Abstract
Background
Oral squamous cell carcinoma (OSCC) has the highest mortality rate among all head and neck cancers and a relatively low five-year survival rate. Generally, the development of an oral mucosal malignancy represents a multistep process beginning with normal oral mucosa epithelium and culminating in OSCC after transitioning through intermediary oral premalignant disorders (OPMDs), during which dysplasia is often observed. Noncoding RNAs (ncRNAs) are RNAs that are not translated into proteins, but still can participate in regulating neoplastic cell behavior. Recently, data have emerged on the role of ncRNAs in the progression of oral mucosal malignant diseases, but the exact mechanisms through which ncRNAs are involved remain to be elucidated.
Conclusions
Knowledge on ncRNAs has added an extra layer of complexity to our understanding of the malignant progression of oral mucosal diseases. The identification of ncRNAs in multiple body fluids as biomarkers may provide new diagnostic options that can be used for the diagnosis and prognosis of OPMDs and OSCC, respectively. Despite overall advances that have been made in cancer treatment, the treatment options for OPMDs and OSCC are still limited. Several studies have shown that ncRNA-based treatment regimens may hold promise as alternative methods for treating OPMDs and OSCC. The use of ncRNAs as therapeutic agents, including miR-155, miR-34 and lncRNA HOTAIR, appear promising.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Oral squamous cell carcinoma (OSCC), characterized by differentiation and a tendency to undergo lymph node metastasis, is the most common malignant tumor in the head and neck region, with over 200,000 newly diagnosed tumors each year [1]. Smoking, alcohol use, betel chewing and HPV infection are considered the major risk factors for OSCC [2, 3]. In most cases, the progression of oral mucosal malignancy requires an extended duration and multiple steps that may or may not involve risk factors, because OSCC may exhibit complex genetic changes and pathologies. Normal oral keratinocytes may be affected by adverse factors, resulting in changes in their intracellular microenvironment and genome, the latter of which can propagate changes during proliferation [4, 5]. These affected oral keratinocyte clones can transform into premalignant diseases and even further deteriorate into invasive OSCC. Most cases of OSCC are preceded by oral premalignant disorders (OPMDs), which are defined as epithelial lesions or disorders that have a high risk for malignant transformation [6,7,8].
Early-stage oral cancers and OPMDs are often subtle and asymptomatic. Therefore, it is important to create and improve tools for detecting early-stage oral cancers and OPMDs. Ample evidence has indicated that noncoding RNAs (ncRNAs) may participate in nearly every step of oral mucosal tumorigenesis, Thus, understanding the functional characteristics of these ncRNAs is essential. DNA alterations and changes in the expression of genes, such as MMP1 and KNG1, are considered promising biomarkers for diagnosing OPMDs and for detecting OSCC, while RACK1 and PA28γ have shown promise as prognostic predictors for OSCC. ncRNAs can be identified in various (pre-)malignant tissues, and it has been found that the specificity of identification can be improved when they are combined with other cancer cell markers [9,10,11]. Here, we discuss the importance of ncRNAs as biomarkers and therapeutic targets in OPMDs and OSCC by systematically reviewing the present literature regarding the emerging roles of ncRNAs. We believe that this review will add to our current understanding of the malignant progression of oral mucosal diseases and to the identification of new clinical tools to be used in the diagnosis and treatment of OPMDs and early-stage OSCC.
2 Characteristics of ncRNAs
It has been shown that only 2% of our DNA encodes proteins, whereas more than 70% is actively transcribed into ncRNAs that may function in gene regulation, mRNA maturation and/or protein synthesis [12]. Although they lack the ability to be translated into proteins, ncRNAs are also subject to alterations that can drive cancer development, in addition to the well-known protein coding gene alterations. ncRNAs include:
MicroRNAs (miRNAs), which are highly conserved ncRNAs approximately 21 nucleotides in length. They are involved in gene regulation by, generally, inhibiting the translation of target genes [13]. However, various functions of miRNAs have been discovered and they can be categorized into eight main types: conventional downregulation of gene expression and seven unconventional functions, i.e., pri-miRNAs coding for peptides, interaction with non-Ago proteins, activation of Toll-like receptors (TLRs), upregulation of gene expression, targeting nuclear ncRNAs, targeting mitochondrial transcripts, and direct activation of transcription (Fig. 1a).
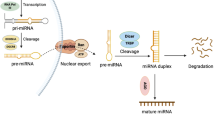
Functions of miRNAs and lncRNAs. a The functions of miRNAs. a: pri-miRNAs coding for peptides: At the start of transcription, a miRNA is generated as a longer pri-miRNA. There is evidence that some pri-miRNAs encode peptides, which are termed miRNA-encoded peptides (miPEPs). These miPEPs play a role in increasing the transcription of their own pri-miRNAs and in enhancing the accumulation of mature miRNAs; b: Downregulating protein expression: miRNAs can post-transcriptionally reduce the expression levels of target proteins by either promoting messenger RNA (mRNA) decay or by blocking mRNA translation; c: Interacting with non-AGO proteins: miRNAs interact with Argonaute (AGO) protein-containing complexes, where they perform conventional repression functions. Examples of miRNAs that can interfere with the function of a specific protein by acting as a RNA decoy have been reported; d: Activating Toll-like receptors: Although direct physical interactions have not yet been proven, an unconventional role for miRNAs has been reported that involves activation of Toll-like receptors (TLRs); e: Upregulating protein expression: miRNAs can directly associate with AGO2 and AU-rich elements (AREs) in target mRNAs to activate translation; f: Targeting nuclear ncRNAs: miRNAs can localize to the nucleus to inhibit the maturation of other miRNAs via direct interactions with their primary transcripts; g: Targeting mitochondrial transcripts: Although no miRNAs have been identified within the mitochondrial genome, miRNAs can translocate into mitochondria to inhibit target gene expression while simultaneously increasing the mRNA expression and protein content of target genes; h: Directly activating transcription: A specific miRNA can act as a transferable nuclear localization element that directs the nuclear enrichment of small ncRNAs by attaching to them and importing them into the nucleus via importin 8, which enables them to bind to promoters and induce transcription of target genes. b The functions of lncRNAs. a: Pre-transcriptional regulation: Pre-transcriptional regulation includes lncRNA regulation, splicing of pre-mRNAs, regulation of translation, recruitment of proteins to stabilize mRNAs by binding to target miRNA sites or binding directly to mRNAs to stabilize them, and serving as precursors or transcriptional hosts for small RNAs (such as miRNAs); b: Post-transcriptional regulation: Post-translational regulation of proteins includes the involvement of lncRNAs in subcellular structure formation, protein transport and localization, and in mediating the formation of protein complexes, where they serve as scaffolds to either promote protein-protein interactions or stabilize proteins; this process may occur in the nucleus or the cytoplasm; c: miRNA sponging: lncRNAs can absorb or “sponge” miRNAs that would (if not bound to a lncRNA) otherwise inhibit the expression of their target mRNA(s); these RNAs have been termed competing endogenous RNAs (ceRNAs); this process can occur in the nucleus or the cytoplasm; d: Epigenetic regulation: Epigenetic regulation can occur when DNA interacts with lncRNAs to regulate histone modifications and chromatin remodeling/folding, which affects the recruitment of transcription factors to target promoters for active transcription. Furthermore, with the formation of RNA/DNA/protein complexes, transcription is inhibited, since the formation of complexes with transcription factors prevents their ability to activate transcription; this occurs only in the nucleus
Long noncoding RNAs (lncRNAs), transcripts longer than 200 bp that are classified according to their functions in pre-transcriptional regulation, post-transcriptional regulation, miRNA sponging, and epigenetic regulation (Fig. 1b) [14].
Circular RNAs (circRNAs), which are ncRNAs with closed loop structures. As previously described, the functions of circRNAs can be categorized into five main types: regulating linear RNA transcription, miRNA sponging, protein sponging, interaction with different proteins, and being translated into peptides [15].
3 Role of ncRNAs in OPMDs
At the beginning of the development of malignant oral mucosal diseases, OPMDs are generally found in the oral cavity. They possess a high risk of malignant transformation into invasive OSCC. Clinically, OPMDs usually appear grossly abnormal and are often accompanied by oral leukoplakia (OLK), oral lichen planus (OLP), oral submucous fibrosis (OSF) or other types of potentially malignant diseases [6]. OPMDs occur as a result of early processes by which both genetic and phenotypic changes, including changes in ncRNAs, accumulate in the normal oral mucosa [16].
3.1 Oral leukoplakia
Oral leukoplakia (OLK) often presents clinically as a ‘white patch’ in the oral cavity, from which 5%–36% of OSCC cases develop [17]. Based on histopathological evaluation, OLK can be classified as nondysplastic or dysplastic. OLK with moderate or severe dysplasia is also called malignantly transformed OLK (mtOLK), which is considered to have a higher risk for progression to carcinoma than OLK with mild dysplasia. It has been found that 3 miRNAs may be significantly dysregulated in mtOLK and can mediate the initiation and development of OLK [18]. By contrast, clinical and histological characteristics cannot distinguish between “progressing” and “non-progressing” cases among nondysplastic or mild to moderately dysplastic OLK. The identification of specific miRNAs may, however, be able to make these distinctions. It was found that the expression of several miRNAs, including miR-10b-3p, was significantly altered in the saliva of patients with progressing mild dysplasia OLK relative to that in patients with non-progressing OLK [19]. Another study indicated that a proposed expression profile of 8 overexpressed miRNAs was related to the progression of OLK to OSCC. In particular, the miR-345/21/181b subset was strongly related to increased lesion severity in the histological progression from premalignant to malignant lesions [20], which may indicate that miRNAs have the potential to predict and distinguish “progressing” from “non-progressing” OLK with comparatively mild dysplasia.
3.2 Oral lichen planus
Oral lichen planus (OLP) is characterized histologically by a subepithelial band-like lymphocytic infiltrate and epithelial basal cell destruction, and it is a common chronic inflammatory disease associated with immunological dysfunction mediated by cells, including CD4+, CD8+ and helper T cells. Furthermore, OLP has a tendency to develop dysplasia and to progress towards malignancy [21, 22]. Chronic inflammation is the main pathological feature of OLP, and many recent studies have revealed roles of ncRNAs in mediating unbalanced immunoreactions in OLP. Upon comparison of miRNA expression profiles in peripheral blood mononuclear cells (PBMCs) from patients with OLP to those from healthy volunteers, it was found that miR-155 was the most downregulated and miR-19a the most upregulated miRNA in the OLP group [23]. By synergistically functioning to induce an imbalance between Th1 and Th2 cells, simultaneous deregulation of miR-155/eNOS and miR-19a/TLR2 was shown to be responsible for an elevated risk of OLP in an in vitro experiment [23]. Furthermore, a positive feedback loop involving miR-155 and IFN-γ was found to potentially contribute to the Th1-dominated immune response in erosive-type OLP, and SOCS1 was identified as the most likely target of miR-155 involved in this feedback loop [24]. Overexpression of lncRNA DQ786243 significantly increased the suppressive function of CD4+ T cells, such as Th1 and Th17, by decreasing the levels of IFN-γ and IL-17 in CD4+ Treg cells. This regulation was observed in peripheral blood of OLP patients, and to occur through the Foxp3/miR-146a/NF-κB axis [25].
Apart from chronic inflammation, dysplasia involving the degeneration of keratinocytes is another pathological feature of OLP. It was found that in an LPS-induced OLP model, miR-125b directly targeted MMP-2, inhibited keratinocyte proliferation and promoted keratinocyte apoptosis through the PI3K/Akt/mTOR signaling pathway [26]. Foxp3 can regulate miR-146a, thereby controlling the proliferation and apoptosis of LPS-treated immortalized human keratinocyte HaCaT cells, and Foxp3/miR-146a has been shown to regulate TRAF6 expression in CD4+ T cells in OLP [27]. Additionally, miR-27b-3p suppresses keratinocyte apoptosis in OLP by targeting cyclin D and regulating BCL2 signaling pathways, suggesting that miRNAs could be potential treatment targets for preventing the processes by which OLP initiates or progresses to OSCC [28]. A recent study revealed that lncRNA MEG3 could induce apoptosis of keratinocytes in OLP by sponging miR-361-5p and promoting the expression of SDHD [29]. Since there is a large body of work on miRNAs in OLP, we summarized their significance in Table 1.
3.3 Oral submucous fibrosis
Oral submucous fibrosis (OSF) has been identified as an OPMD characterized by a burning sensation, blanching and stiffening of the oral cavity, resulting in patients having difficulty opening their mouths and eventually enduring a series of histopathological stages culminating in invasive OSCC. The transformation rate is 7–13% [44, 45]. At present, our knowledge about the exact mechanisms underlying the initiation and development of OSF is still limited, especially regarding the exact roles of ncRNAs in this disease. A study on miRNA expression profiles showed that a total of 11 unique miRNAs were differentially expressed in tissues from OSF patients compared to the normal oral mucosa, suggesting a potential role of these newly identified miRNAs in OSF [46]. In addition, low levels of miR-499a-5p were found to contribute to an increased risk of progression of OSF, due to betel quid chewing, to OSCC [47]. Overexpression of miR-200b abolished arecoline-induced myofibroblast activities and led to downregulation of a-SMA and vimentin by decreasing ZEB2 in OSF [48].
In addition, Zhou et al. identified 687 lncRNAs that were significantly and differentially expressed during OSF progression, including 231 upregulated lncRNAs and 456 downregulated lncRNAs. These lncRNAs were found to be associated with OSCC pathogenesis, and the involved processes to include inflammation signaling, Wnt signaling, angiogenesis, CCKR signaling, integrin signaling, PDGF signaling, p53 signaling and EGFR signaling pathways [49]. These differentially expressed lncRNAs may provide new leads for the study of OSF malignant development and its treatment. Recently, some studies have revealed a potential role for lncRNAs in the development and progression of OSF, which occurs mainly by increasing myofibroblast activities through TGFβ signaling. It was found that lncRNA LINC00974 could promote the development of OSF by increasing myofibroblast activities via elevation in the expression levels of α-SMA, α-1 type I collagen, and fibronectin through TGF-β/Smad signaling [50]. Overexpression of lncRNA GAS5-AS1 was found to regulate arecoline-associated myofibroblast activation to suppress fibrogenesis, which was achieved by blocking upregulation of p-Smad2 by arecoline [51].
Based on these preliminary data, ncRNAs can be identified as a class of essential regulators that promote the progression of high-risk OPMDs to OSCC under certain circumstances. To determine the networks in which ncRNAs participate in OPMDs, more research is needed.
4 Functional characteristics of ncRNAs in OSCC
4.1 Regulation of proliferation and survival
Recent evidence indicates that ncRNAs are significantly correlated with the growth and survival of OSCC cells, because they can disrupt the balance between proliferative signaling and growth suppression in multiple ways (Fig. 2). Receptor signaling can, for example, be deregulated by downregulation of miRNAs whose direct targets are growth factor receptors. OSCC cells with a miR-375low/IGF-1Rhigh signature have considerable proliferation potential [52]. In addition, a lack of OSCC dependence on growth factors may also be due to continual stimulation of downstream signaling owing to ncRNA dysregulation. Notably, low expression levels of miR-138 and miR-1285-3p have been observed in OSCC cells with elevated YAP1 expression and an activated Hippo pathway, and lncRNA RBM5-AS1 has been shown to rescue miR-1285-3p-mediated inhibition of YAP1 by sponging miR-1285-3p. This phenomenon was confirmed in nude mice [53, 54]. Moreover, KLF8 may be recruited by lncRNA AC132217.4 to bind the 3′-UTR of IGF2 mRNA, resulting in upregulation of its expression at the transcriptional level. Furthermore, circR-0007059 has been reported to regulate the proliferation, metastasis and invasion of OSCC cells via the AKT/mTOR pathway, which was confirmed by in vivo experiments [55]. ncRNAs can also influence the regulation of growth factor expression in OSCC cells. It has been found, for example, that miR-338 and miR-23a-3p can significantly decrease the expression of NRP1 and EGFL7, respectively, and that lncRNA OIP5-AS can abolish miR-338-mediated control of NRP1 expression, resulting in OSCC cell proliferation [56,57,58,59]. Modulating cell cycle regulators is another way by which ncRNAs can regulate cell growth in OSCC. It has been found that downregulated miR-145 and miR-155 expression can increase the expression of c-Myc, CDK6 and p27Kip1, thereby preventing cyclin D/E expression in OSCC and helping cells to escape from G1 phase arrest in OSCC [60, 61]. LncRNA CASC2 has been shown to play a similar role in regulating the cell cycle by targeting CDK1 in OSCC [62].
ncRNAs involved in regulating proliferation and survival in OSCC. ncRNAs can mediate the generation of growth factors, promote the presence of growth factor receptors on the cell surface, activate signaling pathways downstream of these receptors and disrupt the secretion of growth factors, leading to a proliferative stimulus of OSCC cells. In the EGF/PI3K/AKT pathway, miR-375 and lncRNA AC1322.17.4/miR-23a-3p inhibit OSCC cell proliferation by reducing the levels of IGFR and IGF/EGF, respectively. cicR-0007059 can promote OSCC cell proliferation via the AKT pathway. In the WNT pathway, miR-27b and sponging of miR-224-5p by lncRNA FTH1P3 inhibit Frizzled at the cell surface, and the lncRNA RBM5-AS1/miR-1285-5p axis can regulate YAP/TAZ to reduce the proliferation of OSCC cells (the colored circles represent circRNA; TF = transcription factor). miR-155 promotes the expression of CDK2/4/6 and Cyclin D2/E by simultaneously inhibiting BCL6 and p21Cip1/p27kip1. circR-100,290 and lncRNA PDA3P sponge miR-29b and miR-185-3p, respectively, which allows OSCC cells to escape the checkpoint by mediating CDK expression. LncRNAs ELF3-AS1 and GAS5, and miR-340 and miR-143 target Glut and mediate glycolysis in OSCC cells. LncRNAs Meg3 and circR-DOCK1 sponge miR-548d-3p and miR-196a-5p, which target the JAK/STAT pathway and BIRC1, respectively, thus regulating apoptosis.
Evidence has indicated that ncRNAs can operate as control nodes that govern cellular decisions to perform certain proliferative or apoptotic activities by regulating their target genes, thus serving as tumor suppressors in OSCC. miR-194 can, for example, decrease cyclin D1 expression and promote p21 expression by inhibiting the PI3K/AKT/FoxO3a signaling pathway via suppression of AGK in OSCC [63]. miR-377 can upregulate its target gene HDAC9 and inhibit apoptosis of OSCC cells in part through its effects on the NR4A1/Nur77 pathways [64]. LncRNAs HOXA11-AS and LINC00958 have been found to inhibit apoptosis of OSCC cells by sponging miRNAs in the miR-98-5p-YBX2 and miR-185-5p/YWHAZ axes, respectively [65, 66]. LncRNA MEG3 plays a role in OSCC as a growth suppressor by regulating the miR-548d-3p/JAK-STAT pathway [67]. It has also been found that circ-DOCK1 and circR-100,290 can regulate BIRC3 and CDK6 by sponging miR-196a-5p and miR-29b, respectively, thereby participating in the regulation of OSCC. This evidence was based on tumor formation experiments in nude mice [68, 69]. However, all of the abovementioned growth-suppressing ncRNAs are expressed at low levels in OSCC. Thus, OSCC cells escape these growth inhibiting processes.
Reprogramming of the energy metabolism network mediated by ncRNAs is an alternative strategy used by OSCC cells to survive. Glut1 has been identified as a popular target by which ncRNAs regulate energy metabolism. It has been suggested that miR-143 targeting hexokinase 2 and miR-340 targeting Glut1 are important for cellular glucose metabolism and proliferation in OSCC, which was confirmed in a xenograft model [70, 71]. Similarly, experimental findings indicated that miR-31-5p-ACOX1 changes the lipid metabolome in OSCC, promoting the expression of certain esters [72]. LncRNAs ELF3-AS1 and GAS5 have been found to promote the proliferation of OSCC cells by regulating Glut1 and reprogramming glucose metabolism [73, 74].
4.2 Acquisition of invasive and metastatic abilities
The contribution of ncRNAs to the acquisition of invasive and metastatic abilities by OSCC cells can be broadly grouped into the following mechanisms (Fig. 3). Aberrant expression of some ncRNAs facilitates the separation of OSCC cells from each other and from the extracellular matrix (ECM), but they increase the adhesion of OSCC cells to basement membranes. Zheng et al., for instance, revealed that TNF-α can inhibit the metastasis of OSCC cells in a miR-765high/EMP3low/p66Shchigh pattern-dependent manner [75]. LncRNA HOTAIR, which is overexpressed in OSCC, can promote malignancy of OSCC by recruiting EZH2 and H3K27me3 to local chromatin, resulting in suppression of E-cadherin expression [76]. Aberrant ncRNA expression can also promote ECM degradation. MMP2 has been found to be upregulated in OSCC cells due to a low expression of miR-29a, which directly targets MMP2. These observations suggest that miR-29a plays an inhibitory role in the invasion and migration of OSCC cells [77]. LncRNA FOXC1 can recruit FOXCUT to local chromatin and promote the expression of MMPs and VEGF-A, resulting in an increased proliferation and migration of OSCC cells [78, 79]. Aberrant expression of other ncRNAs, in turn, can regulate transcription factors whose downstream pathways contribute to the migration and invasion of OSCC cells. It has been found, for example, that overexpression of ΔNp63, a direct target of miR-138-5p, which decreases ΔNp63 expression [80], can reduce miR-138-5p transcription in OSCC cells, which results in a ΔNp63high/miR-138-5plow pattern that is partly the result of positive feedback. This dysregulation increases the growth and migration of OSCC cells. LncRNAs LINC00974 and HCP5 promote OSCC cell migration and invasion via the miR-122/RhoA and miR-140-5p/SOX4 axes, respectively [81]. As mentioned before, lncRNA NEAT1 is a potential biomarker for OLP and a recent study has shown that NEAT1 can act as a ncRNA sponge for miR-365 and promote OSCC cell proliferation and invasion by regulating RGS20 [82]. Dysregulated ncRNAs can also promote surface expression of receptors that contribute to metastasis and invasion of OSCC cells. Yuan et al. reported, for example, that miR-101 inhibits OSCC cell growth, invasion and migration by targeting CXCR7 [83].
ncRNAs involved in acquiring invasive and metastatic abilities in OSCC. ncRNAs can increase the ability of OSCC cells to invade and metastasize by decreasing the number of cell attachments. miR-497 targets Smad7 to mediate TGF-β, and increased miR-765 reduces the expression of adhesion molecules, including EMP-3, ZO-1 and E-cadherin, on the surface of the cell. LncRNA LINC01116 regulates RhoA via miR-122 and participates in the TGF-β pathway. The lncRNA LINC01116/miR-136 axis regulates FN1 and adhesion between cells and the extracellular matrix (ECM) in OSCC. miR-29 increases the expression of the adhesion molecule CX3CL1 in OSCC cells to increase their adhesion to the basement membrane, and it promotes ECM degradation by increasing MMP2 secretion. LncRNAs H1 and FALEC (which sponge miR-21), the lncRNA HCP5/miR-140-5p axis and lncRNA LUCAT1 regulate the expression of the pleiotropic transcription factors EZH2, SOX4 and FOXCUT, respectively. LncRNA FOXC1 can recruit FOXCUT to local chromatin and promote the expression of MMPs. miR-424-5p plays an important role in irregular loop activity, as the SOCS2/STAT5/miR-424-5p axis regulates invasion and metastasis. miR-138-5p targets ΔNp63 and is upregulated by ΔNp63 in OSCC cells.
4.3 Remodeling the tumor microenvironment
The tumor microenvironment (TME) is a complex network composed of a variety of cell types (e.g. endothelial cells, fibroblasts, immune cells) and extracellular components (e.g. cytokines, growth factors, hormones, ECM) that surround tumor cells [84,85,86]. In recent years, many studies have revealed that ncRNAs, as extracellular components, can participate in the regulation of the TME and mediate the interaction between tumor cells and other cells, including endothelial cells, fibroblasts and immune cells, in the TME to promote the development and invasion of tumors.
One of the characteristics of a hostile TME is poor oxygen (O2) and nutrient supply. The strategy for tumor cells to escape the stress from hypoxia and poor nutrient supply is angiogenesis. Current studies have shown that low levels of miR-126 and miR-320 in OSCC cells are involved in activation of angiogenesis and lymphangiogenesis via VEGF-A and NRP1, respectively, which was confirmed by in vivo tumor growth experiments [87, 88]. In addition, Li and colleagues found that depletion of miR-21 in hypoxic OSCC cells results in low miR-21 expression in exosomes, leading to reduced metastasis and invasion. This effect can be rescued by the introduction of miR-21 in exosomes released by OSCC cells grown under normoxic conditions [89]. Similarly, it was shown that when miR-200c-3p was transferred via exosomes, the recipient OSCC cells with a low risk of invasion were prompted towards invasion by disruption of CHD9 and WRN [90]. Moreover, transmission of drug-resistant phenotypes mediated by ncRNAs depends on a similar pattern. Exosomes containing miR-21 from cisplatin-resistant OSCC cells were found to promote chemoresistance by targeting PTEN and PDCD4 in recipient OSCC cells [91]. Additionally, injection of exosomes containing miR-21 from cisplatin-resistant OSCC cells induced cisplatin resistance in OSCC murine models [91].
ncRNAs play an important role in communication between tumor cells and other cell types in the TME. The main stromal cell type in the TME is represented by cancer-associated fibroblasts (CAFs) [92], which were found to be capable of transferring exosomes containing miR-34a-5p to OSCC cells. This miRNA binds to AXL, resulting in inhibition of the AKT/GSK-3β/β-catenin pathway in OSCC cells. Moreover, injection of Cal27 cells with CAF-expressed miR-34a-5p reduced the tumorigenicity of OSCC in vivo [93]. miR-382-5p in exosomes derived from CAFs has been found to promote the migration and invasion of OSCC cells [94]. The immune cells in the TME are essential regulators of interactions between cancer cells and the TME. It was found that miR-29a-3p in exosomes from OSCC promotes M2 subtype polarization by activating SOCS1/STAT6 signaling in tumor-associated macrophages [95]. In addition, a study focusing on miRNA profiles in extracellular vesicles from OSCC cells revealed that oncogenic miRNA could reprogram monocytes via the NF-κB pathway [96].
It appears that the only research reported on the role of ncRNAs in remodeling the TME in OSCC relates to miRNAs. Given the complexity of the regulatory network of the TME and the important roles of lncRNAs and circRNAs in cancer development, we recognize that also these latter RNAs likely play essential roles in remodeling the TME. Several studies have underscored this notion. LncRNA NKILA has, for example, been found to promote immune evasion of tumors by sensitizing T cells to activation-induced cell death in breast cancer [97]. Under hypoxic stress, lncRNA CamK-A activated NF-kB via the Ca2+ signaling pathway and remodeled the TME in breast cancer. Furthermore, the circ_0000977/miR-153 axis has been found to enable a MICA- and HIF1A-mediated immune escape of pancreatic cancer cells from NK cells [98, 99]. LncRNAs and circRNAs in OSCC may have a similar effect in remodeling the TME, which turns them into promising targets for OSCC treatment.
5 ncRNAs: From the bench to the clinic
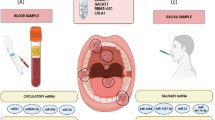
Accumulating evidence indicates that ncRNAs have the potential to be used in diagnosing OPMDs. It has been found, for example, that lncRNA NEAT1, miR-21/184 and miR-145 are deregulated but stably exist in the saliva of patients with OPMDs and OSCC [100, 101]. In addition, it has been found that miR-196 in plasma of patients with OPMDs may serve as a biomarker for early cancer detection, whereas no significant association of miR-196 expression was found with demographic characteristics of patients, including sex, age and smoking status [102]. Another report has indicated that miR-375/21/181b and miR-345 are consistently elevated and related to the degree of lesion severity during disease progression [20]. Zhou et al. found that 687 lncRNAs were significantly and differentially expressed during OSF progression, and that they may serve as additional biomarkers for the diagnosis of OPMDs [46]. The levels of these ncRNAs changed gradually during progressive stages of OSCC, suggesting that at least a subset of these ncRNAs may constitute a signature for OPMD progression.
In considering the application of ncRNAs in the detection of OSCC, miRNAs in serum, plasma and saliva have been proposed for many years to serve as biomarkers for oral cancer. Many studies have reported that several ncRNAs have the potential to be used as noninvasive biomarkers for diagnosing oral cancer. For example, many miRNAs, including miR-136, have been reported to be present at lower levels in saliva of patients with OSCC than in those of normal control subjects [103]. In addition, it has been found that serum levels of lncRNAs AC007271.3, SCCA and TSGF can distinguish patients with OSCC from healthy individuals [104]. Salivary levels of circ-0001874 and circ-001971 have also been tested as early biomarkers for establishing the diagnosis, TNM stage and tumor grade of OSCC [105].
To date, several efficient methods for using ncRNAs to treat OSCC have been reported in experimental animal models. It has, for example, been found that intravenous injection of antisense miR-21 oligonucleotides into OSCC murine models reduced tumor growth [106]. In addition, when cells overexpressing lncRNA AC132217.4 were injected intravenously in nude mice, the mice exhibited an increased number of lung nodules [107]. Moreover, when tumor cells overexpressing circ-AKT3 were injected into nude mice, they showed a decreased ability to form cancers [108]. In 2013, the Texas-based company Mirna Therapeutics began using MRX34, a miR-34 mimic, for treating cancer. It became the first miRNA drug to reach phase 1 trials. However, the trial had to be halted due to unexplained immune-related adverse events. Therefore, RNA-based treatment of OSCC and OPMDs requires additional research before it can be clinically applied.
6 Conclusions and perspectives
A biological role of ncRNAs in various diseases has amply been demonstrated. Many studies have, however of high quality, failed to exactly define the mechanisms of action of ncRNAs. One clear example is a study demonstrating that transfection of a miR-545 mimic into OSCC cells can downregulate RIG-I protein expression, but to substantiate this notion only one plain Western blot image was presented [109]. In addition, most current studies focus on miRNAs and conventional functions of ncRNAs, such as miRNAs downregulating target expression and lncRNAs and circRNAs sponging miRNAs. These focused studies may overlook broader perspectives of ncRNAs. Therefore, increased attention should be devoted to uncovering unconventional regulatory mechanisms related to ncRNAs, especially lncRNAs and circRNAs, in OPMDs and OSCC.
The stable presence of ncRNAs in multiple body fluids has potential as biomarkers and prognostic indicators for malignant OPMDs and OSCC, respectively. As such, they may provide a new type of diagnostic tool in the clinic. A liquid biopsy is representative of a noninvasive method and is less painful. There are many conflicting results regarding the early diagnosis of OSCC because of varying sample resources and demographic differences (such as ethnicity), which may affect the diagnostic efficacy of ncRNAs [110]. Once these problems are addressed, however, these biomarkers may guide the detection, risk assessment, and treatment decisions in patients with OPMDs and OSCC, especially in patients with OPMDs that are at risk for malignant transformation.
Several studies have shown that ncRNA-based treatment regimens have potential as promising alternatives for treating OPMDs and OSCC. MiR-21 and lncRNA UCA1/miR-184 have, for instance, been linked to cisplatin resistance in OSCC, and targeting these ncRNAs in conjunction with cisplatin treatment may improve patient outcome [91, 111]. However, identifying the ideal ncRNA candidates as targets for OPMDs and OSCC is still a major challenge in ncRNA-based treatment. Also other problems, including tissue-specific targeting, potential toxicities and off-target effects, need to be solved before ncRNAs can be used as mainstream therapeutic targets for OPMDs and OSCC. Once these issues are resolved, ncRNAs may increasingly be implemented as alternative treatment options for patients with OSCC and OPMD.
Overall, research on ncRNAs has added an extra layer of complexity to our understanding of the malignant progression of oral mucosal diseases (Table 2). Given the large number of ncRNAs that exist, those reviewed in this article may represent only a small portion of the ncRNAs functionally related to the malignant progression of oral mucosal diseases. Thus, also here additional research is warranted. The lack of adequate treatment targets and administration methods remain major challenges for adequately addressing the malignant progression of oral mucosal diseases in the near future.
Data Availability
The datasets generated and analyzed during the current study are available in the PubMed repository, www.ncbi.nlm.nih.gov/pubmed.
References
F. Bray, J. Ferlay, I. Soerjomataram, R.L. Siegel, L.A. Torre, A. Jemal, Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 68, 394–424 (2018)
S.R. Moore, N.W. Johnson, A.M. Pierce, D.F. Wilson, The epidemiology of mouth cancer: A review of global incidence. Oral Dis. 6, 65–74 (2000)
A.W. Joseph, G. D'Souza, Epidemiology of human papillomavirus-related head and neck cancer. Otolaryngol. Clin. N. Am. 45, 739–764 (2012)
L.L. Feller, R.R. Khammissa, B.B. Kramer, J.J. Lemmer, Oral squamous cell carcinoma in relation to field precancerisation: Pathobiology. Cancer Cell Int. 13, 31 (2013)
C.P. Gan, K.K. Sam, P.S. Yee, N.S. Zainal, B.K.B. Lee, Z.A.A. Rahman, V. Patel, A.C. Tan, R.B. Zain, S.C. Cheong, IFITM3 knockdown reduces the expression of CCND1 and CDK4 and suppresses the growth of oral squamous cell carcinoma cells. Cell. Oncol. 42, 477–490 (2019)
U.U. Malik, S. Zarina, S.R. Pennington, Oral squamous cell carcinoma: Key clinical questions, biomarker discovery, and the role of proteomics. Arch. Oral Biol. 63, 53–65 (2016)
S. Warnakulasuriya, N.W. Johnson, I. van der Waal, Nomenclature and classification of potentially malignant disorders of the oral mucosa. J. Oral Pathol. Med. 36, 575–580 (2007)
L. Ben, Slama, Potentially malignant disorders of the oral mucosa: Terminology and classification. Rev. Stomatol. Chir. Maxillofac. 111, 208–212 (2010)
J.S. Yu, Y.T. Chen, W.F. Chiang, Y.C. Hsiao, L.J. Chu, L.C. See, C.S. Wu, H.T. Tu, H.W. Chen, C.C. Chen, W.C. Liao, Y.T. Chang, C.C. Wu, C.Y. Lin, S.Y. Liu, S.T. Chiou, S.L. Chia, K.P. Chang, C.Y. Chien, S.W. Chang, C.J. Chang, J.D. Young, C.C. Pao, Y.S. Chang, L.H. Hartwell, Saliva protein biomarkers to detect oral squamous cell carcinoma in a high-risk population in Taiwan. Proc. Natl. Acad. Sci. U. S. A. 113, 11549–11554 (2016)
S. Liu, J. Liu, J. Wang, J. Cheng, X. Zeng, N. Ji, J. Li, Q. Chen, RACK1 is an organ-specific prognostic predictor in OSCC. Oral Oncol. 76, 22–26 (2018)
J. Li, X. Feng, C. Sun, X. Zeng, L. Xie, H. Xu, T. Li, R. Wang, X. Xu, X. Zhou, M. Zhou, Y. Zhou, H. Dan, Z. Wang, N. Ji, P. Deng, G. Liao, N. Geng, Y. Wang, D. Zhang, Y. Lin, L. Ye, X. Liang, L. Li, G. Luo, L. Jiang, Z. Wang, Q. Chen, Associations between proteasomal activator PA28gamma and outcome of oral squamous cell carcinoma: Evidence from cohort studies and functional analyses. EBioMedicine 2, 851–858 (2015)
D.R. Zerbino, P. Achuthan, W. Akanni, M.R. Amode, D. Barrell, J. Bhai, K. Billis, C. Cummins, A. Gall, C.G. Giron, L. Gil, L. Gordon, L. Haggerty, E. Haskell, T. Hourlier, O.G. Izuogu, S.H. Janacek, T. Juettemann, J.K. To, M.R. Laird, I. Lavidas, Z. Liu, J.E. Loveland, T. Maurel, W. McLaren, B. Moore, J. Mudge, D.N. Murphy, V. Newman, M. Nuhn, D. Ogeh, C.K. Ong, A. Parker, M. Patricio, H.S. Riat, H. Schuilenburg, D. Sheppard, H. Sparrow, K. Taylor, A. Thormann, A. Vullo, B. Walts, A. Zadissa, A. Frankish, S.E. Hunt, M. Kostadima, N. Langridge, F.J. Martin, M. Muffato, E. Perry, M. Ruffier, D.M. Staines, S.J. Trevanion, B.L. Aken, F. Cunningham, A. Yates, P. Flicek, Ensembl 2018. Nucleic Acids Res. 46, D754–D761 (2018)
M. Gorenchtein, C.F. Poh, R. Saini, C. Garnis, MicroRNAs in an oral cancer context - from basic biology to clinical utility. J. Dent. Res. 91, 440–446 (2012)
F. Kopp, J.T. Mendell, Functional classification and experimental dissection of long noncoding RNAs. Cell 172, 393–407 (2018)
K. Lei, H. Bai, Z. Wei, C. Xie, J. Wang, J. Li, Q. Chen, The mechanism and function of circular RNAs in human diseases. Exp. Cell Res. 368, 147–158 (2018)
T. Yap, C. Seers, K. Koo, L. Cheng, L.J. Vella, A.F. Hill, E. Reynolds, A. Nastri, N. Cirillo, M. McCullough, Non-invasive screening of a microRNA-based dysregulation signature in oral cancer and oral potentially malignant disorders. Oral Oncol. 96, 113–120 (2019)
A. Villa, S. Sonis, Oral leukoplakia remains a challenging condition. Oral Dis. 24, 179–183 (2018)
A. Maimaiti, K. Abudoukeremu, L. Tie, Y. Pan, X. Li, MicroRNA expression profiling and functional annotation analysis of their targets associated with the malignant transformation of oral leukoplakia. Gene 558, 271–277 (2015)
Y. Yang, Y.X. Li, X. Yang, L. Jiang, Z.J. Zhou, Y.Q. Zhu, Progress risk assessment of oral premalignant lesions with saliva miRNA analysis. BMC Cancer 13, 129 (2013)
N.K. Cervigne, P.P. Reis, J. Machado, B. Sadikovic, G. Bradley, N.N. Galloni, M. Pintilie, I. Jurisica, B. Perez-Ordonez, R. Gilbert, P. Gullane, J. Irish, S. Kamel-Reid, Identification of a microRNA signature associated with progression of leukoplakia to oral carcinoma. Hum. Mol. Genet. 18, 4818–4829 (2009)
G. Lodi, M. Carrozzo, S. Furness, K. Thongprasom, Interventions for treating oral lichen planus: A systematic review. Br. J. Dermatol. 166, 938–947 (2012)
M. Mehdipour, M. Shahidi, S. Manifar, S. Jafari, F.M. Abbas, M. Barati, H. Mortazavi, M. Shirkhoda, A. Farzanegan, Z.E. Rankohi, Diagnostic and prognostic relevance of salivary microRNA-21,-125a,-31 and-200a levels in patients with oral lichen planus - a short report. Cell. Oncol. 41, 329–334 (2018)
L. Wang, W. Wu, J. Chen, Y. Li, M. Xu, Y. Cai, MicroRNA microarray-based identification of involvement of miR-155 and miR-19a in development of Oral lichen Planus (OLP) by modulating Th1/Th2 balance via targeting eNOS and toll-like receptor 2 (TLR2). Med. Sci. Monit. 24, 3591–3603 (2018)
J.Y. Hu, J. Zhang, J.Z. Ma, X.Y. Liang, G.Y. Chen, R. Lu, G.F. Du, G. Zhou, MicroRNA-155-IFN-gamma feedback loop in CD4(+)T cells of erosive type Oral lichen Planus. Sci. Rep. 5, 16935 (2015)
J. Wang, X. Zhai, J. Guo, Y. Li, Y. Yang, L. Wang, L. Yang, F. Liu, Long non-coding RNA DQ786243 modulates the induction and function of CD4(+) Treg cells through Foxp3-miR-146a-NF-kappaB axis: Implications for alleviating oral lichen planus. Int. Immunopharmacol. 75, 105761 (2019)
J. Wang, H. Luo, Y. Xiao and L. Wang, miR-125b inhibits keratinocyte proliferation and promotes keratinocyte apoptosis in oral lichen planus by targeting MMP-2 expression through PI3K/Akt/mTOR pathway. Biomed. Pharmacother. 80, 373–380 (2016)
J. Wang, L. Yang, L. Wang, Y. Yang, Y. Wang, Forkhead box p3 controls progression of oral lichen planus by regulating microRNA-146a. J. Cell. Biochem. 119, 8862–8871 (2018)
J. Chen, Y. Wang, G. Du, W. Zhang, T. Cao, L. Shi, Y. Wang, J. Mi and G. Tang, Down-regulation of miRNA-27b-3p suppresses keratinocytes apoptosis in oral lichen planus. J. Cell. Mol. Med. 23, 4326–4337 (2019)
Q. Yang, H. Sun, X. Wang, X. Yu, J. Zhang, B. Guo, S. Hexige, Metabolic changes during malignant transformation in primary cells of oral lichen planus: Succinate accumulation and tumour suppression. J. Cell. Mol. Med. 24, 1179–1188 (2020)
N.A. Ghallab, R.F. Kasem, S.F.A. El-Ghani, O.G. Shaker, Gene expression of miRNA-138 and cyclin D1 in oral lichen planus. Clin. Oral Investig. 21, 2481–2491 (2017)
H. Zheng, S. Li, Reduced miRNA214 expression in oral mucosa contributes to the pathogenesis of oral lichen planus by targeting CD44. Mol. Med. Rep. 17, 1919–1925 (2018)
J. Chen, G. Du, Y. Chang, Y. Wang, L. Shi, J. Mi, G. Tang, Downregulated miR-27b promotes keratinocyte proliferation by targeting PLK2 in oral lichen planus. J. Oral Pathol. Med. 48, 326–334 (2019)
J. Chen, Y. Wang, G. Du, W. Zhang, T. Cao, L. Shi, Y. Wang, J. Mi, G. Tang, Down-regulation of miRNA-27b-3p suppresses keratinocytes apoptosis in oral lichen planus. J. Cell. Mol. Med. 23, 4326–4337 (2019)
D.D. Stasio, L. Mosca, A. Lucchese, D.D. Cave, H. Kawasaki, A. Lombardi, M. Porcelli, M. Caraglia, Salivary mir-27b expression in oral lichen planus patients: A series of cases and a narrative review of litterature. Curr. Top. Med. Chem. 19, 2816–2823 (2019)
W.Y. Zhang, W. Liu, Y.M. Zhou, X.M. Shen, Y.F. Wang, G.Y. Tang, Altered microRNA expression profile with miR-27b down-regulation correlated with disease activity of oral lichen planus. Oral Dis. 18, 265–270 (2012)
W. Shi, J. Yang, S. Li, X. Shan, X. Liu, H. Hua, C. Zhao, Z. Feng, Z. Cai, L. Zhang, D. Zhou, Potential involvement of miR-375 in the premalignant progression of oral squamous cell carcinoma mediated via transcription factor KLF5. Oncotarget 6, 40172–40185 (2015)
Z. Shen, G. Du, Z. Zhou, W. Liu, L. Shi, H. Xu, Aberrant expression of interleukin-22 and its targeting microRNAs in oral lichen planus: A preliminary study. J. Oral Pathol. Med. 45, 523–527 (2016)
J. Dang, Y.Q. Bian, J.Y. Sun, F. Chen, G.Y. Dong, Q. Liu, X.W. Wang, J. Kjems, S. Gao, Q.T. Wang, MicroRNA-137 promoter methylation in oral lichen planus and oral squamous cell carcinoma. J. Oral Pathol. Med. 42, 315–321 (2013)
J.G. Yang, Y.R. Sun, G.Y. Chen, X.Y. Liang, J. Zhang, G. Zhou, Different expression of MicroRNA-146a in peripheral blood CD4(+) T cells and lesions of Oral lichen Planus. Inflammation 39, 860–866 (2016)
K. Danielsson, Y.B. Wahlin, X. Gu, L. Boldrup, K. Nylander, Altered expression of miR-21, miR-125b, and miR-203 indicates a role for these microRNAs in oral lichen planus. J. Oral Pathol. Med. 41, 90–95 (2012)
M. Mehdipour, M. Shahidi, S. Manifar, S. Jafari, F. Mashhadi Abbas, M. Barati, H. Mortazavi, M. Shirkhoda, A. Farzanegan, Z. Elmi Rankohi, Diagnostic and prognostic relevance of salivary microRNA-21, −125a, −31 and -200a levels in patients with oral lichen planus - a short report. Cell. Oncol. 41, 329–334 (2018)
E. Chattopadhyay, R. Singh, A. Ray, R. Roy, N. De Sarkar, R.R. Paul, M. Pal, R. Aich, B. Roy, Expression deregulation of mir31 and CXCL12 in two types of oral precancers and cancer: Importance in progression of precancer and cancer. Sci. Rep. 6, 32735 (2016)
L. Wang, W. Wu, J. Chen, Y. Li, M. Xu and Y. Cai, miR122 and miR199 synergistically promote autophagy in oral lichen planus by targeting the Akt/mTOR pathway, Int. J. Mol. Med. 43, 1373–1381 (2019)
I. van der Waal, Potentially malignant disorders of the oral and oropharyngeal mucosa; terminology, classification and present concepts of management. Oral Oncol. 45, 317–323 (2009)
S.S. Hsue, W.C. Wang, C.H. Chen, C.C. Lin, Y.K. Chen, L.M. Lin, Malignant transformation in 1458 patients with potentially malignant oral mucosal disorders: A follow-up study based in a Taiwanese hospital. J. Oral Pathol. Med. 36, 25–29 (2007)
D. Chickooree, K. Zhu, V. Ram, H.J. Wu, Z.J. He, S. Zhang, A preliminary microarray assay of the miRNA expression signatures in buccal mucosa of oral submucous fibrosis patients. J. Oral Pathol. Med. 45, 691–697 (2016)
Y.Y. Hou, J.H. Lee, H.C. Chen, C.M. Yang, S.J. Huang, H.H. Liou, C.C. Chi, K.W. Tsai, L.P. Ger, The association between miR-499a polymorphism and oral squamous cell carcinoma progression. Oral Dis. 21, 195–206 (2015)
Y.W. Liao, C.C. Yu, P.L. Hsieh, Y.C. Chang, miR-200b ameliorates myofibroblast transdifferentiation in precancerous oral submucous fibrosis through targeting ZEB2. J. Cell. Mol. Med. 22, 4130–4138 (2018)
S. Zhou, Y. Zhu, Z. He, D. Zhang, F. Guo, X. Jian, C. Zhang, Long non-coding RNA expression profile associated with malignant progression of Oral submucous fibrosis. J. Oncol. 2019, 6835176 (2019)
C.Y. Fang, C.C. Yu, Y.W. Liao, P.L. Hsieh, M.Y. Lu, K.C. Lin, C.Z. Wu, L.L. Tsai, LncRNA LINC00974 activates TGF-beta/Smad signaling to promote oral fibrogenesis. J. Oral Pathol. Med. 48, 151–158 (2019)
C.Y. Lin, Y.W. Liao, P.L. Hsieh, M.Y. Lu, C.Y. Peng, P.M. Chu, H.W. Yang, Y.F. Huang, C.C. Yu, C.H. Yu, LncRNA GAS5-AS1 Inhibits Myofibroblasts Activities in Oral Submucous Fibrosis. J. Formos. Med. Assoc. 117, 727–733 (2018)
B. Zhang, Y. Li, D. Hou, Q. Shi, S. Yang, Q. Li, MicroRNA-375 inhibits growth and enhances Radiosensitivity in Oral squamous cell carcinoma by targeting insulin like growth factor 1 receptor. Cell. Physiol. Biochem. 42, 2105–2117 (2017)
R. Xu, G. Zeng, J. Gao, Y. Ren, Z. Zhang, Q. Zhang, J. Zhao, H. Tao, D. Li, miR-138 suppresses the proliferation of oral squamous cell carcinoma cells by targeting Yes-associated protein 1. Oncol. Rep. 34, 2171–2178 (2015)
C. Li, J. Ye, Z. Zhang, Z. Gong, Z. Lin, M. Ding, Long non-coding RNA RBM5-AS1 promotes the aggressive behaviors of oral squamous cell carcinoma by regulation of miR-1285-3p/YAP1 axis. Biomed. Pharmacother. 123, 109723 (2019)
W. Su, Y. Wang, F. Wang, B. Zhang, H. Zhang, Y. Shen, H. Yang, Circular RNA hsa_circ_0007059 indicates prognosis and influences malignant behavior via AKT/mTOR in oral squamous cell carcinoma. J. Cell. Physiol. 234, 15156–15166 (2019)
C. Liu, Z. Wang, Y. Wang, W. Gu, MiR-338 suppresses the growth and metastasis of OSCC cells by targeting NRP1. Mol. Cell. Biochem. 398, 115–122 (2015)
X. Yang, H. Wu, T. Ling, Suppressive effect of microRNA-126 on oral squamous cell carcinoma in vitro. Mol. Med. Rep. 10, 125–130 (2014)
F. Chen, S. Qi, X. Zhang, J. Wu, X. Yang, R. Wang, miR-23a-3p suppresses cell proliferation in oral squamous cell carcinomas by targeting FGF2 and correlates with a better prognosis: miR-23a-3p inhibits OSCC growth by targeting FGF2. Pathol. Res. Pract. 215, 660–667 (2019)
M. Li, J. Ning, Z. Li, Q. Fei, C. Zhao, Y. Ge, L. Wang, Long noncoding RNA OIP5-AS1 promotes the progression of oral squamous cell carcinoma via regulating miR-338-3p/NRP1 axis. Biomed. Pharmacother. 118, 109259 (2019)
Y. Shao, Y. Qu, S. Dang, B. Yao, M. Ji, MiR-145 inhibits oral squamous cell carcinoma (OSCC) cell growth by targeting c-Myc and Cdk6. Cancer Cell Int. 13, 51 (2013)
S. Fu, H.H. Chen, P. Cheng, C.B. Zhang, Y. Wu, MiR-155 regulates oral squamous cell carcinoma Tca8113 cell proliferation, cycle, and apoptosis via regulating p27Kip1. Eur. Rev. Med. Pharmacol. Sci. 21, 937–944 (2017)
H.B. Xing, H.M. Qiu, Y. Li, P.F. Dong, X.M. Zhu, Long noncoding RNA CASC2 alleviates the growth, migration and invasion of oral squamous cell carcinoma via downregulating CDK1. Eur. Rev. Med. Pharmacol. Sci. 23, 4777–4783 (2019)
H. Chi, miR-194 regulated AGK and inhibited cell proliferation of oral squamous cell carcinoma by reducing PI3K-Akt-FoxO3a signaling. Biomed. Pharmacother. 71, 53–57 (2015)
B. Rastogi, A. Kumar, S.K. Raut, N.K. Panda, V. Rattan, N. Joshi, M. Khullar, Downregulation of miR-377 promotes Oral squamous cell carcinoma growth and migration by targeting HDAC9. Cancer Investig. 35, 152–162 (2017)
X. Niu, B. Yang, F. Liu, Q. Fang, LncRNA HOXA11-AS promotes OSCC progression by sponging miR-98-5p to upregulate YBX2 expression. Biomed. Pharmacother. 121, 109623 (2020)
Z. Wang, X. Zhu, P. Dong and J. Cai, Long noncoding RNA LINC00958 promotes the oral squamous cell carcinoma by sponging miR-185-5p/YWHAZ. Life Sci. 242, 116782 (2020)
J. Tan, L. Xiang, G. Xu, LncRNA MEG3 suppresses migration and promotes apoptosis by sponging miR-548d-3p to modulate JAK-STAT pathway in oral squamous cell carcinoma. IUBMB Life 71, 882–890 (2019)
L. Wang, Y. Wei, Y. Yan, H. Wang, J. Yang, Z. Zheng, J. Zha, P. Bo, Y. Tang, X. Guo, W. Chen, X. Zhu, L. Ge, CircDOCK1 suppresses cell apoptosis via inhibition of miR196a5p by targeting BIRC3 in OSCC. Oncol. Rep. 39, 951–966 (2018)
L. Chen, S. Zhang, J. Wu, J. Cui, L. Zhong, L. Zeng, S. Ge, circRNA_100290 plays a role in oral cancer by functioning as a sponge of the miR-29 family. Oncogene 36, 4551–4561 (2017)
P. Xu, Y. Li, H. Zhang, M. Li, H. Zhu, MicroRNA-340 mediates metabolic shift in Oral squamous cell carcinoma by targeting glucose Transporter-1. J. Oral Maxillofac. Surg. 74, 844–850 (2016)
X. Sun, L. Zhang, MicroRNA-143 suppresses oral squamous cell carcinoma cell growth, invasion and glucose metabolism through targeting hexokinase 2. Biosci. Rep. 37, pii: BSR20160404 (2017)
Y.H. Lai, H. Liu, W.F. Chiang, T.W. Chen, L.J. Chu, J.S. Yu, S.J. Chen, H.C. Chen, B.C. Tan, MiR-31-5p-ACOX1 Axis enhances tumorigenic fitness in Oral squamous cell carcinoma via the Promigratory prostaglandin E2. Theranostics 8, 486–504 (2018)
H. Chu, Z. Li, Z. Gan, Z. Yang, Z. Wu, M. Rong, LncRNA ELF3-AS1 is involved in the regulation of oral squamous cell carcinoma cell proliferation by reprogramming glucose metabolism. Onco. Targets Ther 12, 6857–6863 (2019)
C. Gao, C. Ren, Z. Liu, L. Zhang, R. Tang, X. Li, GAS5, a FoxO1-actived long noncoding RNA, promotes propofol-induced oral squamous cell carcinoma apoptosis by regulating the miR-1297-GSK3beta axis. Artif. Cells Nanomed. Biotechnol. 47, 3985–3993 (2019)
Z. Zheng, X. Luan, J. Zha, Z. Li, L. Wu, Y. Yan, H. Wang, D. Hou, L. Huang, F. Huang, H. Zheng, L. Ge, H. Guan, TNF-alpha inhibits the migration of oral squamous cancer cells mediated by miR-765-EMP3-p66Shc axis. Cell. Signal. 34, 102–109 (2017)
Y. Wu, L. Zhang, L. Zhang, Y. Wang, H. Li, X. Ren, F. Wei, W. Yu, T. Liu, X. Wang, X. Zhou, J. Yu, X. Hao, Long non-coding RNA HOTAIR promotes tumor cell invasion and metastasis by recruiting EZH2 and repressing E-cadherin in oral squamous cell carcinoma. Int. J. Oncol. 46, 2586–2594 (2015)
L. Lu, X. Xue, J. Lan, Y. Gao, Z. Xiong, H. Zhang, W. Jiang, W. Song, Q. Zhi, MicroRNA-29a upregulates MMP2 in oral squamous cell carcinoma to promote cancer invasion and anti-apoptosis. Biomed. Pharmacother. 68, 13–19 (2014)
X.P. Kong, J. Yao, W. Luo, F.K. Feng, J.T. Ma, Y.P. Ren, D.L. Wang, R.F. Bu, The expression and functional role of a FOXC1 related mRNA-lncRNA pair in oral squamous cell carcinoma. Mol. Cell. Biochem. 394, 177–186 (2014)
Y. Wang, X. Zhang, Z. Wang, Q. Hu, J. Wu, Y. Li, X. Ren, T. Wu, X. Tao, X. Chen, X. Li, J. Xia, B. Cheng, LncRNA-p23154 promotes the invasion-metastasis potential of oral squamous cell carcinoma by regulating Glut1-mediated glycolysis. Cancer Lett. 434, 172–183 (2018)
Z. Zhuang, Nan Xie, J. Hu, P. Yu, C. Wang, X. Hu, X. Han, J. Hou, H. Huang and X. Liu, Interplay between ΔNp63 and miR-138-5p regulates growth, metastasis and stemness of oral squamous cell carcinoma. Oncotarget 8, 21954-21973 (2017)
Y. Tian, L. Zhong, S. Gao, Y. Yu, D. Sun, X. Liu, J. Ji, Y. Yao, Y. Liu, Z. Jiang, LncRNA LINC00974 Downregulates miR-122 to Upregulate RhoA in Oral squamous cell carcinoma. Cancer Biother. Radiopharm. (2019). https://doi.org/10.1089/cbr.2019.2907
G. Huang, X. He, X.L. Wei, lncRNA NEAT1 promotes cell proliferation and invasion by regulating miR365/RGS20 in oral squamous cell carcinoma. Oncol. Rep. 39, 1948–1956 (2018)
Y. Hui, Y. Li, Y. Jing, J.Q. Feng, Y. Ding, miRNA-101 acts as a tumor suppressor in oral squamous cell carcinoma by targeting CX chemokine receptor 7. Am. J. Transl. Res. 8, 4902–4911 (2016)
S.C. Casey, A. Amedei, K. Aquilano, A.S. Azmi, F. Benencia, D. Bhakta, A.E. Bilsland, C.S. Boosani, S. Chen, M.R. Ciriolo, S. Crawford, H. Fujii, A.G. Georgakilas, G. Guha, D. Halicka, W.G. Helferich, P. Heneberg, K. Honoki, W.N. Keith, S.P. Kerkar, S.I. Mohammed, E. Niccolai, S. Nowsheen, H.P.V. Rupasinghe, A. Samadi, N. Singh, W.H. Talib, V. Venkateswaran, R.L. Whelan, X. Yang, D.W. Felsher, Cancer prevention and therapy through the modulation of the tumor microenvironment. Semin. Cancer Biol. 35(Suppl), S199–S223 (2015)
P. Nilendu, S.C. Sarode, D. Jahagirdar, I. Tandon, S. Patil, G.S. Sarode, J.K. Pal, N.K. Sharma, Mutual concessions and compromises between stromal cells and cancer cells: Driving tumor development and drug resistance. Cell. Oncol. 41, 353–367 (2018)
A. Salmaninejad, S.F. Valilou, A. Soltani, S. Ahmadi, Y.J. Abarghan, R.J. Rosengren, A. Sahebkar, Tumor-associated macrophages: Role in cancer development and therapeutic implications. Cell. Oncol. 42, 591–608 (2019)
T. Sasahira, M. Kurihara, U.K. Bhawal, N. Ueda, T. Shimomoto, K. Yamamoto, T. Kirita, H. Kuniyasu, Downregulation of miR-126 induces angiogenesis and lymphangiogenesis by activation of VEGF-A in oral cancer. Br. J. Cancer 107, 700–706 (2012)
Y.Y. Wu, Y.L. Chen, Y.C. Jao, I.S. Hsieh, K.C. Chang, T.M. Hong, miR-320 regulates tumor angiogenesis driven by vascular endothelial cells in oral cancer by silencing neuropilin 1. Angiogenesis 17, 247–260 (2014)
L. Li, C. Li, S. Wang, Z. Wang, J. Jiang, W. Wang, X. Li, J. Chen, K. Liu, C. Li, G. Zhu, Exosomes derived from hypoxic Oral squamous cell carcinoma cells deliver miR-21 to normoxic cells to elicit a Prometastatic phenotype. Cancer Res. 76, 1770–1780 (2016)
T. Kawakubo-Yasukochi, M. Morioka, M. Hazekawa, A. Yasukochi, T. Nishinakagawa, K. Ono, S. Kawano, S. Nakamura, M. Nakashima, miR-200c-3p spreads invasive capacity in human oral squamous cell carcinoma microenvironment. Mol. Carcinog. 57, 295–302 (2018)
T. Liu, G. Chen, D. Sun, M. Lei, Y. Li, C. Zhou, X. Li, W. Xue, H. Wang, C. Liu, J. Xu, Exosomes containing miR-21 transfer the characteristic of cisplatin resistance by targeting PTEN and PDCD4 in oral squamous cell carcinoma. Acta Biochim. Biophys. Sin. Shanghai 49, 808–816 (2017)
N. Eiro, L. Gonzalez, A. Martinez-Ordonez, B. Fernandez-Garcia, L.O. Gonzalez, S. Cid, F. Dominguez, R. Perez-Fernandez, F.J. Vizoso, Cancer-associated fibroblasts affect breast cancer cell gene expression, invasion and angiogenesis. Cell. Oncol. 41, 369–378 (2018)
Y.Y. Li, Y.W. Tao, S. Gao, P. Li, J.M. Zheng, S.E. Zhang, J. Liang, Y. Zhang, Cancer-associated fibroblasts contribute to oral cancer cells proliferation and metastasis via exosome-mediated paracrine miR-34a-5p. EBioMedicine 36, 209–220 (2018)
L.P. Sun, K. Xu, J. Cui, D.Y. Yuan, B. Zou, J. Li, J.L. Liu, K.Y. Li, Z. Meng, B. Zhang, Cancerassociated fibroblastderived exosomal miR3825p promotes the migration and invasion of oral squamous cell carcinoma. Oncol. Rep. 42, 1319–1328 (2019)
J. Cai, B. Qiao, N. Gao, N. Lin, W. He, Oral squamous cell carcinoma-derived exosomes promote M2 subtype macrophage polarization mediated by exosome-enclosed miR-29a-3p. Am. J. Physiol. Cell Physiol. 316, C731–c740 (2019)
F. Momen-Heravi, S. Bala, Extracellular vesicles in oral squamous carcinoma carry oncogenic miRNA profile and reprogram monocytes via NF-kappaB pathway. Oncotarget 9, 34838–34854 (2018)
D. Huang, J. Chen, L. Yang, Q. Ouyang, J. Li, L. Lao, J. Zhao, J. Liu, Y. Lu, Y. Xing, F. Chen, F. Su, H. Yao, Q. Liu, S. Su, E. Song, NKILA lncRNA promotes tumor immune evasion by sensitizing T cells to activation-induced cell death. Nat. Immunol. 19, 1112–1125 (2018)
L.J. Sang, H.Q. Ju, G.P. Liu, T. Tian, G.L. Ma, Y.X. Lu, Z.X. Liu, R.L. Pan, R.H. Li, H.L. Piao, J.R. Marks, L.J. Yang, Q. Yan, W. Wang, J. Shao, Y. Zhou, T. Zhou, A. Lin, LncRNA CamK-A Regulates Ca(2+)-Signaling-Mediated Tumor Microenvironment Remodeling. Mol. Cell 72, 71–83.e77 (2018)
Z.L. Ou, Z. Luo, W. Wei, S. Liang, T.L. Gao, Y.B. Lu, Hypoxia-induced shedding of MICA and HIF1A-mediated immune escape of pancreatic cancer cells from NK cells: Role of circ_0000977/miR-153 axis. RNA Biol. 16, 1592–1603 (2019)
F. Zahran, D. Ghalwash, O. Shaker, K. Al-Johani, C. Scully, Salivary microRNAs in oral cancer. Oral Dis. 21, 739–747 (2015)
E.A. Gibb, K.S. Enfield, G.L. Stewart, K.M. Lonergan, R. Chari, R.T. Ng, L. Zhang, C.E. MacAulay, M.P. Rosin, W.L. Lam, Long non-coding RNAs are expressed in oral mucosa and altered in oral premalignant lesions. Oral Oncol. 47, 1055–1061 (2011)
Y.C. Lu, J.T. Chang, Y.C. Huang, C.C. Huang, W.H. Chen, L.Y. Lee, B.S. Huang, Y.J. Chen, H.F. Li, A.J. Cheng, Combined determination of circulating miR-196a and miR-196b levels produces high sensitivity and specificity for early detection of oral cancer. Clin. Biochem. 48, 115–121 (2015)
F. Momen-Heravi, A.J. Trachtenberg, W.P. Kuo, Y.S. Cheng, Genomewide Study of Salivary MicroRNAs for Detection of Oral Cancer. J. Dent. Res. 93, 86s–93s (2014)
C. Zhao, H. Zou, J. Wang, J. Shen, H. Liu, A three long noncoding RNA-based signature for Oral squamous cell carcinoma prognosis prediction. DNA Cell Biol. 37, 888–895 (2018)
S.Y. Zhao, J. Wang, S.B. Ouyang, Z.K. Huang, L. Liao, Salivary circular RNAs Hsa_Circ_0001874 and Hsa_Circ_0001971 as novel biomarkers for the diagnosis of Oral squamous cell carcinoma. Cell. Physiol. Biochem. 47, 2511–2521 (2018)
J. Li, H. Huang, L. Sun, M. Yang, C. Pan, W. Chen, D. Wu, Z. Lin, C. Zeng, Y. Yao, P. Zhang, E. Song, MiR-21 indicates poor prognosis in tongue squamous cell carcinomas as an apoptosis inhibitor. Clin. Cancer Res. 15, 3998–4008 (2009)
X. Li, C. Ma, L. Zhang, N. Li, X. Zhang, J. He, R. He, M. Shao, J. Wang, L. Kang, C. Han, LncRNAAC132217.4, a KLF8-regulated long non-coding RNA, facilitates oral squamous cell carcinoma metastasis by upregulating IGF2 expression. Cancer Lett. 407, 45–56 (2017)
X. Xia, X. Li, F. Li, X. Wu, M. Zhang, H. Zhou, N. Huang, X. Yang, F. Xiao, D. Liu, L. Yang, N. Zhang, A novel tumor suppressor protein encoded by circular AKT3 RNA inhibits glioblastoma tumorigenicity by competing with active phosphoinositide-dependent Kinase-1. Mol. Cancer 18, 131 (2019)
G. Yuan, H. Wu, Y. Du, F. He, Tumor suppressor role of microRNA-545 in oral squamous cell carcinoma. Oncol. Lett. 17, 2063–2068 (2019)
L.O. Reis, T.C. Pereira, I. Lopes-Cendes, U. Ferreira, MicroRNAs: A new paradigm on molecular urological oncology. Urology 76, 521–527 (2010)
Z. Fang, J. Zhao, W. Xie, Q. Sun, H. Wang, B. Qiao, LncRNA UCA1 promotes proliferation and cisplatin resistance of oral squamous cell carcinoma by sunppressing miR-184 expression. Cancer Med. 6, 2897–2908 (2017)
S. Hunt, A.V. Jones, E.E. Hinsley, S.A. Whawell, D.W. Lambert, MicroRNA-124 suppresses oral squamous cell carcinoma motility by targeting ITGB1. FEBS Lett. 585, 187–192 (2011)
A. Tiwari, S. Shivananda, K.S. Gopinath, A. Kumar, MicroRNA-125a reduces proliferation and invasion of oral squamous cell carcinoma cells by targeting estrogen-related receptor alpha: Implications for cancer therapeutics. J. Biol. Chem. 289, 32276–32290 (2014)
A. Bufalino, N.K. Cervigne, C.E. de Oliveira, F.P. Fonseca, P.C. Rodrigues, C.C. Macedo, L.M. Sobral, M.C. Miguel, M.A. Lopes, A.F. Paes Leme, D.W. Lambert, T.A. Salo, L.P. Kowalski, E. Graner, R.D. Coletta, Low miR-143/miR-145 cluster levels induce Activin a overexpression in Oral squamous cell carcinomas, which contributes to poor prognosis. PLoS One 10, e0136599 (2015)
Q. Zeng, X. Tao, F. Huang, T. Wu, J. Wang, X. Jiang, Z. Kuang, B. Cheng, Overexpression of miR-155 promotes the proliferation and invasion of oral squamous carcinoma cells by regulating BCL6/cyclin D2. Int. J. Mol. Med. 37, 1274–1280 (2016)
K.H. Shin, S.D. Bae, H.S. Hong, R.H. Kim, M.K. Kang, N.H. Park, miR-181a shows tumor suppressive effect against oral squamous cell carcinoma cells by downregulating K-ras. Biochem. Biophys. Res. Commun. 404, 896–902 (2011)
Z. Cai, X.Y. Hao, F.X. Liu, MicroRNA-186 serves as a tumor suppressor in oral squamous cell carcinoma by negatively regulating the protein tyrosine phosphatase SHP2 expression. Arch. Oral Biol. 89, 20–25 (2018)
L. Li, H.Q. Ma, MicroRNA-216a inhibits the growth and metastasis of oral squamous cell carcinoma by targeting eukaryotic translation initiation factor 4B. Mol. Med. Rep. 12, 3156–3162 (2015)
A. Uesugi, K. Kozaki, T. Tsuruta, M. Furuta, K. Morita, I. Imoto, K. Omura, J. Inazawa, The tumor suppressive microRNA miR-218 targets the mTOR component Rictor and inhibits AKT phosphorylation in oral cancer. Cancer Res. 71, 5765–5778 (2011)
F. Jiang, W. Zhao, L. Zhou, L. Zhang, Z. Liu, D. Yu, miR-222 regulates the cell biological behavior of oral squamous cell carcinoma by targeting PUMA. Oncol. Rep. 31, 1255–1262 (2014)
C.N. Yang, Y.T. Deng, J.Y. Tang, S.J. Cheng, S.T. Chen, Y.J. Li, T.S. Wu, M.H. Yang, B.R. Lin, M.Y. Kuo, J.Y. Ko, C.C. Chang, MicroRNA-29b regulates migration in oral squamous cell carcinoma and its clinical significance. Oral Oncol. 51, 170–177 (2015)
X. Yang, H. Ruan, X. Hu, A. Cao, L. Song, miR-381-3p suppresses the proliferation of oral squamous cell carcinoma cells by directly targeting FGFR2, am. J. Cancer Res. Ther. 7, 913–922 (2017)
H.Y. Peng, S.S. Jiang, J.R. Hsiao, M. Hsiao, Y.M. Hsu, G.H. Wu, W.M. Chang, J.Y. Chang, S.L. Jin, S.G. Shiah, IL-8 induces miR-424-5p expression and modulates SOCS2/STAT5 signaling pathway in oral squamous cell carcinoma. Mol. Oncol. 10, 895–909 (2016)
C.M. Cheng, S.G. Shiah, C.C. Huang, J.R. Hsiao, J.Y. Chang, Up-regulation of miR-455-5p by the TGF-beta-SMAD signalling axis promotes the proliferation of oral squamous cancer cells by targeting UBE2B. J. Pathol. 240, 38–49 (2016)
J. Hu, J.F. Xu, W.L. Ge, MiR-497 enhances metastasis of oral squamous cell carcinoma through SMAD7 suppression. Am. J. Transl. Res. 8, 3023–3031 (2016)
T. Yu, K. Liu, Y. Wu, J. Fan, J. Chen, C. Li, Q. Yang, Z. Wang, MicroRNA-9 inhibits the proliferation of oral squamous cell carcinoma cells by suppressing expression of CXCR4 via the Wnt/beta-catenin signaling pathway. Oncogene 33, 5017–5027 (2014)
Y. Du, Y. Li, H. Lv, S. Zhou, Z. Sun, M. Wang, miR-98 suppresses tumor cell growth and metastasis by targeting IGF1R in oral squamous cell carcinoma. Int. J. Clin. Exp. Pathol. 8, 12252–12259 (2015)
Y.C. Yen, S.G. Shiah, H.C. Chu, Y.M. Hsu, J.R. Hsiao, J.Y. Chang, W.C. Hung, C.T. Liao, A.J. Cheng, Y.C. Lu, Y.W. Chen, Reciprocal regulation of microRNA-99a and insulin-like growth factor I receptor signaling in oral squamous cell carcinoma cells. Mol. Cancer 13, 6 (2014)
T.R. Shao, Z.N. Zheng, Y.C. Chen, Q.Q. Wu, G.Z. Huang, F. Li, W.S. Zeng, X.Z. Lv, LncRNA AC007271.3 promotes cell proliferation, invasion, migration and inhibits cell apoptosis of OSCC via the Wnt/beta-catenin signaling pathway. Life Sci. 239, 117087 (2019)
L. Liu, S.B. Ning, S. Fu, Y. Mao, M. Xiao, B. Guo, Effects of lncRNA ANRIL on proliferation and apoptosis of oral squamous cell carcinoma cells by regulating TGF-beta/Smad pathway. Eur. Rev. Med. Pharmacol. Sci. 23, 6194–6201 (2019)
C. Yao, F. Kong, S. Zhang, G. Wang, P. She, Q. Zhang, Long non-coding RNA BANCR promotes proliferation and migration in oral squamous cell carcinoma via MAPK signaling pathway. J. Oral Pathol. Med. (2019). https://doi.org/10.1111/jop.12968
D. Dai, X.D. Feng, W.Q. Zhu, Y.N. Bao, LncRNA BLACAT1 regulates the viability, migration and invasion of oral squamous cell carcinoma cells by targeting miR-142-5p. Eur. Rev. Med. Pharmacol. Sci. 23, 10313–10323 (2019)
X. Zhang, B. Guo, Y. Zhu, W. Xu, S. Ning, L. Liu, Up-regulation of plasma lncRNA CACS15 distinguished early-stage oral squamous cell carcinoma patient. Oral Dis. (2019). https://doi.org/10.1111/odi.13245
J. Ye, Y. Jiao, LncRNA FAL1 promotes the development of oral squamous cell carcinoma through regulating the microRNA-761/CRKL pathway. Eur. Rev. Med. Pharmacol. Sci. 23, 5779–5786 (2019)
B. Jia, T. Xie, X. Qiu, X. Sun, J. Chen, Z. Huang, X. Zheng, Z. Wang, J. Zhao, Long noncoding RNA FALEC inhibits proliferation and metastasis of tongue squamous cell carcinoma by epigenetically silencing ECM1 through EZH2. Aging (Albany NY) 11, 4990–5007 (2019)
L. Xu, T.J. Hou, P. Yang, Mechanism of lncRNA FEZF1-AS1 in promoting the occurrence and development of oral squamous cell carcinoma through targeting miR-196a. Eur. Rev. Med. Pharmacol. Sci. 23, 6505–6515 (2019)
Y. Hong, H. He, W. Sui, J. Zhang, S. Zhang, D. Yang, Long non-coding RNA H1 promotes cell proliferation and invasion by acting as a ceRNA of miR138 and releasing EZH2 in oral squamous cell carcinoma. Int. J. Oncol. 52, 901–912 (2018)
J. Zhao, X. Bai, C. Feng, X. Shang, Y. Xi, Long non-coding RNA HCP5 facilitates cell invasion and epithelial-Mesenchymal transition in Oral squamous cell carcinoma by miR-140-5p/SOX4 Axis. Cancer Manag. Res. 11, 10455–10462 (2019)
M. Li, J. Ning, Z. Li, J. Wang, C. Zhao, L. Wang, LINC00152 promotes the growth and invasion of oral squamous cell carcinoma by regulating miR-139-5p. Onco. Targets Ther. 11, 6295–6304 (2018)
Y. Jiang, W. Cao, K. Wu, X. Qin, X. Wang, Y. Li, B. Yu, Z. Zhang, X. Wang, M. Yan, Q. Xu, J. Zhang, W. Chen, LncRNA LINC00460 promotes EMT in head and neck squamous cell carcinoma by facilitating peroxiredoxin-1 into the nucleus. J. Exp. Clin. Cancer Res.: CR 38, 365 (2019)
C.Z. Zhang, Long intergenic non-coding RNA 668 regulates VEGFA signaling through inhibition of miR-297 in oral squamous cell carcinoma. Biochem. Biophys. Res. Commun. 489, 404–412 (2017)
Z. Chen, Q. Tao, B. Qiao, L. Zhang, Silencing of LINC01116 suppresses the development of oral squamous cell carcinoma by up-regulating microRNA-136 to inhibit FN1. Cancer Manag. Res. 11, 6043–6059 (2019)
Y. Kong, Y. Feng, Y.Y. Xiao, S.C. Liu, X.G. Li, Q.L. Yang, W.H. Chu, J.G. Liu, LncRNA LUCAT1 promotes rowth, migration, and invasion of oral squamous cell carcinoma by upregulating PCNA. Eur. Rev. Med. Pharmacol. Sci. 23, 4770–4776 (2019)
S.M. Chang, W.W. Hu, Long non-coding RNA MALAT1 promotes oral squamous cell carcinoma development via microRNA-125b/STAT3 axis. J. Cell. Physiol. 233, 3384–3396 (2018)
C.C. Sun, L. Zhang, G. Li, S.J. Li, Z.L. Chen, Y.F. Fu, F.Y. Gong, T. Bai, D.Y. Zhang, Q.M. Wu, D.J. Li, The lncRNA PDIA3P interacts with miR-185-5p to modulate Oral squamous cell carcinoma progression by targeting Cyclin D2. Mol. Ther. Nucleic. Acids 9, 100–110 (2017)
J. Wu, W. Zhao, Z. Wang, X. Xiang, S. Zhang, L. Liu, Long non-coding RNA SNHG20 promotes the tumorigenesis of oral squamous cell carcinoma via targeting miR-197/LIN28 axis. J. Cell. Mol. Med. 23, 680–688 (2019)
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Nos. 81672675, 81872211 and 81621062), the 111 Project of MOE China (Grant No. B14038) and the Sichuan University Innovation and Entrepreneurship Training Program for Undergraduates (No. C2019106455).
Author information
Authors and Affiliations
Contributions
FH wrote the manuscript and designed the figures; CX, HB and KL collected the related references and edited the manuscript; QC and JL provided guidance and revised this manuscript. All authors approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Huang, F., Xin, C., Lei, K. et al. Noncoding RNAs in oral premalignant disorders and oral squamous cell carcinoma. Cell Oncol. 43, 763–777 (2020). https://doi.org/10.1007/s13402-020-00521-9
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13402-020-00521-9