Abstract
Background and aims
The turnover of plant- and microbial- derived carbon (C) plays a significant role in the soil organic C (SOC) cycle. However, there is limited information about the turnover of the recently photosynthesized plant- and soil microbe-derived C in paddy soil.
Methods
We conducted an incubation study with four different 13C–labeled substrates: rice shoots (Shoot-C), rice roots (Root-C), rice rhizodeposits (Rhizo-C), and microbe-assimilated C (Micro-C).
Results
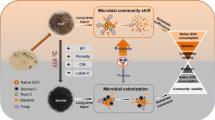
Shoot- and Root-C were initially rapidly transformed into the dissolved organic C (DOC) pool, while their recovery in microbial biomass C (MBC) and SOC increased with incubation time. There were 0.05%, 9.8% and 10.0% of shoot-C, and 0.06%, 15.9% and 16.5% of root-C recovered in DOC, MBC and SOC pools, respectively at the end of incubation. The percentages of Rhizo- and Micro-C recovered in DOC, MBC, and SOC pools slowly decreased over time. Less than 0.1% of the Rhizo- and Micro-C recovered in DOC pools at the end of experiment; while 45.2% and 33.8% of Rhizo- and Micro-C recovered in SOC pools. Shoot- and Root-C greatly increased the amount of 13C–PLFA in the initial 50 d incubation, which concerned PLFA being indicative for fungi and actinomycetes while those assigning gram-positive bacteria decreased. The dynamic of soil microbes utilizing Rhizo- and Micro-C showed an inverse pattern than those using Shoot- and Root-C. Principal component analysis of 13C–PLFA showed that microbial community composition shifted obviously in the Shoot-C and Root-C treatments over time, but that composition changed little in the Rhizo-C and Micro-C treatments.
Conclusions
The input C substrates drive soil microbial community structure and function with respect to carbon stabilization. Rhizodeposited and microbial assimilated C have lower input rates, however, they are better stabilized than shoot- and root-derived C, and thus are preferentially involved in the formation of stable SOC in paddy soils.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Soil carbon (C) turnover is a fundamental process in ecosystem functioning (Ruf et al. 2006). As primary decomposers, soil microorganisms turn over C within the soil via decomposition, immobilization, and resynthesis of organic substances (Paterson et al. 2011). The rates and pathways of soil C turnover are controlled by the composition of the microbial community, which, in turn, is typically governed by C availability (Glanville et al. 2012; Wang et al. 2014b).
Soil microorganisms play a vital role in the formation and maintenance of soil structure, organic matter decomposition, biogeochemical cycling, and plant nutrient availability (Waldrop and Firestone 2004; Falkowski and Godfrey 2008). Because of differences in growth forms, stoichiometric requirements, and functional abilities, soil microbial decomposers are influenced by soil C quantity, quality, and input regularity (Singh et al. 2007; Loeppmann et al. 2016). Soil C is mainly composed of old, indigenous soil organic C (SOC), but also includes recent products of photosynthesis. According to Lu et al. (2002) and Johnson et al. (2006), the latter C is the primary source for soil microbial metabolic and catabolic activity, thus contributing to soil respiration and formation of SOC. However, the composition of these fresh exogenous substrates vary greatly in complexity, ranging from low weight compounds such as amino acids, sugars, and peptides, to recalcitrant polymeric compounds such as cellulose, hemicellulose, lignin, and proteins (Lu et al. 2002; Baumann et al. 2009). Plants are the primary C sources in soil, and deliver C to the soil as plant residues, dominated by cellulose and lignin, which are broken down into smaller units by exoenzymes. However, only a relatively small group of microorganisms such as fungi and actinomycetes produce these enzymes for decomposition of the recalcitrant compounds (Brant et al. 2006; Weintraub et al. 2007). In contrast, plant rhizodeposits, consisting of root exudates, mucilage, sloughed-off cells and tissue, cell lysates, and root debris, are a readily available source of C for soil microorganisms (Kuzyakov and Larionova 2005). In addition to plants, soil photoautotrophic microorganisms, which are able to assimilate CO2, contribute to soil C formation (Yuan et al. 2012; Ge et al. 2013). The microbial biomass C and the metabolites they release into the soil are mainly recycled through further microbial metabolism or stabilized by soil minerals, which may have lower rates of turnover and thus effectively improve soil C storage (Bastida et al. 2013; Schurig et al. 2013). Therefore, quantification of the fate of different C forms in the soil and the microbial role in C turnover is essential for understanding the dynamics of C cycling in paddy soil ecosystems.
Members of the microbial community are differentially involved in the assimilation of plant residue-derived C (Derrien et al. 2014). At the initial stage of plant residue decomposition, easily available and water-soluble C compounds are released into the soil (Berg 1986; Müller et al. 2016). After these substrates are depleted, more-complex compounds are used, a change associated with more-intensive interactions between soil microorganisms (Lu et al. 2002; Dilly et al. 2004; Brant et al. 2006; Baumann et al. 2009). Bacteria dominate crop residue decomposition in the initial phases; gram-positive bacteria use both easily available and recalcitrant compounds, whereas gram-negative bacteria preferentially process low-molecular-weight compounds. Fungi dominate in the later stages (Waldrop and Firestone 2004; Marschner et al. 2011). Saprotrophic fungi produce a wide range of extracellular enzymes, allowing decomposition of the recalcitrant substrates that other organisms are unable to degrade (Boer et al. 2005). Input of low-molecular-weight organic C such as rhizodeposits and sugars could stimulate an opportunistic subset of the bacterial biomass (Fontaine et al. 2011; Li et al. 2015), and the labile substrates can be directly metabolized or recycled for energy production and microbial growth, leading to a decline in microbial abundance, species richness, and diversity (Schurig et al. 2013; Yuan et al. 2016). Accordingly, also for paddy soils differences in substrate quantity, quality, and pathways of C input to soil and the microbial utilization of this C are essential for C turnover and sequestration (Lu et al. 2003; Lal 2004; Ge et al. 2013).
The utilization of the substances in paddy soils are different from upland soils because they are regularly flooded and intermittently irrigated (Kögel-Knabner et al. 2010). Differences in the redox potentials maybe contribute to changes in the physico-chemical soil properties (Nikolausz et al. 2008), thereby affecting the microbial community composition and their succession. Paddy soils are characterized by relatively slow C turnover (Wu et al. 2012); however, only few studies have shown the biological processes determining soil organic matter stabilization and turnover in paddy soil, especially at the microbial level (Gale et al. 2000). Better understanding of the different forms of C input (root, shoot, rhizodeposits, and microbial assimilated C) into paddy soils, their turnover and microbial utilization are critical for understanding global C cycling and the ecological functions of paddy ecosystems. We hypothesized that (1) the turnover of C substrates was affected by the amount of available C and the activity of key microbes in the soil, and (2) the input of shoots and roots into soil could significantly alter the soil autochthonous microbial community because of the complex composition of plants residues, while the microbial community structure would change relatively little during the turnover of rhizodeposited and microbial-assimilated C. We tested these hypothesis by carrying out a 300-day incubation study using 13C–labeled rice plant residues (shoots and roots), rhizodeposits, and microbe-assimilated C in paddy soils. We quantified C turnover and the dynamics of the soil microbial community structure by tracking the 13C–labeled photosynthesized substrates utilized by microbes using 13C–phospholipids fatty acid (PLFA) analysis.
Materials and methods
Study site and basic soil properties
The study site was a rice field (113°19′52″ E, 28°33′04″ N; 80 m above sea level), located at the Changsha Research Station for Agricultural and Environmental Monitoring, Hunan, China. The soil type is typical Stagnic Anthrosol developed from highly weathered granite, and the climate is subtropical with a mean annual temperature of 17.5 °C and yearly rainfall of 1300 mm. Moist soil samples were collected from the plow layer (0–20 cm) and sieved <4 mm to remove visible plant residues. The soil contained 18.1 g kg−1 organic C, 1.8 g kg−1 total nitrogen, and 0.5 g kg−1 total phosphorus, and had a pH of 5.6 (1:2.5, soil/water ratio).
Production of 13C–labeled substrates
Rice growth conditions, 13CO2 labeling, and 13C–labeled substrate collection were as described by Zhu et al. (2016). Briefly, 60 pots were each filled with 1 kg dry soil; of these, 40 pots were planted with three 30-d-old rice (Oryza sativa L. ‘Zhongzao 39’) seedlings each, and the remaining 20 pots were left unplanted. Thirty pots (20 planted and 10 unplanted) were used for 13C labeling in a gas-tight growth chamber system (110 cm length, 250 cm width, 180 cm height), where they were exposed to 13CO2 via fumigation for 18 days (between May 14 and 31, 2013). The surface of each planted pot was covered with black plastic sheeting to prevent algal photosynthesis, thereby ensuring that only the rice shoots were exposed to 13CO2. The unplanted pots were not covered, so that the soil in the pots was directly exposed to 13CO2 and photoautotrophic soil microorganisms could assimilate atmospheric 13CO2. The other 30 pots (20 planted and 10 unplanted) were used for measuring the natural 13C abundance as a control. Because these pots were not fumigated, there was no need to cover their surfaces with black plastic sheeting. Pots were placed outdoors, with unlabeled pots 10–15 m away from 13C–labeled pots. All pots were watered every few days to maintain a water depth of 2–3 cm above the soil surface until harvest. Weeds were manually removed from all pots every week.
The CO2 concentration in the growth chamber was measured using an infrared analyzer (Shsen-QZD, Qingdao, China) and maintained between 360 and 380 μl L−1. 13CO2 was generated through acidification of Na2 13CO3 (1.0 M, 99 atom% 13C; Cambridge Isotope Laboratories, Tewksbury, MA, USA) with H2SO4 (0.5 M) in a beaker placed inside the growth chamber. During labeling, CO2 was released only when the concentration in the chamber was lower than 360 μl L−1. Conversely, when the CO2 concentration in the chamber was higher than 380 μL L−1, a switch diverted the gas flow to pass through CO2 traps (1 M NaOH solution) to absorb excess CO2.
Labeled and unlabeled rice plants and soil were sampled destructively after labeling. Rice shoots were removed from the shoot bases, and roots were separated from the soil by washing with deionized water. Shoots and roots were dried at 60 °C for 48 h and cut into pieces <5 mm. To obtain rhizodeposited 13C, the roots and soil were separated by two steps (slightly modified from Lu et al. 2002). First, roots were separated from soil by washing through a 2-mm sieve with 1.0 L distilled water. Root debris >2-mm in size was collected and combined with the root samples. Then, soil slurries, consisting of soil and wash water, were mixed well and centrifuged at 13,000×g for 20 min, the obtained soil was considered rhizosphere soil. After root debris was removed, well-mixed rhizosphere soil was used in the incubation experiment. Meanwhile, labeled unplanted soils were collected to obtain the 13C originating from assimilation of 13CO2 by soil phototrophic microorganisms. The labeled unplanted soil was mixed thoroughly and used in the incubation experiment. The C content and values of 13C atom percent (atom%) of the four photosynthesized C substrates are shown in our previous study (Zhu et al. 2016).
Incubation study
For the incubation experiment, the following treatments were used: (1) unlabeled soil containing 13C–labeled shoots (Shoot-C), (2) unlabeled soil containing 13C–labeled roots (Root-C), (3) 13C–labeled rhizosphere soils (Rhizo-C), (4) 13C–labeled unplanted soil containing only microbial-assimilated 13C (Micro-C), and (5) unlabeled and unplanted soil (CK). Three additional incubation treatments were carried out as to measure the natural 13C abundance and thus calculate the 13C atom% excess: (1) unlabeled soil containing unlabeled shoots, (2) unlabeled soil containing unlabeled roots, and (3) unlabeled soil containing unlabeled rhizodeposited C. For Shoot-C and Root-C, 0.6 g shoots or roots, respectively, were homogenized with 100 g unlabeled soil (oven-dried weight) with a water content of 50% for a final residue concentration of 6 g kg−1 (dry) soil. For Rhizo-C and Micro-C, 150 g of fresh soil (i.e., 100 g oven dry weight) was directly weighed into 500 mL containers. Samples were then placed into 500-mL containers with 100 mL deionized water to create a 1–2 cm water layer. The excess 13C per 100 g dry soil in the container was 11.4, 5.75, 1.61, and 0.49 mg in the treatments of Shoot-C, Root-C, Rhizo-C, and Micro-C, respectively (Zhu et al. 2016). Each treatment was performed four times and all samples were incubated at 25 °C in the dark for 300 d.
Soil preparation
All four containers from each treatment were destructively sampled at 0, 5, 50, 100, 200, and 300 d. Plant residues were separated from soil by gently shaking in 300 mL deionized water and passing through a 1 mm sieve. The resultant soil slurries, consisting of soil and wash water, were mixed well and centrifuged at 13,000×g for 20 min. Supernatant solutions were filtered through a 0.45-μm fiberglass filter. The organic C contained in the solution was considered water-soluble organic C. The fresh soil slurries were mixed thoroughly and divided into three portions. The soils sampled at 5, 50 and 300 d were immersed immediately in liquid nitrogen and stored at −70 °C until PLFA analysis. Another aliquot was used for determination of microbial biomass C (MBC), and 0.5 M K2SO4-extractable organic C. The total dissolved organic C (DOC) was the sum of the water-soluble organic C and 0.5 M K2SO4-extractable organic C; extractions were carried out consecutively. The remaining portion was air-dried, ground, sieved through a 100 mesh sieve, and used to determine δ13C of SOC and other soil physiochemical properties.
Soil and microbial C and δ13C analysis
The C content of bulk soil, rhizosphere soil, soil containing microbial-assimilated C, and shoot and root samples were analyzed using an automated C/N analyzer (Vario MAX, Elementar Analysensysteme GmbH, Hanau, Germany). Prior to analysis for δ13C, dry samples of all sampled soils, shoots, and roots were ground to a fine powder. The stable C isotope ratios of plant and soil materials were measured using a MAT253 isotope ratio mass spectrometer coupled to an elemental analyzer FLASH HT (ThermoFisher Scientific, Waltham, MA, USA). Soil MBC, together with water-soluble C extracted by 0.5 M K2SO4, were measured by the fumigation extraction method (Wu et al. 1990). Briefly, samples of wet soil (equivalent to about 20 g of oven-dried soil) were amended with 2 ml alcohol-free CHCl3 from the surface of the slurries soil and then additionally fumigated by exposing the soil to alcohol-free CHCl3 vapor for 24 h in a vacuum desiccator (Wu et al. 1990). The residual CHCl3 was removed by vacuuming 5–10 times, each for about 5 min. Then, the wet fumigated and non-fumigated soils were extracted with 80 ml of 0.05 M K2SO4 by shaking at 250 rpm for 30 min. The suspensions were filtered through Whatman No. 42 filter papers. Organic C in the K2SO4 extracts was analyzed by an automated procedure using a total carbon analyzer (Phoenix 8000, Tekmar-Dohrmann Co., USA) and the isotope signature (δ13C) in the extracts was measured by an isotope ratio mass spectrometer (MAT253) equipped with an elemental analyzer (FLASH 2000; Thermo-Fisher Scientific, Waltham, MA, USA) after freeze-drying and the grinding the dried salt with a mortar and pestle to a fine powder.
PLFA extraction and analysis
Soil microbial PLFA were extracted, fractionated, and purified following the methods described by Yuan et al. (2016), which were modified from Buyer et al. (2010). Briefly, soil samples were freeze-dried, ground, and passed through a 2 mm sieve, then approximately 2 g soil was extracted twice using a 22.8-mL single-phase mixture of chloroform:methanol:citrate buffer (1:2:0.8 v/v/v, 0.15 M, pH 4.0). Phospholipids were then separated from neutral lipids and glycolipids on a silica acid column (Supelco, Bellefonte, PA, USA). Methyl nonadecanoate fatty acid (19:0) was added prior to derivatization as an internal standard to quantify the concentrations of phospholipids. Following methylation of the phospholipids, the PLFA methyl esters (PAMES) were separated and identified using a gas chromatograph (GC; N6890; Agilent, Santa Clara, CA, USA) fitted with a MIDI Sherlock microbial identification system (Version 4.5; MIDI, Newark, DE, USA). The δ13C of individual PLFAs were analyzed by gas chromatography combustion isotope ratio mass spectrometry (GC–C–IRMS) using a TRACE GC Ultra gas chromatograph with combustion column attached via a GC Combustion III to a Delta V Advantage isotope ratio mass spectrometer (Thermo Finnigan, Bremen, Germany). An internal standard of 19:0 methyl ester was used for assurance of quantification. The following PLFAs were used as markers for specific groups of bacteria: i14:0, i15:0, a15:0, i16:0, i17:0, and a17:0 for gram-positive bacteria; 16:1ω5c, 17:1ω8c, cy17:0, and cy19:0 for gram-negative bacteria (Zelles 1997, 1999). The 18:1ω9c and 10Me16:0, 10Me17:0, and 10Me18:0 PLFAs were used as fungal and actinomycetes biomarkers, respectively, while non-specific (universal) PLFA were represented by 14:0, 15:0, 16:0, 17:0 and 18:0 straight-chain acids.
Calculations and statistical analysis
The δ13C values of plant residues, rhizodeposits, microbial-assimilated C, bulk soils, DOC, MBC, SOC, and PLFAs were measured as δ (‰) relative to the Pee Dee Belemnite (PDB; 13C, 0.0111802) standard and further expressed as atom percent (atom%), where
The incorporation of 13C (excess 13C) into plant residues, rhizodeposits, microbe-assimilated C, bulk soils, DOC, SOC, and PLFA was calculated as
where (atom% 13C)L and (atom% 13C)UL are the atom% 13C in labeled and unlabeled samples, respectively, and Csample is the C content of each sample.
The 13C incorporated into microbial biomass (13C–MBC) was calculated as the difference in 13C between fumigated and unfumigated soil extracts, divided by a factor of 0.45 (Wu et al. 1990; Ge et al. 2012), according to
where f indicate fumigated and uf unfumigated soil extracts, L indicate extracts from labeled sampled and UL those from unlabeled samples, while Cf and Cuf are the total C contents of the fumigated and unfumigated soil extracts, respectively.
The amount of 13C incorporation in each PLFA (13C excess, mg kg−1) was determined using a mass balance approach:
where (atom% 13C)PLFA,L and (atom% 13C)PLFA,UL indicate PLFA extracts from labeled samples and unlabeled samples, respectively. We calculated the relative 13C distribution (%) in each specific microbial group according to
where 13CPLFA-group is the amount of 13C–PLFA incorporated into the specific microbial group, and ∑13CPLFA is the total amount of 13C–PLFA incorporated into soil microbes.
The percentage of initial substrate-derived 13C that was incorporated (%13CINCORP) into DOC, MBC, SOC, and microbial groups was expressed as the percentage of 13C recovery on each sampling day according to
where 13Csample is the sum of 13C–DOC, 13C–MBC, 13C–SOC, and 13C–PLFA, and initial 13C is the total amount of 13C input in soil.
Analysis of variance (ANOVA) in conjunction with Duncan’s multiple range test (P < 0.05) and correlation analysis were carried out using SPSS 17 (SPSS Inc., Chicago, IL, USA). Principal components analysis (PCA) was performed with CANOCO 5.0 for Windows (Microcomputer Power, Ithaca, NY, USA). Figures were created using Origin 8.5 (OriginLab, Northampton, MA, USA).
Results
Dynamic of carbon substrates in paddy soil
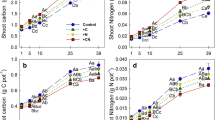
The incorporation of shoot- and root-derived C in the DOC pool reached 0.38% in both treatments at 50 d incubation, and thereafter declined to 0.05% and 0.06%, respectively, after 200 d incubation (Fig. 1a). The incorporation of C from rhizodeposits and microbial-assimilated C into DOC showed similar patterns, but the highest relative proportions of C allocated to DOC were approximately only a quarter of those from Shoot-C and Root-C. Incorporation of the four C substrates into DOC were all similar after 200 d incubation.
Incorporation of initial added 13C into dissolved organic carbon (DOC) (a), microbial biomass carbon (MBC) (b) and total soil organic carbon (SOC) (c) during 300 days incubation. Error bars represent the standard error of the mean (n = 4). Abbreviations: Shoot-C, unlabeled paddy soil containing 13C–labeled shoots; Root-C, unlabeled paddy soil containing 13C–labeled roots; Rhizo-C, labeled paddy soil containing 13C–labeled rice rhizodeposited C; Micro-C, labeled paddy soil containing 13C–labeled microbial-assimilated C
The incorporation of shoot- and root-derived 13C into the MBC pool increased from 7.6% and 8.6%, respectively, at 5 d, to 9.8% and 15.9% at the end of the incubation (Fig. 1b). Microbial-assimilated C incorporation into MBC was the highest of all C sources at all sampling points, and gradually decreased from 23.0% to 18.4% over the entire incubation period. The percentage of rhizodeposited 13C incorporated into MBC was intermediate between that of the Micro-C treatment and those of the shoot- and root-derived C; it decreased linearly during the whole incubation period.
The percentages of 13C incorporation into SOC in the Shoot-C and Root-C treatments were much lower than those of Rhizo-C and Micro-C (Fig. 1c). At the beginning of the incubation 61.0% of the 13C was incorporated into SOC for Rhizo-C, while it was 71.1% for Micro-C. Thereafter, the proportions declined to 45.2% and 33.8%, respectively, at the end of incubation. In contrast, the proportions of shoot- and root-C incorporated into SOC gradually increased to 15.1% and 33.1%, respectively, at 50 d, and then decreased to 10.0% and 16.5%, respectively, at the end of the incubation.
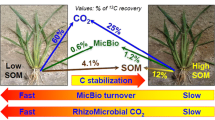
We calculated the recovery of 13C in each C pool based on the distribution of the 13C–labeled substrate into the three soil fractions (Fig. 1a-c) and on the mineralization of the four C substrates as reported in our previous study (Zhu et al. 2016) (Fig. 2). The proportions of 13C recovery were quite different in the four treatments. There was still undecomposed shoot- and root-C after 300 d incubation (37.8% and 9.0%, respectively), which was possibly due to their high contents of recalcitrant components (Baumann et al. 2009). Further, 38.3% and 54.5% of the Shoot-13C and Root-13C, respectively, were emitted as CO2 and CH4, and 12.1% and 19.8% were recovered in SOC. In contrast, the percentages of rhizodeposited C and microbial-assimilated C recovered in SOC (71.9% and 55.6%) and MBC (17.7% and 34.8%) were much greater than in case of shoot- and root-derived C, and only about 10% of the substrate C was emitted as CO2 and CH4 (Fig. 2).
Incorporation of carbon from different substrates into the soil microbial community
Total PLFA ranged from 16.4 to 19.8 mg C kg−1 and decreased gradually over the incubation period in all four treatments. The contents of 13C–PLFAs in the Shoot-C and Root-C treatments were significantly higher than those of Rhizo-C and Micro-C treatments on all sampling days (P < 0.05; Table 1, Table 3). In the Shoot-C and Root-C treatments, the 13C incorporation into PLFA peaked at 112.1 and 139.2 μg C kg−1, respectively, after 50 d incubation and then decreased towards the end of the incubation to 89.5 and 51.8 μg C kg−1. In the Rhizo-C and Micro-C treatments only 38.5 and 11.8 μg C kg−1, respectively, of the 13C was incorporated into PLFA at 5 d, and further decreased to 16.7 and 3.9 μg C kg−1 at 300 d.
The 13C from the four different C substrates were not evenly distributed among microbial groups, indicating that soil microorganisms differed in their uptake and utilization of C substrates (Fig. 3). For all three sampling days and in all four treatments more than 30% of the 13C recovered from the PLFA was incorporated into gram-positive bacteria. But this gradually decreased over time in the Shoot-C and Root-C treatments, whereas it increased in the Rhizo-C and Micro-C treatments. Conversely, in the Shoot-C and Root-C treatments, the 13C recovery from gram-negative bacteria, fungi, and actinomycetes increased over time, while there was an inverse pattern in the Rhizo-C and Micro-C treatments. Furthermore, the amounts of 13C incorporated into universal PLFAs decreased during the 300 d incubation in the Shoot-C and Root-C treatments, but increased in the Rhizo-C and Micro-C treatments.
Percentage of total 13C labelled phospholipid fatty acids (13C–PLFA) recovered from each microbial group in the Shoot-C, Root-C, Rhizo-C, and Micro-C treatments after 5, 50, and 300 days of incubation. PLFA were classified as gram-positive bacteria (G+), gram-negative bacteria (G-), universal, fungi, or actinomycetes. Error bars represent standard error of the mean (n = 4)
The relative incorporation of 13C into specific biomarker C-PLFA was substrate- and incubation time-dependent (Tables 2, 3). In the Shoot-C treatment, the ratios of 13C incorporation into fungi to that into bacteria significantly decreased over time. However, in Rhizo-C and Micro-C treatments these ratios significantly increased to 0.46 and 0.27, respectively, at 50 d and then decreased. Further, the ratio of 13C–PLFA indicative of gram-positive bacteria to those indicative of gram-negative bacteria decreased sharply during the initial 50 d incubation in the Shoot-C and Root-C treatments, while inverse patterns were observed in Rhizo-C and Micro-C treatments. Finally, the ratio of 13C–PLFA assigned to fungi and those assigned to gram-positive bacteria was lower than that of fungi to gram-negative bacteria in all treatments. These ratios decreased with incubation time.
To determine the soil microbial composition in response to the C substrates, total PLFA and 13C–PLFA data were subjected to PCA (Fig. 4). The composition of C sources-dependent microbe communities (13C–PLFA) was clearly grouped by sampling date, whereas the overall soil microbial diversity (PLFA) formed a single group. The 13C utilized by different groups of soil microbes was also substrate-dependent (Fig. 5). PCA of the 13C–PLFA profiles in the Shoot-C and Root-C treated soil samples revealed that the microbial community structure became more scattered with increasing incubation time, while that of the Rhizo-C and Micro-C treated soil samples clustered together and showed no changes with incubation time. This suggests that rather different soil microbial communities degraded the four different C substrates during the 300 d incubation.
Discussion
C turnover of different substrates
The origin and quantity of the input substrates to soil were the primary factors controlling C turnover and distribution in different soil C pools. Initial 13C incorporation into soil C pools was significantly affected by the added C sources and incubation times (P < 0.05; Table 3). The Shoot- and Root-C treatments were one-time C inputs into soil, and there was abundant C and nutrients provided to soil microbes in the initial stage of decomposition (Jingguo and Bakken 1997; Devêvre and Horwáth 2000). As a result, shoot- and root-C were rapidly incorporated into the DOC pool (Fig. 1), which represents a small but labile and rapidly replenished fraction of soil organic C (Jones et al. 2005; Boddy et al. 2007). With the stimulation of the increased DOC, the soil microbes accelerated the decomposition of new input substrates, leading to an increase in the shoot- and root-13C incorporation into MBC in the first 200 d of incubation (Fig. 1). Cotrufo et al. (2013) provided evidence that labile plant C is utilized efficiently by microorganisms and thus becomes the main precursor of stable soil organic matter (SOM) by promoting aggregation and chemical bonding to minerals. This was consistent with our study, where the SOC derived from shoots and roots increased along with the increase of 13C–DOC and 13C–MBC in the first 50 d incubation (Fig. 1). However, after the exhaustion of the water-soluble and labile C, more recalcitrant components of plant residues are utilized (Lu et al. 2003; Williams et al. 2006). After 50 d incubation, percentages of 13C derived from shoots and roots that were incorporated into DOC sharply decreased (Fig. 1), suggesting that the contribution of the remaining recalcitrant components of shoot- and root-C to the available soil C pool had slowed down (Baumann et al. 2009). Subsequently, the activity of soil microbes would be reduced because of the lack of sufficient C and energy. Eventually, less than 20% of the shoot- and root-C were incorporated into SOC, indicating that the active C pools play a pivotal role in plant residue turnover, and that the efficiency of the plant residues assimilation into stable soil C pool is low.
In contrast to shoot- and root-C, rhizodeposited and microbe-assimilated C were more efficiently metabolized by soil microbes or transferred into SOC (Lu et al. 2003; Ge et al. 2013). In the Rhizo-C and Micro-C treatments, the 13C incorporated into DOC, MBC, and SOC gradually decreased over time (Fig. 1), a distinctly different pattern from that of Shoot- and Root-C which prevailed throughout the whole incubation. This indicated that parts of rhizodeposited and microbe-assimilated C existed as relatively labile forms at the beginning of the incubation period (Lu et al. 2003). However, these C inputs were mainly distributed to the MBC and SOC pools over the whole incubation period, indicating that the rhizodeposits and microbial-assimilated C were stabilized by interactions with minerals (Mikutta and Kaiser 2011; Schurig et al. 2013) and intensive internal recycling or cross-feeding (Gunina et al. 2014; Müller et al. 2016).
Utilization of C substrates by microbes in paddy soil
The incorporation of 13C among PLFA describes how C is distributed among the ‘active’ microbial population (Wang et al. 2014b; Yuan et al. 2016). PCA revealed that 13C–PLFA were clearly grouped by sampling date, whereas the overall PLFA formed a single group (Fig. 4). Consequently, input of fresh C to soil more strongly affected the microbes utilizing easily available exogenous C than it did the overall microbial community (Waldrop et al. 2012; Wang et al. 2014b). In the Shoot-C and Root-C treatments, there were distinct differences in the incorporation of plant residue-derived C into the different microbial groups (Table 1, Fig. 3). The dominant ‘active’ microbes, gram-positive bacteria, decreased gradually, while fungi and actinomycetes increased over the three sampling dates (Table 2, Fig. 3). This was consistent with previous reports that recalcitrant litter is often associated with higher fungal and actinomycete biomass, which could be involved in processing recalcitrant compounds such as lignin and even SOC (Waldrop et al. 2012; Spohn et al. 2016). In the Rhizo-C and Micro-C treatments, the recovery of 13C in PLFA decreased over time (Table 1), which was consistent with the pattern of 13C incorporation to MBC and SOC, suggesting that with the decreasing amount of available 13C in the soil the microbes shifted to utilize SOC (Brant et al. 2006; Wang et al. 2014b). The changes of ‘active’ microbes in the treated soil showed an inverse pattern compared with that of the Shoot-C and Root-C treatments (Table 2, Fig. 3). Gram-positive bacteria increased gradually, while gram-negative bacteria, fungi, and actinomycetes decreased over time. Wang et al. (2014a) suggested that Gram-positive bacteria have a primary role in C cycles. Rubino et al. (2010) and Dungait et al. (2011) showed that 13C derived from exudates and glucose was primarily incorporated into gram-positive bacteria rather than gram-negative bacteria. In line with these authors, our observation suggests that gram-positive bacteria play a leading role in utilizing the rhizodeposited and microbe-assimilated C. Alternatively, they may have got enriched during cross-feeding (Seth and Taga 2014).
PCA showed that microbial community composition shifted in the Shoot-C and Root-C treatments across the three sampling points, whereas the community composition change was slight in the Rhizo-C and Micro-C treatments (Fig. 5). The changes in microbial community structure according to decomposition stage (or substrate availability) could be explained by the different life strategies of microorganisms, i.e., the r- and K-strategies (Waldrop and Firestone 2004; Chen et al. 2014). At the initial incubation stage, the substrates supplied sufficient C and nutrients for microbial r-strategies, and in the Shoot-C and Root-C treatments the large amount of available C released to the soil activated the soil microbes and increased the MBC and PLFAs. Corresponding to the recalcitrant C that was dominant at the end of incubation, microbial activity declined and the key functional microbial groups changed. For example, the fungi and actinomycete PLFA increased and the ratio of gram-positive PFLA to gram-negative PFLA decreased (Table 2). This indicates that the input of plant residues into soil affects the activity of autochthonous microbes, as does the higher input amount and varied composition the different substrates offer to soil microbes during the incubation period (Kramer et al. 2012; Wang et al. 2014b). However, the amount of rhizodeposited and microbe-assimilated C did not significantly change over time, and their recovery in MBC and PLFA were also relatively constant (Table 1; Fig. 1). Rhizodeposits consisted mostly of low-molecular-weight sugars and acids that are highly bioavailable and that can be incorporated into the microbial community just a few hours after assimilation aboveground (Shahzad et al. 2015; Yuan et al. 2016). As a result, the rhizodeposits were mainly cycled internally among the soil microbes during incubation, and the relatively small amount of rhizodeposit input into the soil did not significantly stimulate the growth of soil microbes (Waldrop and Firestone 2004; Tian et al. 2013). Therefore, the microbial community composition was only slightly changed during the incubation period. The microbial-assimilated C was mainly in the form of MBC and SOC. After death of the organisms some microbial components are more resistant to decomposition and contribute to the more-stabilized SOC pool (Liang and Balser 2008; Bol et al. 2009). This might explain why the soil microbial community composition did not distinctly change during the incubation of Micro-C treated soil.
General implications for C stabilization of different substrates
Photosynthetic carbon substrates were the main sources of the organic C stabilized in soil. However, their quantity, quality, and input frequency all affect their fates in paddy soil and the efficiency of contributing to soil carbon sequestration (Lal 2004; Pan et al. 2004; Ge et al. 2012). As shown in Fig. 2, the distribution of four typical photosynthetic C substrates in paddy soil were distinct after 300 d incubation. While only 12.1% of shoot C and 16.6% of root C were recovered in SOC, 71.9% of rhizodeposited C and 55.5% of microbial-assimilated C were recovered. This suggests that rhizodeposited C and microbial-assimilated C are more efficient in SOC stabilization. We attribute this to the following reasons: residues of shoots and roots, which often enter the soil simultaneously, are characterized by a high C/N ratio that is beyond of the microbes’ stoichiometric requirements (Sinsabaugh et al. 2013; Alberti et al. 2015). As a result, most of the C from these sources are emitted as greenhouse gases due to being consumed as energy by microbes mining nutrition from SOM (Williams et al. 2006; Kirkby et al. 2013). Moreover, plant litter contains high quantities of lignin, tannin, and other low-energy contents that are usually respired but not used for biomass production (Devêvre and Horwáth 2000; Lu et al. 2003). Rice rhizodeposits, in contrast, mainly contain readily available C substrates that are rapidly utilized by soil microorganisms. Products of microbial resynthesis then finally transform to stable SOC (Pan et al. 2004; Ge et al. 2012), partly by sorption to reactive soil minerals (Xiao et al. 2015). Although microbial-assimilated C has been reported to account for less than 1.0% of the total C fixed in rice paddy soils (Yuan et al. 2012; Ge et al. 2013), it might be efficiently recycled in soil microbes and stabilized by soil minerals, which could have significant implications for long-term field fertility management.
Conclusion
Here, we investigated the microbial utilization of four different types of exogenous C sources in a paddy soil along with the efficiency of SOC formation. Soil microbial community structure and composition are substrate-dependent and driven by substrate availability. Although rice plant residues are widely used and large amounts enter the soil as shoot and root litter, their contribution to SOC formation is inefficient. In contrast, rhizodeposits and microbial-assimilated C have lower input rates, but are better stabilized by microbial metabolism, microbial recycling, and associations with soil minerals, and thus are characterized by high-efficiency soil C sequestration.
References
Alberti G, Vicca S, Inglima I, Belelli-Marchesini L, Genesio L, Miglietta F, Marjanovic H, Martinez C, Matteucci G, Andrea E, Peressotti A, Petrella F, Rodeghiero M, and Cotrufo M (2015) Soil C:N stoichiometry controls carbon sink partitioning between above-ground tree biomass and soil organic matter in high fertility forests. iForest 8:195–206
Bastida F, Torres IF, Hernández T, Bombach P, Richnow HH, García C (2013) Can the labile carbon contribute to carbon immobilization in semiarid soils? Priming effects and microbial community dynamics. Soil Biol Biochem 57:892–902
Baumann K, Marschner P, Smernik RJ, Baldock JA (2009) Residue chemistry and microbial community structure during decomposition of eucalypt, wheat and vetch residues. Soil Biol Biochem 41:1966–1975
Berg B (1986) Nutrient release from litter and humus in coniferous forest soils—a mini review. Scand J Forest Res 1:359–369
Boddy E, Hill PW, Farrar J, Jones DL (2007) Fast turnover of low molecular weight components of the dissolved organic carbon pool of temperate grassland field soils. Soil Biol Biochem 39:827–835
Boer W, Folman LB, Summerbell RC, Boddy L (2005) Living in a fungal world: impact of fungi on soil bacterial niche development. FEMS Microbiol Rev 29:795–811
Bol R, Poirier N, Balesdent J, Gleixner G (2009) Molecular turnover time of soil organic matter in particle-size fractions of an arable soil. Rapid Commun Mass Sp 23:2551–2558
Brant JB, Sulzman EW, Myrold DD (2006) Microbial community utilization of added carbon substrates in response to long-term carbon input manipulation. Soil Biol Biochem 38:2219–2232
Buyer JS, Teasdale JR, Roberts DP, Zasada IA, Maul JE (2010) Factors affecting soil microbial community structure in tomato cropping systems. Soil Biol Biochem 42:831–841
Chen R, Senbayram M, Blagodatsky S, Myachina O, Dittert K, Lin X, Blagodatskaya E, Kuzyakov Y (2014) Soil C and N availability determine the priming effect: microbial N mining and stoichiometric decomposition theories. Glob Chang Biol 20:2356–2367
Cotrufo MF, Wallenstein MD, Boot CM, Denef K, Paul E (2013) The microbial efficiency-matrix stabilization (MEMS) framework integrates plant litter decomposition with soil organic matter stabilization: do labile plant inputs form stable soil organic matter? Glob Chang Biol 19:988–995
Derrien D, Plain C, Courty PE et al (2014) Does the addition of labile substrate destabilise old soil organic matter. Soil Biol Biochem 76:149–160
Devêvre OC, Horwáth WR (2000) Decomposition of rice straw and microbial carbon use efficiency under different soil temperatures and moistures. Soil Biol Biochem 32:1773–1785
Dilly O, Bloem J, Vos A, Munch JC (2004) Bacterial diversity in agricultural soils during litter decomposition. Appl Environ Microb 70:468–474
Dungait JAJ, Kemmitt SJ, Michallon L, Guo S, Wen Q, Brookes PC, Evershed RP (2011) Variable responses of the soil microbial biomass to trace concentrations of 13C-labelled glucose, using 13C-PLFA analysis. Eur J Soil Sci 62:117–126
Falkowski PG, Godfrey LV (2008) Electrons, life and the evolution of Earth's oxygen cycle. Philos T R Soc B Biol Sci 363:2705–2716
Fontaine S, Henault C, Aamor A, Bdioui N, Bloor JMG, Maire V, Mary B, Revaillot S, Maron PA (2011) Fungi mediate long term sequestration of carbon and nitrogen in soil through their priming effect. Soil Biol Biochem 43:86–96
Gale WJ, Cambardella CA, Bailey TB (2000) Root-derived carbon and the formation and stabilization of aggregates. Soil Sci Soc Am J 64:201–207
Ge T, Yuan H, Zhu H, Wu X, Nie S, Liu C, Tong C, Wu J, Brookes P (2012) Biological carbon assimilation and dynamics in a flooded rice – soil system. Soil Biol Biochem 48:39–46
Ge T, Wu X, Chen H, Yuan HZ, Li B, Zhou P, Liu C, Tong C, Brookes P, Wu J (2013) Microbial phototrophic fixation of atmospheric CO2 in China subtropical upland and paddy soils. Geochim Cosmochim Ac 113:70–78
Glanville H, Rousk J, Golyshin P, Jones DL (2012) Mineralization of low molecular weight carbon substrates in soil solution under laboratory and field conditions. Soil Biol Biochem 48:88–95
Gunina A, Dippold MA, Glaser B, Kuzyakov Y (2014) Fate of low molecular weight organic substances in an arable soil: from microbial uptake to utilisation and stabilisation. Soil Biol Biochem 77:304–313
Jingguo W, Bakken LR (1997) Competition for nitrogen during mineralization of plant residues in soil: microbial response to C and N availability. Soil Biol Biochem 29:163–170
Johnson JMF, Allmaras RR, Reicosky DC (2006) Estimating source carbon from crop residues, roots and rhizodeposits using the National Grain-Yield Database. Agron J 98:622–636
Jones DL, Kemmitt SJ, Wright D, Cuttle SP, Bol R, Edwards AC (2005) Rapid intrinsic rates of amino acid biodegradation in soils are unaffected by agricultural management strategy. Soil Biol Biochem 37:1267–1275
Kirkby CA, Richardson AE, Wade LJ, Batten GD, Blanchard C, Kirkegaard JA (2013) Carbon-nutrient stoichiometry to increase soil carbon sequestration. Soil Biol Biochem 60:77–86
Kögel-Knabner I, Amelung W, Cao Z, Fiedler S, Frenzel P, Jahn R, Kalbitz K, Kölbl A, Schloter M (2010) Biogeochemistry of paddy soils. Geoderma 157:1–14
Kramer S, Marhan S, Ruess L, Armbruster W, Butenschoen O, Haslwimmer H, Kuzyakov Y, Pausch J, Scheunemann N, Schoene J, Schmalwasser A, Totsche KU, Walker F, Scheu S, Kandeler E (2012) Carbon flow into microbial and fungal biomass as a basis for the belowground food web of agroecosystems. Pedobiologia 55:111–119
Kuzyakov Y, Larionova AA (2005) Root and rhizomicrobial respiration: a review of approaches to estimate respiration by autotrophic and heterotrophic organisms in soil. J Plant Nut Soil Sci 168:503–520
Lal R (2004) Soil carbon sequestration impacts on global climate change and food security. Science 304:1623–1627
Li J, Li Y, Yang X, Zhang J, Lin Z, Zhao B (2015) Microbial community structure and functional metabolic diversity are associated with organic carbon availability in an agricultural soil. J Integr Agr 14:2500–2511
Liang C, Balser TC (2008) Preferential sequestration of microbial carbon in subsoils of a glacial-landscape toposequence, Dane County, WI, USA. Geoderma 148:113–119
Loeppmann S, Blagodatskaya E, Pausch J, Kuzyakov Y (2016) Substrate quality affects kinetics and catalytic efficiency of exo-enzymes in rhizosphere and detritusphere. Soil Biol Biochem 92:111–118
Lu Y, Watanabe A, Kimura M (2002) Contribution of plant-derived carbon to soil microbial biomass dynamics in a paddy rice microcosm. Biol Fert Soil 36:136–142
Lu Y, Watanabe A, Kimura M (2003) Carbon dynamics of rhizodeposits, root- and shoot-residues in a rice soil. Soil Biol Biochem 35:1223–1230
Marschner P, Umar S, Baumann K (2011) The microbial community composition changes rapidly in the early stages of decomposition of wheat residue. Soil Biol Biochem 43:445–451
Mikutta R, Kaiser K (2011) Organic matter bound to mineral surfaces: resistance to chemical and biological oxidation. Soil Biol Biochem 43:1738–1741
Müller K, Kramer S, Haslwimmer H, Marhan S, Scheunemann N, Butenschön O, Scheu S, Kandeler E (2016) Carbon transfer from maize roots and litter into bacteria and fungi depends on soil depth and time. Soil Biol Biochem 93:79–89
Nikolausz M, Kappelmeyer U, Székely A, Rusznyák A, Márialigeti K (2008) Diurnal redox fluctuation and microbial activity in the rhizosphere of wetland plants. Eur J Soil Biol 44:324–333
Pan G, Li L, Wu L, Zhang X (2004) Storage and sequestration potential of topsoil organic carbon in China's paddy soils. Glob Chang Biol 10:79–92
Paterson E, Sim A, Osborne SM, Murray PJ (2011) Long-term exclusion of plant-inputs to soil reduces the functional capacity of microbial communities to mineralise recalcitrant root-derived carbon sources. Soil Biol Biochem 43:1873–1880
Rubino M, Dungait JAJ, Evershed RP, Bertolini T, Angelis PD, D’Onofrio A, Lagomarsino A, Lubritto C, Merola A, Terrasi F, Cotrufo MF (2010) Carbon input belowground is the major C flux contributing to leaf litter mass loss: evidences from a 13C labelled-leaf litter experiment. Soil Biol Biochem 42:1009–1016
Ruf A, Kuzyakov Y, Lopatovskaya O (2006) Carbon fluxes in soil food webs of increasing complexity revealed by 14C labelling and 13C natural abundance. Soil Biol Biochem 38:2390–2400
Schurig C, Smittenberg RH, Berger J, Kraft F, Woche SK, Goebel MO, Heipieper HJ, Miltner A, Kaestner M (2013) Microbial cell-envelope fragments and the formation of soil organic matter: a case study from a glacier forefield. Biogeochemistry 113:595–612
Seth EC, Taga ME (2014) Nutrient cross-feeding in the microbial world. Front Microbiol 5:1–6
Shahzad T, Chenu C, Genet P, Barot S, Perveen N, Mougin C, Fontaine S (2015) Contribution of exudates, arbuscular mycorrhizal fungi and litter depositions to the rhizosphere priming effect induced by grassland species. Soil Biol Biochem 80:146–155
Singh S, Ghoshal N, Singh KP (2007) Variations in soil microbial biomass and crop roots due to differing resource quality inputs in a tropical dryland agroecosystem. Soil Biol Biochem 39:76–86
Sinsabaugh RL, Manzoni S, Moorhead DL, Richter A (2013) Carbon use efficiency of microbial communities: stoichiometry, methodology and modelling. Ecol Lett 16:930–939
Spohn M, Pötsch EM, Eichorst SA, Woebken D, Wanek W, Richter A (2016) Soil microbial carbon use efficiency and biomass turnover in a long-term fertilization experiment in a temperate grassland. Soil Biol Biochem 97:168–175
Tian J, Dippold M, Pausch J, Blagodatskaya E, Fan M, Li X, Kuzyakov Y (2013) Microbial response to rhizodeposition depending on water regimes in paddy soils. Soil Biol Biochem 65:195–203
Waldrop MP, Firestone MK (2004) Microbial community utilization of recalcitrant and simple carbon compounds: impact of oak-woodland plant communities. Oecologia 138:275–284
Waldrop MP, Harden JW, Turetsky MR, Petersen DG, McGuire AD, Briones MJI, Churchill AC, Doctor DH, Pruett LE (2012) Bacterial and enchytraeid abundance accelerate soil carbon turnover along a lowland vegetation gradient in interior Alaska. Soil Biol Biochem 50:188–198
Wang Q, Wang S, He T, Liu L, Wu J (2014a) Response of organic carbon mineralization and microbial community to leaf litter and nutrient additions in subtropical forest soils. Soil Biol Biochem 71:13–20
Wang Q, Wang Y, Wang S, He T, Liu L (2014b) Fresh carbon and nitrogen inputs alter organic carbon mineralization and microbial community in forest deep soil layers. Soil Biol Biochem 72:145–151
Weintraub M, Scott-Denton L, Schmidt S, Monson R (2007) The effects of tree rhizodeposition on soil exoenzyme activity, dissolved organic carbon, and nutrient availability in a subalpine forest ecosystem. Oecologia 154:327–338
Williams MA, Myrold DD, Bottomley PJ (2006) Carbon flow from 13C-labeled straw and root residues into the phospholipid fatty acids of a soil microbial community under field conditions. Soil Biol Biochem 38:759–768
Wu J, Joergensen RG, Pommerening B, Chaussod R, Brookes PC (1990) Measurement of soil microbial biomass C by fumigation-extraction—an automated procedure. Soil Biol Biochem 22:1167–1169
Wu J, Zhou P, Li L, Su Y, Yuan H, Syers JK (2012) Restricted mineralization of fresh organic materials incorporated into a subtropical paddy soil. J Sci Food Agr 92:1031–1037
Xiao J, Wen Y, Li H, Hao J, Shen Q, Ran W, Mei X, He X, Yu G (2015) In situ visualisation and characterisation of the capacity of highly reactive minerals to preserve soil organic matter (SOM) in colloids at submicron scale. Chemosphere 138:225–232
Yuan H, Ge T, Wu X, Liu S, Tong C, Qin H, Wu M, Wei W, Wu J (2012) Long-term field fertilization alters the diversity of autotrophic bacteria based on the ribulose-1,5-biphosphate carboxylase/oxygenase (RubisCO) large-subunit genes in paddy soil. Appl Microbiol Biot 95:1061–1071
Yuan H, Zhu Z, Liu S, Ge T, Jing H, Li B, Liu Q, Lynn TM, Wu J, Kuzyakov Y (2016) Microbial utilization of rice root exudates: 13C labeling and PLFA composition. Biol Fert Soil 52:615–627
Zelles L (1997) Experimental and theoretical approaches in environmental chemistry phospholipid fatty acid profiles in selected members of soil microbial communities. Chemosphere 35:275–294
Zelles L (1999) Fatty acid patterns of phospholipids and lipopolysaccharides in the characterisation of microbial communities in soil: a review. Biol Fert Soil 29:111–129
Zhu Z, Zeng G, Ge T, Hu Y, Tong C, Shibistova O, He X, Wang J, Guggenberger G, Wu J (2016) Fate of rice shoot and root residues, rhizodeposits, and microbe-assimilated carbon in paddy soil – part 1: decomposition and priming effect. Biogeosciences 13:4481–4489
Acknowledgments
The present study was supported by the National Natural Science Foundation of China (41522107; 41501321), the Strategic Priority Research Program of the Chinese Academy of Sciences (XDB15020401), and the Recruitment Program of High-End Foreign Experts of the State Administration of Foreign Experts Affairs, awarded to Prof. Georg Guggenberger (GDT20164300013). We thank the Public Service Technology Center, Institute of Subtropical Agriculture, Chinese Academy of Sciences for technical assistance.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Peter J. Gregory .
Rights and permissions
About this article
Cite this article
Zhu, Z., Ge, T., Hu, Y. et al. Fate of rice shoot and root residues, rhizodeposits, and microbial assimilated carbon in paddy soil - part 2: turnover and microbial utilization. Plant Soil 416, 243–257 (2017). https://doi.org/10.1007/s11104-017-3210-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-017-3210-4