Abstract
The previous phylogenies of Sordariomycetes by M.E. Barr, O.E. Eriksson and D.L. Hawksworth, and T. Lumbsch and S. Huhndorf, were mainly based on morphology and thus were somewhat subjective. Later outlines by T. Lumbsch and S. Huhndorf, and Maharachchikumbura and co-authors, took into account phylogenetic evidence. However, even these phylogenetic driven arrangements for Sordariomycetes, were somewhat subjective, as the arrangements in trees depended on many variables, such as number of taxa, different gene regions and methods used in the analyses. What is needed is extra evidence to help standardize ranking in the fungi. Estimation of divergence times using molecular clock methods has been proposed for providing additional rational for higher ranking of taxa. Thus, in Sordariomycetes, a divergence period (i.e. 200–300 MYA) can be used as criteria to judge when a group of related taxa evolved and what rank they should be given. In this paper, we provide an updated classification of accepted subclasses, orders of Sordariomycetes and use divergence times to provide additional evidence to stabilize ranking of taxa in the class. We point out and discuss discrepancies where the phylogenetic tree conflicts with the molecular clock.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sordariomycetes is an important class of ascomycetes, mainly characterized by non-lichenized, flask-shaped fruiting bodies (perithecia) and unitunicate asci (Lumbsch 2000; Zhang et al. 2006; Maharachchikumbura et al. 2015, 2016). However, this simple definition could change upon the growth form and habitat. Most members of Xylariomycetidae and some of Sordariomycetidae have dark perithecia, amyloid asci, true paraphyses and periphysate ostioles, while most taxa of Hypocreomycetidae have light coloured perithecia, nonamyloid ascal apical rings (when apical rings are present) and lack true paraphyses. Some groups of Sordariomycetes have cleistothecia (Zhang et al. 2006; Tang et al. 2007; Senanayake et al. 2015). The class Sordariomycetes has a cosmopolitan distribution and accommodates mostly terrestrial taxa, although several taxa can be found in aquatic habitats (Hyde and Jones 1989; Tsui et al. 2000; Ho et al. 2001; Cai et al. 2002; Jones et al. 2009a, b, 2015). They are also pathogens of plants, arthropods and mammals (Sung et al. 2007; Maharachchikumbura et al. 2012, 2015; Hyde et al. 2014, 2016) and have been isolated as endophytes from various plants (Guo et al. 2001; Promputtha et al. 2005), while many are saprobes involved in decomposition and nutrient cycling (Prados-Rosale et al. 2012; Keim et al. 2014; Liu et al. 2015; Ariyawansa et al. 2015; Li et al. 2016).
The classification of Sordariomycetes has changed drastically over the past decades because of the plasticity and variability in phenotype characters (Alexopoulos et al. 1996; Barr 1983, 1987, 1990; Eriksson and Hawksworth 1993). In addition, morphology alone is unable to provide evidence for the natural origins and patterns of evolution among these fungi (Mitchell et al. 1995; Vijaykrishna et al. 2006; Tang et al. 2007). Molecular studies on Sordariomycetes began in the early 1990s and SSU and LSU sequence data was mostly used as markers (Berbee and Taylor 1992; Spatafora and Blackwell 1993; Spatafora 1995). However, SSU and LSU alone, are not sufficient to resolve the most of the groups in Sordariomycetes as they have a low resolution in ranking taxa (Tang et al. 2007).
The higher ranking of Sordariomycetes is not yet stable. Eriksson and Winka (1997) introduced three subclasses, Hypocreomycetidae, Sordariomycetidae and Xylariomycetidae based on morphology (perithecial ascomata, hamathecium composed of paraphyses, ostioles with periphyses and unitunicate or pseudoprotunicate asci) and nrDNA sequence data. However, it has been found that assemblages of protein genes yield a higher phylogenetic resolution, as compared to ribosomal regions (Schoch et al. 2009). In most recent studies, in addition to ribosomal genes, the phylogenetic relationships among Sordariomycetes were investigated using partial translation elongation factor 1-alpha, the second largest subunit of RNA polymerase (RPB2) and beta-tubulin genes (Zhang et al. 2006; Tang et al. 2007). In a revision of Sordariomycetes, Maharachchikumbura et al. (2015) introduced three new subclasses; Diaporthomycetidae, Lulworthiomycetidae, and Meliolomycetidae based on morphology and combined analysis of LSU, SSU, TEF and RPB2 sequence data. According to the outline by Maharachchikumbura et al. (2016), Sordariomycetes currently has six subclasses, 32 orders, 105 families and 1331 genera.
Fossil calibration data and divergence time estimates are being used as additional evidence for the ranking of fungi (Beimforde et al. 2014; Hongsanan et al. 2016; Pérez-Ortega et al. 2016; Samarakoon et al. 2016; Zhao et al. 2016) and studies have shown that Sordariomycetes had a higher speciation process over time, when compared to Dothideomycetes and Leotiomycetes (Wang et al. 2010). In this paper, we provide an updated backbone tree for Sordariomycetes based on the analysis of LSU, SSU, TEF and RPB2 sequence data. Based on the new phylogenies and additional evidence from divergence times published in Hyde et al. (2017), several taxonomic changes to Sordariomycetes are necessary. We therefore provide a list of proposed changes, including new introductions and discuss discrepancies between divergent times and phylogenetic results, when they are inconsistent.
Materials and methods
Phylogenetic analysis
Representative LSU, SSU, TEF1 and RPB2 sequence data from each family in Sordariomycetes and some strains of Leotiomycetes (346 taxa) were downloaded from GenBank to supplement the dataset (Supplementary Table 1). Representative strains from Eurotiomycetes were used as outgroup taxa. The data set was aligned by using MAFFT (Katoh et al. 2009), checked and aligned manually using Bioedit (Hall 1999). Maximum likelihood analysis using RAxML was performed by using raxmlGUIv.0.9b2 (Silvestro & Michalak 2012). The GTRGAMMA model was used in the analysis, the search strategy was set to 1000 rapid bootstrapping. The number of replicates was inferred using the stopping criterion (Pattengale et al. 2009). The trees from analysis were viewed in FigTree (Rambaut 2006). The bootstrap values equal or greater than 50% are given as the first set of numbers above the nodes (Fig. 1).
Molecular clock analysis
Data for the molecular clock analysis of Sordariomycetes is provided in Hyde et al. (2017). We use the data from the molecular clock evidence in this paper and compare it with the phylogenetic tree presented here. The conflicts found between the MCC tree in Hyde et al. (2017) and the phylogenetic tree in this study are discussed below.
Results and discussion
Representative strains of LSU, SSU, TEF1 and RPB2 sequence data of Sordariomycetes and Leotiomycetes were included in the phylogenetic analysis; representative strains of Eurotiomycetes were selected as an outgroup (Figs. 1, 2). Phylogenetic analysis generated by RAxML analysis indicates Sordariomycetes share a common ancestor with Leotiomycetes with high support (94% ML). In the tree, Diaporthomycetidae, Hypocreomycetidae, Lulworthiomycetidae, Sordariomycetidae, Savoryellomycetidae and Xylariomycetidae are well-supported. Meliolomycetidae belongs to Sordariomycetidae, and is no longer treated as subclass. Savoryellomycetidae is introduced formally in this study based on phylogenetic analysis (100% ML) and its stem age (267 MYA) reported in Hyde et al. (2017). The internal classification of each subclass is discussed in this paper based on phylogenetic (this study) and the MCC trees (Hyde et al. 2017). Moreover, some conflicts between the phylogenetic tree in this study and previous studies are also discussed.
Sordariomycetes backbone tree
In this section, we provide a revised phylogenetic arrangement for Sordariomycetes following the previous backbone trees in Maharachchikumbura et al. (2015, 2016). The new backbone tree incorporates six subclasses, 28 orders, and 105 families.
In this paper, we provide an updated phylogenetic assessment of Sordariomycetes based on sequence data from 345 strains. Hyde et al. (2017) provided a molecular clock for Sordariomycetes, thus we use evidence from the phylogenetic analyses and the molecular clock in Hyde et al. (2017) to rationalise the arrangement of Sordariomycetes to family level. The crown node versus stem node age must be considered when reviewing evidence from the molecular clock. The crown node age is affected by the species number used in the analysis and number of base pair differences between species (Zhao et al. 2016; Hyde et al. 2017). Thus, Hyde et al. (2017) used the stem age to evaluate divergence times as used by Zhao et al. (2016, 2017) and Garnica et al. (2016).
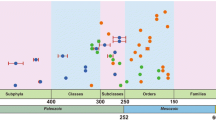
In this study, the divergence for Sordariomycetes is estimated at 341 MYA and this concurs with the recommendation times given for classes at 300 MYA (Hyde et al. 2017). Maharachchikumbura et al. (2016) accepted six subclasses (including Meliolomycetidae), whereas in our phylogeny we resolved six subclasses, but Meliolomycetidae (131 MYA) is reduced to an order of Sordariomycetidae (Fig. 2). In this study, bootstrap and posterior probability values from both analyses, including the stem ages support the status of subclass Diaporthomycetidae (youngest subclass), Hypocreomycetidae, Lulworthiomycetidae, Sordariomycetidae, Savoryellomycetidae and Xylariomycetidae at 250–289 MYA. Although the stem age of Glomerellales is 256 MYA, we retain Glomerellales within the subclass Hypocreomycetidae, due to few extant taxa being used in the tree and its unstable placement. The divergence times for the six subclasses falls within recommended range of ca. 250–300 MYA (Hyde et al. 2017) and provides evidence for the status of these subclasses.
Table 1 provides an updated classification. In the phylogenetic tree, sequence data clusters into 28 well-resolved groups that can be used to represent orders (130–250 MYA). These orders generally correspond to the orders accepted in Maharachchikumbura et al. (2015, 2016), with changes as listed in Table 1. Below we provide notes on any recently introduced taxa and any changes recommended as a result of this paper. There are however, several places where there are discrepancies between the phylogenetic tree and molecular clock results and these are discussed.
Subclass Diaporthomycetidae Senan. et al. 2015
In Diaporthomycetidae, Maharachchikumbura et al. (2016) recognised nine orders, while Crous et al. (2015) introduced Myrmecridiales and Senanayake et al. (2016) introduced Phomatosporales. In the present study, we recognize nine orders (Table 1). The status of Amplistromatales, Calosphaeriales, Diaporthales, Magnaporthales, Ophiostomatales, Phomatosporales and Togniniales are supported with divergence times of 137–188 MYA. Annulatascales and Myrmecridiales fall in the family status, and they share the most common ancestor at 122 MYA. The families in Diaporthomycetidae are mostly well-supported falling in the recommended divergence times of 50–150 MYA.
Magnaporthales Thongk et al. 2009
Pseudohalonectriaceae Hongsanan & K.D. Hyde, fam. nov.
Index Fungorum number: IF553215; Facesoffungi number: FoF 03355
Diagnosis Pseudohalonectriaceae is introduced as a new family within Magnaporthales. The stem age of Pseudohalonectriaceae is 95 MYA. Members in Pseudohalonectria are found in marine and terrestrial or freshwater habitats (Perera et al. 2016).
Sexual morph: Ascomata erumpent to immersed with a protruding neck, cylindrical, periphysate necks, greenish yellow, bright yellow to brown. Neck conical, composed of parallel hyphae, outer hyphae outwardly directed, subglobose with enlarged ends, greenish yellow, periphysate. Peridium multi-layers. Paraphyses tapering towards the apex, thin-walled, attached to ascogenous hyphae. Asci 8-spored, unitunicate, cylindrical to clavate, with a nonamyloid, thimble-shaped, refractive apical apparatus. Ascospores overlapping uniseriate to biseriate, cylindrical or ellipsoidal, straight to curved, usually multi-septate, constricted or not-constricted at the septa, hyaline to slightly coloured and pale brown, pink/orange in mass in some species, smooth-walled (Shearer and Zare-Maivan 1988; Hyde et al. 1999; Perera et al. 2016). Asexual morph: Undetermined.
Type genus: Pseudohalonectria Minoura & T. Muroi 1978
Notes: Pseudohalonectria was introduced without being assigned to an order or family (Minoura and Muroi 1978). The genus was reviewed and added new species by several researchers (see Perera et al. 2016). Shearer (1989) linked hyphomycetous, phialidic anamorph to P. phialidica, however, this species was transferred to Ceratosphaeria (Huhndorf et al. 2008). Thus, the asexual morph of Pseudohalonectria is still questionable. The MCC tree in Hyde et al. (2017), the stem age of Pseudohalonectria falls in the family status (95 MYA), with high support in the phylogenetic (this study) and the MCC trees.
Annulatascales D'souza et al. 2015 and Myrmecridiales Crous 2015
Annulatascales and Myrmecridiales are the youngest orders and have the same common ancestor at 122 MYA, and the molecular clock indicates that they might represent a single order. However, the bootstrap and posterior probability values from phylogenetic and molecular clock analyses do not support their relationships. Therefore, these orders are not synonymised until the placement of Myrmecridiales in phylogenetic tree is stable. Myrmecridiales is a monotypic order in the subclass Diaporthomycetidae and accommodates the family Myrmecridiaceae (Crous et al. 2015). Members of this family are ramichloridium-like taxa with hyaline mycelium, and pale to unpigmented, pimple-like denticles (Arzanlou et al. 2007). Phylogenetically it has affinities to the order Ophiostomatales (Crous et al. 2015). The genus Myrmecridium are saprobes, isolated from leaf litter, soil, plant stems, leaves and roots (Arzanlou et al. 2007; Réblová et al. 2016). Although, Thyridiaceae shares a common ancestor with Annulatascales and Myrmecridiales with a stem age 137 MYA, it comprises a single strain. Thus, we do not raise Thyridiaceae to ordinal status.
Calosphaeriales M.E. Barr 1983
= Jobellisiales M.J. D’souza & K.D. Hyde 2015
Calosphaeriaceae Munk 1957
Pleurostomataceae Réblová, L. Mostert, W. Gams & Crous 2004
Jobellisiaceae Réblová 2008
In the MCC tree, Pleurostomataceae forms a distinct lineage that diverged from the same common ancestor with Calosphaeriaceae at 129 MYA, which falls in the family status. We suggest that Pleurostomataceae should be retained as a family in Calosphaeriales. Based on the divergence time Jobellisiales falls in the ordinal status (146 MYA), however, the placement of this order is unstable as sometimes it clusters with Pleurostomataceae. Thus, we maintain it under order Calosphaeriales to retain the monophyly of the group.
Diaporthales Nannf. 1932 and Tirisporellales Suetrong et al. 2015
Tirisporellales is a monotypic order in the class Sordariomycetes, subclass Diaporthomycetidae and includes a single family Tirisporellaceae. The family was introduced by Suetrong et al. (2015) based on SSU and LSU rDNA sequence analysis and morphological observations and comprises two monotypic genera, Tirisporella and Thailandiomyces. In the MCC analysis, the order Tirisporellales has a divergence time at 112 MYA and shares a common ancestor with Sydowiellaceae and also Pseudovalsariaceae, while Tirisporellales is closely related to Pseudovalsaceae in the phylogenetic tree. The placement of Tirisporellales is not-well supported in both MCC and phylogenetic trees, but falls within Diaporthales. However, Tirisporellaceae is a marine fungal family and it has a hyphomyceteous (asexual morph) which is not found in Diaporthales. Therefore, Tirisporellales is not synonymised under Diaporthales until more data are available.
Gnomoniaceae G. Winter 1886 and Melanconidaceae G. Winter 1886
Melanconidaceae (crown 15.6 MYA) is a family in Diaporthales with a relatively young divergent time (stem age at 52 MYA). In most of the phylogenetic analyses, it forms a sister clade to the well-supported Gnomoniaceae. Taxa belonging to these two families share somewhat similar morphologies (Senanayake et al. pers. comm.). Maharachchikumbura et al. (2016) listed 24 genera under this family based on morphology following Lumbsch and Huhndorf (2010). However, Senanayake et al. (pers. comm.) restrict the family Melanconidaceae for Melanconis. Based on the stem age (52 MYA) which is near to the minimum bounds of family status (50–130 MYA), and this can be changed in other analyses (height 95% HPD = 37–68 MYA) based on many reasons discussed in Hyde et al. (2017), here we synonymize Melanconidaceae under Gnomoniaceae.
Harknessiaceae Crous 2012 Cryphonectriaceae Gryzenh. & M.J. Wingf. 2006 and Schizoparmaceae Rossman et al. 2007
The clade comprising Harknessiaceae, Cryphonectriaceae and Schizoparmaceae has a stem age at 70 MYA, indicating the status of a single family. We however do not follow this here, because the data set used is not sufficient and these families are morphologically diverse.
Lamproconiaceae C. Norphanphoun et al. 2016
Norphanphoun et al. (2016) assigned the genera Lamproconium and Hercospora to a new family Lamproconiaceae in Diaporthales. In the combined ITS and LSU phylogeny, Lamproconiaceae forms a clade basal to Sydowiellaceae and Stilbosporaceae (Norphanphoun et al. 2016). This family is typified by Lamproconium (Lamproconium desmazieri). Lamproconiaceae has molecular clock support as a family in Diaporthales (89 MYA).
Juglanconidaceae Voglmayr & Jaklitsch 2017
= Melansporellaceae C.M. Tian & Z. Du 2017
Based on the sequence analysis of ITS and LSU data, Voglmayr et al. (2017) observed that Melanconis species occurring on Juglandaceae are different from Melanconis sensu stricto, and hence the novel genus Juglanconis was introduced. The genus Juglanconis has no affinities to any of the known families in Diaporthales and therefore, a new family Juglanconidaceae was introduced to accommodate it. This family was introduced late in the writing of this paper and we could not include it in the analysis. Thus, it will be interesting to see if future MCC trees will support its status. Du et al. (2017) introduced Melansporellaceae which is the same as Juglanconidaceae and is herein synonymised.
Distoseptisporaceae K.D. Hyde & McKenzie 2016
Distoseptisporaceae diverged from its sister clade Phomatosporales at 122 MYA in the MCC tree with moderate support, while it clusters with Pseudohalonectria (genera incertae sedis in Diaporthomycetidae) and Myrmecridiaceae in phylogenetic tree, but with low bootstrap support. The split between Distoseptisporaceae and its sister clade could not be considered because different split/divergence events can produce different stem ages, thus we do not raise this family to ordinal level until the placement in the phylogenetic and MCC trees is stable. We presently retained this family in Diaporthomycetidae, family incertae sedis.
Phomatosporales Senanayake et al. 2016
Phomatosporales was introduced by Senanayake et al. (2016) to accommodate the new family Phomatosporaceae with the genera Phomatospora, Lanspora, and the new genus Tenuimurus. LSU, SSU and ITS sequence data showed that Phomatosporaceae formed a distinct clade in Sordariomycetes, which is sister to the order Amplistromatales. The asexual morph of the Phomatosporaceae are Sporothrix-like (Fournier and Lechat 2010). The divergence of the order (stem age, 138 MYA) supports its ordinal status.
Papulosaceae Winka & O.E. Erikss. 2000 Sporidesmiaceae Fr. 1849 and Trichosphaeriaceae G. Winter 1885
These three families were indicated to need an upgrade to ordinal status, under the name Sporidesmiales with a stem age at 186 MYA (Hyde et al. 2017). The species of Sporidesmiaceae show a sister relationship to Papulosaceae and Trichosphaeriaceae in this study, but with low support. In some studies, these families clustered with other families in Diaporthomycetidae. Their status may need revision following further study.
Trichosphaeriales M.E. Barr 1983
Continued use of Trichosphaeriales has not been recommended by Réblová and Gams (2016).
Subclass Hypocreomycetidae O.E. Erikss. & Winka 1997
In Hypocreomycetidae, Maharachchikumbura et al. (2016) recognised ten orders, Yang et al. (2016) introduced the new order Fuscosporellales in Hypocreomycetidae, however, the order is placed in Savoryellomycetidae in this study. There is evidence for six well-resolved orders. The status of Coronophorales, Clavicipitales, Falcocladiales, Hypocreales, Microascales and Torpedosporales are supported with divergence times of 171–241 MYA. There is strong support for the status of the orders Hypocreales, Torpedosporales, Microascales in the MCC tree.
The families in Hypocreomycetidae are mostly well-supported, falling in the recommended divergence times. Graphiaceae in Microascales will need further study to check its ordinal status due to the stem age 166 MYA and its placement is also well-supported in both phylogenetic and MCC trees.
Coronophorales Nannf. 1932
= Melanosporales N. Zhang & M. Blackw. 2007
In the MMC tree, the families listed under Coronophorales in Maharachchikumbura et al. (2016) separate into three main lineages which indicate they may be three separate orders (134–170 MYA). Coronophorales is typified by the monotypic family Coronophoraceae and this is represented by a single strain of Coronophora gregaria which only has TEF and RPB2 sequence data. More species of Coronophora need to be included in analyses to better understand the generic and familial circumscriptions as the present placement of the Coronophoraceae is unstable (Mugambi and Huhndorf 2010). Furthermore, Nitschkiaceae is presently paraphyletic (Mugambi and Huhndorf 2010). We therefore make no further recommendations here. However, the phylogeny supports the inclusion of Melanosporales under Coronophorales to maintain the monophyly of the group. In both the phylogenetic and MCC trees, Melanosporales should be a synonym of Coronophorales, even though the relationship between Melanosporales and other families in Coronophorales has moderate support in the phylogenetic tree. The clade represented by Chaetosphaerellaceae diverged from the above ca 179 MYA with high support in both phylogenetic and MCC trees. We however do not suggest ordinal status for this family until more data are available.
Bionectriaceae Samuels & Rossman 1999 and Nectriaceae Tul. & C. Tul. 1865
According to the literature, the placement of Bionectriaceae is not stable within Hypocreales (Rossman et al. 2001; Liu and Cai 2012; Maharachchikumbura et al. 2016; Su et al. 2016). In the MCC analysis, Niessliaceae is basal to Nectriaceae and sister families, which indicates that Nectriaceae might need raising to ordinal status. However, the placement of Niessliaceae is also unstable (Lombard et al. 2015; Su et al. 2016) and therefore we make no further recommendations.
Clavicipitaceae O.E. Erikss. 1982
= Flammocladiellaceae Crous, L. Lombard & R.K. Schumach. 2015
Flammocladiaceae was introduced by Crous et al. (2015) to accommodate Flammocladiella. In both the phylogenetic and MCC trees this family has poor support as a separate family. This was also shown in the recent study of Su et al. (2016). We therefore synonymize it under Clavicipitaceae.
Nectriaceae Tul. & C. Tul. 1865, Niessliaceae Kirschst. 1939 and Stachybotryaceae L. Lombard & Crous 2014
The families Nectriaceae, Niessliaceae and Stachybotryaceae group and diverge from Clavicipitales 157 MYA in this study, however the placement of Niessliaceae is not well-supported (this study; Lombard et al. 2014), thus these taxa are not dealt with further here until more evidence is available.
Glomerellales Chadef. ex Réblová et al. 2011
The families Australiascaceae, Glomerellaceae, Plectosphaerellaceae and Reticulascaceae are well-supported in both phylogenetic and MCC trees (Fig. 1; Hyde et al. 2017) as in a single order with a stem age of 256 MYA. This age is however, rather old for an order, indicating that Glomerellales should be raised to subclass. There is also support for this in the phylogenetic tree. However, we refrain from introducing a new subclass at this stage because placement of Glomerellales is unstable within Hypocreomycetidae.
Subclass Lulworthiomycetidae Dayarathne et al. 2015
In Lulworthiomycetidae, Maharachchikumbura et al. (2016) recognised three orders, and in this study, there is strong support for the status of Lulworthiales, Koralionastetales in both phylogenetic and MCC trees (289 MYA). The relationship between Pisorisporiales and the above two orders is moderately supported in the phylogenetic tree (Fig. 1), while well-supported in the MCC tree. We therefore include Pisorisporiales in Lulworthiomycetidae. However, as the stem age at 266 MYA falls within that of a subclass, its status may need revision following further study.
Subclass Savoryellomycetidae Hongsanan, K.D. Hyde & Maharachch., subclass nov.
Index Fungorum number: IF553214; Facesoffungi number: FoF03354
Diagnosis The orders Conioscyphales, Pleurotheciales and Savoryellales cluster with strong support in both phylogenetic and MCC trees with a stem age of 268 MYA. This stem age falls in the range of subclass status, we therefore raise Savoryellales to subclass Savoryellomycetidae.
Type order: Savoryellales Boonyuen, Suetrong, Sivichai, K.L. Pang & E.B.G. Jones 2011
Notes: The subclass Savoryellomycetidae, comprises three orders (Conioscyphales, Pleurotheciales and Savoryellales); the MCC tree does support the status of Pleurotheciales with high posterior probability, and stem age at 139 MYA, while it has moderate support in phylogenetic tree. We suggest that the ordinal status of Pleurotheciales can be retained. Conioscyphales is a well-supported order being a sister clade of Pleurotheciales.
Fuscosporellales J. Yang et al. 2016
Based on a combined analysis of LSU, SSU and RPB2 sequence data, Yang et al. (2016) introduced the new order Fuscosporellales with the new family Fuscosporellaceae, as a sister clade to Conioscyphales, Pleurotheciales and Savoryellales. Fuscosporellaceae includes Bactrodesmiastrum and Plagiascoma, and three new genera as Fuscosporella, Mucispora and Parafuscosporella. Members of the Fuscosporellales shared sporodochial conidiomata, monoblastic conidiogenous cells and brown septate conidia (Hernández-Restrepo et al. 2015). The MCC tree supports the status of Fuscosporellales (218 MYA).
Subclass Sordariomycetidae O.E. Erikss. & Winka 1997
= Meliolomycetidae P.M. Kirk & K.D. Hyde 2015
Meliolomycetidae was raised to subclass in Kirk and Hyde (2015) based on the phylogeny shown in Justavino et al. (2015) and this was followed by Maharachchikumbura et al. (2016). However, the support in the phylogenetic tree (Fig. 2) and divergence time in the MCC tree for Meliolomycetidae at 216 MYA does not support its status as a subclass. Therefore, we discard the use of Meliolomycetidae and use only the order Meliolales. In Sordariomycetidae, Maharachchikumbura et al. (2015, 2016) recognised four orders, and in this study, there is evidence for six well-resolved orders. The MCC tree supports the status of Boliniales, Chaetosphaeriales, Coniochaetales, Meliolales (no longer a subclass), Phyllachorales and Sordariales, with stem ages between 145 − 216 MYA.
Coniochaetales Huhndorf, A.N. Mill. & F.A. Fernández 2004
= Cordanales M. Hern.-Rest. & Crous 2015
Notes: In the phylogenetic tree (Fig. 1) Cordanaceae and Coniochaetaceae are sister taxa with a common ancestor at 77 MYA. The status of these families is supported in both trees, however their status as the orders, Coniochaetales and Cordanales is not supported. Cordanales and Coniochaetales should be combined under Coniochaetales based on the most recent common ancestor at 77 MYA and the stem age of the new Coniochaetales is 176 MYA. The ordinal status of Coniochaetales (including Coniochaetaceae and Cordanaceae) is also supported by phylogenetic tree with high support (98% ML).
Cephalothecaceae family incertae sedis in Sordariomycetidae
Although there is moderate support for the placement of Cephalothecaceae in the phylogenetic tree (55% ML), it has high support in the MCC tree. Cephalothecaceae has a divergence time at 175 MYA, thus, the ordinal status of Cephalothecaceae should be considered when we have sufficient data.
Subclass Xylariomycetidae O.E. Erikss. & Winka 1997
Samarakoon et al. (2016) provide evidence for the continuation of Amphisphaeriales and Xylariales as distinct orders in Xylariomycetidae. This is supported in this study, with Amphisphaeriales and Xylariales having diverged 152 MYA, which is in the range of ordinal status. The families in Xylariomycetidae are mostly well-supported falling in the recommended divergence times. There is only one strain was used in this analysis and molecular dating analysis in Hyde et al. (2017) for Melogrammataceae, Hyponectriaceae and Vialaeaceae, however, the divergence time estimates in this study indicate that they can be retained as distinct families.
Amphisphaeriales D. Hawksw. & O.E. Erikss. 1986
This is a well-supported order of Xylariomycetidae both in the phylogenetic analysis and MCC tree. The order contains twelve families Amphisphaeriaceae, Apiosporaceae, Beltraniaceae, Clypeophysalosporaceae, Coniocessiaceae, Hyponectriaceae, Melogrammataceae, Oxydothidaceae, Phlogicylindriaceae, Pseudomassariaceae, Sporodaceae and Vialaeaceae. The MCC tree support was provided for these families, but Coniocessiaceae is placed within Xylariales in Samarakoon et al. (2016). Thus, we do not suggest the placement of Coniocessiaceae, and treat it as family incertae sedis in the subclass Xylariomycetidae.
Oxydothidaceae Konta & K.D. Hyde 2016
Phylogenetic analyses based on combined ITS, LSU and SSU sequence data indicate that Oxydothis species form a distinct lineage related to Vialaeaceae and Coniocessiaceae in the order Xylariales. Hence the new family Oxydothidaceae was introduced by Konta et al. (2016) to accommodate species of Oxydothis which are associated with monocotyledons including Arecaceae (palms), Pandanaceae, Poaceae (Bamboo) and Liliaceae. Species of Oxydothis have been recorded as endophytes, pathogens and saprobes. The MCC tree moderately supports the status of Oxydothidaceae (114 MYA) in Xylariales.
Sporocadaceae Corda 1842
= Bartaliniaceae Wijayaw. et al. 2015
= Discosiaceae Maharachch. & K.D. Hyde 2015
= Pestalotiopsidaceae Maharachch. & K.D. Hyde 2015
= Robillardaceae Crous 2015
Jaklitsch et al. (2016) synonymized Bartaliniaceae, Discosiaceae, Pestalotiopsidaceae and Robillardaceae under Sporocadaceae. Taxa belonging to these families are acervular coelomycetes with the same type of conidiogenesis and conidia (Jaklitsch et al. 2016). The MCC tree indicates that these families have been derived from a common ancestor (67 MYA) and there is not much divergence within the group. Therefore, we follow Jaklitsch et al. (2016) in this paper and consider them synonyms of Sporocadaceae.
Clypeophysalosporaceae Giraldo & Crous 2017
Clypeophysalosporaceae was introduced to accommodate the genera Bagadiella, Clypeophysalospora, Neophysalospora and Plectosphaera which grouped together in an analysis of LSU sequence data (Giraldo et al. 2017). These genera share morphological characteristics of both sexual and asexual morph and are mainly recorded from Eucalyptus spp. in Australia and South Africa. The family is typified by Clypeophysalospora latitans, and Clypeophysalosporaceae forms a sister clade to the family Sporocadaceae within the order Xylariales (Giraldo et al. 2017). The stem age also supports the status of Clypeophysalosporaceae (62 MYA).
Xylariales Nannf. 1932
This is a well-supported order of Xylariomycetidae both in the phylogenetic analysis and MCC tree. The order contains seven families Diatrypaceae, Graphostromataceae, Hypoxylaceae, Lopadostomataceae, Microdochiaceae, Requienellaceae and Xylariaceae. MCC support was provided for these families in Samarakoon et al. (2016), but included Coniocessiaceae which we treat it as family incertae sedis in Xylariomycetidae.
Graphostromataceae M.E. Barr, J.D. Rogers & Y.M. Ju 1993
The family Graphostromataceae was introduced by Barr et al. (1993), but has been synonymized under Xylariaceae (Senanayake et al. 2015; U’Ren et al. 2016). The revision of Wendt et al. (2017) validates its taxonomic status as a family based on the support of the phylogenetic analysis. The MCC tree moderately supports the status of Graphostromataceae (63 MYA) in Xylariales.
Hypoxylaceae DC. 2017
The family Hypoxylaceae was introduced by de Candolle (cf. de Lamarck and de Candolle 1805), but has not been used in modern classifications and treated under Xylariaceae. Multigene phylogeny, morphology and chemotaxonomy used in Wendt et al. (2017) validate and confirm it as a distinct family in Xylariales.
Requienellaceae Boise 1986
Boise (1986) introduced the family Requienellaceae, however, it was not accepted by Eriksson (in Eriksson & Hawksworth 1986) and maintained in Pyrenulaceae. Based on phylogenetic analysis, Jaklitsch et al. (2016) reinstated Requienellaceae as a family of Xylariales. The MCC tree supports the status of Requienellaceae (111 MYA) in Xylariales.
Xylariaceae Tul. & C. Tul. 1863
= Clypeosphaeriaceae G. Winter
Based on the phylogenetic placement of the generic type Clypeosphaeria mamillana within the family Xylariaceae, the use of the family Clypeosphaeriaceae has been discontinued and treated as a synonym of Xylariaceae (Jaklitsch et al. 2016).
Xylariomycetidae family incertae sedis
Cainiaceae J.C. Krug 1978 and Iodosphaeriaceae Locq. 1984
The Cainiaceae and Iodosphaeriaceae clade forms a distinct lineage from the same common ancestor with Amphisphaeriales and Xylariales (this study; Maharachchikumbura et al. 2016), but share the most common ancestor with Xylariales in Hyde et al. (2017) with a divergence time of ca 128 MYA. However, the relationship between Cainiaceae and Iodosphaeriaceae is not well-supported in the phylogenetic and MCC trees, they did not group together in previous studies (Samarakoon et al. 2016a). Thus, we suggest to retain Cainiaceae and Iodosphaeriaceae as families incertae sedis in Xylariomycetidae, until further data are available.
Sordariomycetes, families incertae sedis
Catabotryaceae Petr. 1954
Catabotryaceae diverged from Amplistromatales ca. 165 MYA, however Catabotryaceae has only one strain representing its placement, thus more samples are needed to provide evidence for its ordinal status.
Outlook
The arrangements of higher level taxa resulting from the phylogenetic and molecular clock analyses are not always in agreement. There are many possible reasons for the discrepancies such as using different strains or data set, number of generations and sampling frequency in analysis, models and parameters selection. We have provided some examples of discrepancies between the phylogenetic tree and molecular clock and provide suggestions for the systematic arrangements. We conclude that we cannot use the molecular clock solely as a tool to re-rank Sordariomycetes. The clock must be used in conjunction with the phylogenetic data and other characters in order to derive a stable and natural classification.
The classification of species of Sordariomycetes is far from stabilized as has been shown in several recent studies (Maharachchikumbura et al. 2015, 2016; Senanayake et al. 2016; Jaklitsch et al. 2016; Samarakoon et al. 2016) and this is probably because the group is under represented in terms of collections and sequence data. A greater sampling of taxa in varied habitats, such as freshwater and marine environments, and especially from tropical regions, is needed, and sequence data should be obtained to provide a better understanding of their relationships. In this way, we will eventually derive a stable and natural classification of the class with multifaceted evidence. The molecular clock can be used as additional evidence to derive a stable and natural classification. If both the phylogenetic tree and molecular clock supports the status of a certain rank then this can be followed. However, when there are discrepancies we should not solely follow the molecular clock, but we should take into account all evidence available.
References
Alexopoulos CJ, Mims CW, Blackwell M (1996) Introductory Mycology, 4th edn. Wiley, New York
Ariyawansa HA, Hyde KD, Jayasiri SC, Buyck B, Chethana KWT, Dai DQ, Dai YC, Daranagama DA, Jayawardena RS, Lücking R, Ghobad-Nejhad M, Niskanen T, Thambugala KM, Voigt K, Zhao RL, Li GJ, Doilom M, Boonmee S, Yang ZL, Cai Q, Cui YY, Bahkali AH, Chen J, Cui BK, Chen YY, Monika CD, Dissanayake AJ, Ekanayaka AH, Hashimoto A, Hongsanan S, Jones EBG, Larsson E, Li WJ, Li QR, Liu JK, Luo ZL, Maharachchikumbura SSN, Mapook A, McKenzie EHC, Norphanphoun C, Konta S, Pang KL, Perera RH, Phookamsak R, Phukhamsakda C, Pinruan U, Randrianjohany E, Singtripop C, Tanaka K, Tian CM, Tibpromma S, Abdel-Wahab MA, Wanasinghe DN, Wijayawardene NN, Zhang JF, Zhang H, Abdel-Aziz FA, Wedin M, Westberg M, Ammirati JF, Bulgakov TS, Lima DX, Callaghan TM, Callac P, Chang CH, Coca LF, Dal-Forno M, Dollhofer V, Fliegerová K, Greiner K, Griffith GW, Ho HM, Hofstetter V, Jeewon R, Kang JC, Wen TC, Kirk PM, Kytövuori I, Lawrey JD, Xing J, Li H, Liu ZY, Liu XZ, Liimatainen K, Lumbsch HT, Matsumura M, Moncada B, Moncada S, Parnmen S, de Azevedo Santiago ALCM, Sommai S, Song Y, de Souza CAF, de Souza-Motta CM, Su HY, Suetrong S, Wang Y, Wei SF, Yuan HS, Zhou LW, Réblová M, Fournier J, Camporesi E, Luangsa-ard JJ, Tasanathai K, Khonsanit A, Thanakitpipattana D, Somrithipol S, Diederich P, Millanes AM, Common RS, Stadler M, Yan JY, Li XH, Lee HW, Nguyen TTT, Lee HB, Battistin E, Marsico O, Vizzini A, Vila J, Ercole E, Eberhardt U, Simonini G, Wen HA, Chen XH (2015) Fungal diversity notes 111–252: taxonomic and phylogenetic contributions to fungal taxa. Fungal Divers 75:1–248
Arzanlou M, Groenewald JZ, Gams W, Braun U, Shin HD, Crous PW (2007) Phylogenetic and morphotaxonomic revision of Ramichloridium and allied genera. Stud Mycol 58:57–93
Barr ME (1983) The ascomycetes connection. Mycologia 75:1–13
Barr ME (1987) Prodromus to class Loculoascomycetes. Newell, Amherst
Barr ME (1990) Prodromus to nonlichenized, pyrenomycetous members of class Hymenoascomycetes. Mycotaxon 39:43–184
Barr ME, Rogers JD, Ju YM (1993) Revisionary studies in the Calosphaeriaceae. Mycotaxon 48:529–535
Beimforde C, Feldberg K, Nylinder S, Rikkinen J, Tuovila H, Dörfelt H, Gube M, Jackson DJ, Reitner J, Seyfullah LJ, Schmidt AR (2014) Estimating the phanerozoic history of the Ascomycota lineages: combining fossil and molecular data. Mol Phylogenet Evol 78:386–398
Berbee ML, Taylor JW (1992) Two ascomycete classes based on fruiting-body characters and ribosomal DNA sequence. Mol Biol Evol 9:278–284
Boise J (1986) Requienellaceae, a new family of Loculoascomycetes. Mycologia 78:37–41
Cai L, Tsui CKM, Zhang KQ, Hyde KD (2002) Aquatic fungi from Lake Fuxian, Yunnan, China. Fungal Divers 9:57–70
Crous PW, Wingfield MJ, Guarro J, Hernández-Restrepo M, Sutton DA, Acharya K, Barber PA, Boekhout T, Dimitrov RA, Dueñas M, Dutta AK, Gené J, Gouliamova DE, Groenewald M, Lombard L, Morozova OV, Sarkar J, Smith MTh, Stchigel AM, Wiederhold NP, Alexandrova AV, Antelmi I, Armengol J, Barnes I, Cano-Lira JF, Castañeda Ruiz RF, Contu M, PrR Courtecuisse, da Silveira AL, Decock CA, de Goes A, Edathodu J, Ercole E, Firmino AC, Fourie A, Fournier J, Furtado EL, Geering ADW, Gershenzon J, Giraldo A, Gramaje D, Hammerbacher A, He X-L, Haryadi D, Khemmuk W, Kovalenko AE, Krawczynski R, Laich F, Lechat C, Lopes UP, Madrid H, Malysheva EF, Marín-Felix Y, Martín MP, Mostert L, Nigro F, Pereira OL, Picillo B, Pinho DB, Popov ES, Rodas Peláez CA, Rooney-Latham S, Sandoval-Denis M, Shivas RG, Silva V, Stoilova-Disheva MM, Telleria MT, Ullah C, Unsicker SB, van der Merwe NA, Vizzini A, Wagner H-G, Wong PTW, Wood AR, Groenewald JZ (2015) Fungal Planet description sheets: 320–370. Persoonia 34:167–266
Du Z, Hyde KD, Yang Q, Liang YM, Tian CM (2017) Melansporellaceae: a novel family of Diaporthales (Ascomycota). Phytotaxa 305:191–200
Eriksson OE, Hawksworth DL (1986) Notes on ascomycete systematics. Nos 1–224. Syst Ascomycetum 5:113–174
Eriksson O, Hawksworth DL (1993) Outline of the Ascomycetes—1993. Syst Ascomycetum 12:1–257
Eriksson OE, Winka K (1997) Supraordinal taxa of Ascomycota. Myconet 1:1–16
Fournier J, Lechat C (2010) Phomatospora luteotingens sp. nov., a new aquatic species of Phomatospora from France and Spain. Mycosphere 1:39–43
Garnica G, Riess K, Schön ME, Oberwinkler F, Setaro SD (2016) Divergence times and phylogenetic patterns of Sebacinales, a highly diverse and widespread fungal lineage. PLoS ONE 11(3):e0149531
Giraldo A, Crous PW, Schumacher RK, Cheewangkoon R, Ghobad-Nejhad M, Langer E (2017) The genera of fungi-G3: aleurocystis, Blastacervulus, Clypeophysalospora, Licrostroma, Neohendersonia and Spumatoria. Mycol Prog 16:1–24
Guo LD, Hyde KD, Liew EC (2001) Detection and taxonomic placement of endophytic fungi within frond tissues of Livistona chinensis based on rDNA sequences. Mol Phylogenet Evol 20:1–13
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Hernández-Restrepo M, Gené J, Castañeda-Ruiz RF, Mena-Portales J, Guarro J (2015) Emendation of the genus Bactrodesmiastrum (Sordariomycetes) and description of Bactrodesmiastrum monilioides sp. nov. from plant debris in Spain. Mycol Prog 14:48
Ho WH, Hyde KD, Hodgkiss IJ (2001) Fungal communities on submerged wood from streams in Brunei, Hong Kong and Malaysia. Mycol Res 105:1492–1501
Hongsanan S, Sánchez-Ramírez S, Crous PW, Ariyawansa HA, Zhao RL, Hyde KD (2016) The evolution of fungal epiphytes. Mycosphere 7:1690–1712
Huhndorf SM, Greif M, Miller AN (2008) Two new genera in the Magnaporthaceae, a new addition to Ceratosphaeria and two new species of Lentomitella. Mycologia 100:940–955
Hyde KD, Jones EBG (1989) Intertidal mangrove fungi from Brunei. Lautospora gigantea gen. et sp. nov., a new loculoascomycete from prop roots of Rhizophora spp. Bot Mar 32:479–482
Hyde KD, Taylor JE, Fröhlich J (1999) Two new species of Pseudohalonectria from palms. Mycologia 91:520–524
Hyde KD, Nilsson RH, Alias SA, Ariyawansa HA, Blair JE, Cai L, de Cock AWAM, Dissanayake AJ, Glockling SL, Goonasekara ID, Gorczak M, Hahn M, Jayawardena RS, van Kan JAL, Laurence MH, Lévesque CA, Li XH, Liu JK, Maharachchikumbura SSN, Manamgoda DS, Martin FN, Mckenzie EHC, McTaggart AR, Mortimer PE, Nair PVR, Pawlowska J, Rintoul TL, Shivas RG, Spies CFJ, Summerell BA, Taylor PWJ, Terhem RB, Udayanga D, Vaghefi N, Walther G, Wilk M, Wrzosek M, Xu JC, Yan JY, Zhou N (2014) One stop shop: backbones trees for important phytopathogenic genera: I. Fungal Divers 67:21–125
Hyde KD, Hongsanan S, Jeewon R et al (2016) Fungal diversity notes 367–491: taxonomic and phylogenetic contributions to fungal taxa. Fungal Divers 80:1–270
Hyde KD, Maharachchikumbura SSN, Hongsanan S, Samarakoon MC et al. (2017) The ranking of fungi—a tribute to David L. Hawksworth on his 70th birthday. Fungal Divers 84: (in press)
Jaklitsch WM, Gardiennet A, Voglmayr H (2016) Resolution of morphology-based taxonomic delusions: Acrocordiella, Basiseptospora, Blogiascospora, Clypeosphaeria, Hymenopleella, Lepteutypa, Pseudapiospora, Requienella, Seiridium and Strickeria. Persoonia 37:82–105
Jones EBG, Sakayaroj J, Suetrong S, Somrithipol S, Pang KL (2009a) Classification of marine Ascomycota, anamorphic taxa and Basidiomycota. Fungal Divers 35:1–187
Jones EBG, Zuccaro A, Mitchell J, Nakagiri A, Chatmala I, Pang KL (2009b) Phylogenetic position of freshwater and marine Sigmoidea species: introducing a marine hyphomycete Halosigmoidea gen. nov. (Halosphaeriales). Bot Mar 52:349–359
Jones EBG, Suetrong S, Sakayaroj J, Bahkali AH, Abdel-Wahab MA, Boekhout T, Pang KL (2015) Classification of marine Ascomycota, Basidiomycota, Blastocladiomycota and Chytridiomycota. Fungal Divers 73:1–72
Justavino DR, Kirschner R, Piepenbring M (2015) New species and new records of Meliolaceae from Panama. Fungal Divers 70:73–84
Katoh K, Asimenos G, Toh H (2009) Multiple alignment of DNA sequences with MAFFT. Methods Mol Biol 537:39–64
Keim J, Mishra B, Sharma R, Ploch S, Thines M (2014) Root-associated fungi of Arabidopsis thaliana and Microthlaspi perfoliatum. Fungal Divers 66:99–111
Kirk PM, Hyde KD (2015) Nomenclatural novelties: Paul. M. Kirk. Index Fungorum. 242:1
Konta S, Hongsanan S, Tibpromma S, Thongbai B, Maharachchikumbura SSN, Bahkali AH, Hyde KD, Boonmee S (2016) An advance in the endophyte story: Oxydothidaceae fam. nov. with six new species of Oxydothis. Mycosphere 7:1425–1446
Li GJ, Hyde KD, Zhao RL, Hongsanan S, Abdel-Aziz FA, Abdel-Wahab MA, Alvarado P, Alves-Silva G, Ammirati SF, Ariyawansa HA, Baghela A, Bahkali AH, Beug M, Bhat DJ, Bojantchev D, Boonpratuang T, Bulgakov TS, Camporesi E, Boro MC, Ceska O, Chakraborty D, Chen JJ, Chethana KWT, Chomnunti P, Consiglio G, Cui BK, Dai DQ, Dai YC, Daranagama DA, Das K, Dayarathne MC, De Crop E, De Oliveira RJV, de Souza CAF, de Souza JI, Dentinger BTM, Dissanayake AJ, Doilom M, Drechsler-Santos ER, Ghobad-Nejhad M, Gilmore SP, Góes-Neto A, Gorczak M, Haitjema CH, Hapuarachchi KK, Hashimoto A, He MQ, Henske JK, Hirayama K, Iribarren MJ, Jayasiri SC, Jayawardena RS, Jeon SJ, Jerônimo GH, Jesus AL, Jones EBG, Kang JC, Karunarathna SC, Kirk PM, Konta S, Kuhnert E, Langer E, Lee HS, Lee HB, Li WJ, Li XH, Liimatainen K, Lima DX, Lin CG, Liu JK, Liu XZ, Liu ZY, Luangsa-ard JJ, Lücking R, Lumbsch HT, Lumyong S, Leaño EM, Marano AV, Matsumura M, McKenzie EHC, Mongkolsamrit S, Mortimer PE, Nguyen TTT, Niskanen T, Norphanphoun C, O’Malley MA, Parnmen S, Pawłowska J, Perera RH, Phookamsak R, Phukhamsakda C, Pires-Zottarelli CLA, Raspé O, Reck MA, Rocha SCO, de Santiago ALCMA, Senanayake IC, Setti L, Shang QJ, Singh SK, Sir EB, Solomon KV, Song J, Srikitikulchai P, Stadler M, Suetrong S, Takahashi H, Takahashi T, Tanaka K, Tang LP, Thambugala KM, Thanakitpipattana D, Theodorou MK, Thongbai B, Thummarukcharoen T, Tian Q, Tibpromma S, Verbeken A, Vizzini A, Vlasák J, Voigt K, Wanasinghe DN, Wang Y, GothamieWeerakoon G, AnWen H, Wen TC, Wijayawardene NN, Wongkanoun S, Wrzosek M, Xiao YP, Xu JC, Yan JY, Yang J, Yang SD, Hu Y, Zhang JF, Zhao J, Zhou LW, Peršoh D, Phillips AJL, Maharachchikumbura SSN (2016) Fungal diversity notes 253–366: taxonomic and phylogenetic contributions to fungal taxa. Fungal Divers 78:1–237
Liu F, Cai L (2012) Morphological and molecular characterization of a novel species of Simplicillium from China. Cryptogam Mycol 33:137–144
Liu JK, Hyde KD, Jones EBG, Ariyawansa HA, Bhat DJ, Boonmee S, Maharachchikumbura S, McKenzie EHC, Phookamsak R, Phukhamsakda R, Abdel-Wahab MA, Buyck B, Chen J, Chethana KWT, Singtripop C, Dai DQ, Dai YC, Daranagama DA, Dissanayake AJ, Doliom M, Fan LX, Goonasekara D, Hirayama K, Hongsanan S, Jayasiri SC, Jayawardena RS, Karunarathna SC, Li WJ, Mapook A, Norphanphoun C, Pang KL, Perera RH, Peršoh D, Pinruan U, Senanayake IC, Somrithipol S, Satinee S, Tanaka K, Thambugala KM, Tian Q, Tibpromma S, Udayanga D, Wijayawardene NN, Wanasinghe D, Abdel-Aziz FA, Adamčík S, Bahkali AH, Boonyuen N, Bulgakov T, Callac P, Chomnunti P, Greiner K, Hashimoto A, Hofstetter V, Kang JC, Li XH, Liu ZY, Matumura M, Mortimer PE, Rambold R, Randrianjohany E, Sato G, Indrasutdhi VS, Verbeken A, Brackel W, Wang Y, Wen TC, Xu JC, Yan JY, Zhao RL, Camporesi E (2015) Fungal diversity notes 1–110: taxonomic and phylogenetic contributions to fungal species. Fungal Divers 72:1–197
Lombard L, van der Merwe NA, Groenewald JZ (2014) Lineages in Nectriaceae: re-evaluating the generic status of Ilyonectria and allied genera. Phytopathologia Mediterranea 53:515–532
Lombard L, van der Merwe NA, Groenewald JZ, Crous PW (2015) Generic concepts in Nectriaceae. Stud Mycol 80:189–245
Lumbsch HT (2000) Phylogeny of filamentous ascomycetes. Naturwissenchaften 87:335–342
Lumbsch HT, Huhndorf SM (2010) Myconet volume 14 part one. Outline of ascomycota-2009. Fieldiana Life Earth Sci 1:1–922
Maharachchikumbura SSN, Guo LD, Cai L (2012) A multi-locus backbone tree for Pestalotiopsis, with a polyphasic characterization of 14 new species. Fungal Divers 56:95–129
Maharachchikumbura SN, Hyde KD, Jones EBG, McKenzie EHC, Huang SK, Abdel-Wahab MA, Daranagama DA, Dayarathne M, D’souza MJ, Goonasekara ID, Hongsanan S, Jayawardena RS, Kirk PM, Konta S, Liu JK, Liu Z-Y, Norphanphoun C, Pang KL, Perera RH, Senanayake IC, Shang Q, Shenoy BD, Xiao Y, Bahkali AH, Kang J, Somrothipol S, Suetrong S, Wen T, Xu J (2015) Towards a natural classification and backbone tree for Sordariomycetes. Fungal Divers 72:199–301
Maharachchikumbura SSN, Hyde KD, Jones EBG, McKenzie EHC, Bhat JD, Dayarathne MC, Huang S-K, Norphanphoun C, Senanayake IC, Perera RH, Shang Q-J, Xiao Y, Dsouza MJ, Hongsanan S, Jayawardena RS, Daranagama DA, Konta S, Goonasekara ID, Zhuang WY, Jeewon R, Phillips AJL, Abdel-Wahab MA, Al-Sadi AM, Bahkali AH, Boonmee S, Boonyuen N, Cheewangkoon R, Dissanayake AJ, Kang J, Li Q-R, Liu JK, Liu XZ, Liu Z-Y, Luangsa-ard JJ, Pang K-L, Phookamsak R, Promputtha I, Suetrong S, Stadler M, Wen T, Wijayawardene NN (2016) Families of Sordariomycetes. Fungal Divers 79:1–317
Minoura K, Muroi T (1978) Some freshwater ascomycetes from Japan. Trans Mycol Soc Jpn 19:129–134
Mitchell JI, Roberts PJ, Moss ST (1995) Sequence or structure? A short review on the application of nucleic acid sequence information of fungal taxonomy. Mycologist 9:67–75
Mugambi GK, Huhndorf SM (2010) Multigene phylogeny of the Coronophorales: morphology and new species in the order. Mycologia 102:185–210
Norphanphoun C, Hongsanan S, Doilom M, Bhat DJ, Wen TC, Senanayake IC, Bulgakov TS, Hyde KD (2016) Lamproconiaceae fam. nov. to accommodate Lamproconium desmazieri. Phytotaxa 270:89–102
Pattengale ND, Alipour M, Bininda-Emonds OR, Moret BM, Stamatakis A (2009) How many bootstrap replicates are necessary? In: Annual international conference on research in computational molecular biology. Springer, Berlin, pp 184–200
Perera RH, Maharachchikumbura SSN, Ariyawansa H, Bahkali AH, Jones EBG, Al-Sadi AM, Hyde KD, Liu ZY (2016) Two new Pseudohalonectria species on beech cupules (Fagus sylvatica) and a new genus to accommodate P. suthepensis. Phytotaxa 278:115–131
Pérez-Ortega S, Garrido-Benavent I, Grube M, Olmo R, de los Ríos A (2016) Hidden diversity of marine borderline lichens and a new order of fungi: Collemopsidiales (Dothideomyceta). Fungal Divers 80:285–300
Prados-Rosale RC, Roldán-Rodríguez R, Serena C, López-Berges MS, Josep Guarro J, Martínez-del-Pozo A, Di Pietro A (2012) A PR-1-like protein of Fusarium oxysporum functions in virulence on mammalian host. J Biol Chem 287:21970–21979
Promputtha I, Jeewon R, Lumyong S, McKenzie EHC, Hyde KD (2005) Ribosomal DNA fingerprinting in the identification of non sporulating endophytes from Magnolia liliifera (Magnoliaceae). Fungal Divers 20:167–186
Rambaut A (2006) FigTree. Tree figure drawing tool version 1.3.1. Institute of Evolutionary Biology, University of Edinburgh, Edinburgh. http://tree.bio.ed.ac.uk/software/figtree/
Réblová M, Gams W (2016) A revision of Sphaeria pilosa Pers. and re-evaluation of the Trichosphaeriales. Mycol Prog 15:1–8
Réblová M, Fournier J, Štěpánek V (2016) Two new lineages of aquatic ascomycetes: Atractospora gen. nov. and Rubellisphaeria gen. et sp. nov., and a sexual morph of Myrmecridium montsegurinum sp. nov. Mycol Prog 15:1–18
Rossman AY, McKemy JM, Pardo-Schultheiss RA, Schroers HJ (2001) Molecular studies of the Bionectriaceae using large subunit rDNA sequences. Mycologia 93:100–110
Samarakoon MC, Hyde KD, Promputtha I, Hongsanan S, Ariyawansa HA, Maharachchikumbura SSN, Daranagama DA, Stadler M, Mapook A (2016) Evolution of Xylariomycetidae (Ascomycota: Sordariomycetes). Mycosphere 7:1746–1761
Schoch CL, Crous PW, Groenewald JZ, Boehm EWA, Burgess TI, de Gruyter J, de Hoog GS, Dixon LJ, Grube M, Gueidan C, Harada Y, Hatakeyama S, Hirayama K, Hosoya T, Huhndorf SM, Hyde KD, Jones EGB, Kohlmeyer J, Kruys Å, Li YM, Lücking R, Lumbsch HT, Marvanová L, Mbatchou JS, McVay AH, Miller AN, Mugambi GK, Muggia L, Nelsen MP, Nelson P, Owensby CA, Phillips AJL, Phongpaichit S, Pointing SB, Pujade-Renaud V, Raja HA, Plata ER, Robbertse B, Ruibal C, Sakayaroj J, Sano T, Selbmann L, Shearer CA, Shirouzu T, Slippers B, Suetrong S, Tanaka K, Volkmann- Kohlmeyer B, Wingfield MJ, Wood AR, Woudenberg JHC, Yonezawa H, Zhang Y, Spatafora JW (2009) A class-wide phylogenetic assessment of Dothideomycetes. Stud Mycol 64:1–15
Senanayake IC, Maharachchikumbura SSN, Hyde KD, Bhat JD, Jones EBG, McKenzie EHC, Dai DQ, Daranagama DA, Dayarathne MC, Goonasekara ID, Konta S, Li WJ, Shang QJ, Stadler M, Wijayawardene NN, Xiao YP, Norphanphoun C, Li Q, Liu XZ, Bahkali AH, Kang JC, Wang Y, Wen TC, Wendt L, Xu JC, Camporesi E (2015) Towards unraveling relationships in Xylariomycetidae (Sordariomycetes). Fungal Divers 73:73–144
Senanayake IC, Al-Sadi AM, Bhat JD, Camporesi E, Dissanayake AJ, Lumyong S, Maharachchikumbura SSN, Hyde KD (2016) Phomatosporales ord. nov. and Phomatosporaceae fam. nov., to accommodate Lanspora, Phomatospora and Tenuimurus, gen. nov. Mycosphere 7:628–641
Shearer CA (1989) Pseudohalonectria (Lasiosphaeriaceae), an antagonistic genus from wood in freshwater. Can J Bot 67: 1944–1955
Shearer A, Zare-Maivan H (1988) In vitro hyphal interactions among wood- and leaf-inhabiting Ascomycetes and Fungi Imperfecti from freshwater habitats. Mycologia 80:31–37
Silvestro D, Michalak I (2012) raxmlGUI: a graphical front-end for RAxML. Org Divers Evol 12:335–337
Spatafora JW (1995) Ascomal evolution of filamentous ascomycetes: evidence from molecular. Can J Bot 73:811–815
Spatafora JW, Blackwell M (1993) Molecular systematics of unitunicate perithecial ascomycetes: the Clavicipitales–Hypocreales connection. Mycologia 85:912–922
Su HY, Hyde KD, Maharachchikumbura SSN, Ariyawansa HA, Luo ZL, Promputtha I, Tian Q, Lin CG, Shang QJ, Zhao YC, Chai HM, Liu XY, Bahkali AH, Bhat DJ, McKenzie EHC, Zhou DQ (2016) The families Distoseptisporaceae fam. nov., Kirschsteiniotheliaceae, Sporormiaceae and Torulaceae, with new species from freshwater in Yunnan Province, China. Fungal Divers 80:375–409
Suetrong S, Klaysuban A, Sakayaroj J, Preedanon S, Rung-areerate P, Phongpaichit S, Jones EBG (2015) Tirisporellaceae, a new family in the order Diaporthales (Sordariomycetes, Ascomycota). Cryptogam, Mycol 36:319–330
Sung G-H, Hywel-Jones NL, Sung J-M, Luangsaard JJ, Shresthra B, Spatafora JW (2007) Phylogenetic classification of Cordyceps and the clavicipitaceous fungi. Stud Mycol 57:5–59
Tang AM, Jeewon R, Hyde KD (2007) Phylogenetic utility of protein (RPB2, β-tubulin) and ribosomal (LSU, SSU) gene sequences in the systematics of Sordariomycetes (Ascomycota, Fungi). Antonie Van Leeuwenhoek 91(4):327
Tsui CKM, Hyde KD, Hodgkiss IJ (2000) Biodiversity of fungi on submerged wood in Hong Kong streams. Aquat Microb Ecol 21:289–298
U’Ren JM, Miadlikowska J, Zimmerman NB, Lutzoni F, Stajich JE, Arnold AE (2016) Contributions of North American endophytes to the phylogeny, ecology, and taxonomy of Xylariaceae (Sordariomycetes, Ascomycota). Mol Phylogenet Evol 98:210–232
Vijaykrishna D, Jeewon R, Hyde KD (2006) Molecular taxonomy, origins and evolution of freshwater ascomycetes. Fungal Divers 23:351–390
Voglmayr H, Castlebury LA, Jaklitsch WM (2017) Juglanconis gen. nov. on Juglandaceae, and the new family Juglanconidaceae (Diaporthales). Persoonia 38:136–155
Wang H, Guo S, Huang M, Thorsten LH, Wei J (2010) Ascomycota has a faster evolutionary rate and higher species diversity than Basidiomycota. Sci China Life Sci 53(10):1163–1169
Wendt L, Sir EB, Kuhnert E, Heitkämper Lambert C, Hladki AI, Romero AI, Luangsa-ard JJ, Srikitikulchai P, Peršoh Stadler M (2017) Resurrection and emendation of the Hypoxylaceae, recognised from a multi-gene geneology of the Xylariales. Mycol Prog. doi:10.1007/s11557-017-1303-3
Yang J, Maharachchikumbura SSN, Bhat DJ, Hyde KD, McKenzie EHC, Jones EBG, Al-Sadi AM, Lumyong S (2016) Fuscosporellales, a new order of aquatic and terrestrial Hypocreomycetidae (Sordariomycetes). Cryptogr Mycol 37:449–475
Zhang N, Castlebury LA, Miller AN, Huhndorf SM, Schoch CL, Seifert KA, Rossman AY, Rogers JD, Kohlmeyer J, Volkmann-Kohlmeyer B, Sung GH (2006) An overview of the systematics of the Sordariomycetes based on four-gene phylogeny. Mycologia 98:1076–1087
Zhao RL, Zhou JL, Chen J, Margaritescu S, Sánchez-Ramírez S, Hyde KD, Callac P, Parra LA, Li GJ, Moncalvo JM (2016) Towards standardizing taxonomic ranks using divergence times—a case study for reconstruction of the Agaricus taxonomic system. Fungal Divers 78:239–292
Zhao RL, Li GJ, Sánchez-Ramírez S, Stata M, Moncalvo J-M, Yang ZL, Wu G, Dai YC, He SH, Cui BK, Zhou JL, Wu F, He MQ, Hyde KD (2017) A six-genes phylogenetic overview of Basidiomycota and allied phyla with estimated divergence times of higher taxa and a phyla proteomics perspective. Fungal Divers 84: (in press)
Acknowledgements
K.D. Hyde thanks the Chinese Academy of Sciences, Project Number 2013T2S0030, for the award of Visiting Professorship for Senior International Scientists at Kunming Institute of Botany. The authors extend their appreciation to the International Scientific Partnership Program ISPP at King Saud University for funding this research work through ISPP# 0089. K.D. Hyde would like to thank the Thailand Research Fund (TRF) Grant No. RSA5980068 entitled Biodiversity, phylogeny and role of fungal endophytes on above parts of Rhizophora apiculata and Nypa fruticans and National Research Council of Thailand (NRCT) entitled Diseases of mangrove trees and maintenance of good forestry practice (Grant Number: 60201000201). The authors would like to thank NRCT for the grant to study the Biodiversity, phylogeny and role of fungal endophytes of Pandanaceae (Grant No.: 592010200112).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hongsanan, S., Maharachchikumbura, S.S.N., Hyde, K.D. et al. An updated phylogeny of Sordariomycetes based on phylogenetic and molecular clock evidence. Fungal Diversity 84, 25–41 (2017). https://doi.org/10.1007/s13225-017-0384-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13225-017-0384-2