Abstract
Microspore culture stimulates immature pollen grains to develop into plants via tissue culture and is used routinely in many crop species to produce “doubled haploids”: homozygous, true-breeding lines. However, microspore culture is also often used on material that does not have stable meiosis, such as interspecific hybrids. In this case, the resulting progeny may lose their “doubled haploid” homozygous status as a result of chromosome missegregation and homoeologous exchanges. However, little is known about the frequency of these effects. We assessed fertility, meiosis and genetic variability in self-pollinated progeny sets (the MDL2 population) resulting from first-generation plants (the MDL1 population) derived from microspores of a near-allohexaploid interspecific hybrid from the cross (Brassica napus × B. carinata) × B. juncea. Allelic inheritance and copy number variation were predicted using single nucleotide polymorphism marker data from the Illumina Infinium 60K Brassica array. Seed fertility and viability decreased substantially from the MDL1 to the MDL2 generation. In the MDL2 population, 87% of individuals differed genetically from their MDL1 parent. These genetic differences resulted from novel homoeologous exchanges between chromosomes, chromosome loss and gain, and segregation and instability of pre-existing karyotype abnormalities. Novel karyotype change was extremely common, with 2.2 new variants observed per MDL2 individual. Significant differences between progeny sets in the number of novel genetic variants were also observed. Meiotic instability clearly has the potential to dramatically change karyotypes (often without detectable effects on the presence or absence of alleles) in putatively homozygous, microspore-derived lines, resulting in loss of fertility and viability.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Interspecific hybridization is a key factor in speciation, and has resulted in the formation of diverse agricultural species (Abbott et al. 2013). Although interspecific hybridization can produce new species without chromosome doubling, particularly in animals (Mallet 2007), hybridization in plants is more commonly coupled with polyploidy, such that resulting “allopolyploid” species contain a full set of chromosomes from both parent species (Leitch and Leitch 2008). Successful allopolyploid crops include bread wheat (Triticum aestivum), sisal (Agave sisalana), coffee (Coffea arabica), cotton (Gossypium hirsutum), sugarcane (Saccharum spp.) and oats (Avena sativa) (Leitch and Leitch 2008). Newly formed species may benefit from increased heterozygosity, novel phenotypic variation and better adaptation to new environmental niches (Comai 2005; Udall and Wendel 2006). However, many new interspecific hybrids are genomically unstable, undergoing frequent non-homologous interactions between chromosomes from different subgenomes during meiosis, leading to chromosome loss, instability and infertility (Gaeta and Pires 2010; Song et al. 1995; Szadkowski et al. 2010). Regulation of meiotic stability in newly formed allopolyploids is a key factor affecting whether an allopolyploid speciation event will be successful (Péle et al. 2018).
The Brassica genus contains many economically important crops, such as vegetables, oilseeds and condiments (cabbage, cauliflower, broccoli, pok choi, canola and mustards). The interrelationships between Brassica species evident in “U’s Triangle” (U N, 1935) are an example of natural allopolyploidy events that have generated new species and contributed to differentiation within a genus: the three diploid species Brassica rapa, Brassica nigra and Brassica. oleracea have genome complements of 2n = AA, BB and CC respectively, and formed natural allotetraploids Brassica juncea, Brassica napus and Brassica carinata with 2n = AABB, 2n = AACC and 2n = BBCC, respectively, through pairwise hybridization. Although the combination of the A, B and C genomes in one Brassica species does not occur naturally, a new allohexaploid Brassica (2n = AABBCC) may have potential for increased hybrid vigour and adaptation (Chen et al. 2011). However, a meiotically stable allohexaploid Brassica has yet to become a possibility in agriculture (Chen et al. 2011; Mason and Batley 2015). Most attempts to develop stable allohexaploid Brassica have been largely unsuccessful, with aberrant meiosis and the frequent occurrence of aneuploid plants in the selfed progenies (Iwasa 1964; Zhou et al. 2016). Despite this, some allohexaploids synthesized from the cross B. carinata × B. rapa have shown an increase in meiotic stability and higher proportions of euploid progeny with successive self-pollination and selection in each generation (Tian et al. 2010), while others appear immediately stable upon synthesis (Gupta et al. 2016). Variation for meiotic stability in this material appears strongly genotype-specific: differences between genotypes were also observed for the cross combination (B. napus × B. carinata) × B. juncea (Mwathi et al. 2017).
Microspore culture is utilized widely in many crop species, including Brassica oilseeds, to generate haploid and doubled haploid (DH) homozygous lines and germplasm for breeding and genetic analysis (Jahne and Lorz 1995; Takahira et al. 2011). Haploid plants produced from an F1 combine the two parental genomes, and have only one allele at every locus. Converting sterile haploids into fertile doubled haploids produces pure-breeding homozygous lines which are useful in fixing traits quickly in desirable combinations in a variety (Seymour et al. 2012). Microspore culture has also previously been applied to interspecific hybrids in order to more rapidly fix desirable traits, or to provide a fixed resource for future breeding applications (Geng et al. 2013; Navabi et al. 2010; Yang et al. 2018). However, such approaches may result in unexpected results. One example of this is the strong selective pressures for particular gamete types observed in microspore culture of different interspecific Brassica hybrids (e.g. (Li et al. 2018; Mason et al. 2011; Nelson et al. 2009)). In this study, we aimed to investigate another possibility: that unstable meiosis in lines derived from interspecific hybridisation will result in genetic variation within putatively “fixed” microspore-derived lines. Secondarily, we also aimed to determine if fixed, unbalanced homoeologous exchanges, where a part of a chromosome from one subgenome has been replaced by part of a chromosome from a homoeologous subgenome, are likely to precipitate further genomic instability (Fig. 1). For the purposes of this investigation, we grew up a second generation (MDL2) from seed produced by previously produced microspore-derived lines (MDL1; described in Mason et al. (2014b) and Mason et al. (2015) and known to be meiotically unstable) from a near-allohexaploid hybrid from the cross combination (B. napus × B. carinata) × B. juncea (Mason et al. 2012).
The putative role of microspore culture (with induced chromosome doubling to produce “doubled haploid” lines) in fixing translocations between chromosomes in an unstable allopolyploid. Question marks indicate the unknown chance of non-homologous recombination events occurring during meiosis in microspore-derived progeny. * “unbalanced” translocation event or homoeologous exchange; often referred to as “homoeologous non-reciprocal translocation” or HNRT. # “balanced” translocation event or homoeologous exchange, often referred to as a “homoeologous reciprocal translocation” or HRT. Putatively, “unbalanced” translocations may lead to multivalent formation during meiosis due to the presence of homology between four different chromosomes, subsequently increasing the chance of further karyotype rearrangements or abnormal chromosome segregation events
Materials and methods
Experimental material
The experimental material comprised self-pollinated seeds produced by first-generation microspore-derived plants from a parent near-allohexaploid plant derived from the cross (B. napus × B. carinata) × B. juncea (Fig. 2). These lines were predicted to have unstable meiosis to varying degrees. The production of the parent near-allohexaploid is described in Mason et al. (2012), while the MDL1 population was produced and previously analysed in Mason et al. (2014b) and Mason et al. (2015); the SP1 population of self-pollinated lines used for comparative purposes in this study is described in Mason et al. (2014b). Fourteen microspore-derived plants from the MDL1 population produced more than ten seeds (12–198 seeds); these “MDL2” seeds comprised the experimental population. A total of 6–26 MDL2 seeds per MDL1 were sown, with 2–15 seeds per line sown at each of four different timepoints between March 2015 and July 2016, dictated by survival rates. Seeds from each of the parent genotypes B. napus “Surpass400_024DH”, B. carinata “195923.3.2_01DH” and B. juncea “JN9-04” were planted as controls. Germination and plant growth were carried out under temperature-controlled glasshouse conditions (20–22 °C) at the University of Western Australia, Perth.
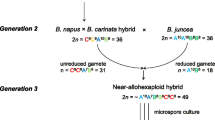
Hybridization between Brassica napus and B. carinata to produce an interspecific F1 hybrid with the genome composition CCAB, followed by crossing of this hybrid to B. juncea (2n = AABB) to produce a near-allohexaploid hybrid via union of an unreduced gamete (n = CCAB) from the interspecific hybrid and a reduced gamete (n = AB) from B. juncea, followed by microspore culture of the near-allohexaploid hybrid (2n = ~ AABBCC) to produce 75 microspore-derived lines (MDL1 population; as described in Mason et al. 2015), then self-pollination of these lines to produce a second generation (MDL2 population)
Pollen fertility, seed set and meiotic chromosome observations
Pollen grains were assessed by squashing anthers in 1% acetocarmine on glass microscope slides. Observations were carried out using a compound light microscope. Pollen grains that were plump and darkly stained were considered viable, whereas unstained and shrivelled pollen were considered non-viable. A total of 300 pollen grains were scored from at least three flowers from each plant. Seed set was also determined at the end of the growing season, after enclosing racemes in pollination bags to encourage self-pollination. Round, filled seeds, including seeds which had germinated prematurely in siliques, were counted in order to estimate self-pollinated seed fertility.
Floral buds were collected during the early morning and fixed in Carnoy’s II solution (ethanol: chloroform: acetic acid 6:3:1) for up to 48 h and subsequently stored in 70% ethanol at 4 °C. Preparation of the anthers involved staining followed by squashing in a drop of 1% acetic acid carmine solution on glass microscope slides. Observations of the pollen mother cells (PMCs) were carried out at metaphase I and II stages of meiosis using a ZEISS light microscope (bright field phase) and images captured using the Axio Vision Imaging system (Release 4.8.1). Meiosis was observed for progeny in the MDL28 and MDL60 progeny sets (10–35 pollen mother cells (PMCs) per plant, average 20), although clear, good quality meiotic slides were difficult to obtain.
DNA extraction and SNP genotyping
The DNeasy® Plant Mini Kit (QIAGEN©) was used for DNA isolation, according to the manufacturer’s instructions. DNA was tested for quality and quantity using comparison to a known size ladder (Thermo Scientific GeneRuler 1 kb DNA Ladder) via 1% agarose gel electrophoresis (1 × TBE buffer) with ethidium bromide staining and through DNA Qubit quantification. Illumina Infinium Brassica 60K SNP array (Illumina Inc., San Diego, USA) genotyping was carried out according to the manufacturer’s instructions.
SNP array data was visualized using Illumina Genome Studio, and analysed according to methods detailed in Mason et al. (2015) and Mason et al. (2017). Markers were clustered based on the publicly available cluster file for the B. napus genome (Clarke et al. 2016) and mapped to the Darmor v8.1 B. napus reference genome (Bayer et al. 2017). SNP markers were selected on the basis of known map positions on the reference genome and polymorphism between the two species parents for each genome (B. napus and B. juncea for the A genome, B. napus and B. carinata for the C genome), with no allele amplification permitted in the A genome from the B. carinata parent and no allele amplification permitted in the C genome from the B. juncea parent. Markers which presented with haplotype calls that did not match their predicted genomic position, or which presented with an excess of heterozygote or null calls relative to expectations, were manually removed. Finally, 7666 markers were selected for analysis: 2971 in the A genome and 4695 in the C genome (Supplementary Table 1).
Copy number variation plots (Supplementary Fig. 1) were generated in R v3.4.3 (R Core Team 2017) based on Log R ratio data for each SNP (Supplementary Table 2) as exported from Illumina Genome Studio (see Mason et al. (2015)) for a detailed description of this method). SNPs with a log R ratio of greater than 0 suggest a higher copy number, while SNPs with a log R ratio of less than 0 suggest a lower copy number. Genomic regions with higher average log R ratios putatively indicate that one or more additional copies of this region are present, while genomic regions with lower average log R ratios putatively indicate one copy of this region is absent (Supplementary Table 2). Deletions of both copies of a genomic region result in more extreme negative log R ratios (less than − 2) and are otherwise identifiable through “no calls” (NCs) in genotype call data (Supplementary Table 1, Supplementary Table 2).
Data analysis and visualization
The aov() and TukeyHSD() functions in base R v3.4.3 (R_Core_Team 2017) were used to assess if significant differences between progeny sets for pollen fertility % and seed set were present using one-way analysis of variance (ANOVA) and Tukey’s Honest Significant Differences test (p < 0.05). R functions hist() and boxplot() were used to generate summary figures for fertility data. Barcharts and other figures were generated using Microsoft Office 2010 (Excel and Powerpoint) (Microsoft Corporation).
Results
Germination, survival and fertility
A total of 250 experimental second-generation microspore-derived line (MDL2) seeds were planted from 14 different first-generation microspore-derived line (MDL1) parents, with 6–26 seeds planted per MDL. Of these 250 seeds, only 95 germinated (38%), and 75 survived to flowering (30%), with an attack of Alternaria blight in one sowing trial accounting for the majority of the difference between germination and survival rates overall. Finally, 42 plants from six lines which germinated at similar timepoints in disease-free environments were genotyped and characterized, of which three were eliminated on the basis of out-pollination based on the genotyping data to leave a total of 39 individuals in the final experimental population (9, 5, 10, 1, 10 and 4 plants from MDL07, MDL23, MDL28, MDL30, MDL60, and MDL64, respectively).
In each of the six lines, self-pollinated seed set was lower on average in the MDL2 progeny generation relative to their MDL1 parent generation; only a single MDL2 plant exceeded its MDL1 parent seed fertility (Fig. 3). Estimated pollen viability was more variable, with MDL1 and MDL2 showing similar results for most progeny sets (Supplementary Fig. 2) with the exception of the most seed-fertile parent MDL07, which showed 73% pollen viability but whose MDL2 progeny showed only 5–25% pollen viability (average 15%). MDL2 progeny sets differed significantly in pollen fertility (p < 0.0001, one-way ANOVA) but not in self-pollinated seed set (p > 0.1, one-way ANOVA). Multivalents and univalents were frequently observed in the MDL2 progeny, indicative of unstable meiosis (Fig. 4).
Genetic variation observed within microspore-derived lines
Despite the fact that microspore-derived lines are not expected to show any genetic variation either between individuals within a line or from one generation to the next, 87% of individuals in our study (34/39 plants) showed at least one genetic difference when compared to their parents, and 100% showed some sort of karyotypic abnormality as assessed by loss or gain of partial or whole chromosomes (Supplementary Table 3). Such events were divided into (a) novel events occurring between the MDL1 and MDL2 generations (i.e. in the MDL1 meiosis), (b) inheritance of abnormalities present in the MDL1 generation to the MDL2 generation without change and (c) segregation of pre-existing chromosomal abnormalities, where MDL2 progeny show different karyotypes to their parent MDL1, but as a result of a pre-existing abnormality in the parent MDL1 (Fig. 5, Supplementary Table 3). The vast majority of events could not be detected on the basis of allele presence/absence results (Supplementary Table 1), but were revealed based on copy number analysis (e.g. presence of a homozygous chromosome or chromosome fragment in one, three or four copies; Supplementary Tables 2 and 3).
Graphed Log R Ratios (from − 1.5 to 1.5) output from Illumina Genome Studio (normalized) for the Illumina Infinium Brassica 60 K SNP array SNPs used to genotype and derive copy-number data for alleles in the Brassica A and C genomes on plant MDL60, which was cultured from a single microspore of a (Brassica napus × B. carinata) × B. juncea allohexaploid hybrid, and for three of its self-pollinated progeny (MDL60_01, MDL60_02 and MDL60_03). Lower than expected allele intensity is shown in orange, and indicates chromosomes and chromosome segments present in only a single copy, whereas higher than expected allele intensity is shown in light blue, and indicates chromosomes and chromosome segments present in three or four copies, while dark blue indicates regions putatively present in the expected two copies
Novel karyotypic abnormalities involving deletion and duplication of whole chromosomes and chromosome segments were extremely prevalent in the MDL2 population: a total of 86 different instances were observed, on 17/19 chromosomes assessed (Fig. 6). This was more than twice as common as genetic changes caused by pre-existing karyotype abnormalities (32 events). A further 135 events of “stable” inheritance of karyotype abnormalities from the MDL1 to the MDL2 were observed, but the majority of these (101/135, 75%) involved the absence of two copies of a particular chromosome or chromosome fragment in the MDL1 parent, which is a fixed event unable to segregate in the progeny (Fig. 7). Both duplication and deletion events involving one chromosome copy were frequently observed (Fig. 7), although novel events involving loss of both chromosome copies either in whole or in part were rare. Significant differences between progeny sets (one-way ANOVA, p = 0.02) were observed in the accumulation of novel karyotypic changes from the MDL1 to the MDL2 generation (Supplementary Fig. 3).
Genome instability from one generation to the next in a population of 39 microspore-derived lines derived from a (Brassica napus × B. carinata) × B. juncea allohexaploid hybrid: presence of novel, inherited, and inherited but segregating (where progeny show different genotypes to their parent, but as a result of a pre-existing abnormality in the parent) deletions or duplications of chromosomes or chromosome segments in the A and C genomes
Homoeologous exchanges and segregation of chromosome abnormalities
Of the 86 putatively new duplications and deletions observed, only 59 (69%) were clear homoeologous exchanges, such that loss of a particular chromosome or chromosomal region was compensated by the presence of an additional chromosome or chromosomal region from a homoeologous chromosome. Excluding instances where the parent plant was missing both copies of a chromosome, two-thirds of events (47/66) involving segregation and inheritance of pre-existing karyotype abnormalities resulted in plants with homoeologous exchanges between primary A- and C-genome homoeologues, as expected. However, the presence and segregation of balanced, undetectable A-C translocations in the parent plant (where an A genome segment and a C genome segment have switched genomic locations, without any corresponding gain or loss of chromosome fragments) could not be ruled out for most events. For instance, eight out of ten individuals in progeny set MDL60 had a novel copy number variant (frequently without detectable homoeologous exchange) present for one or more of homoeologous chromosomes A9, A10 and C9, despite the absence of an observable copy number variant or translocation in the MDL60 parent plant. This suggests the presence of balanced translocations in the MDL60 plant involving these chromosomes that we were unable to observe. Segregation of homoeologous exchanges from the parents also appeared random for some events (e.g. A01/C1 fragments in population MDL28), but biased for others (e.g. inheritance of A10/C09 fragments in population MDL23), but due to the small size of the population and complexity of the possible parent karyotypes, these could not be validated statistically.
Comparisons between MDL2 and pre-existing parent microspore-derived lines and self-pollinated plants from the same allohexaploid parent plant
Due to the availability of previously generated SNP data (Mason et al. 2014b) for both the parent microspore-derived population (68 plants in addition to the six MDL1 parents of the MDL2) and a set of self-pollinated progeny (SP1) from the same parent allohexaploid (53 plants with high-quality data; SP_054, SP058, SP_069, SP_070, SP_072, SP_076, SP_079, SP_083, SP_087, SP_093, SP_116, SP_125 and SP_128 were excluded after visualization revealed high noise levels/poor data quality for these lines, see Supplementary Fig. 1), copy number analysis was also performed to score how often novel karyotypic variants arose in each population type. Once regions of pre-existing translocation, duplication or deletion were excluded from the analysis (from 27 to 28 Mbp on A05, 20.4–28.8 Mbp on A07, 10–20 Mbp on A10, 57.6–61 Mbp on C04, 50.6–52.1 Mbp on C05 and 29.9–43 Mbp on C06, plus single-copy chromosomes A02, A06 and A09), and after correction for the number of meiotic events being observed from the progeny (two for self-pollinated plants; one for first-generation microspore-derived lines), no significant differences were observed in the frequency of novel duplication or deletion events of part or whole chromosomes between the three population types (Fig. 8, Fig. 9).
Estimated percentage of gametes with a novel copy number variant (excluding any pre-existing translocations, duplications or deletions) on each chromosome as assessed from self-pollinated plants (SP1; 53 plants), microspore-derived plants (MDL1; 68 plants) and self-pollinated microspore-derived plants (MDL2; 39 plants) from the same allohexaploid hybrid parent plant resulting from the cross (Brassica napus × B. carinata) × B. juncea
Frequency of novel variants (excluding any pre-existing translocations, duplications or deletions) arising in the A and C genomes of self-pollinated plants (SP1; 53 plants), microspore-derived plants (MDL1; 68 plants) and self-pollinated microspore-derived plants (MDL2; 39 plants) from the same allohexaploid hybrid parent plant resulting from the cross (Brassica napus × B. carinata) × B. juncea. SP1 and MDL2 plants represent the product of two meioses from the parent allohexaploid, while MDL1 plants represent the products of a single meiosis from the parent allohexaploid
Interestingly, some differences in generation of novel genetic variation were observed for particular chromosomes between populations (Fig. 8). For instance, no novel genetic variants were observed for chromosomes C02 and A08 in the MDL2 population, despite the high frequency of novel variants observed for C02 in the SP1 and MDL1 populations. In addition, the frequency of novel variants observed for homoeologous chromosomes A10 and C9 was higher in the MDL2 population, and the MDL1 showed a lower frequency of A07 and C6 variants than the SP1 and MDL2 (Fig. 8). The SP1 population also appeared to have more frequent loss and gain of whole chromosomes than the MDL2 population (Fig. 9).
Fertility and karyotype
There was no clear relationship between the number of non-reciprocal translocations, chromosome loss and duplication events or whether karyotype rearrangements detected were balanced or unbalanced and the fertility of individual MDL2 plants. The plant (MDL60_07) with the highest self-pollinated seed set (104 seeds) and pollen viability (90%) had an additional copy of part of C9 and a missing copy of part of A10 in addition to the inherited loss of chromosome A02, and a balanced translocation event (A03/C3) partially inherited from the parent plant, but larger in size than the event present in the parent plant and other MDL60 progeny. Of the five individuals with putatively unchanged karyotypes relative to their parent MDL, three set no seed, one set 19 seeds and one set 51 seeds (Supplementary Table 3).
Discussion
In this study, we assessed genetic identity, fertility and meiotic behaviour in two generations of microspore-derived allohexaploid lines (MDL1 and MDL2) in order to test whether (1) unstable meiosis produces genetic variation in these putatively homozygous lines and (2) whether pre-existing unbalanced homoeologous translocations precipitate further genomic instability. Unstable meiosis most definitely did result in novel genetic variation: 87% of MDL2 lines in our study were genetically different from their MDL1 parent. Pre-existing unbalanced homoeologous translocations were also found to precipitate further genomic instability, but surprisingly this effect was actually less (0.8 events per plant on average) than the spontaneous generation of novel genetic variants due to unstable meiosis (2.2 events per plant on average). Use of microspore culture on interspecific hybrids or lines with putatively unstable meiosis is often undertaken to fix desirable traits or karyotypes (e.g. Geng et al. 2013; Mason et al. 2015; Navabi et al. 2010; Yang et al. 2018). This approach may still show significant benefits in revealing phenotypic variation or even fixing lines with high meiotic stability in large populations segregating for genetic control of meiosis. Also, novel genetic variation generated through chromosome rearrangements may not always be bad: genomic rearrangements due to homoeology between the A and C genomes are seen frequently in Brassica interspecific hybrids (Nicolas et al. 2007; Nicolas et al. 2012), where they can become an important source of genetic and phenotypic variation (Stein et al. 2017). Additionally, copy number variants are now known to play a greater role in crop plant traits and resistances than previously expected (Dolatabadian et al. 2017; Schiessl et al. 2018). Despite this, the presence of genetic variation within MDLs is almost always undesirable, as disruption of homozygosity defeats the point of producing the MDLs in the first place. Therefore, careful consideration should be taken when producing MDLs from potentially unstable hybrid lines, and extreme caution should be undertaken in assuming homozygosity or “true-breeding” karyotypes in this material.
An average of 2.2 completely new karyotype changes per plant were observed in the MDL2, ranging from 0 to 9 events per plant. Similar numbers of novel karyotype changes were observed in self-pollinated progeny as well as the first-generation microspore-derived lines from the same parent allohexaploid plant, suggesting that the main cause of these novel variants was abnormal meiosis rather than a microspore culture-specific effect. In further support of homoeologous chromosome interactions as the primary mechanism for generation of novel genetic variants in this material, no novel variants were observed for chromosome C2 in the MDL2 population, in which almost all individuals were missing a copy of primary homoeologue A02. Novel variants on chromosome C2 were otherwise extremely frequent in the MDL1 and SP1 populations, for which the parent plant contained one copy of chromosome A02. Although chromosome A08 also showed no novel genetic variants in the MDL2 population, only low frequencies of novel variants were also observed for A08 in the MDL1 and SP1 populations, thus reflecting the low level of homoeologous pairing generally observed for this chromosome and for chromosome C7 (Mason et al. 2014a). A further 0.8 changes per MDL2 plant were caused by segregation or instability of a pre-existing chromosome rearrangement event in the parent MDL1 plant, in addition to 3.5 karyotype abnormalities on average directly inherited from the parent. Our secondary hypothesis that fixed non-reciprocal translocation events in the parent MDL1 would lead to genomic instability in MDL2 progeny containing these events was borne out by our results. However, novel karyotype rearrangements played a much greater role than inherited variants in generating genetic variation from the MDL1 to the MDL2 generation.
Due to the inability of our methods to pick up balanced homoeologous exchanges, or the putatively rarer events involving the more distantly related B genome (Lagercrantz and Lydiate 1996; Mason et al. 2015; Navabi et al. 2011), the numbers of segregating chromosome rearrangement events detected in our study are undoubtedly underestimated. This is also supported by the observation that “novel” variants in the MDL2 population were suspiciously frequent on particular homoeologous chromosomes (e.g. A10/C9 and A07/C6), suggesting the possible presence of balanced homoeologous exchanges in the previous generation parent/s. The presence of mitotic instability in these lines is also possible and may result in novel karyotypic changes arising which are subsequently transmitted through the germline (see Schubert and Lysak (2011) for review). As our experimental population is the product of multiple meioses (Fig. 2), we cannot assess the probability of mitotic instability in this material. Regardless, based on our results, continuous accumulation of genetic variation from one generation to the next is predicted in this material until fertility and viability drop to zero. With strong selection for fertility in large enough populations, it is possible that particular karyotype changes could occur that restore genomic stability: one plant in our population did appear to “recover” a higher degree of fertility relative to its siblings and parent after novel translocation events occurred. Despite this, we suggest that the generally low fertility and viability of these microspore-derived lines is an argument against the induction of doubled-haploids in the early generations of highly unstable interspecific hybrid material.
Only 69% of novel karyotypic changes observed in our MDL2 population appeared to be symmetrical i.e. where a duplication/deletion event is present between regions of primary homoeology in the Brassica A and C genomes, such that the plant retains the same number of copies of the homoeologous region (4A + 0C or 0C + 4A for one region). This is unexpected under normal chromosome segregation and behaviour models for a single non-homologous crossover event between primary homoeologues: 100% of gametes should retain the same number of copies of homoeologous regions under such a scenario (Nicolas et al. 2007), and strong selection for this effect is also expected (Chester et al. 2013; Xiong et al. 2011). In the case of deletions without a visible duplication on a corresponding homoeologue, it is possible that these represent real losses of DNA sequence, as has been reported at low frequencies in synthetic B. napus (Samans et al. 2017). However, it is hard to see how an additional chromosome fragment could be present in these lines unless it also retains a centromere, which is putatively not the case for most duplicated fragments. It is possible that A-B or B-C chromosome pairing and autosyndesis within genomes are responsible for the frequent observation of duplications without deletion events and vice versa in our lines, such that the homoeologous exchange partners are not being detected by our analysis. As an argument against this, the vast majority of non-homologous chromosome interactions in Brassica hybrids are predicted to occur between primary A-C homoeologues, based on previous results (Mason et al. 2010, 2014a, 2015), and our results are similar to observations in unstable synthetic B. napus (2n = AACC; Nicolas et al. (2007)), which has no B genome. Pairing between secondary homoeologues, usually via chromosome pairing within genomes (autosyndesis), also comprises less than 10% of non-homologous chromosome interactions in various hybrid types containing the A and C genomes (Mason et al. 2010, 2014a; Nicolas et al. 2007). Regardless, it is possible that previous studies underestimated frequencies of homeologous exchanges with the B genome, and this possibility should be investigated in future research. Potentially, chromosome pairing disruptions in the form of high frequencies of multivalents and univalents during meiosis (as we observed; see Fig. 3) could also be at least partly responsible for this result. Such chromosome configurations allow for loss or gain of chromosome fragments and whole chromosomes through mis-segregation of chromosomes (e.g. a 3:1 instead of 2:2 segregation of chromosomes into daughter cells) and promote meiotic phenomena such as chromosome laggards which can result in mechanistic loss of chromosomes without non-homologous recombination events taking place (Bretagnolle and Thompson 1995; De Storme and Geelen 2013; De Storme and Mason 2014). Loss or gain of a single chromosome was more common in SP1 than in MDL2 populations (Fig. 9), which may indicate differences between heterozygous (SP1) and homozygous (MDL2) lines in the frequency of crossover formation and formation of univalents if no crossovers are produced. Although this is highly speculative based on our results and requires further validation, Ziolkowski et al. (2015) also observed genotype-specific differences in crossover frequency between heterozygotes and homozygotes in Arabidopsis, supporting this interpretation.
Overall, lower fertility was observed in MDL2 progeny compared to their MDL1 parents based on seed set data, and seed viability in the MDL2 generation was extremely poor. Pollen viability did not show the same trend of decrease from the MDL1 parents to the MDL2 progeny, but this is likely a result of greater tolerance of karyotype abnormalities in the production of viable pollen than in the more complex processes of self-fertilization, embryo development and seed production. The reduced seed fertility of the MDL2 relative to their MDL1 parents supports the idea that homeologous recombination occurring due to polyploidy can destabilize genomes, leading to additional future translocations and thus increasing generational instability (Gaeta and Pires 2010). However, this could also be due to an increasing chance that lethal chromosome rearrangements, duplication or deletion events will occur with increasing numbers of meioses in our study, since we also observed a very high frequency of novel karyotype changes in our MDL2 plants. “Abnormal” or otherwise maladaptive chromosome karyotypes that could normally be eliminated by self-fertilization processes may also be fixed through the microspore culture process. As well, novel chromosome rearrangements when present in only one of two gametes in a self-fertilization event will result in a translocation heterozygote, which may produce progeny without the translocation. Loss of the translocation may also be selected for in the progeny of the translocation heterozygote, resulting in more normal karyotypes through self-fertilization processes. Studies on allohexaploids from B. carinata × B. rapa crosses found mixed results for intergenerational fertility: Howard (1942) observed fertility to increase over several generations, while Iwasa (1964) found fertility to be decreasing across generations. Mwathi et al. (2017) also observed decreased fertility and chromosome loss in B. carinata × B. rapa crosses in the second generation, with variable fertility in heterozygous second-generation allohexaploids from the cross (B. napus × B. carinata) × B. juncea. A study on allohexaploids generated from different combinations found that B. carinata × B. rapa and B. juncea × B. oleracea crosses had higher fertility compared to allohexaploids arising either from natural and synthetic B. napus and B. nigra or from the sequential cross between B. rapa, B. oleracea and B. nigra (Zhou et al. 2016). From the results of these studies, generational fertility and genomic stability in allohexaploid Brassica are very likely to be controlled by particular allelic variants, which would also explain the significant variation in fertility and meiotic stability (accumulation of novel karyotype rearrangements) observed between MDL2 lines in the present study.
References
Abbott R, Albach D, Ansell S, Arntzen JW, Baird SJE, Bierne N, Boughman JW, Brelsford A, Buerkle CA, Buggs R, Butlin RK, Dieckmann U, Eroukhmanoff F, Grill A, Cahan SH, Hermansen JS, Hewitt G, Hudson AG, Jiggins C, Jones J, Keller B, Marczewski T, Mallet J, Martinez-Rodriguez P, Most M, Mullen S, Nichols R, Nolte AW, Parisod C, Pfennig K, Rice AM, Ritchie MG, Seifert B, Smadja CM, Stelkens R, Szymura JM, Vainola R, Wolf JBW, Zinner D (2013) Hybridization and speciation. J Evol Biol 26:229–246
Bayer PE, Hurgobin B, Golicz AA, Chan CKK, Yuan YX, Lee H, Renton M, Meng JL, Li RY, Long Y, Zou J, Bancroft I, Chalhoub B, King GJ, Batley J, Edwards D (2017) Assembly and comparison of two closely related Brassica napus genomes. Plant Biotechnol J 15:1602–1610
Bretagnolle F, Thompson JD (1995) Tansley review no. 78. Gametes with the stomatic (sic) chromosome number: mechanisms of their formation and role in the evolution of autopolypoid plants. New Phytol 129:1–22
Chen S, Nelson MN, Chèvre A-M, Jenczewski E, Li Z, Mason AS, Meng J, Plummer JA, Pradhan A, Siddique KHM, Snowdon RJ, Yan G, Zhou W, Cowling WA (2011) Trigenomic bridges for Brassica improvement. Crit Rev Plant Sci 30:524–547
Chester M, Lipman MJ, Gallagher JP, Soltis PS, Soltis DE (2013) An assessment of karyotype restructuring in the neoallotetraploid Tragopogon miscellus (Asteraceae). Chromosom Res 21:75–85
Clarke WE, Higgins EE, Plieske J, Wieseke R, Sidebottom C, Khedikar Y, Batley J, Edwards D, Meng J, Li R, Lawley CT, Pauquet J, Laga B, Cheung W, Iniguez-Luy F, Dyrszka E, Rae S, Stich B, Snowdon RJ, Sharpe AG, Ganal MW, Parkin IAP (2016) A high-density SNP genotyping array for Brassica napus and its ancestral diploid species based on optimised selection of single-locus markers in the allotetraploid genome. Theor Appl Genet 129:1887–1899
Comai L (2005) The advantages and disadvantages of being polyploid. Nat Rev Genet 6:836–846
De Storme N, Geelen D (2013) Sexual polyploidization in plants--cytological mechanisms and molecular regulation. New Phytol 198:670–684
De Storme N, Mason AS (2014) Plant speciation through chromosome instability and ploidy change: cellular mechanisms, molecular factors and evolutionary relevance. Current Plant Biology 1:10–33
Dolatabadian A, Patel DA, Edwards D, Batley J (2017) Copy number variation and disease resistance in plants. Theor Appl Genet 130:2479–2490
Gaeta RT, Pires JC (2010) Homoelogous recombination in allopolyploids: the polyploid ratchet. New Phytol 186:18–28
Geng XX, Chen S, Astarini IA, Yan GJ, Tian E, Meng J, Li ZY, Ge XH, Nelson MN, Mason AS, Pradhan A, Zhou WJ, Cowling WA (2013) Doubled haploids of novel trigenomic Brassica derived from various interspecific crosses. Plant Cell Tiss Org 113:501–511
Gupta M, Atri C, Agarwal N, Banga SS (2016) Development and molecular-genetic characterization of a stable Brassica allohexaploid. Theor Appl Genet 129:2085–2100
Howard HW (1942) The effect of polyploidy and hybridity on seed size in crosses between Brassica chinensis, B. carinata, amphidiploid B. chinensis-carinata and autotetraploid B. chinensis. J Genet 43:105–119
Iwasa S (1964) Cytogenetic studies on the artificially raised trigenomic hexaploid hybrid forms in the genus Brassica. J Fac Agric Kyushu Univ 13:309–352
Jahne A, Lorz H (1995) Cereal microspore culture. Plant Sci 109:1–12
Lagercrantz U, Lydiate DJ (1996) Comparative genome mapping in Brassica. Genetics 144:1903–1910
Leitch AR, Leitch IJ (2008) Genomic plasticity and the diversity of polyploid plants. Science 320:481–483
Li QF, Chen YG, Yue F, Qian W, Song HY (2018) Microspore culture reveals high fitness of B. napus-like gametes in an interspecific hybrid between Brassica napus and B. oleracea. PLoS One 13:e0193548
Mallet J (2007) Hybrid speciation. Nat Rev 446:279–283
Mason AS, Batley J (2015) Creating new interspecific hybrid and polyploid crops. Trends Biotechnol 33:436–441
Mason AS, Batley J, Bayer PE, Hayward A, Cowling WA, Nelson MN (2014a) High-resolution molecular karyotyping uncovers pairing between ancestrally related Brassica chromosomes. New Phytol 202:964–974
Mason AS, Higgins EE, Snowdon RJ, Batley J, Stein A, Werner C, Parkin IAP (2017) A user guide to the Brassica 60K Illumina Infinium (TM) SNP genotyping array. Theor Appl Genet 130:621–633
Mason AS, Huteau V, Eber F, Coriton O, Yan G, Nelson MN, Cowling WA, Chèvre A-M (2010) Genome structure affects the rate of autosyndesis and allosyndesis in AABC, BBAC and CCAB Brassica interspecific hybrids. Chromosom Res 18:655–666
Mason AS, Nelson MN, Castello M-C, Yan G, Cowling WA (2011) Genotypic effects on the frequency of homoeologous and homologous recombination in Brassica napus × B. carinata hybrids. Theor Appl Genet 122:543–553
Mason AS, Nelson MN, Takahira J, Cowling WA, Moreira Alves G, Chaudhuri A, Chen N, Ragu ME, Dalton-Morgan J, Coriton O, Huteau V, Eber F, Chèvre A-M, Batley J (2014b) The fate of chromosomes and alleles in an allohexaploid Brassica population. Genetics 197:273–283
Mason AS, Takahira J, Atri C, Samans B, Hayward A, Cowling WA, Batley J, Nelson MN (2015) Microspore culture reveals complex meiotic behaviour in a trigenomic Brassica hybrid. BMC Plant Biol 15:173
Mason AS, Yan GJ, Cowling WA, Nelson MN (2012) A new method for producing allohexaploid Brassica through unreduced gametes. Euphytica 186:277–287
Mwathi MW, Gupta M, Atri C, Banga SS, Batley J, Mason AS (2017) Segregation for fertility and meiotic stability in novel Brassica allohexaploids. Theor Appl Genet 130:767–776
Navabi ZK, Stead KE, Pires JC, Xiong Z, Sharpe AG, Parkin IAP, Rahman MH, Good AG (2011) Analysis of B-genome chromosome introgression in interspecific hybrids of Brassica napus x B. carinata. Genetics 187:659–673
Navabi ZK, Strelkov SE, Good AG, Thiagarajah MR, Rahman MH (2010) Brassica B-genome resistance to stem rot (Sclerotinia sclerotiorum) in a doubled haploid population of Brassica napus x Brassica carinata. Can J Plant Pathol 32:237–246
Nelson MN, Mason AS, Castello MC, Thomson L, Yan GJ, Cowling WA (2009) Microspore culture preferentially selects unreduced (2n) gametes from an interspecific hybrid of Brassica napus L. x Brassica carinata Braun. Theor Appl Genet 119:497–505
Nicolas SD, Le Mignon G, Eber F, Coriton O, Monod H, Clouet V, Huteau V, Lostanlen A, Delourme R, Chalhoub B, Ryder CD, Chèvre AM, Jenczewski E (2007) Homeologous recombination plays a major role in chromosome rearrangements that occur during meiosis of Brassica napus haploids. Genetics 175:487–503
Nicolas SD, Monod H, Eber F, Chèvre AM, Jenczewski E (2012) Non-random distribution of extensive chromosome rearrangements in Brassica napus depends on genome organization. Plant J 70:691–703
Péle A, Rousseau-Gueutin M, Chèvre AM (2018) Speciation success of polyploid plants closely relates to the regulation of meiotic recombination. Front Plant Sci 9:907
R_Core_Team (2017) R: A language and environment for statistical computing., 2.10.1 edn. R Foundation for Statistical Computing, Vienna, Austria
Samans B, Chalhoub B, Snowdon RJ (2017) Surviving a genome collision: genomic signatures of allopolyploidization in the recent crop species Brassica napus. Plant Genome 10:3
Schiessl S-V, Katche E, Lhien E, Singh Chawla H, Mason AS (2018) The role of genomic structural variation in the genetic improvement of polyploid crops. The Crop Journal 7:127–140
Schubert I, Lysak MA (2011) Interpretation of karyotype evolution should consider chromosome structural constraints. Trends Genet 27:207–216
Seymour DK, Filiault DL, Henry IM, Monson-Miller J, Ravi M, Pang AD, Comai L, Chan SWL, Maloof JN (2012) Rapid creation of Arabidopsis doubled haploid lines for quantitative trait locus mapping. Proc Natl Acad Sci U S A 109:4227–4232
Song KM, Lu P, Tang KL, Osborn TC (1995) Rapid genome change in synthetic polyploids of Brassica and its implications for polyploid evolution. Proc Natl Acad Sci U S A 92:7719–7723
Stein A, Coriton O, Rousseau-Gueutin M, Samans B, Schiessl SV, Obermeier C, Parkin IAP, Chevre AM, Snowdon RJ (2017) Mapping of homoeologous chromosome exchanges influencing quantitative trait variation in Brassica napus. Plant Biotechnol J 15:1478–1489
Szadkowski E, Eber F, Huteau V, Lodé M, Huneau C, Belcram H, Coriton O, Manzanares-Dauleux MJ, Delourme R, King GJ, Chalhoub B, Jenczewski E, Chèvre AM (2010) The first meiosis of resynthesized Brassica napus, a genome blender. New Phytol 186:102–112
Takahira J, Cousin A, Nelson MN, Cowling WA (2011) Improvement in efficiency of microspore culture to produce doubled haploid canola (Brassica napus L.) by flow cytometry. Plant Cell Tiss Org 104:51–59
Tian E, Jiang Y, Chen L, Zou J, Liu F, Meng J (2010) Synthesis of a Brassica trigenomic allohexaploid (B. carinata × B. rapa) de novo and its stability in subsequent generations. Theor Appl Genet 121:1431–1440
U N (1935) Genome-analysis in Brassica with special reference to the experimental formation of B. napus and peculiar mode of fertilization. Japanese Journal of Botany 7:389–452
Udall JA, Wendel JF (2006) Polyploidy and crop improvement. Crop Sci 46:S3–S14
Xiong ZY, Gaeta RT, Pires JC (2011) Homoeologous shuffling and chromosome compensation maintain genome balance in resynthesized allopolyploid Brassica napus. Proc Natl Acad Sci U S A 108:7908–7913
Yang S, Chen S, Zhang KN, Li L, Yin YL, Gill RA, Yan GJ, Meng JL, Cowling WA, Zhou WJ (2018) A high-density genetic map of an allohexaploid Brassica doubled haploid population reveals quantitative trait loci for pollen viability and fertility. Front Plant Sci 9:1161
Zhou JN, Tan C, Cui C, Ge XH, Li ZY (2016) Distinct subgenome stabilities in synthesized Brassica allohexaploids. Theor Appl Genet 129:1257–1271
Ziolkowski PA, Berchowitz LE, Lambing C, Yelina NE, Zhao X, Kelly KA, Choi K, Ziolkowska L, June V, Sanchez-Moran E, Franklin C, Copenhaver GP, Henderson IR (2015) Juxtaposition of heterozygous and homozygous regions causes reciprocal crossover remodelling via interference during Arabidopsis meiosis. eLife 4:1–29
Acknowledgements
MM’s PhD study was supported by the Research and Higher Degree and the Tuition Scholarships from the University of Queensland, Australia, and by an Australia-India Strategic Research Fund: Biotechnology grant. Dr. Ning Cheng assisted with SNP array preparations. 60K Infinium SNP data imaging analysis was done at the Translational Research Institute (TRI) in Brisbane, Australia. ASM is supported by DFG Emmy Noether grant MA6473/1-1.
Author information
Authors and Affiliations
Contributions
MM conducted the experiments and collected the data, and co-wrote the manuscript and analysed the data with AM. SVS assisted with data analysis and interpretation and visualization of the data. MM, AM and JB contributed to manuscript revisions and provided input into experimental design. AM and JB supervised MM.
Corresponding author
Ethics declarations
The authors declare that this paper complies with all relevant ethical standards.
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Supplementary Figure 1
Graphed Log R Ratios (from −1.5 to 1.5) output from Illumina Genome Studio (normalized) for the Illumina Infinium Brassica 60 K SNP array SNPs used to genotype and derive copy-number data for six allohexaploid lines (MDL07, MDL23, MDL28, MDL30, MDL60 and MDL64) derived from microspore culture of a single hybrid plant of the cross (Brassica napus × B. carinata) × B. juncea and their self-pollinated progeny. (PPTX 6283 kb)
Supplementary Figure 2
Pollen viability in second-generation individuals derived from microspores of a Brassica napus × B. carinata) × B. juncea hybrid. First generation parent pollen viability is indicated with a blue star for each line. Different lowercase letters indicate significant differences in pollen viability between progeny sets (Tukey’s HSD; p < 0.05). Supplementary Figure 3 Number of novel karyotype changes (whole or segmental chromosome duplication and deletion events) in second-generation individuals derived from microspores of a Brassica napus × B. carinata) × B. juncea hybrid. Different lowercase letters indicate significant differences in the number of novel karyotype changes between progeny sets (Tukey’s HSD; p < 0.05). (PDF 220 kb)
ESM 1
Supplementary Table 1: Phased A- and C-genome allele calls from cleaned SNP data for microspore-derived allohexaploid lines (MDLs) from the cross (Brassica napus × B. carinata) × B. juncea and their self-pollinated progeny (highlighted in blue). Supplementary Table 2: Log R Ratio output from Illumina Genome Studio (normalized) for the Illumina Infinium Brassica 60 K SNP array SNPs used to genotype and derive copy-number data for allohexaploid lines derived from self-pollination (SP) and microspore culture (MDL) of a single hybrid plant of the cross (Brassica napus × B. carinata) × B. juncea. Self-pollinated progeny of each microspore derived line are highlighted in pale blue. Putative duplications (values >0.2 or > 0.5) are highlighted in blue, putative deletions of one chromosome copy (values between −0.2/−0.5 and − 2.0) are highlighted in orange, and putative deletions of both chromosome copies are highlighted in red (values < −2.0). Supplementary Table 3: Loss of both chromosome copies for a whole chromosome (del) or part of a chromosome (del – p), loss of one chromosome copy for a whole chromosome (miss) or part of a chromosome (miss – p), and duplication of a whole chromosome (dup) or part of a chromosome (dup – p) of A and C genome chromosomes in allohexaploid lines produced by microspore culture (MDL) and self-pollination (SP) of an allohexaploid hybrid from the cross combination (Brassica napus × B. carinata) × B. juncea), as predicted from Log R Ratios output by Illumina Genome Studio from Illumina Infinium Brassica 60 K SNP array data. (XLSX 23561 kb)
Rights and permissions
About this article
Cite this article
Mwathi, M.W., Schiessl, S.V., Batley, J. et al. “Doubled-haploid” allohexaploid Brassica lines lose fertility and viability and accumulate genetic variation due to genomic instability. Chromosoma 128, 521–532 (2019). https://doi.org/10.1007/s00412-019-00720-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-019-00720-w