Abstract
The current classification system for the recognition of taxonomic ranks among fungi, especially at high-ranking level, is subjective. With the development of molecular approaches and the availability of fossil calibration data, the use of divergence times as a universally standardized criterion for ranking taxa has now become possible. We can therefore date the origin of Ascomycota lineages by using molecular clock methods and establish the divergence times for the orders and families of Dothideomycetes. We chose Dothideomycetes, the largest class of the phylum Ascomycota, which contains 32 orders, to establish ages at which points orders have split; and Pleosporales, the largest order of Dothideomycetes with 55 families, to establish family divergence times. We have assembled a multi-gene data set (LSU, SSU, TEF1 and RPB2) from 391 taxa representing most family groups of Dothideomycetes and utilized fossil calibration points solely from within the ascomycetes and a Bayesian approach to establish divergence times of Dothideomycetes lineages. Two separated datasets were analysed: (i) 272 taxa representing 32 orders of Dothideomycetes were included for the order level analysis, and (ii) 191 taxa representing 55 families of Pleosporales were included for the family level analysis. Our results indicate that divergence times (crown age) for most orders (20 out of 32, or 63%) are between 100 and 220 Mya, while divergence times for most families (39 out of 55, or 71%) are between 20 and 100 Mya. We believe that divergence times can provide additional evidence to support establishment of higher level taxa, such as families, orders and classes. Taking advantage of this added approach, we can strive towards establishing a standardized taxonomic system both within and outside Fungi. In this study we found that molecular dating coupled with phylogenetic inferences provides no support for the taxonomic status of two currently recognized orders, namely Bezerromycetales and Wiesneriomycetales and these are treated as synonyms of Tubeufiales while Asterotexiales is treated as a synonym of Asterinales. In addition, we provide an updated phylogenetic assessment of Dothideomycetes previously published as the Families of Dothideomycetes in 2013 with a further ten orders and 35 families.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Attempts to classify fungi at the higher levels have always been subjective (Liu et al. 2016; Hyde et al. 2017), especially when based solely on morphology and in some cases has resulted in unnecessary personalized conflicts in the literature. Mycologists have made attempts to establish more reliable ways to resolve taxa at the ordinal, familial and generic, as well as species levels (Liu et al. 2016; Jeewon and Hyde 2016; Divakar et al. 2017). Lumbsch and Huhndorf (2010) provided Outline of Ascomycota, which took into account molecular phylogenies, and this was followed by outlines of individual classes which also incorporated both morphology and phylogenetic data (e.g. Families of Dothideomycetes, Hyde et al. 2013; Outline of Dothideomycetes, Wijayawardene et al. 2014; Outline of Sordariomycetes, Maharachchikumbura et al. 2015; Families of Sordariomycetes, Maharachchikumbura et al. 2016). However, there have been many disagreements at various taxonomic levels, as the use of phylogenetic data may also be subjective and depends on the taxa analysed. For example, based on molecular data, Crous et al. (2017) accepted nine families in Botryosphaeriales, while Liu et al. (2016) suggested that this is excessive, as they considered that three of the monotypic families could in fact be treated as genera. This clearly indicates that assigning taxa to specific ranks can largely be subjected to personal interpretations. Therefore, further evidence is needed to try to resolve these problematic groups. Hyde et al. (2017) have reviewed the grouping and ranking of fungi and their classification, and we refer to their review in this paper and do not discuss this further.
The use of divergence times has recently been applied in fungi to support the ranking of taxa, particularly at the higher levels. Zhao et al. (2016) used divergence times to resolve subgenera and sections in the large genus Agaricus. Mapook et al. (2016) used it to support the introduction of the new family Palawaniaceae, while Phukhamsakda et al. (2016) provided additional support for the introduction of the family Longipedicellataceae with evidence from divergence times. Samarakoon et al. (2016) reassessed the Xylariales and reported that the previously discarded Amphisphaeriales evolved between 147 and 168 Mya and should be reinstated as an order. This dating approach is relatively new and is only now being applied to provide additional evidence for ranking taxa (Pérez-Ortega et al. 2016; Zhao et al. 2016, 2017; Hyde et al. 2017; Divakar et al. 2017).
Dothideomycetes is the largest class of Ascomycota, with an estimated 19,000 species (Kirk et al. 2008). The class has a worldwide distribution, ranging from tropical rainforests, to temperate broad-leaved forests, and to deserts, and life modes that are saprobic, pathogenic or endophytic, and they can be found in terrestrial and aquatic habitats (Crous et al. 2006, Boehm et al. 2009; Schoch et al. 2009a; Suetrong et al. 2009; Tanaka et al. 2009; Zhang et al. 2009; Liu et al. 2011, 2012; Pérez-Ortega et al. 2014, 2016; Phookamsak et al. 2014; Jaklitsch et al. 2015; Raja et al. 2015; Tanaka et al. 2015; Bezerra et al. 2017). Despite some recent taxonomic controversies, Dothideomycetes is an ideal group to study and apply divergence time estimates for higher rankings for the following reasons: (1) a broad taxon sampling is possible; (2) classification and phylogeny have been well investigated (Boehm et al. 2009; Crous et al. 2009; Mugambi and Huhndorf 2009; Nelsen et al. 2009; Shearer et al. 2009; Schoch et al. 2009a; Liu et al. 2012; Hyde et al. 2013; Wijayawardene et al. 2014; Jaklitsch et al. 2015); (3) sufficient DNA sequence data are available from different gene regions to allow phylogenetic inferences to be made (Boehm et al. 2009; Crous et al. 2009; Mugambi and Huhndorf 2009; Nelsen et al. 2009; Shearer et al. 2009; Schoch et al. 2009a; Suetrong et al. 2009; Liu et al. 2012, 2014, 2015; Zhang et al. 2012; Ariyawansa et al. 2014, 2015; Boonmee et al. 2014, 2016; Jaklitsch et al. 2015; Raja et al. 2015; Tanaka et al. 2015); (4) morphological classification criteria are well understood (Crous et al. 2006, 2009; Mugambi and Huhndorf 2009; Nelsen et al. 2009; Shearer et al. 2009; Suetrong et al. 2009; Zhang et al. 2012; Hyde et al. 2013); (5) an increase in the number of species being described and establishment of new orders, families and genera that might be subjected to differences in perceptions (Boehm et al. 2009; Tanaka et al. 2009, 2015; Ariyawansa et al. 2014, 2015; Boonmee et al. 2014, 2016; Hongsanan et al. 2014; Liu et al. 2014, 2015; Pérez-Ortega et al. 2014, 2016; Phookamsak et al. 2014, 2015; Jaklitsch et al. 2015; Raja et al. 2015; Thambugala et al. 2015; Tian et al. 2015; Li et al. 2016; Hyde et al. 2016; Bezerra et al. 2017).
In this paper, we provide an updated multi-locus phylogeny of the class Dothideomycetes with sufficient number of representatives of each order and for the order Pleosporales representatives of its families to unravel evolutionary relationships and assess the reliability of molecular dating in assigning taxa to ordinal and familial ranks. In addition, interordinal and interfamilial phylogenetic relationships were investigated to clarify taxonomic ambiguities. Analyses including a broad taxon sampling of the Dothideomycetes covering a wide range of taxa were performed to obtain a reliable time scale that can be used to establish ordinal and familial ranks. The applicability of using divergence time estimates in classification is discussed. We also discuss some cases where there are conflicts between phylogeny and divergence times.
Materials and methods
We use small subunits ribosomal RNA (SSU), large subunit ribosomal RNA (LSU), the translation elongation factor-1 alpha (TEF1) and the second largest subunit of RNA polymerase II (RPB2) were used as implemented in a previous study by Hyde et al. (2013). Sequences were obtained from GenBank following mostly previous publications (e.g. Schoch et al. 2006, 2009a; Suetrong et al. 2009; Zhang et al. 2012; Liu et al. 2012, 2014; Hyde et al. 2013, Pérez-Ortega et al. 2014, 2016; Jaklitsch et al. 2015; Raja et al. 2015; Tanaka et al. 2015; Tian et al. 2015; Bezerra et al. 2017) and these are listed in Supplementary Table 1.
Datasets for each gene (SSU, LSU, TEF1 and RPB2) were aligned separately with MAFFT version 6 (Katoh and Toh 2008) with subsequent manual adjustment in BioEdit 5.0.9 (Hall 1999). The software package jModeltest2.1.1 was used to select the best-fitting models of nucleotide substitution for each gene. The Bayesian information criterion supported the GTR + G + I model as the best fit for SSU, LSU and TEF1 and GTR + G for RPB2. Topological congruence of the four datasets was checked by visual comparison of phylogenetic trees obtained from maximum likelihood-based analysis with RAxML (Stamatakis et al. 2008), and a general time-reversible model (GTR) was applied with a discrete gamma distribution and four rate classes. Fifty thorough ML tree searches were done in RAxML v.7.2.7 under the same model. One thousand non-parametric bootstrap iterations were run with the GTR model and a discrete gamma distribution. And all genes were subsequently combined in a super matrix using BioEdit 5.0.9. Bayesian analyses were performed by using Markov chain Monte Carlo (MCMC) sampling in MrBayes v3.1.2 (Ronquist and Huelsenbeck 2003, Zhaxybayeva and Gogarten 2002) to generate a reasonable starting tree for subsequent analyses of divergence date estimates in BEAST. Four simultaneous Markov chains were run for 10,000,000 generations and trees were sampled every 1000th generation, thus 10,000 trees were obtained. The suitable burn-in phases were determined by inspecting likelihoods and parameters in Tracer version 1.6 (Rambaut et al. 2013). Based on the tracer analysis, the first 1000 trees representing 10%, were discarded as the burn-in phase in the analysis. The remaining trees were used to calculate posterior probabilities in the majority rule consensus tree (critical value for the topological convergence diagnostic set to 0.01).
Divergence time analyses were estimated using BEAST v1.8 (Drummond et al. 2012). Aligned sequence data were partitioned separately for each LSU, SSU, TEF1 and RPB2 data set, and loaded to prepare an XML file constructed with BEAUTI v1.8. Clock and substitution models were set to be unlinked (independently estimated for each gene partition), while the tree prior parameters were set to be linked across partitions (concatenation). An uncorrelated relaxed clock model (Drummond et al. 2006) with a lognormal distribution of rates for each gene estimate was used for the analyses. We used a Yule tree prior, which assumes a constant speciation rate per lineage, and a randomly generated starting tree. The tree prior was shared by all tree models; this consisted of a birth/death incomplete sampling tree prior and was used to model the speciation of nodes in the topology with uniform prior on probability of splits and extinctions. To ensure congruence we carried out the analyses three times for 100 million generations each, and sampling parameters every 10,000 generations. Tracer v.1.6 was used to check the effective sample sizes (ESS), and acceptable values were higher than 200. After removal of a proportion of each run as burn-in the remaining trees were combined in LogCombiner 1.8.0. Maximum clade creditability (MCC) tree was given by summarized data and was estimated in TreeAnnotator 1.8.0, and then visualized using FigTree (Rambaut 2009).
Due to the limited and sporadic fossil records for fungi, it has been difficult to choose a reliable calibration point for the divergence time estimations of any fungal group. Previous molecular study on the porcini used the divergence between Basidiomycota and Ascomycota as a calibration (either 452 Mya according to Taylor and Berbee 2006 or 582 Mya according to Lücking et al. 2009), derived by different researchers using different methods but based on the same 400 million year old fossil, Paleopyrenomycites devonicus. The different estimates were due to the position of this fossil in different subphyla in Ascomycota since it became available and has been used in most fungal evolutionary studies (Heckman et al. 2001; Padovan et al. 2005; Taylor and Berbee 2006; Lücking et al. 2009). Lücking et al. (2009) provided a detailed discussion focusing on the placement of this fossil while recalibrating several earlier studies (Berbee and Taylor 1993; Doolittle et al. 1996; Redecker et al. 2000; Heckman et al. 2001; Padovan et al. 2005) by reassessing the systematic placement of Paleopyrenomycites. Several recent studies have widely used this fossil as the calibration point (Gueidan et al. 2011; Prieto and Wedin 2013; Beimforde et al. 2014; Pérez-Ortega et al. 2016).
In this study, we used one calibration (A1): the divergence between Ascomycota and Basidiomycota, and a normal distribution was applied by setting the mean and the standard deviation to 582.5 and 50.15, respectively. To establish the influence of the calibration points on the results, a secondary calibration (A2) from the order Capnodiales with a constrained age of 100 Mya is used. The mean and the standard deviation were set to 100 and 150 respectively with representation from a fossil Metacapnodiaceae which is hypothesized to have given rise to the common ancestor of the order. All geological time intervals followed Gradstein and Ogg (2004). Mean node age, 95% highest posterior density (HPD) and posterior probability (pp) were mapped on the maximum clade credibility tree.
Results and discussion
Phylogenetic analyses
The combined LSU, SSU, TEF1 and RPB2 gene region data set consist of 391 taxa and 5099 unambiguously aligned sites, 1498 for the LSU, 1617 for the SSU, 856 for TEF1 and 1128 for the RPB2, with Endogone pisiformis (Zygomycota) as the outgroup taxon. Bayesian and ML analyses returned similar topologies with no significant conflicts. The maximum likelihood phylogenies obtained with RAxML (Figs. 1, 2) and BEAST analyses are generally congruent with results reported by other large scale phylogenies of Ascomycota e.g. James et al. (2006) and Schoch et al. (2009b), as well as the most recent broad scale phylogenetic studies of Dothideomycetes (Hyde et al. 2013; Hongsanan et al. 2015; Jaklitsch et al. 2015; Pérez-Ortega et al. 2016). The phylogenetic placement of Orbiliomycetes and Pezizomycetes support the statement that they are the two basal Pezizomycotina classes. Two sister clades were formed and named as the Eurotiomycetes-Lecanoromycetes and Leotiomycetes-Sordariomycetes clades. Phylogenetic relationships among the major groups of Dothideomycetes were recovered with high support and corroborate major classification schemes of previous studies (Schoch et al. 2009a; Zhang et al. 2012; Hyde et al. 2013; Jaklitsch et al. 2015). All accepted Dothideomycetes orders and Pleosporales families were monophyletic with the exception of Asterinales whose taxonomy is discussed later. The 364 strains of Dothideomycetes representing 32 orders, and 114 families segregate into two Dothideomycetes subclasses (Figs. 1, 2) as previously acknowledged based on the presence or absence of pseudoparaphyses (Schoch et al. 2006). The subclass Pleosporomycetidae includes Pleosporales, Mytilinidiales, and Hysteriales and is similar to conclusions from previous publications (Schoch et al. 2009a; Boehm et al. 2009; Shearer et al. 2009; Suetrong et al. 2009; Hyde et al. 2013; Wijayawardene et al. 2014). The subclass Dothideomycetidae comprises Capnodiales, Dothideales and Myriangiales (Schoch et al. 2006, 2009a; Boehm et al. 2009; Zhang et al. 2009; Hyde et al. 2013). Acrospermales, Dyfrolomycetales and Strigulales formed a stable monotypic clade both in the RAxML phylogenetic trees (Figs. 1, 2) and MCC tree (Fig. 3). Natipusillales and Zeloasperisporiales also formed a stable monotypic clade even through their habitats are quite different; Natipusillales is a group of freshwater taxa while Zeloasperisporiales is a group of epiphytic taxa. Taxa of Asterinales are segregated into two different clades, Asterinaceae sensu stricto and Asterinaceae sensu lato, strains of Asterinaceae sensu stricto are sister to the Asterotexiaceae (Asterotexiales) and close to Jahnulales. The species of Asterinaceae sensu lato clustered together with Cladoriellaceae (Cladoriellales), and a new order is probably needed for this group.
The best scoring RAxML Dothideomycetes tree from 391 taxa based on a combined dataset of LSU, SSU, TEF1 and RPB2 sequences. Bootstrap support values for maximum likelihood (ML) >75% are given above the nodes; branches with Bayesian posterior probabilities (PP) above 0.95 are in bold. The original isolate numbers are given after the species names. The tree was rooted with Endogone pisiformis
Maximum clade credibility (MCC) tree with divergence times estimates for Dothideomycetes obtained from a Bayesian approach (BEAST) using single fossil (A1). Numbers at nodes indicate posterior probabilities (pp) for node support; bars correspond to the 95% highest posterior density (HPD) intervals. For estimated median age of nodes, see Table 1. The circles in green indicate the node ages agree with the recommendation made in the paper, and in orange indicate the opposite
An updated treatment of orders of Dothideomycetes and families of Pleosporales which takes into account divergence times is given in Table 1.
Changes in the classification of Dothideomycetes
We present an updated phylogeny of Dothideomycetes including all the accepted orders of which ten were recently established (Pérez-Ortega et al. 2014; Guatimosim et al. 2015; Hongsanan et al. 2015; Jaklitsch et al. 2015; Raja et al. 2015; Pérez-Ortega et al. 2016; Bezerra et al. 2017) since Hyde et al. (2013). The phylogeny presented herein supports 32 orders and 114 families and this includes the 22 orders and 64 families accepted in Hyde et al. (2013).
Divergence times estimates
In the divergence time analysis, although most parameters rapidly reached stationary, the first half of sampled trees (10,000 trees) were considered as part of the burn-in phase and further excluded. The 10,000 remaining trees were summarized in TreeAnnotator. The maximum clade credibility (MCC) tree with divergence estimates obtained through BEAST was topologically similar to those recovered by Bayesian and ML procedures regarding most of the major lineages within Dothideomycetes. The resulting chronograms are shown in Figs. 3, 4 with bars representing 95% confidence intervals for each node. The mean dates for the divergence of Dothideomycetes recovered mostly agreed with reported estimates (Prieto and Wedin 2013; Beimforde et al. 2014, Hyde et al. 2017) and confidence intervals largely overlapped, and the age for the crown of Dothideomycetes is 366 (400–492) Mya.
Maximum clade credibility (MCC) tree with divergence times estimates for Pleosporales obtained from a Bayesian approach (BEAST) using a single fossil (A1). Numbers at nodes indicate posterior probabilities (pp) for node support; bars correspond to the 95% highest posterior density (HPD) intervals. For estimated median age of nodes, see Table 2. The circles in green indicate the node ages agree with the recommendation made in the paper, and in orange indicate the opposite
Generally, two particular informative issues are discussed and indicated in the maximum clade credibility (MCC) tree; one is the stem node age and the other is crown node age. The crown group refers to a collection of species consisting of the living representatives of the collection together with their ancestors back to their most recent common ancestor as well as all of that ancestor’s descendants (WIKIPEDIA: https://en.wikipedia.org/wiki/Crown_group). The stem group refers to those taxa descended from the point where an ancestral taxon split into two sister groups to the point at which a further split gave rise to an extant crown group. Therefore, the stem age of any given groups of taxa is always older than the crown age (Zhao et al. 2016). The species richness of the group, the net diversification rate, the timescale, and model setup can affect the length of branch between stem ancestor and crown clades (McPeek and Brown 2007; Zhao et al. 2016). As a case study, we use the crown age as priority with the stem age as further support, wherever the recommendations are made. Both the crown age and stem age are given and considered in this study.
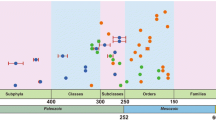
In the maximum clade credibility (MCC) trees in Figs. 3 and 4, the various ages of different orders and families are shown. The age for the crown of most Dothideomycetes orders (20 out of 32, or 63%) are between 100 to 220 Mya, while it is younger for the orders Abrothallales (39 Mya), Asterotexiales (23 Mya), Bezerromycetales (14 Mya), Cladoriellales (6 Mya), Lichenoconiales (9 Mya), Lichenotheliales (46 Mya), Wiesneriomycetales (40 Mya) and Zeloasperisporiales (54 Mya), and the divergence times (stem age) of most orders are between 130 and 310 Mya. The detailed divergence time (stem versus crown age) of the orders are listed in Table 1. The age for the crown of most Pleosporales families (39 out of 55, or 71%) are between 20 and 100 Mya. The detailed divergence times (stem versus crown age) of the families of Pleosporales are listed in Table 2. Divergence times can therefore provide additional evidence for the status of new higher level taxa, or when such taxa e.g. families, orders and classes, are established. Below we recommend how to use divergence times to recognize subclasses, orders and families in the class Dothideomycetes. We suggest that these times should be used as additional evidence for the introduction or acceptance of families, orders and subclasses.
Recommendations for using divergence times to recognize subclasses, orders and families in the class Dothideomycetes
-
1.
Subclasses should have evolved in the range of 235 and 250 Mya (crown age) and 260–322 Mya (stem age).
-
2.
Order should have evolved in the range between 100 and 220 Mya (crown age) and 130 and 310 Mya (stem age).
-
3.
Families should have evolved in the range between 20 and 100 Mya (crown age).
Taxonomic changes
In this section, we make changes to the higher ranking of Dothideomycetes based on our additional evidence from the maximum clade credibility (MCC) trees. We also discuss cases where DNA sequence data can show how the present understanding of some families and orders may be incorrect.
Asterinales M.E. Barr ex D. Hawksw. & O.E. Erikss., Syst. Ascom. 5(1): 177 (1986)
Synonyms
Asterotexiales Firmino, O.L. Pereira & Crous [as ‘Asterotexiales’], Persoonia 35: 238 (2015)
The family Asterotexiaceae was introduced for the monotypic genus Asterotexis with A. cucurbitacearum as the type species (Guatimosim et al. 2015), and the order Asterotexiales was established for this single family. Guatimosim et al. (2015) reported a close phylogenetic relationship between Asterotexiales and Jahnulales. In their analyses, however, they did not include taxa from Asterinaceae sensu stricto, but only Asterinaceae sensu lato (Hyde et al. 2016). There is confusion surrounding strains of Asterinaceae as taxa cluster into two distinct, but well-separated lineages. This discrepancy is thought to result from direct sequencing and thus more than one taxon being sequenced. Hyde et al. (2016) suggested that the group clustering near Asterotexiaceae were Asterinaceae species, while those clustering near Parmulariaceae were likely to be fungi associated with Asterinaceae. In our phylogenetic and BEAST analyses, we included taxa from Asterinaceae sensu stricto and sensu lato. Asterotexiaceae clustered together with strains from Asterinaceae sensu stricto and Jahnulales and is phylogenetically distinct from Asterinaceae sensu lato. We therefore treat Asterotexiales as a synonym of Asterinales. The age for the Asterinales (Asterinaceae sensu stricto and Asterotexiales) crown group is estimated at 133 Mya (stem age 227 Mya) which supports its status as an order.
Botryosphaeriales C.L. Schoch, Crous & Shoemaker., Mycologia 98: 1050 (2006)
Previous studies by Liu et al. (2016) on Botryosphaeriales reported that the order originated around 103 Mya in the Cretaceous period, and Botryosphaeriaceae (which includes Pseudofusicoccumaceae and Endomelanconiopsisaceae) and Phyllostictaceae lineages separated around 87 Mya. These dates coincide closely with the emergence of Botryosphaeriales (114 Mya) and divergence times of these same two families (72 Mya) as estimated in the present study thus supporting the status of Botryosphaeriales as an order and Botryosphaeriaceae and Phyllostictaceae as families. Liu et al. (2016) suggested that Botryosphaeriaceae, Phyllostictaceae and Melanopsaceae clearly warrant separate families, but they considered that separate families for Aplosporellaceae, Planistromellaceae, Saccharataceae and Septorioideaceae are difficult to justify. The crown ages for the Botryosphaeriaceae, Phyllostictaceae, Planistromellaceae and Saccharataceae groups estimated here are 44, 27, 25 and 28 Mya respectively (stem ages 52, 50, 85 and 114 Mya), which suggests that these constitute families. However, the divergence time of Saccharataceae and Septorioidaceae (28 Mya) point towards these being two genera in Saccharataceae. Planistromellaceae and Melanopsaceae diverged at 85 Mya and this confirms the recommendation of Liu et al. (2016) that they are two separate families. The present results indicate that the families Aplosporellaceae, Endomelanconiopsisaceae, Saccharataceae and Septorioideaceae may not represent familial ranks but instead should be considered as genera. However, the status of all families currently included in Botryosphaeriales remains unclear and should be re-evaluated in an in-depth study with greater taxon-sampling.
Floricolaceae Thambugala, Kaz. Tanaka & K.D. Hyde., Fungal Diversity 74: 244 (2015)
Thambugala et al. (2015) provided an account of Lophiostomataceae (family of Pleosporales) and related families and genera. Using a combined analysis of LSU, SSU, ITS and TEF1 sequences data of 109 taxa they introduced a new family Floricolaceae which formed a well-resolved distinct lineage. The 28 taxa used in the tree for this family formed nine distinct groups that were supported by morphology and molecular data. Thambugala et al. (2015) therefore introduced six genera in the family with Floricola as the type. Jaklitsch et al. (2016) however, introduced an epitype for the type species of Teichospora which is the type of Teichosporaceae and synonymized Floricolaceae, and they also synonymized all genera that Thambugala et al. (2015) had accepted under Floricolaceae and thus the family became monotypic. In our phylogeny (Fig. 2), the inclusion of only six strains, and some of the genera (e.g. Asymmetrispora and Ramusculicola) proposed by Thambugala et al. (2015), are well-supported. However, when we weigh in additional evidence from the maximum clade credibility (MCC) tree (Fig. 4), these strains appear to have evolved from 26 to 89 Mya (crown age), which is much older than the average for genera of Ascomycota or Basidiomycota (Hyde et al. 2017). Therefore, the use of these genera in Floricolaceae is justified here.
Tubeufiales Boonmee & K.D. Hyde., Fungal Diversity 68: 245 (2014)
Synonyms
Bezerromycetales J.D.P. Bezerra, C.M. Souza-Motta & Crous, Mycol Prog 16 (4): 301 (2017)
Wiesneriomycetales J.D.P. Bezerra, R.J.V. Oliveira, C.M. Souza-Motta, J.Z. Groenewald & Crous, Mycol Prog 16 (4): 305 (2017)
The orders Bezerromycetales and Wiesneriomycetales were introduced by Bezerra et al. (2017) based on a phylogenetic study of endophytic fungi from the cactus Tacinga inamoena in a Brazilian tropical dry forest. Both orders were established as monotypic with a single family, and one family (Wiesneriomycetaceae) was assigned to Tubeufiales. In our maximum clade credibility (MCC) tree (Fig. 3), the orders Bezerromycetales and Wiesneriomycetales are shown to have diverged recently with the age for their crown groups at 14 and 40 Mya respectively (stem ages 147 and 186 Mya) and it is more likely to be given familial rank which in line with our recommendations between 20 and 100 Mya (crown age). Our recommendation for orders is that they should be more than 100 Mya and this is not the case. Therefore, we synonymize Bezerromycetales and Wiesneriomycetales under Tubeufiales, which is the oldest name. The age for the crown of Tubeufiales is about 189 Mya (stem age 268 Mya), which supports its ordinal status.
Importance of accurately determining the fossil age
The justification of the age and phylogenetic position of the key fossils is an important step in calibrating a node in a divergence dating analysis and can be the core data for the application of divergence times for the classification. Fortunately, there are many studies that have led to credible calibrations and reliable divergence dates (Gueidan et al. 2011; Prieto and Wedin 2013; Beimforde et al. 2014; Pérez-Ortega et al. 2016). This provides us with the possibility to apply divergence times as further evidence to better resolve controversies in fungal classification.
Several studies have shown that using multiple fossil constraints is one possible way to improve the accuracy of molecular dating (Graur and Martin 2004; Hedges and Kumar 2004; Rutschmann et al. 2007; Sauquet et al. 2012). However, the lack of well preserved and identifiable fungal fossils is a limiting factor in dating studies. Another approach is to use secondary calibrations obtained from previous studies (Prieto and Wedin 2013; Beimforde et al. 2014; Pérez-Ortega et al. 2016). In this study, we performed molecular dating analyses using a single relevant fossil, and a secondary calibration point. The estimates obtained based on different setups give similar results with an average ratio of 2.2 Mya (Order, Fig. 3) and 1.3 Mya (Family, Fig. 4) younger ages from analyses A1 and A2 (trees not shown) respectively. Both analyses are based on previous studies with a robust fossil record using five fossil calibration points (Beimforde et al. 2014; Pérez-Ortega et al. 2016). By comparing the inferred divergence dates with the widely accepted dates, we provide insight into using divergence times as additional evidence in higher ranking taxa (order and family) within the class Dothideomycetes and order Pleosporales as selected study groups. Although this could be seen as a circular argument, the use of divergence times can in reality serve as an additional criterion to justify fungal taxonomy conclusions and phylogeny.
Conclusions
We provide four examples where divergence times do not support the status of families and orders. Liu et al. (2016) also discussed the distinction of three genera as distinct families in Botryosphaeriales which was not supported in the MCC trees. These are examples where the distinctiveness of introduced higher taxa using phylogenies should be reinforced by further evidence e.g. divergence times.
Molecular clocks calibrated using fossils are important tools in estimating the timing of evolutionary events in fossil-poor groups. However, when fossil evidence is limited and there are considerable differences in substitution rates change between lineages, it is difficult to establish reliable divergence time estimates. The status of fossils and analysis, an awareness of the phylogeny, fossils and the clock will help to align expectations for fungal evolution. However, it will be many years before we can obtain natural classification for fungi. As more data and approaches become available and fossil and phylogenetic evidence becomes more reliable, we can obtain a better-characterized divergence dating with few limitations.
The major contribution of this paper is an updated phylogeny of the class Dothideomycetes to order level and the order Pleosporales to the family level. The addition of the maximum clade credibility (MCC) tree to support the phylogenetic conclusions show that some orders and some families are not supported and are therefore synonymized. A major conclusion is that any inference from phylogenetic trees depends on the taxa used. For example, Hyde et al. (2016) reported strain MFLUCC 15-1248 as a collection of Neoacanthostigma septoconstrictum as it had strong support in their phylogenetic tree. However, with the addition of ten extra strains in the genus, this strain clustered with a new species, N. brownispora with good support and the strain was renamed. Therefore, any interpretation of phylogenetic trees should be treated with caution as it depends entirely on the taxa chosen to build it. We predict that clade ages will improve the definitions (ranking) of higher taxa and understanding of phylogenetic relationships. Given that the rates of molecular evolution vary with the molecular markers used (ribosomal versus protein coding ones), future studies can show how this can affect calibration as well as estimate evolutionary rates for specific genes and their impact on dating. The results obtained in phylogenetic and molecular clock studies are currently the best hypotheses using present methodologies and data; however, additional data, taxa and/or new methodologies, may result in modified conclusions.
References
Ariyawansa HA, Tanaka K, Thambugala KM, Phookamsak R, Tian Q, Camporesi E, Hongsanan S, Monkai J, Wanasinghe DN, Mapook A (2014) A molecular phylogenetic reappraisal of the Didymosphaeriaceae (=Montagnulaceae). Fungal Divers 68:69–104
Ariyawansa HA, Thambugala KM, Manamgoda DS, Jayawardena R, Camporesi E, Boonmee S, Wanasinghe DN, Phookamsak R, Hongsanan S, Singtripop C, Chukeatirote E (2015) Towards a natural classification and backbone tree for Pleosporaceae. Fungal Divers 71:85–139
Beimforde C, Feldberg K, Nylinder S, Rikkinen J, Tuovila H, Dörfelt H, Gube M, Jackson DJ, Reitner J, Seyfullah LJ (2014) Estimating the Phanerozoic history of the Ascomycota lineages: combining fossil and molecular data. Mol Phylogenet Evol 78:386–398
Berbee ML, Taylor JW (1993) Dating the evolutionary radiations of the true fungi. Can J Bot 71:1114–1127
Bezerra JD, Oliveira RJ, Paiva LM, Silva GA, Groenewald JZ, Crous PW, Souza-Motta CM (2017) Bezerromycetales and Wiesneriomycetales ord. nov. (class Dothideomycetes), with two novel genera to accommodate endophytic fungi from Brazilian cactus. Mycol Prog 16(4):1297–1309
Boehm E, Mugambi G, Miller A, Huhndorf S, Marincowitz S, Spatafora J, Schoch C (2009) A molecular phylogenetic reappraisal of the Hysteriaceae, Mytilinidiaceae and Gloniaceae (Pleosporomycetidae, Dothideomycetes) with keys to world species. Stud Mycol 64:49–83
Boonmee S, Rossman AY, Liu JK, Li WJ, Dai DQ, Bhat JD, Jones EG, McKenzie EH, Xu JC, Hyde KD (2014) Tubeufiales, ord. nov., integrating sexual and asexual generic names. Fungal Divers 68:239–298
Boonmee S, D’souza MJ, Luo Z, Pinruan U, Tanaka K, Su H, Bhat DJ, McKenzie EHC, Jones EBG, Taylor JE, Phillips AJL, Hirayama K, Eungwanichayapant PD, Hyde KD (2016) Dictyosporiaceae fam. nov. Fungal Divers 80:457–482
Crous PW, Slippers B, Wingfield MJ, Rheeder J, Marasas WFO, Philips AJL, Alves A, Burgess TI, Barber PA, Groenewald JZ (2006) Phylogenetic lineages in the Botryosphaeriacea. Stud Mycol 55:235–253
Crous PW, Schoch C, Hyde K, Wood A, Gueidan C, De Hoog G, Groenewald J (2009) Phylogenetic lineages in the Capnodiales. Stud Mycol 64:17–47
Crous PW, Slippers B, Groenewald JZ, Wingfield MJ (2017) Botryosphaeriaceae: systematics, pathology, and genetics. Fungal Biol 121:305–306
Divakar PK, Crespo A, Kraichak E, Leavitt SD, Singh G, Schmitt G, Lumbsch HT (2017) Using a temporal phylogenetic method to harmonize familyand genus-level classification in the largest clade of lichen-forming fungi. Fungal Divers. doi:10.1007/s13225-017-0379-z
Doolittle RF, Feng DF, Tsang S, Cho G, Little E (1996) Determining divergence times of the major kingdoms of living organisms with a protein clock. Science 271:470
Drummond AJ, Ho SYW, Phillips MJ, Rambaut A (2006) Relaxed phylogenetics and dating with confidence. PLoS Biol 4(5):e88
Drummond AJ, Suchard MA, Xie D, Rambaut A (2012) Bayesian phylogenetics with BEAUti and BEAST 1.7. Mol Biol 29:1969–1973
Gradstein F, Ogg J (2004) Geologic time scale 2004—why, how, and where next! Lethaia 37:175–181
Graur D, Martin W (2004) Reading the entrails of chickens: molecular timescales of evolution and the illusion of precision. Trends Genet 20:80–86
Guatimosim E, Firmino AL, Bezerra JL, Pereira OL, Barreto RW, Crous PW (2015) Towards a phylogenetic reappraisal of Parmulariaceae and Asterinaceae (Dothideomycetes). Persoonia 35:230–241
Gueidan C, Ruibal C, de Hoog GS, Schneider H (2011) Rock-inhabiting fungi originated during periods of dry climate in the late Devonian and middle Triassic. Fungal Biol 115:987–996
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucl Acids Symp Ser 41:95–98
Heckman DS, Geiser DM, Eidell BR, Stauffer RL, Kardos NL, Hedges SB (2001) Molecular evidence for the early colonization of land by fungi and plants. Science 293:1129–1133
Hedges SB, Kumar S (2004) Precision of molecular time estimates. Trends Genet 20:242–247
Hongsanan S, Li YM, Liu JK, Hofmann T, Piepenbring M, Bhat JD, Boonmee S, Doilom M, Singtripop C, Tian Q, Mapook A, Zeng XY, Bahkali AH, Xu JC, Mortimer PE, Wu HX, Yang JB, Hyde KD (2014) Revision of genera in Asterinales. Fungal Divers 68:1–68
Hongsanan S, Tian Q, Bahkali AH, Yang JB, McKenzie EH, Chomnunti P, Hyde KD (2015) Zeloasperisporiales ord. nov., and two new species of Zeloasperisporium. Cryptogam Mycol 36:301–317
Hyde KD, Jones EBG, Liu JK, Ariyawansa H, Boehm E, Boonmee S, Braun U, Chomnunti P, Crous PW, Dai DQ, Diederich P, Dissanayake A, Doilom M, Doveri F, Hongsanan S, Jayawardena R, Lawrey JD, Li YM, Liu YX, Lucking R, Monkai J, Muggia L, Nelsen MP, Pang KL, Phookamsak R, Senanayake IC, Shearer CA, Suetrong S, Tanaka K, Thambugala KM, Wijayawardene NN, Wikee S, Wu HX, Zhang Y, Aguirre-Hudson B, Alias SA, Aptroot A, Bahkali AH, Bezerra JL, Bhat DJ, Camporesi E, Chukeatirote E, Gueidan C, Hawksworth DL, Hirayama K, De Hoog S, Kang JC, Knudsen K, Li WJ, Li XH, Liu ZY, Mapook A, McKenzie EHC, Miller AN, Mortimer PE, Phillips AJL, Raja HA, Scheuer C, Schumm F, Taylor JE, Tian Q, Tibpromma S, Wanasinghe DN, Wang Y, Xu JC, Yacharoen S, Yan JY, Zhang M (2013) Families of Dothideomycetes. Fungal Divers 63:1–313
Hyde KD, Hongsanan S, Jeewon R, Bhat DJ, McKenzie EHC, Jones EBG, Phookamsak R, Ariyawansa HA, Boonmee S, Zhao Q, Abdel-Aziz FA, Abdel-Wahab MA, Banmai S, Chomnunti P, Cui BK, Daranagama DA, Das K, Dayarathne MC, de Silva NI, Dissanayake AJ, Doilom M, Ekanayaka AH, Gibertoni TB, Góes-Neto A, Huang SK, Jayasiri SC, Jayawardena RS, Konta S, Lee HB, Li WJ, Lin CG, Liu JK, Lu YZ, Luo ZL, Manawasinghe IS, Manimohan P, Mapook A, Niskanen T, Norphanphoun C, Papizadeh M, Perera RH, Phukhamsakda C, Richter C, de Santiago ALCMA, Drechsler-Santos ER, Senanayake IC, Tanaka K, Tennakoon TMDS, Thambugala KM, Tian Q, Tibpromma S, Thongbai B, Vizzini A, Wanasinghe DN, Wijayawardene NN, Wu HX, Yang J, Zeng XY, Zhang H, Zhang JF, Bulgakov TS, Camporesi E, Bahkali AH, Amoozegar AM, Araujo-Neta LS, Ammirati JF, Baghela A, Bhatt RP, Bojantchev S, Buyck B, da Silva GA, de Lima CLF, de Oliveira RJV, de Souza CAF, Dai YC, Dima B, Duong TT, Ercole E, Mafalda-Freire F, Ghosh A, Hashimoto A, Kamolhan S, Kang JC, Karunarathna SC, Kirk PM, Kytövuori I, Lantieri A, Liimatainen K, Liu ZY, Liu XZ, Lücking R, Medardi G, Mortimer PE, Nguyen TTT, Promputtha I, Raj KNA, Reck MA, Lumyong S, Shahzadeh-Fazeli SA, Stadler M, Soudi MR, Su HY, Takahashi T, Tangthirasunun N, Uniyal P, Wang Y, Wen TC, Xu JC, Zhang ZK, Zhao YC, Zhou JZ, Zhu L (2016) Fungal diversity notes 367–490: taxonomic and phylogenetic contributions to fungal taxa. Fungal Divers 80:1–270
Hyde KD, Maharachchikumbura SSN, Hongsanan S, Samarakoon MC, Lücking R, Pem D, Harishchandra D, Jeewon R, Zhao RL, Xu JC, Liu JK, Al-Sadi AM, Bahkali AH, Elgorban AM (2017) The ranking of fungi—a tribute to David L. Hawksworth on his 70th birthday. Fungal Divers. doi:10.1007/s13225-017-0383-3
Jaklitsch WM, Fournier J, Dai DQ, Hyde KD, Voglmayr H (2015) Valsaria and the Valsariales. Fungal Divers 73:159–202
Jaklitsch WM, Olariaga I, Voglmayr H (2016) Teichospora and the Teichosporaceae. Mycol Prog 15:1–20
James TY, Kauff F, Schoch CL, Matheny PB, Hofstetter V, Cox CJ, Celio G, Gueidan C, Fraker E, Miadlikowska J, Lumbsch HT, Rauhut A, Reeb V, Arnold AE, Amtoft A, Stajich JE, Hosaka K, Sung GH, Johnson D, O’Rourke B, Crockett M, Binder M, Curtis JM, Slot JC, Wang Z, Wilson AW, Schüssler A, Longcore JE, O’Donnell K, Mozley- Standridge S, Porter D, Letcher PM, Powell MJ, Taylor JW, White MM, Griffith GW, Davies DR, Humber RA, Morton JB, Sugiyama J, Rossman AY, Rogers JD, Pfister DH, Hewitt D, Hansen K, Hambleton S, Shoemaker RA, Kohlmeyer J, Volkmann- Kohlmeyer B, Spotts RA, Serdani M, Crous PW, Hughes KW, Matsuura K, Langer E, Langer G, Untereiner WA, Lücking R, Büdel B, Geiser DM, Aptroot A, Diederich P, Schmitt I, Schultz M, Yahr R, Hibbett DS, Lutzoni F, McLaughlin DJ, Spatafora JW, Vilgalys R (2006) Reconstructing the early evolution of Fungi using a six-gene phylogeny. Nature 19:818–822
Jeewon R, Hyde KD (2016) Establishing species boundaries and new taxa among fungi: recommendations to resolve taxonomic ambiguities. Mycosphere 7:1669–1677
Katoh K, Toh H (2008) Recent developments in the MAFFT multiple sequence alignment program. Brief Bioinform 9:286–298
Kirk PM, Ainsworth GC, Cannon PF, Minter DW (2008) Ainsworth & Bisby’s dictionary of the fungi, 10th edn. CAB International, Wallingford
Li GJ, Hyde KD, Zhao RL, Hongsanan S, Abdel-Aziz FA, Abdel-Wahab MA, Alvarado P, Alves-Silva G, Ammirati JF, Ariyawansa HA, Baghela A, Bahkali AH, Beug M, Bhat DJ, Bojantchev D, Boonpratuang T, Bulgakov TS, Camporesi E, Boro MC, Ceska O, Chakraborty D, Chen JJ, Chethana KWT, Chomnunti P, Consiglio G, Cui BK, Dai DQ, Dai YC, Daranagama DA, Das K, Dayarathne MC, Crop ED, De Oliveira RJV, de Souza CAF, de Souza JI, Dentinger BTM, Dissanayake AJ, Doilom M, Drechsler-Santos ER, Ghobad-Nejhad M, Gilmore SP, Goes-Neto A, Gorczak M, Haitjema GH, Hapuar-achchi KK, Hashimoto A, He MQ, Henske JK, Hirayama K, Iribarren MJ, Jayasiri SC, Jayawardena RS, Jeon SJ, Jerônimo GH, Jesus AL, Jones EBG, Kang JC, Karunarathna SC, Kirk PM, Konta S, Kuhnert E, Langer E, Lee HS, Lee HB, Li WJ, Li XH, Liimatainen K, Lima DX, Lin CG, Liu JK, Liu XZ, Liu ZY, Luangsa-ard JJ, Lücking R, Lumbsch HT, Lumyong S, Leaño EM, Marano AV, Matsumura M, McKenzie EHC, Mongkol-samrit S, Mortimer PE, Nguyen TTT, Niskanen T, Norphan-phoun C, O’Malley MA, Parnmen S, Pawłowska J, Perera RH, Phookamsak R, Phukhamsakda C, Pires-Zottarelli CLA, Raspé O, Reck MA, Rocha SCO, de Santiago ALCMA, Senanayake IC, Setti L, Shang QJ, Singh SK, Sir EB, Solomon KV, Song J, Srikitikulchai P, Stadler M, Suetrong S, Takahashi H, Takahashi T, Tanaka K, Tang LP, Thambugala KM, Thanakitpipattana D, Theodorou MK, Thongbai B, Thummarukcharoen T, Tian Q, Tibpromma S, Verbeken A, Vizzini A, Vlasák J, Voigt K, Wanasinghe DN, Wang Y, Weerakoon G, Wen HA, Wen TC, Wijayawardene NN, Wongkanoun S, Wrzosek M, Xiao YP, Xu JC, Yan JY, Yang J, Yang SD, Hu Y, Zhang JF, Zhao J, Zhou LW, Persoh D, Phillips AJL, Maharachchikumbura SSN (2016) Fungal diversity notes 253–366: taxonomic and phylogenetic contributions to fungal taxa. Fungal Divers 78:1–237
Liu JK, Phookamsak R, Jones EBG, Zhang Y, Ko-Ko TW, Hu HL, Boonmee S, Doilom M, Chukeatirote E, Bahkali AH, Wang Y, Hyde KD (2011) Astrosphaeriella is polyphyletic, with species in Fissuroma gen. nov., and Neoastrosphaeriella gen. nov. Fungal Divers 51:135–154
Liu JK, Phookamsak R, Doilom M, Wikee S, Li YM, Ariyawansha H, Boonmee S, Chomnunti P, Dai DQ, Bhat JD, Romero AI, Zhuang WY, Monkai J, Jones EBG, Chukeatirote E, Ko-Ko TW, Zhao YC, Wang Y, Hyde KD (2012) Towards a natural classification of Botryosphaeriales. Fungal Divers 57:149–210
Liu JK, Phookamsak R, Dai DQ, Tanaka K, Jones EBG, Xu JC, Chukeatirote E, Hyde KD (2014) Roussoellaceae, a new pleosporalean family to accommodate the genera Neoroussoella gen. nov., Roussoella and Roussoellopsis. Phytotaxa 181:001–033
Liu JK, Hyde KD, Jones EBG, Ariyawansa HA, Bhat DJ, Boonmee S, Maharachchikumbura SSN, McKenzie EHC, Phookamsak R, Phukhamsakda C, Shenoy BD, Abdel-Wahab MA, Buyck B, Chen J, Chethana KWT, Singtripop C, Dai DQ, Dai YC, Daranagama DA, Dissanayake AJ, Doilom M, D’souza MJ, Fan XL, Goonasekara ID, Hirayama K, Hongsanan S, Jayasiri SC, Jayawardena RS, Karunarathna SC, Li WJ, Mapook A, Norphanphoun C, Pang KL, Perera RH, Peršoh D, Pinruan U, Senanayake IC, Somrithipol S, Suetrong S, Tanaka K, Thambugala KM, Tian Q, Tibpromma S, Udayanga D, Wijayawardene NN, Wanasinghe D, Wisitrassameewong K, Zeng XY, Abdel-Aziz FA, Adamčík S, Bahkali AH, Boonyuen N, Bulgakov T, Callac P, Chomnunti P, Greiner K, Hashimoto A, Hofstetter V, Kang JC, Lewis J, Li XH, Liu XZ, Liu ZY, Matsumura M, Mortimer PE, Rambold G, Randrianjohany E, Sato G, Sri-Indrasutdhi V, Tian CM, Verbeken A, von Brackel W, Wang Y, Wen TC, Xu JC, Yan JY, Zhao RL, Camporesi E (2015) Fungal diversity notes 1–110: taxonomic and phylogenetic contributions to fungal species. Fungal Divers 72:1–197
Liu NG, Ariyawansa HA, Hyde KD, Maharachchikumbura SSN, Zhao RL, Phillips AJL, Jayawardena RS, Thambugala KM, Dissanayake AJ, Wijayawardene NN, Liu JK, Liu ZY, Jeewon R, Jones EBG, Jumpathong J (2016) Perspectives into the value of genera, families and orders in classification. Mycosphere 7:1649–1668. doi:10.5943/mycosphere/7/11/3
Lücking R, Huhndorf S, Pfister DH, Plata ER, Lumbsch HT (2009) Fungi evolved right on track. Mycologia 101:810–822
Lumbsch HT, Huhndorf SM (2010) Myconet volume 14. Part one. Outline of Ascomycota—2009. Fieldiana (Life Earth Sci) 1:1–42
Maharachchikumbura SSN, Hyde KD, Jones EBG, McKenzie EHC, Huang SK, Abdel-Wahab MA, Daranagama DA, Dayarathne M, D’souza MJ, Goonasekara ID, Hongsanan S, Jayawardena RS, Kirk PM, Konta S, Liu JK, Liu ZY, Norphanphoun C, Pang KL, Perera RH, Senanayake IC, Shang QJ, Shenoy BD, Xiao YP, Bahkali AH, Kang JC, Somrothipol S, Suetrong S, Wen TC, Xu JC (2015) Towards a natural classification and backbone tree for Sordariomycetes. Fungal Divers 72:199–301
Maharachchikumbura SSN, Hyde KD, Jones EBG, McKenzie EHC, Bhat DJ, Dayarathne MC, Huang SK, Norphanphoun C, Senanayake IC, Perera RH, Shang QJ, Xiao YP, D’souza MJ, Hongsanan S, Jayawardena RS, Daranagama DA, Konta S, Goonasekara ID, Zhuang WY, Jeewon R, Phillips AJL, Abdel-Wahab MA, Al-Sadi AM, Bahkali AH, Boonmee S, Boonyuen N, Cheewangkoon R, Dissanayake AJ, Kang JC, Li QR, Liu JK, Liu XZ, Liu ZY, Luangsa-ard JJ, Pang KL, Phookamsak R, Promputtha I, Suetrong S, Stadler M, Wen TC, Wijayawardene NN (2016) Families of Sordariomycetes. Fungal Divers 79:1–317
Mapook A, Hyde KD, Hongsanan S, Phukhamsakda C, Li JF, Boonmee S (2016) Palawaniaceae fam. nov., a new family (Dothideomycetes, Ascomycota) to accommodate Palawania species and their evolutionary time estimates. Mycosphere 7:1732–1745
McPeek MA, Brown JM (2007) Clade age and not diversification rate explains species richness among animal taxa. Am Nat 169:E000
Mugambi GK, Huhndorf SM (2009) Molecular phylogenetics of Pleosporales: Melanommataceae and Lophiostomataceae re-circumscribed (Pleosporomycetidae, Dothideomycetes, Ascomycota). Stud Mycol 64:103–121
Nelsen M, Lücking R, Grube M, Mbatchou J, Muggia L, Plata ER, Lumbsch H (2009) Unravelling the phylogenetic relationships of lichenised fungi in Dothideomyceta. Stud Mycol 64:135–144
Padovan ACB, Sanson GF, Brunstein A, Briones MR (2005) Fungi evolution revisited: application of the penalized likelihood method to a Bayesian fungal phylogeny provides a new perspective on phylogenetic relationships and divergence dates of Ascomycota groups. J Mol Evol 60:726–735
Pérez-Ortega S, Suija A, Crespo A, de los Ríos A (2014) Lichenicolous fungi of the genus Abrothallus (Dothideomycetes: Abrothallales ordo nov.) are sister to the predominantly aquatic Janhulales. Fungal Divers 64:295–304
Pérez-Ortega S, Garrido-Benavent I, Grube M, Olmo R, de los Ríos A (2016) Hidden diversity of marine borderline lichens and a new order of fungi: Collemopsidiales (Dothideomyceta). Fungal Divers 80:285–300
Phookamsak R, Liu JK, McKenzie EH, Manamgoda DS, Ariyawansa H, Thambugala KM, Dai DQ, Camporesi E, Chukeatirote E, Wijayawardene NN, Bahkali AH, Mortimer PE, Xu JC, Hyde KD (2014) Revision of Phaeosphaeriaceae. Fungal Divers 68:159–238
Phookamsak R, Norphanphoun C, Tanaka K, Dai DQ, Luo ZL, Liu JK, Su HY, Bhat DJ, Bahkali AH, Mortimer PE, Xu JC, Hyde KD (2015) Towards a natural classification of Astrosphaeriella-like species; introducing Astrosphaeriellaceae and Pseudoastrosphaeriellaceae fam. nov and Astrosphaeriellopsis, gen. nov. Fungal Divers 74:143–197
Phukhamsakda C, Hongsanan S, Ryberg M, Ariyawansa H, Chomnunti P, Bahkali A, Hyde KD (2016) The evolution of Massarineae with Longipedicellataceae fam. Nov. Mycosphere 7:1713–1731
Prieto M, Wedin M (2013) Dating the diversification of the major lineages of Ascomycota (Fungi). PLoS ONE 8:e65576
Raja HA, El-Elimat T, Oberlies NH, Shearer CA, Miller AN, Tanaka K, Hashimoto A, Fournier J (2015) Minutisphaerales (Dothideomycetes, Ascomycota): a new order of freshwater ascomycetes including a new family, Minutisphaeraceae, and two new species from North Carolina, USA. Mycologia 107:845–862
Rambaut A (2009) FigTree 1.2.2. http://tree.bio.ed.ac.uk/software/figtree/
Rambaut A, Suchard M, Drummond AJ (2013) Tracer 1.6 http://tree.bio.ed.ac.uk/software/tracer/
Redecker D, Kodner R, Graham LE (2000) Glomalean fungi from the Ordovician. Science 289:1920–1921
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572
Rutschmann F, Eriksson T, Salim KA, Conti E (2007) Assessing calibration uncertainty in molecular dating: the assignment of fossils to alternative calibration points. Syst Biol 56:591–608
Samarakoon M, Hyde KD, Promputtha I, Hongsanan S, Ariyawansa HA, Maharachchikumbura SSN, Daranagama A, Stadler M, Mapook A (2016) Evolution of Xylariomycetidae (Ascomycota: Sordariomycetes). Mycosphere 7:1746–1791
Sauquet H, Ho SYW, Gandolfo MA, Jordan GJ, Wilf P, Cantrill DJ, Bayly MJ, Bromham L, Brown GK, Carpenter RJ, Lee DM, Murphy DJ, Sniderman JMK, Udovicic F (2012) Testing the impact of calibration on molecular divergence times using a fossil-rich group: the case of Nothofagus (Fagales). Syst Biol 61:289–313
Schoch CL, Shoemaker RA, Seifert KA, Hambleton S, Spatafora JW, Crous PW (2006) A multigene phylogeny of the Dothideomycetes using four nuclear loci. Mycologia 98:1041–1052
Schoch CL, Crous PW, Groenewald JZ, Boehm EWA, Burgess TI, de Gruyter J, de Hoog GS, Dixon LJ, Grube M, Gueidan C, Harada Y, Hatakeyama S, Hirayama K, Hosoya T, Huhndorf SM, Hyde KD, Jones EBG, Kohlmeyer J, Kruys Å, Li YM, Lücking R, Lumbsch HT, Marvanová L, Mbatchou JS, McVay AH, Miller AN, Mugambi GK, Muggia L, Nelsen MP, Nelson P, Owensby CA, Phillips AJL, Phongpaichit S, Pointing SB, Pujade-Renaud V, Raja HA, Rivas Plata E, Robbertse B, Ruibal C, Sakayaroj J, Sano T, Selbmann L, Shearer CA, Shirouzu T, Slippers B, Suetrong S, Tanaka K, Volkmann-Kohlmeyer B, Wingfield MJ, Wood AR, Woudenberg JHC, Yonezawa H, Zhang Y, Spatafora JW (2009a) A class-wide phylogenetic assessment of Dothideomycetes. Stud Mycol 64:1–15
Schoch CL, Sung GH, López-Giráldez F, Townsend JP, Miadlikowska J, Hofstetter V, Robbertse B, Mathen PB, Kauff F, Wang Z, Gueidan CC, Andrie RM, Trippe K, Ciufetti LM, Wynns A, Fraker E, Hodkinson BP, Bonito G, Groenewald JZ, Arzanlou M, De-Hoog GS, Crous PW, Hewitt D, Pfister DH, Peterson K, Gryzenhout M, Wingfield MJ, Aptroot A, Suh SO, Blackwell M, Hillis DM, Griffith GW, Castlebury LA, Rossman AY, Lumbsch HT, Lücking R, Büdel B, Rauhut A, Diederich P, Ertz D, Geiser DM, Hosaka K, Inderbitzin P, Kohlmeyer J, Volkmann-Kohlmeyer B, Mostert L, O’Donnell K, Sipman H, Rogers J, Shoemaker RA, Sugiyama J, Summerbell RC, Untereiner W, Johnston PR, Stenroos S, Zuccaro A, Dyer PS, Crittenden PD, Cole MS, Hansen K, Trappe JM, Yahr R, Lutzoni FO, Spatafora JW (2009b) The Ascomycota tree of life: a phylum-wide phylogeny clarifies the origin and evolution of fundamental reproductive and ecological traits. Syst Biol 58:224–239
Shearer CA, Raja HA, Miller AN, Nelson P, Tanaka K, Hirayama K, Marvanová L, Hyde KD, Zhang Y (2009) The molecular phylogeny of freshwater Dothideomycetes. Stud Mycol 64:145–153
Stamatakis A, Hoover P, Rougemont J (2008) A rapid bootstrap algorithm for the RAxML web servers. Syst Biol 57:758–771. doi:10.1080/10635150802429642
Suetrong S, Schoch CL, Spatafora JW, Kohlmeyer J, Volkmann-Kohlmeyer B, Sakayaroj J, Phongpaichit S, Tanaka K, Hirayama K, Jones EBG (2009) Molecular systematics of the marine Dothideomycetes. Stud Mycol 64:155–173
Tanaka K, Hirayama K, Yonezawa H, Hatakeyama S, Harada Y, Sano T, Shirouzu T, Hosoya T (2009) Molecular taxonomy of bambusicolous fungi: Tetraplosphaeriaceae, a new pleosporalean family with Tetraploa-like anamorphs. Stud Mycol 64:175–209
Tanaka K, Hirayama K, Yonezawa H, Sato G, Toriyabe A, Kudo H, Hashimoto A, Matsumura M, Harada Y, Kurihara Y (2015) Revision of the Massarineae (Pleosporales, Dothideomycetes). Stud Mycol 82:75–136
Taylor JW, Berbee ML (2006) Dating divergences in the Fungal Tree of Life: review and new analyses. Mycologia 98:838–849
Thambugala KM, Hyde KD, Tanaka K, Tian Q, Wanasinghe DN, Ariyawansa HA, Jayasiri SC, Boonmee S, Camporesi E, Hashimoto A, Hirayama K, Schumacher RK, Promputtha I, Liu ZY (2015) Towards a natural classification and backbone tree for Lophiostomataceae, Floricolaceae, and Amorosiaceae fam. nov. Fungal Divers 74:199–266
Tian Q, Liu JK, Hyde KD, Wanasinghe DN, Boonmee S, Jayasiri SC, Luo ZL, Taylor JE, Phillips AJ, Bhat DJ, Li WJ, Ariyawansa H, Thambugala KM, Jones EBG, Chomnunti P, Bahkali AH, Xu JC, Camporesi E (2015) Phylogenetic relationships and morphological reappraisal of Melanommataceae (Pleosporales). Fungal Divers 74:267–324
Wijayawardene NN, Crous PW, Kirk PM, Hawksworth DL, Boonmee S, Braun U, Dai DQ, Dsouza MJ, Diederich P, Dissanayake A, Doilom M, Hongsanan S, Jones EBG, Groenewald JZ, Jayawar-dena R, Lawrey JD, Liu JK, Lücking R, Madrid H, Manamgoda DS, Muggia L, Nelsen MP, Phookamsak R, Suetrong S, Tanaka K, Thambugala KM, Wanasinghe DN, Wikee S, Zhang Y, Aptroot A, Ariyawansa HA, Bahkali AH, Bhat DJ, Gueidan C, Chomnunti P, De Hoog GS, Knudsen K, Li WJ, McKenzie EHC, Miller AN, Phillips AJL, Piątek M, Raja HA, Shivas RS, Slippers B, Taylor JE, Tian Q, Wang Y, Woudenberg JHC, Cai L, Jaklitsch WM, Hyde KD (2014) Naming and outline of Dothideomycetes–2014 including proposals for the protection or suppression of generic names. Fungal Divers 69:1–55
Zhang Y, Schoch CL, Fournier J, Crous PW, De Gruyter J, Woudenberg JHC, Hirayama K, Tanaka K, Pointing SB, Spatafora JW, Hyde KD (2009) Multi-locus phylogeny of Pleosporales: a taxonomic, ecological and evolutionary re-evaluation. Stud Mycol 64:85–102
Zhang Y, Crous PW, Schoch CL, Hyde KD (2012) Pleosporales. Fungal Divers 53:1–221
Zhao RL, Zhou JL, Chen J, Margaritescu S, Sánchez-Ramirez S, Hyde KD, Callac P, Parra LA, Li GJ, Moncalvo JM (2016) Towards standardizing taxonomic ranks using divergence times-a case study for reconstruction of the Agaricus taxonomic system. Fungal Divers 78:239–292. doi:10.1007/s13225-016-0357-x
Zhao RL, Li GJ, Sánchez-Ramírez S, Stata M, Moncalvo J-M, Yang ZL, Wu G, Dai YC, He SH, Cui BK, Zhou JL, Wu F, He MQ, Hyde KD (2017) A six-genes phylogenetic overview of Basidiomycota and allied phyla with estimated divergence times of higher taxa and a phyloproteomics perspective. Fungal Divers. doi:10.1007/s13225-017-0381-5
Zhaxybayeva O, Gogarten JP (2002) Bootstrap, Bayesian probability and maximum likelihood mapping: exploring new tools for comparative genome analyses. BMC Genomics 3:4
Acknowledgements
This work was funded by grants of the National Natural Science Foundation of China (NSFC 31210103919; 31600032; 31360015), Science and Technology Foundation of Guizhou Province (LH [2015]7061) and the Research of Featured Microbial Resources and Diversity Investigation in Southwest Karst area (Project No. 2014FY120100.). Jian-Kui Liu thanks Dr. Bang Feng (Kunming Institute of Botany, Chinese Academy of Sciences, Kunming, China) for his valuable help with phylogenetic analysis. Dr. Rui-Lin Zhao and Dr. H.A. Ariyawansa are thanked for their valuable suggestions. Dr. Hong Luo is thanked for commenting the manuscript. K.D. Hyde thanks the Chinese Academy of Sciences, Project Number 2013T2S0030, for the award of Visiting Professorship for Senior International Scientists at Kunming Institute of Botany. K.D. Hyde also extends his appreciation to the Thailand Research Fund (TRF) Grant (RSA5980068) and National Research Council of Thailand (NRCT) for Grants (60201000201; 592010200112). Alan JL Phillips acknowledges the support from Biosystems and Integrative Sciences Institute (BioISI, FCT/UID/Multi/04046/2013).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Liu, JK., Hyde, K.D., Jeewon, R. et al. Ranking higher taxa using divergence times: a case study in Dothideomycetes. Fungal Diversity 84, 75–99 (2017). https://doi.org/10.1007/s13225-017-0385-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13225-017-0385-1