Abstract
The soluble components of rhizodeposition—root exudates—are the most important sources of readily available carbon (C) for rhizosphere microorganisms. The first steps of exudate utilization by microorganisms define all further flows of root C in the soil, including recycling and stabilization. Nevertheless, most studies have traced root exudates C much later after its initial uptake by microorganisms. To understand microbial uptake and utilization of rice root exudates, we traced 13C incorporated into microbial groups by 13C profiles of phospholipid fatty acids (PLFAs) within a short time (6 h) after 13CO2 pulse labeling. Labeling was conducted five times during three growth stages: active root growth (within the 21 days after transplanting), rapid shoot growth (37 and 45 days), and rapid reproduction (53 and 63 days). 13C was quickly assimilated throughout the rhizosphere microorganism, and the incorporation rate increased with rice maturity. Despite low redox conditions in paddy soil, fungi outcompeted bacteria in utilizing the root exudates. At all growth stages, fungal PLFAs (18:2 w6, 9c/18:0) showed the highest 13C levels, whereas actinomycete PLFAs (16:0 10-methyl) showed the lowest 13C incorporation. Principal component analysis revealed that the rhizosphere microbial community differed among rice growth stages, whereas the whole microbial community remained stable. In conclusion, the rapid incorporation of carbon from root exudates into microorganisms in paddy soils depends on the growth stage of the rice plant and is the first step of C utilization in rice rhizosphere, further defining C utilization and stabilization.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Rhizodeposits consist of a wide range of compounds, including root exudates, mucilage, sloughed-off cells and tissue, cell lysates and root debris (Kuzyakov and Larionova 2005; Gregory 2006). Exudates are photosynthetic products released continuously and passively from roots, and account for 5–30 % of the total photosynthetically fixed C in the soil (Kuzyakov and Domanski 2000; Kuzyakov et al. 2003; Nguyen 2003). The main components of root exudates are soluble sugars ( Fischer et al. 2010; Palacios et al. 2014; Gunina and Kuzyakov 2015), which serve as a primary energy source and an important source of C for soil microorganisms in the vicinity of growing roots (Högberg et al. 2008; Werth and Kuzyakov 2010; Pii et al. 2015). Photosynthates assimilated in leaves are actively loaded through phloem to the roots and passively released into the rhizosphere. The time interval between photoassimilation in leaves and direct transport to roots can be predicted from phloem transport rates, which range from 0.5 to 1.0 m h−1, and vary between species and growth stages (Thompson and Holbrook 2003; Nobel 2005; Barnard et al. 2007; Kuzyakov and Gavrichkova 2010), depending on environmental conditions. In contrast, the release of exudates from roots is very fast and occurs nearly without any time delay (Gavrichkova and Kuzyakov 2012). The amount and quality of exudates are strongly controlled by photosynthesis and depend on plant species, age, and external biotic and abiotic factors (Gavrichkova and Kuzyakov 2012; Ge et al. 2012; Meng et al. 2013). In general, the amount of root exudate is highest during fast vegetative growth and decreases with plant age (Yevdokimov et al. 2006; Garbeva et al. 2008).
Many researchers have analyzed belowground C partitioning through rhizodeposition and the incorporation of that C into microbial biomass (Paterson et al. 2007; Hannula et al. 2012; Benesch et al. 2015). Several molecular and biochemical methods have been developed over the past decade to study the diversity of soil microorganisms, most of which are unknown and unculturable (Ganzert et al. 2011; Shade et al. 2012). For example, the composition of microbial phospholipid fatty acids (PLFAs) mirrors the general composition of soil and aquatic microbial communities (Lu et al. 2007; Abraham 2014). Stable C isotope techniques coupled with PLFA analysis allows study of both the function and structure of microbial communities. 13C-PLFA reveals information about the portion of the microbial community that uses 13C-labeled substrates (Webster et al. 2006; Dungait et al. 2011; Yao et al. 2015).
Nearly all 13C-PLFA studies have been conducted in dryland soils and with crops such as maize, wheat, ryegrass, or barley to determine of the amount of root-derived C entering the soil and its subsequent microbial utilization. 13C pulse-chase labeling combined with PLFA analysis of ryegrass has shown that the responses of microbial communities to rhizodeposition depend on plant age (Butler et al. 2004; Philippot et al. 2013). The role of rhizodeposition in the relationship between the composition of microbial community and photosynthesis in paddy soils has been evaluated in only a few studies (Tian et al. 2013). Even fewer studies have focused on such fast process as deposition of root exudates and implications for associated microbial communities in rice paddies.
Paddy soils cultivated with rice (Oryza sativa L.) make up the largest anthropogenic wetlands on earth (Kögel-Knabner et al. 2010) and are a primary food source for nearly half of the world’s population (FAO 2011). Paddy ecosystems have great potential for sequestering atmospheric CO2 (Lal 2004) and are very sensitive to climate change (Ge et al. 2012, 2015). Paddy soils are different from upland soils because they are regularly flooded and intermittently irrigated. Rice is cultivated under continuous flooding and anaerobic conditions, but the rhizosphere of rice plants can have physicochemical conditions that differ from those found in the absence of roots. The roots of rice plants promote diffusion of atmospheric oxygen into the rhizosphere. This oxygenation capability of rice roots in the rhizosphere is largely affected by the age and phenological stage of the plant (Doran et al. 2006). Differences in the rhizosphere redox potentials also contribute to changes in the physicochemical soil properties (Nikolausz et al. 2008), thereby affecting the microbial composition and the succession of microbial communities. Paddy soils are characterized by relatively slow C turnover; however, only few studies have shown the biological processes determining soil organic matter stabilization and turnover in paddy soil, especially at the microbial level (Sollins et al. 1996; Gale et al. 2000). Better understanding of the release of root C by rice, its distribution and transformation belowground, and its microbial utilization are critical for understanding global C cycling and the ecological functions of paddy ecosystems.
We tested two hypotheses: (1) root exudates and their utilization by microbes change during plant growth, and (2) rhizosphere fungi outcompete bacteria for initial uptake of root exudates because of their larger biomass and because the diffusion of atmospheric oxygen by rice roots favors fungal growth. We performed 13C pulse-chase labeling of wetland rice at three growth stages under field conditions. To test the first hypothesis, the pattern of 13C photosynthate uptake into shoot, roots, total soil organic matter, and microbial biomass were measured and compared during the growth stages. To test the second hypothesis, the incorporation of exudates into different microbial groups was traced by analyzing PLFAs shortly after 13C assimilation aboveground.
Materials and methods
Soils and sampling
Typical Stagnic Anthrosol soil, developed from highly weathered granite, was collected from a rice field (113° 19′ 52″ E, 28° 33′ 04″ N, 80 m a.s.l.) located at the Changsha Research Station for Agricultural and Environmental Monitoring. The site has a subtropical climate with an annual mean temperature of 17.5 °C, 1300 mm rainfall, 16.63 h day−1 sunshine, and a frost-free period of 274 days. The soil at the research station was derived from quaternary red clay with a bulk density (BD) of 1.31 g cm−3. Moist soil samples were collected from the plow layer (0–20 cm) and sieved (<4 mm) to remove coarse plant residues. The soil consisted of 7.51 % clay, 68.4 % silt, and 24.1 % sand. The soil contained 18.1 g kg−1 organic C, 1.80 g kg−1 total N, 0.43 g kg−1 total P, and had a pH of 5.56 (1:2.5, soil/water ratio). The extractable nutrients and other basic properties of this soil are listed in Table 1.
Rice growth conditions and 13CO2 labeling
Rice growth conditions and 13CO2 labeling were as described by Ge et al. (2012, 2015) with some modifications. Briefly, three 30-day-old rice seedlings (Oryza sativa L.; two-line hybrid rice Zhongzao 39; average dry weight 0.10 g per plant) were transplanted into each of 30 pots (filled with 1 kg soil; dry weight) on April 29, 2013. An initial basal fertilizer, consisting of (NH4)2SO4, calcium superphosphate, and KCl, was applied at a rate of 60 mg N, 20 mg P, and 80 mg K kg−1 of soil and mixed thoroughly. Deionized water was used for irrigation, and a 1–2-cm water layer was maintained above the soil surface throughout the growing season. Any weeds in the soil were carefully removed by hand. Control soil was held outside of the labeling chambers.
Fifteen pots were used for labeling: 13CO2 pulse labeling was performed five times at 21, 37, 45, 52, and 63 days after transplantation, covering the complete growth period of rice plants (Lu et al. 2002a, 2007). These times correspond to the three typical rice growth stages (rapid root growth stage (up to 21 days after transplantation), rapid shoot growth (between 37 and 45 days after transplantation), and reproduction (between 52 and 63 days after transplantation)). The plants in three independent pots were labeled on each date. Plants were labeled with 13CO2 for 6 h (09:00–15:00) in an automatically controlled gas-tight growth chamber system (area 110 cm × 250 cm, height 180 cm). No specific precautions, such as leak checks, were taken to ensure that shoots and roots were isolated because downward diffusion of 13CO2 into acid floodwater (pH < 6.0) was negligible (Lu et al. 2002a). The 13CO2 used in the experiment was generated through reaction of NaH13CO3 (99 atom% 13C, Cambridge Isotope Laboratories, Inc.) and HCl (1 M) in a plastic beaker placed inside the growth chamber (Shsen-QZD, Qingdao, China). During labeling, CO2 was released into the chamber only when the CO2 concentration in the chamber was lower than 360 μL L−1. At CO2 concentrations greater than 380 μL L−1, a switch diverted gas flow so that it passed through CO2 traps (NaOH solution), where excess CO2 was absorbed. The other 15 pots and a control soil without plant were prepared as references for natural 13C abundance to calculate the 13C atom % excess and were placed outdoors, 10–15 m away from labeled plants. The soil properties and the δ13C signature did not change in the control soil during the test period.
Sampling and harvesting
At the end of the 13C pulse labeling, the plants and soils from three independent pots were destructively sampled after the measurements of soil redox potentials (Eh) by using a platinum electrode against a standard Ag/AgCl electrode (Mettler Toledo. Switzerland), at soil depths of 5–10 cm. This corresponded to 6 h after the start of assimilation. At harvest, the shoots were cut off at the stem base, allowing for separation of the roots, shoots, and soil components. Because roots and fungi share several of common PLFA biomarkers, it was necessary to make sure of the PLFA profiles obtained and their δ13C values were representative of the soil microbial community rather than plant material. To do this, any soil adhering to the roots was removed as described by Butler et al. (2002), suspended in 40 ml of 50 mM phosphate buffer (pH 6.8), centrifuged at 8400×g for 5 min, and the supernatant was decanted to remove the root cell contamination (the floatable, plant-derived debris). To assess whether this method was effective enough to remove most of the fine living roots from soils, 15 g of soil from each sampling pot was examined under a dissecting microscope (SM-1BSZZ-64S, AmScope, USA). If the fine roots were not found in samples, the sieving-centrifugation procedure was considered effective enough to remove most of the fine living roots in comparison with visual separation under a microscope. If not, the fine living roots were removed manually after the resuspension and centrifugation. All obtained fresh soil samples were mixed thoroughly and divided into three portions. One portion was immersed immediately in liquid nitrogen and stored at −70 °C for future PLFA analysis. Another was used immediately for determination of soil Mn (II), Fe (II), and microbial biomass C. The remaining portion was air-dried, ground, sieved through a 100 mesh, and used for determining the 13C-soil organic carbon (SOC) and other soil physiochemical properties. Visible dead roots were included in the total root mass because it was impossible to separate fragments of dead roots from living roots. Therefore, all analyses were carried out with mixed (living and dead) root material (Yevdokimov et al. 2006). Shoot and root materials were dried in an oven for 72 h at 70 °C, weighed, and pulverized.
The soil physicochemical properties and 13C in plant–soil–microbial system
The C content of independent soil, shoot, and root samples were analyzed using an automated C/N analyzer (Vario MAX, Elementar Analysensysteme GmbH, Germany). The soil cation exchange capacity (CEC) was measured by titration (Rhoades 1982), total P by NaOH fusion and colorimetric determination (Olsen and Somers 1982), total K by NaOH fusion flame photometry (Allen 1989), and available N by alkali hydrolytic diffusion (Keeney and Nelson 1982). Olsen P was extracted using sodium bicarbonate and determined using the Mo-Sb colorimetry method (Colwell 1963). Available K was extracted by ammonium acetate and determined by flame photometry (Knudsen et al. 1982). Soil exchangeable Mn (II) was extracted by NH4OAc (1 M at pH 6.0) and determined by atomic absorption spectroscopy (Ragnarsdottir and Hawkins 2006), and available Fe (II) was extracted by HCl (0.5 M) and determined by 1,10-phenanthroline colorimetric assay (Fadrus and Malý 1975).
Prior to analysis for δ13C, dry samples of the soil, shoots, and roots were ground to a fine powder. The stable C isotope ratios of plant and soil materials were measured using a MAT253 isotope ratio mass spectrometer coupled with an elemental analyzer FLASH HT (ThermoFisher Scientific, USA). Soil microbial biomass C (MBC) together with dissolved organic carbon (DOC) extracted by 0.5 M K2SO4 were measured by the fumigation extraction method (Wu et al. 1990) with subsequent δ13C analysis.
PLFA extraction and analysis
Twelve soil samples, representing the three typical rice growth stages, were selected for soil microbial PLFA analysis. The soil microbial PLFAs were extracted, fractionated, and purified according to the method described by Wu et al. (2009), with minor modifications. Briefly, approximately 2 g of freeze-dried soil was extracted twice using 22.8 ml single-phase chloroform/methanol/citrate buffer system (1:2:0.8 v/v/v; pH 4.0). The phospholipids were then separated from neutral lipids and glycolipids using silica acid columns (Supelco, Bellefonte, PA, USA). Phospholipids were methylated using mild alkaline methanolysis (White et al. 1979). Following methylation of the phospholipids, the PLFA methyl esters (FAME) were separated and identified using a gas chromatograph (N6890; Agilent, Santa Clara, CA, USA) fitted with a MIDI Sherlock microbial identification system (Version 4.5; MIDI, Newark, DE, USA). For quantification of the phospholipids, methyl nonadecanoate fatty acid (19:0) was added prior to derivatization as an internal concentration standard. Microbial phospholipid fatty acid markers (Tunlid and White 1992; Zelles et al. 1992; Frostegård et al. 1993; Olsson et al. 1999; Hill et al. 2000; Grayston et al. 2001; Priha et al. 2001 ) are shown in Table 2. The remaining fatty acids were not assigned to any microbial group, including monounsaturated fatty acids (16:1 w7c/16:1 w6c, 16:1 w5c, 18:1 w9c, 18:1 w7c, 20:1 w9c), cyclopropane fatty acids (17:0 cyclo, 19:0 cyclo w8c), anteiso (15:0 anteiso,17:0 anteiso), and iso (14:0 iso,15:0 iso,16:0 iso.17:0 iso). A Trace GC Ultra gas chromatograph (GC) with a combustion column attached via GC Combustion III to a Delta V Advantage isotope ratio mass spectrometer (IRMS; Thermo Finnigan, Germany) was used to analyze the δ13C of individual PLFAs. The equipment and running conditions were as described in detail by Thornton et al. (2011).
Calculations
13C incorporation into plant and soil samples was calculated according to Eq. (1).
where Atomic 13 C% L and Atomic 13 C% UL are atomic 13C% in labeled and unlabeled samples, respectively, and C is the total C content of samples.
The 13C-MBC was calculated as the difference in 13C between fumigated and non-fumigated soil extracts, divided by a factor of 0.45, as shown in Eq. 2 (Lu et al. 2002b).
where Atomic 13 C% f, uf, L, and UL indicate fumigated soil extracts, non-fumigated soil extracts, extracts from labeled samples, and extracts from unlabeled samples, respectively. C f and C uf are the total C contents of the fumigated and non-fumigated soil extracts.
The amount of 13C incorporation into each PLFA (mg kg−1) was determined using a mass balance approach:
where Atomic 13 C% PLFA, L, and UL indicate PLFA, extracts from labeled samples, and extracts from unlabeled samples, respectively. We calculated the relative 13C distribution in each specific microbial group according to the following equation (Tian et al. 2013):
where 13CPLFA-group is the amount of 13C-PLFA incorporated into each specific microbial group and ∑13 C PLFA is the total amount of 13C-PLFA incorporated into soil microbes.
The 13C incorporated into shoots, roots, and soil pools was expressed as the percentage of 13C recovery on each sampling day. The total 13C recovery after each sampling in the rice–soil system was calculated as the sum of the 13C in shoots, roots, and soil (Tian et al. 2013).
Statistical analysis
Data were processed using Excel 2010 for the means and standard errors. After normality testing on the data with the W-test, multiple comparison tests were made using one-way ANOVA followed by Duncan’s multiple range test (P < 0.05). Correlation analyses were done using the Pearson correlation method with significance defined at the 0.05 level unless otherwise stated. All analyses were performed using SPSS 13.0 software for Windows XP.
Principal components analysis (PCA) and redundancy analysis (RDA) were performed with CANOCO 5.0 for Windows (Microcomputer Power, Ithaca, NY, USA) based on the PLFA profiles during each rice growth stage to identify the effects of soil physiochemical parameters and root exudates, including 13C-SOC, 13C-root, and soil extractable nutrients on the soil rhizosphere microbial communities.
Results
The soil physiochemical properties at rice stages
Rice growth had an effect on the soil chemical properties (Table 1). Eh increased during active root and shoot growth and then slightly decreased during the reproductive stage; a similar trend was found for Fe (II) content, which significantly increased at the root and shoot growth stages and then remained constant through the reproductive stage. The available N and K contents decreased during the experiment, while the Olsen P content increased during the first 21 days (root growth) and decreased until day 45 (shoot growth). Other soil physiochemical parameters such as CEC and Mn (II) contents did not greatly vary (P > 0.05) during the rice growth period.
Shoot and root biomass and allocation of 13C photosynthates in the plant–soil system
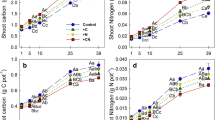
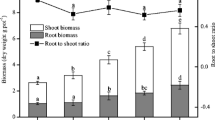
Shoot and root biomass increased with rice growth (from 1.6 to 11.8 and 1.2 to 5.5 g C per pot, respectively; Fig. 1): overall, biomass of both shoots and roots increased throughout the life of the plant until final harvest (63 days after transplantation). Allocation of assimilated 13C in shoots and roots was depended on the rice growth stage. The amount of 13C in shoots and roots ranged from 18.6 to 163 and from 2.9 to 13.7 mg C per pot, respectively. The 13C in roots and shoots increased with rice growth (Fig. 1). The 13C-SOC content ranged from 2.7 to 6.9 mg C per pot and increased as plants grew, with the greatest increase occurring as shoot growth transitioned to reproductive growth (Fig. 2). The 13C incorporated into DOC (13C-DOC) was 9.2–12.6 μg C per pot and 13C-MBC was 6395–1310 μg C per pot. The increases in 13C-MBC occurred in concert with plant growth and, as for DOC, there was no significant difference during the plant growth period (Fig. 2). Correlation parameters showed significant correlation among 13C-SOC, 13C-MBC, and 13C in the roots (P < 0.01; Fig. 3). Some soil extractable nutrient variables and plant 13C-photosynthetic products were also significantly correlated (Table 3). Available K was significantly correlated with 13C in the shoots, root, SOC, and MBC, and a significant correlation was also observed between Available N and 13C-SOC (see Table 3).
Relationships between a root biomass and soil organic 13C (13C-SOC), b root biomass and microbial biomass 13C (13C-MBC), and c 13C-SOC and 13C-MBC in paddy soils sampled at five time points during growth of rice plants. Samples were collected within 1 h of 13C pulse labeling of rice plants. Points represent individual tested soil means
Incorporation of rhizo-C in total PLFAs and microbial groups
Total PLFAs and 13C incorporation into PLFAs ranged from 46.4 to 53.1 mg C kg−1 and from 107 to 165 μg C kg−1, respectively (Table 4) and the recovery of 13C-PLFAs for three of the sampling days differed significantly (P < 0.05; Table 4).
The 13C from root exudates was not evenly distributed among microbial groups, indicating that they differed in C uptake (Fig. 4a). For all three sampling days selected for soil microbial PLFA analysis, more than 40 % of 13C recovered from the PLFAs was incorporated into fungi (18:2w6, 9c, 18:1w9c). Large amounts of 13C were also incorporated into Gram-negative bacteria (16:1w7, 18:1w7c, cy17:0, cy18:0; Fig. 3b), which contained about 15 % of the total 13C-PLFA. The actinomycetes (16:0 10-methyl) and mycorrhizal (AM) fungi (16:1w5c) only accounted for about 1 % of total 13C-PLFAs, and the lowest percentage of 13C-PLFA was incorporated into actinomycetes at day 63 (less than 0.5 %).
To determine the effects of rhizodeposition on microbial composition, PLFA and 13C in PLFA data were subjected to PCA. The first two axes explained 91.3 % of the variance. The composition of rhizodeposit-dependent microbe communities (13C-PLFA) was clearly grouped by sampling date, whereas the overall soil microbial diversity (PLFA) formed a single group (Fig. 5a). Furthermore, RDA results revealed that the soil Fe (II) and Olsen P values were the most significantly correlated with the rhizodeposit-dependent microbial composition (P = 0.002 by the Monte Carlo permutation test). However, Mn (II) levels were not significantly associated with the rhizodeposit-dependent microbial composition (Fig. 5b).
Principal components (a) and redundancy analysis (b) of the 13C in phospholipid fatty acids (13C-PLFA) profiles of the rhizosphere microbial composition in the soil at three rice growth stages. Arrows indicate the variables. The direction of an arrow indicates the steepest increase of the variable, and the length indicates the strength relative to other variables
Discussion
13C allocation in plant soil between growth stages of rice
For the high sensitivity of the PLFA-stable isotope probing (SIP) technique, soil sampling analysis of 13C in microbial groups was performed only 6 h after the start of 13C assimilation in order to investigate the early stages of root exudate utilization by rhizosphere microorganisms (Wiesenberg et al. 2010). Because of the short period, only a small part of the 13C assimilated by the plants was dispersed belowground and incorporated into microbial PLFA. The distribution of photosynthates in plant–soil systems is affected by plant variety and growth stage (Kuzyakov and Cheng 2001; Lu et al. 2002a). In the present study, the total assimilated 13C in this plant–soil system increased with rice age, but the proportion of 13C incorporated into the roots decreased from 12 to 7 %. The highest rate of 13C incorporation into SOC (10 %) occurred in the first growth stage and promoted microbial activity and growth (13C-MBC, 3 %) by providing readily available substrates. The roots of young rice grew relatively quickly and, therefore, were a greater C sink than those of mature rice plants. Moreover, a shorter transport pathway in younger rice decreases the time interval between photoassimilation in leaves and release of root exudates compared to mature plants (Kuzyakov and Gavrichkova 2010; Gavrichkova et al. 2011). Therefore, young roots drew more photosynthates belowground than the roots of maturing rice. This is likely related to the greater C sink activity observed in the roots of younger plants than of older plants (Lu et al. 2002a).
Carbon (13C or 14C) incorporated into SOC and MBC during pulse or continuous labeling is positively correlated with rice biomass measured in greenhouse experiments (Ge et al. 2012). Pausch et al. (2013) successfully estimated the rhizodeposition-C in maize using 14C labeling and suggested that rhizodeposition could be quantified at field scale by assuming a constant ratio of rhizodeposited C to root C. Thus, to estimate rhizodeposition under field conditions, it is necessary to use the linear relationship between rice biomass and 13C-SOC with adjustment of its incorporation into MBC and DOC for the portion respired as CO2. This will be verified in further studies on upland and paddy soils by examining more soil types, plant species, and depending on environmental conditions (Roth et al. 2013; He et al. 2015).
Incorporation of rhizodeposits into microbial groups at three growth stages of rice
Root exudates diffuse from root surfaces to microorganisms (Darrah 1991; Hafner et al. 2014). These easily available organic substances are utilized by rhizosphere microorganisms within a few minutes (Fischer et al. 2010; Moran-Zuloaga et al. 2015). Other relevant rhizodeposition processes, such as the sloughing off of cells or the death of root hairs and fine roots, take much longer than exudation (days, rather than minutes; Kuzyakov and Domanski 2002; Högberg and Read 2006; Paterson et al. 2009) and could not have played a significant role within the 6-h labeling period used in this study. Most of the 13C was recovered from fungal PLFAs (16:0, 18:2w6, or 18:1w9c; PLFAs 18:2w6, 9c, and 18:1w9c) on all three dates used for PLFA analysis (Fig. 4). This is consistent with the pulse labeling results of Wang et al. (2016).
An enormous diversity of fungi can be found worldwide that have varied ecologies and life cycle strategies (Brundrett 2002, De Boer et al. 2005; Amaya-Carpio et al. 2009). Rice plants are continually involved in mycorrhizal associations under upland conditions. However, infection is rare under submerged conditions owing to the anoxic environment. In spite of this, some mycorrhizae belonging to the genus Glomus do survive in waterlogged conditions. Moreover, fungi were observed in the rhizosphere soil. Because the rhizosphere is well aerated by rice root parenchyma, fungi are not limited by anoxic conditions common in non-rhizosphere soil. With rice growth, dissolved nutrients in the soil decreased while reducing substances, such as Fe (II) increased. In the rhizosphere, fungal hyphae have very large surface areas, preferentially allowing fungi to mine for nutrients and oxygen (diffusion from rice plant roots) compared to bacteria and Archaea. Further, mycorrhizal associations allow organics to be absorbed directly from roots, before they can be released into the soil. A significant amount of 13C was recovered from fungal PLFAs, which indicates that fungi might play a preeminent role in the utilization of root exudates. Recent studies have shown that fungi also play much more significant roles in C and N cycling under subanoxic soil conditions (Qin et al. 2014; Chen et al. 2015).
Effect of photosynthate inputs on soil microbial community composition
Rhizosphere microbial populations are significantly affected by photosynthate inputs and depend on plant communities and their growth stages (Göttlicher et al. 2006). However, this is not always true since soil properties can be more important than plant effects, particularly if the plant has been growing in the soil for a few years. Whether soil characteristics or specific plant properties are more important in shaping the bacterial community structure of the rhizosphere has been a topic of considerable debate (De Ridder-Duine et al. 2005; Delmont et al. 2014). Little research has been performed thus far to ascertain the effects of root exudates on the microbial community composition involved in plant-derived C cycling. Our study indicated that the allocation of photosynthesized 13C into the soil affected the composition of the rhizo-coupled microbial community (Fig. 5). The changes in allocation of photosynthesized 13C into the paddy soil ecosystem that occurred with growth stage resulted in obvious variations in the composition of the rhizodeposit-dependent microbial community. However, the whole soil microbial community composition did not change much during the study. Consequently, photosynthesized C inputs more easily affect the rhizo-coupled microbes than the overall microbial community. Among the physicochemical properties measured, multivariate statistical analysis showed that the major influences on the rhizodeposit-dependent microbial community were the soil Fe (II) and Olsen P values coupled with other soil extractable nutrients. However, the greatest influencing factors were changes in the soil Fe (II), Olsen P values, or Eh and 13C-SOC that depended on the growth stage of the rice plant. In this study, the paddy soil pots were irrigated with deionized water, with a 2–3-cm water layer maintained above the soil surface throughout the growing season. Therefore, we suspect that the redox potentials and reduced forms of some elements (e.g., Fe (II)) were mainly affected by the rice roots. In the rhizosphere, organic compounds released from the rice plant root contribute to oxygen consumption mediated by heterotrophs, while atmospheric oxygen is diffused by the roots to lower the concentrations of these toxic-reduced substances. Both of the above processes are highly dependent on rice root age (Nawaz et al. 2012). Rice root uptake of nutrients from the soil solution differs during rice plant growth stages, from germination to harvest, and generally most of the nutrient uptake occurs before flowering (Ramanathan and Krishnamoorthy 1973). Correlation analysis also showed significant dependence between some of nutrients, microbial biomass, and 13C-photosynthetic products (Table 3). Therefore, we expected that the physiological changes within the rice root through the stages of plant growth modify soil physiochemical conditions, in turn affecting the formation of the distinct rhizo-coupled microbial communities.
Conclusions
Rice rhizodeposits were incorporated into microbial community already 6 h after assimilation aboveground. Fungi appeared to outcompete bacteria for initial uptake of root exudates and are therefore responsible for the first step of microbial utilization of root-derived C in soil. Rhizodeposition can be well assessed under field conditions based on the root biomass, and therefore, the suggested approach can be used to estimate belowground C input and contributions to C sequestration. 13C-PLFA analysis coupled with 13C pulse labeling was a promising and effective approach for examining microbial dynamics associated with rhizosphere C cycling. Since the poor phylogenetic discrimination of the PLFA technique, the application of this methodology coupled with molecular techniques (RNA-SIP) in further studies will allow clarifying the taxonomy of fungi and other microbes involved in root-derived C cycle. These couplings of isotopic approaches with biomarkers have the potential to greatly enhance our knowledge of rhizosphere processes by focusing on the members actively involved in the cycling of C and nutrients within complex soil system.
References
Abraham WR (2014) Applications and impacts of stable isotope probing for analysis of microbial interactions. Appl Microbiol Biotechnol 98:4817–4828
Allen SE (1989) Chemical analysis of ecological material. Blackwell, Oxford, p 386
Amaya-Carpio L, Davies FT, Fox T, He C (2009) Arbuscular mycorrhizal fungi and organic fertilizer influence photosynthesis, root phosphatase activity, nutrition, and growth of Ipomoea carnea ssp. fistulosa. Photosynthetica 47:1–10
Barnard RL, Salmon Y, Kodama N, Sörgel K, Holst J, Rennenberg H, Gessler A, Buchmann N (2007) Evaporative enrichment and time lags between δ18O of leaf water and organic pools in a pine stand. Plant Cell Environ 30:539–550
Benesch M, Glaser B, Dippold M, Zech W (2015) Soil microbial C and N turnover under Cupressus lusitanica and natural forests in southern Ethiopia assessed by decomposition of 13C- and 15N-labelled litter under field conditions. Plant Soil 388:133–146
Brundrett MC (2002) Coevolution of roots and mycorrhizas of land plants. New Phytol 154:275–304
Butler JL, Williams MA, Bottomley PJ, Myrold DD (2002) Microbial community dynamics associated with rhizosphere carbon flow. Appl Environ Microbiol 69:6793–6800
Butler JL, Bottomley PJ, Griffith SM, Myrold DD (2004) Distribution and turnover of recently fixed photosynthate in ryegrass rhizospheres. Soil Biol Biochem 36:371–382
Chen H, Mothapo NV, Shi W (2015) Soil moisture and pH control relative contributions of fungi and bacteria to N2O production. Microb Ecol 69:180–191
Colwell JD (1963) The estimation of phosphorus fertilizer requirements of wheat in Southern New South Wales by soil analysis. Anim Reprod Sci 3:190–197
Darrah PR (1991) Measuring the diffusion coefficient of rhizosphere exudates in soil I. The diffusion of non-sorbing compounds. J Soil Sci 42:413–420
De Boer W, Folman LB, Summerbell RC, Boddy L (2005) Living in a fungal world: impact of fungi on soil bacterial niche development. FEMS Microbiol Rev 29:795–811
De Ridder-Duine AS, Kowalchuk GA, Klein Gunnewiek PJA, Smant W, van Veen JA, de Boer W (2005) Rhizosphere bacterial community composition in natural stands of Carex arenaria (sand sedge) is determined by bulk soil community composition. Soil Biol Biochem 37:349–357
Delmont TO, Francioli D, Jacquesson S, Laoudi S, Nesme MJ, Ceccherini MT, Nannipieri P, Simonet P, Vogel TM (2014) Microbial community development and unseen diversity recovery in inoculated sterile soil. Biol Fertil Soils 50:1069–1076
Doran G, Eberbach P, Helliwell S (2006) The impact of rice plant roots on the reducing conditions in flooded rice soils. Chemosphere 63:1892–1902
Dungait JAJ, Kemmitt SJ, Michallon L, Guo S, Wen Q, Brookes PC, Evershed RP (2011) Variable responses of the soil microbial biomass to trace concentrations of 13C-labelled glucose, using 13C-PLFA analysis. Eur J Soil Sci 62:117–126
Fadrus H, Malý J (1975) Rapid extraction-photometric determination of traces of iron (II) and iron (III) in water with 1,10-phenanthroline. Anal Chim Acta 77:315–316
FAO (2011) Rice Market Monitor. April 2011. Vol XIV issue no. 2. P2. http://www.fao.org/docrep/014/am491e/am491e00.pdf
Fischer H, Ingwersen J, Kuzyakov Y (2010) Microbial uptake of low-molecular-weight organic substances out-competes sorption in soil. Eur J Soil Sci 61:504–513
Frostegård A, Bååth E, Tunlid A (1993) Shifts in the structure of soil microbial communities in limed forests as revealed by phospholipid fatty acid analysis. Soil Biol Biochem 25:723–730
Gale WJ, Cambardella CA, Bailey TB (2000) Root-derived carbon and the formation and stabilization of aggregates. Soil Sci Soc Am J 64:201–207
Ganzert L, Lipski A, Hubberten H, Wagner D (2011) The impact of different soil parameters on the community structure of dominant bacteria from nine different soils located on Livingston Island, South Shetland Archipelago, Antarctica. FEMS Microbiol Ecol 76:476–491
Garbeva PV, Van Elsas JD, van Veen JA (2008) Rhizosphere microbial community and its response to plant species and soil history. Plant Soil 302:19–32
Gavrichkova O, Kuzyakov Y (2012) Direct phloem transport and pressure concentration waves in linking shoot and rhizosphere activity. Plant Soil 351:23–30
Gavrichkova O, Proietti S, Moscatello S, Portarena S, Battistelli A, Matteucci G, Brugnoli E (2011) Short-term natural δ13C variations in pools and fluxes in a beech forest: the transfer of isotopic signal from recent photosynthates to soil respired CO. Biogeosciences 8:2403–2437
Ge TD, Yuan HZ, Zhu HH, Wu XH, Nie SA, Liu C, Tong CL, Wu J, Brookes P (2012) Biological carbon assimilation and dynamics in a flooded rice–soil system. Soil Biol Biochem 48:39–46
Ge T, Liu C, Yuan H, Zhao Z, Wu X, Zhu Z, Brookes P, Wu J (2015) Tracking the photosynthesized carbon input into soil organic carbon pools in a rice soil fertilized with nitrogen. Plant Soil 392:17–25
Göttlicher GS, Steinmann K, Betson N, Högberg P (2006) The dependence of soil microbial activity on recent photosynthate from trees. Plant Soil 287:85–94
Grayston SJ, Griffith GS, Mawdsley JL, Campbell CD, Bardgett RD (2001) Accounting for variability in soil microbial communities of temperate upland grassland ecosystems. Soil Biol Biochem 33:533–551
Gregory PJ (2006) Roots, rhizosphere and soil: the route to a better understanding of soil science? Eur J Soil Sci 57:2–12
Gunina A, Kuzyakov Y (2015) Sugars in soil and sweets for microorganisms: review of origin, content, composition and fate. Soil Biol Biochem 90:87–100
Hafner S, Wiesenberg GLB, Stolnikova E, Merz K, Kuzyakov Y (2014) Spatial distribution and turnover of root-derived carbon in alfalfa rhizosphere depending on top- and subsoil properties and mycorrhization. Plant Soil 380:101–115
Hannula SE, Boschker HTS, Boer WD, Veen JAV (2012) 13C-pulse-labeling assessment of the community structure of active fungi in the rhizosphere of a genetically starch-modified potato (Solanum tuberosum) cultivar and its parental isoline. New Phytol 194:784–799
He Y, Siemens J, Amelung W, Goldbach H, Wassmann R, Alberto MCR, Lücke A, Lehndorff E (2015) Carbon release from rice roots under paddy rice and maize–paddy rice cropping. Agric Ecosyst Environ 210:15–24
Hill GT, Mitkowski NA, Aldrich-Wolfe L, Emele LR, Jurkonie DD, Ficke A, Maldonado-Ramireza S, Lyncha ST, Nelsona EB (2000) Methods for assessing the composition and diversity of soil microbial communities. Appl Soil Ecol 15:25–36
Högberg P, Read DJ (2006) Towards a more plant physiological perspective on soil ecology. Trends Ecol Evol 21:548–554
Högberg P, Högberg MN, Göttlicher SG, Betson NR, Keel SG, Metcalfe DB, Campbell C, Schindlbacher A, Hurry V, Lundmark T, Linder S, Näsholm T (2008) High temporal resolution tracing of photosynthate carbon from the tree canopy to forest soil microorganisms. New Phytol 177:220–228
Keeney DR, Nelson DW (1982) Nitrogen inorganic forms. In: Page AL, Miller RH, Keeney DR (eds) Methods of soil analysis. Agronomy monograph 9 part 2, 2nd edn. American Society of Agronomy, Madison, pp 643–698
Knudsen D, Peterson GA, Pratt PF (1982) Lithium, sodium, and potassium. In: Page AL, Miller RH, Keeny DR (Eds) Methods of soil analysis. Part 2. Chemical and Microbiological Properties, 2nd ed. Agronomy No. 9. American Society of Agronomy, Madison, WI, pp 225–246
Kögel-Knabner I, Amelung W, Cao Z, Fiedler S, Frenzel P, Jahn R, Kalbitz K, Kölbl A, Schloter M (2010) Biogeochemistry of paddy soils. Geoderma 157:1–14
Kuzyakov Y, Cheng W (2001) Photosynthesis controls of rhizosphere respiration and organic matter decomposition. Soil Biol Biochem 33:1915–1925
Kuzyakov Y, Domanski G (2000) Carbon input by plants into the soil. Review. J Plant Nutr Soil Sci 163:421–431
Kuzyakov Y, Domanski G (2002) Model for rhizodeposition and CO2 efflux from planted soil and its validation by 14C pulse labeling of ryegrass. Plant Soil 239:87–102
Kuzyakov Y, Gavrichkova O (2010) Review: time lag between photosynthesis and carbon dioxide efflux from soil: a review of mechanisms and controls. Glob Chang Biol 16:3386–3406
Kuzyakov Y, Larionova AA (2005) Root and rhizomicrobial respiration: a review of approaches to estimate respiration by autotrophic and heterotrophic organisms in soil. J Plant Nutr Soil Sci 168:503–520
Kuzyakov Y, Leinweber P, Sapronov D, Eckhardt K (2003) Qualitative assessment of rhizodeposits in non-sterile soil by analytical pyrolysis. J Plant Nutr Soil Sci 166:719–723
Lal R (2004) Offsetting China’s CO2 emissions by soil carbon sequestration. Clim Chang 65:263–275
Lu Y, Watanabe A, Kimura M (2002a) Input and distribution of photosynthesized carbon in a flooded rice soil. Glob Biogeochem Cycles 16:32-1–32-8
Lu Y, Watanabe A, Kimura M (2002b) Contribution of plant-derived carbon to soil microbial biomass dynamics in a paddy rice microcosm. Biol Fertil Soils 36:136–142
Lu Y, Abraham W, Conrad R (2007) Spatial variation of active microbiota in the rice rhizosphere revealed by in situ stable isotope probing of phospholipid fatty acids. Environ Microbiol 9:474–481
Meng F, Dungait JAJ, Zhang X, He M, Guo Y, Wu W (2013) Investigation of photosynthate-C allocation 27 days after 13C-pulse labeling of Zea mays. at different growth stages. Plant Soil 1–2:755–764
Moran-Zuloaga D, Dippold MA, Glaser B, Kuzyakov Y (2015) Organic nitrogen uptake by plants: reevaluation by position-specific labeling of amino acids. Biogeochemistry 125:1–16
Nawaz MF, Bourrie G, Gul S, Trolard F, Ahmad I, Tanvir MA, Mouret J (2012) Impacts of rice plant roots on the variation in electro-physico-chemical properties of soil waters. Pak J Bot 44:1891–1896
Nguyen C (2003) Rhizodeposition of organic C by plant: mechanisms and controls. Agronomie 23:375–396
Nikolausz M, Kappelmeyer U, Székely A, Rusznyák A, Márialigeti K, Kästner (2008) Diurnal redox fluctuation and microbial activity in the rhizosphere of wetland plants. Eur J Soil Biol 44:324–333
Nobel PS (2005) Physicochemical and environmental plant physiology, 3rd edn. Elsevier Academic Press, New York
Olsen SR, Somers LE (1982) Phosphorus. In: Page AL, Miller RH, Keene DR (eds) Methods of soil analysis, vol 2. Soil Science Society of America, Madison, pp 403–448
Olsson PA, Thingstrup I, Jakobsen I, Bååth E (1999) Estimation of the biomass of arbuscular mycorrhizal fungi in a linseed field. Soil Biol Biochem 31:1879–1887
Palacios OA, Bashan Y, De-Bashan LE (2014) Proven and potential involvement of vitamins in interactions of plants with plant growth-promoting bacteria—an overview. Biol Fertil Soils 50:415–432
Paterson E, Gebbing T, Abel C, Sim A, Telfer G (2007) Rhizodeposition shapes rhizosphere microbial community structure in organic soil. New Phytol 173:600–610
Paterson E, Midwoo AJ, Millard P (2009) Through the eye of the needle: a review of isotope approaches to quantify microbial processes mediating soil carbon balance. New Phytol 184:19–33
Pausch J, Tian J, Riederer M, Kuzyakov Y (2013) Estimation of rhizodeposition at field scale: upscaling of a 14C labeling study. Plant Soil 364:273–285
Philippot L, Raaijmakers J, Lemanceau P, Putten VDWH (2013) Going back to the roots: the microbial ecology of the rhizosphere. Nat Rev Microbiol 11:789–799
Pii Y, Mimmo T, Tomasi N, Terzano R, Cesco S, Crecchio C (2015) Microbial interactions in the rhizosphere: beneficial influences of plant growth-promoting rhizobacteria on nutrient acquisition process. A review. Biol Fertil Soils 51:403–415
Priha O, Grayston SJ, Hiukka R, Pennanen T, Smolander A (2001) Microbial community structure and characteristics of the organic matter in soils under Pinus sylvestris, Picea abies and Betula pendula at two forest sites. Biol Fertil Soils 33:17–24
Qin H, Wang H, James Strong P, Li Y, Xu Q, Wu Q (2014) Rapid soil fungal community response to intensive management in a bamboo forest developed from rice paddies. Soil Biol Biochem 68:177–184
Ragnarsdottir KV, Hawkins DP (2006) Bioavailable copper and manganese in soils from Iceland and their relationship with scrapie occurrence in sheep. J Geochem Explor 88:228–234
Ramanathan K, Krishnamoorthy K (1973) Nutrient uptake by paddy during the main three stages of growth. Plant Soil 39:29–33
Rhoades JD (1982) Cation exchangeable capacity. In: Page AL, Miller RH, Keene DR (eds) Methods of soil analysis: part 2—chemical and microbiological properties, 2nd edn. American Society of Agronomy, Inc. and Soil Science Society of America, Inc, Madison, pp 149–165
Roth PJ, Lehndorff E, Hahn A, Frenzel P, Amelung W (2013) Cycling of rice rhizodeposits through peptide-bound amino acid enantiomers in soils under 50 and 2000 years of paddy management. Soil Biol Biochem 65:227–235
Shade A, Hogan CS, Klimowicz AK, Linske M, McManus PS, Handelsman J (2012) Culturing captures members of the soil rare biosphere. Environ Microbiol 14:2247–2252
Sollins P, Homann P, Caldwell BA (1996) Stabilization and destabilization of soil organic matter: mechanisms and controls. Geoderma 74:65–105
Thompson MV, Holbrook MN (2003) Scaling phloem transport: water potential equilibrium and osmoregulatory flow. Plant Cell Environ 26:1561–1577
Thornton B, Zhang Z, Mayes RW, Hogberg MN, Midwood AJ (2011) Can gas chromatography combustion isotope ratio mass spectrometry be used to quantify organic compound abundance? Rapid Commun Mass Sp 25:2433–2438
Tian J, Dippold M, Pausch J, Blagodatskaya E, Fan M, Li X, Kuzyakov Y (2013) Microbial response to rhizodeposition depending on water regimes in paddy soils. Soil Biol Biochem 65:195–203
Tunlid A, White DC (1992) Biochemical analysis of biomass community structure, nutritional status and metabolic activity of microbial communities in soil. Soil Biol Biochem 7:229–262
Wang J, Chapmanc SJ, Yao H (2016) Incorporation of 13C-labelled rice rhizodeposition into soil microbial communities under different fertilizer applications. Appl Soil Ecol 101:11–19
Webster G, Watt LC, Rinna J, Fry JC, Evershed RP, Parkes RJ, Weightman AJ (2006) A comparison of stable-isotope probing of DNA and phospholipid fatty acids to study prokaryotic functional diversity in sulfate-reducing marine sediment enrichment slurries. Environ Microbiol 8:1575–1589
Werth M, Kuzyakov Y (2010) 13C fractionation at the root–microorganisms–soil interface: a review and outlook for partitioning studies. Soil Biol Biochem 42:1372–1384
White DC, Davis WM, Nickels JS, King JD, Bobbie RJ (1979) Determination of the sedimentary microbial biomass by extractible lipid phosphate. Oecologia 40:51–62
Wiesenberg GLB, Gocke M, Kuzyakov Y (2010) Fast incorporation of root-derived lipids and fatty acids into soil–evidence from a short term multiple 14CO2 pulse labelling experiment. Org Geochem 41:1049–1055
Wu J, Joergensen RG, Pommerening B, Chaussod R, Brookes PC (1990) Measurement of soil microbial biomass C by fumigation-extraction—an automated procedure. Soil Biol Biochem 22:1167–1169
Wu Y, Ding N, Wang G, Xu J, Wu J, Brookes PC (2009) Effects of different soil weights, storage times and extraction methods on soil phospholipid fatty acid analyses. Geoderma 150:171–178
Yao H, Chapman SJ, Thornton B, Paterson E (2015) 13C-PLFA: a key to open the soil microbial black box? Plant Soil 392:3–15
Yevdokimov I, Ruser R, Buegger F, Marx M, Munch JC (2006) Microbial immobilisation of 13 C rhizodeposits in rhizosphere and root-free soil under continuous 13C-labelling of oats. Soil Biol Biochem 38:1202–1211
Zelles L, Bai QY, Beck T, Beese F (1992) Signature fatty acids in phospholipids and lipopolysaccharides as indicators of microbial biomass and community structure in agricultural soils. Soil Biol Biochem 24:317–323
Acknowledgments
This study was financially supported by the National Natural Science Foundation of China (41430860; 41301275), NSFC Research Fund for International Young Scientists (41450110432), Royal Society Newton Advanced Fellowship (NA150182), China-ASEAN Talented Young Scientists Program (Myanmar—14-003), international cooperation and regional science and technology of Hunan Province (2015WK3044), and the Recruitment Program of High-end Foreign Experts of the State Administration of Foreign Experts Affairs awarded to Y. K. (GDW20144300204).
Author information
Authors and Affiliations
Corresponding author
Additional information
Hongzhao Yuan and Zhenke Zhu contributed equally to this work.
Rights and permissions
About this article
Cite this article
Yuan, H., Zhu, Z., Liu, S. et al. Microbial utilization of rice root exudates: 13C labeling and PLFA composition. Biol Fertil Soils 52, 615–627 (2016). https://doi.org/10.1007/s00374-016-1101-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00374-016-1101-0