download citation
DOI: https://doi.org/10.1007/978-1-0716-1315-3_2
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-1314-6
Online ISBN: 978-1-0716-1315-3
eBook Packages: Springer Protocols