Abstract
Key message
Development of wheat- D. villosum 1V#4 translocation lines; physically mapping the Glu - V1 and Gli - V1 / Glu - V3 loci; and assess the effects of the introduced Glu - V1 and Gli - V1 / Glu - V3 on wheat bread-making quality.
Abstract
Glu-V1 and Gli-V1/Glu-V3 loci, located in the chromosome 1V of Dasypyrum villosum, were proved to have positive effects on grain quality. However, there are very few reports about the transfer of the D. villosum-derived seed storage protein genes into wheat background by chromosome manipulation. In the present study, a total of six CS-1V#4 introgression lines with different alien-fragment sizes were developed through ionizing radiation of the mature female gametes of CS––D. villosum 1V#4 disomic addition line and confirmed by cytogenetic analysis. Genomic in situ hybridization (GISH), chromosome C-banding, twelve 1V#4-specific EST–STS markers and seed storage protein analysis enabled the cytological physical mapping of Glu-V1 and Gli-V1/Glu-V3 loci to the region of FL 0.50–1.00 of 1V#4S of D. villosum. The Glu-V1 allele of D. villosum was Glu-V1a and its coded protein was V71 subunit. Quality analysis indicated that Glu-V1a together with Gli-V1/Glu-V3 loci showed a positive effect on protein content, Zeleny sedimentation value and the rheological characteristics of wheat flour dough. In addition, the positive effect could be maintained when specific Glu-V1 and Gli-V1/Glu-V3 loci were transferred to the wheat genetic background as in the case of T1V#4S-6BS·6BL, T1V#4S·1BL and T1V#4S·1DS translocation lines. These results showed that the chromosome segment carrying the Glu-V1 and Gli-V1/Glu-V3 loci in 1V#4S of D. villosum had positive effect on bread-making quality, and the T1V#4S-6BS·6BL and T1V#4S·1BL translocation lines could be useful germplasms for bread wheat improvement. The developed 1V#4S-specific molecular markers could be used to rapidly identify and trace the alien chromatin of 1V#4S in wheat background.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The end-product quality of common wheat (Triticum aestivum L.) is mainly determined by its seed storage proteins (SSPs), including glutenins and gliadins. Gliadins are monomeric and play important roles in determining extensibility properties of gluten dough, whereas glutenins are polymeric, consisting of high-molecular-weight (HMW) and low-molecular-weight (LMW) subunits, which can form complex polymeric proteins by inter-chain disulphide bonds and impart dough viscoelasticity (Payne et al. 1980, 1987; Shewry and Tatham 1990; Shewry et al. 2003). In bread wheat, glutenins and gliadins are encoded by loci located in homoeologous groups 1 and 6 chromosomes (McIntosh et al. 2008). The Glu-1 loci for high-molecular-weight glutenin subunits (HMW-GSs) are in the long arms of group 1 chromosomes; while the Gli-1 locus for gliadins and Glu-3 for low-molecular-weight glutenin subunits (LMW-GSs) are in the short arms. HMW-GSs explained up to 70 % of the variation in bread-making performance among European wheat cultivars (Branlard and Dardevet 1985; Payne et al. 1987, 1988), although they only account for approximately 12 % of the total protein in the endosperm of common wheat (Halford et al. 1992). Even though the HMW-GSs are such important determinants for processing quality, the number of subunits associated with good quality remains rather limited (Garg et al. 2009b). Screening for positive effect HMW-GS in wheat-related species is therefore important for improving bread-making quality. By using PCR-based methods, some HMW-GS alleles from wheat-related species in tribe Triticeae have been characterized (Mackie et al. 1996; Wan et al. 2002; Cao et al. 2007; Liu et al. 2008, 2012; Garg et al. 2009a; Niu et al. 2011; Li et al. 2013). But only a few reported the successful transfer of alien chromatin segment carrying the SSP genes into wheat background by chromosome manipulation (Li et al. 2013).
Dasypyrum villosum (L.) Candargy (Dv), also known as Haynaldia villosa (L.) Schur, which is native to the northeastern part of the Mediterranean regions and southwest Asia, is an annual diploid wild relative (2n = 2x = 14, genome VV) of common wheat (Agnieszka 2006). In addition to having good resistances to several fungal pathogens that cause severe diseases in wheat (Chen et al. 1995; Zhang et al. 2005; Qi et al. 2011), V-genome also contains genes that can increase the amount of SSP and gluten strength (Shewry et al. 1987, 1991; De Pace et al. 2001; Zhao et al. 2010; Vaccino et al. 2010; Dong et al. 2013). It has been proved that the wheat-D. villosum alien chromosome lines involving chromosomes 1V, 4V or 6V had higher protein contents than wheat, but only those lines having chromosome 1V, which carry the complex loci coding for HMW-GS (Glu-V1), LMW polymeric prolamin proteins (Glu-V3), sulfur-poor (ω-type) and sulfur-rich (γ-type) monomeric prolamins (Gli-V1) have improved grain quality (De Pace et al. 2001; Gradzielewska 2006; Zhao et al. 2010; Vaccino et al. 2010). Zhong and Qualset (1993) identified 14 alleles for Glu-V1 coding for null, single and two HMW protein subunits. As a result, different homoeologous chromosome 1V could differ not only for the allele at the Glu-V1 locus, but also for the alleles at the Gli-V1/Glu-V3 loci. Thus, D. villosum is also an important resource for the improvement of wheat grain quality.
The development of wheat-D. villosum alien introgression lines involving 1V, with Glu-V1 and Gli-V1/Glu-V3 loci carrying chromosome fragments transferred into wheat background may provide useful genetic resources for quality improvement. The objectives of this study are to develop wheat-D. villosum alien translocation lines involving 1V, map the Glu-V1 and Gli-V1/Glu-V3 loci physically to specific chromosome region, and assess the effects of the introduced Glu-V1 and Gli-V1/Glu-V3 on wheat quality. Our results will extend the current understandings of the chromosome location of the HMW-GS gene family and be regarded as a prerequisite for the utilization of Glu-V1 and Gli-V1/Glu-V3 loci in wheat breeding for the improvement of the end-use quality.
Materials and methods
Plant materials
The plant materials used in this study included T. aestivum cv. Chinese Spring (CS), D. villosum parental line (accession no. 91C43, the donor of Glu-V1 and Gli-V1/Glu-V3 loci) T. durum cv. (accession no. 1286), T. durum cv. (1286)-D. villosum (91C43) amphiploid (AABBVV), a complete set of wheat-D. villosum disomic addition lines (The V chromosomes of this stock were numbered #4 by De Pace et al. 2011) in the CS background (Zhang et al. 2013) and two telosomic addition lines 1V#4L and 1V#4S. These cytogenetic materials were all developed and maintained at the Cytogenetic Institute, Nanjing Agricultural University (CINAU), China. Other genetic stocks including three nulli-tetrasomic (N1AT1D, N1BT1D, N1DT1B) and six ditelosomic (Dt, Dt1AS, Dt1AL, Dt1BS, Dt1BL, Dt1DS and Dt1DL) stocks, which were kindly provided by the Wheat Genetics and Genomics Resource Centre, Kansas State University, Manhattan, KS, USA, were also used in this study.
Irradiation of female gametes
The adult plants, with the mature female gametes of the DA1V#4 were irradiated by 60CO-γ ray at the dosages of 1,200 Rad with the dose rate of 160 Rad/min before flowering. The irradiated spikes were emasculated and covered with paper bags, the florets were pollinated with normal fresh matured pollens of CS 2–3 days after emasculation.
Cytogenetic analysis
The procedures for chromosome C-banding were according to Gill et al. (1991) and that for GISH followed Chen et al. (1995). To characterize the translocated chromosomes, dual-color fluorescence in situ hybridization (FISH) was employed using repetitive DNA clones, i.e., pSc119.2, pAs1, 45S rDNA and total genomic DNA of D. villosum as probes. Dual-color FISH can therefore not only differentiate the wheat and D. villosum chromatin, but also determine the identities of particular wheat chromosomes. The genomic DNA of D. villosum was labeled with fluorescein-12-dUTP, and used to detect the chromosome fragment of 1V#4. The clone pAs1 from Aegilops tauschii Coss. was used to detect most of the D-genome chromosomes, and the clone pSc119.2 could identify the B-genome chromosomes (Mukai et al. 1993). The 45S rDNA probe (pTa71) was used to detect the chromosomal nucleolus organizing region (Nor), and the Nor-B2 (6BS) and Nor-B1 (1BS) loci of common wheat (Mukai et al. 1991) and Nor-V1 (1VS) loci of D. villosum (Zhang et al. 2013) can be visualized. The clones of pSc119.2, pAs1 and pTa71 were all labeled with digoxigenin-11-dUTP (Roche Diagnostics GmbH, Germany). After hybridization with the probes, chromosomes were treated with FITC-avidin antibody, stained with DAPI and visualized under an Olympus BX60 fluorescence microscope and photographed with a SPOT Cooled Color Digital Camera.
Molecular markers analysis
For the development of the 1V#4 arm-specific EST–PCR markers, a total of 250 EST-based primers from different regions of each arm of the wheat group 1 chromosome were used to screen the polymorphism among CS, T. durum cv. 1286, T. durum cv. 1286-D. villosum amphiploid (AABBVV), a set of wheat-D. villosum disomic addition lines (DA1V#4–7V#4) and telosomic addition lines 1V#4L and 1V#4S. The primers were designed using the ConservedPrimers 2.0 software (Frank et al. 2009). Five previously reported 1V#4 arm-specific EST–PCR markers (CINAU23, CINAU24, CINAU25, CINAU26 and CINAU27) were also used in this study (Wang et al. 2007) (Table 1). Genomic DNA was isolated from young leaves following the procedure of Doyle and Doyle (1990). PCR amplifications were conducted in a 25-μL reaction mixture containing 1× Taq DNA polymerase buffer, 0.8 mmol/L MgCl2, 0.8 mmol/L dNTPs, 200 μmol/L primers, 2 units DNA polymerase and 50-ng genomic DNA as template. The samples were denaturated at 94 °C for 5 min and subjected to 34 cycles of 30 s of denaturation at 94 °C, 45 s annealing at T m and 2.0-min elongation at 72 °C. A final cycle with an extension of 10 min at 72 °C was applied to complete the reactions. The PCR products were analyzed on 8 % polyacrylamide gels in 1× TBE buffer.
Protein extraction from the grain and SDS-PAGE
The composition of glutenin subunit (HMW-GS and LMW-GS) in the half-seed endosperm (the other half with the embryo was used to germinate for collecting the seed–roots for cytogenetic analysis) was determined by SDS-PAGE according to Garg et al. (2009a). A 10-μL sample was loaded onto an SDS-PAGE gel, formed by a 12.5 % gradient separating gel (pH 8.5) and a 4.0 % stacking gel (pH 6.8). Gliadins were extracted by 50 % v/v isopropanol and separated by 17.5 % SDS-PAGE. Separation was carried out at a constant current of 6 mA for 14–16 h. The gel was stained by gently shaking in 0.25 % w/v Coomassie Brilliant Blue R250, 12.5 % trichloroacetic acid for 10 h and then rinsed with distilled water.
Evaluation of grain quality
All the genetic stocks were grown in the greenhouse at Jiangpu Experimental Station of Nanjing Agriculture University, China, during 2010–2011 and 2011–2012 in a randomized plot design. Each plot with three replications had six 1.2-m rows with 0.25 m between-row space and 0.1 m between plants within row space. Field management followed local practices.
Grain samples were harvested and cleaned prior to conditioning and milling. Grain protein content (14 % moisture) was determined by near-infrared spectroscopy (NIRS) analysis using an Instalab 610 instrument (Newport Scientific Scales and Services Pty Ltd). Zeleny sedimentation tests were performed according to AACC method 56–61A. Dough rheological parameters, dough development time (DT) and dough stability time (ST) were measured by Farinograph 810104 (Brabender, Germany) according to AACC method 54–21 (1987). Analysis of variance was performed on original data for grain quality values using the SAS 8.2 system.
Results
Development of the wheat-D. villosum introgression lines involving chromosome 1V#4
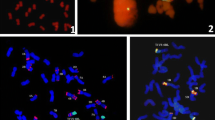
Six CS-1V#4 introgression lines with different alien-fragment sizes were identified from 146 M1 plants by GISH and C-banding analyses. These plants were backcrossed using CS as the recurrent parent and then self-fertilized. Homozygous translocation lines were identified in the BC2F2 population and characterized by GISH on chromosome preparations at metaphases of both mitosis of the root tip cells and meiosis of the pollen mother cells (Fig. 1). Among them, NAU1V-1 (2n = 44) was a 1V#4S deletion line with the breakpoint at FL0.50 (Fig. 1a). NAU1V-2 (2n = 42) was a large-fragment translocation, involving 1V#4 (Fig. 1b), and NAU1V-3 (2n = 42) was a small alien-fragment terminal translocation (Fig. 1c), in which 42 chromosomes paired as 21 bivalents at meiotic metaphase I and the translocation chromosomes formed a ring bivalent revealed by GISH (Fig. 1d), suggesting that NAU1V-3 was a homozygous translocation. Three lines, NAU1V-4, NAU1V-5 and NAU1V-6, all had chromosome numbers as 2n = 42 and were homozygous whole-arm translocations, having a pair of 1V#4S·W or 1V#4L·W, respectively (Fig. 1e–j).
Mitotic and meiotic GISH analysis of the wheat-1V#4 introgression lines. GISH using D. villosum total genomic DNA labeled with digoxigenin-11-dUTP as probe (the D. villosum chromatin fluoresced yellow-greenish and the wheat chromatin fluoresced red). a Somatic metaphase of line NAU1V-1(2n = 44), which contained 42 wheat chromosomes and a pair of 1V#4 deletion chromosomes (yellowish-green color); b GISH patterns of NAU1V-2 line (2n = 42) with a pair of wheat-1V#4 large-fragment translocation chromosomes; c GISH patterns of NAU1V-3 (2n = 42), which included 40 wheat chromosomes and one pair of wheat-1V#4 small-fragment terminally translocated chromosomes; d PMCs at meiotic metaphase I of NAU1V-3, where translocated chromosomes formed a ring bivalent. e, f Mitotic and meiotic GISH patterns of NAU1V-4 (2n = 42) with 21 II (ring), which included a pair of wheat-1V#4 whole-arm translocated chromosomes. g, h Mitotic and meiotic GISH patterns of NAU1V-5 (2n = 42) with 21 II containing a pair of wheat-1V#4 whole-arm translocated chromosomes. i, j Mitotic and meiotic GISH patterns of NAU1V-6 (2n = 42) formed 21 II with a pair of wheat-1V#4 whole-arm translocated chromosomes (color figure online)
Characterization of the wheat-D. villosum 1V#4 translocation chromosomes by GISH/FISH
Zhang et al. (2013) observed that 45S rDNA loci were located to the terminal of 1VS (Nor-V1) in D. villosum. By the dual color of GISH/FISH using total genomic DNA of D. villosum and 45S rDNA as probes, we found that the Nor-V1 loci were absent in NAU1V-1, NAU1V-2 and NAU1V-6, which implied that the distal region of 1VS#4 was all missing in these lines. Whereas, the Nor-V1 loci were present in lines NAU1V-3, NAU1V-4 and NAU1V-5 (Fig. 2). In addition, the identities of wheat chromosomes involved in these translocation chromosomes were characterized by sequential chromosome C-banding and FISH. Referring to the standard C-banding pattern (Gill et al. 1991) and FISH pattern for pSc119.2 and pAs1 (Mukai et al. 1993) established for wheat variety CS, it was found that the wheat chromosomes involved in the translocation with 1V#4 in NAU1V-3, NAU1V-4, NAU1V-5 and NAU1V-6 were 6B, 1B and 1D (NAU1V-5 and NAU1V-6), respectively (Fig. 2). Hence, the NAU1V-3 was a small terminal translocation and designated T1VS#4-6BS·6BL (Fig. 2), while the NAU1V-4, NAU1V-5 and NAU1V-6 were whole-arm translocations and designated T1VS#4·1BL, T1VS#4·1DS and T1VL#4·1DL, respectively (Fig. 2).
C-banding and FISH patterns of wheat-1V#4 translocated chromosomes. Dual-color FISH using a pSc119.2, pAs1 or 45S labeled with digoxigenin-11-dUTP (red) and total genomic DNA of D. villosum labeled with fluorescein-12-dUTP (green) as probes. The 45S rDNA signals are near the tip of the short arm of chromosome 1V, while secondary constriction of Nor-V1 in wheat-1V#4 introgression lines could not be observed, demonstrating that the expression of Nor-V1 could be suppressed by the Nors of wheat in these lines (color figure online)
Characterization of the wheat-D. villosum 1V#4 translocations by molecular marker analysis
Seven out of the 250 EST–PCR primer pairs (2.8 %) amplified polymorphism products between CS and the wheat-D. villosum disomic addition line DA1V#4. By amplification in two telosomic addition lines 1VL#4 and 1VS#4, four were assigned to the short arm (1VS#4) and three to the long arm (1VL#4) (Table 1). Further amplification of the above seven markers in the nulli-/tetrasomic and ditelosomic stocks of homoeologous group 1 chromosomes of CS showed that all were located in the corresponding homoeologous wheat group 1 chromosome arms of 1V#4 (part of the amplification results were shown in Fig. 3). Together with the five previously reported 1V#4 specific markers, a total of 12 makers were used to detect the alien segments present in the six wheat-D. villosum 1V#4 translocation lines (Table 2). The results showed that in NAU1V-1 and NAU1V-2, all the diagnostic bands of seven 1VL#4- and two 1VS#4-specific molecular markers were present. However, the diagnostic bands of the remaining three 1V#4S-specific markers (CINAU27, 1EST1112 and 1EST1118) were absent (Table 2), further confirming these two lines did not have the terminal fragment of 1V#4S. In NAU1V-3, only three of the five 1V#4S-specific molecular markers could amplify the diagnostic bands (Table 2), confirmed that NAU1V-3 was a small alien-fragment translocation involving the terminal fragment of 1V#4S. In NAU1V-4 and NAU1V-5, all the five 1V#4S-specific markers were present, whereas the 1BS fragment or the 1DL fragment was absent (Fig. 3), confirming that NAU1V-4 was a T1VS#4·1BL translocation and NAU1V-5 was a T1VS#4·1DS translocation. In NAU1V-6, the 1VL#4-specific bands of CINAU23, CINAU24, CINAU25, CINAU26, 1EST235, 1EST425 and 1EST493 were present, but 1DS-specific bands were absent (Fig. 3), supporting that NAU1V-6 was a T1VL#4·1DL translocation, which was also consistent with the cytogenetic results.
Electrophoresis patterns of 1V#4-specific EST–PCR markers. The straight line indicates the fragment that can be assigned to a certain chromosome. The 1V#4L-specific fragments amplified by the three EST primers (1EST235, 1EST425 and 1EST493) were all present in NAU1V-6, and 1VS-specific fragments amplified by another three EST primers (1EST1112, 1EST1118 and 1EST1150) were all present in NAU1V-4 and NAU1V-5. The 1BS fragments were absent in N1BT1D and NAU1V-4. The 1DL fragments were absent in N1DT1B and NAU1V-5, while 1DS fragments were absent in NAU1V-6
Seed storage protein constitutions of the wheat-D. villosum 1V#4 translocations by SDS-PAGE
SDS-PAGE analysis indicated that the D. villosum and DA1V#4 had one extra HMW-GS, which was a fast-moving HMW-GS and below the 1Bx7 of CS (Fig. 4a). According to the designation system for the Glu-V1 alleles proposed by Zhong and Qualset (1993), based on the SDS-PAGE migration distance of D. villosum HMW-GS relative to those (7 + 8 subunits) of CS, the Glu-V1 allele of D. villosum in the current study was Glu-V1a and its coded protein was the V71 subunit. This subunit was also present in lines NAU1V-3, NAU1V-4 and NAU1V-5, but absent in NAU1V-1 and NAU1V-6 (Fig. 4b, c). Comparing the SDS-PAGE patterns of DA1V#4 and NAU1V-1, the Glu-V1a locus was cytologically mapped to the distal region of 1VS, immersed in the bin of FL 0.50–1.00. It was found that the 2 + 12 subunits located in the 1DL was absent in line NAU1V-5 (Fig. 4b), confirming NAU1V-5 was a T1VS#4·1DS translocation line. In NAU1V-3 and DA1V#4 there were two additional intensive subunits (indicated by arrows in Fig. 4c), which were absent in NAU1V-1 in the region of wheat LMW-GS. The result suggested that the additional LMW-GSs encoded by the orthologous Glu-V3 locus were cytologically mapped to the distal region of 1VS. Similarly, the additional gliadin subunits (showed by arrows in Fig. 4d) encoded by the orthologous Gli-V1 locus were also cytologically mapped to the distal region of 1VS, immersed in the bin of FL 0.50–1.00.
Seed storage protein of the wheat-1V#4 introgression lines. a SDS-PAGE of CS-DA1V#4–7V#4 addition lines. SDS-PAGE showing a additional HMW-GS (V71) of D. villosum in the homoeologous group 1 addition line (DA 1V#4), the mobility of the subunit V71 coded by Glu-V1a is greater than that of subunit 7 of CS. b SDS-PAGE of the whole-arm translocation lines. Glu-D1-encoded bands 1Dx2 and 1Dy12 bands in NAU1V-5 are lost. c, d Sequential SDS-PAGE separation of glutenins(c) and gliadins (d) from the same kernels. Glu-V3 and Gli-V1 locus-specific subunits of D. villosum indicated by arrowheads and arrows, respectively, are all present in DA1V#4 and NAU1V-3 lines but absent in 1V#4-deletion line NAU1V-1
Grain quality of the wheat-D. villosum 1V#4 translocations
The results of grain quality of the chromosome 1V#4 variations were shown in Table 3. In the 2 years, the seed protein concentration (SPC) and the values of Zeleny sedimentation value (ZSV), dough development time (DT) and dough stability time (ST) of DA1V#4 were all significantly higher than those of CS and NAU1V-1. In addition, the bread-making quality characteristics of the three translocation lines NAU1V-3, NAU1V-4 and NAU1V-5 were all better than those of their recipient parent CS. These indicated that the introduction of the Glu-V1a, Glu-V3 and Gli-V1 loci in 1V#4S into wheat led to improved bread-making quality. By comparison, the values of ZSV, DT and ST of T1V#4S·1DS translocation line NAU1V-5 were lower than those of T1V#4S-6BS·6BL translocation line NAU1V-3 and T1V#4S·1BL translocation line NAU1V-4. The differences for SPC and gluten strength among different translocation lines carrying the same Glu-V1a, Glu-V3 and Gli-V1 loci of D. villosum showed the wheat chromosome involving in these translocated chromosomes may have effects on grain quality.
Discussion
The use of alien genetic resources from a wide collective of exotic Triticeae species is important for the genetic improvement of agronomic traits, including the end-use quality of wheat. Ionizing radiation using the wheat-alien species amphiploid, addition, substitution and whole-arm translocation lines as initial materials is the most preferred choice for the induction of alien chromosome translocation, which is the best carrier for the useful alien genes (Sears and Gustafson 1993; Chen et al. 1995, 2005, 2008; Friebe et al. 1996; Zhang et al. 2010, 2012). In the present study, by 60CO-γ ray irradiation of the adult plants of the disomic addition line (DA1V#4), various chromosome structural aberrants including deletion, large alien-fragment translocation, small alien-fragment translocation and whole-arm translocation were achieved (Fig. 1; Table 2). Cytological studies indicated that no abnormal behavior was observed in the meiotic process of pollen mother cells of the translocation chromosomes in NAU1V-3 and NAU1V-4. The major agronomic characters of T1V#4S-6BS·6BL (NAU1VS-3) and T1V#4S·1BL (NAU1V-4) translocation lines, such as plant height and flowering time, were similar as the recipient variety CS (date was not shown), showing that the translocation chromosomes, T1V#4S-6BS·6BL and T1V#4S·1BL, had no obvious influence on agronomic performances. We proposed that this was due to the good genetic compensation of 6A/6D or 1VS/1AS/1DS homologous genes to those genes located on the terminal of 6BS and 1BS, which were lost in T1V#4S-6BS·6BL and T1V#4S·1BL translocation line, respectively. It could be concluded that T1V#4S-6BS·6BL small alien translocation line and T1V#4S·1BL compensating translocation line carrying the Glu-V1, Glu-V3 and Gli-V1 loci of D. villosum could be used to improve the quality of bread wheat. While, some of the sterile spikelets were observed in T1V#4S·1DS translocation line (NAU1V-5), leading to a decreasing seeds per spike. This may be due to the genetic disequilibrium when chromosome 1DL was substituted by chromosome 1VS#4S. Therefore, development of bread wheat cultivars using NAU1V-5 as the donor of the Glu-V1, Glu-V3 and Gli-V1 loci may have a negative effect on the yield.
In genus Triticum and Aegilops, genetic studies confirmed that the genes controlling the HMW-GS were located on the long arms of the group 1 chromosomes; all ω-gliadins, most of the γ-gliadins and a few of β-gliadins were encoded by the Gli-1 loci on the short arms of homoeologous chromosome 1, which was tightly linked to the Glu-3 loci coding for LMW-GS (McIntosh et al. 2008). Chromosomes of 1V of D. villosum and 1A, 1B, 1D of wheat are homeologous chromosomes. Theoretically, the prolamin genes on these chromosomes should be synteny; that is to say, Glu-V1 and Glu-V3/Gli-V1 loci should be located on 1VL and 1VS, respectively. Vaccino et al. (2010) confirmed that the Glu-V3/Gli-V1 loci were synteny to wheat’s Glu-B3/Gli-B1 loci because they were equal crossing-over in CS × V63 lines. They also argued that the Glu-V1 loci was located on chromosome 1VL based on the HMW-GS band of D. villosum carrying in the line 09.CS 1B-1V (T1BL-1V#1L·1V#1S). Zhao et al. (2010) developed the Robertsonian translocation lines T1V#3L·1DS and T1V#3S·1DL (this V chromosomes of this stock were numbered #2 by De Pace et al. 2011). Recently, Dong et al. (2013) analyzed their HMW-GS components and confirmed that the HMW-GS gene of D. villosum was located on both 1V#3L and 1V#3S, and the gene expression level might be higher in T1V#3S·1DL than in T1V#3L·1DS. In the present study, the six developed CS-1V#4 introgression lines and two telosomic addition lines 1V#4L and 1V#4S were confirmed by molecular cytogenetics analysis. Dual-color FISH, chromosome C-banding, 12 1V#4-specific EST–STS markers and SSP analysis enabled the cytological physical mapping of Glu-V1 and Gli-V1/Glu-V3 loci to the region of FL 0.50–1.00 of 1V#4S of D. villosum. The chromosome location of Gli-V1/Glu-V3 loci was consistent with the result of Vaccino et al. (2010), and the Glu-V3/Gli-V1 loci of 1V#4S could be homoeologous to Glu-3/Gli-1 loci of common wheat. However, the finding of a HMW-GS gene located on the short arm of homoeologous chromosome 1 in tribe Triticeae breaks the traditional understandings. The different results of Glu-V1 locus’ chromosome location in Vaccino et al. (2010), Dong et al. (2013) and the present study may be due to the polymorphism of the Glu-V1 locus in different accessions of D. villosum. It is possible that the HMW-GS genes on 1V#4S (this study) and 1V#3S (Dong et al. 2013) in D. villosum had independent origins due to independent translocations of ancestral gene from the long arm of chromosome 1. The degenerate primers, P1: 5-ATGGCTAAGCGGc/tTa/gGTCCTCTTTG and P2: 5-CTATCACTGGCTa/gGCCGACAATGCG, which were previously used to isolate the homologous HMW-GS genes from wheat and its relative species (Xia et al. 2003; Sun et al. 2006; Liu et al. 2008; Garg et al. 2009a; Gao et al. 2010; Li et al. 2013) were used to isolate the homologous genes from D. villosum in the present study. PCR amplification employed a high fidelity LA Taq polymerase (TaKaRa Biotechnology) with a GC buffer provided for GC-rich template. However, we were not able to find any amplification product in the D. villosum accession 91C43, possibly due to the specific structure of HMW-GS gene sequence in D. villosum. Therefore, the wheat-1V#4S translocation lines carrying the Glu-V1 and Glu-V3/Gli-V1 loci could provide novel germplasm for the improvement of bread wheat quality.
The distinct differences for SPC and the values of ZSV, dough development time (DT) and dough stability time (ST) values between the three lines NAU1V-3, NAU1V-4 and NAU1V-5 could be attributed in part to the interaction of prolamins encoded by loci of CS and 1V#4S. T1V#4S-6BS·6BL (NAU1V-3) translocation line had significantly lower positive effects on SPC compared to DA1V#4. This may due to the missing of the Gli-B2 loci located in the 6BS. The T1V#4S·1BL (NAU1V-4) translocation line also had significantly positive effects on gluten strength when 1V#4S replaced 1BS, suggesting a higher positive effect on quality owing to the glutens and gliadins encoded at the Glu-V1 and Gli-V1/Glu-V3 loci of 1V#4S than that of Gli-1/Glu-3 loci of 1BS. The similar results were also observed when the Gli-B1/Glu-B3 loci were substituted by Glu-V1 and Gli-V1/Glu-V3 loci in the line 09.CS 1B-1V (T1BL-1V#1L·1V#1S) (Vaccino et al. 2010).The Glu-D1 locus on 1DL shows a drastic positive effect on gluten strength compared with the euploid CS (Rogers et al. 1990). Although the Glu-D1 locus is absent in T1V#4S·1DS translocation line NAU1V-5 (Fig. 4b), the quality is also increased compared to the CS, suggesting a higher positive effect on gluten strength of 1V#4S than that of 1DL.
Extensive studies on SSPs in tribe Triticeae have been carried out and revealed genetic diversity among wheat and its relative species. SSPs of rye (Secale cereale) substituted for wheat SSPs, for example, T1BL·1RS translocation line, resulted in inferior wheat quality (Lukaszewski 1993). Whereas, SSPs located on chromosome 1E of Thinopyrum elongatum had positive effects in addition lines but not in substitution lines for chromosome 1D of wheat (Garg et al. 2009a). In contrast, SSPs of Th. intermedium have positive effects, even in case of substitution for chromosome 1D of wheat (Cao et al. 2007). De Pace et al. (2001) have confirmed that SSPs located on chromosome 1V of D. villosum had large positive effects of bread quality in addition line CS+1V#1, substitution lines CS·1V#1 (1A) and CS·1V#1 (1B). Vaccino et al. (2010) observed the similar results when the Gli-B1/Glu-B3 loci were substituted by Glu-V1 and Gli-V1/Glu-V3 loci in the line 09.CS 1B-1V (T1BL-1V#1L·1V#1S). While, Dong et al. (2013) suggested that T1DS·1V#3L had a significant negative effect on wheat dough strength, but T1DL·1V#3S significantly improved bread-making quality. Our results also suggested that SSPs located on chromosome 1V#4S of D. villoum had positive effects in addition lines and translocation lines. The significant positive effects of quality in the wheat having D. villoum chromosome 1VS are believed to be due to D. villoum alleles at the Glu-V1 and Gli-V1/Glu-V3 loci. Although, Vaccino et al. (2010) argue that the improved bread-making quality was attributed to the Glu-V1 introgressed in CS because replacing Gli-B1/Glu-B3 loci by the Gli-V1/Glu-V3 loci affected end-use grain quality by decreasing the specific SDS sedimentation volume and increasing the water absorption properties of the flour. We proposed that the interaction effect of Glu-V1 between Gli-V1/Glu-V3 loci may also have contribution on grain quality because HMW and LMW subunits can form complex polymeric proteins by inter-chain disulphide bonds and impart dough viscoelasticity (Shewry et al. 2003). Also, whether the effect on grain quality of Glu-V1 locus only is better than that of the interaction of three loci needs further study. The small segmental translocation line (T1V#4S-6BS·6BL, NAU1V-3) and the whole-arm translocation line (T1V#4S·1BL, NAU1V-4), which had significantly improved and stably inherited dough strength, could be used directly by wheat breeders. In addition, the whole-arm translocation line T1V#4S·1BL could be used as initial materials to induce small alien chromosome translocations only carries the Glu-V1 locus between 1V#4S and short arms of group 1 using CSph1b mutant. The five 1V#4S-specific molecular markers developed could be used to detect the alien chromatin of 1V#4S in wheat background.
References
Agnieszka G (2006) The genus Dasypyrum—part 2. Dasypyrum villosum—a wild species used in wheat improvement. Euphytica 152:441–454
Blanco A, Resta P, Simeone R, Parmar S, Shewry PR, Sabelli P, Lafiandra D (1991) Chromosomal location of seed storage protein genes in the genome of Dasypyrum villosum (L.) Candargy. Theor Appl Genet 82:358–362
Branlard G, Dardevet M (1985) Diversity of grain protein and bread wheat quality. II. Correlation between high molecular weight subunits of glutenin and flour quality characteristics. J Cereal Sci 3:345–354
Cao S, Xu H, Li Z, Wang X, Wang D, Zhang A, Jia X, Zhang X (2007) Identification and characterization of a novel Ag.intermedium HMW-GS gene from T. aestivum to Ag. intermedium addition lines TAI-I series. J Cereal Sci 45:293–301
Chen PD, Qi LL, Zhou B, Zhang SZ, Liu DJ (1995) Development and molecular cytogenetic analysis of wheat-Haynaldia villosa 6VS/6AL translocation lines specifying resistance to powdery mildew. Theor Appl Genet 91:1125–1128
Chen PD, Liu WX, Yuan JH (2005) Development and characterization of wheat-Leymus racemosus translocation lines with resistance to Fusarium Head Blight. Theor Appl Genet 111:941–948
Chen SW, Chen PD, Wang XE (2008) Inducement of chromosome translocation with small alien segments by irradiating mature female gametes of the whole arm translocation line. Sci China Ser C Life Sci 51(4):346–352
De Pace C, Snidaro D, Ciaffi M, Vittori D, Ciofo A, Cenci A, Tanzarella OA, Qualset CO, Scarascia Mugnozza GT (2001) Introgression of Dasypyrum villosum chromatin into common wheat improves grain protein quality. Euphytica 117:67–75
De Pace C, Vaccino P, Cionini G, Pasquini M, Bizzarri M, Qualset CO (2011) Dasypyrum. In: Kole C (ed) Wild crop relatives: genomic and breeding resources, cereals, vol 1, chapter 4. Springer, Heidelberg, pp 185–292
Dong J, Yang H, Zhao WC, LI XY, Chen QJ, Gao X (2013) Agronomic traits and grain quality of Chinese Spring–Dasypyrum villosum translocation lines T1DL·1VS and T1DS·1VL. Acta Agron Sin 39(8):1386–1390
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12(1):13–15
Frank MY, Naxin H, Yong QG, Gerard RL, Dvorak J, Anderson OD (2009) ConservedPrimers 2.0: a high-throughput pipeline for comparative genome referenced intron-flanking PCR primer design and its application in wheat SNP discovery. BMC Bioinformatics 10:331–341
Friebe B, Jiang J, Raupp WJ (1996) Characterization of wheat-alien translocations conferring resistance to diseases and pest: current status. Euphytica 91:59–87
Gao X, Liu SW, Sun Q, Xia GM (2010) High frequency of HMW-GS sequence variation through somatic hybridization between Agropyron elongatum and common wheat. Planta 231:245–250
Garg M, Tanaka H, Ishikawa N, Takata K, Yanaka M, Tsujimoto H (2009a) Agropyron elongatum HMW-glutenins have a potential to improve wheat end-product quality through targeted chromosome introgression. J Cereal Sci 50:358–363
Garg M, Tanaka H, Tsujimoto H (2009b) Exploration of Triticeae seed storage proteins for improvement of wheat end product quality. Breed Sci 59:519–528
Gill BS, Friebe B, Endo TR (1991) Standard karyotype and nomenclature system for description of chromosome bands and structural aberrations in wheat (Triticum aestivum). Genome 34:830–839
Halford NG, Field JM, Blair H, Urwin P, Moore K, Robert L, Thompson R, Flavell RB, Tatham AS, Shewry PR (1992) Analysis of HMW glutenin subunits encoded by chromosome 1A of bread wheat (Triticum aestivum L.) indicates quantitative effects on grain quality. Theor Appl Genet 83:373–378
Ishikawa G, Yonemaru J, Saito M, Nakamura T (2007) PCR based landmark unique gene (PLUG) markers effectively assign homoeologous wheat genes to A, B and D genomes. BMC Genomics 8:135–147
Li GR, Liu C, Li CH, Zhao JM, Zhou L, Dai G, Yang EN, Yang ZJ (2013) Introgression of a novel Thinopyrum intermedium St-chromosome-specific HMW-GS gene into wheat. Mol Breed. doi:10.1007/s11032-013-9838-8
Liu SW, Gao X, Xia GM (2008) Characterizing HMW-GS alleles of decaploid Agropyron elongatum in relation to evolution and wheat breeding. Theor Appl Genet 116:325–334
Liu S, Zhu X, Tan Y, Liu S (2012) Isolation and characterization of Glu-1 genes from the St genome of Pseudoroegneria libanotica. Gene 499:154–159
Lukaszewski AJ (1993) Comparative flour quality and protein characteristics of 1BL/1RS, and 1AL/1RS wheat-rye translocation lines. J Cereal Sci 17:95–106
Mackie AM, Lagudah ES, Sharp PJ, Lafiandra D (1996) Molecular and biochemical characterization of HMW glutenin subunits from T. tauschii and the D genome of hexaploid wheat. J Cereal Sci 23:213–225
McIntosh RA, Yamazaki Y, Dubcovsky J, Rogers WJ, Morris CF, Somers DJ, Appels R, Devos KM (2008) Catalogue of gene symbols for wheat. In: Proceedings of the 11th international wheat genetic symposium, Sydney University of Sydney Press, Australia
Montebove L, De Pace C, Jan CC, Scarascia-Mugnozza GT, Qualset CO (1987) Chromosomal location of isozyme and seed storage protein genes in Dasypyrum villosum (L.) Candargy. Theor Appl Genet 73:836–845
Mukai Y, Endo TR, Gill BS (1991) Physical mapping of the 18S·26S rRNA multigene family in common wheat: identification of a new locus. Chromosoma 100:71–78
Mukai Y, Nakaharya Y, Yamamotmo M (1993) Simultaneous discrimination of the three genomes in hexaploid wheat by multicolor fluorescence in situ hybridization using total genomic and highly repeated DNA probes. Genome 36:489–494
Niu ZX, Klindworth DL, Wang RRC, Jauhar PP, Larkin PJ, Xu SS (2011) Characterization of HMW glutenin subunits in Thinopyrum intermedium, Th.bessarabicum, Lophopyrum elongatum, Aegilops markgrafii, and their addition lines in wheat. Crop Sci 51:667–677
Payne PI, Law CN, Mudd EE (1980) Control by homoeologous group 1 chromosomes of the high-molecular-weight subunits of glutenin, a major protein of wheat endosperm. Theor Appl Genet 58:113–120
Payne PI, Nightingale MA, Krattiger AF, Holt LM (1987) The relationship between HMW glutenin subunit composition and the breadmaking quality of British grown wheat varieties. J Sci Food Agric 40:51–65
Payne PI, Holt LM, Krattiger AF, Carrillo JM (1988) Relationship between seed quality characteristics and HMW glutenin subunit composition determined using wheats grown in Spain. J Cereal Sci 7:229–235
Pedersen C, Langridge P (1997) Identification of the entire chromosome complement of bread wheat by two-colour FISH. Genome 40:589–593
Qi LL, Chen PD, Liu DJ, Gill BS (1999) Homoeologous relationships of Haynaldia villosa chromosomes with those of Triticum aestivum as revealed by RFLP analysis. Genes Genet Syst 74:77–82
Qi LL, Pumphrey MO, Friebe B, Zhang P, Qian C, Bowden RL, Rouse MN, Jin Y, Gill BS (2011) A novel Robertsonian translocation event leads to transfer of a stem rust resistance gene (Sr52) effective against race Ug99 from Dasypyrum villosum into bread wheat. Theor Appl Genet 123:159–167
Rogers WJ, Rickatson JM, Sayers EJ, Law CN (1990) Dosage effects of chromosomes of homoeologous groups 1 and 6 upon bread-making quality in hexaploid wheat. Theor Appl Genet 80:281–287
Sears ER (1953) Addition of the genome of Haynaldia villosa–Triticum aestivum. Am J Bot 40:168–174
Sears ER, Gustafson JP (1993) Use of radiation to transferalien chromosome segments to wheat. Crop Sci 33:897–901
Shewry PR, Tatham AS (1990) The prolamin storage proteins of cereal seeds: structure and evolution. Biochem J 267:1–12
Shewry PR, Parmar S, Pappin DJC (1987) Characterization and genetic control of the prolamins of Haynaldia villosa: relationship to cultivated species of the Triticeae (rye, wheat and barley). Biochem Genet 25:309–325
Shewry PR, Sabelli PA, Parmar S, Lafiandra D (1991) λ-Type prolamins are encoded by genes on chromosomes 4Ha and 6Ha of Haynaldia villosa Schur (syn. Dasypyrum villosum L.). Biochem Genet 29:207–211
Shewry PR, Halford NG, Tatham AS, Popineau Y, Lafiandra D, Belton PS (2003) The high molecular weight subunits of wheat glutenin and their role in determining wheat processing properties. Adv Food Nut Res 45:219–302
Sun X, Hu SL, Liu X, Qian WQ, Hao ST, Zhang AM, Wang DW (2006) Characterization of the HMW glutenin subunits from Aegilops searsii and identification of a novel variant HMW glutenin subunit. Theor Appl Genet 113:631–641
Vaccino P, Banfi R, Corbellini M, De Pace C (2010) Broadening and improving the wheat genetic diversity for end-use grain quality by introgression of chromatin from the wheat wild relative Dasypyrum villosum. Crop Sci 50:528–540
Wan Y, Wang D, Shewry PR, Halford NG (2002) Isolation and characterization of five novel high molecular weight subunit of glutenin genes from Triticum timopheevi and Aegilops cylindrica. Theor Appl Genet 104:828–839
Wang CM, Feng YG, Zhuang LF, Cao YP, Qi ZJ, Bie TD, Cao AZ, Chen PD (2007) Screening of chromosome-specific markers for chromosome 1R of Secale cereale, 1V of Haynaldia villosa and 1Rk# 1 of Roegneria kamoji. Acta Agron Sin 33:1741–1747
Wang MJ, Zhang Y, Lin ZS, Ye XG, Yuan YP, Ma W, Xin ZY (2010) Development of EST–PCR markers for Thinopyrum intermedium chromosome 2Ai#2 and their application in characterization of novel wheat-grass recombinants. Theor Appl Genet 121:1369–1380
Xia GM, Xiang FN, Zhou AF, Wang H, Chen HM (2003) Asymmetric somatic hybridization between wheat (Triticum aestivum L.) and Agropyron elongatum (Host) Nevishi. Theor Appl Genet 107:299–305
Zhang Q, Li Q, Wang X, Wang H, Lang S, Wang Y, Wang S, Chen P, Liu D (2005) Development and characterization of a Triticum aestivum–Haynaldia villosa translocation line T4VS/4DL conferring resistance to wheat spindle streak mosaic virus. Euphytica 145:317–320
Zhang RQ, Cao YP, Wang XE, Feng YG, Chen PD (2010) Development and characterization of a Triticum aestivum–D. villoum T5VS.5DL translocation line with soft grain texture. J Cereal Sci 51:220–225
Zhang RQ, Wang XE, Chen PD (2012) Molecular and cytogenetic characterization of a small alien-segment translocation line carrying the softness genes of Haynaldia villosa. Genome 55:639–646
Zhang W, Zhang RQ, Feng YG, Bie TD, Chen PD (2013) Distribution of highly repeated DNA sequences in Haynaldia villosa and its application in the identification of alien chromatin. Chin Sci Bull 58:890–897
Zhao WC, Qi LL, Gao X, Zhang GS, Dong J, Chen QJ, Friebe B, Gill BS (2010) Development and characterization of two new Triticum aestivum–Dasypyrum villosum Robertsonian translocation lines T1DS.1V#3L and T1DL.1V#3S and their effect on grain quality. Euphytica 175:343–350
Zhong GY, Qualset CO (1993) Allelic diversity of high-molecular-weight glutenin protein subunits in natural populations of Dasypyrum villosum (L.) Candargy. Theor Appl Genet 86:851–858
Acknowledgments
This study was supported by the National Natural Science Foundation of China (No. 31101142), the “863” high-Tech project (No. 2011AA10010201), the Ph.D Programs Foundation of Ministry of Education of China (No. 20110097120040) and the Project Funded by the Priority Academic Program Development of Jiangsu Higher Education Institutions (PAPD).
Conflict of interest
All the authors have no conflict of interest and agree with published.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Pat. Heslop-Harrison.
Rights and permissions
About this article
Cite this article
Ruiqi, Z., Mingyi, Z., Xiue, W. et al. Introduction of chromosome segment carrying the seed storage protein genes from chromosome 1V of Dasypyrum villosum showed positive effect on bread-making quality of common wheat. Theor Appl Genet 127, 523–533 (2014). https://doi.org/10.1007/s00122-013-2244-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-013-2244-0