Abstract
Key message
The 13 MlNAC genes could respond to various abiotic stresses, suggesting their crucial roles in stress response. Overexpression of MlNAC2 in Arabidopsis led to improved drought tolerance.
Abstract
NAC (NAM, ATAF1/2 and CUC2) proteins are plant-specific transcription factors that play crucial roles in plant development, growth and stress responses. In this study, 13 stress-responsive NAC genes were identified from Miscanthus lutarioriparius. Full-length cDNA sequences were obtained for 11 MlNAC genes, which were phylogenetically classified into six subfamilies. Sequence alignment revealed the highly conserved NAC domain in the N-terminus of these MlNACs, while the C-terminus was highly divergent. We performed quantitative real-time RT-PCR to examine the expression profiles of MlNAC genes in different tissues including root, rhizome, mature stem, young stem, leaf and sheath. The 13 MlNAC genes displayed distinct tissue-specific patterns in six tissues examined. To gain further insight into their roles in response to abiotic stresses, expressions of MlNAC genes were analyzed under different stresses and hormone treatments including salt, drought, cold, wounding, abscisic acid, Methyl jasmonate and salicylic acid. The 13 MlNAC genes could respond to at least five stress treatments, and over 100-fold variations in transcript levels of MlNAC1, MlNAC2, MlNAC4, and MlNAC12 were observed in salt, drought and MeJA treatments, which indicated that MlNACs play crucial roles in stress response. Crosstalk among various abiotic stress and hormone responses was also discussed based on the expression of MlNAC genes. Overexpression of MlNAC2 in Arabidopsis (Col-0) led to improved drought tolerance. The water loss rate was significantly lower, and the recovery rate after a 60-min dehydration stress treatment was significantly higher in the MlNAC2 overexpression lines than the control.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the natural environment, plants constantly suffer from various biotic and abiotic stresses. Plants respond and adapt to these stresses to survive under stress conditions at the molecular and cellular levels as well as at the physiological and biochemical levels. A considerable number of stress-related genes are induced during the response and adaptation to diverse stresses. Accumulating evidence indicates that transcription factors NAC (NAM/ATAF1,2/CUC2), bZIP, WRKY, and AP2/EREBP genes play important regulatory roles in stress response by regulating gene expression and signal transduction (Christianson et al. 2010; Jaglo-Ottosen et al. 1998; Jung et al. 2008; Lu et al. 2009; Shinozaki and Yamaguchi-Shinozaki 1997).
The NAC gene superfamily is one of the largest plant-specific transcription factor families. NAC proteins contain a conserved DNA-binding domain in the N-terminal region and a diversified activation domain in C-terminus (Ooka et al. 2003). There are 117 members in Arabidopsis, 151 in rice, 152 in soybean, 147 in foxtail millet, 110 from potato, 163 in poplar and 74 in grapevine, respectively (Hu et al. 2010; Le et al. 2011; Nuruzzaman et al. 2010; Puranik et al. 2013; Singh et al. 2013; Wang et al. 2013). NAC proteins participate in various cellular processes, including shoot apical meristem development (Aida et al. 1997; Hibara et al. 2003; Nikovics et al. 2006), embryo development (Duval et al. 2002), lateral root development (He et al. 2005; Xie et al. 2000), leaf senescence (Guo and Gan 2006), formation of secondary cell wall (Mitsuda et al. 2005), as well as biotic and abiotic stress responses (Delessert et al. 2005; Fujita et al. 2004; He et al. 2005; Mao et al. 2014; Nakashima et al. 2007; Takasaki et al. 2010; Tran et al. 2004; Wang et al. 2009; Wu et al. 2009; Xue et al. 2011). It has been reported that ANAC019, ANAC055, and ANAC072 were induced by drought, salinity and abscisic acid (ABA) treatments in Arabidopsis. Overexpression of these genes enhanced tolerance to drought and salt stresses (Tran et al. 2004). ATAF1 from Arabidopsis was shown to play positive roles in responding to drought stress, the transgenic plants displayed enhanced tolerance to drought but led to dwarf stature and short primary roots (Wu et al. 2009). In rice, more and more stress-responsive NAC genes, such as SNAC1 (Hu et al. 2006), OsNAC6/SNAC2 (Nakashima et al. 2007), OsNAC5 (Takasaki et al. 2010), ONAC045 (Zheng et al. 2009), were also reported to be involved in abiotic stress responses. Overexpression of SNAC1 significantly improved drought and salt tolerance in transgenic rice with no adverse effects on grain yield (Hu et al. 2006). OsNAC5 was induced by various abiotic and hormone treatments such as drought, cold, salinity, ABA, and methyl jasmonate (MeJA). Overexpression of OsNAC5 in transgenic plants showed improved tolerance to high salinity with no adverse effects on plant growth in normal conditions, indicative of potential use to improve stress tolerance in molecular breeding (Takasaki et al. 2010). Recently, stress-responsive NAC genes were also reported in wheat. Overexpression of stress-responsive NAC genes TaNAC67 and TaNAC69 from wheat resulted in enhanced tolerance to drought and salinity stresses (Mao et al. 2014; Xue et al. 2011).
Plants have evolved a complex signaling network to adapt to adverse conditions. There is overwhelming evidence that extensive crosstalk and interactions exist in abiotic stress and hormone signaling (Fujita et al. 2006; Nakashima et al. 2012; Narusaka et al. 2004; Nuruzzaman et al. 2013). As one of the first reported NAC genes, ATAF1 was shown to be involved in both abiotic and biotic stress responses. The expression of ATAF1 was induced by wounding, pathogen attack, drought, and ABA (Jensen et al. 2008; Lu et al. 2007; Wu et al. 2009). Overexpression of ATAF1 enhanced the tolerance to drought stress as well as increased resistance to pathogen attack in transgenic Arabidopsis (Wu et al. 2009), while the penetration resistance upon pathogen attack was reduced in ataf1 mutant (Jensen et al. 2008). OsNAC6/SNAC2, the closest ortholog of ATAF1 in rice, was induced by both biotic and abiotic stresses. Overexpression of OsNAC6/SNAC2 in rice led to enhanced tolerances to abiotic stress including drought, salinity and cold, as well as increased resistance to blast disease (Hu et al. 2008; Nakashima et al. 2007). RD26/ANAC072, a dehydration-responsive NAC gene in Arabidopsis, was induced by ABA, MeJA, salicylic acid (SA), as well as by salt and drought stresses, which suggests it might play a role in the regulation of both abiotic and hormone signaling (Fujita et al. 2004).
Miscanthus lutarioriparius, a perennial rhizomatous grass, is one of Miscanthus species native to East Asia (Chen and Renvoize 2006). Recently, it has attracted much attention as a bioenergy crop because of its high photosynthetic efficiency and biomass productivity, as well as wide adaptability to different environments (Brosse et al. 2012). Genomic and small RNA sequencing of Miscanthus × giganteus (Mxg) demonstrated that the coding fraction of the Mxg genome shares a high level of sequence identity with Sorghum (Sorghum bicolor), thus making the sorghum genome as a promising reference for Miscanthus genus (Swaminathan et al. 2010).
The studies on NAC genes were mainly carried out in model plant species Arabidopsis and rice, there are no reports on M. lutarioriparius NAC transcription factors to date. In the present study, expression profiles of 13 NAC genes from M. lutarioriparius in different tissues and under various abiotic stresses and hormone treatments were analyzed. The crosstalk among different abiotic stresses and hormone treatments was also discussed. This study may lay a foundation for further improvement of stress tolerance in Miscanthus by genetic engineering.
Materials and methods
Plant materials
The present study was conducted on M. lutarioriparius, a homogenous diploid (2n = 2x = 38) species represented by the genotype from the Shanghai Institute for Biological Sciences of the Chinese Academy of Sciences. New plants were propagated via rhizomes in a growth chamber under 16-h light/8-h dark photoperiod at 28 °C with relative humidity of 65 %. Up to the six-leaf stage, the root, rhizome, mature stem, young stem, leaf, and sheath were collected and divided separately to investigate the tissue-specific expression patterns.
Arabidopsis was grown in a controlled growth chamber at 22 °C with 60 % relative humidity under 16-h light/8-h dark photoperiod.
Stress treatment
The artificial stress treatments were described as following on the rhizome propagated plants at six-leaf stage (Supplemental Fig. 1). For drought stress treatment, plants were first gently removed from the soil, followed by washing out of the soil, and then dehydrated on filter paper at room temperature at 65 % relative humidity. For treatment of salt stress, the similar plant removing and root washing operations were conducted, and then followed by soaking in 150-mM NaCl solution. For cold stress, plants were transferred to a cold chamber with light at 4 °C for continuing growth. Plants sprayed with 100-μM ABA, 100-μM MeJA and 1.5-mM SA dissolved in 0.1 % (v/v) ethanol were set as the hormone treatments, while 0.1 % (v/v) ethanol was considered as the control treatment. Wounding treatment was achieved by cutting leaves with clean scissors. At 0, 1, 3, 6, 12, and 24 h after treatment conducted, the second leaves from the top were collected, immediately frozen in liquid nitrogen and then stored at −80 °C.
Isolation of MlNAC cDNAs
NAC domain proteins from Arabidopsis thaliana and Oryza sativa L., characterized as being involved in stress responses, were used as queries for orthologous genes in Sorghum using the Phytozome database (http://www.phytozome.net). Based on the orthologous genes identified in Sorghum, specific primers were designed to isolate the full-length MlNAC cDNAs with the primer sequences shown in Table 1.
Total RNA was isolated using the CTAB method with minor modifications (Chang et al. 1993). RNA integrity and purity was monitored by 2 % agarose gel electrophoresis and ethidium bromide staining. Reverse transcription was performed with approximately 2 μg RNA that was predigested with DNase I (Promega, Madison, WI, USA) using the RevertAid First Strand cDNA Synthesis Kit (MBI, Fermentas) according to the manufacturer’s instructions.
The full open reading frame (ORF) of MlNAC genes was amplified by PCR using the cDNA mentioned above as template. Gel-purified PCR products were cloned into the pMD19-T vector, and then sequenced by Shanghai Sunny Biotechnology Co, Ltd.
Sequence alignment and phylogenetic analysis
Multiple sequence alignment of protein sequences was performed by the Clustal X (version 2.0) program and the BioEdit program (version 7.0). The unrooted phylogenetic tree was constructed with MEGA 5.0 using Neighbor-Joining (NJ), and the bootstrap test was carried out with 1,000 replicates (Tamura et al. 2011). The conserved motifs in the MlNAC protein sequences were identified using multiple expectation maximization for motif elicitation (MEME) (http://www.meme.sdsc.edu) (Bailey et al. 2006). The analysis parameters were set as following: number of repetitions, any; maximum number of motifs, 20; and the optimum motif widths were constrained to between 6 and 200 residues. Structural motif annotation was performed using the Pfam (http://www.pfam.sanger.ac.uk) and SMART (http://www.smart.embl-heidelberg.de) databases (Finn et al. 2006; Letunic et al. 2004).
Quantitative real-time RT-PCR (RT-qPCR) analysis
Primers for RT-qPCR reactions were designed by Beacon Designer v7.0 (Premier Biosoft International, Palo Alto, California, USA) with optimal melting temperatures of 58–60 °C, primer lengths of 20–24 bp and amplicon lengths of 100–200 bp (Supplemental Table 2). RT-qPCR was performed in triplicate with a LightCycler® 480 Detection System (Roche, Germany) using SYBR Premix ExTaq (TaKaRa, Toyoto, Japan). Each 20-μl PCR reaction contained 1 μl cDNA, 10 μl of 2 × SYBR Premix, 200 nM of each primer. The PCR amplification was conducted with an initial denaturation of 95 °C for 10 s followed by 40 cycles of 95 °C for 5 s, 60 °C for 1 min. Melting curve analysis was performed at the end of each PCR run (60–95 °C, 0.06 °C/s). Baseline and threshold cycles (Ct) were analyzed by LightCycler® 480 Software (release 1.5.0.). Relative gene expressions were calculated with respect to internal reference gene of MlACT11 using the relative 2−∆∆Ct method (Livak and Schmittgen 2001).
Generation of transgenic Arabidopsis plants
The sequencing confirmed open reading frame (ORF) of MlNAC2 was cloned into the pEarleyGate100 plasmid, which was controlled by a Cauliflower mosaic virus 35S (CaMv 35S) promoter (Earley et al. 2006). The constructs were introduced into Agrobacterium tumefaciens GV3101 by electroporation, and then transformed into Arabidopsis Col-0 through the floral dip method (Clough and Bent 1998). Seeds of the T 0 and T 1 generation were sown on MS agar medium plates containing glufosinate (50 mg/l, Sigma-Aldrich). Homozygous T 3 generation transgenic lines were harvested individually for further analysis.
Dehydration assay
For drought treatment, sterilized transgenic Arabidopsis seeds were sown on vertically oriented ½ MS medium plates to grow for 10 days, and then transferred to soil to grown for 2 weeks. Overexpression lines and WT plants were dehydrated in air for 60 min and photographed at 0, 30 and 60 min, respectively. After drought stress, seedlings were re-watered, and the recovery rates were counted after 2 h. Drought stress treatment was conducted at conditions of 22 °C with 60 % humility. The experiment was repeated three times, and 50 plants were used for each replication.
Water loss measurement
Five leaves per individual of OX-5, OX-9 and WT plants grown in soil for 2 weeks at normal conditions (22 °C, 60 % humility, 16-h light/8-h dark photoperiod) were excised, and fresh weight was determined at designated time intervals to determine water loss rates. Three replications were performed and the means were calculated.
Statistical analysis
Statistical testing was performed with SPSS software (V13.0, SPSS Inc. USA).
Results
Isolation of MlNAC cDNAs and sequence analysis
These 13 MlNAC genes were designated as MlNAC1 to MlNAC13 (Table 1). With the exception of MlNAC3 and MlNAC6, all full-length cDNAs were obtained. The sequences of these 11 full-length MlNACs have been submitted to GeneBank under the accession numbers KM017001 to KM017011. Sequence alignment showed that the N-terminal region of NAC proteins contained the highly conserved NAC domain (Fig. 1), which was further divided into five subdomains (A-E) according to the previous study (Ooka et al. 2003).
Sequence alignment of 11 full-length MlNACs with NAP, NAM, NAC1, ATAF1, and ATAF2 from Arabidopsis. The five conserved subdomains (a–e) of NAC are indicated by lines above the sequence. Note that the full-length coding sequences of MlNAC3 and MlNAC6 were not obtained and were excluded in sequence alignment
Phylogenetic analysis
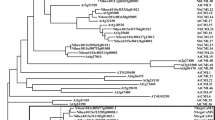
To investigate the phylogenetic relationship between the MlNAC proteins and other NAC proteins, an unrooted phylogenetic tree was constructed from alignments of the full-length protein sequences from 11 MlNACs and 23 orthologous NACs from A. thaliana and rice by MEGA 5.0 using the Neighbor-Joining (NJ) method (Fig. 2 and Supplemental Table 3). As shown in Fig. 2a, 34 NACs were divided into six distinct subfamilies. As the NAC members exhibited an interspersed distribution in the majority of subfamilies, it could be inferred that the expansion of NAC genes occurred before the divergence of Miscanthus, Arabidopsis, and rice. MlNAC10 was distinguished from other NAC members and formed an individual clade with a robust bootstrap value support (95 %). In addition, the expressions of 13 MlNAC genes under different abiotic stresses conducted in this study (see below) and the stress-inducible NAC genes reported in previous studies were listed for a comparison. As shown in Fig. 2b, NAC genes clustered into the same subgroup exhibited similar expression patterns under abiotic stress treatments. For example, the members in subgroup I, including ATAF1, OsNAC5, OsNAC6/SNAC2, MlNAC1 and MlNAC5 were all induced by salt, drought stresses and hormone (ABA and MeJA) treatments.
Phylogenetic relationship and expression profiles of MlNACs and their homologs from Arabidopsis and rice. a Phylogenetic relationship of NAC in Miscanthus lutarioriparius, Arabidopsis thaliana and rice. The six subfamilies are shown in different color rectangles. The phylogenetic tree was generated using MEGA5.0 from 1,000 bootstrap replicates. b Expression patterns of NAC genes under different stress conditions. Black dot indicates an induction. Members of NAC protein from M. lutarioriparius were denoted in red
Conserved motifs in MlNAC proteins
To gain further insight into the diversification of NAC genes in M. lutarioriparius, putative motifs were predicted by the MEME program and 20 distinct motifs were identified (Fig. 3). However, most of the putative motifs remain to be annotated since no homologs were found when searching against Pfam and SMART databases (Finn et al. 2006; Letunic et al. 2004). The details of the 20 putative motifs are available in Supplemental Fig. 2. In this study, motif 3, 8, 1, 2, and 10 specified the NAC subdomain A to E, respectively. But among the MlNACs surveyed, only MlNAC1, MlNAC2, MlNAC4, MlNAC5, and MlNAC12 had these five motifs in the typical NAC DNA-binding domains. As expected, closely related genes in the phylogenetic tree had common motif compositions as expected (Fig. 3), indicating the functional similarity among NAC proteins within the same subfamily. However, specific motifs were identified for each subfamily, e.g., motif 14 for subfamily I, motifs 7, 9, and 12 for subfamily II, and motif 6 for subfamily VI. These specific motifs may confer a unique functional role to MlNAC genes. In spite of the high divergence in C-terminal regions of MlNACs, at least eight conserved motifs were still identified.
Tissue-specific expression patterns of MlNAC genes
We investigated the expression patterns of MlNAC genes in root, rhizome, mature stem, young stem, leaf, and sheath of M. lutarioriparius under normal conditions using RT-qPCR analysis (Fig. 4). According to the data, the MlNAC genes displayed distinct tissue-specific patterns in different tissues examined. For example, MlNAC1, MlNAC5 and MlNAC13 genes were predominantly expressed in rhizome, whereas MlNAC6 and MlNAC8 genes were weakly expressed. MlNAC2, MlNAC9 and MlNAC12 genes were exclusively expressed at higher levels in leaf and sheath. MlNAC11 was more highly expressed in root, rhizome, and sheath compared to mature stem, young stem and leaf. Furthermore, all MlNAC genes except MlNAC10 exhibited higher expression in mature stem than in young stem.
Expression profiles of MlNAC genes in six different tissues by RT-qPCR analysis. The normalized relative quantity of root was set up as “1”. The y-axis demonstrates the relative fold-change of gene expression compared with root. The x-axis indicates different tissues, Ro root, Rh rhizome, MS mature stem, YS young stem, L leaf, S sheath
Responses of MlNAC genes to abiotic stresses and hormone treatments
RT-qPCR analysis was further performed to detect the expression patterns of 13 MlNAC genes under abiotic stress and hormone treatments. We considered those genes with an expression ratio of threefold higher than the untreated control for at least one time point in each stress treatment as stress-inducible genes (Teige et al. 2004). All MlNAC genes examined were up-regulated by salt, drought, MeJA, and SA treatments. The transcript abundance of four genes (MlNAC1, MlNAC2, MlNAC4, MlNAC12) and three genes (MlNAC1, MlNAC11, MlNAC12) exhibited more than 100-fold increases following salt and drought treatments, respectively (Figs. 5, 6). Seven MlNACs (MlNAC1, 4, 5, 10–13) were up-regulated following cold treatment (Fig. 7), among which, MlNAC11 and MlNAC13 were the most dramatically regulated with more than tenfold increase after treatment for 24 h, following by MlNAC10 with about a 16-fold increase. In contrast, the expression of MlNAC3, MlNAC7 and MlNAC9 was down-regulated in cold treatment. Eleven MlNAC genes were slightly up-regulated following ABA treatment, among which, eight genes increased more than sixfold, whereas MlNAC4 and MlNAC10 were down-regulated (Fig. 8). The expression of MlNAC2, MlNAC4 and MlNAC12 was at least 100-fold increased after MeJA treatment. All MlNAC genes surveyed were up-regulated by MeJA as well as SA treatments (Figs. 9, 10), suggesting a positive interaction between MeJA and SA, which will be further discussed in the following section. The expressions of all MlNAC genes surveyed except MlNAC11 showed significant increases in wounding treatment (Fig. 11).
RT-qPCR analysis of MlNACs’ expression under salt stress. The normalized relative quantity of 0 h was set up as “1”. The y-axis demonstrates the relative fold-change of gene expression compared with 0 h. The x-axis indicates treatment time, 0, 1, 3, 6, 12, and 24 h. Data represent means of three replicates. Error bars represent standard deviation. The statistical significance of gene expression was evaluated using Student’s t test. The asterisks indicated significant differences (**P < 0.01, and *P < 0.05)
Expression profiles of MlNAC genes under drought stress by RT-qPCR analysis. The normalized relative quantity of 0 h was set up as “1”. The y-axis demonstrates the relative fold-change of gene expression compared with 0 h. The x-axis indicates treatment time, 0, 1, 3, 6, 12, and 24 h. Data represent means of three replicates. Error bars represent standard deviation. The statistical significance of gene expression was evaluated using Student’s t test. The asterisks indicated significant differences (**P < 0.01, and *P < 0.05)
RT-qPCR analysis of MlNACs expression under cold stress. The normalized relative quantity of 0 h was set up as “1”. The y-axis demonstrates the relative fold-change of gene expression compared with 0 h. The x-axis indicates treatment time, 0, 1, 3, 6, 12, and 24 h. Data represent means of three replicates. Error bars represent standard deviation. The statistical significance of gene expression was evaluated using Student’s t test. The asterisks indicated significant differences (**P < 0.01, and *P < 0.05)
Expression profiles of MlNAC genes under ABA application by RT-qPCR analysis. The normalized relative quantity of 0 h was set up as “1”. The y-axis demonstrates the relative fold-change of gene expression compared with 0 h. The x-axis indicates treatment time, 0, 1, 3, 6, 12 and 24 h. Data represent means of three replicates. Error bars represent standard deviation. The statistical significance of gene expression was evaluated using Student’s t test. The asterisks indicated significant differences (**P < 0.01, and *P < 0.05)
RT-qPCR analysis of MlNACs’ expression under MeJA application. The normalized relative quantity of 0 h was set up as “1”. The y-axis demonstrates the relative fold-change of gene expression compared with 0 h. The x-axis indicates treatment time, 0, 1, 3, 6, 12 and 24 h. Data represent means of three replicates. Error bars represent standard deviation. The statistical significance of gene expression was evaluated using Student’s t test. The asterisks indicated significant differences (**P < 0.01, and *P < 0.05)
Expression profiles of MlNAC genes under SA application by RT-qPCR analysis. The normalized relative quantity of 0 h was set up as “1”. The y-axis demonstrates the relative fold-change of gene expression compared with 0 h. The x-axis indicates treatment time, 0, 1, 3, 6, 12 and 24 h. Data represent means of three replicates. Error bars represent standard deviation. The statistical significance of gene expression was evaluated using Student’s t test. The asterisks indicated significant differences (**P < 0.01, and *P < 0.05)
RT-qPCR analysis of MlNACs’ expression under wounding stress. The normalized relative quantity of 0 h was set up as “1”. The y-axis demonstrates the relative fold-change of gene expression compared with 0 h. The x-axis indicates treatment time, 0, 1, 3, 6, 12 and 24 h. Data represent means of three replicates. Error bars represent standard deviation. The statistical significance of gene expression was evaluated using Student’s t test. The asterisks indicated significant differences (**P < 0.01, and *P < 0.05)
Crosstalk between different stresses
Venn diagrams were drawn to indicate the possible crosstalk between different stresses surveyed (Fig. 12). Stress-inducible genes were classified into groups on the basis of their expression patterns. Results of the present study support the viewpoint that extensive crosstalk between the abiotic stress and hormone stress-signaling exists. We identified 13, 13, 7, and 11 MlNAC genes as salt-, drought-, cold-, and ABA-inducible genes, respectively. Analysis of overlapping on the Venn diagram displayed that all 13 MlNAC genes were induced by both salt and drought treatments. Eleven MlNAC genes, accounting for 85 % of the genes tested, were identified as drought- and ABA-inducible genes. Seven (54 %) MlNAC genes were induced by both cold and drought treatments and five (38 %) MlNAC genes responded to both cold and ABA treatments. All MlNACs surveyed were up-regulated under MeJA, SA and ABA treatments except MlNAC4 and MlNAC10.
Venn diagrams showing the classification of genes up-regulated by salt, drought and cold stresses and ABA, MeJA and SA application identified on the basis of RT-qPCR analyses. The number of genes classified in each region is indicated. a Intersection of genes that were up-regulated by salt, drought, cold stresses and ABA application. b Intersection of genes that were up-regulated by ABA application with those that were either up-regulated by MeJA and SA application
Performance of MlNAC2 overexpression plants in transgenic Arabidopsis under dehydration treatment
To further characterize the roles of MlNAC genes in abiotic stress, we selected MlNAC2 for further functional analysis in transgenic Arabidopsis. In our study, we showed that MlNAC2 was induced by various abiotic stress treatments (Figs. 5, 6, 7, 8, 9, 10, 11), so we are prompting to see if the overexpression of MlNAC2 in transgenic Arabidopsis can enhance plant tolerance to abiotic stresses. The expression levels of MlNAC2-overexpressing lines were examined by RT-qPCR (Fig. 13a). As the expression levels of MlNAC2 in OX-5 and OX-9 were relatively higher than the others, these two lines were selected for the dehydration assay to examine their drought tolerance. To assess the water retention ability, two transgenic lines (OX-5 and OX-9) and WT were analyzed for water loss rates during a 60-min dehydration. According to the data shown in Fig. 13b, OX-5 and OX-9 plants showed significantly lower water loss rates than WT under the dehydration stress. In addition, the time when WT began to wither and exhibited a dark green color was earlier than the transgenic plants. After the dehydration treatment for 60 min, both OX-5 and OX-9 plants were only slightly withered, while the leaves of WT were already severely curled and withered. After re-watering for 2 h, 52 % (26/50) to 56 % (28/50) of transgenic plants recovered from the dehydration stress compared to 34 % (17/50) of the WT controls (Fig. 13c). These data suggested that MlNAC2 might play a positive role in drought stress tolerance.
Dehydration assays of MlNAC2-overexpressing plants. a Expression levels of MlNAC2 in different transgenic Arabidopsis lines. Gene expression levels of MlNAC2 in different transgenic lines were determined by RT-qPCR. OX2, OX-5, OX-8, OX-9 and OX-12, MlNAC2 transgenic lines. The expression of MlNAC2 in OX-2 was regarded as a standard for its lower expression level. b Water loss from detached leaves of OX-5, OX-9 and WT plants. Each data point represents the mean of three independent experiments. The difference of the MlNAC2 overexpression plants to WT was evaluated using Student’s t test. The asterisks indicated significant differences (*P < 0.05). c Phenotypes of OX-5, OX-9 and WT under dehydration stress
Discussion
In this study, we identified 13 stress-responsive NAC genes from M. lutarioriparius, a potential bioenergy crop. Sequence alignment indicated that MlNAC proteins have conserved NAC domains which were further divided into five subdomains (A–E) as previously reported (Ernst et al. 2004; Ooka et al. 2003). Subdomains C and D are involved in DNA binding, subdomain A plays a role in the formation of a functional dimer and subdomains B and E might contribute to the functional diversity of NAC genes (Chen et al. 2011; Duval et al. 2002; Ernst et al. 2004; Kikuchi et al. 2000; Ooka et al. 2003). Additionally, the C-terminal transcription activation region was extremely diversified, which was reported to serve as either activator or repressor and may sometimes possess protein-binding activity (Chen et al. 2011; Ernst et al. 2004; Ooka et al. 2003).
NAC genes with similar function showed a tendency to fall into one subgroup. For instance, in subfamily I, two of the well-characterized members ATAF1 and OsNAC6/SNAC2 were shown to play important roles in both biotic and abiotic stresses (Hu et al. 2008; Lu et al. 2007; Nakashima et al. 2007; Wu et al. 2009). Subfamily II was a monocot-specific subfamily, which might play a role in monocot-specific responses to stress (Ooka et al. 2003). Subfamily III contained NAC proteins that function in the regulation of flowering time, cold stress response (e.g., ANAC035) (Yoo et al. 2007) and salt stress response (e.g., ONAC063) (Yokotani et al. 2009). In subfamily IV, only NAC1 from rice was well characterized, which was shown to promote lateral root development (Guo et al. 2005).
NAC genes are involved in diverse processes of plant growth and development. The molecular properties of eight NAC genes in rice (OsNAC1 to OsNAC8) were investigated, and each of these genes has a unique tissue-specific expression pattern (Kikuchi et al. 2000). In this study, the majority of MlNAC genes showed distinct tissue-specific expression patterns, suggesting they probably play diverse functional roles in M. lutarioriparius development. In contrast, the expression of MlNAC3, MlNAC7, MlNAC10 and MlNAC13 was ubiquitous in the tissues examined, which indicated that they might participate in various developmental processes in M. lutarioriparius.
Previous studies demonstrated that NAC genes function at the convergence point of abiotic and biotic stress signaling (reviewed by Nuruzzaman et al. 2013). For example, the expression of ATAF1 was induced by drought, salt, ABA, MeJA, SA and pathogen infection (Jensen et al. 2008; Lu et al. 2007; Wu et al. 2009). In rice, the closest ortholog of ATAF1, OsNAC6/SNAC2, was induced by drought, salt, cold, ABA and MeJA treatments (Hu et al. 2008; Nakashima et al. 2007). Overexpression of ATAF1 and OsNAC6/SNAC2 led to enhanced tolerance to drought and salinity stresses as well as increased disease resistance (Hu et al. 2008; Jensen et al. 2008; Lu et al. 2007; Nakashima et al. 2007; Wu et al. 2009), which indicated that ATAF1 and OsNAC6/SNAC2 play dual roles in both abiotic and biotic stresses. In the present study, all MlNACs surveyed were induced by abiotic stress and hormone treatments. As shown in Fig. 12a, there seemed to be more extensive crosstalk between salt, drought and ABA stress-signaling processes than between drought and cold stress or ABA and cold stress-signaling processes. Additionally, all ABA-regulated MlNAC genes were also regulated by salt, drought or cold treatments in the present study, which indicated that MlNACs might participate in abiotic stress signaling through an ABA-dependent pathway.
As previous reports suggested, ABA, MeJA and SA are involved in the NAC-mediated hormone-signaling pathway (Takasaki et al. 2010; Wang et al. 2009; Wu et al. 2009). Therefore, we analyzed expression patterns of MlNAC genes under treatments with these three hormones. The result showed that all 13 MlNAC genes were up-regulated by MeJA and SA treatments and all MlNAC genes except MlNAC4 and MlNAC10 were up-regulated by ABA treatment. These results indicate that MlNAC genes may play a role in hormone-signaling pathways. On the other hand, accumulating evidence demonstrated that NAC genes might play a positive or negative role in plant disease resistance. For example, overexpression of ATAF1 resulted in enhanced drought and salt tolerance as well as susceptibility to pathogens infection in Arabidopsis (Wu et al. 2009). Transgenic rice plants overexpressing OsNAC6/SNAC2 exhibited enhanced tolerance to drought, salt and cold stresses as well as increased resistance to blast disease (Hu et al. 2008; Nakashima et al. 2007). The plant hormones MeJA and SA are the major molecules participating in defense responses in plants (Anderson et al. 2004). Our data showed that all MlNAC genes surveyed were up-regulated by MeJA and SA, implying the potential role of Miscanthus NAC genes in plant defense responses.
Phylogenetic analysis revealed that 13 MlNAC proteins were classified into six subfamilies. MlNAC2 exhibited the highest (80 %) sequence identity to SNAC1, which belongs to subfamily II. In this subfamily, OsNAC3 and OsNAC4 and SNAC1 were strongly induced by various abiotic stress and hormone treatments (Fig. 2b). Overexpression of SNAC1 in rice significantly enhanced plant drought and salt tolerance, while exerted no adverse effects on plant growth and grain yield in normal conditions (Hu et al. 2006). In our study, we showed that the expression of MlNAC2 was up-regulated by various abiotic stress treatments (Figs. 5, 6, 7, 8, 9, 10, 11). Consequently, we further studied the functional roles of MlNAC2 in abiotic stress response in transgenic Arabidopsis. Not unexpectedly, overexpression of MlNAC2 in Arabidopsis led to enhanced drought tolerance. Both morphological and physiological data demonstrated that the transgenic lines, OX-5 and OX-9, were more tolerant to drought than WT plants. Since the expression of MlNAC2 was significantly up-regulated by ABA treatment, which suggested that MlNAC2 might participate in an ABA-dependent signaling pathway in response to drought stress. Moreover, the growth of transgenic plants overexpressing MlNAC2 was similar to that of WT, despite previous reports that overexpression of several NAC genes has adverse effects on plant growth and development (Fujita et al. 2004; Takasaki et al. 2010; Wu et al. 2009). Our study indicated that MlNAC2 might have the potential for improving the drought tolerance of crops.
In addition, the expression of four genes, MlNAC1, MlNAC5, MlNAC12 and MlNAC13, was induced by all seven treatments examined in the present study, indicating their potential for utilization in genetic manipulation to improve abiotic stress tolerance of Miscanthus.
Conclusion
In summary, 13 MlNAC genes were identified in this study. Based on gene expression analyses, these MlNAC genes were shown to participate in diverse abiotic stress responses in M. lutarioriparius. Furthermore, substantial crosstalk between different stresses was also discovered in M. lutarioriparius. Overexpression MlNAC2 in Arabidopsis conferred increased tolerance to drought stress. Future studies should focus on the functional annotation of these MlNAC genes to reveal whether they have the potential to improve stress tolerance of Miscanthus.
References
Aida M, Ishida T, Fukaki H, Fujisawa H, Tasaka M (1997) Genes involved in organ separation in Arabidopsis: an analysis of the cup-shaped cotyledon mutant. Plant Cell 9:841–857
Anderson JP, Badruzsaufari E, Schenk PM, Manners JM, Desmond OJ, Ehlert C, Maclean DJ, Ebert PR, Kazan K (2004) Antagonistic interaction between abscisic acid and jasmonate-ethylene signaling pathways modulates defense gene expression and disease resistance in Arabidopsis. Plant Cell 16:3460–3479
Bailey TL, Williams N, Misleh C, Li WW (2006) MEME: discovering and analyzing DNA and protein sequence motifs. Nucleic Acids Res 34:W369–W373
Brosse N, Dufour A, Meng XZ, Sun QN, Ragauskas A (2012) Miscanthus: a fast-growing crop for biofuels and chemicals production. Biofuels, Bioprod Bioref 6:580–598
Chang SJ, Puryear J, Cairney J (1993) A simple and efficient method for isolating RNA from pine trees. Plant Mol Biol Report 11:113–116
Chen SL, Renvoize SA (2006) Miscanthus. Flora of China. Science Press, Beijing, pp 581–583
Chen QF, Wang Q, Xiong LZ, Lou ZY (2011) A structural view of the conserved domain of rice stress-responsive NAC1. Protein Cell 2:55–63
Christianson JA, Dennis ES, Llewellyn DJ, Wilson IW (2010) ATAF NAC transcription factors: regulators of plant stress signaling. Plant Signal Behav 5:428–432
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
Delessert C, Kazan K, Wilson IW, Van Der Straeten D, Manners J, Dennis ES, Dolferus R (2005) The transcription factor ATAF2 represses the expression of pathogenesis-related genes in Arabidopsis. Plant J 43:745–757
Duval M, Hsieh TF, Kim SY, Thomas TL (2002) Molecular characterization of AtNAM: a member of the Arabidopsis NAC domain superfamily. Plant Mol Biol 50:237–248
Earley KW, Haag JR, Pontes O, Opper K, Juehne T, Song K, Pikaard CS (2006) Gateway-compatible vectors for plant functional genomics and proteomics. Plant J 45:616–629
Ernst HA, Olsen AN, Larsen S, Lo Leggio L (2004) Structure of the conserved domain of ANAC, a member of the NAC family of transcription factors. EMBO Rep 5:297–303
Finn RD, Mistry J, Schuster-Bockler B, Griffiths-Jones S, Hollich V, Lassmann T, Moxon S, Marshall M, Khanna A, Durbin R, Eddy SR, Sonnhammer ELL, Bateman A (2006) Pfam: clans, web tools and services. Nucleic Acids Res 34:D247–D251
Fujita M, Fujita Y, Maruyama K, Seki M, Hiratsu K, Ohme-Takagi M, Tran LS, Yamaguchi-Shinozaki K, Shinozaki K (2004) A dehydration-induced NAC protein, RD26, is involved in a novel ABA-dependent stress-signaling pathway. Plant J 39:863–876
Fujita M, Fujita Y, Noutoshi Y, Takahashi F, Narusaka Y, Yamaguchi-Shinozaki K, Shinozaki K (2006) Crosstalk between abiotic and biotic stress responses: a current view from the points of convergence in the stress signaling networks. Curr Opin Plant Biol 9:436–442
Guo YF, Gan SS (2006) AtNAP, a NAC family transcription factor, has an important role in leaf senescence. Plant J 46:601–612
Guo HS, Xie Q, Fei JF, Chua NH (2005) MicroRNA directs mRNA cleavage of the transcription factor NAC1 to downregulate auxin signals for Arabidopsis lateral root development. Plant Cell 17:1376–1386
He XJ, Mu RL, Cao WH, Zhang ZG, Zhang JS, Chen SY (2005) AtNAC2, a transcription factor downstream of ethylene and auxin signaling pathways, is involved in salt stress response and lateral root development. Plant J 44:903–916
Hibara K, Takada S, Tasaka M (2003) CUC1 gene activates the expression of SAM-related genes to induce adventitious shoot formation. Plant J 36:687–696
Hu HH, Dai MQ, Yao JL, Xiao BZ, Li X, Zhang QF, Xiong LZ (2006) Overexpressing a NAM, ATAF, and CUC (NAC) transcription factor enhances drought resistance and salt tolerance in rice. Proc Natl Acad Sci USA 103:12987–12992
Hu HH, You J, Fang YJ, Zhu XY, Qi ZY, Xiong LZ (2008) Characterization of transcription factor gene SNAC2 conferring cold and salt tolerance in rice. Plant Mol Biol 67:169–181
Hu RB, Qi G, Kong YZ, Kong DJ, Gao Q, Zhou GK (2010) Comprehensive analysis of NAC domain transcription factor gene family in Populus trichocarpa. BMC Plant Biol 10:145
Jaglo-Ottosen KR, Gilmour SJ, Zarka DG, Schabenberger O, Thomashow MF (1998) Arabidopsis CBF1 overexpression induces COR genes and enhances freezing tolerance. Science 280:104–106
Jensen MK, Hagedorn PH, de Torres-Zabala M, Grant MR, Rung JH, Collinge DB, Lyngkjaer MF (2008) Transcriptional regulation by an NAC (NAM-ATAF1,2-CUC2) transcription factor attenuates ABA signalling for efficient basal defence towards Blumeria graminis f. sp. hordei in Arabidopsis. Plant J 56:867–880
Jung C, Seo JS, Han SW, Koo YJ, Kim CH, Song SI, Nahm BH, Choi YD, Cheong JJ (2008) Overexpression of AtMYB44 enhances stomatal closure to confer abiotic stress tolerance in transgenic Arabidopsis. Plant Physiol 146:623–635
Kikuchi K, Ueguchi-Tanaka M, Yoshida KT, Nagato Y, Matsusoka M, Hirano HY (2000) Molecular analysis of the NAC gene family in rice. Mol Genet Genomics 262:1047–1051
Le DT, Nishiyama R, Watanabe Y, Mochida K, Yamaguchi-Shinozaki K, Shinozaki K, Tran LS (2011) Genome-wide survey and expression analysis of the plant-specific NAC transcription factor family in soybean during development and dehydration stress. DNA Res 18:263–276
Letunic I, Copley RR, Schmidt S, Ciccarelli FD, Doerks T, Schultz J, Ponting CP, Bork P (2004) SMART 4.0: towards genomic data integration. Nucleic Acids Res 32:D142–D144
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-delta delta C(T)) Method. Methods 25:402–408
Lu PL, Chen NZ, An R, Su Z, Qi BS, Ren F, Chen J, Wang XC (2007) A novel drought-inducible gene, ATAF1, encodes a NAC family protein that negatively regulates the expression of stress-responsive genes in Arabidopsis. Plant Mol Biol 63:289–305
Lu GJ, Gao CX, Zheng XN, Han B (2009) Identification of OsbZIP72 as a positive regulator of ABA response and drought tolerance in rice. Planta 229:605–615
Mao XG, Chen SS, Li A, Zhai CC, Jing RL (2014) Novel NAC transcription factor TaNAC67 confers enhanced multi-abiotic stress tolerances in Arabidopsis. PLoS ONE 9:e84359
Mitsuda N, Seki M, Shinozaki K, Ohme-Takagi M (2005) The NAC transcription factors NST1 and NST2 of Arabidopsis regulate secondary wall thickenings and are required for anther dehiscence. Plant Cell 17:2993–3006
Nakashima K, Tran LS, Van Nguyen D, Fujita M, Maruyama K, Todaka D, Ito Y, Hayashi N, Shinozaki K, Yamaguchi-Shinozaki K (2007) Functional analysis of a NAC-type transcription factor OsNAC6 involved in abiotic and biotic stress-responsive gene expression in rice. Plant J 51:617–630
Nakashima K, Takasaki H, Mizoi J, Shinozaki K, Yamaguchi-Shinozaki K (2012) NAC transcription factors in plant abiotic stress responses. Biochim Biophys Acta 1819:97–103
Narusaka Y, Narusaka M, Seki M, Umezawa T, Ishida J, Nakajima M, Enju A, Shinozaki K (2004) Crosstalk in the responses to abiotic and biotic stresses in Arabidopsis: analysis of gene expression in cytochrome P450 gene superfamily by cDNA microarray. Plant Mol Biol 55:327–342
Nikovics K, Blein T, Peaucelle A, Ishida T, Morin H, Aida M, Laufs P (2006) The balance between the MIR164A and CUC2 genes controls leaf margin serration in Arabidopsis. Plant Cell 18:2929–2945
Nuruzzaman M, Manimekalai R, Sharoni AM, Satoh K, Kondoh H, Ooka H, Kikuchi S (2010) Genome-wide analysis of NAC transcription factor family in rice. Gene 465:30–44
Nuruzzaman M, Sharoni AM, Kikuchi S (2013) Roles of NAC transcription factors in the regulation of biotic and abiotic stress responses in plants. Front Microbiol 4:248
Ooka H, Satoh K, Doi K, Nagata T, Otomo Y, Murakami K, Matsubara K, Osato N, Kawai J, Carninci P, Hayashizaki Y, Suzuki K, Kojima K, Takahara Y, Yamamoto K, Kikuchi S (2003) Comprehensive analysis of NAC family genes in Oryza sativa and Arabidopsis thaliana. DNA Res 10:239–247
Puranik S, Sahu PP, Mandal SN, VS B, Parida SK, Prasad M (2013) Comprehensive genome-wide survey, genomic constitution and expression profiling of the NAC transcription factor family in foxtail millet (Setaria italica L.). PLoS ONE 8:e64594. doi:10.1371/journal.pone.0064594
Shinozaki K, Yamaguchi-Shinozaki K (1997) Gene expression and signal transduction in water-stress response. Plant Physiol 115:327–334
Singh AK, Sharma V, Pal AK, Acharya V, Ahuja PS (2013) Genome-wide organization and expression profiling of the NAC transcription factor family in potato (Solanum tuberosum L.). DNA Res 20:403–423
Swaminathan K, Alabady MS, Varala K, De Paoli E, Ho I, Rokhsar DS, Arumuganathan AK, Ming R, Green PJ, Meyers BC, Moose SP, Hudson ME (2010) Genomic and small RNA sequencing of Miscanthus × giganteus shows the utility of sorghum as a reference genome sequence for Andropogoneae grasses. Genome Biol 11:R12
Takasaki H, Maruyama K, Kidokoro S, Ito Y, Fujita Y, Shinozaki K, Yamaguchi-Shinozaki K, Nakashima K (2010) The abiotic stress-responsive NAC-type transcription factor OsNAC5 regulates stress-inducible genes and stress tolerance in rice. Mol Genet Genomics 284:173–183
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Teige M, Scheikl E, Eulgem T, Doczi R, Ichimura K, Shinozaki K, Dangl JL, Hirt H (2004) The MKK2 pathway mediates cold and salt stress signaling in Arabidopsis. Mol Cell 15:141–152
Tran LS, Nakashima K, Sakuma Y, Simpson SD, Fujita Y, Maruyama K, Fujita M, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2004) Isolation and functional analysis of Arabidopsis stress-inducible NAC transcription factors that bind to a drought-responsive cis-element in the early responsive to dehydration stress 1 promoter. Plant Cell 16:2481–2498
Wang XE, Basnayake BMVS, Zhang HJ, Li GJ, Li W, Virk N, Mengiste T, Song FM (2009) The Arabidopsis ATAF1, a NAC transcription factor, is a negative regulator of defense responses against necrotrophic fungal and bacterial pathogens. Mol Plant Microbe In 22:1227–1238
Wang N, Zheng Y, Xin HP, Fang LC, Li SH (2013) Comprehensive analysis of NAC domain transcription factor gene family in Vitis vinifera. Plant Cell Rep 32:61–75
Wu YR, Deng ZY, Lai JB, Zhang YY, Yang CP, Yin BJ, Zhao QZ, Zhang L, Li Y, Yang CW, Xie Q (2009) Dual function of Arabidopsis ATAF1 in abiotic and biotic stress responses. Cell Res 19:1279–1290
Xie Q, Frugis G, Colgan D, Chua NH (2000) Arabidopsis NAC1 transduces auxin signal downstream of TIR1 to promote lateral root development. Genes Dev 14:3024–3036
Xue GP, Way HM, Richardson T, Drenth J, Joyce PA, McIntyre CL (2011) Overexpression of TaNAC69 leads to enhanced transcript levels of stress up-regulated genes and dehydration tolerance in bread wheat. Mol Plant 4:697–712
Yokotani N, Ichikawa T, Kondou Y, Matsui M, Hirochika H, Iwabuchi M, Oda K (2009) Tolerance to various environmental stresses conferred by the salt-responsive rice gene ONAC063 in transgenic Arabidopsis. Planta 229:1065–1075
Yoo SY, Kim Y, Kim SY, Lee JS, Ahn JH (2007) Control of flowering time and cold response by a NAC-domain protein in Arabidopsis. PLoS ONE 2:e642
Zheng XN, Chen B, Lu GJ, Han B (2009) Overexpression of a NAC transcription factor enhances rice drought and salt tolerance. Biochem Biophys Res Commun 379:985–989
Acknowledgments
This study was supported by grants from the National High-Tech Research and Development Program of China (2011AA100209), and National Key Technology Support Program of China (2013BAD22B01). We thank Amy Mason (Forage Improvement Division, the Samuel Roberts Noble Foundation, USA) for her critical review on the manuscript.
Conflict of interest
We declare that we have no conflict of interest.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Communicated by Marcelo Menossi.
L. Ji and R. Hu contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ji, L., Hu, R., Jiang, J. et al. Molecular cloning and expression analysis of 13 NAC transcription factors in Miscanthus lutarioriparius . Plant Cell Rep 33, 2077–2092 (2014). https://doi.org/10.1007/s00299-014-1682-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-014-1682-8