Abstract
Compact plant growth is an economically important trait for many crops. In practice, compactness is frequently obtained by applying chemical plant growth regulators. In view of sustainable and environmental-friendly plant production, the search for viable alternatives is a priority for breeders. Co-cultivation and natural transformation using rhizogenic agrobacteria result in morphological alterations which together compose the Ri phenotype. This phenotype is known to exhibit a more compact plant habit, besides other features. In this review, we highlight the use of rhizogenic agrobacteria and the Ri phenotype with regard to sustainable plant production and plant breeding. An overview of described Ri lines and current breeding applications is presented. The potential of Ri lines as pre-breeding material is discussed from both a practical and legal point of view.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plant breeding is a continuous process in which the creation of novelty and new traits are primary goals (Louwaars et al. 2012; Morgan and Funnell 2018). Classical breeding by intra- and interspecific crosses has been used to obtain new traits and develop new varieties (Kuligowska et al. 2016). This strategy meets several restrictions related to time investment, available diversity in the gene pool, and fertility barriers (Horn 2002; Long et al. 2018). Agrobacterium-mediated transformation can be used to obtain new traits in crops that are not present in the existing gene pool (Casanova et al. 2005). However, this technique is often associated with the use of genetic vectors that typically results in a genetically modified organism (GMO) and is therefore limited in terms of its applicability for practical plant breeding in Europe (Gelvin 2003). Another strategy to increase variability in plants is Agrobacterium-mediated transformation by the use of wild type rhizogenic agrobacteria (Otten 2018a).
Rhizogenic agrobacteria are a group of bacteria with a pathogenic lifestyle and root-inducing ability, and consists of strains belonging to the Rhizobium radiobacter and Rhizobium rhizogenes species both carrying the root-inducing (Ri) plasmid (pRi) (Bosmans et al. 2017; White and Nester 1980b). The Ri plasmid confers pathogenicity to the bacteria and contains the mechanism to initiate a horizontal gene transfer event by which a part of the Ri plasmid, namely the transfer DNA (T-DNA), is transferred to the host plant (Chilton et al. 1982; Lacroix and Citovsky 2016). After successful transfer and integration of the T-DNA genes in the nuclear DNA, the host plant will undergo changes that are instigated by the foreign DNA with the primary symptom being the proliferation of roots from the infection site. This phenomenon was first described by Riker et al. (1930) as infectious or hairy roots. It was later understood that the hairy roots are synthesizing and excreting specific compounds, termed opines, that serve to create an ecological niche in which nutrients are provided for the agrobacteria (Moore et al. 1997; Petit et al. 1983). The fact that opine production is plasmid encoded suggested that also the rhizogenic response of infected tissues could be evoked by pRi T-DNA-specific genes (White et al. 1982). Multiple genes involved in hairy root formation were later identified and documented to have strong morphogenic effects (Huffman et al. 1984; Spena et al. 1987). These genes were consequently defined as “plast” genes by Levesque et al. (1988). The hairy roots, because of their genetic stability and vigorous growth in hormone-free media, became a popular tool for plant metabolic engineering and studying root biology in the form of hairy root cultures (Hamill and Lidgett 1997; Shanks and Morgan 1999). Tepfer (1984) found that, following pRi T-DNA integration, plant regeneration was possible, resulting in a modified phenotype of the regenerated plant termed the hairy root or Ri phenotype.
This knowledge was applied to obtain phenotypically distinct plants with useful agronomical traits. Initially, this was used to create composite plants consisting of a transformed root system and wild type aerial parts (Lambert and Tepfer 1992). Regeneration from transformed tissues resulted in the development of complete natural transgenic plants, which also allows sexual transmission of the phenotype (Costantino et al. 1984; Ooms et al. 1985; Tepfer 1984). In some plant species, such as Nicotiana glauca, Nicotiana tomentosiformis, and Ipomoea batatas, spontaneous regeneration from hairy roots under natural circumstances occurred, and the resulting genotypes were characterized (Chen et al. 2014; Kyndt et al. 2015; White et al. 1983). The natural occurrence of gene transfer to plants combined with the plethora of useful traits makes the technique of using wild type rhizogenic agrobacteria very promising in terms of practical plant breeding applications (Casanova et al. 2005; Otten 2018b; Tempé and Schell 1977). Moreover, Ri lines of Kalanchoe blossfeldiana have already been successfully applied in commercial plant breeding (Christensen et al. 2014; Lütken et al. 2012b).
The global trend towards a more sustainable plant production comes with many challenges. Plant breeders today are faced with restrictions on the use of plant growth regulators, crop protection substances, and GMO techniques. The search for viable alternatives is a priority. In this work, we highlight the potential of wild type rhizogenic agrobacteria in the light of sustainable plant production, how this technique can lead to the creation of novel phenotypes in a non-GMO setting, and its application in practical plant breeding.
Introduction of pRi oncogenes in crops
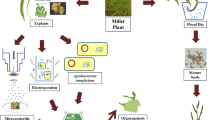
Transformation of plants using rhizogenic strains carrying a wild type Ri plasmid often relies on an in vitro co-cultivation protocol (Christensen and Müller 2009a; Karami 2008). The process is started by a selection of a wild type genotype that will be transformed (Fig. 1a). Ideally, an axenic culture of the plant is established, allowing fast propagation and year-round availability of plant material for protocol development. Specific parts, called explants, are harvested from the plantlets and used for further inoculation. Different types of explants vary in their susceptibility to bacterial infection, thus, multiple explant types are commonly tested (Cao et al. 2009; Ooi et al. 2013; Weber and Bodanese-Zanettini 2011). In some crops, the plant tissue culture can be quite challenging. For this reason, the tissue culture step is sometimes omitted by direct inoculation of surface-sterilized plant material or by the use of in vitro-germinated seedlings (Hegelund et al. 2017; Mugnier 1988). The second major component of the transformation protocol is the selection of a virulent rhizogenic strain and the preparation of an inoculum in a standardized way. Host range studies of R. rhizogenes have exhibited significant differences in plant susceptibility and revealed a specific strain relationship in terms of virulence with the plant host (De Cleene and De Ley 1981; Porter and Flores 1991). Therefore, multiple strains are often compared in terms of their virulence and transformation potential. In addition to the type of strain, also the preparation and density of the bacterial inoculum are important parameters influencing the success rate of co-cultivation and transformation (Desmet et al. 2019). Next, explants are submersed in a bacterial suspension for a fixed amount of time before being transferred to a co-cultivation medium. During the co-cultivation, the bacteria will infect the explants and transfer of pRi T-DNA occurs. Afterwards, the explants are submitted to an antibiotic treatment, which serves to eliminate residual R. rhizogenes from the explants. Successfully transformed explants will produce hairy roots from the sites of infection that serve as a source material for shoot regeneration (Tepfer 1984). Since no selectable markers are used, regenerated shoots are individually tested to confirm the transgenic nature. Transgenic-regenerated shoots are clonally propagated as unique Ri lines and are, after acclimatization to greenhouse conditions, further grown to facilitate phenotypic evaluation. Ri lines with interesting morphological features can then be further used in plant breeding (Fig. 1b).
Schematic overview of (a) natural pRi transformation. Numbers 1–4 depict steps in which optimization is needed: (1) selection of explant type and pre-culture; (2) selection of the bacterial strain and inoculum preparation; (3) method of inoculation and co-cultivation; (4) hairy root culture and regeneration. b application in plant breeding (WT, wild type; Ri, Ri line, plant derived from hairy root tissue; P, parental lines; R0, plants regenerated from hairy root tissue carrying pRi T-DNA genes; R1, first generation progeny of an Ri plant, ratios given apply to offspring of a single locus T-DNA insertion Ri line)
Ri plasmid
Rhizogenic agrobacteria carry an extrachromosomal circular DNA structure referred to as the Ri plasmid. This virulence plasmid is essential for the pathogenicity of the bacterium and contains several regions with high homology to the tumor inducing (Ti) plasmid (pTi) of tumorigenic agrobacteria (Huffman et al. 1984; White and Nester 1980b). Ri plasmids are classified as mega-plasmids because of their relative large size, ranging from 180 to 250 kbp (Petersen et al. 1989; Suzuki et al. 2009). Different types of plasmids are classified based on the type of opine produced by transformed tissues (Moore et al. 1997). To date, four different opine and Ri plasmid (pRi) types have been described: agropine (pRiA4 of type strain A4), cucumopine (pRi2659 of type strain NCPPB2659), mannopine (pRi8196 of type strain NCIB8196), and mikimopine (pRi1724 of type strain MAFF301724) (Fig. 2) (Vladimirov et al. 2015). Parts of the plasmids with high similarity that are not transferred to the host plant include the virulence region (Hooykaas et al. 1984; White and Nester 1980a), the origin of replication (Jouanin et al. 1985), and the opine catabolism region (Moore et al. 1997). The transferred part of the Ri plasmid, the T-DNA, carries genes for opine biosynthesis, genes necessary for the initiation of hairy roots, among which the root oncogenic loci (rol genes) and other genes of unidentified function, i.e., open reading frames (ORFs) (Sinkar et al. 1987; Slightom et al. 1986). The T-DNA of agropine-type strains is physically split into the two parts: TL-DNA and TR-DNA. The TL-DNA of agropine strains shares homology with the T-DNA of cucumopine, mannopine, and mikimopine strains, which consists of a single T-DNA (Jouanin 1984). In comparison, the TR-DNA shares homology with pTi T-DNA in loci responsible for agropine synthesis and the auxin biosynthesis genes (aux1 and aux2 of pRi being homologous to tms1 and tms2 of pTi) (Camilleri and Jouanin 1991).
The phenotypic effects observed in Ri lines are the direct consequence of transfer, integration, and expression processes of pRi T-DNA to the host plant cell (Costantino et al. 1994). Upon successful completion of these steps, many of the T-DNA genes and ORFs will cause strong morphogenic effects that originate from underlying shifts in plant hormone metabolism. Through the last almost 40 years, numerous studies have addressed the phenotypes of the rol genes and ORFs in relation to the morphological changes observed in transformed plants. In comparison, the knowledge of the mechanisms per se and the protein functions mostly remain elusive. Many of the rol genes and ORFs lead to hormonal changes and altered cell division in plants. Some of the overall mechanisms are briefly described in the following. The rolB gene is the most well-characterized gene and known as a strong denominator of root induction as well as promoting floral meristems. Several modes of action have been proposed for the gene: tyrosine phosphatase (Filippini et al. 1996), indoxyl-beta-glucosidase stimulation of auxin binding to membranes (Estruch et al. 1991) as well as interaction with a 14-3-3 protein and subsequent localization in the nucleus (Moriuchi et al. 2004). rolC has been shown to have cytokinine beta-glucosidase activity. In respect to plant growth, ORF13 has been demonstrated to induce callus on carrot roots (Fründt et al. 1998), and when overexpressed, it led to extreme dwarfing as observed in Arabidopsis (Kodahl et al. 2016). Furthermore, rolB, rolC, and ORF13 resulted in dark green leaves. Moreover, leaf wrinkling was described for rolA, rolB, rolBTR, and ORF13. In this respect, it can be speculated that there is a connection with cell division. In contrast, the function of rolD has been elucidated; it encodes a functional cyclodeaminase converting ornithine to proline, and it has been described as a stress-related osmoprotectant (Trovato et al. 2001). It seems that the rol genes and ORFs act synergistically in altering the transformed plants phenotypically (Spena et al. 1987; White et al. 1985). Further studies are needed to unravel the exact mechanisms of the rol genes and ORFs. One interesting avenue to elaborate on the functions of rol genes and ORFs will be targeting their corresponding gene products and studying protein interactions. For more thorough information of the mechanisms, we refer to Britton et al. (2008); Bulgakov (2008); Costantino et al. (1994); Mauro et al. (2017); Nilsson and Olsson (1997); and Otten (2018a).
Rhizogenic agrobacteria and plant breeding
Ri phenotype
Plants regenerated from hairy roots have an altered phenotype, which is commonly referred to as the “Ri phenotype” or “hairy root (HR) phenotype” (Christey 2001). Features of the Ri phenotype originally listed by Tepfer (1984) include wrinkled leaves, increased rooting ability, reduced apical dominance, and changes in flower morphology. Numerous studies have since been performed to obtain naturally transformed plants for which similar traits have been described. In the current study, an extensive literature review was conducted to compile a list of described Ri phenotypes. However, only studies which met the following criteria were selected: (1) a rhizogenic strain with a wild type Ri plasmid was used for transformation, (2) full plant regeneration from hairy root tissue was described, and (3) a clear description of the obtained Ri phenotype was given. A total of 91 studies provided a clear description of the Ri phenotype of 80 distinct plant species. From the 95 descriptions, several commonly recurring alterations of plant morphology can be derived. Based on the original description of Tepfer (1984), four definitive Ri traits were defined that influence different parts of the plant: (1) compact growth, (2) altered leaf morphology, (3) altered root morphology, and (4) altered flowering and fertility. Each definitive trait is broadly defined, such that a gradient of variation is encompassed (Table 1).
The most prominent Ri trait is a degree of compact growth resulting from multiple morphological changes. Plants with shorter internodes display a reduced total plant height. Also, increased branching and enhanced axillary bud growth, alongside with reduced apical dominance, further contribute to the compact growth habit of the plant. The change in compactness is one-directional, i.e., all described Ri plants have a similar or more compact growth habit compared with their wild type control. Shorter internodes of Ri lines have been reported in a wide range of plant species such as Catharanthus roseus, Prunus avium x Prunus pseudocerasus, Allocasuarina verticillata, and Lavandula x intermedia (Choi et al. 2004; Gutièrrez-Pesce et al. 1998; Phelep et al. 1991; Tsuro and Ikedo 2011). The reduction in apical dominance is observed through increased branching in Ri lines of Centaurium erythraea and Hyoscyamus muticus (Piatczak et al. 2006; Sevón et al. 1997). However, in some cases, the changes in terms of plant architecture have a compensating effect on each other. For example, a Plumbago indica Ri line exhibited longer shoots; however, the increase in individual shoot length was compensated for by a decrease in internodal length, hence maintaining a similar degree of compactness (Gangopadhyay et al. 2010). A comparable compensatory mechanism for the number of internodes and their individual length has been observed for the Ri phenotype of Datura arborea (Giovannini et al. 1997). However, for K. blossfeldiana, Mecardonia sp. and Medicago sativa, this compensation does not occur, and a more compact plant phenotype was obtained (Christensen et al. 2008; Pérez de la Torre et al. 2018; Spanò et al. 1987).
Altered leaf morphology is the second most frequent observed change in Ri phenotypes. In most cases, this consists of strong wrinkling of the leaves, coincided by changes in leaf size. In addition to this, differences in leaf pigmentation are common. Several reports correlate the coloration of darker green leaves with higher chlorophyll content (Jaziri et al. 1994; Koike et al. 2003); however, also more pale and yellowish leaves have been observed (Mehrotra et al. 2013). Discoloration and strong wrinkling effects could potentially have an effect on the growth, photosynthetic capacity, and yield of Ri plants, especially when this phenotype progresses to be more extreme as the plant ages (Giovannini et al. 1997; Webb et al. 1990). More recently, Rugini et al. (2015) investigated chlorophyll a and b content in leaves of P. avium x P. pseudocerasus Ri lines and found no significant difference compared with wild type plants. In some plant species, a change in the number of leaves per Ri plant has been noted: an increased number of leaves was observed in the species Pelargonium fragrans, Pelargonium odoratissimus, Pelargonium quercifolia, Pelargonium graveolens, and Ajuga reptans (Pellegrineschi and Davolio-Mariani 1996; Saxena et al. 2007; Tanaka and Matsumoto 1993), whereas a decreased number of leaves were found for Digitalis purpurea and Rhaponticum carthamoides (Koga et al. 2000a; Skała et al. 2019). The leaf number increase is developmentally correlated with the number of nodes and branching pattern. However, for D. purpurea, also increased branching was observed (Koga et al. 2000a). An important difference to note here is that both incidences of reported decrease in the number of leaves per plant relate to rosette forming species.
The third foremost trait groups changes to the root morphology of Ri plants. In two-thirds of the Ri phenotypes, a clear alteration in root morphology was reported. This trait relates back to the biological essence of the hairy root syndrome (Moore et al. 1979; Nester 2015). Infected hosts are, by natural means of T-DNA insertion, altered to produce differentiated, neoplastic roots (Costantino et al. 1994). The implication is that hairy roots carry a genetically underlying basis that enhances adventitious root formation. Hairy roots are characterized by fast growth, a high degree of lateral branching, and plagiotropic development (Christey 2001; Tepfer 2016). Likewise, in Ri plants, this genetic basis leads to enhanced rooting ability as observed in many Ri phenotypes, both in vitro as in greenhouse or field conditions. This more extensive root system has pronounced implications for the applicability of Ri plants. Firstly, the increase in rooting ability can lead to efficient vegetative propagation and better adaptation to ex vitro conditions (Casanova et al. 2005). Moreover, Ri plants with better developed root systems are promising in terms of sustainable plant production/agriculture, because of improved water and nutrient management, with for example better drought tolerance (Tepfer 2016). Secondly, increases in root biomass are commonly reported, a trait especially beneficial for root crops or when roots are used for extracting specific metabolites (Chaudhuri et al. 2006; Gangopadhyay et al. 2010; Majumdar et al. 2011; Pérez de la Torre et al. 2018; Sevón et al. 1997).
A fourth major change that can be expected encompasses changes in flower morphology, flowering onset and fertility of the plant. While decreases in flower size have been reported in Ri lines of Rudbeckia hirta, Brassica napus, Rehmannia elata, Ipomoea trichocarpa, and P. graveolens (Daimon and Mii 1995; Hegelund et al. 2018; Kim et al. 2012; Otani et al. 1996; Saxena et al. 2007), an increase in flower size has not been reported. Another compensatory mechanism seems to be present in terms of flowering, with the size and number of flowers per plant being inversely correlated, i.e., reduced flower size generally coincides with an increase in the number of flowers. However, Hosokawa et al. (1997) reported Ri lines of Gentiana triflora x Gentiana scabra to have an increased number of flowers without apparent changes to the size of individual flowers. In this case, the increase in flower number is consequential of increased branching and a reduction in apical dominance. The onset of flowering is also influenced by the presence of pRi T-DNA, with delayed flowering being the most prominent change. Earlier flowering has only been observed for Helichrysum stoechas and G. scabra Ri lines (Giovannini 2006; Suginuma and Akihama 1995).
An important point to address is phenotypic variability of Ri lines within the same plant species. Pronounced morphological differences in Ri lines of G. triflora x G. scabra (Hosokawa et al. 1997), P. avium x P. pseudocerasus (Gutièrrez-Pesce et al. 1998; Rugini et al. 2015), and B. napus (Guerche et al. 1987; Hegelund et al. 2018) have been reported. Similarly, the fertility of Antirrhinum majus Ri lines varied (Handa 1992a; Hoshino and Mii 1998). The range of phenotypic variation is useful in a plant breeding program; different Ri lines can be used to introduce specific traits in crops. Since it is known that hairy roots are clonal of origin (Chilton et al. 1982; Van Sluys and Tempé 1989), differences in Ri phenotype severity have been ascribed to several influencing variables, such as (1) the plant species and genotype, (2) the bacterial strain used for transformation, and (3) the genetic details of the transformation event itself (Christensen and Müller 2009a; Gelvin 2017; Koncz and Schell 1992). Genetic details which render Ri lines a unique character include the following: the inserted copy number of pRi oncogenes, complete or truncated transfer of the T-DNA, expression levels of the transferred genes, and positional integration effects (Golds et al. 1991; Hänisch ten Cate et al. 1990; Lütken et al. 2012a; Sun et al. 1991).
The importance of the rhizogenic phenotype in breeding applications
Natural pRi T-DNA transformation, through underlying genetic changes, results in plants that display the rhizogenic phenotype. Since wild type strains are used in this technique, the processing, transfer, and integration of the T-DNA are governed by natural means. This implies that the transformation event is pleiotropic in its effects on the host plant. Multiple phenotypic traits are affected by one transformation event, leading to changes which, depending on the point of view of the breeder and the specific use of the plant species, can be interpreted as wanted or unwanted changes (Christey 2001).
The importance of plant architecture and its modification through plant breeding has long been recognized, with compact plant architecture being a desirable trait in agriculture and horticulture (Coyne 1980; Hammerschlag and Smigocki 1998; Hauptli et al. 1990; Lütken et al. 2012a). In many crops, such as legume and cereal grains and sugarcane, compactness and branching pattern are important traits in relationship to lodging and productivity (Berry et al. 2004; Heath et al. 1994; Pribil et al. 2007). In ornamentals, compact growth is inherently linked to plant quality and economic viability of the plant product (Bergstrand 2017; Rademacher 2015). Ri lines of K. blossfeldiana with increased branching, reduced apical dominance, and shorter internodes have already been successfully applied in relation to commercial plant breeding to enhance compact growth (Christensen and Müller 2009a; Lütken et al. 2012b). Likewise, Koga et al. (2000a) created D. purpurea Ri lines to obtain dwarf cultivars. Alterations of leaf color, flower morphology, or the number of flowers can improve the ornamental value of plants (Casanova et al. 2005; Kim et al. 2012). The increased rooting ability of Ri lines has already been used to facilitate rooting of Malus x domestica cuttings (Lambert and Tepfer 1992). On the other hand, changes such as severe leaf wrinkling could negatively impact the photosynthetic capacity of crop canopy or reduce the visual appeal of ornamentals. For generatively propagated crops, reduced fertility is unwanted. In addition, a delay in flowering time could negatively affect production time for many ornamentals (Lütken et al. 2012b).
Due to the fact that in Ri lines, both desired as undesired traits can occur; a plant breeding approach based on introgression of specific Ri traits should be used. The implementation of Ri lines in existing breeding programs can offer an elegant way to obtain a phenotype with specific Ri traits, while maintaining the initial commercial/breeding value of the line after subsequent back-crossing (Christensen et al. 2008; Otten 2018b). Moreover, it has been well established that the inserted T-DNA is transmissible through meiosis and that the Ri phenotype is inheritable, mainly in a dominant Mendelian fashion (David et al. 1984; Durand-Tardif et al. 1985). Therefore, the segregation ratio of pRi T-DNA inserts is sufficiently high to allow further breeding.
To date, a limited number of studies have investigated pRi T-DNA inheritance in detail. Out of the aforementioned references with described Ri phenotypes, those with (1) a clear description of a breeding step with an Ri line as one of the parents and (2) described inheritance of the Ri phenotype were selected. In addition to this, the breeding with Ri lines described by Durand-Tardif et al. (1985), Limami et al. (1998), and Lütken et al. (2012b) was included (Table 2). Similarly to the generation nomenclature of plants derived from genetic modification, the generations are designated as R0, R1, and R2 (Yin et al. 2004). From the described R1 and R2 generations, it is clear that inheritance of pRi T-DNA takes place, regardless of whether the fact that progeny was obtained by either selfing or crossing. It is generally accepted that T-DNA insertion leads to plants that are hemizygous for the locus of insertion, both for events in which single or multiple copies (as direct or inverted repeats) are inserted at the same locus (Gelvin 2017; Sridevi et al. 2006). Thus, an R0 plant with a single locus T-DNA insert will transmit this in a dominant Mendelian way with a phenotype/genotype ratio of 3:1/1:2:1 in the case of selfing and 1:1/1:1 for a backcross with a wild type parent (Fig. 1b). Segregation can be confirmed by either phenotypic or molecular methods. Since opines are known to diffuse through plant tissue and variability of the Ri phenotype is common (Hegelund et al. 2017; Zhan et al. 1988), molecular techniques such as polymerase-chain reaction (PCR) and Southern hybridization are preferred to confirm segregation ratios.
Expected Mendelian segregation for single and double locus insert Ri lines has been observed for Nierembergia scoparia and B. napus, respectively (Godo et al. 1997; Guerche et al. 1987; Jouanin et al. 1987). However, segregation distortion was observed in R1 generations of Daucus carota, Calibrachoa excellens, and A. majus (David et al. 1984; Gennarelli et al. 2009; Handa 1992a) (Table 2).
Plant genotype, Ri phenotype, and segregation are linked and mediated through T-DNA structure and expression. The Ri phenotype in itself, as a direct consequence of the transformation event and therefore the R0 genotype, is not always sufficient to fully explain segregation and heredity. Copy number and the number of insertion sites both influence segregation; as the number of insertion sites increases, so does the complexity of segregation patterns (Jouanin et al. 1987). Additionally, homozygosity for the Ri locus is thought not to lead to direct changes in Ri phenotype (Durand-Tardif et al. 1985; Tepfer 1984), but it is still desired from a breeding point of view because of its influence on segregation ratios (Martin-Tanguy et al. 1990; Passricha et al. 2016). In summary, differences in segregation ratios and subsequent variability of the Ri phenotype can be explained by the nature of the recombination and the genetic makeup of the R0 line used to create the progenies. Detailed molecular characterization of R0 lines thus provides highly relevant information for the breeding process.
Legislation and legal aspects
In respect to legislation, the first and most important issue to address is whether modified or unmodified bacterial strains of R. rhizogenes are used. Application of modified strains, i.e., strains where recombination of nucleic acids has been made, results in GMOs according to legislation in the European Union (EU) (European Union 2001, Directive 2001/18/EC, Annex I A, part 1§1). Hence, gene insertions, deletions, and genome editing such as Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR) combined with a CRISPR-associated protein 9 (Cas9), CRISPR/Cas9, all fall under the GMO definition. Consequently, using transformation with modified strains is not applicable as a commercial breeding method in the EU.
Interestingly, the same directive also describes exemptions from the rule defining which techniques do not result in a GMO in the EU; natural processes such as conjugation, transduction, and transformation belong to this category (European Union 2001, Directive 2001/18/EC, Annex I A, part 2§2). Based on the definition of the latter, usage and application of unmodified bacterial strains have also been termed natural transformation (Christensen and Müller 2009b; Christensen et al. 2008; Lütken et al. 2012a). This, therefore, represents an interesting avenue for potential commercial application based on natural transformation. The natural transformation concept is supported by several studies showing that a number of wild plants actually contain remnants of R. rhizogenes T-DNAs, i.e., they have been transformed naturally during plant speciation. To this group belongs, e.g., tobacco, sweet potato and several species of Linaria (Kovacova et al. 2014; Kyndt et al. 2015; Matveeva et al. 2012; Suzuki et al. 2002). A recent study even indicates that an additionally 23 species contain bacterial T-DNA (Matveeva and Otten 2019).
In Denmark, authorities have confirmed that plants obtained through the biotechnological method of using unmodified strains of R. rhizogenes are non-GMOs (Lütken et al. 2012a). Furthermore, in Japan, unmodified strains of R. rhizogenes similarly do not fall under the GMO definition (Mishiba et al. 2006). In the USA, plants transformed with potential plant pathogens have to be evaluated by the Animal and Plant Health Inspection Service (APHIS) under the United States Department of Agriculture (USDA). The criteria are described under APHIS’s Plant Protection Act 7 CFR part 340, “Introduction of Organisms and Products Altered or Produced Through Genetic Engineering Which Are Plant Pests or Which There Is Reason To Believe Are Plant Pests.” Based on this, APHIS stated that Kalanchoe obtained following natural transformation does not fall under the term “altered or produced through genetic engineering,” and hence, the plants do not require specific regulations as they do not fall under the Plant Protection Act 7 CFR part 340.
In respect to breeders’ rights of plants derived following natural transformation with R. rhizogenes, one strategy is to protect naturally transformed plant as new cultivars. This can apply if the plant clearly exhibits new and distinctive traits, which has to be described and registered officially. Moreover, it must be possible to propagate the plant consistently. However, one might be aware that the introduction of rol genes in a protected plant is actually leading to the creation of an essentially derived variety (EDV) which cannot be protected itself (Krieger et al. 2019).
Another strategy has also been pursued to protect interspecific hybrids of Kalanchoe transformed with unmodified R. rhizogenes strains with the aim of obtaining the desired intermediate plant height. The plants produced with this method are covered by a patent (EP2698432A1, Christensen et al. 2014). The disclosure of the patent specifically includes Kalanchoe interspecific hybrid plants, and considers rol-transformation in Kalanchoe species and hybrids. The methodology discloses production of rol-transformed Kalanchoe interspecific hybrid plants, as well as resultant rol-transformed Kalanchoe interspecific hybrid plants with novel phenotypes.
Conclusion
In terms of practical breeding applications, co-cultivation with rhizogenic agrobacteria has potential to deliver pre-breeding material. Having efficient tissue culture protocols for transformation and regeneration ensures the creation of many Ri lines. Thorough genetic identification of lines from unique transformation events will facilitate the implementation in conventional and molecular plant breeding. The exact segregation ratio of the pRi T-DNA and its accompanying phenotype, based on molecular characterization, is essential for accurate application in breeding programs and allows breeders to make informed decisions. To date, many of the studies involving the transformation of plants using wild type rhizogenic agrobacteria and the Ri phenotype end without the continuation to the plant breeding process. Furthermore, in those cases for which Ri breeding is studied, the majority of the generated knowledge is mostly based on qualitative observations that, while valuable, are at best indicative of the usefulness of Ri lines in plant breeding. There is a need for studies with well-defined Ri lines that integrate both phenotypic and molecular based approaches in a range of crops. Such knowledge and detailed examples will help the implementation of Ri breeding on a larger scale.
References
Banerjee S, Zehra M, Gupta MM, Kumar S (1997) Agrobacterium rhizogenes-mediated transformation of Artemisia annua: production of transgenic plants. Planta Med 63:467–468. https://doi.org/10.1055/s-2006-957737
Behera PR, Jena RC, Das A, Thirunavoukkarasu M, Chand PK (2016) Genetic stability and coumarin content of transformed rhizoclones and regenerated plants of a multi-medicinal herb, Hybanthus enneaspermus (L.) F. Muell. Plant Growth Regul 80:103–114. https://doi.org/10.1007/s10725-016-0145-3
Bergstrand KJI (2017) Methods for growth regulation of greenhouse produced ornamental pot- and bedding plants - a current review. Folia Hortic 29:63–74. https://doi.org/10.1515/fhort-2017-0007
Berry P, Sterling M, Spink J, Baker C, Sylvester-Bradley R, Mooney S, Tams A, Ennos A (2004) Understanding and reducing lodging in cereals. Advances in Agronomy 84:217–271. https://doi.org/10.1016/S0065-2113(04)84005-7
Bertoli A, Giovannini A, Ruffoni B, Di Guardo A, Spinelli G, Mazzetti M, Pistelli L (2008) Bioactive constituent production in St. John’s Wort in vitro hairy roots. Regenerated plant lines. J Agric Food Chem 56:5078–5082. https://doi.org/10.1021/jf0729107
Bosmans L, Moerkens R, Wittemans L, De Mot R, Rediers H, Lievens B (2017) Rhizogenic agrobacteria in hydroponic crops: epidemics, diagnostics and control. Plant Pathol 66:1043–1053. https://doi.org/10.1111/ppa.12687
Brillanceau M-H, David C, Tempé J (1989) Genetic transformation of Catharanthus roseus G. Don by Agrobacterium rhizogenes. Plant Cell Rep 8:63–66. https://doi.org/10.1007/BF00716839
Britton MT, Escobar MA, Dandekar AM (2008) The oncogenes of Agrobacteriumtumefaciens and Agrobacterium rhizogenes.In: Tzfira T, CitovskyV (eds) Agrobacterium: from biology to biotechnology. Springer New York, New York, pp 523–563
Bulgakov VP (2008) Functions of rol genes in plant secondary metabolism. Biotechnol Adv 26:318–324. https://doi.org/10.1016/j.biotechadv.2008.03.001
Camilleri C, Jouanin L (1991) The TR-DNA region carrying the auxin synthesis genes of the Agrobacterium rhizogenes agropine-type plasmid pRiA4: nucleotide sequence analysis and introduction into tobacco plants. Mol Plant-Microbe Interact 4:155–162
Cao D, Hou W, Song S, Sun H, Wu C, Gao Y, Han T (2009) Assessment of conditions affecting Agrobacterium rhizogenes-mediated transformation of soybean. Plant Cell Tissue Organ Cult 96:45–52. https://doi.org/10.1007/s11240-008-9458-x
Casanova E, Trillas MI, Moysset L, Vainstein A (2005) Influence of rol genes in floriculture. Biotechnol Adv 23:3–39. https://doi.org/10.1016/j.biotechadv.2004.06.002
Celma RC, Palazón J, Cusidó RM, Piñol MT, Keil M (2001) Decreased scopolamine yield in field-grown Duboisia plants regenerated from hairy roots. Planta Med 67:249–253. https://doi.org/10.1055/s-2001-12006
Chaudhuri KN, Ghosh B, Tepfer D, Jha S (2006) Spontaneous plant regeneration in transformed roots and calli from Tylophora indica: changes in morphological phenotype and tylophorine accumulation associated with transformation by Agrobacterium rhizogenes. Plant Cell Rep 25:1059–1066. https://doi.org/10.1007/s00299-006-0164-z
Chen K, Dorlhac De Borne F, Szegedi E, Otten L (2014) Deep sequencing of the ancestral tobacco species Nicotiana tomentosiformis reveals multiple T-DNA inserts and a complex evolutionary history of natural transformation in the genus Nicotiana. Plant J 80:669–682. https://doi.org/10.1111/tpj.12661
Chilton M-D, Tepfer DA, Petit A, David C, Casse-Delbart F, Tempé J (1982) Agrobacterium rhizogenes inserts T-DNA into the genomes of the host plant root cells. Nature 295:432–434. https://doi.org/10.1038/295432a0
Choi PS, Kim YD, Choi KM, Chung HJ, Choi DW, Liu JR (2004) Plant regeneration from hairy-root cultures transformed by infection with Agrobacterium rhizogenes in Catharanthus roseus. Plant Cell Rep 22:828–831. https://doi.org/10.1007/s00299-004-0765-3
Christensen B, Müller R (2009a) The use of Agrobacterium rhizogenes and its rol-genes for quality improvement in ornamentals. Eur J Hortic Sci 74:275–287
Christensen B, Müller R (2009b) Kalanchoe blossfeldiana transformed with rol genes exhibits improved postharvest performance and increased ethylene tolerance. Postharvest Biol Technol 51:399–406. https://doi.org/10.1016/j.postharvbio.2008.08.010
Christensen B, Sriskandarajah S, Serek M, Müller R (2008) Transformation of Kalanchoe blossfeldiana with rol-genes is useful in molecular breeding towards compact growth. Plant Cell Rep 27:1485–1495. https://doi.org/10.1007/s00299-008-0575-0
Christensen E, Müller R, Lütken H, Hegelund J, Jensen L, Christensen B, Bollerup E, Nielsen K (2014) Agrobacterium rhizogenes transformation and expression of rol genes in Kalanchoë
Christey MC (2001) Invited review: use of Ri-mediated transformation for production of transgenic plants. Vitr Cell Dev Biol - Plant 37:687–700. https://doi.org/10.1079/IVP2001203
Christey MC, Sinclair BK (1993) Field-testing of kapeti kale regenerated from Agrobacterium-induced hairy roots. New Zeal J Agric Res 36:389–392. https://doi.org/10.1080/00288233.1993.10417738
Costantino P, Spanò L, Pomponi M, Benvenuto E, Ancora G (1984) The T-DNA of Agrobacterium rhizogenes is transmitted through meiosis to the progeny of hairy root plants. J Mol Appl Genet 2:465–470
Costantino P, Capone I, Cardarelli M, De Paolis A, Mauro ML, Trovato M (1994) Bacterial plant oncogenes: the rol genes’ saga. Genetica 94:203–211. https://doi.org/10.1007/BF01443434
Coyne D (1980) Modification of plant architecture and crop yield by breeding. HortScience 15:244–247
Daimon H, Mii M (1995) Plant regeneration and thiophene production in hairy root cultures of Rudbeckia hirta L. used as an antagonistic plant to nematodes. Japanese J Crop Sci 64:650–655. https://doi.org/10.1626/jcs.64.650
David C, Tempé J (1988) Genetic transformation of cauliflower (Brassica oleracea L. var. botrytis) by Agrobacterium rhizogenes. Plant Cell Rep 7:88–91. https://doi.org/10.1007/BF00270111
David C, Chilton M-D, Tempé J (1984) Conservation of T-DNA in plants regenerated from hairy root cultures. Nat Biotechnol 2:73–76. https://doi.org/10.1038/nbt0184-73
De Cleene M, De Ley J (1981) The host range of infectious hairy-root. Bot Rev 47:147–194. https://doi.org/10.1007/BF02868853
Desmet S, De Keyser E, Van Vaerenbergh J, Baeyen S, Van Huylenbroeck J, Geelen D, Dhooghe E (2019) Differential efficiency of wild type rhizogenic strains for rol gene transformation of plants. Appl Microbiol Biotechnol 103:6657–6672. https://doi.org/10.1007/s00253-019-10003-0
di Guardo A, Cellarova E, Koperdakova J, Pistelli L, Ruffoni B, Allavena A, Giovannini A (2003) Hairy root induction and plant regeneration in Hypericum perforatum L. J Genet Breed 57:269–278
Durand-Tardif M, Broglie R, Slightom J, Tepfer D (1985) Structure and expression of Ri T-DNA from Agrobacterium rhizogenes in Nicotiana tabacum. J Mol Biol 186:557–564. https://doi.org/10.1016/0022-2836(85)90130-5
Estruch JJ, Schell J, Spena A (1991) The protein encoded by the rolB plant oncogene hydrolyses indole glucosides. EMBO J 10:3125–3128
European Union (2001) Directive 2001/18/EC of the European Parliament and of the Council of 12 March 2001 on the deliberate release into the environment of genetically modified organisms and repealing Council Directive 90/220/EEC (OJ L 106 17.04.2001 p. 1). In: Sands P, Galizzi P (eds) Documents in European Community Environmental Law. Cambridge University Press, Cambridge, pp 787–836
Filippini F, Rossi V, Marin O, Trovato M, Costantino P, Downey PM, Lo Schiavo F, Terzi M (1996) A plant oncogene as a phosphatase. Nature 379:499–500
Fründt C, Meyer AD, Ichikawa T, Meins F (1998) A tobacco homologue of the Ri-plasmid orf13 gene causes cell proliferation in carrot root discs. Mol Gen Genet 259:559–568. https://doi.org/10.1007/s004380050849
Gangopadhyay M, Chakraborty D, Bhattacharyya S, Bhattacharya S (2010) Regeneration of transformed plants from hairy roots of Plumbago indica. Plant Cell Tissue Organ Cult 102:109–114. https://doi.org/10.1007/s11240-010-9702-z
Gelvin SB (2003) Agrobacterium-mediated plant transformation: the biology behind the “gene-jockeying” tool. Microbiol Mol Biol Rev 67:16–37. https://doi.org/10.1128/MMBR.67.1.16
Gelvin SB (2017) Integration of Agrobacterium T-DNA into the plant genome. Annu Rev Genet 51:195–217. https://doi.org/10.1146/annurev-genet-120215-035320
Gennarelli MC, Hagiwara JC, Tosto D, Álvarez MA, Borja M, Escandóni AS (2009) Genetic transformation of Calibrachoa excellens via Agrobacterium rhizogenes: changing morphological traits. J Hortic Sci Biotechnol 84:305–311. https://doi.org/10.1080/14620316.2009.11512522
Giovannini A (2006) Tissue culture, cell culture and genetic transformation by wild type Agrobacterium rhizogenes in mediterranean Helichrysum. In: de Silva J (ed) Floriculture, Ornamental and Plant Biotechnology Advances and Tropical Issues II. Global Science Books, Ikenobe, pp 222–226
Giovannini A, Allavena A, Pecchioni N (1996) Genetic transformation of lisianthus (Eustoma grandiflorum Griseb.) by Agrobacterium rhizogenes. J Genet Breed 50:35–40
Giovannini A, Pecchioni N, Rabaglio M, Allavena A (1997) Characterization of ornamental Datura plants transformed by Agrobacterium rhizogenes. Vitr Cell Dev Biol 33:101–106. https://doi.org/10.1007/s11627-997-0004-z
Giovannini A, Mascarello C, Pipino L, Nostro A (2008) Agrobacterium rhizogenes - mediated transformation in mediterranean Helichrysum. Transgenic Plant J 2:54–61
Godo T, Tsujii O, Ishikawa K, Mii M (1997) Fertile transgenic plants of Nierembergia scoparia Sendtner obtained by a mikimopine type strain of Agrobacterium rhizogenes. Sci Hortic (Amsterdam) 68:101–111. https://doi.org/10.1016/S0304-4238(96)00964-8
Golds T, Lee J, Husnain T, Ghose T, Davey M (1991) Agrobacterium rhizogenes mediated transformation of the forage legumes Medicago sativa and Onobrychis viciifolia. J Exp Bot 42:1147–1157. https://doi.org/10.1093/jxb/42.9.1147
Guerche P, Jouanin L, Tepfer D, Pelletier G (1987) Genetic transformation of oilseed rape (Brassica napus) by the Ri T-DNA of Agrobacterium rhizogenes and analysis of inheritance of the transformed phenotype. Mol Gen Genet 206:382–386. https://doi.org/10.1007/BF00428875
Gutièrrez-Pesce P, Taylor K, Muleo R, Rugini E (1998) Somatic embryogenesis and shoot regeneration from transgenic roots of the cherry rootstock Colt (Prunus avium x P. pseudocerasus) mediated by pRi 1855 T-DNA of Agrobacterium rhizogenes. Plant Cell Rep 17:574–580. https://doi.org/10.1007/s002990050445
Hamill JD, Lidgett AJ (1997) Hairy root cultures - opportunities and key protocols for studies in metabolic engineering. In: Doran P (ed) Hairy roots: culture and applications. Harwood Academic Publishers, Amsterdam, pp 1–29
Hamill JD, Rhodes MJC (1988) A spontaneous, light independent and prolific plant regeneration response from hairy roots of Nicotiana hesperis transformed by Agrobacterium rhizogenes. J Plant Physiol 133:506–509. https://doi.org/10.1016/S0176-1617(88)80046-4
Hammerschlag FA, Smigocki AC (1998) Growth and in vitro propagation of peach plants transformed with the shooty mutant strain of Agrobacterium tumefaciens. HortScience 33:897–899. https://doi.org/10.21273/HORTSCI.33.5.897
Handa T (1992a) Genetic transformation of Antirrhinum majus L. and inheritance of altered phenotype induced by Ri T-DNA. Plant Sci 81:199–206. https://doi.org/10.1016/0168-9452(92)90043-L
Handa T (1992b) Regeneration and characterization of prairie gentian (Eustoma grandiflorum) plants transformed by Agrobacterium rhizogenes. Plant tissue Cult Lett 9:10–14. https://doi.org/10.5511/plantbiotechnology1984.9.10
Handa T, Sugimura T, Kato E, Kamada H, Takayanagi K (1995) Genetic transformation of Eustoma grandiflorum with rol genes. Acta Hortic. https://doi.org/10.17660/ActaHortic.1995.392.25
Hänisch ten Cate CH, Ennik E, Roest S, Ramulu SK, Dijkhuis P, de Groot B (1988) Regeneration and characterization of plants from potato root lines transformed by Agrobacterium rhizogenes. Theor Appl Genet 75:452–459. https://doi.org/10.1007/BF00276749
Hänisch ten Cate CH, Loonen AEHM, Ottaviani MP, Ennik L, van Eldik G, Stiekema WJ (1990) Frequent spontaneous deletions of Ri T-DNA in Agrobacterium rhizogenes transformed potato roots and regenerated plants. Plant Mol Biol 14:735–741. https://doi.org/10.1007/BF00016506
Hauptli H, Katz D, Thomas BR, Goodman RM (1990) Biotechnology and crop breeding for sustainable agriculture. In: Edwards C, Lal R, Madden P, Miller R, House G (eds) Sustainable agricultural systems. Soil and Water Conservation Society, Ankeny, pp 141–156
Heath MC, Pilbeam CJ, McKenzie BA, Hebblethwaite PD (1994) Plant architecture, competitive ability and crop productivity in food legumes with particular emphasis on pea (Pisum sativum L.) and faba bean (Vicia faba L.). In: Muehlbauer F, Kaiser W (eds) Expanding the production and use of cool season food legumes. Current plant science and biotechnology in agriculture, vol 19. Springer, Dordrecht, pp 771–790
Hegelund JN, Lauridsen UB, Wallström SV, Müller R, Lütken H (2017) Transformation of Campanula by wild type Agrobacterium rhizogenes. Euphytica 213:2–9. https://doi.org/10.1007/s10681-017-1845-0
Hegelund JN, Liang C, Lauridsen UB, Kemp O, Lütken H, Müller R (2018) Increasing genetic variability in oilseed rape (Brassica napus) – genotypes and phenotypes of oilseed rape transformed by wild type Agrobacterium rhizogenes. Plant Sci 271:20–26. https://doi.org/10.1016/j.plantsci.2018.03.003
Henzelyová J, Čellárová E (2018) Modulation of naphthodianthrone biosynthesis in hairy root-derived Hypericum tomentosum regenerants. Acta Physiol Plant 40:82–12. https://doi.org/10.1007/s11738-018-2664-1
He-Ping S, Yong-Yue L, Tie-Shan S, Eric TPK (2011) Induction of hairy roots and plant regeneration from the medicinal plant Pogostemon Cablin. Plant Cell Tissue Organ Cult 107:251–260. https://doi.org/10.1007/s11240-011-9976-9
Hooykaas PJJ, Hofker M, den Dulk-Ras H, Schilperoort RA (1984) A comparison of virulence determinants in an octopine Ti plasmid, a nopaline Ti plasmid, and an Ri plasmid by complementation analysis of Agrobacterium tumefaciens mutants. Plasmid 11:195–205. https://doi.org/10.1016/0147-619X(84)90026-X
Horn W (2002) Breeding methods and breeding research. In: Vainstein A (ed) Breeding for ornamentals: classical and molecular approaches. Springer Netherlands, Dordrecht, pp 47–83
Hoshino Y, Mii M (1998) Bialaphos stimulates shoot regeneration from hairy roots of snapdragon (Antirrhinum majus L.) transformed by Agrobacterium rhizogenes. Plant Cell Rep 17:256–261. https://doi.org/10.1007/s002990050388
Hosokawa K, Matsuki R, Oikawa Y, Yamamura S (1997) Genetic transformation of gentian using wild-type Agrobacterium rhizogenes. Plant Cell Tissue Organ Cult 51:137–140. https://doi.org/10.1023/A:1005962913491
Hosoki T, Shiraishi K, Kigo T, Ando M (1989) Transformation and regeneration of ornamental kale (Brassica oleracea var. acephala DC) mediated by Agrobacterium rhizogenes. Sci Hortic (Amsterdam) 40:259–266. https://doi.org/10.1016/0304-4238(89)90118-0
Huang Y, Diner A, Karnosky D (1991) Agrobacterium rhizogenes-mediated genetic transformation and regeneration of a conifer: Larix decidua. Vitr Cell Dev Biol 27:201–207
Huffman GA, White FF, Gordon MP, Nester EW (1984) Hairy-root-inducing plasmid: physical map and homology to tumor-inducing plasmids. J Bacteriol 157:269–276
Jaziri M, Yoshimatsu K, Homès J, Shimomura K (1994) Traits of transgenic Atropa belladonna doubly transformed with different Agrobacterium rhizogenes strains. Plant Cell Tissue Organ Cult 38:257–262. https://doi.org/10.1007/BF00033885
Jia H, Zhao B, Wang X, Wang Y (2008) Agrobacterium rhizogenes-mediated transformation and regeneration of the Apocynum venetum. Chin J Biotechnol 24:1723–1728. https://doi.org/10.1016/S1872-2075(08)60071-0
Jouanin L (1984) Restriction map of an agropine-type Ri plasmid and its homologies with Ti plasmids. Plasmid 12:91–102. https://doi.org/10.1016/0147-619X(84)90055-6
Jouanin L, Vilaine F, D’Enfert C, Casse-Delbart F (1985) Localization and restriction maps of the replication origin regions of the plasmids of Agrobacterium rhizogenes strain A4. Mol Gen Genet 201:370–374. https://doi.org/10.1007/BF00331325
Jouanin L, Guerche P, Pamboukdjian N, Tourneur C, Delbart FC, Tourneur J (1987) Structure of T-DNA in plants regenerated from roots transformed by Agrobacterium rhizogenes strain A4. Mol Gen Genet 206:387–392
Kang HJ, Anbazhagan VR, You XL, Moon HK, Yi JS, Choi YE (2006) Production of transgenic Aralia elata regenerated from Agrobacterium rhizogenes-mediated transformed roots. Plant Cell Tissue Organ Cult 85:187–196. https://doi.org/10.1007/s11240-005-9070-2
Karami O (2008) Factors affecting Agrobacterium-mediated transformation of plants. Transgenic Plant J 2:127–137
Kim YS, Kim YK, Xu H, Uddin MR, Park NI, Kim HH, Chae SC, Park SU (2012) Improvement of ornamental characteristics in Rehmannia elata through Agrobacterium rhizogenes-mediated transformation. Plant Omics 5:376–380
Kodahl N, Müller R, Lütken H (2016) The Agrobacterium rhizogenes oncogenes rolB and ORF13 increase formation of generative shoots and induce dwarfism in Arabidopsis thaliana (L.) Heynh. Plant Sci 252:22–29. https://doi.org/10.1016/j.plantsci.2016.06.020
Koga M, Hirashima K, Nakahara T (2000a) The transformation system in foxglove (Digitalis purpurea L.) using Agrobacterium rhizogenes and traits of the regenerants. Plant Biotechnol 17:99–104. https://doi.org/10.5511/plantbiotechnology.17.99
Koga M, Hirashima K, Nakahara T (2000b) Genetic transformation in Vaccaria pyramidata using Agrobacterium rhizogenes. Plant Biotechnol 17:163–166. https://doi.org/10.5511/plantbiotechnology.17.163
Koike Y, Hoshino Y, Mii M, Nakano M (2003) Horticultural characterization of Angelonia salicariifolia plants transformed with wild-type strains of Agrobacterium rhizogenes. Plant Cell Rep 21:981–987. https://doi.org/10.1007/s00299-003-0608-7
Koncz C, Schell J (1992) Exploitation of Agrobacterium tumefaciens. In: Russo V, Ottolenghi S (eds) Development. Springer Berlin, Heidelberg, pp 217–224. https://doi.org/10.1007/978-3-642-77043-2_16
Koperdáková J, Komarovská H, Košuth J, Giovannini A, Čellárová E (2009) Characterization of hairy root-phenotype in transgenic Hypericum perforatum L. clones. Acta Physiol Plant 31:351–358. https://doi.org/10.1007/s11738-008-0241-8
Kovacova V, Zluvova J, Janousek B, Talianova M, Vyskot B (2014) The evolutionary fate of the horizontally transferred agrobacterial mikimopine synthase gene in the genera Nicotiana and Linaria. PLoS One 9:1–23. https://doi.org/10.1371/journal.pone.0113872
Krieger E, De Keyser E, De Riek J (2019) Do new breeding techniques in ornamentals and fruits lead to essentially derived varieties? Front Plant Sci 10:1612. https://doi.org/10.3389/fpls.2019.01612
Kuligowska K, Lütken H, Müller R (2016) Towards development of new ornamental plants: status and progress in wide hybridization. Planta 244:1–17. https://doi.org/10.1007/s00425-016-2493-7
Kyndt T, Quispe D, Zhai H, Jarret R, Ghislain M, Liu Q, Gheysen G, Kreuze JF (2015) The genome of cultivated sweet potato contains Agrobacterium T-DNAs with expressed genes: an example of a naturally transgenic food crop. Proc Natl Acad Sci 112:5844–5849. https://doi.org/10.1073/pnas.1419685112
Lacroix B, Citovsky V (2016) Transfer of DNA from bacteria to eukaryotes. MBio 7:1–9. https://doi.org/10.1128/mbio.00863-16
Lambert C, Tepfer D (1992) Use of Agrobacterium rhizogenes to create transgenic apple trees having an altered organogenic response to hormones. Theor Appl Genet 85:105–109. https://doi.org/10.1007/BF00223851
Lee MH, Yoon ES, Jeong JH, Choi YE (2004) Agrobacterium rhizogenes-mediated transformation of Taraxacum platycarpum and changes of morphological characters. Plant Cell Rep 22:822–827. https://doi.org/10.1007/s00299-004-0763-5
Levesque H, Delepelaire P, Rouzé P, Slightom J, Tepfer D (1988) Common evolutionary origin of the central portions of the Ri TL-DNA of Agrobacterium rhizogenes and the Ti T-DNAs of Agrobacterium tumefaciens. Plant Mol Biol 11:731–744. https://doi.org/10.1007/BF00019514
Limami M, Sun LY, Douat C, Helgeson J, Tepfer D (1998) Natural genetic transformation by Agrobacterium rhizogenes: annual flowering in two biennials, belgian endive and carrot. Plant Physiol 118:543–550
Long C, Chen Z, Zhou Y, Long B (2018) The role of biodiversity and plant conservation for ornamental breeding. In: Van Huylenbroeck J (ed) Ornamental Crops. Handbook of Plant Breeding. Springer Cham, Switzerland, pp 1–12. https://doi.org/10.1007/978-3-319-90698-0_1
Louwaars N, Dons H, Van Overwalle G, Raven H, Arundel A, Eaton D, Nelis A (2012) Breeding business: plant breeder's rights and patent rights in the plant breeding business. In: Improving Agricultural Knowledge and Innovation Systems: OECD Conference Proceedings, OECD Publishing, Paris, pp 263–277. https://doi.org/10.1787/9789264167445-en
Lütken H, Clarke JL, Müller R (2012a) Genetic engineering and sustainable production of ornamentals: current status and future directions. Plant Cell Rep 31:1141–1157. https://doi.org/10.1007/s00299-012-1265-5
Lütken H, Wallström SV, Jensen EB, Christensen B, Müller R (2012b) Inheritance of rol-genes from Agrobacterium rhizogenes through two generations in Kalanchoë. Euphytica 188:397–407. https://doi.org/10.1007/s10681-012-0701-5
Majumdar S, Garai S, Jha S (2011) Genetic transformation of Bacopa monnieri by wild type strains of Agrobacterium rhizogenes stimulates production of bacopa saponins in transformed calli and plants. Plant Cell Rep 30:941–954. https://doi.org/10.1007/s00299-011-1035-9
Martin-Tanguy J, Tepfer D, Paynot M, Burtin D, Heisler L, Martin C (1990) Inverse relationship between polyamine levels and the degree of phenotypic alteration induced by the root-inducing, left-hand transferred DNA from Agrobacterium rhizogenes. Plant Physiol 92:912–918. https://doi.org/10.1104/pp.92.4.912
Matveeva TV, Otten L (2019) Widespread occurrence of natural genetic transformation of plants by Agrobacterium. Plant Mol Biol 101:415–437. https://doi.org/10.1007/s11103-019-00913-y
Matveeva TV, Bogomaz DI, Pavlova OA, Nester EW, Lutova LA (2012) Horizontal gene transfer from genus Agrobacterium to the plant Linaria in nature. Mol Plant-Microbe Interact 25:1542–1551. https://doi.org/10.1094/MPMI-07-12-0169-R
Mauro ML, Costantino P, Bettini PP (2017) The never ending story of rol genes: a century after. Plant Cell Tissue Organ Cult 131:201–212. https://doi.org/10.1007/s11240-017-1277-5
McInnes E, Morgan AJ, Mulligan BJ, Davey MR (1991) Phenotypic effects of isolated pRiA4 TL-DNA rol genes in the presence of intact TR-DNA in transgenic plants of Solanum dulcamara L. J Exp Bot 42:1279–1286. https://doi.org/10.1093/jxb/42.10.1279
Mehrotra S, Goel MK, Rahman LU, Kukreja AK (2013) Molecular and chemical characterization of plants regenerated from Ri-mediated hairy root cultures of Rauwolfia serpentina. Plant Cell Tissue Organ Cult 114:31–38. https://doi.org/10.1007/s11240-013-0302-6
Mishiba KI, Nishihara M, Abe Y, Nakatsuka T, Kawamura H, Kodama K, Takesawa T, Abe J, Yamamura S (2006) Production of dwarf potted gentian using wild-type Agrobacterium rhizogenes. Plant Biotechnol 23:33–38. https://doi.org/10.5511/plantbiotechnology.23.33
Momčilović I, Grubišić D, Kojić M, Nešković M (1997) Agrobacterium rhizogenes-mediated transformation and plant regeneration of four Gentiana species. Plant Cell Tissue Organ Cult 50:1–6. https://doi.org/10.1023/A:1005880802231
Moore L, Warren G, Strobel G (1979) Involvement of a plasmid in the hairy root disease of plants caused by Agrobacterium rhizogenes. Plasmid 2:617–626. https://doi.org/10.1016/0147-619X(79)90059-3
Moore LW, Chilton WS, Canfield ML (1997) Diversity of opines and opine-catabolizing bacteria isolated from naturally occurring crown gall tumors. Appl Environ Microbiol 63:201–207
Morgan E, Funnell K (2018) Limonium. In: Van Huylenbroeck J (ed) Ornamental Crops. Handbook of Plant Breeding. Springer Cham, Switzerland, pp 513–52. https://doi.org/10.1007/978-3-319-90698-0_21
Moriuchi H, Okamoto C, Nishihama R, Yamashita I, Machida Y, Tanaka N (2004) Nuclear localization and interaction of rolB with plant 14-3-3 proteins correlates with induction of adventitious roots by the oncogene rolB. Plant J 38:260–275. https://doi.org/10.1111/j.1365-313X.2004.02041.x
Mugnier J (1988) Establishment of new axenic hairy root lines by inoculation with Agrobacterium rhizogenes. Plant Cell Rep 7:9–12. https://doi.org/10.1007/BF00272966
Nester EW (2015) Agrobacterium: nature’s genetic engineer. Front Plant Sci 5:1–16. https://doi.org/10.3389/fpls.2014.00730
Nilsson O, Olsson O (1997) Getting to the root: the role of the Agrobacterium rhizogenes rol genes in the formation of hairy roots. Physiol Plant 100:463–473. https://doi.org/10.1034/j.1399-3054.1997.1000307.x
Noda T, Tanaka N, Mano Y, Nabeshima S, Ohkawa H, Matsui C (1987) Regeneration of horseradish hairy roots incited by Agrobacterium rhizogenes infection. Plant Cell Rep 6:283–286. https://doi.org/10.1007/BF00271999
Oksman-Caldentey KM, Kivelä O, Hiltunen R (1991) Spontaneous shoot organogenesis and plant regeneration from hairy root cultures of Hyoscyamus muticus. Plant Sci 78:129–136. https://doi.org/10.1016/0168-9452(91)90169-9
Ooi CT, Syahida A, Stanslas J, Maziah M (2013) Efficiency of different agrobacterium rhizogenes strains on hairy roots induction in Solanum mammosum. World J Microbiol Biotechnol 29:421–430. https://doi.org/10.1007/s11274-012-1194-z
Ooms G, Karp A, Burrell MM, Twell D, Roberts J (1985) Genetic modification of potato development using Ri T-DNA. Theor Appl Genet 70:440–446. https://doi.org/10.1007/BF00273752
Otani M, Mii M, Handa T, Kamada H, Shimada T (1993) Transformation of sweet potato (Ipomoea batatas (L.) Lam.) plants by Agrobacterium rhizogenes. Plant Sci 94:151–159. https://doi.org/10.1016/0168-9452(93)90016-S
Otani M, Shimada T, Kamada H, Teraya H, Mii M (1996) Fertile transgenic plants of Ipomoea trichocarpa Ell. induced by different strains of Agrobacterium rhizogenes. Plant Sci 116:169–175. https://doi.org/10.1016/0168-9452(96)04394-4
Otten L (2018a) The Agrobacterium phenotypic plasticity (plast) genes. In: Gelvin S. (ed) Agrobacterium Biology. Current Topics in Microbiology and Immunology, Springer Cham, Switzerland, pp 375–419. https://doi.org/10.1007/82_2018_93
Otten L (2018b) How Agrobacterium, a natural genetic engineer, became a tool for modern agriculture. In: Kuntz M (ed) Advances in Botanical Research, Academic Press, Cambridge. pp 17–44. https://doi.org/10.1016/bs.abr.2017.11.002
Pal A, Swain SS, Mukherjee AK, Chand PK (2013) Agrobacterium pRI TL-DNA rolB and TR-DNA opine genes transferred to the spiny amaranth (Amaranthus spinosus L.), a nutraceutical crop. Food Technol Biotechnol 51:26–35
Passricha N, Saifi S, Khatodia S, Tuteja N (2016) Assessing zygosity in progeny of transgenic plants: current methods and perspectives. J Biol Methods 3:46. https://doi.org/10.14440/jbm.2016.114
Pellegrineschi A, Davolio-Mariani O (1996) Agrobacterium rhizogenes-mediated transformation of scented geranium. Plant Cell Tissue Organ Cult 47:79–86. https://doi.org/10.1007/BF02318969
Peres LEP, Morgante PG, Vecchi C, Kraus JE, Van Sluys MA (2001) Shoot regeneration capacity from roots and transgenic hairy roots of tomato cultivars and wild related species. Plant Cell Tissue Organ Cult 65:37–44. https://doi.org/10.1023/A:1010631731559
Pérez de la Torre MC, Fernández P, Greppi JA, Coviella MA, Fernández MN, Astigueta F, Mata DA, Trupkin SA (2018) Transformation of Mecardonia (Plantaginaceae) with wild-type Agrobacterium rhizogenes efficiently improves compact growth, branching and flower related ornamental traits. Sci Hortic (Amsterdam) 234:300–311. https://doi.org/10.1016/j.scienta.2018.02.047
Petersen SG, Stummann BM, Olesen P, Henningsen KW (1989) Structure and function of root-inducing (Ri) plasmids and their relation to tumor-inducing (Ti) plasmids. Physiol Plant 77:427–435. https://doi.org/10.1111/j.1399-3054.1989.tb05664.x
Petit A, David C, Dahl GA, Ellis JG, Guyon P, Casse-Delbart F, Tempé J (1983) Further extension of the opine concept: plasmids in Agrobacterium rhizogenes cooperate for opine degradation. Mol Gen Genet 190:204–214. https://doi.org/10.1007/BF00330641
Phelep M, Petit A, Martin L, Duhoux E, Tempé J (1991) Transformation and regeneration of a nitrogen-fixing tree, Allocasuarina verticillata Lam. Nat Biotechnol 9:461–466. https://doi.org/10.1038/nbt0591-461
Piatczak E, Krolicka A, Wysokinska H (2006) Genetic transformation of Centaurium erythraea Rafn by Agrobacterium rhizogenes and the production of secoiridoids. Plant Cell Rep 25:1308–1315. https://doi.org/10.1007/s00299-006-0155-0
Porter JR, Flores H (1991) Host range and implications of plant infection by Agrobacterium rhizogenes. Crit Rev Plant Sci 10:387–421. https://doi.org/10.1080/07352689109382318
Pribil M, Hermann S, Dun G, Ngo C, O’Neill S, Wang L, Bonnett G, Chandler P, Beveridge C, Lakshmanan P (2007) Altering sugarcane shoot architecture through genetic engineering: prospects for increasing cane and sugar yield. Proc Aust Soc Sugar Cane Technol 29:251–257
Rademacher W (2015) Plant growth regulators: backgrounds and uses in plant production. J Plant Growth Regul 34:845–872. https://doi.org/10.1007/s00344-015-9541-6
Rawat JM, Rawat B, Mehrotra S (2013) Plant regeneration, genetic fidelity, and active ingredient content of encapsulated hairy roots of Picrorhiza kurrooa Royle ex Benth. Biotechnol Lett 35:961–968. https://doi.org/10.1007/s10529-013-1152-3
Riker A, Banfield W, Wright W, Keitt G, Sagen H (1930) Studies on infectious hairy root of nursery apple trees. J Agric Res 41:507–540
Rugini E, Silvestri C, Cristofori V, Brunori E, Biasi R (2015) Ten years field trial observations of ri-TDNA cherry Colt rootstocks and their effect on grafted sweet cherry cv Lapins. Plant Cell Tissue Organ Cult 123:557–568. https://doi.org/10.1007/s11240-015-0860-x
Saitou T, Kamada H, Harada H (1991) Isoperoxidase in hairy roots and regenerated plants of horseradish (Armoracia lapathifolia). Plant Sci 75:195–201. https://doi.org/10.1016/0168-9452(91)90234-Y
Satheeshkumar K, Jose B, Soniya EV, Seeni S (2009) Isolation of morphovariants through plant regeneration in Agrobacterium rhizogenes induced hairy root cultures of Plumbago rosea L. Indian J Biotechnol 8:435–441
Saxena G, Banerjee S, Laiq-Ur-Rahman VPC, Mallavarapu GR, Kumar S (2007) Rose-scented geranium (Pelargonium sp.) generated by Agrobacterium rhizogenes mediated Ri-insertion for improved essential oil quality. Plant Cell Tissue Organ Cult 90:215–223. https://doi.org/10.1007/s11240-007-9261-0
Sevón N, Dräger B, Hiltunen R, Oksman-Caldentey KM (1997) Characterization of transgenic plants derived from hairy roots of Hyoscyamus muticus. Plant Cell Rep 16:605–611. https://doi.org/10.1007/s002990050287
Shanks J, Morgan J (1999) Plant “hairy root” culture. Curr Opin Biotechnol 10:151–155. https://doi.org/10.1016/S0958-1669(99)80026-3
Sinkar VP, White FF, Gordon MP (1987) Molecular biology of Ri-plasmid – a review. J Biosci 11:47–57. https://doi.org/10.1007/BF02704657
Skała E, Picot L, Bijak M, Saluk-Bijak J, Szemraj J, Kicel A, Olszewska MA, Sitarek P (2019) An efficient plant regeneration from Rhaponticum carthamoides transformed roots, enhanced caffeoylquinic acid derivatives production in pRi-transformed plants and their biological activity. Ind Crop Prod 129:327–338. https://doi.org/10.1016/j.indcrop.2018.12.020
Slightom JL, Durand-Tardif M, Jouanin L, Tepfer D (1986) Nucleotide sequence analysis of TL-DNA of Agrobacterium rhizogenes agropine type plasmid. Identification of open reading frames. J Biol Chem 261:108–121
Spanò L, Mariotti D, Pezzotti M, Damiani F, Arcioni S (1987) Hairy root transformation in alfalfa (Medicago sativa L.). Theor Appl Genet 73:523–530. https://doi.org/10.1007/BF00289189
Spena A, Schmülling T, Koncz C, Schell JS, Kayani WK (1987) Independent and synergistic activity of rol A, B and C loci in stimulating abnormal growth in plants. EMBO J 6:3891–3899
Sridevi G, Parameswari C, Rajamuni P, Veluthambi K (2006) Identification of hemizygous and homozygous transgenic rice plants in T1 generation by DNA blot analysis. Plant Biotechnol 23:531–534. https://doi.org/10.5511/plantbiotechnology.23.531
Suginuma C, Akihama T (1995) Transformation of gentian with Agrobacterium rhizogenes. Acta Hortic. https://doi.org/10.17660/ActaHortic.1995.392.18
Sun LY, Touraud G, Charbonnier C, Tepfer D (1991) Modification of phenotype in Belgian endive (Cichorium intybus) through genetic transformation by Agrobacterium rhizogenes: conversion from biennial to annual flowering. Transgenic Res 1:14–22. https://doi.org/10.1007/BF02512992
Suzuki K, Yamashita I, Tanaka N (2002) Tobacco plants were transformed by Agrobacterium rhizogenes infection during their evolution. Plant J 32:775–787. https://doi.org/10.1046/j.1365-313X.2002.01468.x
Suzuki K, Tanaka K, Yamamoto S, Kiyokawa K, Moriguchi K, Yoshida K (2009) Ti and Ri Plasmids. In: Schwartz E (ed) Microbiol Megaplasmids. Springer Berlin Heidelberg, Berlin, pp 133–147
Tanaka N, Matsumoto T (1993) Regenerants from Ajuga hairy roots with high productivity of 20-hydroxyecdysone. Plant Cell Rep 13:87–90. https://doi.org/10.1007/BF00235296
Tao R, Handa T, Tamura M, Sugiura A (1994) Genetic transformation of Japanese persimmon (Diospyros kaki L.) by Agrobacterium rhizogenes wild type strain A4. J Japanese Soc Hortic Sci 63:283–289
Taylor BH, Amasino RM, White FF, Nester EW, Gordon MP (1985) T-DNA analysis of plants regenerated from hairy root tumors. Mol Gen Genet 201:554–557. https://doi.org/10.1007/BF00331355
Tempé J, Schell J (1977) Is crown-gall a natural instance of gene transfer? In: Legocki A (ed) Translation of natural and synthetic polynucleotides. Poznan Agricultural University, Poznan, pp 415–420
Tepfer D (1984) Transformation of several species of higher plants by Agrobacterium rhizogenes: sexual transmission of the transformed genotype and phenotype. Cell 37:959–967. https://doi.org/10.1016/0092-8674(84)90430-6
Tepfer D (2016) DNA transfer to plants by Agrobacterium rhizogenes: a model for genetic communication between species and biospheres. In: Jha S (ed) Transgenesis and secondary metabolism. Springer International Publishing, Cham, pp 1–41
Trovato M, Maras B, Linhares F, Costantino P (2001) The plant oncogene rolD encodes a functional ornithine cyclodeaminase. Proc Natl Acad Sci USA 98:13449–13453. https://doi.org/10.1073/pnas.231320398
Tsuro M, Ikedo H (2011) Changes in morphological phenotypes and essential oil components in lavandin (Lavandula×intermedia Emeric ex Loisel.) transformed with wild-type strains of Agrobacterium rhizogenes. Sci Hortic (Amsterdam) 130:647–652. https://doi.org/10.1016/j.scienta.2011.08.011
Van Sluys MA, Tempé J (1989) Behavior of the maize transposable element activator in Daucus carota. Mol Gen Genet 219:313–319. https://doi.org/10.1007/BF00261193
Verma P, Khan SA, Masood N, Manika N, Sharma A, Verma N, Luqman S, Mathur AK (2017) Differential rubisco content and photosynthetic efficiency of rol gene integrated Vinca minor transgenic plant: correlating factors associated with morpho-anatomical changes, gene expression and alkaloid productivity. J Plant Physiol 219:12–21. https://doi.org/10.1016/j.jplph.2017.09.004
Vishwakarma KS, Mohammed SI, Chaudhari AR, Salunkhe NS, Maheshwari VL (2017) Micropropagation and Agrobacterium rhizogenes mediated transformation studies in Mucuna pruriens (L.) DC. Indian J Nat Prod Resour 8:172–178
Vladimirov IA, Matveeva TV, Lutova LA (2015) Opine biosynthesis and catabolism genes of Agrobacterium tumefaciens and Agrobacterium rhizogenes. Russ J Genet 51:121–129. https://doi.org/10.1134/S1022795415020167
Wang YM, Wang JB, Luo D, Jia JF (2001) Regeneration of plants from callus tissues of hairy roots induced by Agrobacterium rhizogenes on Alhagi pseudoalhagi. Cell Res 11:279–284. https://doi.org/10.1038/sj.cr.7290097
Watase I, Sudo H, Yamazaki M, Saito K (2004) Regeneration of transformed Ophiorrhiza pumila plants producing camptothecin. Plant Biotechnol 21:337–342. https://doi.org/10.5511/plantbiotechnology.21.337
Webb K, Jones S, Robbins M, Minchin F (1990) Characterization of transgenic root cultures of Trifolium repens, Trifolium pratense and Lotus corniculatus and transgenic plants of Lotus corniculatus. Plant Sci 70:243–254. https://doi.org/10.1016/0168-9452(90)90139-F
Weber RLM, Bodanese-Zanettini MH (2011) Induction of transgenic hairy roots in soybean genotypes by Agrobacterium rhizogenes-mediated transformation. Pesqui Agropecu Bras 46:1070–1075
White FF, Nester EW (1980a) Relationship of plasmids responsible for hairy root and crown gall tumorigenicity. J Bacteriol 144:710–720
White FF, Nester EW (1980b) Hairy root: plasmid encodes virulence traits in Agrobacterium rhizogenes. J Bacteriol 141:1134–1141
White FF, Ghidossi G, Gordon MP, Nester EW (1982) Tumor induction by Agrobacterium rhizogenes involves the transfer of plasmid DNA to the plant genome. Proc Natl Acad Sci U S A 79:3193–3197. https://doi.org/10.1073/pnas.79.10.3193
White FF, Garfinkel DJ, Huffman GA, Gordon MP, Nester EW (1983) Sequences homologous to Agrobacterium rhizogenes T-DNA in the genomes of uninfected plants. Nature 301:348–350. https://doi.org/10.1038/301348a0
White FF, Taylor BH, Huffman GA, Gordon MP, Nester EW (1985) Molecular and genetic analysis of the transferred DNA regions of the root-inducing plasmid of Agrobacterium rhizogenes. J Bacteriol 164:33–44
Yamakawa Y, Chen L-H (1996) Agrobacterium rhizogenes mediated transformation of kiwifruit (Actinidia deliciosa) by direct formation of adventitious buds. J Japanese Soc Hortic Sci 64:741–747. https://doi.org/10.2503/jjshs.64.741
Yamashita H, Daimon H, Akasaka-Kennedy Y, Masuda T (2004) Plant regeneration from hairy roots of apple rootstock, Malus prunifolia Borkh. var. ringo Asami, strain Nagano No. 1, transformed by Agrobacterium rhizogenes. J Japanese Soc Hortic Sci 73:505–510. https://doi.org/10.2503/jjshs.73.505
Yang D, Choi Y (2000) Production of transgenic plants via Agrobacterium rhizogenes- mediated transformation of Panax ginseng. Plant Cell Rep 19:491–496. https://doi.org/10.1007/s002990050761
Yazawa M, Suginuma C, Ichikawa K, Kamada H, Akihama T (1995) Regeneration of transgenic plants from hairy root of kiwi fruit (Actinidia deliciosa) induced by Agrobacterium rhizogenes. Japanese J Breed 45:241–244. https://doi.org/10.1270/jsbbs1951.45.241
Yin Z, Pląder W, Malepszy S (2004) Transgene inheritance in plants. J Appl Genet 45:127–144
Yoshimatsu K, Shimomura K (1992) Transformation of opium poppy (Papaver somniferum L.) with Agrobacterium rhizogenes MAFF03-01724. Plant Cell Rep 11:132–136. https://doi.org/10.1007/BF00232165
Yoshimatsu K, Shimomura K, Yamazaki M, Saito K, Kiuchi F (2003) Transformation of Ipecac (Cephaelis ipecacuanha) with Agrobacterium rhizogenes. Planta Med 69:1018–1023. https://doi.org/10.1055/s-2003-45149
Zhan XC, Jones DA, Kerr A (1988) Regeneration of flax plants transformed by Agrobacterium rhizogenes. Plant Mol Biol 11:551–559. https://doi.org/10.1007/BF00017455
Zhang GN, Jia JF, Hao JG, Wang XR, He T (2008) Plant regeneration from mesophyll protoplasts of Agrobacterium rhizogenes-transformed Astragalus melilotoides. Biol Plant 52:373–376. https://doi.org/10.1007/s10535-008-0078-4
Zhou YQ, Duan HY, Zhou CE, Li JJ, Gu FP, Wang F, Zhang ZY, Gao ZM (2009) Hairy root induction and plant regeneration of Rehmannia glutinosa Libosch. f. hueichingensis Hsiao via Agrobacterium rhizogenes-mediated transformation. Russ J Plant Physiol 56:224–231. https://doi.org/10.1134/S1021443709020113
Funding
This study was funded by VLAIO (LA grant number 150889).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Desmet, S., Dhooghe, E., De Keyser, E. et al. Rhizogenic agrobacteria as an innovative tool for plant breeding: current achievements and limitations. Appl Microbiol Biotechnol 104, 2435–2451 (2020). https://doi.org/10.1007/s00253-020-10403-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-020-10403-7