Abstract
Ryegrass was pulse-labeled with enriched 13CO2 for 18 h, followed by dynamic photosynthetic-carbon (13C) quantification in the plant (shoot, root), soil aggregates (three size classes), and microbial phospholipids fatty acids (PLFA-SIP) in soil amended with or without 700 °C-pyrolyzed biochar. We observed that biochar led to no difference of 13C allocation in shoot or root but reduced 88.7% of total 13C in soil, with decreased incorporation by 92.8% (macroaggregates), 94.5% (microaggregates), and 84.1% (silt-clays), respectively, compared to biochar-unamended soil. Meanwhile, biochar exerted negative effects on fungal relative abundance but led to positive impacts on that of bacteria, e.g., it reduced root-associated fungi (i.e., 16:1ω5c) and fungal-assimilated 13C (from averagely 71.2 ng C g−1 soil to 26.3 ng C g−1 soil after biochar application). The enhanced bacteria/fungi could be driven by biochar-mediated pH increase that relieved acid stress to bacteria. Co-occurrence network confirmed that biochar addition favored bacteria to compete with fungi, leading to decreased aggregation and stability (indicated by reduced normalized mean weight diameter) due to less fungal entangling with aggregates, thus exposing the rhizodeposits to bacterial (i.e., actinomycetes) decomposition. The correlation analysis further evidenced that fungal abundance was associated with 13C accumulation in soil aggregates, while bacterial relative abundance especially that of actinomycetes was negatively correlated with 13C accumulation. Random forest modeling (RF) supported the contributions of fungi to 13C-sequestration compared to bacteria. Taken together, we concluded that less stabilization of rhizodeposits in the biochar-amended soil was due to changes in microbial community, particularly the balance of fungi-bacteria and their interactions with soil physicochemical properties, i.e., aggregation and pH.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Rhizodeposits represent the labile source of carbon (C) and contribute to the formation of soil organic matter (SOM) (Pausch and Kuzyakov 2018). Photosynthetic-C input into grassland under long-term planting ranges from 0.11 to 3.04 t C ha−1 year−1 (Soussana et al. 2004). Within 4 weeks, up to 11% of the photosynthetic-C of willows was found in SOM, demonstrating a close link and great aboveground photosynthesis contributions to belowground C storage (Neergaard et al. 2002). The quality and quantity of rhizodeposits in soil depends on plant species (Ladygina and Hedlund 2010) and phenology (Epron et al. 2011; Meng et al. 2013), agriculture practices (Luo et al. 2018), soil properties such as aggregate size fractions (Fahey et al. 2013), and microbial community (Kaiser et al. 2015).
Rhizodeposits are easily available C and energy for soil microorganisms. On an average, 0.54% of 13C was found in microbial biomass C (MBC) immediately after 13CO2 pulse labeling of flooded rice, and up to 0.41% of 13C remained in MBC pools at the end of the growing season in the rhizosphere (Lu et al. 2002). Considering the microbial process mediated root-derived C sequestration, it is also essential to identify the principal groups associated with roots and primary recipients of root-derived organics (Jin et al. 2013). A study showed that the 13C incorporation into the microbial community was detected in fungal PLFA biomarkers, ranging from 14.1 to 39.6% of total 13C-PLFAs at different rates of nitrogen (N) application (Ge et al. 2017). Rhizodeposits can be differently utilized by microbial communities depending upon the traits of each microbial group on C and nutrient requirements (Malik et al. 2019; Peduruhewa et al. 2020), with consequences for rhizodeposits stabilization and decomposition (Hartmann et al. 2009). Soil abiotic properties, such as pH, SOM content, and physical structure, can affect the composition of soil microbial community and consequent consumption of rhizodeposits. In this regard, (i) combination of resources (e.g., C content) and stress (e.g., low pH) were found to be the main contributors to soil microbial community and subsequent C decomposition (Malik et al. 2019); (ii) aggregation of different size classes regulates C processes via harboring unique microbial community composition and their functions (Caravaca et al. 2005; Kumar et al. 2017), as well as determining the accessibility of microbe to C sources. For instance, Brookes et al. (2017) revealed that the spatial arrangement of the aggregate structure affected the bio-accessibility of C to consumers, thus largely regulated SOC content.
The fate of root-derived C in various aggregate size classes has gained increasing attention recently (Fang et al. 2016; Zhang et al. 2015), as the dynamics of aggregates elucidate mechanisms involved in SOM formation, decomposition and consequent storage (Six et al. 2002). Aggregate size classes exert effects on the processes of both mineralization (Blaud et al. 2012) and stabilization of organic matter (Qiao et al. 2014). According to the hierarchical order of aggregation, free primary soil particles and silt-clays (< 0.02 mm) are cemented together into microaggregates (< 0.25 mm) by persistent binding agents (i.e., metal-cation complexes or oxides). These stable microaggregates in turn are bound with transient (plant or microbial derived polysaccharides) and temporary (roots and fungal hyphae) agents. Furthermore, these intra-aggregates and agents together exert a binding capacity on macroaggregate formation (> 0.25 mm) (Jastrow 1996; Tisdall and Oades 1982). As the hierarchical order and binding agents, microaggregate stability is stronger and less vulnerable to soil management in contrast with macroaggregates (Hassink et al. 1997). Therefore, longer turnover time of new-fixed C was estimated in microaggregates than in macroaggregates through more stable physicochemical protections (Bailey et al. 2013; Fang et al. 2016). For example, after 1.5 years of 13C-labeled wheat residues application, the microaggregates still accounted for the highest amount (about 50%) of the plant-derived C (Angers et al. 1997). Macroaggregate plays significant role in the early period since receiving new C input into soil (Six et al. 2002). Therefore, it is important to understand the spatial distribution of root-derived C that entering into different aggregate size classes.
Biochar is widely applied in agro-ecosystem to sustain SOM stocks (Sohi 2012; Luo et al. 2017b). Biochar benefiting C sequestration is due to recalcitrant nature per se, and it can persist in soil up to thousand years (Glaser et al. 2009; Singh et al. 2012). Additionally, biochar could indirectly stabilize non-biochar C, e.g., root-derived C, by enhancing the formation of mineral-organic associations (Weng et al. 2017). Also, biochar was found to enhance decomposition of plant straw, rhizodeposits and SOM through changing microbial community composition (Maestrini et al. 2015; Luo et al. 2016). Although several studies reported the processes and mechanisms of rhizodeposits decomposition/stabilization via biochar-induced changes in soil abiotic and biotic properties, photosynthetic-C distribution within plant-soil-microbe continuum, particularly rhizodeposits assimilation in microbial community and aggregates in biochar-amended soil is still limited (Keith et al. 2015; Weng et al. 2017; Whitman et al. 2014).
Here, we performed 13CO2 pulse-labeling on ryegrass-soil system that treated with 0% and 3% (by dry-soil weight) biochar, to quantify the allocation of photosynthetic-C in plants (shoots, roots) and soil (three aggregate size fractions), during the elongation stage at four sampling times (2, 5, 9, and 16 days after labeling). The study also aimed to investigate the composition of main microbial groups and their roles in utilizing rhizodeposits by using PLFA-SIP technique, thus fully assessing abiotic and microbial mechanisms underlying plant-C sequestration in biochar incorporated soils. We hypothesized that (i) the allocation of photosynthetic-C within plant-soil-microbe pools varied due to changes of biochar in soil properties; (ii) biochar amendment enhanced soil aggregates stability, hence increased ryegrass-derived C retention in macroaggregate; (iii) soil microbial community and their assimilation of rhizodeposits differ due to biochar-induced changes in soil abiotic variables, e.g., pH.
Materials and methods
Soil and materials
Top soil (0–20 cm) was sampled from a mulberry experimental field, Zhejiang Province, China (30° 16′ N, 120° 11′ E). The mean annual temperature is 16.2 °C with an annual rainfall 1500 mm. Soil was passed through a 2-mm sieve to remove visible plant residues and stones. Soil pH was determined by suspension in a 1:2.5 soil solution (0.01 M CaCl2). Total C and N were measured by an analyzer with a built-in dry combustion device (LECO, St. Joseph, MI, USA). The 13C abundance (δ13C) was determined with an isotope ratio mass spectrometry (DELTA V plus IRMS, Bremen, Germany). The soil was classified as sandy clay loam according to the US soil classification (Soil Survey Staff 2014); it contained: sand 63.8%, silt 20.4%, clay 15.4%. The initial δ13C value of the soil was −24.9‰ and pH at 5.8. The soil contained 12.5 g kg−1 total C and 0.9 g kg−1 total N. Biochar was derived from swine manure feedstock as described by Dai et al. (2014), pyrolyzed in a muffle furnace (Yizhong Electricity Furnace Inc., Shanghai, China) under oxygen limited conditions at a heating rate of 26 °C min−1 and kept up to 700 °C for 8 h. After being cooled under ambient condition, the biochar was grounded <0.2 mm and collected. Biochar pH (9.6) was measured in deionized water at a ratio of 1:5 (w/w) biochar/water by an ISFET electrode with a tip diameter of 1.2 mm (SevenExcellence™ pH, Mettler Toledo Inc., Switzerland) after 1 h solution equilibrium (Zhang et al. 2016). Biochar contained 56.7 total C (%), 0.67 total N (%), and had a natural δ13C abundance of −29.8‰. Additionally, biochar contained 138.7 mg kg−1 Zn, 36.5 mg kg−1 Cu, and 3.5 mg kg−1 As. The aromatic-C accounted for 53.2% of the chemical functional groups. The basic properties are given in SI Table 1.
Experimental layout
Two treatments included biochar-unamended (BC-0%) and biochar-amended (BC-3%) soils. Biochar was homogenously applied in sieved soils as 3% of soil weight (oven-dry basis) (Bruun et al. 2011). The experiment conducted 32 labeled pots (two treatments each with four replicates and four sampling points), and the same number of pots were unlabeled. Polyvinyl chloride (PVC) pots (height 10.5 cm, diameter 11.3 cm) were filled with 324 g dry soil. Black film covered the pots to avoid the influence of autotrophic microorganism. The soil water content was adjusted to 60% of water holding capacity, and it was controlled gravimetrically during the experiment.
Ryegrass (Lolium perenne L.) seeds were sterilized with 30% H2O2 solution for 30 min; then, they were washed with deionized water thoroughly. To hasten germination, seeds were soaked in 25 °C deionized water in the dark for 12 h before being sown in pots. Emergence occurred 1 week after sowing, weak leaves were cut, and shoots were clipped 4 cm uniformly at the tillering stage of ryegrass (4 weeks after sowing). Each pot contained 30 seedlings with a similar growth before labeling.
Ryegrass growth conditions and 13CO2 pulse labeling
The light period lasted for 14 h and the temperature was 28 °C during the day and 20 °C during the night and relative humidity was 70% in the greenhouse (Chen et al. 2016). Ryegrass seedlings were put in an air-tight perspex chamber (length 1.75 m, width 0.8 m, height 1 m) in the greenhouse, then an 18 h-13CO2 pulse labeling was performed during their elongation stage according to Lu et al. (2002). The 13CO2 was generated by an acid-base reaction mixing 3 M excess H2SO4 and 1 M Na213CO2 (≥ 99% atom, Cambridge Isotope Laboratories, Inc., USA) solution. A fan was fixed at the top of the chamber to homogenize the gas, and the thermometer was installed to detect the internal temperature. The CO2 concentration was monitored by a portable CO2 detector (SMART-CO2, Shenzhen, China), and new 13CO2 was produced through regulating the acid-base reaction. Unlabeled pots were kept 20 m away from the labeling chamber.
Plants and soils from independent pots were destructively sampled on 2, 5, 9, and 16 days after labeling. The first sampling started on January 25, 2018, shoots were cut off at the soil surface, and visible roots were separated manually through a 0.5-mm sieve with deionized water. Soil sticking to roots was removed as modifications from Luo et al. (2018): soil was suspended in 40 mL CaCl2 buffer (0.01 M, pH 6.2) and centrifuged at 8400×g for 5 min (Yuan et al. 2016). Then decanted the supernatant to separate the plant-derived debris from soil, i.e., picked out floatable fine roots then washed with deionized water. Subsequently, the remaining soil sediments were purified by twice resuspension in deionized water, centrifuged, collected, and mixed thoroughly. Finally, all shoots and roots were individually oven-dried at 65 °C for 72 h, weighed and ball-milled <0.25 mm. All fresh soil samples were divided into three portions of subsamples: one portion was immediately stored at −80 °C for PLFA extraction; one portion was stored at 4 °C for aggregate size fractionation; the remaining portion was freeze-dried for 72 h, grounded, sieved through a 100 mesh to determine total C/N content and 13C-values.
Aggregate size fractionation
Soil aggregates of three size classes were separated by the optimal-moisture sieving method (Dorodnikov et al. 2009). The soil samples were spread on a thin layer in a ventilation hood and air-dried to 25% WHC. Then 20-g soil sub-samples (dry mass) were transferred to a nest of sieves (2, 0.25, and 0.053 mm). The separation was achieved by an electric swing-machine (Electricity analyzer Inc., Hangzhou, China) moving the sieves up and down 4 cm at 30 rounds per minute for 30 min. Thereafter, macroaggregates (0.25–2 mm) were collected from the surface of a 0.25-mm sieve, while microaggregates (0.053–0.25 mm) passed through the 0.25-mm sieve and collected on surface of 0.053-mm sieve. The remaining material passed through the 0.053-mm sieve and was categorized as silt-clays (< 0.053 mm). All size classes were weighed, freeze-dried overnight for 13C and total C content analysis.
PLFAs extraction and analysis
The PLFAs were extracted and measured according to Luo et al. (2017a). The freeze-dried soil samples (2 g) were extracted by a 22.8-mL single-phase (0.15 M, pH 4.0) mixture of chloroform-methanol-citrate buffer (1:2:0.8). Then phospholipids were separated from neutral lipids and glycolipids on a silica acid column (Supelco, Bellefonte, PA, USA). Following methylation of the phospholipids, the methyl esters were extracted in n-hexane and N2 gas-dried. Methyl nonadecanoate fatty acid (19:0) was added before derivatization, as an internal concentration standard to quantify the phospholipids. Using a gas chromatograph installed with a flame ionization detector (GC-FID, Agilent Technologies, USA) and fitted with a MIDI Sherlock microbial identification system (Version 4.5; MIDI, Newark, DE, USA), the PLFA biomarkers were identified into five groups eventually: fungi, actinomycetes, gram-negative (G−) bacteria, gram-positive (G+) bacteria, and general FAMEs (fatty acids methyl esters). The 16:0 and 18:0 PLFA cannot be assigned to specific functional groups of microorganisms (Dungait et al. 2013), which occur generally in the all living cells, being defined as the general FAME (Tavi et al. 2013). Detailed nomenclature of PLFAs was described in SI Table 2. To determine the 13C incorporation into specific biomarker in response to biochar application, the δ13C value of individual PLFAs (PLFA-SIP) was measured using a Trace GC Ultra gas chromatograph with combustion column attached via a GC Combustion III to a Delta V Advantage isotope ratio mass spectrometer (Thermo Finnigan, Germany) (Wang et al. 2016).
Calculations and statistics
The 13C abundance of shoots, roots, soil aggregate size classes, and PLFA biomarker were measured as δ13C (‰) values relative to the standard Pee Dee Belemnite, and values were calculated as artificial-labeled atom percent (%) (Zhu et al. 2017). Then, the 13C incorporation into individual C pools (mg 13C kg−1 soil) was calculated as reported by Atere et al. (2017), according to the following equation:
where the δ13C values of plant, soil, and PLFAs samples were measured as δ (‰) relative to the Pee Dee Belemnite (PDB; 13C, 0.0111802) standard and further expressed as atom percent (atom%).
where (atom%13C)s,L and (atom%13C)s,UL are 13C percent (%) of labeled and unlabeled samples, respectively, Cs is the C content of each sample (mg C kg−1 soil).
13C isotope analysis of PLFAs was used to explore which microbial group utilized the plant-C (Ge et al. 2017; Luo et al. 2017a). The amount of 13C incorporation in each PLFA (ng 13C g−1 soil) was determined using a mass balance approach:
where (atom%13C)PLFA,L and (atom%13C)PLFA,UL are the 13C percent (%) of labeled and unlabeled PLFA biomarkers, respectively. CPLFA is the C content of each PLFA biomarker (ng C g−1 soil). The relative abundance of each individual group was expressed as the percentage of each PLFA-C in total PLFAs-C according to the following equation:
where CPLFA-Group is the sum of the C-PLFA of each microbial group, and ∑ CPLFAs is the total C-PLFA amount.
Partial calculation and modeling of microbial data were based on two treatments with three replicates, including Pearson’s correlation analysis, the redundancy analysis (RDA) and the RF modeling. Pearson’s correlation analysis was conducted by SPSS 20.0 (p < 0.05) (SPSS, Inc., Chicago, IL, USA), which individually assessed the correlation between the relative abundance of each microbial group and 13C allocation into plant shoot, root, the whole soil, and three aggregate fractions. To reveal environment factors influencing the composition of main microbial groups and their assimilation of 13C, RDA was performed using the RDA package of R (Comprehensive R Archive Network) (Kambura et al. 2016). This canonical community ordination method explored the relationship between 13C-PLFA amount and soil properties (pH, total C/TC, total N/TN, water content/WC, dissolved organic C/DOC) with great fitness (r2 > 0.5, p < 0.05). To assess the significant predictors of integrate systems in the 13C belowground allocation quantitatively, biotic- and abiotic-predictors were analyzed by the RF modeling via random forest package (Chen et al. 2019); biotic-predictors were microbial relative abundance, root biomass and bacterial stress; abiotic-predictors were selected based on principal factors from RDA and aggregation stability (the normalized mean weight diameter of aggregation).
To explore the biochar-induced influence on connections within and between individual microbial groups, a co-occurrence network analysis (Conet) was performed (based on all extracted PLFA biomarkers) using Cytoscape version 3.5 (Shannon et al. 2003). The biochar-unamended and biochar-amended soil were separately analyzed with twelve samples (with 3 replicates each time, 4 sampling times during plant growth). Firstly, the Conet pattern was constructed based on the PLFA-C. Secondly, the dissimilarity threshold to the maximum value of the Kullback-Leibler dissimilarities (KLD) matrix and the Spearman’s correlation threshold to 0.9 were calculated (p < 0.05). Thirdly, each edge was set at 1000 iterations of bootstrap distribution under the permutations of randomization. The specific p value was united through the Brown’s method, and was adjusted according to Benjamini and Hochberg (1995) to diminish the probability of false-positive results. Finally, ran and generated networks, the nodes of networks represented specific microbial groups, the edges indicated paired and significant correlations between nodes. Network visualization was conducted in Cytoscape.
All statistical analyses of all non-microbial data were considered the mean of four replicates with standard deviations and were performed by SPSS 20.0. One-way ANOVA (analysis of variance) was used to analyze data variance. Normality and homogeneity of variances were tested using the Shapiro-Wilk and Levene’s test, respectively. The Tukey post hoc test was applied to identify the significance of data in each treatment during four sampling points. The Student’s t test was used to evaluate the significant differences of means pairwise between BC-0% and BC-3% treatment (p < 0.05).
Results
Allocation of photosynthetic-C in ryegrass-soil system
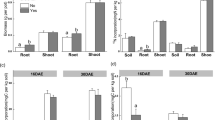
Biochar application significantly decreased shoot biomass on day 2 and day 16 compared to the biochar-unamended soil, respectively (p < 0.05) (SI Fig. 2a), and reduced root biomass from 2 to 9 days. The 13C of shoot was the highest among pools of shoot, root and soil, but it gradually decreased from 10.5 to 8.9 mg C pot−1 during growth (p > 0.05) (Fig. 1a). The 13C of root had a similar trend as that of shoot in both treatments over time (Fig. 1c). Based on time-average, biochar did not result in a significant difference in the 13C-pool of shoot and root (Fig. 1b, d). Less 13C was incorporated in both shoot and root from 2 to 9 days in biochar-enriched soils, whereas the opposite trend occurred on day 16 (p > 0.05).
Photosynthetic-13C allocation in ryegrass-soil system on 2, 5, 9, and 16 days after labeling of shoots (a, b); roots (c, d); macroaggregates (e, f); microaggregates (g, h); and silt-clay fractions (i, j). Treatments include biochar-unamended (BC-0%) and biochar-amended (BC-3%) soil. Asterisks above the bars indicate significant differences pairwise between treatments at each time point. Different letters indicate statistically significant differences in a treatment with time. Error bars represent the standard error of the mean (n = 4)
Biochar significantly decreased the 13C incorporation into macroaggregate by 92.8%, into microaggregate by 94.5% and into silt-clays by 84.1%, compared to soil without biochar (Fig. 1f, h, j). In biochar-unamended treatments, 13C allocation in all aggregates significantly decreased from 2 to 5 days, with reduction in macroaggregate (27.5%), microaggregates (52.7%), and silt-clays (81.4%) (p < 0.05) (Fig. 1e, g, i). Both biochar-amended and -unamended soils had similar trend of 13C incorporation in aggregate size fractions since day 5 (p < 0.05).
Composition of main microbial groups and network interactions
The highest relative abundance was observed in fungal biomarker, accounting for 36.3% of total PLFAs on average (Fig. 2a). Biochar addition decreased the relative abundance of fungal PLFAs by 21.0% on average (p < 0.05), and increased that of G+, G− bacteria and actinomycetes by 33.9%, 28.8%, and 69.7%, respectively. Significant declines were found in environmental stress indicators for bacteria after the biochar amendment, i.e., both stress 1 (cy17:0/16:1ω7c) and stress 2 (cy19:0/18:1ω7c) decreased by 15.8% and 26.7% on day 2 (p < 0.05), respectively (Fig. 2b).
Relative abundance of PLFA biomarkers represent main microbial groups in total PLFAs (a); biochar-induced bacterial stress (shown as the ratio of cyclopropyl with its precursor PLFA-C) (b). Treatments include biochar-unamended (BC-0%) and biochar-amended (BC-3%) soil. Asterisks above the bars indicate significant differences pairwise between treatments at each time point; error bars represent the standard error of the mean (n = 3)
The co-occurrence networks showed that the application of biochar modified the interactions among microbial communities including general FAME, fungi, actinomycetes, G+ and G− bacteria (Fig. 3c, d). The blue and red edges represented significantly positive and negative correlations between pairs of groups, respectively. The network analysis of the biochar-unamended soils showed 53.3% of negative links between fungi and bacterial groups, i.e., G− bacteria and actinomycetes (Fig. 3c, SI Table 5). In the presence of biochar, the number of edges representing positive and negative connections decreased, the structure of network became loose (Fig. 3d), and the mutual-exclusion only existed between fungi-bacteria connections; meanwhile, the fungal percentage decreased while the bacterial percentage increased (p < 0.05).
Photosynthetic-13C incorporation into microbial PLFAs in biochar-unamended (a) and biochar-amended (b) soil, error bars represent the standard error of the mean (n = 3). Co-occurrence network of microbial communities based on Spearman’s threshold (0.9) in biochar-unamended (c) and biochar-amended soil (d), nodes with different color represent individual group; node size is in proportion to microbial relative abundance; solid blue and dotted red lines represent positive and negative connections, respectively. Redundancy analysis of 13C-PLFA profiles of soil samples in two treatments (biochar with 0% and 3% application rate) on 2, 5, 9, and 16 days after labeling (e). Random forest analysis for important biotic and abiotic predictors of 13C allocation in soil (f)
13C incorporation into microbial groups
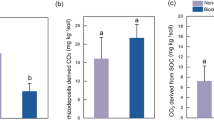
The 13C incorporation into microbial groups in both treatments ranked as fungi > general FAME > G− bacteria > G+ bacteria > actinomycetes (Fig. 3a, b). Fungal 13C-PLFA was dominant in both treatments during the entire period. The 13C incorporation into fungi peaked (141.2 ng C g−1) on day 2 without biochar and decreased with time (Fig. 3a). Application of biochar decreased 13C within most microbial groups especially in fungi between day 2 and day 5 (p < 0.05) (Fig. 3b; SI Table 3). From 9 to 16 days, the 13C incorporation into all groups was similar between biochar-amended and biochar-unamended soils.
Correlations between soil abiotic, biotic variables, and rhizodeposits stabilization
Redundancy analysis (RDA) revealed the correlation between 13C-PLFAs and soil properties (pH, TC, TN, WC, DOC) (Fig. 3e). The first and second canonical axis explained 63.7% and 11.1% of the variation to 13C-PLFA, respectively. The longest arrow indicated that pH was the main factor in the axis RDA1, and it was positively correlated with biochar-amended samples. The axis RDA2 separated the biochar-unamended samples with time, and clustered by soils sampled at day 9 and day 16.
The RF model explained 76.3% of the total variance (Fig. 3e). It indicated fungal relative abundance accounted for 3.2% of the increase of mean square error (InMSE) and represented as the most important biotic predictor for photosynthetic-13C allocation. Consistent with the RDA analysis, TC, TN, and pH were the primary abiotic predictors for 13C utilization by microbial community and stabilization within aggregates (above 7.0% of InMSE).
The Pearson’s correlation coefficients revealed significant relations between 13C allocation belowground and relative abundance of microbial groups (Table 1). The relative abundance of fungi and general FAME were positively correlated with 13C amounts in bulk soil and aggregate size classes, whereas the actinomycetes relative abundance was negatively correlated with 13C amount in soils especially within macroaggregates (p < 0.01).
Discussion
Distribution of photosynthetic-C in plant and soil
Compared with the biochar-unamended soil, the averaged 13C allocation in plant pools (shoot and root) were not changed significantly (p > 0.05) after biochar addition, but much less of 13C was allocated in the soil amended with biochar (Fig. 1). Biochar could not directly affect the 13CO2 flow in leaf photosynthesis, but likely it exerted “indirect influences” on fate of photosynthates in soil via changes in nutrients, pH, and subsequent microbial community. For instance, the altered 13C allocation into soil was likely due to reduced root biomass and less input of root-derived organics into soil for nutrients exchange (Foster et al. 2016), as biochar-mediated increase in nutrient availability (SI Fig. 5). It was reported that insufficient nutrients enhanced root metabolic costs in the secretion of rhizodeposits as C source to trade-off N (Kumar et al. 2016). In this regard, Mellado-Vázquez et al. (2016) showed that biochar addition decreased the abundance of exudate-utilizing fungi. Consistently, biochar decreased fungal PLFA and their 13C assimilation, which also mirrored the decreased rhizodeposits as C sources for microorganisms. Moreover, the mean biomass of shoot was significantly reduced after biochar application (SI Fig. 2), indicating that biochar negatively influenced ryegrass growth likely because it contained toxicants (SI Table 1). This was reported previously (Anyanwu et al. 2018; Revell et al. 2012). However, we cannot distinguish whether the surplus of nutrients or toxic effects inhibited rhizodeposition processes because it is required direct measurement of rhizodeposits.
Biochar effect on allocation of rhizodeposit-13C into soil aggregates
A significant decrease of 13C allocation within three aggregates occurred after biochar addition (Fig. 1f, h, j). This might be due to (i) the reduced root exudation in nutrients sufficient soils; and (ii) the decreased soil aggregation (indicating potential C loss), due to less root exudates which acted as gluing agents for aggregation formation (Tripathi et al. 2014). Components of root exudates, such as polysaccharides, can bind microaggregates to form macroaggregates (Six et al. 2004). Indeed, the normalized mean weight diameter (NMWD) value, an indicator of aggregate stability (Kumar et al. 2017), decreased slightly after biochar application, indicating the negative effects of biochar on aggregation (SI Fig. 4). Contrary to our results, some previous studies suggested a biochar-mediated increase in aggregates formation and stability (Herath et al. 2013; Burrell et al. 2016). The field biochar application promoted the formation of soil organo-mineral microstructures after 9.5 years, thus resulting in an increased recovery of root-13C by 20% (Weng et al. 2017). However, the effects of biochar on aggregate formation depend on times, rates, and types of biochar application and soil types, i.e., texture. Biochar used in this study was produced at 700 °C, and it contained higher aromatic-C content than those produced under lower temperatures (Jie et al. 2015; Novotny et al. 2015). A negative correlation between aromatic fraction of biochar and soil aggregates stability was observed (Sarker et al. 2018), suggesting that the percentage of aromatic-C negatively affected aggregation (Fig.4).
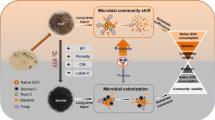
Conceptual diagram of fate of rhizodeposit-13C, as regulated by microbial composition and aggregation stability, determined the 13C allocation. The biochar application reduced rhizodeposit-13C exudation and root growth under excess nutrition: (a) biochar improved bacterial (13C-mineralization associated groups) growth via changing soil TC,TN contents and especially pH, thus larger proportion of bacteria decomposed more 13C; (b) biochar application diminished aggregation thus giving fewer fine-roots that occluded with microbe-available rhizodeposits in macroaggregates, causing less abundant fungi (13C-sequestration associated groups) than in biochar-unamended soil. Consequently, less 13C was redistributed to inner size fractions, which was mineralized by bacterial degradation
Less input of rhizodeposits in the biochar-amended soil decreased aggregation, leading root-derived 13C vulnerable to microbial attack. Since plant-derived CO2 was not quantified in the present study, future work should consider the full fate of photosynthetic-C flow through pools in the plant-soil-atmosphere system, including rhizodeposits respiration.
Effects of biochar on the composition of main microbial groups and bacteria fungi ratio
The relative abundance of G+, G− bacteria and actinomycetes increased in biochar-amended soils during all the sampling times (Fig. 2a). This indicated biochar interacted positively with these bacterial microbiotas in the ryegrass-soil system. The G+ bacteria abundance increased in biochar-amended soils likely due to the ability of this group to decompose the aromatic components (Santos et al. 2012). Actinobacteria were the dominant phyla in soil after biochar addition (Yu et al. 2018; Zhou et al. 2019). Abundance and diversity of bacteria can increase in soils with increasing pH from 4.9 to 7.5 (Nicol et al. 2008), and bacteria can be active in a wide range of pH from 3.6 to 8.9 (Tripathi et al. 2018). The mechanisms underlying the increased relative abundance of bacteria with biochar application include (i) the alleviated acid stress by the biochar-mediated pH increase (Martinsen et al. 2015), as the relative abundance of the bacterial stress indicators, i.e., stress 1 (cy17:0/16:1ω7c) and stress 2 (cy19:0/18:1ω7c), both declined after biochar application (Fig. 2b). The indicator of stress 1 is associated with the physiological stress on bacteria, and the indicator of stress 2 depends on bacterial activity (Macdonald et al. 2004); (ii) biochar pore spaces may have generated suitable habitats for bacteria, with tiny bacteria adhered to biochar surfaces, being less susceptible to leaching (Lehmann et al. 2011); (iii) biochar resulted in a more suitable microbial habitat because it can offer C sources, nutrients, space, etc., and thus, promoting microbial, i.e., actinobacteria, colonization (Luo et al. 2013).
Therefore, biochar amendment caused opposite effects on fungi, i.e., the relative abundance of fungi decreased with biochar application especially in the early time compared to bacteria (Fig. 2a). Network analysis revealed less co-existence (positive interactions) than mutual-exclusion (negative interactions) of microorganisms in biochar-amended soils, with more negative links between fungi and bacteria (Fig. 3c, d). Biochar application loosed the network structure, especially exacerbated mutual-exclusion between fungi and bacteria. The shifted composition of main microbial groups (bacteria versus fungi) by biochar addition (Fig. 2a) was likely due to biochar-mediated changes in soil C, nutrients, C/N ratio and pH (Fig. 3e). The biochar enhanced pH, increased abundance of bacteria that outcompeted fungi as the former were more sensitive to pH changes than fungi (Herold et al. 2012; Rousk et al. 2010). The C/N ratio of biochar per se and corresponding changes in soil C/N ratio could have affected the fungi: bacteria ratio (Rousk et al. 2013). Root biomass might have been limited either by biochar induced toxicity (Mellado-Vázquez et al. 2016) or by over-nutrition (Hagemann et al. 2017). The reduction of root biomass decreased colonization of root-associated microorganisms especially fungi (Deyn et al. 2011; Kusliene et al. 2014). For example, the abundances of mycorrhizal fungi decreased with biochar-mediated higher N or/and P availability (Lehmann et al. 2011).
Microbial utilization of rhizodeposit-13C
Translocation of 13C into PLFA biomarkers occurred rapidly, peaking on the first sampling time (day 2), and then decreased (Fig. 3a, b). The RDA plot showed clear separation of the 13C-PLFA biomarkers with time (Fig. 3e). Early incorporation of the photosynthetic-13C into microbial PLFAs was consistent with Chaudhary and Dick (2016), who revealed the C from photosynthesis was rapidly transported belowground within 2 days after pulse labeling. Arbuscular mycorrhizal fungi (AMF), represented by 16:1ω5c biomarkers (Olsson et al. 1997; Ladygina and Hedlund 2010), could have assimilated grass rhizodeposited-13C after 9 h of 13CO2 labeling (Dorodnikov et al. 2009), whereas the bacterial uptake of 13C was delayed (Denef et al. 2007, 2009), likely because bacteria used C-leaking of hyphae or fungal necromass-C rather than the direct utilization of fresh root-exudates (Jin and Evans 2010).
Rhizodeposit-13C incorporation into PLFA biomarkers showed different ability of rhizodeposits utilization between bacteria and fungi (Fig. 3a, b). Our results were consistent with other observations that fungi dominated in soil-ryegrass (Kusliene et al. 2014; Rinnan and Baath 2009), with the highest 13C-PLFA content up to 44.4% (Fig. 3a). Fungi could have assimilated labile exudates directly from roots via an extensive hyphal network before than bacteria (Yuan et al. 2016; Berruti et al. 2013). Biochar significantly decreased 13C incorporation into soil microorganisms particularly fungal 13C-PLFAs (Fig. 3b), confirming what reported by Mellado-Vázquez et al. (2016). Therefore, we confirmed that fungi adapted more quickly and used rhizodeposits earlier than bacteria (Fig. 3a), and the addition of biochar decreased C sequestration.
Contributions to rhizodeposits sequestration via abiotic and biotic attributes
The random forest modeling attributed the stabilization of photosynthetic-13C in soil to abiotic factors (organic C content, nutrient content, pH and aggregates) and biotic variables (fungal and bacterial abundance) (Fig. 3f). Biochar reduced 13C stabilization within aggregate size classes (84–94%) due to the changes in abiotic variables or shifted bacteria to fungi relationships with implication in the fungal hyphae entanglement of aggregation and pH-mediated bacterial mineralization of rhizodeposits, respectively.
The biochar-mediated decrease in fungal abundance decreased aggregate stability, as fungal hyphae are involved in the aggregate formation by acting as binding agents (Gupta and Germida 2015). The decrease in fungal abundance was due to competition by bacteria, reduced root-fungal interactions, diminished fungal use of rhizodeposit-C (by 68.3%), and the aggregation processes, with the latter reducing the physicochemical protection of rhizodeposit-13C. Noticeably, a positive Pearson’s correlation (r2 = 0.721, p < 0.01) was found between fungal relative abundance and 13C allocation belowground.
In addition, biochar significantly decreased 13C incorporation into soil microorganisms except in actinomycetes (Fig. 3b). The increased relative abundance of actinomycetes by biochar has been already reported, and attributed to the better habitat, the decreased acid stress and the increased C and nutrients (Khodadad et al. 2011; Luo et al. 2013). The enrichment of actinomycetes by biochar likely increased rhizodeposits mineralization (Peduruhewa et al. 2020), as supported by the negative correlation of actinomycetes relative abundance and 13C allocation in aggregates, especially within macroaggregates (Table 1). Overall, biochar amendment decreased soil aggregate formation and stability by decreasing fungal abundance, but enhanced relative abundance of bacteria and their utilization of rhizodeposit-13C, with the consequent decrease of 13C in soil aggregates.
Conclusions
This study investigated influence of high temperature pyrolyzed (700 °C) biochar on the allocation of photosynthetic-C within microbial groups inhabiting ryegrass-soil. Biochar amendment decreased 13C stabilization by 84–94% in soil aggregate size classes likely due to (i) biochar-mediated decrease in aggregate formation and stability due to declined fungal abundance with the consequent less root-hyphae-aggregate interactions; (ii) biochar increased pH with enhanced positive effects on relative abundance and activity of bacteria, especially actinobacteria. This study improved the knowledge on C sequestration by biochar by better understanding interactions among root, bacteria, fungi, and aggregates. In particular, biochar caused less stabilization of rhizodeposits in soil via diminished fungi-promoted aggregation and enhanced bacterial abundance.
References
Angers DA, Recous S, Aita C (1997) Fate of carbon and nitrogen in water-stable aggregates during decomposition of 13C 15N-labelled wheat straw in situ. Eur J Soil Sci 48:295–300. https://doi.org/10.1111/j.1365-2389.1997.tb00549.x
Anyanwu IN, Alo MN, Onyekwere AM, Crosse JD, Nworie O, Chamba EB (2018) Influence of biochar aged in acidic soil on ecosystem engineers and two tropical agricultural plants. Ecotoxicol Environ Saf 153:116–126. https://doi.org/10.1016/j.ecoenv.2018.02.005
Atere CT, Ge T, Zhu Z, Tong C, Jones DL, Shibistova O, Guggenberger G, Wu J (2017) Rice rhizodeposition and carbon stabilisation in paddy soil are regulated via drying-rewetting cycles and nitrogen fertilisation. Biol Fertil Soils 53:407–417. https://doi.org/10.1007/s00374-017-1190-4
Bailey VL, McCue LA, Fansler SJ, Boyanov MI, DeCarlo F, Kemner KM, Konopka A (2013) Micrometer-scale physical structure and microbial composition of soil macroaggregates. Soil Biol Biochem 65:60–68. https://doi.org/10.1016/j.soilbio.2013.02.005
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc B 57:289–300. https://doi.org/10.1111/j.2517-6161.1995.tb02031.x
Berruti A, Borriello R, Lumini E, Scariot V, Bianciotto V, Balestrini R (2013) Application of laser microdissection to identify the mycorrhizal fungi that establish arbuscules inside root cells. Front Plant Sci 4:135. https://doi.org/10.3389/fpls.2013.00135
Blaud A, Lerch TZ, Chevallier T, Nunan N, Chenu C, Brauman A (2012) Dynamics of bacterial communities in relation to soil aggregate formation during the decomposition of 13C-labelled rice straw. Appl Soil Ecol 53:1–9. https://doi.org/10.1016/j.apsoil.2011.11.005
Brookes PC, Chen Y, Chen L, Qiu G, Luo Y, Xu J (2017) Is the rate of mineralization of soil organic carbon under microbiological control? Soil Biol Biochem 112:127–139. https://doi.org/10.1016/j.soilbio.2017.05.003
Bruun EW, Müller-Stöver D, Ambus P, Hauggaard-Nielsen H (2011) Application of biochar to soil and N2O emissions: potential effects of blending fast-pyrolysis biochar with anaerobically digested slurry. Eur J Soil Sci 62:581–589. https://doi.org/10.1111/j.1365-2389.2011.01377.x
Burrell LD, Zehetner F, Rampazzo N, Wimmer B, Soja G (2016) Long-term effects of biochar on soil physical properties. Geoderma 282:96–102. https://doi.org/10.1016/j.geoderma.2016.07.019
Caravaca F, Alguacil MM, Torres P, Roldan A (2005) Plant type mediates rhizospheric microbial activities and soil aggregation in a semiarid Mediterranean salt marsh. Geoderma 124(3–4):375–382. https://doi.org/10.1016/j.geoderma.2004.05.010
Chaudhary DR, Dick RP (2016) Identification of metabolically active rhizosphere microorganisms by stable isotopic probing of PLFA in switchgrass. Soil Sci Plan 47:2433–2444. https://doi.org/10.1080/00103624.2016.1243704
Chen L, Brookes PC, Xu J, Zhang J, Zhang C, Zhou X, Luo Y (2016) Structural and functional differentiation of the root-associated bacterial microbiomes of perennial ryegrass. Soil Biol Biochem 98:1–10. https://doi.org/10.1016/j.soilbio.2016.04.004
Chen L, Jiang Y, Liang C, Luo Y, Xu Q, Han C, Zhao Q, Sun B (2019) Competitive interaction with keystone taxa induced negative priming under biochar amendments. Microbiome 7:77. https://doi.org/10.1186/s40168-019-0693-7
Dai Z, Brookes PC, He Y, Xu J (2014) Increased agronomic and environmental value provided by biochars with varied physiochemical properties derived from swine manure blended with rice straw. J Agr Food Chem 62:10623–10631. https://doi.org/10.1021/jf504106v
Denef K, Bubenheim H, Lenhart K, Vermeulen J, Van Cleemput O, Boeckx P, Müller C (2007) Community shifts and carbon translocation within metabolically-active rhizosphere microorganisms in grasslands under elevated CO2. Biogeosciences 4:769–779. https://doi.org/10.5194/bg-4-769-2007
Denef K, Roobroeck D, Wadu MCWM, Lootens P, Boeckx P (2009) Microbial community composition and rhizodeposit-carbon assimilation in differently managed temperate grassland soils. Soil Biol Biochem 41:144–153. https://doi.org/10.1016/j.soilbio.2008.10.008
Deyn GBD, Quirk H, Oakley S, Ostle N, Bardgett RD (2011) Rapid transfer of photosynthetic carbon through the plant-soil system in differently managed species-rich grasslands. Biogeosciences 8:1131–1139. https://doi.org/10.5194/bg-8-1131-2011
Dorodnikov M, Blagodatskaya E, Blagodatsky S, Fangmeier A, Kuzyakov Y (2009) Stimulation of r- vs. K-selected microorganisms by elevated atmospheric CO2 depends on soil aggregate size. Fems Microbiol Ecol 69:43–52. https://doi.org/10.1111/j.1574-6941.2009.00697.x
Dungait JAJ, Kemmitt SJ, Michallon L, Guo S, Wen Q, Brookes PC, Evershed RP (2013) The variable response of soil microorganisms to trace concentrations of low molecular weight organic substrates of increasing complexity. Soil Biol Biochem 64:57–64. https://doi.org/10.1016/j.soilbio.2013.03.036
Epron D, Ngao J, Dannoura M, Bakker M, Zeller B, Bazot S, Bosc A, Plain C, Lata J, Priault P, Barthes L, Loustau D (2011) Seasonal variations of belowground carbon transfer assessed by in situ 13CO2 pulse labelling of trees. Biogeosciences 8:1153–1168. https://doi.org/10.5194/bgd-8-885-2011
Fahey TJ, Yavitt JB, Sherman RE, Groffman PM, Wang GL (2013) Partitioning of belowground C in young sugar maple forest. Plant Soil 367:379–389. https://doi.org/10.1007/s11104-012-1459-1
Fang Y, Singh BP, Badgery W, He X (2016) In situ assessment of new carbon and nitrogen assimilation and allocation in contrastingly managed dryland wheat crop–soil systems. Agric Ecosyst Environ 235:80–90. https://doi.org/10.1016/j.agee.2016.10.010
Foster EJ, Hansen N, Wallenstein M, Cotrufo MF (2016) Biochar and manure amendments impact soil nutrients and microbial enzymatic activities in a semi-arid irrigated maize cropping system. Agric Ecosyst Environ 233:404–414. https://doi.org/10.1016/j.agee.2016.09.029
Ge T, Li B, Zhu Z, Hu Y, Yuan H, Dorodnikov M, Davey LJ, Wu J, Kuzyakov Y (2017) Rice rhizodeposition and its utilization by microbial groups depends on N fertilization. Biol Fertil Soils 53:37–48. https://doi.org/10.1007/s00374-016-1155-z
Glaser B, Parr M, Braun C, Kopolo G (2009) Biochar is carbon negative. Nat Geosci 2:2–2. https://doi.org/10.1038/ngeo395
Gupta VVSR, Germida JJ (2015) Soil aggregation: influence on microbial biomass and implications for biological processes. Soil Biol Biochem 80:A3–A9. https://doi.org/10.1016/j.soilbio.2014.09.002
Hagemann N, Joseph S, Schmidt HP, Kammann CI, Harter J, Borch T, Young RB, Varga K, Taherymoosavi S, Elliott KW, McKenna A, Albu M, Mayrhofer C, Obst M, Conte P, Dieguez-Alonso A, Orsetti S, Subdiaga E, Behrens S, Kappler A (2017) Organic coating on biochar explains its nutrient retention and stimulation of soil fertility. Nat Commun 8:1089. https://doi.org/10.1038/s41467-017-01123-0
Hartmann A, Schmid M, Tuinen DV, Berg G (2009) Plant-driven selection of microbes. Plant Soil 321:235–257. https://doi.org/10.1007/s11104-008-9814-y
Hassink J, Whitmore AP, Kuba J (1997) Size and density fractionation of soil organic matter and the physical capacity of soils to protect organic matter. Eur J Agron 7:189–199. https://doi.org/10.1016/S0378-519X(97)80025-6
Herath HMSK, Camps-Arbestain M, Hedley M (2013) Effect of biochar on soil physical properties in two contrasting soils: an Alfisol and an Andisol. Geoderma 209-210:188–197. https://doi.org/10.1016/j.geoderma.2013.06.016
Herold MB, Baggs EM, Daniell TJ (2012) Fungal and bacterial denitrification are differently affected by long-term pH amendment and cultivation of arable soil. Soil Biol Biochem 54:25–35. https://doi.org/10.1016/j.soilbio.2012.04.031
Jastrow JD (1996) Soil aggregate formation and the accrual of particulate and mineral-associated organic matter. Soil Biol Biochem 28:665–676. https://doi.org/10.1016/0038-0717(95)00159-X
Jie Z, Jia L, Liu R (2015) Effects of pyrolysis temperature and heating time on biochar obtained from the pyrolysis of straw and lignosulfonate. Bioresour Technol 176:288–291. https://doi.org/10.1016/j.biortech.2014.11.011
Jin VL, Evans RD (2010) Microbial 13C utilization patterns via stable isotope probing of phospholipid biomarkers in Mojave Desert soils exposed to ambient and elevated atmospheric CO2. Glob Chang Biol 16:2334–2344. https://doi.org/10.1111/j.1365-2486.2010.02207.x
Jin J, Wang GH, Liu JD, Yu ZH, Liu XB, Herbert SJ (2013) The fate of soyabean photosynthetic carbon varies in Mollisols differing in organic carbon. Eur J Soil Sci 64:500–507. https://doi.org/10.1111/ejss.12030
Kaiser C, Kilburn MR, Clode PL, Fuchslueger L, Koranda M, Cliff JB, Solaiman ZM, Murphy DV (2015) Exploring the transfer of recent plant photosynthates to soil microbes: mycorrhizal pathway vs direct root exudation. New Phytol 205:1537–1551. https://doi.org/10.1111/nph.13138
Kambura AK, Mwirichia RK, Kasili RW, Karanja EN, Makonde HM, Boga HI (2016) Bacteria and archaea diversity within the hot springs of lake magadi and little magadi in Kenya. BMC Microbiol 16:136. https://doi.org/10.1186/s12866-016-0748-x
Keith A, Singh B, Dijkstra FA (2015) Biochar reduces the rhizosphere priming effect on soil organic carbon. Soil Biol Biochem 88:372–379. https://doi.org/10.1016/j.soilbio.2015.06.007
Khodadad CLM, Zimmerman AR, Green SJ, Uthandi S, Foster JS (2011) Taxa-specific changes in soil microbial community composition induced by pyrogenic carbon amendments. Soil Biol Biochem 43:385–392. https://doi.org/10.1016/j.soilbio.2010.11.005
Kumar A, Kuzyakov Y, Pausch J (2016) Maize rhizosphere priming: field estimates using 13C natural abundance. Plant Soil 409:87–97. https://doi.org/10.1007/s11104-016-2958-2
Kumar A, Dorodnikov M, Splettstößer T, Kuzyakov Y, Pausch J (2017) Effects of maize roots on aggregate stability and enzyme activities in soil. Geoderma 306:50–57. https://doi.org/10.1016/j.geoderma.2017.07.007
Kusliene G, Rasmussen J, Kuzyakov Y, Eriksen J (2014) Medium-term response of microbial community to rhizodeposits of white clover and ryegrass and tracing of active processes induced by 13C and 15N labelled exudates. Soil Biol Biochem 76:22–33. https://doi.org/10.1016/j.soilbio.2014.05.003
Ladygina N, Hedlund K (2010) Plant species influence microbial diversity and carbon allocation in the rhizosphere. Soil Biol Biochem 42:162–168. https://doi.org/10.1016/j.soilbio.2009.10.009
Lehmann J, Rillig MC, Thies J, Masiello CA, Hockaday WC, Crowley D (2011) Biochar effects on soil biota - a review. Soil Biol Biochem 43:1812–1836. https://doi.org/10.1016/j.soilbio.2011.04.022
Lu Y, Watanabe A, Kimura M (2002) Contribution of plant-derived carbon to soil microbial biomass dynamics in a paddy rice microcosm. Biol Fertil Soils 36:136–142. https://doi.org/10.1007/s00374-002-0504-2
Luo Y, Durenkamp M, De Nobili M, Lin Q, Devonshire BJ, Brookes PC (2013) Microbial biomass growth, following incorporation of biochars produced at 350 °C or 700 °C, in a silty-clay loam soil of high and low pH. Soil Biol Biochem 57:513–523. https://doi.org/10.1016/j.soilbio.2012.10.033
Luo Y, Yu Z, Zhang K, Xu J, Brookes PC (2016) The properties and functions of biochars in forest ecosystems. J Soils Sediments 16:2005–2020. https://doi.org/10.1007/s11368-016-1483-5
Luo Y, Lin Q, Durenkamp M, Dungait AJ, Brookes PC (2017a) Soil priming effects following substrates addition to biochar-treated soils after 431 days of pre-incubation. Biol Fertil Soils 53:315–326. https://doi.org/10.1007/s00374-017-1180-6
Luo Y, Zang H, Yu Z, Chen Z, Gunina A, Kuzyakov Y, Xu J, Zhang K, Brookes PC (2017b) Priming effects in biochar enriched soils using a three-source-partitioning approach: 14C labelling and 13C natural abundance. Soil Biol Biochem 106:28–35. https://doi.org/10.1016/j.soilbio.2016.12.006
Luo Y, Zhu Z, Liu S, Peng P, Xu J, Brookes PC, Ge T, Wu J (2018) Nitrogen fertilization increases rice rhizodeposition and its stabilization in soil aggregates and the humus fraction. Plant Soil 445:125–135. https://doi.org/10.1007/s11104-018-3833-0
Macdonald LM, Paterson E, Dawson LA, McDonald AJS (2004) Short-term effects of defoliation on the soil microbial community associated with two contrasting Lolium perenne cultivars. Soil Biol Biochem 36:489–498. https://doi.org/10.1016/j.soilbio.2003.11.001
Maestrini B, Nannipieri P, Abiven S (2015) A meta-analysis on pyrogenic organic matter induced priming effect. Glob Chang Biol 7:577–590. https://doi.org/10.1111/gcbb.12194
Malik AA, Martiny JBH, Brodie EL, Martiny AC, Treseder KK, Allison SD (2019) Defining trait-based microbial strategies with consequences for soil carbon cycling under climate change. ISME J 14:1–9. https://doi.org/10.1038/s41396-019-0510-0
Martinsen V, Alling V, Nurida N, Mulder J, Hale S, Ritz C, Rutherford D, Heikens A, Breedveld G, Cornelissen G (2015) pH effects of the addition of three biochars to acidic Indonesian mineral soils. Soil Sci Plant Nutr 61:821–834. https://doi.org/10.1080/00380768.2015.1052985
Mellado-Vázquez PG, Lange M, Bachmann D, Gockele A, Karlowsky S, Milcu A, Piel C, Roscher C, Roy J, Gleixner G (2016) Plant diversity generates enhanced soil microbial access to recently photosynthesized carbon in the rhizosphere. Soil Biol Biochem 94:122–132. https://doi.org/10.1016/j.soilbio.2015.11.012
Meng F, Dungait JAJ, Zhang X, He M, Guo Y, Wu W (2013) Investigation of photosynthate-C allocation 27 days after 13C-pulse labeling of Zea mays L. at different growth stages. Plant Soil 373:755–764. https://doi.org/10.1007/s11104-013-1841-7
Neergaard AD, Porter JR, Gorissen A (2002) Distribution of assimilated carbon in plants and rhizosphere soil of basket willow (Salix viminalis L.). Plant Soil 245:307–314. https://doi.org/10.1023/a:1020414819264
Nicol GW, Leininger S, Schleper C, Prosser JI (2008) The influence of soil pH on the diversity, abundance and transcriptional activity of ammonia oxidizing archaea and bacteria. Environ Microbiol 10:2966–2978. https://doi.org/10.1111/j.1462-2920.2008.01701.x
Novotny EH, Maia CMBDF, Carvalho MTDM, Madari BE (2015) Biochar: pyrogenic carbon for agricultural use - a critical review. Rev Bras Cienc Solo 39:321–344. https://doi.org/10.1590/01000683rbcs20140818
Olsson PA, Baath E, Jakobsen I (1997) Phosphorous effects on the mycelium and storage structures of an arbuscular mycorrhizal fungus as studied in the soil and roots by analysis of fatty acid signatures. Appl Environ Microb 63:3531–3538. https://doi.org/10.1128/aem.63.9.3531-3538.1997
Pausch J, Kuzyakov Y (2018) Carbon input by roots into the soil: quantification of rhizodeposition from root to ecosystem scale. Glob Chang Biol 24:1–12. https://doi.org/10.1111/gcb.13850
Peduruhewa HJ, Gunina A, Tao L, Zhu Z, Kuzyakov Y, Lukas VZ, Guggenberger G, Shen C, Yu G, Singh BP, Pan S, Luo Y, Xu J (2020) Rusty sink of rhizodeposits and associated keystone microbiomes. Soil Biol Biochem 147:107840. https://doi.org/10.1016/j.soilbio.2020.107840
Qiao Y, Miao S, Li N, Han X, Zhang B (2014) Spatial distribution of rhizodeposit carbon of maize (Zea mays L.) in soil aggregates assessed by multiple pulse 13C labeling in the field. Plant Soil 375:317–329. https://doi.org/10.1007/s11104-013-1932-5
Revell KT, Maguire RO, Agblevor FA (2012) Influence of poultry litter biochar on soil properties and plant growth. Soil Sci 177:402–408. https://doi.org/10.1097/ss.0b013e3182564202
Rinnan R, Baath E (2009) Differential utilization of carbon substrates by bacteria and fungi in tundra soil. Appl Environ Microb 75:3611–3620. https://doi.org/10.1128/AEM.02865-08
Rousk J, Bååth E, Brookes PC, Lauber CL, Lozupone C, Caporaso JG, Knight R, Fierer N (2010) Soil bacterial and fungal communities across a pH gradient in an arable soil. ISME J 4:1340–1351. https://doi.org/10.1038/ismej.2010.58
Rousk J, Dempster DN, Jones DL (2013) Transient biochar effects on decomposer microbial growth rates: evidence from two agricultural case-studies. Eur J Soil Sci 64:770–776. https://doi.org/10.1111/ejss.12103
Santos F, Torn MS, Bird JA (2012) Biological degradation of pyrogenic organic matter in temperate forest soils. Soil Biol Biochem 51:115–124. https://doi.org/10.1016/j.soilbio.2012.04.005
Sarker TC, Incerti G, Spaccini R, Piccolo A, Mazzoleni S, Bonanomi G (2018) Linking organic matter chemistry with soil aggregate stability: insight from 13C NMR spectroscopy. Soil Biol Biochem 117:175–184. https://doi.org/10.1016/j.soilbio.2017.11.011
Shannon P, Markiel A, Ozier O, Baliga N, Wang J, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504. https://doi.org/10.1101/gr.1239303
Singh BP, Cowie AL, Smernik RJ (2012) Biochar carbon stability in a clayey soil as a function of feedstock and pyrolysis temperature. Environ Sci Technol 46:11770–11778. https://doi.org/10.1021/es302545b
Six J, Conant RT, Paul EA, Paustian K (2002) Stabilization mechanisms of soil organic matter: implications for C-saturation of soils. Plant Soil 241:155–176. https://doi.org/10.1023/a:1016125726789
Six J, Bossuyt H, Degryze S, Denef K (2004) A history of research on the link between (micro)aggregates, soil biota, and soil organic matter dynamics. Soil Till Res 79:7–31. https://doi.org/10.1016/j.still.2004.03.008
Sohi SP (2012) Carbon storage with benefits. Science 338:1034–1035. https://doi.org/10.1126/science.1225987
Soil Survey Staff (2014) Keys to soil taxonomy, 12th edn. USDA-Natural Resources Conservation Service, Washington, D.C.
Soussana JF, Loiseau P, Vuichard N, Ceschia E, Balesdent J, Chevallier T, Arrouays D (2004) Carbon cycling and sequestration opportunities in temperate grasslands. Soil Use Manag 20:219–230. https://doi.org/10.1111/j.1475-2743.2004.tb00362x
Tavi NM, Martikainen PJ, Lokko K, Kontro M, Wild B, Richter A, Biasi C (2013) Linking microbial community structure and allocation of plant-derived carbon in an organic agricultural soil using 13CO2 pulse-chase labelling combined with 13C-PLFA profiling. Soil Biol Biochem 58:207-215. https://doi.org/10.1016/j.soilbio.2012.11.013
Tisdall JM, Oades JM (1982) Organic matter and water-stable aggregates in soils. J Soil Sci 33:141–163. https://doi.org/10.1111/j.1365-2389.1982.tb01755.x
Tripathi R, Nayak AK, Bhattacharyya P, Shukla AK, Shahid M, Raja R, Panda BB, Mohanty S, Kumar A, Thilagam VK (2014) Soil aggregation and distribution of carbon and nitrogen in different fractions after 41 years long-term fertilizer experiment in tropical rice-rice system. Geoderma 213:280–286. https://doi.org/10.1016/j.geoderma.2013.08.031
Tripathi BM, Stegen JC, Kim M, Dong K, Adams JM, Lee YK (2018) Soil pH mediates the balance between stochastic and deterministic assembly of bacteria. ISME J 12:1072–1083. https://doi.org/10.1038/s41396-018-0082-4
Wang J, Chapman SJ, Yao H (2016) Incorporation of 13C-labelled rice rhizodeposition into soil microbial communities under different fertilizer applications. Appl Soil Ecol 101:11–19. https://doi.org/10.1016/j.apsoil.2016.01.010
Weng Z, Van Zwieten L, Singh BP, Tavakkoli E, Joseph S, Macdonald LM, Rose TJ, Rose MT, Kimber SWL, Morris S, Cozzolino D, Araujo JR, Archanjo BS, Cowie A (2017) Biochar built soil carbon over a decade by stabilizing rhizodeposits. Nat Clim Chang 7:371–376. https://doi.org/10.1038/nclimate3276
Whitman T, Enders A, Lehmann J (2014) Pyrogenic carbon additions to soil counteract positive priming of soil carbon mineralization by plants. Soil Biol Biochem 73:33–41. https://doi.org/10.1016/j.soilbio.2014.02.009
Yu Z, Chen L, Pan S, Li Y, Kuzyakov Y, Xu J, Brookes PC, Luo Y (2018) Feedstock determines biochar-induced soil priming effects by stimulating the activity of specific microorganisms. Eur J Soil Sci 69:521–534. https://doi.org/10.1111/ejss.12542
Yuan H, Zhu Z, Liu S, Ge T, Jing H, Li B, Liu Q, Lynn TM, Wu J, Kuzyakov Y (2016) Microbial utilization of rice root exudates: 13C labeling and PLFA composition. Biol Fertil Soils 52:615–627. https://doi.org/10.1007/s00374-016-1101-0
Zhang H, Ding W, Luo J, Bolan N, Yu H (2015) The dynamics of glucose-derived 13C incorporation into aggregates of a sandy loam soil following two-decade compost or inorganic fertilizer amendments. Soil Till Res 148:14–19. https://doi.org/10.1016/j.still.2014.11.010
Zhang K, Chen L, Li Y, Brookes PC, Xu J, Luo Y (2016) The effects of combinations of biochar, lime, and organic fertilizer on nitrification and nitrifiers. Biol Fertil Soils 53:77–87. https://doi.org/10.1007/s00374-016-1154-0
Zhou G, Xu X, Qiu X, Zhang J (2019) Biochar influences the succession of microbial communities and the metabolic functions during rice straw composting with pig manure. Bioresour Technol 272:10–18. https://doi.org/10.1016/j.biortech.2018.09.135
Zhu Z, Ge T, Hu Y, Zhou P, Wang T, Shibistova O, Guggenberger G, Su Y, Wu J (2017) Fate of rice shoot and root residues, rhizodeposits, and microbial assimilated carbon in paddy soil - part 2: turnover and microbial utilization. Plant Soil 416(1–2):24–257. https://doi.org/10.1007/s11104-017-3210-4
Acknowledgments
This study was supported financially by the National Natural Science Foundation of China (41520104001; 41671233) and the Fundamental Research Funds for the Central Universities (2019QNA6012). The contribution of Amit Kumar was supported by the German Academic Exchange Service (DAAD). We acknowledge the Ningbo Urban Environment Observation and Research Station, Chinese Academy of Sciences for technical assistance. We thank anonymous reviewers and the editor for constructive comments which significantly improved the quality of this paper.
Funding
This study was supported financially by the National Natural Science Foundation of China (41520104001; 41671233) and the Fundamental Research Funds for the Central Universities (2019QNA6012). The contribution of Amit Kumar was supported by the German Academic Exchange Service (DAAD).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Highlights
• Biochar gave no difference of photosynthetic-13C allocation in plant shoot and root but reduced 13C into aggregates
• The relative abundance of bacteria increased owing to the reduced stress by biochar shifted soil variables, like pH.
• Biochar suppressed root-associated fungi due to reducing root biomass and rhizodeposits.
• Less 13C accumulation into aggregates by biochar might be due to the decreased fungi abundance that diminished aggregation thus leaving rhizodeposits unprotected by bacterial utilization.
• Biochar exerted opposite effect on fungal-bacteria growth and their interaction within aggregates could determine rhizodeposit-C stabilization.
Supplementary information
ESM 1
(DOCX 16577 kb)
Rights and permissions
About this article
Cite this article
Chen, Z., Kumar, A., Fu, Y. et al. Biochar decreased rhizodeposits stabilization via opposite effects on bacteria and fungi: diminished fungi-promoted aggregation and enhanced bacterial mineralization. Biol Fertil Soils 57, 533–546 (2021). https://doi.org/10.1007/s00374-020-01539-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00374-020-01539-9