Abstract
A symmetric somatic hybridization was performed to combine the protoplasts of tall wheatgrass (Agropyron elongatum) and bread wheat (Triticum aestivum). Fertile regenerants were obtained which were morphologically similar to tall wheatgrass, but which contained some introgression segments from wheat. An SDS-PAGE analysis showed that a number of non-parental high-molecular weight glutenin subunits (HMW-GS) were present in the symmetric somatic hybridization derivatives. These sequences were amplified, cloned and sequenced, to deliver 14 distinct HMW-GS coding sequences, eight of which were of the y-type (Hy1–Hy8) and six x-type (Hx1–Hx6). Five of the cloned HMW-GS sequences were successfully expressed in E. coli. The analysis of their deduced peptide sequences showed that they all possessed the typical HMW-GS primary structure. Sequence alignments indicated that Hx5 and Hy1 were probably derived from the tall wheatgrass genes Aex5 and Aey6, while Hy2, Hy3, Hx1 and Hy6 may have resulted from slippage in the replication of a related biparental gene. We found that both symmetric and asymmetric somatic hybridization could promote the emergence of novel alleles. We discussed the origination of allelic variation of HMW-GS genes in somatic hybridization, which might be the result from the response to genomic shock triggered by the merger and interaction of biparent genomes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The high-molecular weight glutenin subunits (HMW-GS) are a set of well-conserved endosperm proteins synthesized in the grain of wheat and related grasses (Lawrence and Shepherd 1981; Shewry et al. 2003). In hexaploid wheat, they are encoded by the Glu-1 homoeoloci located on the long arms of chromosomes 1A, 1B and 1D, with each locus comprising a pair of tightly linked genes encoding the x-type (Glu-1-1) and the y-type (Glu-1-2) subunits (Lawrence and Shepherd 1981; Payne 1987). Qualitative and quantitative variation in the HMW-GS is associated with 45–70% of the variation in bread-making performance of European wheat, even though they only represent about 10% of grain protein (Branlard and Dardevet 1985; Payne et al. 1987, 1988). Because of their importance for wheat quality improvement, a substantial number of Glu-1 genes have been cloned (Forde et al. 1985; Sugiyama et al. 1985; Thompson et al. 1985; Halford et al. 1987; Anderson and Greene 1989; Anderson et al. 1989; Halford et al. 1992). Sequence analysis of these genes has shown that each contains a long repetitive region, flanking two highly conserved terminal non-repetitive domains. The repetitive region includes tripeptides, hexapeptides and nonapeptides, with the tripeptide motif restricted to the x-type subunits.
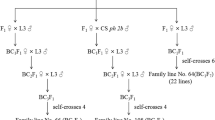
Although HMW-GS are clearly important for the determination of end-use quality, the number of high quality alleles is rather limited within the bread wheat genepool. Thus, some effort has been made to transfer alleles from related species, using either a wide crossing (Zhou et al. 1995) or a somatic hybridization approach. Our focus has been to take advantage of the latter route. We have so far succeeded in fusing the protoplasts of the bread wheat cultivar Jinan 177 (JN177) with UV-irradiated protoplasts of tall wheatgrass (Agropyron elongatum (Host) Nevski [Thinopyrum ponticum]) (Liu SW et al. 2007; Liu H et al. 2009). We have also attempted symmetric somatic hybridization, in which the tall wheatgrass protoplasts were not UV irradiated. Regenerant plants of this latter protoplast fusion resembled the tall wheatgrass parent, but inherited several introgression segments from wheat (Cui et al. 2009). Selections CU and XI (each derived from a single fusion cell) were particularly fertile. Here, we have investigated whether any novel HMW-GS alleles are present in these somatic hybrid regenerants.
Materials and methods
Plant materials
The plant material used in these experiments consisted of the tall wheatgrass and bread wheat biparents of the somatic fusion, five R3 (third generation following the regeneration of the primary somatic hybrid) lines R3CU1, R3CU2, R3CU3, R3XI1 and R3XI2. Karyotypic analysis indicated that the chromosome number of the in vitro cultured tall wheatgrass cells ranged from 60 to 70, while about 80% of R3CU1–R3CU3 and R3XI1–R3XI2 cells carried 66–70 chromosomes. Cui et al. (2009) showed, by a combination of cytological and marker analyses, that the wheat chromosome segments were introgressed into A. elongatum chromosomes in the genomes of R1–R3 regenerants. Both JN177 and the introgression lines were grown in a greenhouse separated from other wheat cultivars to avoid uncontrolled outcrossing.
SDS-PAGE analysis of HMW-GS
The HMW-GS content of JN177 was obtained by SDS-PAGE analysis (Feng et al. 2004) of a crude protein extract of an embryo-less half grains, while those of tall wheatgrass and the introgression lines were obtained from extracts of the whole seed, as described by Mackie et al. (1996).
Cloning and characterizing of HMW-GS genes
Genomic DNA was extracted from the introgression line seedlings using the CTAB method (Murray and Thompson 1980). As the HMW-GS genes are intron less, genomic DNA was used as a PCR template to amplify the entire coding region. A pair of degenerate primers (P1: 5′-ATGGCTAAGCGGc/tTa/gGTCCTCTTTG and P2: 5′-CTATCACTGGCTa/gGCCGACAATGCG) was designed on the basis of published DNA sequences. P1 includes the HMW-GS start codon, and P2 includes the two conserved tandem stop codons. PCR amplification employed a high fidelity LA Taq polymerase (TaKaRa Biotechnology, Dalian, China) with a GC buffer provided for GC-rich template. The amplification profile consisted of a denaturation step (95°C/5 min), followed by 28 cycles of 94°C/40 s, 68°C/4 min, and ending with an extension step (72°C/7 min). The amplicon was recovered from a 1% agarose gel, cloned into the pMD18-T vector (TaKaRa Biotechnology, Dalian, China), and transformed into E. coli DH10B competent cells. Sequencing was performed commercially (Invitrogen, Shanghai, China). Both amplification and cloning were repeated at least three times to minimize the risk of amplification and/or sequencing errors. Sequence analyses were carried out using the MEGA software package v3.1 (Kumar et al. 2004) along with standard programs available from NCBI (http://www.ncbi.nlm.nih.gov/Tools/) and EBI (http://www.ebi.ac.uk/Tools/sequence.html).
Bacterial expression of HMW-GS sequences
To express the mature introgression line HMW-GS proteins in E. coli, two sets of PCR primers (PF/PR1 and PF/PR2) were designed to amplify the sequences while excluding their signal peptides, and at the same time introducing cloning sites. The sequence of PF was 5′-ACCCATATGGAAGGTGAGGCCTCT-3′, that of PR1 was 5′-CTAGAATTCCTATCACTGGCTGGCCGA-3′ (for Hy4, Hy3, Hy7, Hy8) and that of PR2 was 5′-CTAGAATTCCTATCACTGGCTAGCCGA-3′ (for Hy6). PF contains an NdeI site and both PR1 and PR2 an EcoRI site. The amplicons were cloned into the expression vector pET-24a (Novagen, Shanghai, China), and transformed into E. coli BL21 (DE3) pLysS competent cells (Promega, Shanghai, China). Heterologous expression was induced using standard methods (Sambrook et al. 1989) and proteins were extracted by dissolving cells in SDS-PAGE sample buffer for SDS-PAGE analysis (Wan et al. 2002).
Results
The HMW-GS content of the biparents and the introgression lines
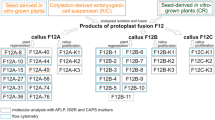
The HMW-GS content of JN177 is 1Bx7.1 + 1By9.1; 1Dx2.1 + 1Dy12.1, while that of the tall wheatgrass consists of nine distinct subunits. The HMW-GS composition of the five introgression lines R3CU1–R3CU3 and R3XI1–R3XI2 was overall very similar to that of the tall wheatgrass, although a small number of novel subunits could be identified (Fig. 1). A gel separation of the amplicons derived from each of the five introgression lines was shown in Fig. 2. After restriction enzyme digestion mapping and terminal DNA sequencing, we confirmed that at least 14 distinct sequences had been amplified from the introgression lines (designated as Hx1–Hx6 and Hy1–Hy8, according to their type and length).
Expression of the HMW-GS alleles in bacterial cells
Five of the cloned sequences with intact ORFs were successfully expressed in E. coli, namely pET-Hy3, pET-Hy4 pET-Hy6, pET-Hy7 and pET-Hy8. The SDS-PAGE mobility of four of these (Hy4, Hy6, Hy3 and Hy8) was similar to that of equivalent subunits extracted from tall wheatgrass seeds, but there was no match between the proteins directed by pET-Hy7 and any tall wheatgrass seed-extracted protein (Fig. 3).
SDS-PAGE analysis of heterologously expressed HMW-GS proteins. M protein molecular weight marker, lane 1 JN177, lane 2 tall wheatgrass, lane 3 bacteria harboring pET-Hy4 without IPTG induction, lanes 4–8 expression of the modified sequences of Hy4, Hy6, Hy3, Hy7 and Hy8. The HMW-GS gene-directed proteins induced by IPTG are indicated by arrows
Characteristic of derived amino acid sequences of HMW-GS alleles
The deduced peptide sequences of the 14 HMW-GS genes shared the expected primary structure. Each consisted of a 21 residues signal peptide, a conserved N-terminal region, a central repetitive domain and a conserved C-terminal region. The N-terminal regions of five of the eight y-type subunits include 105 residues, while this length in Hy1, Hy4 and Hy5 was 104, 76 and 59 residues, respectively (Table 1). N-terminal regions of Hy1 lacked a glutamine residue when compared with Hy2, Hy3, Hy6, Hy7 and Hy8. This glutamine residue is also present in all the known x-type subunits. The conserved C-terminal regions of all the 14 subunits comprise 42 residues, and their central repetitive region included both hexapeptide and nonapeptide motifs; the six x-type subunits also contained the diagnostic GQQ tripeptide motif. Differences between these subunits and those already known in wheat lie mostly in single residue substitutions, and the insertion/deletion of repeat motifs in central repetitive region. The deduced peptide lengths of these subunits varied from 817 (Hx1) to 295 (Hy7 and Hy8) residues (Table 1). The Hy7 and Hy8 are also two of the smallest known HMW-GS.
Relationships between HMW-GS sequences
A phylogenetic tree was assembled from the alignment of the full-length nucleotide sequences of the 14 HMW-GS genes and the HMW-GS genes from JN177 and tall wheatgrass (Liu SW et al. 2007, 2008) (Fig. 4). As expected, the y-type genes were separated from the x-type ones. The eight y-type and the six x-type sequences each clustered into three clades. The Hy1 sequence resembled that of tall wheatgrass Aey6, and was distantly related to the remaining seven y-type sequences, while Hy6 was more similar to Aey10 than to any of the other introgression line alleles. The other six y-type alleles of the introgression lines fell into three subgroups. Hy2 and Hy3 shared a close relationship with Aey8, while Hy7 was similar to Hy8, as were Hy4 and Hy5. Of the six x-type alleles, Hx1 was similar to Aex2, Hx5 to Aex5 and quite closely related to Hx2, Hx3 and Hx4, Hx6 was an outlier within the x-type clade.
Phylogenetic analysis of the HMW-GS sequences in some bread wheat/tall wheatgrass symmetric somatic hybridization derivatives and their parents. The phylogenetic tree was constructed according to the full-length DNA sequences using the MEGA software package v3.1. Hy1–Hy8 and Hx1–Hx6 came from the somatic hybridization derivatives; Aey1–Aey10 and Aex1–Aex5 came from tall wheatgrass; 1Bx7.1, 1By9.1, 1Dx2.1 and 1Dy12.1 came from JN177
Discussion
Asymmetric somatic hybridization between bread wheat and UV-irradiated tall wheatgrass is known to generate wheat-like introgression lines (Xia et al. 2003; Wang et al. 2005), among which a deal of allelic variation for the HMW-GS genes has been identified (Liu SW et al. 2007; Liu H et al. 2009). Symmetric somatic hybridization involving the same biparents has produced fertile regenerants which more resemble tall wheatgrass in phenotype, but whose genomes still contain some introgressed wheat segments (Cui et al. 2009). Therefore, we obtained a contrary introgression line of wheat/tall wheatgrass, which is favorable for exploring the variation of HMW-GS in different introgression lines via symmetric or asymmetric somatic hybridization.
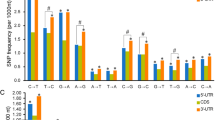
The Hx5 and Hy1 sequences each differed by only a small number of single nucleotides from a tall wheatgrass sequence (Aex5 and Aey6, respectively), and they had no close match with any of the HMW-GS sequences present in another parent JN177. Thus, it is likely that both were inherited from the tall wheatgrass parent, suffering some point mutation as a result of the somatic hybridization process. Similarly, Hy2 and Hy3 resembled Aey8, but for the presence of additional repeat motifs and a few single-nucleotide polymorphisms. Hx1 and Hy6 resembled Aex2 and Aey10, respectively. When compared with Aex2, Hx1 gained three additional repeats but lost one, while, compared to Aey10, Hy6 gained one and lost two (Fig. 5). Possibly, therefore, these four Glu-1 alleles may have derived via slippage of their corresponding parental gene during replication. The reason for why we have not found the origin of the other six novel HMW-GS sequences of the introgression lines might be that A. elongatum was a cross-pollinating species and there were plenty of the Glu-1 alleles in A. elongatum and we have only obtained a limited number of Glu-1 alleles from them (Liu SW et al. 2008).
The addition or deletion of repeat motifs is thought to be an effective source of variation (Wells 1996), while Anderson and Greene (1989) have proposed that the evolution of HMW-GS genes proceeds via a combination of single base changes, deletions or additions within a repeat, single repeat changes and deletions or duplications of blocks of repeats. The formation of some novel hybrid genes was inosculated with the mechanism mentioned by Anderson and Greene (1989). Thus, the forms of novel HMW-GS alleles generated in these introgression lines are consistent with that of naturally emerging ones, although the process of their formation appears to be accelerated by the somatic hybridization procedure.
Both the present symmetric hybridization experiments, as well as those based on the asymmetric hybridization (Liu SW et al. 2007; Liu H et al. 2009) have produced regenerants carrying a number of novel HMW-GS alleles. Although the regenerants from JN177 callus have also been shown to produce novel HMW-GS alleles, the somaclonal mutation rate is much lower (Feng et al. 2004). Although some of the somatic hybridization-induced alleles may have arisen through somaclonal variation, it seems likely that many resulted from an interaction between the biparental genomes and/or the process of protoplast fusion itself; in the case of the asymmetric hybridization products, an additional source of variation is provided by the pre-fusion UV-irradiation treatment (Liu H et al. 2009). The analysis of certain newly synthesized alloploids has shown that when two genomes are united in a single nucleus, some instability ensues, which results in the elimination of genomic DNA sequences, the alteration of cytosine methylation patterns, and the reactivation of retrotransposons (Shaked et al. 2001; Ozkan et al. 2001; Madlung et al. 2002, 2005). Similar instability is, therefore, not unexpected in a somatic hybrid, and this has been demonstrated in wheat/tall wheatgrass combinations in the form of variation at microsatellite sequences, the elimination of DNA sequences, changes in the pattern of cytosine methylation and silencing or activation of homoeologous alleles (unpublished). The wide hybridization of different genomes might trigger a genomic shock that lead to these responses and it fit McClintock’s view about genomic shock response that “initiates a highly programmed sequence of events within the cell that serves to cushion the effect of the shock” (McClintock 1984). Therefore, the response to genomic shock triggered by the merger and interaction of biparent genomes might be mainly responsible for the sequence variation in the introgression lines.
In conclusion, we have shown here that the variation in the HMW-GS sequences can be induced by symmetric as well as asymmetric somatic hybridization. It is possible that some of the novel alleles may make a positive contribution to the wheat end-use quality and is favorable to the investigation of genome variation and evolution.
Abbreviations
- AFLP:
-
Amplification fragment length polymorphism
- GISH:
-
Genome in situ hybridization
- HMW-GS:
-
High-molecular weight glutenin subunit
- SDS-PAGE:
-
Sodium dodecyl sulfate-polyacrylamide gel electrophoresis
References
Anderson OD, Greene FC (1989) The characterization and comparative analysis of high-molecular-weight glutenin genes from genomes A and B of a hexaploid bread wheat. Theor Appl Genet 77:689–700
Anderson OD, Greene FC, Yip RE, Halford NG, Shewry PR, Malpica-Romero J-M (1989) Nucleotide sequences of the two high-molecular-weight glutenin genes from the D-genome of a hexaploid bread wheat, Triticum aestivum L. cv Cheyenne. Nucleic Acids Res 17:461–462
Branlard G, Dardevet M (1985) Diversity of grain protein and bread wheat quality. II. Correlation between high molecular weight subunits of glutenin and flour quality characteristics. J Cereal Sci 3:345–354
Cui H, Yu Z, Deng J, Gao X, Sun Y, Xia G (2009) Introgression of bread wheat chromatin into tall wheatgrass via somatic hybridization. Planta 229:323–330
Feng DS, Xia GM, Zhao SY, Chen FG (2004) Two quality-associated HMW glutenin subunits in a somatic hybrid line between Triticum aestivum and Agropyron elongatum. Theor Appl Genet 110:136–144
Forde J, Malpica J-M, Halford NG, Shewry PR, Anderson OD, Greene FC, Miflin BJ (1985) The nucleotide sequence of a HMW glutenin subunit gene located on chromosome 1A of wheat (Triticum aestivum L.). Nucleic Acids Res 13:6817–6832
Halford NG, Forde J, Anderson OD, Greene FC, Shewry PR (1987) The nucleotide and deduced amino acid sequences of an HMW glutenin subunit gene from chromosome 1B of bread wheat (Triticum aestivum L.) and comparison with those of genes from chromosomes 1A and 1D. Theor Appl Genet 75:117–126
Halford NG, Field JM, Blair H, Urwin P, Moore K, Robert L, Thompson R, Flavell RB, Tatham AS, Shewry PR (1992) Analysis of HMW glutenin subunits encoded by chromosome 1A of bread wheat (Triticum aestivum L.) indicates quantitative effects on grain quality. Theor Appl Genet 83:373–378
Kumar S, Tamura K, Nei M (2004) MEGA3: integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief Bioinform 5:150–163
Lawrence GJ, Shepherd KW (1981) Inheritance of glutenin protein subunits of wheat. Theor Appl Genet 60:333–337
Liu SW, Zhao SY, Chen FG, Xia GM (2007) Generation of novel high quality HMW-GS genes in two introgression lines of Triticum aestivum/Agropyron elongatum. BMC Evol Biol 7:76
Liu SW, Xin Gao, Xia GM (2008) Characterizing HMW-GS alleles of decaploid Agropyron elongatum in relation to evolution and wheat breeding. Theor Appl Genet 116:325–334
Liu H, Liu SW, Xia GM (2009) Generation of high frequency of novel alleles of the high molecular weight glutenin in somatic hybridization between bread wheat and tall wheatgrass. Theor Appl Genet 118:1193–1198
Mackie AM, Lagudah ES, Sharp PJ, Lafiandra D (1996) Molecular and biochemical characterization of HMW glutenin subunits from T. tauschii and the D genome of hexaploid wheat. J Cereal Sci 23:213–225
Madlung A, Masuelli RW, Watson B, Reynolds SH, Davison J, Comai L (2002) Remodeling of DNA methylation and phenotypic and transcriptional changes in synthetic Arabidopsis allotetraploids. Plant Physiol 129:733–746
Madlung A, Tyagi AP, Watson B, Jiang H, Kagochi T, Doerge RW, Martienssen R, Comai L (2005) Genomic changes in synthetic Arabidopsis polyploids. Plant J 41:221–230
McClintock B (1984) The significance of responses of the genome to challenge. Science 226:792–801
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8:4321–4325
Ozkan H, Levy AA, Feldman M (2001) Allopolyploidy-induced rapid genome evolution in the wheat (Aegilops–Triticum) group. Plant Cell 13:1735–1747
Payne PI (1987) Genetics of wheat storage proteins and the effect of allelic variation on breadmaking quality. Annu Rev Plant Physiol 38:141–153
Payne PI, Nightingale MA, Krattiger AF, Holt LM (1987) The relationship between HMW glutenin subunit composition and the breadmaking quality of British grown wheat varieties. J Sci Food Agric 40:51–65
Payne PI, Holt LM, Krattiger AF, Carrillo JM (1988) Relationship between seed quality characteristics and HMW glutenin subunit composition determined using wheats grown in Spain. J Cereal Sci 7:229–235
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory, New York
Shaked H, Kashkush K, Ozkan H, Feldman M, Levy AA (2001) Sequence elimination and cytosine methylation are rapid and reproducible responses of the genome to wide hybridization and allopolyploidy in wheat. Plant Cell 13:1749–1759
Shewry PR, Halford NG, Tatham AS, Popineau Y, Lafiandra D, Belton PS (2003) The high molecular weight subunits of wheat glutenin and their role in determining wheat processing properties. Adv Food Nut Res 45:219–302
Sugiyama T, Rafalski A, Peterson D, Söll D (1985) A wheat HMW glutenin subunit gene reveals a highly repeated structure. Nucleic Acids Res 13:8729–8737
Thompson RD, Bartels D, Harberd NP (1985) Nucleotide sequence of a gene from chromosome 1D of wheat encoding a HMW-glutenin subunit. Nucleic Acids Res 13:6833–6846
Wan Y, Wang D, Shewry PR, Halford NG (2002) Isolation and characterization of five novel high molecular weight subunit of glutenin genes from Triticum timopheevi and Aegilops cylindrica. Theor Appl Genet 104:828–839
Wang J, Xiang FN, Xia GM (2005) Agropyron elongatum chromatin localization on the wheat chromosomes in an introgression line. Planta 211:277–286
Wells RD (1996) Molecular basis of genetic instability of triplet repeats. J Biol Chem 271:2875–2878
Xia GM, Xiang FN, Zhou AF, Wang H, Chen HM (2003) Asymmetric somatic hybridization between wheat (Triticum aestivum L.) and Agropyron elongatum (Host) Nevishi. Theor Appl Genet 107:299–305
Zhou HP, Li B, Li ZS (1995) The study of breeding blue-grain gene translocation of wheat (in Chinese with English Abstract). Acta Bot Boreal-Occident Sin 15:125–128
Acknowledgments
This research was supported by the National Natural Science Foundation of China No. 30871320 and National 863 High Technology Research and Development Project 2006AA100102, the National Key Technology R&D Program 2007BAD59B06 and the National Transgenic Project 2009ZX08002-014B.
Author information
Authors and Affiliations
Corresponding author
Additional information
X. Gao and S. W. Liu contributed equally to the work.
Rights and permissions
About this article
Cite this article
Gao, X., Liu, S.W., Sun, Q. et al. High frequency of HMW-GS sequence variation through somatic hybridization between Agropyron elongatum and common wheat. Planta 231, 245–250 (2010). https://doi.org/10.1007/s00425-009-1040-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-009-1040-1