Abstract
This study provides new insights on the phylogenetic position of the lichenicolous fungal genus Abrothallus based on six molecular markers (nuSSU, nuLSU, mtSSU, RPB1, RPB2 and TEF-α). In a broad-scale analysis, we detected high support for inclusion of the genus within Dothideomycetes. A further analysis provided support for Abrothallus as a member of the subclass Pleosporomycetidae as a sister group of Jahnulales, an order of aquatic Dothideomycetes. Given the exclusive characters of this group of apotheciate fungi within the Dothidiomycetes, a new monotypic order Abrothallales is here introduced together with the new family Abrothallaceae. In a multi-locus analysis (based on the six loci indicated above plus ITS) restricted to 12 putative Abrothallus species, two clearly separated clades were observed: one comprising species growing on lichens of the families Parmeliaceae and Ramalinaceae, and the second including species that live on lichens of the order Peltigerales and the family Cladoniaceae.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Although many fungal species are known to live on lichen thalli (Lawrey and Diederich 2003; Arnold et al. 2009; Peršoh and Rambold 2012; U’Ren et al. 2012), only those that show recognizable sexual or asexual reproductive structures are denoted lichenicolous fungi. The heterogeneity of lichenicolous fungi is manifested through both their phylogenetic diversity and their life-strategies, ranging from aggressive parasitism to commensalism and saprophytism including several intermediate modes of life (Lawrey and Diederich 2003). Despite varying interest for lichenicolous fungi over the past two centuries, the introduction of new research techniques has prompted several studies designed to revisit these much-specialized enigmatic fungi (e.g. the comprehensive review by Lawrey and Diederich 2003). Within the past 10 years, the number of studies based on molecular methods has constantly grown and this has provided new insight into the origin of their life-strategies, speciation and population patterns, and host-specificity (e.g. Sikaroodi et al. 2001; Peršoh and Rambold 2002; Millanes et al. 2011; Lawrey et al. 2012; Werth et al. 2013).
Despite intensified research efforts, phylogenetic relationships remain obscure for numerous lichenicolous genera. One such case is the genus Abrothallus, which is one of the most easily recognizable fungal genera that exclusively grows on lichens (Fig. 1). The genus was introduced to accommodate the single species A. bertianus by De Notaris (1845), and although this species was first described as lichenized, its lichenicolous nature was soon to be established (De Notaris 1846; Tulasne 1852; Montagne 1856). The genus is well-characterized by its: (1) more-or-less globose apothecioid ascomata, often covered with green or golden pruina, (2) bitunicate asci with four to eight, 2- to 4-celled brown asymmetric ascospores with evident ornamentation, (3) ramified-anastomosed interascal filaments, (4) epiphymenial layer with granulose pigments, which commonly dissolve in potassium hydroxide, and (5) its pycnidial Vouauxiomyces-type anamorph (Diederich 2004; Suija 2006; Pérez-Ortega et al. 2011). Reports of the genus have spanned all continents except Antarctica, and representatives of the genus have been described to grow on a wide variety of foliose and fruticose lichens especially those belonging to the families Parmeliaceae and Lobariaceae, but also on species of Nephromataceae, Stereocaulaceae, Cladoniaceae, Ramalinaceae and Pannariaceae.
The genus Abrothallus shows no clear similarity to any other ascomycete genus, prompting several hypotheses about its phylogenetic relationships. Before its parasitic nature was discovered (De Notaris 1846; Tulasne 1852; Montagne 1856), the genus was classified together with lichens as a member of tribe Coccocarpiae of the order Gymnocarpi (Montagne 1851) or suborder Biatorinae of Lecideaceae (Körber 1855; Lindsay 1857). The subsequent authors ascribed it to the order Bulgariacei (= Bulgariaceae, Helotiales) (Montagne 1856) or suborder Phacidiae (= Phacidiaceae, Helotiales) (Saccardo 1889) based on its apparent lack of apothecial margin. Rehm (1896) considered it a member of the Patellariaceae (then Dermatales), a designation supported by several investigators (e.g. Lindau 1897; Vouaux 1913; von Keissler 1929; Kutorga and Hawksworth 1997). However, based on detailed ultrastructural studies, Bellemère et al. (1986) suggested its affinity to Arthoniales, as proposed earlier by Jatta (1911), who included the genus in the tribe Arthoniae, and later followed by Nannfeldt (1932). Fink (1935), who synonymized Abrothallus with Buelliella, proposed its relationship with Buelliaceae due to its dark-coloured ascospores. None of these hypotheses has been satisfactory addressed using molecular methods, and the status of the genus is currently Ascomycetes incertae sedis (Lumbsch and Huhndorf 2010). In preliminary work by Granberg (2001), the phylogenetic adscription of Abrothallus was tested using ribosomal DNA markers (nuSSU, nuLSU), yet results were inconclusive.
Characteristic morphological and anatomical features of the lichenicolous genus Abrothallus. a Greenish pruina on the apothecia of A. secedens; b Convex apothecia of A. acetabuli; c Cross-section of the apothecium of A. acetabuli; d Scanning electron microscope image of a cross-section of A. parmeliarum growing on Parmelia saxatilis; e Paraphyses of A. parmeliarum in phloxin; f Mature ascus with mature ascospores; g Young ascus; h Conidiogenous cells with conidiospores of A. bertianus; i Ascospore with one septum in A. bertianus; j Ascospore with three septa in A. suecicus; k Ascospores divided into two part-spores in A. secedens; l Surface view of pycnidia of A. bertianus on the thallus of Melanelixia fuliginosa; m Cross-section of a pycnidium of A. bertianus observed by scanning electron microscopy; n Detail of the rugose surface of ascospores of A. parmeliarum observed by scanning electron microscopy; o Conidiospores of A. bertianus. Scale bars: a, b, c, d, l: 100 μm; e, f, g, o: 10 μm; h, i, j, k, n: 5 μm
The intra-generic taxonomy of the genus Abrothallus has also been a topic of controversy because of the different weight given to characters such as host-specificity or the reaction of somatic hyphae with Lugol’s solution. Hence, the existence of both broad (Lindsay 1857; von Keissler 1929; Hawksworth 1983) and narrow definitions (Kotte 1909; Clauzade et al. 1989; Santesson et al. 2004) has lead to the recognition of certain taxa only by some authors. For instance, Suija (2006) showed that the colour of the crystalline layer above the hymenium, the pruinosity of the ascomata and Lugol’s solution reacting somatic hyphae are the most reliable characters for species recognition among 2-celled species of the genus.
This study was designed to resolve the phylogenetic relationships of Abrothallus within the Ascomycota through multi-locus analysis. Once this had been established, we addressed relationships within the genus by examining seven loci.
Materials and methods
Taxon sampling and morpho-anatomical characterization
This study was performed on 14 specimens of Abrothallus covering its widest possible host range: three specimens growing on Peltigerales (on Pseudocyphellaria and Nephroma), one specimen on Cladonia, two on Ramalina and eight on Parmeliaceae (Parmelinopsis, Parmelia, Parmelina, Parmotrema, Pleurosticta and Usnea). Voucher specimens were deposited in TU, NY and MA (Table 1).
Specimens were examined using standard microscopy techniques. For colour tests, we used c. 10 % potassium hydroxide (KOH; K) and c. 50 % nitric acid (HNO3; N); Lugol’s solution (I) was used to examine fungal structure reactions. Morphometric measurements were performed in water and values recorded as the minimum–(average)–maximum. Portions of air-dried thalli were gold sputter-coated and observed by electron microscopy with back-scattered electron imaging (FEI INSPECT) at the facilities of the MNCN (CSIC, Madrid). Nomarski differential interference contrast (DIC) micrographs were captured using a Zeiss® AX10 microscope and Zeiss® AxioCam digital camera.
Molecular methods
Molecular analysis
DNA extraction, amplification and sequencing
For DNA extraction, apothecia were removed from the host thallus with the help of a razor blade or forceps and immersed in a sterile water drop. The lower section of each apothecium, including most of the hypothecium, was carefully removed using a razor blade under the dissecting microscope and transferred to a 1.5 ml test tube. The number of apothecia used for DNA extraction was 1 to 12. DNA was extracted using the DNEasy Plant Mini Kit (Qiagen®) and High Pure PCR Template Preparation Kit (Roche Applied Science®) following the manufacturer’s instructions with minor modifications.
To determine the phylogenetic position of Abrothallus and infer relationships among taxa within the genus, we amplified six nuclear genomic regions (ITS, nuLSU, nuSSU, TEF-α, RPB1 and RPB2) and one mitochondrial locus (mtSSU). The following primers were used: ITS1F (Gardes and Bruns 1993) and ITS4 (White et al. 1990) for ITS; nuSSU97A, nSSU131, nSSU1088r (Kauff and Lutzoni 2002) or NS1 (White et al. 1990) and NS24 (Gargas and Taylor 1992) for nuSSU; LR0R (Rehner and Samuels 1994), LR1R (Moncalvo et al. 1993), LR3r, LR7 and LR5 (Vilgalys and Hester 1990) for nuLSU; RPB1-Af (Stiller and Hall 1997) and RPB1-Cr (Matheny et al. 2002) or RPB1-Afasc and RPB1-6R1asc (Hofstetter et al. 2007) for RPB1; fRPB2-5F and fRPB2-7cR (Liu et al. 1999) for RPB2; TEF983F and TEF2218R (initially obtained from S. Rehner: http://ocid.nacse.org/research/deephyphae/EF1primer.pdf) for TEF-α; and mrSSU1 and mrSSU3R (Zoller et al. 1999) for the mtSSU region.
PCR reactions were prepared in a 25 μl final volume containing either 1.25 (10 μM) or 1 μl (20 μM) of each primer, 17.5 μl or 13 μl of distilled water and 5 μl or 10 μl of the DNA template. PuReTaq Ready-To-Go PCR beads (GE Health Care, Amersham Biosciences, 2004) were added to the mix according to the manufacturer’s instructions.
PCR amplifications were performed in a GeneAmp PCR System 2400 (Applied Biosystems®) or Eppendorf Mastercycler® thermal cyclers. PCR (performed by SPO) conditions were as follows: an initial 4 min heating step at 94 °C, followed by 35 cycles of 1.15 min at 94 °C, 1.30 min at 52 °C (for RPB1, RPB2, nuLSU, nuSSU and mtSSU) and 1.45 min at 72 °C. After a final extension step of 7 min at 72 °C, the samples were stored at 4 °C. Conditions for amplification of the ITS and TEF-α regions were an initial step of 3 cycles at an annealing temperature of 54 °C, followed by 30 cycles with the annealing temperature set at 48 °C. PCR protocols (performed by AS) provided by Hofstetter et al. (2007) were used for RPB1; Liu et al. (1999) for RPB2; Rehner (http://ocid.nacse.org/research/deephyphae/EF1primer.pdf) for TEF-α, and Zoller et al. (1999) for mtSSU. For nuLSU and ITS, conditions were an initial 3 min heating step at 95 °C, followed by 35 cycles of 30 s at 95 °C, 30 s at 55 °C and 1 min at 72 °C; the final extension step was 10 min. at 72 °C followed by storage for 10 min at 6 °C. For nuSSU, conditions were an initial 1 min heating step at 95 °C, followed by 30 cycles of 1 min at 95 °C, 1 min at 55 °C, 2 min at 72 °C, and a final 5 min extension step at 72 °C. When needed, ‘touchdown’ (Don et al. 1991) preceded the PCR cycle. PCR products were purified using Exo-Sap enzymes (GE Healthcare, Freiburg, Germany) or Ultra Clean PCR Clean-up (Mo-Bio, Ca U.S.A). Both complementary strands were sequenced in a Macrogen Inc. system (Seoul, Korea) using the same primer set as for PCR amplification with the exception of ITS for which ITS5 (White et al. 1990) was used instead of ITS1F in some cases. Contigs were assembled using Sequencher 4.7 software (GeneCodes Corp., Ann Arbor, MI, USA).
Sequence alignment and phylogenetic analysis
In the first analysis, a six-locus (nuSSU, nuLSU, mtSSU, RPB1, RPB2 and TEF-α) dataset was compiled in which sequences from five specimens of Abrothallus were aligned with sequences retrieved from GenBank covering main groups of the phylum Ascomycota. As a starting point, we chose the dataset compiled by Schoch et al. (2009b), which is currently the largest dataset available for Ascomycota. 58 ingroup taxa (including five Abrothallus taxa) were used together with four outgroup taxa representing Basidiomycota (Supplementary Material, Table S1). For the second analysis, a dataset of five loci (nuSSU, nuLSU, RPB1, RPB2 and TEF-α) was built using Abrothallus sequences aligned with representatives of the best-known groups of Dothideomycetes following Schoch et al. (2009a). 126 ingroup taxa (including five species of Abrothallus) as well as six outgroup taxa (Arthoniomycetes) were used in this analysis (Supplementary Material, Table S2). For both datasets, we only used specimens with the highest number of available markers. For the third analysis, a seven-locus dataset (nuSSU, nuLSU, mtSSU, RPB1, RPB2, TEF-α and ITS) was used to reconstruct phylogenetic relationships within the genus. In total, 14 specimens of Abrothallus were used representing most of the lichen families on which the genus has been reported (Accession numbers are available in Table 1).
All alignments were carried out using Muscle v3.6 (Edgar 2004) and were manually optimized using Bioedit v.7.0.9 (Hall 1999). Gblocks 0.91b (Castresana 2000) was used to remove ambiguously aligned regions and large gaps. To select the best nucleotide substitution model for subsequent analyses, we used jModelTest (Posada 2008). The models selected for each genome region and dataset according to the Akaike Information Criterion (AIC, Akaike 1974) are provided in Supplementary Material, Table S3.
The maximum likelihood (ML) analysis was implemented in RAxML v7.3.2 (Randomized Accelerated Maximum Likelihood) (Stamatakis 2006) on the Cipres Science Gateway v3.3 webportal (http://www.phylo.org/index.php/portal/) (Stamatakis et al. 2008). ML searches were performed using the GTRGAMMA model; for each gene a different partition was set; for the protein coding markers (RPB1, RPB2, TEF-α), each position of the codon was set in a different partition. For the three datasets, 12, 11 and 13 partitions were used respectively. Bootstrap support was obtained through the rapid bootstrapping algorithm in 1, 000 replications. Bayesian analyses were those implemented in MrBayes v3.1.2 (Ronquist and Huelsenbeck 2003). Substitution models used for each partition and analysis are given in Supplementary Material, Table S3. Settings included two parallel runs with eight-chain runs over 8M generations. Sampling was performed after every 200th step; the first 10 % of saved data was discarded as ‘burnin’. Convergence of the parameters was established using Tracer v.1.4 (Rambaut and Drummond 2007). Phylogenetic trees were visualized in FigTree v. 1.3.1 (http://tree.bio.ed.ac.uk/) and Adobe Illustrator CS2® was used for artwork.
Results
Our six-locus based analysis of Abrothallus sequences within the framework of Ascomycota provided the first insight into the phylogenetic adscription of the genus (Supplementary Material, Fig. S1). The recovered relationships among higher taxa within Ascomycetes were similar to those reported by Schoch et al. (2009a, b). The most striking difference was the position of Eurotiomycetes, which rather than as a sister group to Lecanoromycetes + Lichinomycetes as in Schoch et al. (2009a, b) appeared basal to several classes including Dothideomycetes, Sordariomycetes, Leotiomycetes, Lecanoromycetes and Arthoniomycetes. These relationships were, however, not statistically supported in either study. The genus Abrothallus was included within Dothideomycetes with high statistical support (both BP and PP). In this first analysis, Abrothallus appeared as a sister group to Botryosphaeria, but with low statistical support.
Subsequent analyses provided further information on the systematic position of the genus. General relationships between the families and orders of Dothideomycetes were similar to those recovered by Schoch et al. (2009a). Only slight differences were detected in the relationships among some groups. For instance, the positions of Botryosphaeriaceae and Patellariales differed but, as before, these relationships were not statistically supported in either case. Our data assigned Abrothallus to a sister group of the order Jahnulales (Fig. 2) and this relationship was well-supported by PP and BP. In turn, Jahnulales and Abrothallus appeared as sister groups to Patellariales. Many authors have included Abrothallus in this latter order (Lindau 1897; Vouaux 1913; von Keissler 1929; Kutorga and Hawksworth 1997). The enigmatic taxon Farlowiella carmichaeliana (Berk.) Sacc., recovered in previous studies as basal to Jahnulales (Schoch et al. 2006), appeared as a well-supported (BP and PP) sister group to the clade made up of Jahnulales, Abrothallus and Patellariales.
Six-locus phylogeny (50 % majority rule consensus tree) depicting phylogenetic relationships among orders and families within Dothideomycetes including Abrothallus species. Branches in bold indicate PP ≥ 95 % and ML bootstrap values ≥ 75 %. Asterisks represent branches supported only in one of two analyses
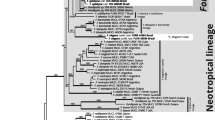
In our seven-marker based phylogenetic reconstruction including 14 specimens of the genus, relationships among the 11 putative species showed an interesting pattern (Fig. 3). First, we observed a clear division of species based on host taxonomy. Two well-supported clades (both by PP and BP) were recovered. The first of these clades included species living on lichens of the order Peltigerales: A. secedens Wedin & R. Sant. (Wedin 1994) on Pseudocyphellaria sp., and undescribed species growing on Nephroma species (Suija et al., in prep.). As sister to this clade appeared specimen NY01041884 whose characters matched those described for A. cladoniae R. Sant. & D. Hawksw. (Hawksworth 1990), i.e. small ascomata (135–(184.5)–270 μm, n = 10) with small ascospores 6–(7.4)–8 × 2.5–(3.1)–3.5 μm (n = 19) which form part-spores already within the ascus.
Seven-locus phylogeny (50 % majority rule consensus tree, midpoint rooting) depicting phylogenetic relationships among 12 Abrothallus species (14 specimens). Branches in bold indicate PP ≥ 95 % and ML bootstrap values ≥ 75 %. Hosts are provided within brackets and host families are shown on the right of the figure
The second large clade included specimens growing on members of the lichen-forming family Parmeliaceae and on the genus Ramalina. The characters of specimen TU45449 (host R. fraxinea) and SPO 304 (host R. cf. protecta) matched those described for A. suecicus (Kirschst.) Nordin (Nordin 1964) including the presence of 4-celled ascospores (Fig. 1j). In turn this clade, whose synapomorphic characters are host of Parmeliaceae and the presence of 2-celled ascospores, is subdivided into two groups. The first of these groups includes two specimens growing on Usnea species. The characters of both specimens match with description of A. usneae var. usneae Rabenh. (Etayo and Osorio 2004; Etayo and van den Boom 2006) i.e. the ascospores are 8.4–(10.3)–11.7 × 3.9–(4.5)–5.0 μm (n = 20; TU45810) and (8–(9.7)–12 × 3–(4.7)–6.0 μm (n = 20; SPO306) in size, and the hymenium contains violet granules. The second clade includes specimens growing on several hosts (Parmelia saxatilis, P. sulcata, Parmelina tiliacea, Parmelinopsis horrescens, Pleurosticta acetabulum, and Parmotrema sp.). Molecular differences among these specimens were small as indicated by the short branches and low statistical support for relationships among them. However, differences were indeed detected in morphological-anatomical characters (size of the ascomata and ascospores, presence/absence of greenish pruina, somatic hyphae reactive to Lugol’s, epihymenial pigments reactive to K, and conidiospore size).
Discussion
Dothideomycetes is the largest class within Ascomycota comprising more than 19, 000 species in 105 families, including saprotrophic, plant pathogenic, fungicolous, and symbiotic fungi (Kirk et al. 2008; Hyde et al. 2013). While in the traditional sense, the class was reserved for taxa with closed (perithecioid) ascolocular fruitbodies and I– bitunicate asci, in the era of molecular taxonomy, the concept of this class has been expanded (e.g. Hyde et al. 2013; Schoch et al. 2006, 2009a). Today, the Dothideomycetes include a small proportion of fungi bearing other types of ascomata – apothecia, hysterothecia, thyriothecia or cleistothecia (e.g. Kirk et al. 2008; Schoch et al. 2009a; Hyde et al. 2013) indicating convergency in fruiting body evolution (e.g. Berbee and Taylor 1992; Hibbett et al. 2007; Lumbsch and Huhndorf 2007; Schoch et al. 2006, 2009a; Zhuang and Liu 2012). Our multi-locus analysis revealed that the obligately lichenicolous genus Abrothallus, whose phylogenetic position has been much debated (e.g. Nannfeldt 1932; Bellemère et al. 1986), is one of the few genera with apothecioid ascomata within the Dothideomycetes. ‘Discothecium’ is a special term which was introduced to designate such ascomata which mimic apothecia but contain bitunicate asci (Korf 1973).
The ontogeny of the ascomata in Abrothallus is ascolocular (Schaechtelin and Werner 1927) which together with having ‘jack-in-the-box’-type asci supports the inclusion of the genus into Dothideomycetes. Moreover, the genus clusters within the subclass Pleosporomycetidae, which is distinguished from the subclass Dothideomycetidae according to the presence of pseudoparaphyses in a hamathecium (Lumbsch and Lindemuth 2001; Schoch 2006, 2009a). To date, the nature of interascal filaments in the hamathecium of Abrothallus remains unclear. They have often been described as paraphyses (e.g. de Notaris 1846; Kotte 1909; Diederich 2004; Suija 2006), but in a few descriptions they have been designated paraphysoids (Nordin 1964; Hawksworth 1983; Hafellner 1994; Wedin 1994) or have not been named at all (Hafellner et al. 2008). Therefore the development of the hamathecium in this genus requires further investigation.
Our results clearly rule out prior hypotheses suggesting an inclusion of Abrothallus within the order Arthoniales (= Arthoniomycetes) or Patellariales (e.g. Rehm 1896; Bellemère et al. 1986). In an early attempt to resolve the phylogenetic relationships of Abrothallus, the nuSSU marker pointed to an unsupported relationship with Dothideales. However, in the combined analysis of nuLSU and nuSSU, in which unfortunately representatives of Dothideomycetes were not included, an ambiguous relationship was observed. Thus, Abrothallus appeared either basal to Arthoniales plus Sordariomycetes or basal with respect to a large group of Ascomycetes (Granberg 2001). Our results show that within Dothideomycetes, the genus is sister to the order Jahnulales, which comprises mainly aquatic (both freshwater and marine) saprotrophic ascomycetes (Pang et al. 2002). Jahnulales are characterized by having stalked or sessile dimorphic ascomata, asymmetric 2-celled hyaline to brown ascospores filled with lipid guttules and with gelatinous appendages or sheaths (Pang et al. 2002; Shearer et al. 2009). Judging from the descriptions and illustrations by Pang et al. (2002), several species of Jahnulales have ascospores which by their external appearance resemble ascospores of Abrothallus. In the recent revision of the class Dothideomyctes by Hyde et al. (2013), the Jahnulales were related with the newly decribed orders Dyfrolomycetales, Strigulales, and Acrospermales. None of the orders share phenotypical and ecological characters similar to Abrothallus. Therefore, given its different anatomical, morphological and ecological characters, and that the genus Abrothallus constitutes a distinct phylogenetic lineage, our proposal of a new monotypic order Abrothallales and family Abrothallaceae seems justified.
Taxonomy
Abrothallales Pérez-Ortega & Suija ord. nov
MycoBank no. MB806050
Type genus: Abrothallus De Not., Mem Reale Accad Sci Torino ser. 2 10:351–355 (1845)
Diagnosis: The monotypic order which contains lichenicolous species with apothecioid ascomata belongs to the Dothideomycetes subclass Pleosporomycetidae based on the multi-locus analysis. The inclusion to the Dothideomycetes is supported by the ascolocular type of ascoma ontogeny (Schaechtelin and Werner 1927) and by the presence of bitunicate asci. The new order is sister to the Jahnulales, which comprises mainly aquatic species. Differences between the new order and Jahnulales rely on the different ascoma type and ecologically by different nutrition mode and substrate type.
Description: Mycelium immersed in the host thallus, somatic hyphae either I+ violet or I–. Ascomata apothecioid, protruding through the host cortex, sessile or partly immersed, spherical to flattened, often with a greenish or yellowish pruina (Fig. 1a, b, d). Excipulum much reduced, composed of short, dichotomously ramified hyphae. Epihymenium brown- or red-granulose (Fig. 1c); granules usually dissolve in K. Hypothecium light to dark brown, consisting of oblong cells covered with a brown pigment. Hamathecium composed of thick-walled, unequally dichotomously branched and anastomosed, septate interascal filaments (possibly pseudoparaphyses sensu Kirk et al. 2008), sometimes slightly swollen at the apex (Fig. 1e); hymenial gel I–. Asci bitunicate, functionally fissitunicate, broadly to narrowly clavate, I–, comprised of four to eight ascospores (Fig. 1f, g); in some species these break into part-spores (Fig. 1k) inside the ascus. Ascospores initially hyaline, later brown, often verruculose, one-, two- or three-septate, asymmetric in shape (Fig. 1i, j). Anamorph common, pycnidial, black, immersed or semi-immersed in host thallus, with a small ostiole (Fig. 1l, m), Vouauxiomyces-type; wall of the pycnidium textura angularis, composed of thick-walled isodiametric cells. Conidiophores missing. Conidiogenous cells percurrently proliferating, ampulliform to lageniform (Fig. 1h), lining the cavity of the pycnidium, hyaline, smooth-walled. Conidia holoblastic, clavate to obpyriform, hyaline (Fig. 1n, o), smooth to very slightly echinulate, in muscilage.
Distribution: cosmopolitan
Ecology: obligately lichenicolous; hosts belong to a variety of fruticose and foliose macrolichens.
The new order Abrothallales includes the single family Abrothallaceae.
Abrothallaceae Pérez-Ortega & Suija fam. nov.
MycoBank no. MB806051
Type genus: Abrothallus De Not., Mem Reale Accad Sci Torino ser. 2 10:351–355 (1845)
Characters as in Abrothallales.
References
Akaike H (1974) A new look at the statistical model identification. IEEE Trans Autom Control 19(6):716–723
Arnold AE, Miadlikowska J, Higgins KL et al (2009) A phylogenetic estimation of trophic transition networks for ascomycetous fungi: are lichens cradles of symbiotrophic fungal diversification? Syst Biol 58:283–297
Bellemère A, Malherbe MC, Chacun H, Hafellner J (1986) Étude ultrastructurale des asques et des ascospores chez les espèces lichénicoles non lichénisées Abrothallus bertianus de Not. et A. parmeliarum (Sommerf.) Nyl. Cryptog Mycolog 7:47–85
Berbee ML, Taylor JW (1992) Two ascomycete classes based on fruiting-body characters and ribosomal DNA sequence. Mol Biol Evol 9(2):278–284
Castresana J (2000) Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol 17:540–552
Clauzade G, Diederich P, Roux C (1989) Nelikenigintaj fungoj likenlogaj. Ilustrita determinlibro. Bull Soc Linn Provence Numéro Spécial 1:1–42
De Notaris G (1845) Abrothallus novum lichenum genus. Mem Reale Accad Sci Torino ser. 2, 10:351–355
De Notaris G (1846) Frammenti lichenografici di un lavoro inedito. G Bot Ital 1:174–224
Diederich P (2004) Abrothallus. In: Nash TH III, Ryan BD, Diederich P, Gries C, Bungartz F (eds) Lichen Flora of the Greater Sonoran Desert Region, vol 2, Lichens Unlimited. Arizona State University, Tempe, pp 626–630
Don R, Cox P, Wainwright B, Baker K, Mattick J (1991) ‘Touchdown’ PCR to circumvent spurious priming during gene amplification. Nucleic Acids Res 19(14):4008
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32(5):1792–1797
Etayo J, Osorio HS (2004) Algunos hongos liquenícolas de Sudamérica, especialmente del Uruguay. Comun Bot Mus Nac Hist Nat Antrop 6(129):1–19
Etayo J, van den Boom PG (2006) Some lichenicolous fungi from Guatemala, with the description of a new species. Herzogia 19:191–197
Fink B (1935) The Lichen Flora of the United States. Completed for Publication by Joyce Hedrick. University of Michigan Press, Ann Arbor
Gardes M, Bruns TD (1993) ITS primers with enhanced specificity for basidiomycetes—application to the identification of mycorrhizae and rusts. Mol Ecol 2:113–118
Gargas A, Taylor JW (1992) Polymerase chain reaction (PCR) primers for amplifying and sequencing 18S rDNA from lichenized fungi. Mycologia 84:589–592
Granberg Å (2001) The phylogenetic relationships of Abrothallus, a genus of lichenicolous fungi. Degree thesis, Umeå University
Hafellner J (1994) Beiträge zu einem Prodromus der lichenicolen Pilze Österreichs und angrenzender Gebiete. I. Einige neue oder seltene Arten. Herzogia 10:1–28
Hafellner J, Herzog G, Mayrhofer H (2008) Zur Diversität von lichenisierten und lichenicolen Pilzen in den Ennstaler Alpen (Österreich: Steiermark, Oberösterreich). Mitt Naturwiss Ver Steiermark 137:131–204
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Hawksworth DL (1983) A key to the lichen-forming, parasitic, parasymbiotic and saprophytic fungi occurring on lichens in the British Isles. Lichenologist 15:1–44
Hawksworth DL (1990) Notes on British lichenicolous fungi: VI. Notes Roy Bot Gard Edinburgh 46(3):391–403
Hibbett DS, Binder M, Bischoff JF et al (2007) A higher-level phylogenetic classification of the Fungi. Mycol Res 111(5):509–547
Hofstetter V, Miadlikowska J, Kauff F, Lutzoni F (2007) Phylogenetic comparison of protein-coding versus ribosomal RNA-coding sequence data: a case study of the Lecanoromycetes (Ascomycota). Mol Phylogenet Evol 44:412–426
Hyde KD, Jones EBG, Liu J-K, Ariyawansa H et al (2013) Families of dothideomycetes. Fungal Divers. doi:10.1007/s13225-013-0263-4
Jatta A (1911) Lichenes (end) Flora Italica Cryptogama, Firenze Pars III. Fasc 4–6:461–958
Kauff F, Lutzoni F (2002) Phylogeny of the Gyalectales and Ostropales (Ascomycota, Fungi): among and within order relationships based on nuclear ribosomal RNA small and large subunits. Mol Phylogenet Evol 25:138–156
Kirk PM, Cannon PF, Stalpers JA, Minter DW (eds) (2008) Dictionary of fungi, 10th edn. Commonwealth Scientific and Industrial Research Organisation, UK
Körber GW (1855) Systema Lichenum Germaniae. Trewendt & Granier, Breslau, Germany
Korf RP (1973) Discomycetes and Tuberales. In: Ainsworth GC, Sparrow FK, Sussman AS (eds) The fungi: an advanced treatise, vol 4A. Academic, London, pp 249–319
Kotte I (1909) Einige neue Fälle von Nebensymbiose (Parasymbiose). Zentralbl Bakteriol Parasitenkd II 24:74–93
Kutorga E, Hawksworth DL (1997) A reassessment of the genera referred to the family Patellariaceae (Ascomycota). Syst Ascomycetum 15(1–2):1–110
Lawrey JD, Diederich P (2003) Lichenicolous fungi: interactions, evolution, and biodiversity. Bryologist 106(1):81–120
Lawrey JD, Diederich P, Nelsen MP, Freebury C, Van Den Broeck D, Sikaroodi M, Ertz D (2012) Phylogenetic placement of lichenicolous Phoma species in the Phaeosphaeriaceae (Pleosporales, Dothideomycetes). Fungal Divers 55:195–213
Lindau G (1897) Pezizineae. In: Engler A, Prantl K (eds) Die Natürlichen Pflanzenfamilien I. W. Engelmann, Leipzig, pp 178–242
Lindsay WL (1857) Monograph of the genus Abrothallus (De Notaris and Tulasne emend.). Q J Microsc Sci 5:27–63
Liu YJ, Whelen S, Hall BD (1999) Phylogenetic relationships among ascomycetes: evidence from an RNA polymerase II subunit. Mol Biol Evol 16:1799–1808
Lumbsch HT, Huhndorf SM (2007) Whatever happened to the pyrenomycetes and loculoascomycetes?. Mycol Res 111:1064–1074
Lumbsch HT, Huhndorf SM (2010) Myconet volume 14. Fieldiana 1:1–64
Lumbsch HT, Lindemuth R (2001) Major lineages of Dothideomycetes (Ascomycota) inferred from SSU and LSU rDNA sequences. Mycol Res 105:901–908
Matheny PB, Liu YJ, Ammirati JF, Hall BD (2002) Using RPB1 sequences to improve phylogenetic inference among mushrooms (Inocybe, Agaricales). Am J Bot 89:688–698
Millanes AM, Diederich P, Ekman S, Wedin M (2011) Phylogeny and character evolution in the jelly fungi (Tremellomycetes, Basidiomycota, Fungi). Mol Phylogenet Evol 61(1):12–28
Moncalvo JM, Rehner SA, Vilgalys R (1993) Systematics of Lyophyllum section Difformia based on evidence from culture studies and ribosomal DNA sequences. Mycologia 85:788–794
Montagne C (1851) Morphologischer Grundriss der Familie der Flechten. Graeger, Halle, Germany
Montagne C (1856) Cryptogamia Guyanensis seu plantarum cellularium in Guyana gallica annis 1835–1849 a cl. Leprieur collectarum enumeratio universalis. suite. Ann Sci Nat 16:47–81
Nannfeldt JA (1932) Studien über die Morphologie und Systematik der Nicht-Lichenisieren inoperculaten Discomycetes. Nova Acta Regiae Soc Sci Ups 8(2):1–368
Nordin I (1964) Abrothallus suecicus, a common lichenicolous fungus. Sven Bot Tidskr 58(1):225–232
Pang K-L, Abdel-Wahab MA, Sivichai S, El-Sharaouney HM, Jones EBG (2002) Jahnulales, (Dothideomycetes, Ascomycota): a new order of lignicolous freshwater ascomycetes. Mycol Res 106(9):1031–1042
Pérez-Ortega S, Suija A, de los Ríos A (2011) The connection between Abrothallus and its anamorph state Vouauxiomyces established by Denaturing Gradient Gel Electrophoresis (DGGE). Lichenologist 43(3):277–279
Peršoh D, Rambold G (2002) Phacopsis—a lichenicolous genus of the family Parmeliaceae. Mycol Prog 1(1):43–55
Peršoh D, Rambold G (2012) Lichen-associated fungi of the Letharietum vulpinae. Mycol Prog 11(3):753–760
Posada D (2008) jModelTest: phylogenetic model averaging. Mol Biol Evol 25(7):1253–1256
Rambaut A, Drummond AJ (2007) Tracer v1.4. Available from http://beast.bio.ed.ac.uk/Tracer
Rehm H (1896) Ascomyceten: Hysteriaceen und Discomyceten. In: Rabenhorst’s Kryptogamen-Flora von Deutschland, Oesterreich und der Schweiz, 1(3):209–336
Rehner S, Samuels GJ (1994) Taxonomy and phylogeny of Gliocladium analyzed from nuclear large subunits ribosomal DNA sequences. Mycol Res 98:625–634
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Saccardo PA (1889) Sylloge fungorum. Published by the author, Pavia
Santesson R, Moberg R, Nordin A, Tønsberg T, Vitikainen O (2004) Lichen-forming and lichenicolous fungi of Fennoscandia. Museum of Evolution, Uppsala University, Uppsala
Schaechtelin J, Werner RG (1927) Développement et biologie de l’Abrothallus parmeliarum Smft. Bull Soc Mycol France 42:233–243
Schoch CL, Shoemaker RA, Seifert KA, Hambleton S, Spatafora JW, Crous PW (2006) A multigene phylogeny of the Dothideomycetes using four nuclear loci. Mycologia 98(6):1041–1052
Schoch CL, Crous PW, Groenewald JZ et al (2009a) A class-wide phylogenetic assessment of Dothideomycetes. Stud Mycol 64:1–15
Schoch CL, Sung GH, López-Giráldez F et al (2009b) The Ascomycota tree of life: a phylum-wide phylogeny clarifies the origin and evolution of fundamental reproductive and ecological traits. Syst Biol 58(2):224–239
Shearer CA, Raja HA, Miller AN et al (2009) The molecular phylogeny of freshwater Dothideomycetes. Stud Mycol 64:145–153
Sikaroodi M, Lawrey JD, Hawksworth DL, DePriest PT (2001) The phylogenetic position of selected lichenicolous fungi: Hobsonia, Illosporium, and Marchandiomyces. Mycol Res 105(4):453–460
Stamatakis A (2006) RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22(21):2688
Stamatakis A, Hoover P, Rougemont J (2008) A rapid bootstrap algorithm for the RAxML web servers. Syst Biol 57(5):758–771
Stiller J, Hall BD (1997) The origin of red algae: implications for plastid evolution. PNAS 94:4520–4525
Suija A (2006) Variation of morphological characters in the lichenicolous ascomycete genus Abrothallus. Ann Bot Fenn 43:193–204
Tulasne L-R (1852) Mémoire pour servir à l’histoire organographique et physiologique des lichens. Ann Sci Nat 17:5–128
U’Ren JM, Lutzoni F, Miadlikowska J, Laetsch A, Arnold AE (2012) Host and geographic structure of endophytic and endolichenic fungi at a continental scale. Am J Bot 99(5):898–914
Vilgalys R, Hester M (1990) Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J Bacteriol 172:4239–4246
von Keissler K (1929) Die Flechtenparasiten Deutschlands, Österreichs und der Schweiz mit Berücksichtigung der übrigen Länder Europas sowie der angrenzenden Meeresgebiete. Leipzig, Germany
Vouaux L (1913) Synopsis des champignons parasites de Lichens (Suite). Bull Soc Mycol France 29:33–128, 399–444, 447–494
Wedin M (1994) New and noteworthy lichenicolous fungi from southernmost South America. Lichenologist 26(3):301–310
Werth S, Millanes AM, Wedin M, Scheidegger C (2013) Lichenicolous fungi show population subdivision by host species but do not share population history with their hosts. Fungal Biol 117(1):71–84
White TJ, Bruns TD, Lee SB, Taylor JW (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols—a guide to methods and applications. Academic, San Diego, pp 315–322
Zhuang W-Y, Liu C-Y (2012) What an rRNA secondary structure tells about phylogeny of fungi in Ascomycota with emphasis on evolution of major types of ascus. PLoS ONE 7(10):e47546
Zoller S, Scheidegger C, Sperisen C (1999) PCR primers for the amplification of mitochondrial small subunit ribosomal DNA of lichen-forming ascomycetes. Lichenologist 31(5):511–516
Acknowledgments
We thank Tiiu Tõrra, Ede Leppik and Martin Kukwa for collecting the specimens and the curator of NY for granting specimens loan. AS would like to thank Kadri Vaidla, Kadi Jairus, Peeter Laas for performing the lab work. We would like to thank the staff of the microscopy facility of the Museo Nacional de Ciencias Naturales (CSIC, Madrid) for their technical assistance, and Ana Burton for revising the English. SPO and AdR were supported by the grant CTM2012-38222-C02-02 from the Spanish Ministry of Economy and Competitiveness. We thank Maria José Malo (Madrid) for her technical assistance during lab work. The work by AS was supported by a grant awarded by the Estonian Science Foundation no. GP1LM7321, target-financing project SF0180012s09 and the European Regional Development Fund (Centre of Excellence FIBIR).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Fig. S1
Six-locus phylogeny (50 % majority rule consensus tree) depicting phylogenetic relationships among classes of Ascomycota including Abrothallus species. Branches in bold indicate PP ≥ 95 % and ML bootstrap values ≥ 75 %. (JPEG 89 kb)
Table S1
Accession numbers corresponding to taxa from different classes of Ascomycota used in the phylogenetic analyses. (DOCX 23 kb)
Table S2
Accession numbers corresponding to taxa from Dothideomycetes and Arthoniomycetes used in the phylogenetic analyses. (DOC 217 kb)
Table S3
Nucleotide substitution models used for each partition in the three different analyses run in this study. (DOCX 10.7 KB)
Rights and permissions
About this article
Cite this article
Pérez-Ortega, S., Suija, A., Crespo, A. et al. Lichenicolous fungi of the genus Abrothallus (Dothideomycetes: Abrothallales ordo nov.) are sister to the predominantly aquatic Janhulales. Fungal Diversity 64, 295–304 (2014). https://doi.org/10.1007/s13225-013-0269-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13225-013-0269-y