Abstract
NAC (NAM, ATAF1/2 and CUC2) proteins represent a major plant-specific transcription factor (TF) family and are expressed in various developmental stages and tissues. In this study, 204 NAC genes, including 17 membrane-bound TFs, were identified in the Brassica rapa (Chinese cabbage) genome. The gene structure and motif compositions obtained suggest that the BrNAC gene family is classified broadly into eight groups. The predicted Chinese cabbage NAC genes were distributed in ten chromosomes at various densities but mainly in chromosomes 3 and 10 (14 and 14 %, respectively). A comprehensive analysis of NAC family genes in Chinese cabbage, rice and Arabidopsis was also performed. Phylogenetic analysis of BrNACs, along with their Arabidopsis and rice counterparts, divided the proteins into two major groups (A and B) and 19 subgroups. Homologous, paralogous, and orthologous searches of Chinese cabbage and Arabidopsis revealed gene loss during genome triplication in Chinese cabbage. Finally, the expression of 55 selected Chinese cabbage BrNAC genes was analyzed under different developmental stages and abiotic stress conditions (heat, cold, ABA, GA, and PEG). The selected genes were classified into three types (constitutive expression, no-expression, and period-specific expression) according to their expression profiles, which indicate that the NAC genes are involved in various aspects of the physiological and developmental processes of Chinese cabbage. Under different abiotic stress conditions, several members of BrNACs were upregulated by various abiotic stresses. Our study provides a very useful reference for functional analysis of members of BrNAC gene family in Chinese cabbage.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Chinese cabbage (Brassica rapa), a very important vegetable, is widely cultivated and consumed globally. Numerous studies on Chinese cabbage have been carried out in the last several decades (Wang et al. 2011; Kim et al. 2013). However, various environmental stresses, including high and low temperature, salinity and drought stress, limit the growth and productivity of Chinese cabbage (Abe et al. 2011; Ahmed et al. 2012). Thus, improved tolerance to these stresses may significantly increase Chinese cabbage production. Tolerance of or susceptibility to these stresses depends on the ability of the plant to express a set of genes whose expression is often regulated by specific transcription factors (TFs). These TFs include members of the AP2/ERF, bZIP, NAC, MYB, and WRKY (Berri et al. 2009; Le et al. 2011; Du et al. 2012; Xiong et al. 2013; Alves et al. 2013) families, each of which has a dedicated binding site for activating or repressing the expression of target genes (Udvardi et al. 2007). These genes often function together as a large gene family in plants, with different members of the same gene family participating in different stress responses (Tran and Mochida 2010).
The NAC (NAM, ATAF1/2 and CUC2) genes are plant-specific transcription factors. Numerous NAC members have been observed in the model plant Arabidopsis thaliana (117) (Kawaura et al. 2008), Nicotiana tabacum (152) (Rushton et al. 2008), Glycine max (152) (Le et al. 2011), Oryza sativa (151) (Nuruzzaman et al. 2010), Vitis vinifera (79) (Wang et al. 2013) and Populus trichocarpa (163) (Hu et al. 2010). The N-terminal regions of NAC proteins share a conserved DNA-binding region, which is divided into five subdomains (A–E), whereas the C-terminal region that contains a transcriptional regulatory domain (Ooka et al. 2003) is highly diversified. Some NAC proteins, referred to as NAC with transmembrane motif 1-like, also contain α-helical transmembrane motifs (TMs) at their C-terminal end (Kim et al. 2007a, b, 2010). These features allow them to activate or repress a variety of downstream genes for governing multiple cellular or molecular processes. The NAC transcription factors are multifunctional proteins that serve a variety of biological functions, such as the development of shoot apical meristems (Souer et al. 1996; Aida et al. 1999), lateral root development (Xie et al. 2002), embryonic and floral development (Sablowski and Meyerowitz 1998; Aida et al. 1999), stress-induced flowering (Kim et al. 2007a; Yoo et al. 2007), leaf senescence (Lee et al. 2012), cell cycle regulation (Kim et al. 2006; Willemsen et al. 2008), hormone signaling (Xie et al. 2002; Fujita et al. 2004; Kim et al. 2006; Jensen et al. 2008), and seed development (Sperotto et al. 2009).
Previous studies have shown that many members of the NAC gene family respond to adversities and hormone treatments and that some show tissue-specific expression patterns (Fujita et al. 2004). For example, ANAC019, ANAC055, and ANAC072 are well-characterized stress-responsive Arabidopsis NAC genes, and overexpressing transgenic plants exhibit improved stress tolerance compared with the wild type (Tran et al. 2004; Bu et al. 2008; Jensen et al. 2010). In soybean, the NAC genes GmNAC11 and GmNAC20 are involved in responses to salt and freezing stresses, and overproduction results in significantly enhanced salt and freezing tolerance (Hao et al. 2011). NAC genes could also be induced by cold temperature in B. napus (Hegedus et al. 2003). In wheat, NAC gene participates in transferring nutrient remobilization from leaves to developing seeds (Uauy et al. 2006). The NAC proteins regulate plant responses against various biotic stresses. Lin et al. (2007) found that OsNAC19 is upregulated by Magnaporthe grisea infection, which suggests that OsNAC19 is involved in rice defense response. NAC proteins are involved in responses to viral infections during plant vegetative development (Xie et al. 1999).
The recent completion of high-quality sequencing of the B. rapa genome (http://brassicadb.org/brad/index.php) provides an excellent opportunity for genome-wide analysis of all of the genes belonging to specific gene families. We identified 204 BrNAC genes in Chinese cabbage by database search and classified these genes according to reported genes. We also performed a phylogenetic analysis, mapped genes in ten Chinese cabbage chromosomes, identified membrane-bound proteins, and carried out expression analysis at various developmental stages, environmental stresses, and hormone treatments. The data generated will provide leads for the functional characterisation of Chinese cabbage NAC TFs for stress-response improvement and may be used for the identification and characterisation of NAC TFs in other species.
Materials and Methods
Genome-Wide Identification of the NAC Gene Family in B. rapa
To identify members of the BrNAC gene family in B. rapa, all files related to B. rapa genome sequence data were downloaded from the Brassica Database sharing site (http://brassicadb.org/brad/index.php, 1.5 version). We analyzed the domains of all of the B. rapa proteins using a Hidden Markov Model (HMM) profile of the NAM domain PF02365 retrieved from Pfam 26.0 (http://Pfam.sanger.ac.uk/) with an expected value (e-value) cut-off of 1.0. All identified protein models were subjected to Pfam analysis to confirm the presence of the NAM domain with an e-value cut-off of 1e-3. Moreover, membrane-bound BrNAC proteins were identified using TMHMM Server v. 2.0 (http://www.cbs.dtu.dk/services/TMHMM/).
Chromosomal Location, Gene Structure, and Motif Analysis of the NAC Genes in B. rapa
The specific positions of each BrNAC gene on genome were retrieved from the Brassica Database. The distribution of genes on the chromosomes was plotted using Perl scripts, and the gene structures of the NAC transcription factors were generated using Gene Structure Display Server (GSDS: http://gsds.cbi.pku.edu.cn/).
The conserved motifs in full-length NAC proteins were identified using Multiple Expectation Maximization for Motif Elicitation (MEME) v. 4.9.0 (http://meme.nbcr.net/meme3/meme.html). Analysis was performed with the following parameters: number of repetitions, any; maximum number of motifs, 20; and optimum width of the motif, ≥50. Discovered MEME motifs (≤1E-30) were searched in the InterPro database with InterProScan (Quevillon et al. 2005).
Phylogenetic Analysis and Gene Duplication
To study the phylogenetic relationship of B. rapa NAC proteins with their counterparts in Arabidopsis and rice, full-length NAC protein sequences were respectively retrieved from TAIR10 (http://www.arabidopsis.org) and RGAP7 (http://rice.plantbiology.msu.edu/) as described. Multiple sequence alignment of the full-length protein sequences was performed using the ClustalW program with default parameters. A phylogenetic tree was plotted using MEGA4 software by the neighbor-joining method with 1,000 bootstrap replicates.
To search for potentially duplicated NAC genes in Chinese cabbage and Arabidopsis, OrthoMCL (http://www.orthomcl.org/cgi-bin/OrthoMclWeb.cgi) software was used (Li et al. 2003). The relationships between orthologous and paralogous genes in these two species were shown using the Circos program (Krzywinski et al. 2009).
Plant Material, Growth, and Stress Treatments
Seeds of Chinese cabbage (B. rapa accession ‘Chiifu-401-42’) were germinated on moistened gauze at 25 °C after immersion in water for 2 days at 37 °C and subsequently grown hydroponically on gauze in Petri dishes at 25 °C under continuous light (approximately 2,500 lx). To study the expression patterns of BrNAC genes in different developmental stages, leaves from five developmental stages (the five-leaf, rosette, mature, bud, and flowering stages) were harvested.
Four-week-old seedlings were subjected to various treatments. For gibberellic acid (GA) and abscisic acid (ABA) treatments, the seedlings were sprayed with solutions containing 100 μM GA and 100 μM ABA, respectively. For cold and heat treatments, seedlings were maintained at 4 and 38 °C, respectively, for the indicated durations. For drought treatment, seedling roots were immersed in solutions containing 20 % PEG 6,000 at 25 °C with 60 % humidity. Leaves were harvested at the indicated durations (0, 4, and 12 h) after the initiation of each treatment. Unstressed plants were maintained as controls. After the treatments, seedlings were immediately frozen in liquid nitrogen and stored at −80 °C until RNA isolation. To obtain precise and reproducible results, each of the above experiments was repeated three times.
RNA Extraction and Real-Time Quantitative Reverse Transcription PCR
Total RNA was isolated from 100 mg of frozen tissue according to the method used by Longeman et al. (1987). DNA contamination was removed from the RNA samples with RNase-free DNase I. The total RNA (2 μg) was used to synthesize the first-strand cDNA using a PrimeScript First Strand cDNA Synthesis Kit (Takara, China).
The quantitative reverse transcription PCR (qRT-PCR) primers were designed from non-conserved regions of the genes using Primer 3 v. 0.4.0 software with default parameters (Supplementary Table S1). qRT-PCR assays were performed with three biological and three technical replicates using a real-time PCR system (Applied Biosytems, USA). The B. rapa ACTIN gene served as the internal control. Each reaction was performed in a total volume of 20 μL containing 10 mM of each primer, 50 ng of cDNA, and 10 μL of 2× Power SYBR Green PCR master mix (Applied Biosystems). The PCR amplification conditions included an initial heat-denaturing step at 95 °C for 3 min and 40 cycles of 95 °C for 20 s, 55 °C for 20 s and 72 °C for 20 s. Fluorescence was measured at the end of each cycle. Melting-curve analysis was performed by heating the PCR product from 65 to 95 °C. The expression data for the B. rapa NAC genes are presented as relative units after normalization against the B. rapa ACTIN gene using the 2−ΔΔCT method.
Results
Identification of BrNAC Genes
To identify BrNAC proteins in the B. rapa genome, an HMM search was performed using the HMM profile of the NAM domain. This search resulted in the identification of 208 BrNAC proteins. Subsequently, the protein sequences were confirmed by SMART and Pfam searches for the presence of the NAM domain with an e-value cut-off of 1e-3. Finally, 204 NAC proteins were identified and used for further analysis because of exclusion of four proteins with either no N-terminal NAM domain or with an e-value of >1e-3. The number of NAC proteins in Chinese cabbage (204 members) was greater than that in other plants (not over 200 members). Detailed information on BrNAC genes can be found in Supplementary Table S2.
Genome Distribution and Gene Duplication of BrNAC Genes
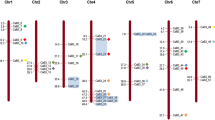
To determine the genomic distribution of BrNAC genes, the DNA sequence of each BrNAC gene was used to search the B. rapa genome database using BLASTN. The physical map positions of 204 BrNAC genes on 10 B. rapa chromosomes were identified. However, one BrNAC gene could not be anchored on any of the B. rapa chromosomes (Bra039353 anchored in Scaffold000164) (Fig. 1 and Supplementary Table S2). Each of the 10 B. rapa chromosomes contained some BrNAC genes but the distribution was uneven (Fig. 1). BrNAC genes were present in all regions on a single chromosome (i.e., at the telomeric ends, near the centromere and in between) and could be distributed individually or in clusters. Chromosomes 3 and 10 contained the largest number of BrNAC genes comprising 29 (14 %) and 29 (14 %) members, respectively, whereas chromosome 4 contained only 7 members (∼3.4 %, Fig. 1).
Chromosomal mapping of the Chinese cabbage NAC transcription factor gene family. Graphical representation of the locations of the putative BrNAC genes on each Chinese cabbage chromosome. The scale bar represents a chromosomal distance of 35.0 Mb. The chromosome number (Chr01–Chr10) is indicated at the bottom of each chromosome
Many members of a gene family could possibly evolve because of the flexibility provided by events of whole genome duplications. Gene duplication, either segmental or in tandem, has been found in several plant TF families, such as MYB and F-box, as well as in NAC (Jain et al. 2007; Cannon et al. 2004; Nuruzzaman et al. 2010). Arabidopsis is the most popular model plant species, and the functions of some Arabidopsis NAC genes have been well characterized. Therefore, we created a comparative syntenic map between Chinese cabbage and Arabidopsis genomes (Fig. 2) to further study the origin, evolutionary history, and putative function of Chinese cabbage BrNAC genes. According to this syntenic map, 124 BrNAC–AtNAC gene pairs were located in 95 pairs of duplicated genomic regions between the Chinese cabbage and Arabidopsis genomes (Supplementary Table S3), which indicates that many of the Chinese cabbage BrNAC genes and their Arabidopsis NAC counterparts maybe derived from a common ancestor. In addition, 30 and 60 paralogous NAC gene pairs were identified in Arabidopsis and Chinese cabbage, respectively (Supplementary Table S3).
Gene duplication and synteny analysis of NAC genes between Chinese cabbage and Arabidopsis. Ten Chinese cabbage (A01 to A10) and five Arabidopsis chromosome (Chr1 to Chr5) maps were based on the orthologous and paralogous pair positions, and demonstrate highly conserved synteny. The red lines represent the orthologous NAC genes between Chinese cabbage and Arabidopsis. The green and blue lines represent the paralogous NAC genes in Chinese cabbage and Arabidopsis, respectively. Colored lines connecting two chromosomal regions denote syntenic regions of genome
Membrane-Associated BrNAC Subfamily
NAC membrane-bound TFs (MTFs) have received much attention as regulators in both biotic and abiotic stress signaling pathways. Using TMHMM Server v. 2.0, we identified 17 (∼8.3 %) BrNAC proteins containing α-helical TMs (Fig. 3 and Supplementary Table S4). Similar to Arabidopsis and rice NAC MTFs, all of the identified BrNAC MTFs also contained a single TM at their C-terminal except Bra012470 (Fig. 3).
Phylogenetic and Motif Analysis of BrNAC Proteins
To clarify the phylogenetic relationships among the BrNAC genes, phylogenetic trees were constructed using BrNAC domain sequences, and an unrooted tree was generated by ClustalW (Thompson et al. 1997) by the neighbor-joining method (Saitou and Nei 1987) and bootstrap analysis (1,000 replicates). The tree was analyzed and illustrated using MEGA software v. 4 (Tamura et al. 2007). The conserved motifs were predicted by the MEME program (Fig. 4 and Supplementary Fig. S1). In general, NAC proteins clustered in together shared similar motif composition. Most of the conserved motifs were found within the N-terminal NAC domain, which indicates that these motifs may be essential for the function of NAC proteins.
Conserved motif compositions of BrNAC proteins. Multiple sequence alignment of 204 full-length BrNAC proteins was done using ClustalW, and the phylogenetic tree was constructed using MEGA4 by the neighbor-joining method with 1,000 bootstrap replicates. Schematic representation of the conserved motifs in the BrNAC proteins was revealed by MEME analysis. Gray lines represent the non-conserved sequences, and each motif is represented by a box numbered at the bottom. The length of protein can be estimated using the scale at the bottom
To study the evolutionary relationship among the NAC TFs in different plant species, another unrooted phylogenetic tree was constructed by the same method used to align OsNAC (rice, monocot) and AtNAC (Arabidopsis, dicot) protein sequences (Fig. 5). Examination of this tree revealed that the BrNAC family may be classified into two major groups (A and B). Groups A and B were further divided into 7 and 12 subgroups, respectively. Group A consisted of 45 BrNAC genes, of which 15 belonged to the A1 subgroup, 2 to A2, 5 to A3, 1 to A4, 5 to A5, 15 to A6 and 2 to A7. Group B consisted of 159 BrNAC genes, of which 30 members belonged to the B1 subgroup, 19 to B2, 5 to B3, 6 to B4, 15 to B5, 5 to B6, 11 to B7, 15 to B8, 9 to B9, 20 to B10, 12 to B11, and 12 to B12 (Fig. 5 and Supplementary Table S5). The NAC members of groups A and B showed high homology with the Arabidopsis and rice NACs. In addition, most subgroups were shared among the NACs from Arabidopsis, rice, and Chinese cabbage, although some species-specific subgroups were also observed. For example, subgroup B3 did not include NAC members from Arabidopsis but included NAC members from Chinese cabbage and rice (Fig. 5 and Supplementary Table S5).
Phylogenetic among the NAC TFs of Chinese cabbage, Arabidopsis, and rice. Multiple sequence alignment of full-length NAC proteins was done using ClustalW, and the phylogenetic tree was constructed using MEGA4 by the neighbor-joining method with 1,000 bootstrap replicates. BrNAC proteins were allocated to two distinct groups (A and B). The tree was divided into 7 phylogenetic subgroups in group A, designated as A1 to A7, 12 phylogenetic subgroups in group B, designated as B1 to B12. The detailed genes in each subgroup were shown in Supplementary Table S5
Expression Patterns of Selected BrNAC Genes in Different Developmental Stages
To determine the function of BrNAC genes in Chinese cabbage in different developmental stages, 55 candidate genes widely representing all of the subfamilies were selected based on sequence similarity comparisons and phylogenetic analyses described in the previous sections. The expression patterns of 55 BrNAC genes in various vegetative and reproductive stages of Chinese cabbage ‘Chiifu’ were investigated by qRT-PCR. Generally, the expression patterns of BrNAC genes can be classified into three types according to their expression profiles (constitutive expression, no-expression, and period-specific expression) (Fig. 6a).
Heat map representation of BrNAC genes in various growth stages (a) and in response to abiotic stresses (b). a Heat map was generated using cluster software. The bar at the bottom of the heat map represents relative expression values, thereby values 0, 1, and 2 represent no-expression, slightly upregulated, and strongly upregulated expression, respectively. The name of the gene is written on the right of the heat map. b Heat map was generated using cluster software. The relative expression values were log2 transformed. The bar at the bottom of the heat map represents relative expression values, thereby values −1, 0, and 1 represent downregulated, unaltered, and upregulated expression, respectively. The name of the gene is written on the right of the heat map
Seventeen BrNAC genes (including Bra000709, Bra005804, and Bra029201) showed a constitutive expression pattern in all of the examined developmental stages (Fig. 7). These genes are closely related to metabolism and cell growth. By contrast, several genes (including Bra033305, Bra013272, Bra007640, Bra002857, and Bra002605) were not expressed in all developmental stages (Fig. 6a). These BrNACs either transcribe at a level too low for detection or have spatial and temporal expression patterns not covered by RT-PCR. The remaining BrNAC genes showed a generally more differentiated expression pattern in the different developmental stages. Bra003998 and Bra004385 were only expressed in reproductive stages, whereas Bra035712 and Bra016098 transcripts were only detected in vegetative stages (Fig. 7).
The relative expression ratio of seven representative BrNAC genes in various growth stages. Relative expression ratios in different stage samples (1, 2, 3, 4, and 5 represent five-leaf stage, rosette stage, mature stage, bud stage, and flowering stage, respectively) have been calculated with reference (actin gene). Different capital letters indicate significant differences between growth stages in the same cultivar (P < 0.05). The name of the gene is written on the top of each bar diagram (error bars indicate standard deviation)
Expression Patterns of Selected BrNAC Genes Under Different Abiotic Conditions
Gene expression patterns can provide crucial clues for determining gene function. To investigate the role of NAC genes in Chinese cabbage under diverse environmental stresses, above 55 genes were subjected to quantitative expression analysis in response to heat, cold, ABA, GA and PEG during short (4 h) and long (12 h) treatment durations. The heat map representation for expression in response to abiotic stresses is shown in Fig. 6b. Overall, qRT-PCR analysis demonstrated that all of the genes displayed variations in their expression behavior in response to one or more stresses during the course of the experiments (Fig. 6b). In all five stress treatments, six genes were significantly induced (including Bra003244 and Bra011037). However, nearly half of the total genes were repressed (including Bra002403 and Bra006019) in all stress treatments (Fig. 8).
The relative expression ratio of four representative BrNAC genes in different abiotic conditions. Relative expression ratios in different stage samples (CK, before treatment; ABA; PEG; GA; Cold and Heat, under stress treatments for 4 and 12 h) have been calculated with reference (actin gene). Different capital letters indicate significant differences between growth stages in the same cultivar (P < 0.05). The name of the gene is written on the top of each bar diagram (error bars indicate standard deviation)
In the temperature treatments, 12 and 9 BrNAC genes (including Bra026595 and Bra003244) continuously increased after heat and cold treatments, respectively (Fig. 9). However, several genes (13 BrNAC genes under heat treatment and 12 BrNAC genes under cold treatment, including Bra034118) initially increased at 4 h and subsequently decreased significantly at the transcriptional level at 12 h; by contrast, the Bra027596 gene was repressed at 4 h and subsequently increased in expression at 12 h (Fig. 9). The remaining BrNAC genes were downregulated (Fig. 6b).
The relative expression ratio of representative BrNAC genes in different abiotic conditions. Relative expression ratios in different stage samples (CK, before treatment; Cold and Heat, under stress treatments for 4 and 12 h; ABA and GA, under stress treatments for 4 and 12 h) have been calculated with reference (actin gene). Different capital letters indicate significant differences between growth stages in the same cultivar (P < 0.05). The name of the gene is written on the top of each bar diagram (error bars indicate standard deviation)
Plant hormones are crucial in the regulation of different plant processes, such as signaling and expression during abiotic and biotic stresses. We attempted to evaluate the expression pattern of these 55 genes under various hormone treatments (Fig. 6b). Several genes were exclusively induced (including Bra000192 and Bra033306) or repressed (including Bra000709 and Bra001365) in all treatments (Fig. 9). These genes may contribute to parts of a general hormonal response rather than a treatment-specific response. For example, Bra033305 and Bra037914 were differentially induced by ABA and GA at 4 and 12 h (Fig. 9).
After PEG treatment, six BrNAC genes (including Bra004385 and Bra011037) increased in expression from 4 to 12 h (Fig. 10). Ten BrNAC genes (including Bra005804 and Bra015750) increased at 4 h and then exhibited significant downregulation at the transcriptional level at 12 h (Fig. 10). Bra000282 and Bra027596 were repressed at 4 h and then displayed induced expression at 12 h treatment (Fig. 10). All other BrNAC genes were repressed (Fig. 6b).
The relative expression ratio of six representative BrNAC genes in PEG condition. Relative expression ratios in different stage samples (CK, before treatment; PEG under stress treatment for 4 and 12 h) have been calculated with reference (actin gene). Different capital letters indicate significant differences between growth stages in the same cultivar (P < 0.05). The name of the gene is written on the top of each bar diagram (error bars indicate standard deviation)
Discussion
A total of 204 BrNAC genes were identified through genome-wide analysis. The NAC gene family in Chinese cabbage is by far the largest compared with other plant species. For example, only 117 NAC genes in Arabidopsis, 151 in rice, 79 in grape, 180 in apple, 152 in soybean, 152 in tobacco, and 163 in poplar have been found; these findings indicate that the NAC genes in Chinese cabbage have expanded (Kawaura et al. 2008; Rushton et al. 2008; Hu et al. 2010; Nuruzzaman et al. 2010; Le et al. 2011; Wang et al. 2013). The presence of additional NAC genes in the Chinese cabbage genome may reflect the involvement of these genes in complicated transcriptional regulation processes in this species.
Comparative genomic analysis confirmed that Chinese cabbage underwent genome triplication since its divergence from Arabidopsis (Wang et al. 2011). Among the orthologous gene pairs of Chinese cabbage and Arabidopsis, we found that each Arabidopsis NAC gene had one to four Chinese cabbage orthologous genes, demonstrating that a few NAC transcription factors in Chinese cabbage underwent duplication accompanied by genome triplication. However, the gene number in the Chinese cabbage genome was notably less than three times the Arabidopsis gene number, revealing that gene loss also occurred during polyploid speciation. Based on the results of comparative genomic analysis, we were able to deduce the function of several Chinese cabbage BrNAC genes according to their Arabidopsis homologs, which could largely facilitate our research on the roles of BrNAC genes in Chinese cabbage. Most plants have experienced one or more rounds of ancient polyploidy. From Supplementary Table S3, NAC genes in B. rapa are over retained after WGT. Three subgenomes, the least fractionated blocks (LF), the medium fractionated blocks (MF1) and the most fractionated blocks (MF2) were defined according to the proportions of genes retained in each of these subgenomes relative to Arabidopsis (Wang et al. 2011). Compared with Arabidopsis, our analysis confirms that a gene duplication event occurred in the NAC gene family of Chinese cabbage. The distribution of BrNACs among the three subgenomes is biased, mainly in LF subgenomes (46.08 %, Supplementary Table S6).
To examine the diversity in Chinese cabbage NAC genes, ten conserved motifs were predicted by the MEME program (Fig. 4). In general, NAC proteins clustered in the same subgroup shared a similar motif composition (Fig. 4). Most of the conserved motifs were found within the N-terminal of the NAC domain. Such motif sequences conservation or variation between the proteins specifies the functional equivalence or diversification, respectively, with respect to the various aspects of biological functions (Puranik et al. 2012). As earlier reported, members of a particular subfamily showed a tendency to have comparable motif composition (Fang et al. 2008; Pinheiro et al. 2009; Hu et al. 2010), and certain motifs were found to be deleted or duplicated within particular clades. To further examine the phylogenetic relationships among the NAC proteins in Chinese cabbage, Arabidopsis and rice, a comprehensive phylogenetic analysis was performed from alignments of the NAC protein sequences. According to the clade support values, topology of the tree, and classification of Arabidopsis and rice, BrNAC family is classified into two major groups (A and B), including 19 subgroups (Fig. 5). Subgroup B3 did not include any NAC of Arabidopsis, which indicates that NACs may have been either lost in Arabidopsis or acquired in Chinese cabbage and rice after the divergence of Arabidopsis from their last common ancestor and that these NACs may have specialized roles in Chinese cabbage and rice. In similar studies, phylogenetic analysis divided apple, poplar, and soybean NACs into 15, 10, and 6 subgroups, respectively (Hu et al. 2010; Le et al. 2011). The current observations indicate that NAC proteins in Chinese cabbage (19 subgroups) are more diverse than those in apple, poplar and soybean. Close association of BrNAC families with their counterparts in other plants may be an implication of sequence conservation and evidence to their similar function in plant roles. Such phylogeny-based function prediction has been well applied for prediction of stress-responsive NAC proteins in other species like rice, Arabidopsis and soybean (Fang et al. 2008; Ooka et al. 2003; Nuruzzaman et al. 2010; Le et al. 2011).
NAC MTFs have been implicated in plant responses to abiotic stress (Kim et al. 2007b, 2010; Lee et al. 2012). Similar to Arabidopsis and rice NAC MTFs (Kim et al. 2007b), all of the identified BrNAC MTFs also contained a single TM at their C-terminal, except Bra012470, which contained a TM at its N-terminal (Fig. 3). NAC MTFs have been known as transcription regulators to be activated via posttranslational modifications under environmental stresses (Puranik et al. 2013). A previous study revealed that GmNAC013 and GmNAC136 contain two TMs (Le et al. 2011). Other studies also indicate that a number of the functionally characterized NAC MTFs are closely related to plant responses to environmental stresses (Saeed et al. 2003; Fujita et al. 2004). Thus, BrNAC MTFs may follow a similar path of nuclear localization and downstream stress-responsive gene expression only after their membrane anchors have been trimmed by proteases following adverse environmental cues.
Gene expression patterns can provide important clues for gene function. According to the phylogenetic analysis, we chose 55 BrNAC genes to examine the expression in leaves of different developmental stages using RT-PCR (Fig. 6a). Among constitutively expressed genes, Bra000709 (Fig. 7, ortholog of AT4G10350), which controls cell wall maturation processes, is closely related to metabolism and cell growth. These genes are necessary in all developmental stages (Bennett et al. 2010). Among no-expression genes, Bra000192 (Fig. 6, ortholog of AT2G27300) is expressed in a tissue-enriched manner in roots and regulates the growth of lateral roots (Kim et al. 2007a), Bra002802 (Fig. 6, ortholog of AT5G56620) is only expressed in the hypocotyl (Schmid et al. 2005), Bra007640 (Fig. 6, ortholog of AT3G61910) participates in regulation of the secondary thickening of anther walls (Mitsuda et al. 2005) and Bra010077 (Fig. 6, ortholog of AT5G62380) is expressed in the presence of auxin, cytokinin, and brassinosteroids (Ohashi-Ito et al. 2010). However, in our research, leaves were used as samples for RT-PCR and no-expression BrNACs may have spatial or induced expression patterns not covered by the method. These observations indicate that various BrNACs may be associated with diversified functions similar to their Arabidopsis orthologs, for example, among period-specific expressed genes, Bra003998 and Bra004385 (Fig. 7, ortholog of AT1G69490) are expressed in the floral primordia and upregulated by AP3 and PI, which indicates that these genes may be involved in the regulation of flower development (Sablowski and Meyerowitz 1998). The specific expression profiling of BrNACs might enable the combinatorial usage of BrNACs in transcriptional regulation of different growth periods or tissues, whereas ubiquitously expressed BrNACs may regulate a broad set of genes. For example, OsNAC10 expressed in both roots and panicles and induced by drought and salinity, when overexpressed, improved root growth, increased grain yield significantly under drought conditions (Jeong et al. 2010). But, the function of these BrNAC genes requires further verification.
Adverse environmental conditions, such as heat, cold, and drought, can cause irreversible damage to the growth and productivity of Chinese cabbage (Wang et al. 2011; Kim et al. 2013). NAC proteins have also been shown to regulate a variety of plant processes by mediating hormone signaling (Fujita et al. 2004). Thus, using qRT-PCR combined with sequence similarity comparisons and phylogenetic analyses, we identified 6 BrNAC genes (including Bra003244 and Bra011037, Fig. 8) that responded to all five abiotic stress treatments (heat, cold, drought, ABA, and GA) and determined these genes to be common stress-response genes. These genes may be a part of a general stresses response rather than being treatment-specific. For example, Bra002403 and Bra006019 (Fig. 8) were repressed in all stress treatments. In a previous study, NAC proteins positively regulated stress responses by activating stress-related genes (Jensen et al. 2008). NAC proteins have also been shown to negatively regulate defense responses by suppressing defense-related gene expression (Wang et al. 2009). Some genes, such as Bra015750, Bra034119, Bra036520, and Bra039915, exhibit different expression patterns under different stress conditions (Fig. 6b), which may indicate that these genes are involved in the communication between different signal transduction pathways. Hormones are extensively involved in influencing signaling response by acting in conjunction with or in opposition to each other for maintaining the cellular homeostasis (Fujita et al. 2006). The NAC TFs form a complex but interesting group as important arbitrators of this process (Puranik et al. 2012). The highly differential expression patterns of BrNAC genes observed in this study may be utilized to improved plants performance under stressful conditions.
Finally, a complete analysis of the NAC family in Chinese cabbage was presented, and the relationship between this family and abiotic stress response was emphasized. The real-time PCR of many NAC transcription factors showed period-specific and tissue-specific expression. Comparative analysis of BrNACs with their respective Arabidopsis orthologs helped predict the potential functions of several BrNAC proteins. Together, although the function of BrNACs remain to be further investigated, the above results also indicate that some given members of the BrNAC gene family may be involved in different abiotic and developmental stages and increase our knowledge about the involvement of the NAC transcription factor in plant resistance and growth.
References
Abe H, Narusaka Y, Sasaki I, Hatakeyama K, Shin-I S, Narusaka M, Fukami-Kobayashi K, Matsumoto S, Kobayashi M (2011) Development of full-length cDNAs from Chinese cabbage (Brassica rapa Subsp. pekinensis) and identification of marker genes for defence response. DNA Res 18:277–289
Ahmed NU, Park JI, Seo MS, Kumar TS, Lee IH, Park BS, Nou IS (2012) Identification and expression analysis of chitinase genes related to biotic stress resistance in Brassica. Mol Biol Rep 39:3649–3657
Aida M, Ishida T, Tasaka M (1999) Shoot apical meristem and cotyledon formation during Arabidopsis embryogenesis: interaction among the CUP-SHAPED COTYLEDON and SHOOT MERISTEMLESS genes. Development 126:1563–1570
Alves MS, Dadalto SP, Gonçalves AB, De Souza GB, Barros VA, Fietto LG (2013) Plant bZIP transcription factors responsive to pathogens: a review. Int J Mol Sci 14:7815–7828
Bennett T, van den Toorn A, Sanchez-Perez GF, Campilho A, Willemsen V, Snel B, Scheres B (2010) SOMBRERO, BEARSKIN1, and BEARSKIN2 regulate root cap maturation in Arabidopsis. Plant Cell 22:640–654
Berri S, Abbruscato P, Faivre-Rampant O, Brasileiro AC, Fumasoni I, Satoh K, Kikuchi S, Mizzi L, Morandini P, Pè ME, Piffanelli P (2009) Characterization of WRKY co-regulatory networks in rice and Arabidopsis. BMC Plant Biol 9:120
Bu Q, Jiang H, Li CB, Zhai Q, Zhang J, Wu X, Sun J, Xie Q, Li C (2008) Role of the Arabidopsis thaliana NAC transcription factors ANAC019 and ANAC055 in regulating jasmonic acid-signaled defense responses. Cell Res 18:756–767
Cannon SB, Mitra A, Baumgarten A, Young ND, May G (2004) The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana. BMC Plant Biol 4:10
Du H, Yang SS, Liang Z, Feng BR, Liu L, Huang YB, Tang YX (2012) Genome-wide analysis of the MYB transcription factor superfamily in soybean. BMC Plant Biol 12:106
Fang Y, You J, Xie K, Xie W, Xiong L (2008) Systematic sequence analysis and identification of tissue-specific or 321 stress-responsive genes of NAC transcription factor family in rice. Mol Genet Genomics 280:535–546
Fujita M, Fujita Y, Maruyama K, Seki M, Hiratsu K, Ohme-Takagi M, Tran LS, Yamaguchi-Shinozaki K, Shinozaki K (2004) A dehydration-induced NAC protein, RD26, is involved in a novel ABA-dependent stress-signaling pathway. Plant J 39:863–876
Fujita M, Fujita Y, Noutoshi Y, Takahashi F, Narusaka Y et al (2006) Crosstalk between abiotic and biotic stress responses: a current view from the points of convergence in the stress signaling networks. Curr Opin Plant Biol 9:436–442
Hao YJ, Wei W, Song QX, Chen HW, Zhang YQ, Wang F, Zou HF, Lei G, Tian AG, Zhang WK, Ma B, Zhang JS, Chen SY (2011) Soybean NAC transcription factors promote abiotic stress tolerance and lateral root formation in transgenic plants. Plant J 68:302–313
Hegedus D, Yu M, Baldwin D, Gruber M, Sharpe A, Parkin I, Whitwill S, Lydiate D (2003) Molecular characterization of Brassica napus NAC domain transcriptional activators induced in response to biotic and abiotic stress. Plant Mol Biol 53:383–397
Hu R, Qi G, Kong Y, Kong D, Gao Q et al (2010) Comprehensive analysis of NAC domain transcription factor gene family in Populus trichocarpa. BMC Plant Biol 10:145
Jain M, Nijhawan A, Arora R, Agarwal P, Ray S, Sharma P, Kapoor S, Tyagi AK, Khurana JP (2007) F-Box proteins in rice. Genome-wide analysis, classification, temporal and spatial gene expression during panicle and seed development, and regulation by light and abiotic stress. Plant Physiol 143:1467–1483
Jensen MK, Hagedorn PH, de Torres-Zabala M, Grant MR, Rung JH, Collinge DB, Lyngkjaer MF (2008) Transcriptional regulation by an NAC (NAMATAF1,2-CUC2) transcription factor attenuates ABA signalling for efficient basal defence towards Blumeria graminis f. Sp. Hordei in Arabidopsis. Plant J 56:867–880
Jensen MK, Kjaersgaard T, Nielsen MM, Galberg P, Petersen K, O’Shea C, Skriver K (2010) The Arabidopsis thaliana NAC transcription factor family: structure-function relationships and determinants of ANAC019 stress signaling. Biochem J 426:183–196
Jeong JS, Kim YS, Baek KH et al (2010) Root-specific expression of OsNAC10 improves drought tolerance and grain yield in rice under field drought conditions. Plant Physiol 153:185–197
Kawaura K, Mochida K, Ogihara Y (2008) Genome-wide analysis for identification of salt-responsive genes in common wheat. Funct Integr Genom 8:277–286
Kim YS, Kim SG, Park JE, Park HY, Lim MH, Chua NH, Park CM (2006) A membrane bound NAC transcription factor regulates cell division in Arabidopsis. Plant Cell 18:3132–3144
Kim SG, Kim SY, Park CM (2007a) A membrane associated NAC transcription factor regulates salt-responsive flowering via flowering locus t in Arabidopsis. Planta 226:647–654
Kim SY, Kim SG, Kim YS, Seo PJ, Bae M, Yoon HK, Park CM (2007b) Exploring membrane-associated NAC transcription factors in Arabidopsis: implications for membrane biology in genome regulation. Nucleic Acids Res 35:203–213
Kim SG, Lee S, Seo PJ, Kim SK, Kim JK, Park CM (2010) Genome-scale screening and molecular characterization of membrane-bound transcription factors in Arabidopsis and rice. Genomics 95:56–65
Kim YB, Li X, Kim SJ, Kim HH, Lee J, Kim H, Park SU (2013) MYB transcription factors regulate glucosinolate biosynthesis in different organs of Chinese cabbage (Brassica rapa ssp. pekinensis). Molecules 18:8682–8695
Krzywinski M, Schein J, Birol I, Connors J, Gascoyne R, Horsman D, Jones SJ, Marra MA (2009) Circos: an information aesthetic for comparative genomics. Genome Res 19:1639–1645
Le DT, Nishiyama R, Watanabe Y, Mochida K, Yamaguchi-Shinozaki K, Shinozaki K, Tran LS (2011) Genome-wide survey and expression analysis of the plant-specific NAC transcription factor family in soybean during development and dehydration stress. DNA Res 18:263–276
Lee S, Seo PJ, Lee HJ, Park CM (2012) A NAC transcription factor NTL4 promotes reactive oxygen species production during drought-induced leaf senescence in Arabidopsis. Plant J 70:831–844
Li L, Stoeckert CJ Jr, Roos DS (2003) OrthoMCL: identification of ortholog groups for eukaryotic genomes. Genome Res 13:2178–2189
Lin R, Zhaom W, Mengm X, Wang M, Peng Y (2007) Rice gene OsNAC19 encodes a novel NAC-domain transcription factor and responds to infection by Magnaporthe grisea. Plant Sci 172:120–130
Longeman J, Schell J, Willmitzer L (1987) Improved method for the isolation of RNA from plant tissues. Anal Biochem 163:16–20
Mitsuda N, Seki M, Shinozaki K, Ohme-Takagi M (2005) The NAC transcription factors NST1 and NST2 of Arabidopsis regulate secondary wall thickenings and are required for anther dehiscence. Plant Cell 17:2993–3006
Nuruzzaman M, Manimekalai R, Sharoni AM, Satoh K, Kondoh H, Ooka H, Kikuchi S (2010) Genome-wide analysis of NAC transcription factor family in rice. Gene 465:30–44
Ohashi-Ito K, Oda Y, Fukuda H (2010) Arabidopsis VASCULAR-RELATED NAC-DOMAIN6 directly regulates the genes that govern programmed cell death and secondary wall formation during xylem differentiation. Plant Cell 22:3461–3473
Ooka H, Satoh K, Doi K, Nagata T, Otomo Y, Murakami K, Matsubara K, Osato N, Kawai J, Carninci P, Hayashizaki Y, Suzuki K, Kojima K, Takahara Y, Yamamoto K, Kikuchi S (2003) Comprehensive analysis of NAC family genes in Oryza sativa and Arabidopsis thaliana. DNA Res 10:239–247
Pinheiro GL, Marques CS, Costa MD, Reis PA, Alves MS et al (2009) Complete inventory of soybean NAC transcription factors: sequence conservation and expression analysis uncover their distinct roles in stress response. Gene 444:10–23
Puranik S, Sahu PP, Srivastava PS, Prasad M (2012) NAC proteins: regulation and role in stress tolerance. Trends Plant Sci 17:1360–1385
Puranik S, Sahu PP, B VS, Mandal SN, Parida SK, Prasad M (2013) Comprehensive genome-wide survey, genomic constitution and expression profiling of the NAC transcription factor family in foxtail millet (Setaria italica L.). PLoS One 8(5):64594
Quevillon E, Silventoinen V, Pillai S, Harte N, Mulder N, Apweiler R, Lopez R (2005) InterProScan: protein domains identifier. Nucleic Acids Res 33:W116–W120
Rushton PJ, Bokowiec MT, Han S, Zhang H, Brannock JF, Chen X, Laudeman TW, Timko MP (2008) Tobacco transcription factors: novel insights into transcriptional regulation in the Solanaceae. Plant Physiol 147:280–295
Sablowski RW, Meyerowitz EM (1998) A homolog of NO APICAL MERISTEM is an immediate target of the floral homeotic genes APETALA3/PISTILLATA. Cell 92:93–103
Saeed AI, Sharov V, White J, Li J, Liang W, Bhagabati N, Braisted J, Klapa M, Currier T, Thiagarajan M, Sturn A, Snuffin M, Rezantsev A, Popov D, Ryltsov A, Kostukovich E, Borisovsky I, Liu Z, Vinsavich A, Trush V, Quackenbush J (2003) TM4: a free, open-source system for microarray data management and analysis. Biotechniques 34:374–378
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Schmid M, Davison TS, Henz SR, Pape UJ, Demar M, Vingron M, Schölkopf B, Weigel D, Lohmann JU (2005) A gene expression map of Arabidopsis thaliana development. Nat Genet 37:501–506
Souer E, van Houwelingen A, Kloos D, Mol J, Koes R (1996) The no apical meristem gene of Petunia is required for pattern formation in embryos and flowers and is expressed at meristem and primordial boundaries. Cell 85:159–170
Sperotto RA, Ricachenevsky FK, Duarte GL, Boff T, Lopes KL, Sperb ER, Grusak MA, Fett JP (2009) Identification of upregulated genes in flag leaves during rice grain filling and characterization of OsNAC5, a new ABA-dependent transcription factor. Planta 230(5):985–1002
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Tran LSP, Mochida K (2010) Identification and prediction of abiotic stress responsive transcription factors involved in abiotic stress signaling in soybean. Plant Signal Behav 5:227–255
Tran LS, Nakashima K, Sakuma Y, Simpson SD, Fujita Y, Maruyama K, Fujita M, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2004) Isolation and functional analysis of Arabidopsis stress-inducible NAC transcription factors that bind to a drought responsive cis-element in the early responsive to dehydration stress 1 promoter. Plant Cell 16:2481–2498
Uauy C, Distelfeld A, Fahima T, Blechl A, Dubcovsky J (2006) A NAC gene regulating senescence improves grain protein, zinc, and iron content in wheat. Science 314:1298–1301
Udvardi MK, Kakar K, Wandrey M, Montanari O, Murray J, Andriankaja A, Zhang JY, Benedito V, Hofer JM, Chueng F, Town CD (2007) Legume transcription factors: global regulators of plant development and response to the environment. Plant Physiol 144:538–549
Wang X, Basnayake BM, Zhang H, Li G, Li W, Virk N, Mengiste T, Song F (2009) The Arabidopsis ATAF1, a NAC transcription factor, is a negative regulator of defense responses against necrotrophic fungal and bacterial pathogens. Mol Plant Microbe Interact 22:1227–1238
Wang X, Wang H, Wang J, Sun R, Wu J, Liu S, Bai Y, Mun JH, Bancroft I, Cheng F (2011) The genome of the mesopolyploid crop species Brassica rapa. Nat Genet 43:1035–1039
Wang N, Zheng Y, Xin H, Fang L, Li S (2013) Comprehensive analysis of NAC domain transcription factor gene family in Vitis vinifera. Plant Cell Rep 32:61–75
Willemsen V, Bauch M, Bennett T, Campilho A, Wolkenfelt H, Xu J, Haseloff J, Scheres B (2008) The NAC domain transcription factors FEZ and SOMBRERO control the orientation of cell division plane in Arabidopsis root stem cells. Dev Cell 15:913–922
Xie Q, Sanz-Burgos AP, Guo H, Garcia JA, Gutierrez C (1999) GRAB proteins, novel members of the NAC domain family, isolated by their interaction with a geminivirus protein. Plant Mol Biol 39:647–656
Xie Q, Guo HS, Dallman G, Fang S, Weissman AM, Chua NH (2002) SINAT5 promotes ubiquitinrelated degradation of NAC1 to attenuate auxin signals. Nature 419:167–170
Xiong AS, Jiang HH, Zhuang J, Peng RH, Jin XF, Zhu B, Wang F, Zhang J, Yao QH (2013) Expression and function of a modified AP2/ERF transcription factor from Brassica napus enhances cold tolerance in transgenic Arabidopsis. Mol Biotechnol 53:198–206
Yoo SY, Kim Y, Kim SY, Lee JS, Ahn JH (2007) Control of flowering time and cold response by a NAC domain protein in Arabidopsis. PLoS ONE 2:e642
Acknowledgments
This work was supported by the National Natural Science Foundation of China (31330067 and 31301782), the National High Technology Research and Development Program of China (863 Program, No. 2012AA100101), the Fundamental Research Funds for the Central Universities of China (KYZ201111), the Natural Science Foundation of Jiangsu Province (BK20130673).
Author information
Authors and Affiliations
Corresponding author
Additional information
Tongkun Liu and Xiaoming Song contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Fig. S1
(DOC 177 kb)
Supplementary Table S1
(DOC 130 kb)
Supplementary Table S2
(XLS 48 kb)
Supplementary Table S3
(XLS 29 kb)
Supplementary Table S4
(DOC 29 kb)
Supplementary Table S5
(XLSX 16 kb)
Supplementary Table S6
(XLS 37 kb)
Rights and permissions
About this article
Cite this article
Liu, T., Song, X., Duan, W. et al. Genome-Wide Analysis and Expression Patterns of NAC Transcription Factor Family Under Different Developmental Stages and Abiotic Stresses in Chinese Cabbage. Plant Mol Biol Rep 32, 1041–1056 (2014). https://doi.org/10.1007/s11105-014-0712-6
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-014-0712-6