Abstract
NAC transcription factors are involved in many biological processes via regulation of downstream target gene expression and play essential roles in regulation of plant growth and improving plant tolerance to abiotic stress. NAC transcription factors have been studied in various species, but little information is available regarding these factors in sorghum. Genome-wide investigation of potentially abiotic stress related sorghum NAC-type genes was performed. A total of 145 non-redundant NAC genes (SbNAC1 –SbNAC145) were identified in the sorghum genome. These genes were distributed unevenly across the 10 chromosomes, and were divided into 16 groups based on sequence similarity. Gene structure analysis indicated that most SbNAC genes contained three exons and two introns, and had ten putative conserved motifs. Phylogenetic analysis indicated that the SbNAC genes with similar motif distributions were clustered into the same branch. Seven SbNAC genes, which were grouped into the stress-related subgroup, were isolated and have been confirmed to have transcriptional activity in yeast. SbNAC genes showed differential expression patterns over time in response to dehydration, salinity, cold, and phytohormone abscisic acid stress treatments, thus suggesting essential roles in plant responses to abiotic stress. In the germination stage, SbNAC56 overexpression transgenic lines exhibited significantly enhanced hypersensitivities to ABA, NaCl and d-Mannitol. This may infer that SbNAC56 may play essential roles in plants response to abiotic stresses in ABA dependent signaling pathway. Here, we present a comprehensive overview of the SbNAC genes and provide a foundation for future functional research regarding their biological roles in sorghum stress tolerance, even in the regulation of plant growth.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the natural environment, plants are constantly exposed to a variety of abiotic and biotic stresses, such as salinity, drought, cold, wounding, and pathogen infection, which restrict plant growth and development (Ji et al. 2014). To cope with these adverse conditions, plants have evolved multiple adaptive strategies at the physiological, biochemical, and molecular levels (Broun 2004). Accumulating evidence indicates that numerous transcription factors (TFs), such as NAC (NAM, ATAF1/2 and CUC2), MYB, AP2/EREB, and bZIP, play essential roles in stress response via binding to the cis-acting elements in the promoter regions of target genes (Guo et al. 2015).
NAC proteins comprise one of the largest families of plant-specific TFs and have been considered the key regulators in plant responses to abiotic and biotic stresses (Zhao et al. 2016). NAC proteins share a common highly conserved DNA-binding domain in the N-terminal region, which consists of five subdomains (A–E). In contrast, a quite divergent sequence in the C-terminus was considered to be the transactivation domain (Ooka et al. 2003).
Increased evidence indicated NAC TFs play essential roles in plant growth and development. Moreover, functional studies of NAC proteins have demonstrated that they are involved in responses to drought, salinity, and cold stresses. For example, ANAC019, ANAC055, and ANAC072 expression in Arabidopsis was induced by salinity, drought, and the phytohormone abscisic acid (ABA), respectively. Overexpression of these three NAC genes markedly enhanced the drought stress tolerance in transgenic Arabidopsis (Tran et al. 2004). Furthermore, ANAC096 has been reported to be involved in the drought stress response through binding to ABA-responsive elements in the promoter regions of target genes (Xu et al. 2013). In wheat, the expression of TaNAC67 was induced by ABA, as well as by salt and drought stresses, which suggested that it may play important roles in the abiotic stress response in the ABA signaling pathway (Mao et al. 2014). Overexpression of wheat TaNAC2, TaNAC2a, and TaNAC69 significantly improved abiotic stress tolerance in transgenic plants (Mao et al. 2012; Xue et al. 2006, 2011). Under conditions of extreme temperature, ABA, gibberellic acid, and polyethylene glycol (PEG) stress treatments, 6 of 55 selected NAC genes in Chinese cabbage were significantly induced (Liu et al. 2014). Solyc07g063410, Solyc04g009440, Solyc01g094490, and Solyc10g047060 in tomato were significantly upregulated by low temperature, PEG, and high salinity treatments (Su et al. 2015). In addition, several NAC members have been identified in chickpea. Overexpression of CarNAC3, CarNAC6 and CarNAC4 in chickpea significantly enhanced salt and drought tolerance in transgenic plants (Movahedi et al. 2015; Yu et al. 2016). Drought, methyl jasmonic acid (MeJA), and salicylic acid (SA) stress treatments increased the transcription levels of 13 Miscanthus NAC genes. MlNAC5 and MlNAC9 conferred enhanced drought stress tolerance and ABA hypersensitivity in transgenic Arabidopsis (Yang et al. 2015; Zhao et al. 2016). Recently, a stress-responsive NAC gene, ZmNAC55, was identified in maize and reportedly functions in drought stress tolerance (Mao et al. 2016). Salinity, cold, wounding, MeJA, and ABA induced the expression of PpNAC2 and PpNAC3, which clustered with Arabidopsis ATAF1 and ATAF2, respectively. A miniature inverted repeat transposable element (MITE) insertion in the ZmNAC111 promoter negatively regulated drought stress tolerance in maize (Pascual et al. 2015; Mao et al. 2015).
Sorghum originated in northern Africa, now it is a tropical crop with a good ecological plasticity, being grown worldwide in the semiarid areas of Africa, India, Southeastern Asia and Australia (Endre et al. 2017; Kebede et al. 2001; Oprea et al. 2015). Its tolerance to drought and extreme temperatures make it is a typical drought-resistant model crop, which is important for production and research. Sorghum expresses a series of stress-related genes under various environmental stresses. To date, little information regarding NAC genes in sorghum has been reported, although NAC TFs are involved in regulating both abiotic and biotic stress responses, which pose challenges to the characterization of NAC family proteins in sorghum. In the present study, 145 SbNAC members were identified in sorghum and classified according to their amino acid sequence identity. Seven of them were isolated from sorghum XGL-1. Phylogenetic analysis, gene structure, chromosomal locations, transactivation activity, prokaryotic expression, expression profiles, and seeds germination were examined. This study will provide a basis for future clarification of their functions in sorghum stress responses.
Materials and methods
Plant material and growth conditions
The inbred sorghum line XGL-1 was provided by the Institute of Crop Sciences, Chinese Academy of Agricultural Sciences. Compared to other tested sorghum genotypes, sorghum XGL-1 was considered as an outstanding drought stress resistant genotype according to our prior trial information in Xinjiang province. XGL-1 provided a relative ideal genetic background to clarify the molecular mechanisms of drought stress resistance.
Seeds were rinsed several times. Plants were grown at 28 °C with 16 h of light, and 22 °C with 8 h of darkness in a greenhouse.
Identification and bioinformatics analysis
Two approaches were used to identify the all putative NAC proteins in sorghum. First, the amino acid sequences of SbNAC proteins were downloaded from the Plant Transcription Factor Database version 4 (PlantTFDB; http://planttfdb.cbi.pku.edu.cn/). Then, “NAC” was used as a query to search against the Joint Genome Institute (JGI) annotation database (https://phytozome.jgi.doe.gov/).
Multiple alignments were performed with Clustal X (ver. 1.83) (Thompson et al. 1997). A phylogenetic tree was constructed by the neighbor-joining method with 1000 replicates on each node using MEGA 4.0 (Saitou 1987; Tamura et al. 2007). The structures of SbNAC genes were analyzed by Gene Structure Display Server 2.0 (http://gsds1.cbi.pku.edu.cn/). MEME online software (version 4.11; http://meme-suite.org/) was used to analyze the amino acid motifs that are represented by colored boxes.
Theoretical molecular weight (Mw) was calculated with online software (http://web.expasy.org/compute_pi/). TMHMM (http://www.cbs.dtu.dk/services/TMHMM/) was used to predict the transmembrane (TM) domains. Chromosome location images were generated by MapChart software (version 2.2). Potential SbNAC gene pairs were identified with multiple sequence alignment.
Abiotic stress treatments
Seeds of the inbred sorghum line XGL-1 were grown in pots filled with vermiculite. For cold stress treatment, sorghum seedlings at the 3-leaf stage were transferred to a low-temperature (4 °C) growth chamber. For salt, drought, and ABA stress treatments, sorghum seedlings were transferred to Hoagland solution supplemented with 200 mM NaCl, 20% PEG, and 100 mM ABA, respectively. Shoots were separately harvested at 0, 1, 3, 6, and 24 h. Plant samples were frozen in liquid nitrogen and stored at −80 °C until analysis.
RNA extraction and quantitative real-time PCR analyses
Total RNA was extracted using TRIzol™ reagent (Invitrogen, Carlsbad, CA) according to the manufacturer’s protocol. The first-strand cDNA was synthesized using a Thermo Scientific RevertAid™ First Strand cDNA Synthesis Kit (#K1622; Waltham, MA). qRT-PCR was performed using SYBR® Ex Taq™ (Takara, Dalian, China) and carried out with a LightCycler® 96 SW 1.1 PCR Detection System (Roche Diagnostics, Penzberg, Germany). The 18SrRNA gene was used as an internal control. The relative expression level for each gene was calculated with the 2−ΔΔCT method. These experiments were repeated with at least three biological replicates.
Transactivation activity analysis
Transactivation activity analysis was performed in the yeast strain AH109 (Clontech, Palo Alto, CA). The full-length cDNAs were amplified by PCR with forward and reverse primers containing EcoRI and BamHI sites, respectively, and inserted into the corresponding sites of the pGBKT7 vector. Subsequently, the pGBKT7 vector (control) and pGBKT7—SbNAC6/SbNAC26/SbNAC46/SbNAC56 were separately transformed into yeast strain AH109. The transformants were spotted on synthetic dropout (SD) medium SD/-Trp, SD/-Trp/-His, SD/-Trp/-His/10 mM (20 mM/30 mM) 3-AT and SD/-Trp/-His/-Ade for 3 days at 30 °C, and β-galactosidase (X-Gal) activity was measured as described by Jin (Jin et al. 1997).
Prokaryotic expression of SbNACs
To investigate the functions of the SbNAC genes, the full-length cDNAs of SbNAC6, SbNAC26, and SbNAC56 were inserted into the pET40b (+) vector to construct fusion vectors. The empty pET40b (+) vector was used as a negative control. The constructs were transformed into Rosetta (DE3) cells. The fusion proteins could be induced by 0.5 mmol/L IPTG for 8 h at 23 °C. The proteins were identified by SDS-PAGE as described by Li et al. (2012).
Arabidopsis transformation and seeds germination assay
To analyze its biological functions, SbSNAC56-pCambia3301 recombinant vector was transformed into Arabidopsis Col-0 wild-type plants via Agrobacterium tumefaciens strain GV3101 and T3 homozygous were obtained. The WT and SbSNAC56 overexpression transgenic lines were screened on MS medium plates that containing NaCl (150, 175 Mm), d-Mannitol (350, 400 mM), phytohormone ABA (1.5, 2.0 µM). The germination ratio of seeds was calculated. All experiments were repeated at least three times, and 45–50 seeds were used in each comparison.
Results
Identification of NAC genes in sorghum
The SbNAC members in sorghum genome DNA were identified in the PlantTFDB version 4. Initially, 180 putative NAC transcription factor genes were obtained. Then, 46 redundant SbNAC proteins were determined by multiple sequence alignment. Furthermore, “NAC” was used as a key word to search for NAC domain-containing proteins in the U.S. Department of Energy (DOE) JGI database, and 53 putative NAC transcription factor genes were obtained. Finally, 145 non-redundant SbNAC proteins, including seven membrane-bound proteins, were identified and used for further study (Fig. S1).
Specific primers were designed to amplify the full-length cDNAs of seven stress-related SbNAC genes: SbNAC6, SbNAC17, SbNAC26, SbNAC46, SbNAC56, SbNAC58, and SbNAC73 (GenBank accession numbers: KY348374, KY348378, KY348375, KY348376, KY348377, KY348379, and KY348380, respectively), which ranged in size from 873 to 1209 bp and encoded peptides of 290 to 402 amino acids (Table S1).
Phylogenetic, structural, and motif analyses of SbNACs
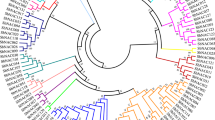
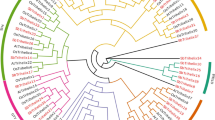
To clarify the phylogenetic relationships among the SbNAC genes, an unrooted phylogenetic tree was constructed. SbNAC members were clustered together and showed similar motif distributions. Most conserved motifs were found within the N-terminal region of the NAC domain, which suggested that these motifs may be essential for the biological function of NAC proteins. In addition, some well-known stress-responsive NAC genes were analyzed in the phylogenetic tree. Based on the results, sorghum NAC genes could be divided into 16 subgroups. According to the phylogenetic relationship, SbNAC6, SbNAC26, and SbNAC56 belonged to the OsNAC3 subfamily, SbNAC58 belonged to the ATAF subfamily, and SbNAC46, SbNAC17, and SbNAC73 were clustered into the NAP subfamily (Fig. 1). And SbNAC6 was defined as the paralogous gene of SbSNAC1 in sorghum (the sequence similarity is 99.38%). Previous studies have confirmed that OsNAC3, ATAF, and NAP are a stress-related subgroup of the NAC superfamily.
Subsequently, we performed gene structure analysis to support the phylogeny reconstruction (Fig. 2a, b). Consistent with the results of phylogenetic analysis, SbANAC genes that clustered in the same subgroup showed similar gene structures, especially with regard to the numbers of exons and introns.
Phylogenetic relationship, gene structure analysis, and motif distributions of SbNAC genes. a A phylogenetic tree was constructed by the neighbor-joining method with 1000 replicates on each node. b Exons and introns are indicated by green rectangles and gray lines, respectively. c Amino acid motifs in the SbNAC proteins (1–10) are represented by colored boxes. The black lines indicate relative protein lengths. (Color figure online)
To further determine the diversification of the NAC family in sorghum, 10 distinct conserved motifs were identified in sorghum NAC proteins (Fig. 2c). The NAC members with a close phylogenetic relationship shared common motif compositions. This gene structural similarity inferred similar biological functions in NAC proteins.
Chromosomal location and gene duplication of SbNACs
A total of 145 SbNAC genes were mapped on sorghum chromosomes (Chr) and showed an uneven distribution. Each of the SbNACs’ name was given according to its physical position from the top to the bottom on the sorghum chromosomes 1 to 10. Chr1 and Chr3 contained the largest number of SbNAC genes (24 and 22 genes, respectively), while Chr8 contained the least (9 genes). Chr4 and Chr5 contained 10 SbNAC genes each (Fig. 3). As shown in Fig. 3, 19 (13%) paralogs have been identified in the sorghum NAC gene family, indicating the evolutionary relationship of these NAC members. Gene duplication was confirmed to play a major role in the expansion of gene families.
Transactivation activity analysis of selected SbNAC genes in yeast. Transformants were streaked on SD/-Trp, SD/-Trp/-His, SD/-Trp/-His+10 mM (20/30 mM 3-AT), and SD/-Trp/-His/-Ade for examination of growth. a β-galactosidase activity assay was performed using X-α-Gal staining. b The β-galactosidase activity was measured using o-nitrophenyl-β-d-galactopyranoside (ONPG) as a substrate. The error bars indicate standard deviation (n = 3)
The prokaryotic expression of SbNAC6, SbNAC26 and SbNAC56. a pET40b-SbNAC6, b pET40b-SbNAC26, c pET40b-SbNAC56. M markers; lane 1 pET-40b vector was non-induced; lane 2 pET-40b induced by IPTG for 8 h; lane 3 SbNACs non-induced; lane 4 SbNACs induced for 8 h at 23 °C; lane 5 SbNACs supernatant after ultrasonication induced for 8 h at 23 °C; lane 6 SbNACs inclusion body induced for 8 h at 23 °C
Performance of SbNAC56 overexpression lines germinated under ABA and osmotic treatments stress. Seeds were germinated on MS medium containing NaCl (150, 175 Mm), d-Mannitol (350, 400 mM) and ABA (1.5, 2.0 μM) for six days. The germination rates were calculated after planting on the plates. Data represent means ± SD of three replicates
SbNACs exhibit transactivation activity
NAC-type proteins are characterized by a conserved N-terminal DNA-binding domain and transcriptional activation or repression region in the C-terminal (Hao et al. 2011; Ooka et al. 2003). Multiple sequence alignment revealed that the putative transcriptional activation region located in the C-terminus of these seven SbNAC proteins, which belong to OsNAC3, ATAF and NAP subfamily, respectively. To determine the SbNAC proteins transactivation activity, four NAC genes were selected randomly and the recombinant plasmids pGBKT7–SbNAC6/SbNAC26/SbNAC46/SbNAC56 were transformed into yeast strain AH109. Different transformants expressing fusion proteins in the yeast strain grew well on SD/-Trp, SD/-Trp/-His, SD/-Trp/-His/10 mM (20/30 mM) 3-AT, and SD/-Trp/-His/-Ade medium. The four NAC proteins showed transactivation activity. In contrast, transformants with the empty pGBKT7 vector (negative control) failed to grow. The results of yeast-based reporter and X-gal activity assays confirmed that SbNAC6, SbNAC26, SbNAC46, and SbNAC56 had transcriptional activation activity in yeast (Fig. 4a).
The β-galactosidase activity was measured using o-nitrophenyl-β-d-galactopyranoside (ONPG) as a substrate. Compared with the negative control (pGBKT7), four SbNAC proteins showed enhanced β-galactopyranoside activity in yeast Y187, among which, SbNAC56 showed the highest activity (Fig. 4b).
Expression of prokaryotic recombinant plasmid
Prokaryotic expression analyses indicated that the fusion proteins expressing pET40b-SbNAC6/SbNAC26/SbNAC56 were induced by IPTG treatment. SDS-PAGE analysis indicated that the Mw of these three SbNAC proteins were approximately 35.09, 31.90, and 34.76 kDa, respectively.
Expression patterns of SbNAC genes under different stress treatments
To explore the sorghum NAC gene functions in abiotic stress responses, the expression patterns of seven selected SbNAC genes, that localized in stress-related subgroups OsNAC3, ATAF and NAP, were analyzed. SbNAC genes were expressed at relatively higher levels under ABA and cold stress treatments (Fig. 5c, d).
As shown in Fig. 5a, the seven SbNAC genes were not significantly induced by control treatment (Hoagland solution). SbNAC17, SbNAC46, SbNAC56, SbNAC58, and SbNAC73 were strongly induced by both PEG and ABA treatments. The transcript level peaked at 6 h, and was then downregulated at 24 h (Fig. 5b, c). Under conditions of cold stress, the expression levels of these selected SbNACs were commonly upregulated, among which, SbNAC17 and SbNAC73 were induced with relative higher expression levels (Fig. 5d). SbNAC17, SbNAC46, and SbNAC73 exhibited a sustained dehydration stress response, and reached the highest level at 24 h (Fig. 5e). The transcripts of SbNACs were all increased in response to salt stress and reached the highest level at 3 h, then decreased at 6 h; among them, SbNAC6, SbNAC26, and SbNAC58 showed the highest transcript levels (Fig. 5f).
SbNAC56 transgenic plants were hypersensitive to ABA and osmotic stresses
Abscisic acid (ABA) is synthesized under drought stress conditions (Shinozaki et al. 2003) and exogenous application of ABA could induce a number of dehydration responsive genes (Zhu 2002). The expression of SbNAC56 was strongly induced by ABA, PEG and NaCl treatments (Fig. 5c). This may infer that SbNAC56 may play essential roles in plants response to osmotic stresses in ABA dependent signaling pathway. To confirm this hypothesis, 35S::SbNAC56 transgenic plants were generated. Under 1.5 and 2.0 μM ABA stress treatment, the germination rates of the WT and 35S::SbNAC56 transgenic lines were 54 and 4%, 48 and 4%, respectively. Whereas no significant differences were detected on ABA-free medium. Similar differences could also be observed under NaCl (150, 175 mM), d-Mannitol (350, 400 mM) treatments(Fig. 7). The significantly increased hypersensitivity to ABA and osmotic stresses indicated that SbNAC56 may work in ABA dependent signaling pathways in response to osmotic stresses.
Discussion
To date, numerous NAC proteins have been identified in multiple species, including 104 in tomato (Xie et al. 2002), 204 in Chinese cabbage (Liu et al. 2014), 79 in grape (Wang et al. 2013), 163 in poplar (Hu et al. 2010), 157 in maize, 151 in rice, and 117 in Arabidopsis (Nuruzzaman et al. 2010). These proteins were shown to play essential roles in diverse aspects of plant growth and development, including modulation of leaf senescence (Guo and Gan 2006), embryogenesis (Souer et al. 1996), lateral root development (He et al. 2005), and secondary wall synthesis (Mitsuda et al. 2005),JUB1 represses GA3ox1 and DWARF4 (DWF4), two key members involved in gibberellin (GA) and brassinosteroid (BR) biosynthesis, and regulated plant growth (Shahnejat-Bushehri et al. 2016, 2017). In Arabidopsis, NST2 together with SND1 and NST1 regulate the deposition of secondary walls in fibers of stems (Rui and Zheng. 2015). MiRNA164-directed cleavage of Arabidopsis NAC1 affects lateral root development. Meanwhile, overexpression of maize ZmNAC1, a homologous gene of Arabidopsis NAC1, positively regulated lateral root development in Arabidopsis (Guo et al. 2005; Jing et al. 2012).
In addition, NAC proteins involved in plant defense responses to abiotic stresses. In this paper, a total of 145 SbNAC genes were identified through genome-wide analysis. Of these, 120 SbNAC genes had at least 1 intron, with the majority containing 2 introns (Fig. 2b). Putative conserved motifs were predicted, and 10 motifs were identified to explore the structural characteristics of the SbNACs. The conserved motifs were located in the N-terminal region of NAC proteins. Based on the results of phylogenetic analysis of SbNACs and some typical stress-related NAC TFs, seven predicted NAC genes were grouped into OsNAC3, NAP, and ATAF subfamilies (Fig. 1), which are reportedly involved in abiotic stress responses. In general, the NAC proteins with close phylogenetic relationships showed similar motif compositions. This will contribute to studies of the biological functions of NAC proteins (Fig. 2a, c).
Chromosomal location analysis indicated that SbNAC members were distributed on all 10 sorghum chromosomes. Some NAC genes often occurred in complexes. For example, SbNAC genes were clustered at the ends of the chromosome arms, including Chr2, Chr4, Chr5, and Chr6 (Fig. 3). Gene duplication may occur after the divergence of different species, and 19 NAC gene duplications were detected in sorghum. In this study, SbNAC6 and SbNAC26 were connected by a dashed line, as well as SbNAC58 and SbNAC129. These duplicated genes showed significantly increased transcript levels under multiple abiotic stresses. Based on syntenic analysis, these NAC paralogs were retained and fixed with similar biological functions.
β-Galactosidase activity was examined to explore the transcriptional activation activity of sorghum NAC gene products. It has been reported that wheat TaNAC4, a homolog of SbNAC56, functions as a transcriptional activator. Significantly increased transcriptional activity of SbNAC56 was particularly noticeable (Fig. 4a, b). While, SbNAC6 performed increased β-galactosidase activity, suggesting the enhanced transcriptional activation (Fig. 4b). For its homolog gene in sorghum, SbSNAC1 was confirmed to have relative higher β-galactosidase activity in the C terminus than in the other deletion regions. This indicated that SbNAC6 functions as a transcriptional activator. To some extent, these stress-responsive SbNACs may have disparate regulatory mechanisms.
Prokaryotic expression analyses were presented to illustrate the Mw of these three SbNAC proteins. As shown in Fig. 6, the theoretical Mw values were consistent with the results of the SDS-PAGE analyses. For example, the theoretical Mw of pET40b-SbNAC6 was 35.42 kDa, while SDS-PAGE indicated 35.09 kDa.
The phytohormone ABA is synthesized under many conditions of environmental stress, including drought, cold, and high soil salinity (Wang et al. 2015). In this study, combined with phylogenetic analysis, we identified seven abiotic stress-responsive NAC genes in sorghum, among which SbNAC17, SbNAC46, and SbNAC73 were significantly upregulated by various abiotic stresses and ABA treatment (Fig. 5). In comparison with the control treatment, the expression levels of these seven SbNAC genes were commonly upregulated by at least 10-fold under conditions of ABA treatment (Fig. 5c). This suggested that these NAC members may play important roles in regulation of abiotic stress responses in the ABA signaling pathway. Under conditions of PEG, cold and salt stress treatments, the expression patterns of the selected SbNACs were indicative of the divergence in regulatory mechanisms (Fig. 5b, d–f). SbNAC6, a homolog of ZmSNAC1 in maize and SbSNAC1 in sorghum, both of which performed similar expression patterns (Lu et al. 2012, 2013), was also induced by salinity, dehydration, cold and ABA. Overexpression of ZmSNAC1 and SbSNAC1 enhanced the dehydration and drought stress tolerance in transgenic Arabidopsis. Previous studies inferred that SbNAC6 is likely to play a potential role in drought stress tolerance. Similarly, the transcript level of SbNAC73, an ortholog of wheat TaNAC69-1, was significantly increased by drought and salt treatments (Fig. 1) (Baloglu et al. 2012). The divergent expression patterns suggested that SbNACs may have various roles in regulation of abiotic stress signaling pathways, and may therefore be excellent candidates for improving abiotic stress resistance.
In this study, we detected the sensitivity to ABA at seed germination stage. Compared to the WT plants, the germination of transgenic plants was significantly inhibited by ABA, NaCl and d-Mannitol. These finding suggest that SbNAC56 may function as a transcriptional activator in ABA-dependent signaling pathway in plant growth, development and response to osmotic stresses.
Overall, in order to mine the drought responsive NAC genes in sorghum and exam the potentially biological functions, we presented detailed descriptions of the sorghum NAC genes. Based on the results of phylogenetic analysis, the 145 SbNAC TFs, along with some well-known stress-responsive NAC TFs in model plants, wheat, and soybean, were classified into 16 groups. Comprehensive analysis of the 145 SbNACs was presented and seven stress related candidates were isolated. Of these, four selected NAC proteins were confirmed to be transcriptional activators. Furthermore, the expression profiles of these seven SbNACs strongly implied diversity in their expression patterns under conditions of abiotic stress in the ABA-dependent pathway. The molecular mechanisms of sorghum NAC genes in abiotic stresses response will be examined in further studies. This work revealed the genomic landscape and expression patterns of seven sorghum NAC genes associated stress response. It can also provide the theoretical basis and candidate gene resources for crops genetic improvement, which will lay the foundation for further function analyses and breeding new varieties of crops.
Abbreviations
- NAC :
-
NAM, ATAF and CUC transcription factor
- TF :
-
Transcription factor
- TM :
-
Transmembrane
- qRT-PCR :
-
Quantitative real-time polymerase chain reaction
- SD :
-
Synthetic dropout
- Chr :
-
Chromosomes
References
Baloglu MC, Oz MT, Oktem HA et al (2012) Expression analysis of TaNAC69-1 and TtNAMB-2, wheat NAC family transcription factor genes under abiotic stress conditions in durum wheat (Triticum turgidum). Plant Mol Biol Rep 30:1246–1252
Broun P (2004) Transcription factors as tools for metabolic engineering in plants. Curr Opin Plant Biol 7:202–209
Endre A, Bantte K (2017) Evaluation and association mapping of agronomic traits for drought tolerance in sorghum [Sorghum bicolor (L.) Moench]. Afr J Biotechnol 16:631–642
Guo YF, Gan SS (2006) AtNAP, a NAC family transcription factor, has an important role in leaf senescence. Plant J 46:601–612
Guo HS, Xie Q, Fei JF (2005) MicroRNA directs mRNA cleavage of the transcription factor NAC1 to down regulate auxin signals for Arabidopsis lateral root development. Plant Cell 17:1376–1386
Guo WW, Zhang JX, Zhang N et al (2015) The wheat NAC transcription factor TaNAC2L is regulated at the transcriptional and post-translational levels and promotes heat stress tolerance in transgenic Arabidopsis. PloS ONE 10:e0135667
Hao YJ, Wei W, Song QX et al (2011) Soybean NAC transcription factors promote abiotic stress tolerance and lateral root formation in transgenic plants. Plant J 68:302–313
He XJ, Mu RL, Cao WH et al (2005) AtNAC2, a transcription factor downstream of ethylene and auxin signaling pathways, is involved in salt stress response and lateral root development. Plant J 44:903–916
Hu RB, Qi G, Kong YZ et al (2010) Comprehensive analysis of NAC domain transcription factor gene family in Populus trichocarpa. BMC Plant Biol 10:1–23
Ji L, Hu RB, Jiang JX et al (2014) Molecular cloning and expression analysis of 13 NAC transcription factors in Miscanthus lutarioriparius. Plant Cell Rep 33:2077–2092
Jin L, Tran DQ, McLachlan JA et al (1997) Several synthetic chemicals inhibit progesterone receptor-mediated transactivation in yeast. Biochem Biophys Res Commun 233:139–146
Jing L, Guo G, Guo W et al (2012) miRNA164-directed cleavage of ZmNAC1 confers lateral root development in maize (Zea mays L.). BMC Plant Biol 12:220
Kebede H, Subudhi PK, Rosenow DT et al (2001) Quantitative trait loci influencing drought tolerance in grain sorghum (Sorghum bicolor L. Moench). Theor Appl Genet 103:266–276
Li B, Ni ZY, Li XD et al (2012) Construction of prokaryotic expression vector, protein purification and identification of GhCOMT1 gene from Gossypium hirsutum L. Acta Bot Boreali-Occidential Sinica 32:1971–1976
Liu TK, Song XM, Duan WK et al (2014) Genome-wide analysis and expression patterns of NAC transcription factor family under different developmental stages and abiotic stresses in Chinese cabbage. Plant Mol Biol Rep 32:1041–1056
Lu M, Ying S, Zhang DF et al (2012) A maize stress-responsive NAC transcription factor, ZmSNAC1, confers enhanced tolerance to dehydration in transgenic Arabidopsis. Plant Cell Rep 31:1701–1711
Lu M, Zhang DF, Shi YS et al (2013) Expression of SbSNAC1, a NAC transcription factor from sorghum, confers drought tolerance to transgenic Arabidopsis. Plant Cell Tissue Organ Cult 115:443–455
Mao X, Zhang H, Qian X et al (2012) TaNAC2, a NAC-type wheat transcription factor conferring enhanced multiple abiotic stress tolerances in Arabidopsis. J Exp Bot 63:2933–2946
Mao XG, Chen SS, Li A et al (2014) Novel NAC transcription factor TaNAC67 confers enhanced multi-abiotic stress tolerances in Arabidopsis. PloS ONE 9:e84359
Mao HD, Qin F, Wang HW et al (2015) A transposable element in a NAC gene is associated with drought tolerance in maize seedlings. Nat Commun. doi:10.1038/ncomms9326
Mao HD, Yu LJ, Han R et al (2016) ZmNAC55, a maize stress-responsive NAC transcription factor, confers drought resistance in transgenic Arabidopsis. Plant Physiol Biochem 105:55–66
Mitsuda N, Seki M, Shinozaki K et al (2005) The NAC transcription factors NST1 and NST2 of Arabidopsis regulate secondary wall thickenings and are required for anther dehiscence. Plant Cell 17:2993–3006
Movahedi A, Zhang JX, Yin TM et al (2015) Functional analysis of two orthologous NAC, genes, CarNAC3, and CarNAC6, from Cicer arietinum, involved in abiotic stresses in poplar. Plant Mol Biol Rep 33:1539–1551
Nuruzzaman M, Manimekalai R, Sharoni AM et al (2010) Genome-wide analysis of NAC transcription factor family in rice. Gene 465:30–44
Ooka H, Satoh K, Doi K et al (2003) Comprehensive analysis of NAC family genes in Oryza sativa and Arabidopsis thaliana. DNA Res 10:239–247
Oprea CA, Marin DI, Bolohan C (2015) Influence of some technological factors on grain sorghum (Sorghum bicolor (L.) Moench var. Eusorghum) yield grown under the conditions of Southeastern Romania. Agrolife Sci J 4:123–130
Pascual MB, Cánovas FM, Ávila C et al (2015) The NAC transcription factor family in maritime pine (Pinus Pinaster): molecular regulation of two genes involved in stress responses. BMC Plant Biol 15:1–15
Rui QZ, Zheng HY (2015) The Arabidopsis NAC transcription factor NST2 functions together with SND1 and NST1 to regulate secondary wall biosynthesis in fibers of inflorescence stems. Plant Signal Behav 10:e989746
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Shahnejat-Bushehri S, Tarkowska D, Sakuraba Y et al (2016) Arabidopsis NAC transcription factor JUB1 regulates GA/BR metabolism and signaling. Nat Plants. doi:10.1038/nplants.2016.13
Shahnejat-Bushehri S, Allu AD, Mehterov N et al (2017) Arabidopsis NAC transcription factor JUNGBRUNNEN1 exerts conserved control over gibberellin and brassinosteroid metabolism and signaling genes in tomato. Front Plant Sci. doi:10.3389/fpls.2017.00214
Shinozaki K, Yamaguchi-Shinozaki K, Seki M (2003) Regulatory network of gene expression in the drought and cold stress responses. Curr Opin Plant Biol 6:410–417
Souer E, Van HA, Kloos D et al (1996) The no apical meristem gene of Petunia is required for pattern formation in embryos and flowers and is expressed at meristem and primordia boundaries. Cell 85:159–170
Su HY, Zhang SZ, Yin YL et al (2015) Genome-wide analysis of NAM-ATAF1, 2-CUC2 transcription factor family in Solanum lycopersicum. J Plant Biochem Biotechnol 24:176–183
Tamura K, Dudley J, Nei M, Kumar S et al (2007) MEGA4: molecular evolutionary genetics analysis (b) software version 4.0. Mol Biol Evol 24:1596–1599
Thompson JD, Gibson TJ, Plewniak F et al (1997) The CLUSTALX windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nuclic Acids Res 25:4876–4882
Tran L, Nakashima K, Sakuma Y et al (2004) Isolation and functional analysis of Arabidopsis stress-inducible NAC transcription factors that bind to a drought-responsive cis-element in the early responsive to dehydration stress 1 promoter. Plant Cell 16:2481–2498
Wang N, Zheng Y, Xin HP et al (2013) Comprehensive analysis of NAC domain transcription factor gene family in Vitis vinifera. Plant Cell Rep 32:61–75
Wang ZY, Gehring C, Zhu J et al (2015) The Arabidopsis Vacuolar Sorting Receptor1 is required for osmotic stress-induced abscisic acid biosynthesis. Plant Physiol 167:137–152
Xie Q, Guo HS, Dallman G et al (2002) SINAT5 promotes ubiquitin-related degradation of NAC1 to attenuate auxin signals. Nature 419:167–170
Xu ZY, Kim SY, Hyeon YD et al (2013) The Arabidopsis NAC transcription factor ANAC096 cooperates with bZIP-type transcription factors in dehydration and osmotic stress responses. Plant Cell 25:4708–4724
Xue GP, Bower NI, Mcintyre CL et al (2006) TaNAC69 from the NAC superfamily of transcription factors is up-regulated by abiotic stresses in wheat and recognises two consensus DNA-binding sequences. Funct Plant Biol 33:43–57
Xue GP, Way HM, Richardson T et al (2011) Overexpression of TaNAC69 leads to enhanced transcript levels of stress up-regulated genes and dehydration tolerance in bread wheat. Mol Plant 4:697–712
Yang XW, Wang XY, Ji L et al (2015) Overexpression of a Miscanthus lutarioriparius NAC gene MlNAC5 confers enhanced drought and cold tolerance in Arabidopsis. Plant Cell Rep 34:943–958
Yu XW, Liu YM, Wang S et al (2016) CarNAC4, a NAC-type chickpea transcription factor conferring enhanced drought and salt stress tolerances in Arabidopsis. Plant Cell Rep 35:1–15
Zhao X, Yang XW, Pei SQ et al (2016) The miscanthus NAC transcription factor MlNAC9 enhances abiotic stress tolerance in transgenic Arabidopsis. Gene 586:158–169
Zhu JK (2002) Salt and drought stress signal transduction in plants. Annu Rev Plant Biol 53:247–273
Acknowledgements
The work was supported by National Natural Science Foundation of China (Grant No. 31501314), DaBeiNong young teachers’ Scientific Research Fund Project (Grant No. 1014115004/025), State Key laboratory of Crop Biology foundation (Grant No. 2015KF08) and the Scientific Research Project of Beijing Educational Committee (Grant No. KM201510020003).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kadier, Y., Zu, Yy., Dai, Qm. et al. Genome-wide identification, classification and expression analysis of NAC family of genes in sorghum [Sorghum bicolor (L.) Moench]. Plant Growth Regul 83, 301–312 (2017). https://doi.org/10.1007/s10725-017-0295-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-017-0295-y