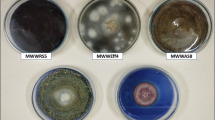
Abstract
Diverse groups of microorganisms have inhabited this earth, which use different types of sources for energy and growth. Industries revolutionize the lifestyle of humankind, which affects negatively the ecosystem. Synthetic dyes impart fabulous colors to cloth, food, paper, and cosmetics. Due to their xenobiotic nature, they are mostly insurmountable for degradation and also toxic. Most of them are washed off during the various processes and mixed in the industrial effluents. Microorganisms have enzymatic system for the decolorization of dyes or simply they can adsorb them on their surface. Several genera of algae, bacteria, and fungi have developed a system to use these unwanted compounds in the water. They can also biotransform or degrade them into non-toxic products. Degradation of the dyes depends upon their toxicity and chemical structure and the type of strain used. Some species were found to be efficient against a variety of dyes at a high concentration level. The present review describes the diversity of three genera Chlorella, Pseudomonas, and Aspergillus of thallophytes for the degradation and decolorization of various dyes in industrial effluents and also the use of integrated approach of different consortia or other treatments for their application in wastewater treatment plants.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
13.1 Introduction
Dyes are synthetic or natural compounds used to color or change the shade of any substance. From the beginning natural dyes from plants were used, but the invention of synthetic dyes by the British chemist William Perkin (1856) from coal tar revolutionized the chemical industry. During the next few decades, production of synthetic dyes has been popularized due to their use in every sector of industries. Dyes are used in food products, paper and textile industry, tanning, cosmetics, pharmaceutical, etc. Commercial products use colors to attract the customers. Due to their high usage, they are concentrated in our environment as xenobiotic compounds. The major share of production goes to textile industry which uses more than 10,000 types of dyes, and most are used as excess levels with 1000 tonnes per annum. About 10–25% is lost at some stage in the dyeing process, and approximately 2–20% is discharged as effluents in water and soil (Carmen and Daniel 2012). They are highly toxic, if not disposed properly as most of them are washed off in the effluents of these industries and reach the water and soil bodies. Dyes and by-products cause environmental, esthetic, and health problems. Dyes can be categorized as disperse, basic, acid, direct, and reactive dyes (Asgher 2012). The breakdown of chromophore groups (azo or anthraquinone) from dyes leads to the formation of toxic compounds (Katheresan et al. 2018). They break down in the form of several carcinogenic or mutagenic forms (aromatic compounds, benzidine, naphthalene, etc.) and cause serious health problems in the food chain. With the time, xenobiotic compounds accumulate in Mother Nature and become problematic for every type of organism. They are mostly degraded or adsorbed by microorganisms, but sometimes become recalcitrant in nature because of insolubility, absence of transporting enzymes, and non-accessibility as substrates (Godheja et al. 2016).
The thallophytes are a group of non-mobile organisms which included algae, bacteria, fungi, and lichens. This group of organisms inhabited the earth in almost all types of conditions like hot springs, volcanoes, and Arctic and Antarctic regions. A variety of microorganisms can tolerate these conditions as well as adapt themselves for their survival. The xenobiotics or industrial effluents make the natural water bodies more acidic and also disturb the growth of biota. Some species of the group were found capable of removing the color from industrial effluents by adsorption or biodegradation or biotransformation or mineralization (Chang et al. 2001a). As compared to chemicophysical treatments, biological degradation of dyes is always cost-effective and also can remove the toxic amines in the effluents, and further the combination of both treatments can produce better results (Hai et al. 2007). The exploration of the diversity and deciphering the underlying mechanism of adaptability will be helpful to make the positive planning to transform the worst environmental conditions (Rampelotto 2010). In the present chapter, we have summarized three different genera, Chlorella (algae), Pseudomonas (bacteria), and Aspergillus (fungi), implicated in the natural degradation of dyes in industrial effluents and the underlying mechanism of decolorization.
13.2 Algae
Algae are a group of aquatic microorganisms having photosynthetic machinery and ca. 50,000 species adapted to various ecological conditions (Xu et al. 2006). They come under the group of thallophytes as due to undifferentiated roots, stems, and leaves. The major commercially available groups of microalga are Chlorophyta, Dinophyta, Haptophyta, Rhodophyta, and Stramenopiles (Heimann and Huerlimann 2015). The microalgal genera studied for the biotreatment of industrial wastewater are Spirogyra, Oscillatoria, Spirulina, Scenedesmus, Cosmarium, etc. (Fazal et al. 2018) Among these groups, Chlorella taxa have been majorly investigated for the treatment of various types of industrial effluents (Banat et al. 1997; Munoza and Guieysse 2006; Safi et al. 2014).
13.2.1 Chlorella
The genus is spherical shaped single cell green algae. It is widely used in the field of productions of biofuels, cosmetics, food, and pigments and wastewater treatments (de Andrade and de Andrade 2017). Industrial wastewater contains dyes and nutrients used by algal community for their growth, which can be used as a sustainable approach for biodiesel production and bioremediation (Fazal et al. 2018). The two species, i.e., C. vulgaris and C. pyrenoidosa, were well documented by various authors for the treatment of effluents of textile industry (Table 13.1).
The first report of degradation of azo dyes by Chlorella was given by Jinqi and Houtian (1992). They tested 30 azo compounds for the decolorization process and found removal percentage in the range of 5–100%. The most easily degradable dye was Direct Blue 71 (100%), and Methyl Red was not decolorized from the medium. The azoreductase enzyme was found to be responsible for the bioconversion of aniline intermediate into carbon dioxide. The same type of degradation product was confirmed by Acuner and Dilek (2004) while studying C. vulgaris for the decolorization of Tectilon Yellow 2G. Sinha et al. (2016) reported the degradation of many industrial pollutants by C. pyrenoidosa NCIM 2738-based photobioreactor. The organism was able to decolorize the dye completely within 2.16 days and also improved the water quality.
The dyes can be degraded into simpler products, or simply they can be adsorbed by the microalgae. Adsorption capacity of microalgae can vary for different dyes and their initial concentration (Aksu and Tezer 2005). The initial pH of the solution was a determining factor for the proper biosorption of the dyes, and it can also vary with the specific dyes. Aksu and Tezer (2005) found that the highest uptake of vinyl sulphone-type reactive dyes occurred at pH 2.0 by dried C. vulgaris, while Daneshvar et al. (2007) demonstrated that basic pH was more favorable for the decolorization of Malachite Green. Similar results were observed by Tsai and Chen (2010) by altering the pH from 3.0 to 11.0. To attain the highest uptake of cationic dyes, the surface should acquire more negative charge which is only possible at this pH. The functional groups, i.e., hydroxyl and carbonyl groups, present on the surface of microalgae help them for the biosorption of dyes (Horník et al. 2013). The optimal temperature range for the dye uptake by Chlorella lies between 25 and 35 °C; however, a wide range has little effect on the biosorption (Tsai and Chen 2010).
The continuous lighting conditions used in the case of mixed culture of algae (13 taxa including Chlorella) removed 80% color within 30 days as compared to 60% after 60 days of exposure under simulated field lighting conditions from the pulping effluent (Dilek et al. 1999). El-Sheekh et al. (2009) tested C. vulgaris among five taxa of microalgae for the removal of basic fuchsin, basic cationic, G-Red, Methyl Red, and Orange II. The most susceptible dyes were basic cationic and basic fuchsin. C. vulgaris removed 43.7 and 59.12% of Orange II and G-Red dyes. The G-Red dye acts as an inducer of the azoreductase enzyme and increases the activity up to 72.25%. Kousha et al. (2013) compared the biosorption activity for Malachite Green of the same species against Scenedesmus quadricauda. They considered the different parameters like dye concentration, contact time, algae amount, and pH. The maximum dye removal was done by C. vulgaris (91.61%) as compared to the latter one (73.49%). Similarly, Lebron et al. (2018) recorded maximum elimination of Methylene Blue by C. vulgaris (98.20%) as compared to Spirulina maxima (94.19%). Recently, Zhao et al. (2018) evaluated the effectiveness of wastewater treatment by C. vulgaris, C. zofingiensis, and Scenedesmus sp. in terms of the activity of photosystem II, nutrient loading, and lipid productivity. C. zofingiensis shows higher absorption capability, productivity, and efficiency as compared to the other two species, even in worse environmental conditions.
The immobilized form of microalgae has more advantages over the free cell suspension for the elimination of heavy metals and xenobiotics in wastewater (Luan et al. 2006). Chu et al. (2009) investigated the immobilized C. vulgaris UMACC 001 (1% κ-carrageenan and 2% sodium alginate) for the treatment of three dyes and textile wastewater. The algae immobilized on 2% sodium alginate has higher color removal efficiency for the textile wastewater and dyes. The immobilized form is more stable, easy to harvest, and protected from the direct exposure to toxicity as compared to free cells. Later, Gao et al. (2011) also found the same results for the removal of nonylphenol using the same type of matrix. Horník et al. (2013) investigated the biosorption capacity of dried biomass of C. pyrenoidosa immobilized in polyurethane foam. The process of sorption of cationic dyes (Thioflavin T and Malachite Green) depends upon the preliminary concentration of dyes, flow rate of solution through the column, bed height, and biomass concentration. The simple or modified polyurethane-based adsorbent has been reported as an efficient sorbent for the elimination of dyes from wastewater (Sultan 2017).
Apart from the treatment of dyes, the genus has been also directly tested for the exclusion of xenobiotics directly from the textile wastewater. The organism utilizes textile wastewater for its growth and also removes the color in the range of 41.8–50.0% as reported by Lim et al. (2010). It also reduces phosphate, nitrate content, BOD, and COD from the effluents. The dried biomass was found more efficient as a biosorbent than wet algal biomass, due to its high binding affinity and large surface area. It can be cultured in the wastewater for color and COD removal and biomass production (El-Kassas and Mohamed 2014; Pathak et al. 2015; Tao et al. 2017). The integrated approach for the treatment of wastewater and production of biomass, lipids, biofuels, bioelectricity, etc. is the promising application of Chlorella in the industry (Logroño et al. 2017; Wang et al. 2017; Fazal et al. 2018). Malla et al. (2015) tested C. minutissima for biodiesel production and nutrient removal from primary and tertiary treated wastewater. The species removed TDS (90–98%), N (70–80%), P (60–70%), and K (45–50%) from the wastewater within 12 days. Zheng et al. (2017) demonstrated the enhanced production of biofuel by using kelp waste extracts combined with acetate in C. sorokiniana.
Seo et al. (2015) used oxidized dye wastewater composed of Methylene Blue and Methyl Orange for the harvesting of algae. The exposed amine groups of oxidized dyes act as amine-based coagulants. Daneshvar et al. (2018) investigated the feasibility of cultivation of C. vulgaris in a combination of aquaculture and pulp effluents. The carbohydrate, lipid, and protein percentage was very much high in the microalgae from the wastewater as compared to Bold’s Basal Medium (BBM) solution. Another aspect of the use of microalgae and textile dyeing sludge was proved by Peng et al. (2015), as the combination of the duo improved char catalytic effect and increased the combustion process for the decomposition of textile dyeing sludge residue at high temperature (530–800 °C).
Undoubtedly, the discharge of the dyes into the aquatic ecosystem causes serious threats for the growth of many microorganisms. Toxicity studies of many dyes on Chlorella have been done by many workers (Hanan 2008; Qian et al. 2008; Hernández-Zamora et al. 2014; Kanhere et al. 2014; Xu et al. 2015). The deteriorated metabolic activity, growth rate, respiration, and photosynthesis efficiency of C. vulgaris were observed due to the direct exposure of Congo Red (Hernández-Zamora et al. 2014). After the bioremoval of the effluents by the species, the influents were less toxic to the primary consumer (Daphnia magna) of the aquatic ecosystem (Hernández-Zamora et al. 2015). Kanhere et al. (2014) observed genotoxic and cytotoxic effects of Malachite Green on C. pyrenoidosa in the form of altered cell morphology, high oxidative stress, DNA damage, and cell death. The growth was inhibited in a dosage-dependent manner, and D. magna ingest the dye even at very low concentrations. Thus, there would be the same type of negative effects on the other aquatic organisms.
13.3 Bacteria
The prominent genera of bacteria explored by different workers are Aeromonas, Bacillus, Escherichia, Eubacterium, Citrobacter, Pseudomonas, Sphingomonas, and Staphylococcus (Rafii et al. 1990; Bumpus 1995; Banat et al. 1997; Keck et al. 1997; Sugiura et al. 1999; Nakanishi et al. 2001; Coughlin et al. 2003). Several anaerobic bacteria produce azoreductase for the degradation of dyes and produced metabolites. Biochemical and molecular characterization has shown that the enzyme presumably a flavin reductase or FMN-dependent NADH-azoreductase or tetrameric NADPH-dependent flavoprotein, as found from Sphingomonas, Escherichia, and Staphylococcus, respectively (Nakanishi et al. 2001; Suzuki et al. 2001; Chen et al. 2005). Bacteria can degrade the xenobiotic compounds in either aerobic or anaerobic or both conditions. Many strains of Pseudomonas have degraded them into non-hazardous products and simultaneously utilized the dyes for their growth (Pandey and Upadhyay 2006). The next section of the chapter reviews the diversity of different species/strains of Pseudomonas capable of degrading dyes in industrial effluents (Table 13.2).
13.3.1 Pseudomonas
Several workers have isolated the azoreductase enzyme from different species of bacteria implicated in the deterioration of azo dyes (Michaels and Lewis 1985; Zhipei and Huifang 1991; Yatome et al. 1990; Hu 1994; Bumpus 1995; Banat et al. 1997). The bacteria utilize them as a source of carbon and nitrogen. However, in the case of RP2B dye, it only acts as an inducer rather than as a growth substrate in the case of P. luteola (Hu 1998). The enzyme was found to be substrate specific, and the susceptibility of the bacterial attack depends on the substitution of the chemical and charged group at specific positions (Zimmermann et al. 1982; Yatome et al. 1990; Ben Mansour et al. 2009a). The degradation reaction of azo dyes into aromatic amines was fully catalyzed by the enzyme under anaerobic conditions, but to produce complete inorganic compounds, aerobic conditions are needed (Zhipei and Huifang 1991; Idaka et al. 1987a, b).
Zimmermann et al. (1982) isolated oxygen-insensitive azoreductase from Pseudomonas KF46, able to degrade the aromatic amines and complete mineralization of carboxy-Orange II. Nachiyar and Rajkumar (2004, 2005) proposed the mechanism of systematic elimination of Navitan Fast Blue S5R by the oxygen-insensitive enzyme, purified from P. aeruginosa. The intermediate metabolites of the dye may have undergone further oxidative deamination/decarboxylation and further enter the TCA cycle to release carbon dioxide. One of the intermediates formed in this study, i.e., metalinic acid, was further degraded into aniline and β-ketoadipic acid (Nachiyar et al. 2007). Işik and Sponza (2003) used aerobic and anaerobic conditions to study the color removal efficiency of Pseudomonas sp. They found that decolorization of Direct Black 38 and Congo Red was 83% and 100% under anaerobic incubation while 74% and 76% under microaerophilic conditions. The aerobic degradation occurs by the action of lignin peroxidase, tyrosinase, and laccase as reported by Kalme et al. (2007b) in P. desmolyticum NCIM 2112. Further, they purified laccase enzyme from the species and demonstrated the asymmetric breakdown of azo bond and that the specificity depends on the position of amino, hydroxyl, and sulfonic group in a dye. The decolorization rate is less when hydroxyl group and sulfonic group are at meta position or charged carboxyl group at ortho position to the azo bond (Nigam et al. 1996; Chen 2006; Kalme et al. 2007b, 2009). The presence of electron-withdrawing groups or absence of charged groups also enhances the rate of decolorization as stated by Hsueh and Chen (2007, 2008) in P. luteola. The toxicity of dyes depends on the type of azo bond, molecular structure, functional groups, and types of intermediates or degraded products. The lesser the toxicity of the dye, the easier will be the decolorization. Chen (2002) tested the toxicity of three reactive dyes against P. luteola (Acid Yellow, Black B, and Red 22). The Reactive Red 22 was easily decolorized, while Reactive Black B was highly toxic as it contains two azo bonds. As in this study decolorization is not growth-associated, the viability of the cells is the important criterion for the metabolism and expression of enzymes. Alternatively the cells can go for biosorption rather than decolorization.
Various authors have also isolated the laccase enzyme from different strains/species of Pseudomonas and showed its applicability in the elimination of synthetic dyes in industrial effluents (Telke et al. 2009; Kuddus et al. 2013; Wang et al. 2012). Phugare et al. (2011) purified a highly active enzyme, i.e., veratryl alcohol oxidase, from P. aeruginosa BCH. The enzyme has specificity for wide varieties of substrates and decolorizes seven dyes (Methyl Orange, Rubine 3GP, Congo Red, Remazol Black, Red HE7B, Red HE8B, and Red HE3B) in the range of 85–100%. One of the dyes, i.e., Remazol Black, was decolorized completely within 6 h and degraded into 7-diazenyl-naphathalene-1-ol and naphthalene-1,2,7-triol. Kalyani et al. (2011) reported a heme-containing peroxidase enzyme isolated from Pseudomonas sp. for the symmetric cleavage of Methyl Orange into N,N-dimethyl-1,4-benzenediamine and an intermediate 4-aminobenzenesulfonic acid. The intermediate formed was further degraded into aniline.
Toxicity analysis of the decolorized dyes should be done either by elucidating the structure of the degraded products by FTIR, GC-MS, HPLC, and NMR techniques or by using different organisms or cell lines. Several authors have checked the genotoxicity/cytoxicity/mutagenic potential of the metabolites formed by Pseudomonas during the remediation of industrial effluents (Adedayo et al. 2004; Pandey and Upadhyay 2006; Kalme et al. 2007a; Kalyani et al. 2009). Perei et al. (2001) isolated an aerobic bacterium called P. paucimobilis from the contaminated sites for the effective degradation of mutagenic metabolite sulfanilic acid. During the degradation of Orange 52, Violet 7, and Acid Yellow 17 by P. putida mt-2, genotoxic metabolites were found high in static cultures as compared to shaken conditions (Ben Mansour et al. 2007). Later on the authors demonstrated that the amines were mutagenic formed under static conditions, which later on vanished during shaken incubation. Further, the metabolite 4′-aminoacetanilide exhibited maximum mutagenicity, while 5-acetamido-2-amino-1-hydroxy-3,6-naphthalene disulfonic acid shows less effect due to presence of sulfonic groups (Ben Mansour et al. 2009b). Telke et al. (2012) tested the toxicity assays of p-dihydroperoxybenzene, 2-hydroxy-7-aminonaphthol-3-sulfonic acid, and 3,6-dihydroxy benzoic acid, metabolites formed during biodegradation of Direct Brown MR by Pseudomonas sp. LBC1. The textile effluents and the dye were more toxic to Vigna radiata and Sorghum bicolor as compared to the biodegraded metabolites.
In the case of Methyl Orange, there wasn’t any kind of removal under aerobic conditions by P. putida mt-2 (Thao et al. 2013). So an immobilized bacterial system can solve the problem for oxygen-sensitive decolorization by creating miniature anoxic environment and complementarily increasing the biomass concentration and providing mechanical strength, feasibility of continuous processing, low-cost recovery, and reusability of biocatalyst (Stormo and Crawford 1992; Park and Chang 2000; Chang et al. 2001a). Puvaneshwari et al. (2002) studied the effective role of immobilized P. fluorescens on sodium alginate for the degradation of Direct Blue (71%) and Direct Red (82%). Chen and Lin (2007) used silicate/alginate sol-gel beads of P. luteola for the decolorization of Reactive Red 22. The rate of decolorization of the free cells decreased, while the immobilized system was static after five repeated batch cycles. Tuttolomondo et al. (2014) reported the biodegradation of Methyl Orange, Benzyl Orange, and Remazol Black by immobilized Pseudomonas sp. in sol-gel silica matrices due to higher expression of extracellular enzymes. The encapsulation directly protects the bacteria from toxic conditions and consecutively increases the production of enzymes involved in degradation. Pseudomonas sp. DY1 immobilized in the fungi (A. oryzae) cellular mass shows 96% decolorization in the batch cycle, still after 16 days. Inhibition test confirmed that the activity of the pellets was mainly due to the bacteria, demonstrating their stable and long-term usability for the dye treatment (Yang et al. 2011a, b). Recently, Roy et al. (2018) used immobilized Pseudomonas sp. in fly ash for the biodegradation of Reactive Yellow. The highest removal percentage (98.72%) was recorded in Pseudomonas sp. on fly ash as compared to sorption by fly ash (88.51%) and degradation by species (92.62%).
The activated carbon in combination with P. luteola was found to be very much effective for the adsorption and biodegradation of Reactive Red 22 (Lin and Leu 2008). Selvakumar et al. (2010) use electro-oxidation and bio-oxidation by P. aeruginosa for the removal of color from textile effluent having Procion Blue 2G dye. Later the treated effluents have been treated with photo-oxidation to remove the bacteria, so that water can be recycled. Similarly, Srinivasan et al. (2011) combined the sonolysis pretreatment with post-biological treatment by the mutant strain of P. putida in the case of Tectilon Yellow 2G.
The studies on the optimization of the conditions like temperature, pH, presence of organic compounds, carbon and nitrogen source, concentration range of dyes, and aerobic or anaerobic or both conditions are very much necessary, depending on the nature of the dye to be treated by Pseudomonas. Yu et al. (2001) observed that presence of nitrate at concentration 1000 mg/L inhibits the process completely, while increase in the temperature from 10 to 35 °C enhances the decolorization rate of Pseudomonas strain GM3. Chang et al. (2001b) found that tryptone and yeast extract enhances the decolorization process of Reactive Red 22, while retarded by the added glucose concentration and dissolved oxygen. The activity of azoreductase enzyme isolated from cell-free extract also depends upon the growth phase of bacteria. Lodato et al. (2007) proved that depletion of dye can be achieved irrespective of the initial concentration by changing the aerobic-anaerobic operating conditions. In the aerobic conditions, growth of Pseudomonas sp. OX1 can be achieved, while in the anaerobic conditions, depletion of dye takes place. Similarly, Lin et al. (2010) observed complete mineralization of Reactive Blue 13 by Pseudomonas sp. L1 in the same conditions. Joe et al. (2011) investigated the optimal conditions needed for Remazol Black B dye by P. aeruginosa CR-25. The maximum rate of removal occurs at 37 °C, pH7 with supplementation of peptone, yeast extract, glucose and fructose as nitrogen and carbon sources under static conditions. The same results have been observed under the above-said conditions by other workers using different species of Pseudomonas (Kalyani et al. 2008; Telke et al. 2009; Thao et al. 2013). Kumar Garg et al. (2012) showed that supplementation of ammonium sulfate (0.1%, w/v) and glucose (0.4% w/v) improved the decolorization of Orange II. Mishra and Maiti (2018) demonstrated that yeast extract has positive effect, while peptone and glucose have negative effect on the decolorization of Reactive Red 21 by P. aeruginosa 23N1. This may be due to the fact that species must have utilized peptone and glucose as primary sources of nitrogen and carbon rather than the dye molecule. Recently, Hashem et al. (2018) isolated a pH-tolerant P. aeruginosa KY284155 with high decolorization rate for Remazol Black B. With the addition of iron, magnesium, and yeast extract in the medium, the degradation rate was further accelerated. The heavy metals and salts at high concentrations in the medium have inhibitory effects on the decolorization of dyes (Gopinath et al. 2011). Some strains of P. aeruginosa were very effective in the degradation of reactive azo dyes even in the presence of heavy metals like lead, zinc, cadmium, and chromium (Maqbool et al. 2016; Hafeez et al. 2018).
The majority of the studies done in Pseudomonas were related to biodegradation of the dyes, but few authors have also studied the adsorption phenomena for the management of industrial effluents. Du et al. (2012) compared the adsorption capacity of live and heat-treated Pseudomonas sp. strain DY1 biomass for Acid Black 172. The heat-treated cells have high adsorption due to increased permeability and denatured intracellular proteins. Deepa et al. (2013) showed that 4 to 9 pH and 1 to 1000 mM NaCl concentrations have insignificant effect on the adsorption rate of Direct Red by P. putida. Later on, Arunarani et al. (2013) proved the same type of effect on the adsorption of Acid Blue 93 and Basic Violet 3 by the same taxa due to pH and salts. Liu et al. (2017) extracted a biosurfactant from P. taiwanensis L1011 and utilized it to accelerate the chemical and biological decolorization of Congo Red and Amaranth, respectively. Recently, Iqbal et al. (2018) developed a novel biosorbent using P. aeruginosa USM-AR2 cells immobilized on mesoporous rice husk ash silica (RHA-SiO2).
There is a lot of variability for the potential of degradation of dyes within the different genera of bacteria. Hu (1996) compared the adsorption efficiency of Aeromonas, Bacillus, Escherichia, Pseudomonas, and Staphylococcus for four reactive azo dyes. The dead biomass of the three genera exhibits higher adsorption capacity in the order of Aeromonas > Pseudomonas > Escherichia. Nachiyar and Rajkumar (2003) tested three species (P. aeruginosa, P. fluorescens, and P. putida) for the decolorization of Navitan Fast Blue S5R and found that P. aeruginosa exhibited maximum efficiency (72–92%) within 72 h. Silveira et al. (2009) compared 4 species (P. oleovorans, P. putida, P. cepacia, and P. aeruginosa) for the efficiency of decolorization of 14 commercial textile dyes. Among them, P. aeruginosa and P. oleovorans were more capable to decolorize ten textile dyes. The mixed consortia of Pseudomonas, Acinetobacter, Escherichia, Enterobacter, Aspergillus, and Actinobacteria were also found to significantly decolorize or degrade different kinds of azo dyes (Kadam et al. 2011; Yang et al. 2011a, b; Patel et al. 2012; Khan et al. 2014; Isaac et al. 2015; Kuppusamy et al. 2017; Sathishkumar et al. 2017).
Pseudomonas genus was also studied for the biotreatment of triphenylmethane dyes, used extensively as biological or dermatological agent, and in various processes in the food, medical, and textile industry (Sarnaik and Kanekar 1995, 1999; Yatome et al. 1981, 1990; Lin et al. 2004; Wu et al. 2009). Malachite Green and Crystal Violet dyes were extensively studied by several researchers (El-Naggar et al. 2004; Chen et al. 2007; Li et al. 2009; Huan et al. 2010; Kalyani et al. 2012; Chaturvedi et al. 2013). Enhancement of degradation of triphenylmethane dyes can be attained by adding glucose and sucrose as cosubstrates and heavy metals in the medium (Oranusi and Ogugbue 2005). Kalyani et al. (2012) showed that aminopyrine N-demethylase, MG reductase, and laccase enzymes were induced in P. aeruginosa NCIM 2074 and degraded Malachite Green into a non-toxic product. The same category of enzymes was also found to degrade heavy amounts of the dye (1800 mg/L) in P. mendocina (Chaturvedi et al. 2013). Li et al. (2009) isolated a strain of Pseudomonas sp. MDB-1 from water of an aquatic hatchery, capable of degrading various triphenylmethane dyes. Later on, tmr2 gene encoding the enzyme (triphenylmethane reductase) was also fully characterized responsible for the biodegradation (Huan et al. 2010; Li et al. 2009). Zabłocka-Godlewska et al. (2014) compared SDz3 and Sz6 strains of P. fluorescens for the biodegradation of mixture containing triphenylmethane (Brilliant Green) and azo (Evans Blue) dyes. The strain Sz6 was able to degrade the dyes faster in shaken/semistatic conditions, and maximum removal (95.4%) was achieved in the case of Brilliant Green.
Various species of Pseudomonas were also reported for the removal of other xenobiotic compounds used for the preparation of dyes. The compounds include phenol by P. putida DSM 548, Pseudomonas CF600, and P. stutzeri (Sá and Boaventura 2001; Moharikar and Purohit 2003; Pazarlioğlu and Telefoncu 2005; Nowak and Mrozik 2018; Singh et al. 2018); 4-aminophenol by Pseudomonas ST-4 (Afzal Khan et al. 2006); pyridine by Pseudomonas sp. PI2 (Mohan et al. 2003); naphthalene and p-cresol by P. putida and P. gessardii LZ-E (Huang et al. 2016a, b; Izmalkova et al. 2013; Surkatti and El-Naas 2014); chloroanilines by P. putida T57 (Nitisakulkan et al. 2014); polycyclic aromatic hydrocarbons by P. stutzeri (Álvarez et al. 2015); polynuclear aromatic hydrocarbons by P. plecoglossicida PB1 and Pseudomonas sp. PB2 (Nwinyi et al. 2016); and phenanthrene by P. stutzeri JP1 and P. mendocina NR802 (Mangwani et al. 2014; Kong et al. 2017).
13.4 Fungi
Many genera of fungi were also explored for the color removal from industrial effluents, especially actinomycetes and basidiomycetes (Chivukula and Renganathan 1995; McMullan et al. 2001). These organisms produce extracellular enzymes (laccase, peroxidases, and azoreductase) to catalyze dealkylation, oxidation, and hydroxylation reactions for the metabolism of dyes (Goszczynski et al. 1994). Most of the work was done for white rot fungus (Phanerochaete), as they are capable to degrade the majority of the azo dyes (Bumpus 1995; Banat et al. 1997; Cripps et al. 1990). The other fungal genera reported for the biodegradation of xenobiotic compounds are Streptomyces, Lenzites, Coriolopsis, Neurospora, Penicillium, Pleurotus, Trichoderma, and Trametes (Paszczynski et al. 1992; Chao and Lee 1994; Knapp and Newby 1999; Saparrat et al. 2014; He et al. 2018; Naraian et al. 2018; Pandey et al. 2018). The brown rot fungus (Aspergillus) has also shown potential to biodegrade a variety of toxic xenobiotic compounds and for the biotreatment of wastewater (Ali et al. 2010; Abd El-Rahim et al. 2017; Gomaa et al. 2011). Recently, Ning et al. (2018) reported biodegradation of 15 dyes by Aspergillus flavus A5p1 in a range of 61.7–100.0%. So there is always a need to explore the different strains/species of the Aspergillus for the degradations of the wide varieties of dyes (Table 13.3).
13.4.1 Aspergillus
The genus is composed of 340 species, widespread in diverse habitats, and reported as a pathogen, spoils food materials, and produces mycotoxins (Bennett and Klich 2003; Houbraken et al. 2016). They reproduce by asexual reproduction via conidiophores. The key to identify or classify various species of the genus is based on the size, color, and arrangement of asexual spores of conidiophores. Some species are associated with serious health problems like allergic bronchopulmonary aspergillosis, liver cancer (consumption of food containing mycotoxins), etc. (Hedayati et al. 2007). Most of the species are also used to produce beneficial products (enzymes, food fermenters, antibiotics, etc.) in biotechnology industry (Samson et al. 2014). To mention some of the species with beneficial/harmful effects are A. flavus (aflatoxin), A. fumigatus (cellulose, xylanase), A. niger (homologous or heterologous proteins), A. oryzae, A. sojae (food fermentation), A. tamari (Japanese soya sauces), and A. terreus (lovastatin, terrein) (Park et al. 2017). The present section reviews the diversity found within the Aspergillus species for the elimination of hazardous dyes from the industrial effluents (Table 13.3).
Initial studies for the wastewater treatment were mainly focused on the white rot fungus group, as they have lignin-degrading enzymes for the oxidation of organic compounds (Bumpus and Aust 1987). Aspergillus genus (brown rot fungi) was also explored for the removal of dyes in the industrial effluents. Ryu and Weon (1992) analyzed four species of Aspergillus (six strains) and one species of Phanerochaete (two strains) for the biodegradation of three azo dyes and stated that the former genus was much more effective in the process. Mainly two processes for the treatment of dyes in the solution or synthetic effluents were studied extensively, either biosorption or biodegradation (Conatao and Corso 1996; Fu and Viraraghavan 2000, 2002a; Sumathi and Manju 2000; Zope et al. 2007; Esmaeili and Kalantari 2011; Almeida and Corso 2014). The biosorption of dyes was influenced by their chemical structure and functional group on the surface of fungus (Fu and Viraraghavan 2002b, 2003). Parshetti et al. (2007) observed faster adsorption rate in A. ochraceus in the shaking conditions. The treatment of Aspergillus species with immobilization beads, autoclaving, and specific compounds also accelerates the process of decolorization (Wang and Hu 2007; Wang et al. 2008; Patel and Suresh 2008). Yang et al. (2011a, b) demonstrated higher biosorption capacity in the CDAB (cetyldimethylammonium bromide) modified biomass of A. oryzae. The same type of result was seen by Huang et al. (2016a, b) while investigating the effect of heavy salts, metals, and SDS on the adsorption kinetics of chemically modified (cetyltrimethylammonium bromide) A. versicolor. They found a close relationship between low pH (2.0) and heavy metals on the biosorption rate. The chemical modification increases the surface area and functional groups. Naskar and Majumder (2017) used response surface methodology for A. niger and demonstrated that adsorption rate depends upon the concentration of biomass, temperature, and pH of the solution. Further, they also revealed that amine and carboxyl groups play an important role in dye sorption along with electrostatic interactions. The same type of phenomena was observed by the authors using different dyes and the same species (Xiong et al. 2010; Mahmoud et al. 2017). The high temperature and low pH range (1–3) in the solution speed up the uptake of the dyes, as the biosorption is mostly endothermic (Akar et al. 2009). This type of condition increases the kinetic energy and diffusion rate (Ramya et al. 2007; Aksu and Karabayır 2008; Abdallah and Taha 2012). Contradictory to this, other authors reported optimal temperature (28–30 °C) and pH (5) as much more favorable condition for the biodegradation of azo dyes (Ali et al. 2007a, b; Ameen and Alshehrei 2017; Sharma et al. 2009) by four Aspergillus spp. The nutritional condition needs to be standardized as sources of nitrogen and carbon in the medium, as they are also a detrimental factor for the rate of dye removal (Kaushik and Malik 2010, 2011). Gomaa et al. (2017) demonstrated the role of calcium chloride as stress response in A. niger and high removal efficiency for commercial dye Malachite Green.
The live fungal strains were extensively studied for the decolorization of dyes from industrial effluents; however, some workers used pellets and dead biomass for the process and found promising results as compared to the living strains (Abdallah and Taha 2012; Abdel Ghany and Al Abboud 2014; Lu et al. 2017). The formation of bioflocculants and silver and zinc oxide nanoparticles using different Aspergillus spp. has also the potential for the color removal from industrial effluents (Deng et al. 2005; Muthu Kumara Pandian et al. 2016; Kalpana et al. 2018a, b). Copete-Pertuz et al. (2019) demonstrated that A. terreus in combination with Trichoderma viride can act as a co-inducer for Leptosphaerulina sp. ligninolytic enzyme activity and improved removal of Reactive Black 5 dye.
Survey of literature reveals that most of the studies were related to the biosorption mechanism rather than the degradation. The metabolites formed during degradation process are shown in Table 13.3. The enzymes involved in the biodegradation were laccase, manganese peroxides, and lignin-modifying enzymes, which mineralize synthetic lignin of dyes (Ali and El-Mohamedy 2012; Hasanin et al. 2019). Azoreductase is one of the key enzymes found in the degradation pathways of the organism. Ameen and Alshehrei (2017) found laccase and azoreductase to be involved in the degradation of Reactive Red 120 into sodium 2-aminobenzenesulfonate. Tamayo-Ramos et al. (2012) characterized three forms of laccase-like multicopper oxidase enzymes having high catalytic activity for several phenolic compounds and synthetic dyes. The optimization process for the high production and activity of laccase enzyme has been done for several Aspergillus species. The factors associated are pH, temperature, carbon and nitrogen sources, inoculum size, etc. (Jin and Ning 2013; Benghazi et al. 2013; Kumar et al. 2016). Recently, Abd El-Rahim et al. (2017) isolated 18 strains belonging to 6 species from the wastewater sample and evaluated them against 20 azo dyes. The most resistant dye was Fast Green azo dye, and easily degradable dyes were Direct Violet and Methyl Red. The decolorization process was enhanced by glucose supplementation, and the limiting factor was a nitrogen source, as in its absence the strains were unable to produce lignin peroxidase enzyme. The high pH has been also shown to be related to the low formation of residual products (Ali et al. 2007a, b).
The different Aspergillus species have shown very much diversity in the biodegradation of various dyes. Anastasi et al. (2009) compared five species of mitosporic fungi (Penicillium, Cladosporium, and Aspergillus) for the removal of nine industrial and two model dyes. They found that A. ochraceus and A. flavus were efficient for the decolorization of all the dyes tested and one species, i.e., A. ochraceus, causes over 90% decolorization against simulated effluents. Similarly, other workers found the maximum potential of Aspergillus as compared to Penicillium (Ali et al. 2010; Gomaa et al. 2011; Ali and El-Mohamedy 2012). Khalaf (2008) tested the effectiveness of Spirogyra sp. (green algae) and A. niger against the reactive dye (Synozol) in textile wastewater. The autoclaved biomass of the both species exhibited 88% and 85% dye removal, respectively. Some species have higher absorption capacity, but still they lack the ability to degrade them into non-toxic metabolites (Almeida and Corso 2014).
The degraded products should be checked for the toxicity assays, as decolorization does not always lead to the absence of toxicity, rather forming incomplete toxic metabolites (Almeida and Corso 2014). The extracellular enzymes were found to degrade triphenylmethane dye by stepwise demethylation into non-toxic N-demethylated products (Kumar et al. 2011, 2012). Andleeb et al. (2012) investigated the toxicity of degraded products formed during biodegradation of Drimarene Blue dye by A. flavus. As compared to dye treatment, the germination and morphological characteristics in Lolium perenne were somewhat near to the untreated. Similarly, Parshetti et al. (2007) observed that germination of Phaseolus mungo was high or near to control in comparison to the Malachite Green treatment.
13.5 Conclusion
The treatment of industrial effluents with cost-effective methods is the urgent need of the society. The literature shows that aerobic and anaerobic conditions were well utilized by algae, bacteria, and fungi for the management of dyes. The effluents also serve as a growth substrate or also can be used to extract biomass. The integrated approach of remediation as successive treatment along with extraction of enzymes, lipids, and biofuels seems to be the best practice for sustainable development. The mixed consortium of best strains of algae, bacteria, and fungi should be tested for the degradation of toxic dyes. Genetically engineered strains may be used for the degradation of toxic amines in the severe environmental conditions. Toxicity assays clearly show which strain is best for the future applications to clear the water for recycling.
References
Abd El-Rahim WM, Moawad H, Abdel Azeiz AZ et al (2017) Optimization of conditions for decolorization of azo-based textile dyes by multiple fungal species. J Biotechnol 260:11–17
Abdallah R, Taha S (2012) Biosorption of methylene blue from aqueous solution by nonviable Aspergillus fumigatus. Chem Eng J 195-196:69–76
Abdel Ghany TM, Al Abboud MA (2014) Capacity of growing, live and dead fungal biomass for safranin dye decolourization and their impact on fungal metabolites. Aus J Basic Appl Sci 8:489–499
Acuner E, Dilek F (2004) Treatment of tectilon yellow 2G by Chlorella vulgaris. Process Biochem 39:623–631
Adedayo O, Javadpour S, Taylor C et al (2004) Decolourization and detoxification of methyl red by aerobic bacteria from a wastewater treatment plant. World J Microbiol Biotechnol 20:545–550
Afzal Khan S, Hamayun M, Ahmed S (2006) Degradation of 4-aminophenol by newly isolated Pseudomonas sp. strain ST-4. Enzym Microb Technol 38:10–13
Akar ST, Akar T, Cabu A (2009) Decolorization of a textile dye, RR198 by Aspergillus parasiticus fungal biosorbent. J Chem Eng 2:399–405
Aksu Z, Karabayır G (2008) Comparison of biosorption properties of different kinds of fungi for the removal of Gryfalan Black RL metal-complex dye. Bioresour Technol 99:7730–7741
Aksu Z, Tezer S (2005) Biosorption of reactive dyes on the green alga Chlorella vulgaris. Process Biochem 40:1347–1361
Ali NF, El-Mohamedy RSR (2012) Microbial decolourization of textile waste water. J Saudi Chem Soc 16(2):117–123
Ali N, Hameed A, Ahmed S, Khan AG (2007a) Decolorization of structurally different textile dyes by Aspergillus niger SA1. World J Microbiol Biotechnol 24(7):1067–1072
Ali N, Ikramullah, Lutfullah G et al (2007b) Decolorization of acid red 151 by Aspergillus niger SA1 under different physicochemical conditions. World J Microbiol Biotechnol 24:1099–1105
Ali N, Hameed A, Ahmed S (2010) Role of brown-rot fungi in the bioremoval of azo dyes under different conditions. J Microbiol 4:907–915
Almeida EJR, Corso CR (2014) Comparative study of toxicity of azo dye Procion Red MX-5B following biosorption and biodegradation treatments with the fungi Aspergillus niger and Aspergillus terreus. Chemosphere 112:317–322
Álvarez MS, Rodríguez A, Sanromán MÁ et al (2015) Simultaneous biotreatment of polycyclic aromatic hydrocarbons and dyes in a one-step bioreaction by an acclimated Pseudomonas strain. Bioresour Technol 198:181–188
Ameen F, Alshehrei F (2017) Biodegradation optimization and metabolite elucidation of Reactive Red 120 by four different Aspergillus species isolated from soil contaminated with industrial effluent. Ann Microbiol 67:303–312
Anastasi A, Prigione V, Casieri L et al (2009) Decolourisation of model and industrial dyes by mitosporic fungi in different culture conditions. World J Microbiol Biotechnol 25:1363–1374
Andleeb S, Atiq N, Robson GD, Ahmed S (2012) An investigation of anthraquinone dye biodegradation by immobilized Aspergillus flavus in fluidized bed bioreactor. Environ Sci Pollut Res 19(5):1728–1737
Arunarani A, Chandran P, Ranganathan BV et al (2013) Bioremoval of basic violet 3 and Acid Blue 93 by Pseudomonas putida and its adsorption isotherms and kinetics. Colloids Surf B: Biointerfaces 102:379–384
Asgher M (2012) Biosorption of reactive dyes: a review. Water Air Soil Pollut 223:2417. https://doi.org/10.1007/s11270-011-1034-z
Banat IM, Nigam P, McMullan G et al (1997) The isolation of thermophilic bacterial cultures capable of textile dyes decolorization. Environ Int 23:547–551
Ben Mansour H, Corroler D, Barillier D et al (2007) Evaluation of genotoxicity and pro-oxidant effect of the azo dyes: acids yellow 17, violet 7 and orange 52, and of their degradation products by Pseudomonas putida mt-2. Food Chem Toxicol 45:1670–1677
Ben Mansour H, Corroler D, Barillier D et al (2009a) Influence of the chemical structure on the biodegradability of acids yellow 17, violet 7 and orange 52 by Pseudomonas putida. Ann Microbiol 59:9–15
Ben Mansour H, Mosrati R, Corroler D et al (2009b) In vitro mutagenicity of Acid Violet 7 and its degradation products by Pseudomonas putida mt-2: correlation with chemical structures. Environ Toxicol Pharmacol 27:231–236
Benghazi L, Record E, Suárez A, Gomez-Vidal JA, Martínez J, de la Rubia T (2013) Production of the Phanerochaete flavido-alba laccase in Aspergillus niger for synthetic dyes decolorization and biotransformation. World J Microbiol Biotechnol 30(1):201–211
Bennett JW, Klich M (2003) Mycotoxins. Clin Microbiol Rev 16:497–516
Bidisha C, Sreeranjani R, Shaik A et al (2006) Bioaccumulation and biosorption of drimarene red dye by Aspergillus foetidus. Int J Environ Pollut 28:517–533
Bouras HD, Yeddou AR, Bouras N (2017) Biosorption of Congo red dye by Aspergillus carbonarius M333 and Penicillium glabrum Pg1: kinetics, equilibrium and thermodynamic studies. J Taiwan Inst Chem E 80:915–923
Bumpus JA (1995) Microbial degradation of azo dyes. In: Singh VP (ed) Biotransformations: microbial degradation of health risk compounds. Elsevier Science, Amsterdam, pp 157–176
Bumpus JA, Aust SD (1987) Biodegradation of environmental pollutants by the white rot fungus Phanerochaete chrysosporium: involvement of the lignin degrading system. Bio Essays 6:166–170
Carmen Z, Daniel S (2012) Textile organic dyes—characteristics, polluting effects and separation/elimination procedures from industrial effluents—a critical overview, organic pollutants ten years after the Stockholm convention, Tomasz Puzyn and Aleksandra Mostrag-Szlichtyng, IntechOpen, doi: 10.5772/32373. Available from: https://www.intechopen.com/books/organic-pollutants-ten-years-after-the-stockholm-convention-environmental-and-analytical-update/textile-organic-dyes-characteristics-polluting-effects-and-separation-elimination-procedures-from-in
Chang JS, Chou C, Chen SY (2001a) Decolorization of azo dyes with immobilized Pseudomonas luteola. Process Biochem 36:757–763
Chang JS, Chou C, Lin YC et al (2001b) Kinetic characteristics of bacterial azo-dye decolorization by Pseudomonas luteola. Water Res 35:2841–2850
Chao WL, Lee SL (1994) Decoloration of azo dyes by three white rot fungi: influence of carbon source. World J Microbiol Biotechnol 10:556–559
Chaturvedi V, Bhange K, Bhatt R et al (2013) Biodetoxification of high amounts of malachite green by a multifunctional strain of Pseudomonas mendocina and its ability to metabolize dye adsorbed chicken feathers. J Environ Chem Eng 1:1205–1213
Chen BY (2002) Understanding decolorization characteristics of reactive azo dyes by Pseudomonas luteola: toxicity and kinetics. Process Biochem 38:437–446
Chen BY (2006) Toxicity assessment of aromatic amines to Pseudomonas luteola: chemostat pulse technique and dose–response analysis. Process Biochem 41:1529–1538
Chen JP, Lin YS (2007) Decolorization of azo dye by immobilized Pseudomonas luteola entrapped in alginate–silicate sol–gel beads. Process Biochem 42:934–942
Chen H, Hopper SL, Cerniglia CE (2005) Biochemical and molecular characterization of an azoreductase from Staphylococcus aureus, a tetrameric NADPH—dependent flavoprotein. Microbiology 151:1433–1441
Chen CC, Liao HJ, Cheng CY et al (2007) Biodegradation of Crystal Violet by Pseudomonas putida. Biotechnol Lett 29:391–396
Chivukula M, Renganathan V (1995) Phenolic azo dye oxidation by laccase from Pyricularia oryzae. Appl Environ Microbiol 61:4374–4377
Chu WL, See YC, Phang SM (2009) Use of immobilised Chlorella vulgaris for the removal of colour from textile dyes. J Appl Phycol 21:641. https://doi.org/10.1007/s10811-008-9396-3
Conatao M, Corso CR (1996) Studies of adsorptive interaction between Aspergillus niger and the reactive azo dye procion blue MX-G. Ecletica Quim 21:97–102
Copete-Pertuz LS, Alandete-Novoa F, al PJ (2019) Enhancement of ligninolytic enzymes production and decolourising activity in Leptosphaerulina sp. by co–cultivation with Trichoderma viride and Aspergillus terreus. Sci Total Environ 646:1536–1545
Coughlin MF, Kinkle BK, Bishop PL (2003) High performance degradation of azo dye acid orange 7 and sulfanilic acid in a laboratory scale reactor after seeding with cultured bacterial strains. Water Res 37:2757–2763
Cripps C, Bumpus JA, Aust SD (1990) Biodegradation of azo and heterocyclic dyes by Phanerochaete chrysosporium. Appl Environ Microbiol 56:1114–1118
Daneshvar N, Khataee AR, Rasoulifard MH et al (2007) Biodegradation of dye solution containing malachite green: optimization of effective parameters using Taguchi method. J Hazard Mater 143:214–219
Daneshvar E, Antikainen L, Koutra E et al (2018) Investigation on the feasibility of Chlorella vulgaris cultivation in a mixture of pulp and aquaculture effluents: treatment of wastewater and lipid extraction. Bioresour Technol 255:104–110
de Andrade CJ, de Andrade LM (2017) An overview on the application of genus Chlorella in biotechnological processes
Deepa K, Chandran P, Sudheer Khan S (2013) Bioremoval of Direct Red from aqueous solution by Pseudomonas putida and its adsorption isotherms and kinetics. Ecol Eng 58:207–213
Deng S, Yu G, Ting YP (2005) Production of a bioflocculant by Aspergillus parasiticus and its application in dye removal. Colloids Surf B: Biointerfaces 44:179–186
Dilek FB, Taplamacioglu HM, Tarlan E (1999) Colour and AOX removal from pulping effluents by algae. Appl Microbiol Biotechnol 52:585–591
Du LN, Yang YY, Li G et al (2010) Optimization of heavy metal-containing dye Acid Black 172 decolorization by Pseudomonas sp. DY1 using statistical designs. Int Biodeterior Biodegrad 64:566–573
Du LN, Wang B, Li G et al (2012) Biosorption of the metal-complex dye Acid Black 172 by live and heat-treated biomass of Pseudomonas sp. strain DY1: kinetics and sorption mechanisms. J Hazard Mater 205-206:47–54
El-Kassas HY, Mohamed LA (2014) Bioremediation of the textile waste effluent by Chlorella vulgaris. Egypt J Aquat Res 40:301–308
El-Naggar MA, El-Aasar SA, Barakat KI (2004) Bioremediation of crystal violet using air bubble bioreactor packed with Pseudomonas aeruginosa. Water Res 38:4313–4322
El-Sheekh MM, Gharieb MM, Abou-El-Souod GW (2009) Biodegradation of dyes by some green algae and cyanobacteria. Int Biodeterior Biodegradation 63:699–704
Esmaeili A, Kalantari M (2011) Bioremoval of an azo textile dye, Reactive Red 198, by Aspergillus flavus. World J Microbiol Biotechnol 28:1125–1131
Fazal T, Mushtaq A, Rehman F et al (2018) Bioremediation of textile wastewater and successive biodiesel production using microalgae. Renew Sustain Energy Rev 82:3107–3126
Fu YZ, Viraraghavan T (2000) Removal of a dye from aqueous solution by the fungus Aspergillus niger. Water Qual Res J Can 35:95–111
Fu Y, Viraraghavan T (2002a) Removal of Congo Red from an aqueous solution by fungus Aspergillus niger. Adv Environ Res 7:239–247
Fu Y, Viraraghavan T (2002b) Dye biosorption sites in Aspergillus niger. Bioresour Technol 82:139–145
Fu Y, Viraraghavan T (2003) Column studies for biosorption of dyes from aqueous solutions on immobilized Aspergillus niger fungal biomass. Water South Africa 29:465–472
Gao QT, Wong YS, Tam NFY (2011) Removal and biodegradation of nonylphenol by immobilized Chlorella vulgaris. Bioresour Technol 102:10230–10238
Godheja J, Shekhar SK, Siddiqui SA et al (2016) Xenobiotic compounds present in soil and water: a review on remediation strategies. J Environ Anal Toxicol 6:5. https://doi.org/10.4172/2161-0525.1000392
Gomaa OM, Momtaz OA, Kareem HAE et al (2011) Isolation, identification, and biochemical characterization of a brown rot fungus capable of textile dye decolorization. World J Microbiol Biotechnol 27:1641–1648
Gomaa OM, Selim NS, Wee J et al (2017) RNA Seq analysis of the role of calcium chloride stress and electron transport in mitochondria for malachite green decolorization by Aspergillus niger. Fungal Genet Biol 105:1–7
Gopinath KP, Kathiravan MN, Srinivasan R et al (2011) Evaluation and elimination of inhibitory effects of salts and heavy metal ions on biodegradation of Congo red by Pseudomonas sp. mutant. Bioresour Technol 102:3687–3693
Goszczynski S, Paszczynski A, Pasti-Grigsby MB et al (1994) New pathway for degradation of sulfonated azo dyes by microbial peroxidases of by Phanerochaete chrysosporium and Streptomyces chromofuscus. J Bacteriol 176:1339–1347
Hafeez F, Farheen H, Mahmood F et al (2018) Isolation and characterization of a lead (Pb) tolerant Pseudomonas aeruginosa strain HF5 for decolorization of reactive red-120 and other azo dyes. Ann Microbiol 68:943–952
Hai FI, Yamamoto K, Fukushi K (2007) Hybrid treatment systems for dye wastewater. Crit Rev Environ Sci Technol 37:315–377
Hanan HO (2008) Algal decolorization and degradation of monoazo and diazo dyes. Pak J Biol Sci 11:1310–1316
Hasanin MS, Darwesh OM, Matter IA et al (2019) Isolation and characterization of non-cellulolytic Aspergillus flavus EGYPTA5 exhibiting selective ligninolytic potential. Biocatal Agri Biotechnol 17:160–167
Hashem RA, Samir R, Essam TM et al (2018) Optimization and enhancement of textile reactive Remazol black B decolorization and detoxification by environmentally isolated pH tolerant Pseudomonas aeruginosa KY284155. AMB Express 8:83. https://doi.org/10.1186/s13568-018-0616-1
He X, Song C, Li Y et al (2018) Efficient degradation of Azo dyes by a newly isolated fungus Trichoderma tomentosum under non-sterile conditions. Ecotox Environ Safety 150:232–239
Hedayati MT, Pasqualotto AC, Warn PA et al (2007) Aspergillus flavus: human pathogen, allergen and mycotoxin producer. Microbiology 153:1677–1692
Heimann K, Huerlimann R (2015) Microalgal classification: major classes and genera of commercial microalgal species. In: Se-Kwon K (ed) Handbook of marine microalgae: biotechnolgy advances. Academic Press, London, UK, pp 25–41
Hernández-Zamora M, Perales-Vela HV, Flores-Ortíz CM et al (2014) Physiological and biochemical responses of Chlorella vulgaris to Congo red. Ecotoxicol Environ Saf 108:72–77
Hernández-Zamora M, Cristiani-Urbina E, Martínez-Jerónimo F et al (2015) Bioremoval of the azo dye Congo Red by the microalga Chlorella vulgaris. Environ Sci Pollut Res Int 22:10811–10823
Horník M, Šuňovská A, Partelová D et al (2013) Continuous sorption of synthetic dyes on dried biomass of microalga Chlorella pyrenoidosa. Chem Pap 67:254–264
Houbraken J, Samson RA, Yilmaz N (2016) Taxonomy of Aspergillus, Penicillium and Talaromyces and its significance for biotechnology. In de Vries RP, Gelber IB, Andersen MR (eds), Aspergillus and Penicillium in the post-genomic era (pp. 1-16). Caister, UK, Academic Press
Hsueh CC, Chen BY (2007) Comparative study on reaction selectivity of azo dye decolorization by Pseudomonas luteola. J Hazard Mater 141:842–849
Hsueh CC, Chen BY (2008) Exploring effects of chemical structure on azo dye decolorization characteristics by Pseudomonas luteola. J Hazard Mater 154:703–710
Hu TL (1994) Decolourization of reactive azo dyes by transformation of Pseudomonas luteola. Bioresour Technol 49:47–51
Hu TL (1996) Removal of reactive dyes from aqueous solution by different bacterial genera. Water Sci Technol 34:89–95
Hu TL (1998) Degradation of azo dye RP2B by Pseudomonas luteola. Water Sci Technol 38:229–306
Huan M, Lian-Tai L, Cai-Fang Y et al (2010) Biodegradation of malachite green by strain Pseudomonas sp. K9 and cloning of the tmr2 gene associated with an ISPpu12. World J Microbiol Biotechnol 27:1323–1329
Huang H, Wu K, Khan A et al (2016a) A novel Pseudomonas gessardii strain LZ-E simultaneously degrades naphthalene and reduces hexavalent chromium. Bioresour Technol 207:370–378
Huang J, Liu D, Lu J et al (2016b) Biosorption of reactive black 5 by modified Aspergillus versicolor biomass: kinetics, capacity and mechanism studies. Colloids Surf A Physicochem Eng Aspect 492:242–248
Idaka E, Ogawa T, Horitsu H (1987a) Reductive metabolism of aminoazobenzenes by Pseudomonas cepacia. Bull Environ Contam Toxicol 39:100–107
Idaka E, Ogawa T, Horitsu H (1987b) Oxidative pathway after reduction of p-aminoazobenzene by Pseudomonas cepacia. Bull Environ Contam Toxicol 39:108–113
Iqbal A, Sabar S, Mun-Yee MK et al (2018) Pseudomonas aeruginosa USM-AR2/SiO 2 biosorbent for the adsorption of methylene blue. J Environ Chem Eng 6:4908–4916
Isaac P, Martínez FL, Bourguignon N et al (2015) Improved PAHs removal performance by a defined bacterial consortium of indigenous Pseudomonas and actinobacteria from Patagonia, Argentina. Int Biodeterior Biodegradation 101:23–31
Işik M, Sponza DT (2003) Effect of oxygen on decolorization of azo dyes by Escherichia coli and Pseudomonas sp. and fate of aromatic amines. Process Biochem 38:1183–1192
Izmalkova TY, Sazonova OI, Nagornih MO, Sokolov SL, Kosheleva IA, Boronin AM (2013) The organization of naphthalene degradation genes in Pseudomonas putida strain AK5. Res Microbiol 164(3):244–253
Jin X, Ning Y (2013) Laccase production optimization by response surface methodology with Aspergillus fumigatus AF1 in unique inexpensive medium and decolorization of different dyes with the crude enzyme or fungal pellets. J Hazard Mater 262:870–877
Jinqi L, Houtian L (1992) Degradation of azo dyes by algae. Environ Pollut 75:273–278
Joe J, Kothari RK, Raval CM, Kothari CR (2011) Decolourization of textile dye Remazol black B by Pseudomonas aeruginosa CR-25 isolated from the common effluent treatment plant. J Bioremed Biodegrade 2:118. https://doi.org/10.4172/2155-6199.1000118
Kadam AA, Telke AA, Jagtap SS et al (2011) Decolorization of adsorbed textile dyes by developed consortium of Pseudomonas sp. SUK1 and Aspergillus ochraceus NCIM-1146 under solid state fermentation. J Hazard Mater 189:486–494
Kalme S, Ghodake G, Govindwar S (2007a) Red HE7B degradation using desulfonation by Pseudomonas desmolyticum NCIM 2112. Int Biodeterior Biodegrad 60:327–333
Kalme SD, Parshetti GK, Jadhav SU et al (2007b) Biodegradation of benzidine based dye Direct Blue-6 by Pseudomonas desmolyticum NCIM 2112. Bioresour Technol 98:1405–1410
Kalme S, Jadhav S, Jadhav M et al (2009) Textile dye degrading laccase from Pseudomonas desmolyticum NCIM 2112. Enzym Microb Technol 44:65–71
Kalpana VN, Kataru BAS, Sravani N et al (2018a) Biosynthesis of zinc oxide nanoparticles using culture filtrates of Aspergillus niger: antimicrobial textiles and dye degradation studies. Open Nano 3:48–55
Kalpana VN, Kataru BAS, Sravani N (2018b) Biosynthesis of zinc oxide nanoparticles using culture filtrates of Aspergillus niger: antimicrobial textiles and dye degradation studies. OpenNano 3:48–55
Kalyani DC, Patil PS, Jadhav JP et al (2008) Biodegradation of reactive textile dye Red BLI by an isolated bacterium Pseudomonas sp. SUK1. Bioresour Technol 99:4635–4641
Kalyani DC, Telke AA, Jadhav JP et al (2009) Ecofriendly biodegradation and detoxification of Reactive Red 2 textile dye by newly isolated Pseudomonas sp. SUK1. J Hazard Mater 163:735–742
Kalyani DC, Phugare SS, Shedbalkar UU et al (2011) Purification and characterization of a bacterial peroxidase from the isolated strain Pseudomonas sp. SUK1 and its application for textile dye decolorization. Ann Microbiol 61:483–491
Kalyani DC, Telke AA, Surwase SN et al (2012) Effectual decolorization and detoxification of triphenylmethane dye malachite green (MG) by Pseudomonas aeruginosa NCIM 2074 and its enzyme system. Clean Techn Environ Policy 14:989–1001
Kang Y, Xu X, Pan H, Tian J, Tang W, Liu S (2017) Decolorization of mordant yellow 1 using. TS-A CGMCC 12964 by biosorption and biodegradation. Bioengineered 9(1):222–232
Kanhere J, Gopinathan R, Banerjee J (2014) Cytotoxicity and genotoxicity of malachite green on non-target aquatic organisms: Chlorella pyrenoidosa and Daphnia magna. Water Air Soil Pollut 225:2134. https://doi.org/10.1007/s11270-014-2134-3
Katheresan V, Kansedo J, Lau SY (2018) Efficiency of various recent wastewater dye removal methods: a review. J Environ Chem Eng 6:4676–4697
Kaushik P, Malik A (2010) Effect of nutritional conditions on dye removal from textile effluent by Aspergillus lentulus. World J Microbiol Biotechnol 26(11):1957–1964
Kaushik P, Malik A (2011) Process optimization for efficient dye removal by Aspergillus lentulus FJ172995. J Hazard Mater 185(2–3):837–843
Keck A, Klein J, Kudlich M et al (1997) Reduction of azo dyes by redox mediators originating in the naphthalene sulfonic acid degradation pathway of Sphingomonas ssp. Strain BN6. Appl Environ Microbiol 63:3684–3690
Khalaf MA (2008) Biosorption of reactive dye from textile wastewater by non-viable biomass of Aspergillus niger and Spirogyra sp. Bioresour Technol 99:6631–6634
Khambhaty Y, Mody K, Basha S (2012) Efficient removal of Brilliant Blue G (BBG) from aqueous solutions by marine Aspergillus wentii: kinetics, equilibrium and process design. Ecol Eng 41:74–83
Khan Z, Jain K, Soni A et al (2014) Microaerophilic degradation of sulphonated azo dye- Reactive Red 195 by bacterial consortium AR1 through co-metabolism. Int Biodeterior Biodegradation 94:167–175
Knapp JS, Newby PS (1999) The decolourisation of a chemical industry effluent by white rot fungi. Water Res 33:575–577
Kong J, Wang H, Liang L et al (2017) Phenanthrene degradation by the bacterium Pseudomonas stutzeri JP1 under low oxygen condition. Int Biodeterior Biodegradation 123:121–126
Kousha M, Farhadian O, Dorafshan S et al (2013) Optimization of malachite green biosorption by green microalgae—Scenedesmus quadricauda and Chlorella vulgaris: application of response surface methodology. J Taiwan Inst Chemical E 44:291–294
Kuddus M, Joseph B, Wasudev Ramteke P (2013) Production of laccase from newly isolated Pseudomonas putida and its application in bioremediation of synthetic dyes and industrial effluents. Biocat Agri Biotechnol 2:333–338
Kumar Garg S, Tripathi M, Singh SK et al (2012) Biodecolorization of textile dye effluent by Pseudomonas putida SKG-1 (MTCC 10510) under the conditions optimized for monoazo dye orange II color removal in simulated minimal salt medium. Int Biodeterior Biodegrad 74:24–35
Kumar CG, Mongolla P, Sheik AB et al (2011) Decolorization and biotransformation of triphenylmethane dye, methyl violet, by Aspergillus sp. isolated from Ladakh, India. J Microbiol Biotechnol 21:267–273
Kumar CG, Mongolla P, Joseph J, Sarma VUM (2012) Decolorization and biodegradation of triphenylmethane dye, brilliant green, by Aspergillus sp. isolated from Ladakh, India. Process Biochem 47(9):1388–1394
Kumar R, Kaur J, Jain S, Kumar A (2016) Optimization of laccase production from Aspergillus flavus by design of experiment technique: partial purification and characterization. J Genet Eng Biotechnol 14(1):125–131
Kuppusamy S, Sethurajan M, Kadarkarai M et al (2017) Biodecolourization of textile dyes by novel, indigenous Pseudomonas stutzeri MN1 and Acinetobacter baumannii MN3. J Environ Chem Eng 5:716–724
Lebron YAR, Moreira VR, Santos LVS et al (2018) Remediation of methylene blue from aqueous solution by Chlorella pyrenoidosa and Spirulina maxima biosorption: equilibrium, kinetics, thermodynamics and optimization studies. J Environ Chem Eng 6:6680–6690
Li L, Hong Q, Yan X et al (2009) Isolation of a malachite green-degrading Pseudomonas sp. MDB-1 strain and cloning of the tmr2 gene. Biodegradation 20:769–776
Lim SL, Chu WL, Phang SM (2010) Use of Chlorella vulgaris for bioremediation of textile wastewater. Bioresour Technol 101:7314–7322
Lin YH, Leu JY (2008) Kinetics of reactive azo-dye decolorization by Pseudomonas luteola in a biological activated carbon process. Biochem Eng J 39:457–467
Lin SF, Yu P, Lin YM (2004) Study on decolorization of malachite green by a Pseudomonas aeruginosa. J Fujian Norm Univ 20:72–75
Lin J, Zhang X, Li Z et al (2010) Biodegradation of Reactive blue 13 in a two-stage anaerobic/aerobic fluidized beds system with a Pseudomonas sp. isolate. Bioresour Technol 101:34–40
Liu C, You Y, Zhao R et al (2017) Biosurfactant production from Pseudomonas taiwanensis L1011 and its application in accelerating the chemical and biological decolorization of azo dyes. Ecotoxicol Environ Saf 145:8–15
Lodato A, Alfieri F, Olivieri G et al (2007) Azo-dye conversion by means of Pseudomonas sp. OX1. Enzym Microb Technol 41:646–652
Logroño W, Pérez M, Urquizo G et al (2017) Single chamber microbial fuel cell (SCMFC) with a cathodic microalgal biofilm: a preliminary assessment of the generation of bioelectricity and biodegradation of real dye textile wastewater. Chemosphere 176:378–388
Lu T, Zhang Q, Yao S (2017) Efficient decolorization of dye-containing wastewater using mycelial pellets formed of marine-derived Aspergillus niger. Chin J Chem Eng 25:330–337
Luan TG, Jin J, Chan SMN et al (2006) Biosorption and biodegradation of tributyltin (TBT) by alginate immobilized Chlorella vulgaris beads in several treatment cycles. Process Biochem 41:1560–1565
Mahmoud MS, Mostafa MK, Mohamed SA (2017) Bioremediation of red azo dye from aqueous solutions by Aspergillus niger strain isolated from textile wastewater. J Environ Chem Eng 5:547–554
Malla FA, Khan SA, Rashmi et al (2015) Phycoremediation potential of Chlorella minutissima on primary and tertiary treated wastewater for nutrient removal and biodiesel production. Ecol Eng 75:343–349
Mangwani N, Shukla SK, Rao TS (2014) Calcium-mediated modulation of Pseudomonas mendocina NR802 biofilm influences the phenanthrene degradation. Colloids Surf B: Biointerfaces 114:301–309
Maqbool Z, Hussain S, Ahmad T et al (2016) Use of RSM modeling for optimizing decolorization of simulated textile wastewater by Pseudomonas aeruginosa strain ZM130 capable of simultaneous removal of reactive dyes and hexavalent chromium. Environ Sci Pollut Res 23:11224–11239
Mathur M, Gola D, Panja R, Malik A, Ahammad SZ (2018) Performance evaluation of two Aspergillus spp. for the decolourization of reactive dyes by bioaccumulation and biosorption. Environ Sci Pollut Res 25(1):345–352
McMullan G, Meehan C, Conneely A et al (2001) Microbial decolourisation and degradation of textile dyes. Appl Microbiol Biotechnol 56:81–87
Michaels GB, Lewis DL (1985) Sorption and toxicity of azo and triphenylmethane dyes to aquatic microbial populations. Environ Toxicol Chem 4:45–50
Mishra S, Maiti A (2018) Optimization of process parameters to enhance the bio-decolorization of Reactive Red 21 by Pseudomonas aeruginosa 23N1. Int J Environ Sci Technol 16:6685–6698. https://doi.org/10.1007/s13762-018-2023-1
Mohan SV, Sistla S, Guru RK et al (2003) Microbial degradation of pyridine using Pseudomonas sp. and isolation of plasmid responsible for degradation. Waste Manag 23:167–171
Moharikar A, Purohit HJ (2003) Specific ratio and survival of Pseudomonas CF600 as co-culture for phenol degradation in continuous cultivation. Int Biodeterior Biodegrad 52:255–260
Munoza R, Guieysse B (2006) Algal–bacterial processes for the treatment of hazardous contaminants: a review. Water Res 40:2799–2815
Muthu Kumara Pandian A, Karthikeyan C, Rajasimman M (2016) Isotherm and kinetic studies on nano-sorption of malachite green onto Aspergillus flavus mediated synthesis of silver nano particles. Environ Nanotechnol Monitor Manag 6:139–151
Nachiyar CV, Rajkumar GS (2003) Degradation of a tannery and textile dye, Navitan Fast Blue S5R by Pseudomonas aeruginosa. World J Microbiol Biotechnol 19:609–614
Nachiyar CV, Rajkumar GS (2004) Mechanism of Navitan fast Blue S5R degradation by Pseudomonas aeruginosa. Chemosphere 57:165–169
Nachiyar CV, Rajkumar GS (2005) Purification and characterization of an oxygen insensitive azoreductase from Pseudomonas aeruginosa. Enzym Microb Technol 36:503–509
Nachiyar CV, Vijayalakshmi K, Muralidharan D et al (2007) Mineralization of metanilic acid by Pseudomonas aeruginosa CLRI BL22. World J Microbiol Biotechnol 23:1733–1738
Nakanishi M, Yatome C, Ishida N et al (2001) Putative ACP phosphodiesterase gene encodes an azoreductase. J Biol Chem 49:46394–46399
Naraian R, Kumari S, Gautam RL (2018) Biodecolorization of brilliant green carpet industry dye using three distinct Pleurotus spp. Environ Sustain 1:141–148
Naskar A, Majumder R (2017) Understanding the adsorption behaviour of acid yellow 99 on Aspergillus niger biomass. J Mol Liq 242:892–899
Nigam P, Banat IM, Singh D et al (1996) Microbial process for the decolorization of textile effluent containing azo, diazo and reactive dyes. Process Biochem 31:435–442
Ning C, Qingyun L, Aixing T et al (2018) Decolorization of a variety of dyes by Aspergillus flavus A5p1. Bioprocess Biosyst Eng 41:511–518
Nitisakulkan T, Oku S, Kudo D et al (2014) Degradation of chloroanilines by toluene dioxygenase from Pseudomonas putida T57. J Biosci Bioeng 117:292–297
Nowak A, Mrozik A (2018) Degradation of 4-chlorophenol and microbial diversity in soil inoculated with single Pseudomonas sp. CF600 and Stenotrophomonas maltophilia KB2. J Environ Manag 215:216–229
Nwinyi OC, Ajayi OO, Amund OO (2016) Degradation of polynuclear aromatic hydrocarbons by two strains of Pseudomonas. Braz J Microbiol 47:551–562
Oranusi NA, Ogugbue CJ (2005) Effect of cosubstrates on primary biodegradation of triphenylmethane dyes by Pseudomonas sp. Afr J Appl Zool Environ Biol 7:38–44
Pandey BV, Upadhyay RS (2006) Spectroscopic characterization and identification of Pseudomonas fluorescens mediated metabolic products of Acid Yellow-9. Microbiol Res 161:311–315
Pandey RK, Tewari S, Tewari L (2018) Lignolytic mushroom Lenzites elegans WDP2: laccase production, characterization, and bioremediation of synthetic dyes. Ecotox Environ Safety 158:50–58
Park JK, Chang HN (2000) Microencapsulation of microbial cells. Biotechnol Adv 18:303–319
Park HS, Jun SC, Han KH et al (2017) Diversity, application, and synthetic biology of industrially important Aspergillus fungi. Adv Appl Microbiol 100:161–202
Parshetti GK, Kalme SD, Gomare SS (2007) Biodegradation of reactive blue-25 by Aspergillus ochraceus NCIM-1146. J Biotechnol 98:3638–3642
Paszczynski A, Pasti-Grigsby MB, Goszczynski S et al (1992) Mineralization of sulfonated azo dyes and sulfanilic acid by Phanerochaete chrysosporium and Streptomyces chromofuscust. Appl Environ Microbiol 58:3598–3604
Patel R, Suresh S (2008) Kinetic and equilibrium studies on the biosorption of reactive black 5 dye by Aspergillus foetidus. Bioresour Technol 99:51–58
Patel Y, Mehta C, Gupte A (2012) Assessment of biological decolorization and degradation of sulfonated di-azo dye Acid Maroon V by isolated bacterial consortium EDPA. Int Biodeterior Biodegrad 75:187–193
Pathak VV, Kothari R, Chopra A et al (2015) Experimental and kinetic studies for phycoremediation and dye removal by Chlorella pyrenoidosa from textile wastewater. J Environ Manag 163:270–277
Pazarlioğlu NK, Telefoncu A (2005) Biodegradation of phenol by Pseudomonas putida immobilized on activated pumice particles. Process Biochem 40:1807–1814
Peng X, Ma X, Xu Z (2015) Thermogravimetric analysis of co-combustion between microalgae and textile dyeing sludge. Bioresour Technol 180:288–295
Perei K, Rakhely G, Kiss I et al (2001) Biodegradation of sulfanilic acid by Pseudomonas paucimobilis. Appl Microbiol Biotechnol 55:101–107
Phugare SS, Waghmare SR, Jadhav JP (2011) Purification and characterization of dye degrading of veratryl alcohol oxidase from Pseudomonas aeruginosa strain BCH. World J Microbiol Biotechnol 27:2415–2423
Puvaneshwari N, Muthukrishnan J, Gunasekaran P et al (2002) Biodegradation of benzidine based azodyes direct red and direct blue by the immobilized cells of Pseudomonas fluorescens D41. Indian J Exp Biol 40:1131–1136
Qian HF, Chen W, Sheng GD et al (2008) Effects of glufosinate on antioxidant enzymes, subcellular structure, and gene expression in the unicellular green alga Chlorella vulgaris. Aquat Toxicol 88:301–307
Rafii F, Franklin W, Cerniglia CE (1990) Azoreductase activity of anaerobic bacteria isolated from human intestinal microflora. Appl Environ Microbiol 56:2146–2151
Rampelotto PH (2010) Resistance of microorganisms to extreme environmental conditions and its contribution to astrobiology. Sustain For 2:1602–1623
Ramya M, Anusha B, Kalavathy S et al (2007) Biodecolorization and biodegradation of Reactive Blue by Aspergillus spp. Afr J Biotechnol 6:1441–1445
Rani B, Kumar V, Singh J, Bisht S, Teotia P, Sharma S, Kela R (2014) Bioremediation of dyes by fungi isolated from contaminated dye effluent sites for bio-usability. Braz J Microbiol 45(3):1055–1063
Roy U, Sengupta S, Banerjee P et al (2018) Assessment on the decolourization of textile dye (Reactive Yellow) using Pseudomonas sp. immobilized on fly ash: response surface methodology optimization and toxicity evaluation. J Environ Manag 223:185–195
Ryu BH, Weon YD (1992) Decolorization of Azo Dyes by Aspergillus sojae B-10. J Microbiol Biotechnol 2:215–219
Sá CS, Boaventura RA (2001) Biodegradation of phenol by Pseudomonas putida DSM 548 in a trickling bed reactor. Biochem Eng J 9:211–219
Safi C, Zebib B, Merah O et al (2014) Morphology, composition, production, processing and applications of Chlorella vulgaris: a review. Renew Sust Energ Rev 35:265–278
Samson RA, Visagie CM, Houbraken J et al (2014) Phylogeny, identification and nomenclature of the genus Aspergillus. Studies Myco 78:141–173
Saparrat MCN, Balatti PA, Arambarri AM et al (2014) Coriolopsis rigida, a potential model of white-rot fungi that produce extracellular laccases. J Ind Microbiol Biotechnol 41:607–617
Sarnaik S, Kanekar P (1995) Bioremediation of colour of methyl violet and phenol from a dye-industry waste effluent using Pseudomonas spp. isolated from factory soil. J Appl Bacteriol 79:459–469
Sarnaik S, Kanekar P (1999) Biodegradation of methyl violet by Pseudomonas mendocina MCM B-402. Appl Microbiol Biotechnol 52:251–254
Sathishkumar K, Sathiyaraj S, Parthipan P et al (2017) Electrochemical decolorization of methyl red by RuO2 -IrO2 -TiO2 electrode and biodegradation with Pseudomonas stutzeri MN1 and Acinetobacter baumannii MN3: an integrated approach. Chemosphere 183:204–211
Selvakumar KV, Basha CA, Prabhu HJ et al (2010) The potential of free cells of Pseudomonas aeruginosa on textile dye degradation. Bioresour Technol 101:2678–2684
Seo YH, Park D, Oh YK et al (2015) Harvesting of microalgae cell using oxidized dye wastewater. Bioresour Technol 192:802–806
Sharma P, Singh L, Dilbaghi N (2009) Response surface methodological approach for the decolorization of simulated dye effluent using Aspergillus fumigatus fresenius. J Hazard Mater 161:1081–1086
Silveira E, Marques PP, Silva SS et al (2009) Selection of Pseudomonas for industrial textile dyes decolourization. Int Biodeterior Biodegrad 63:230–235
Singh U, Arora NK, Sachan P (2018) Simultaneous biodegradation of phenol and cyanide present in coke-oven effluent using immobilized Pseudomonas putida and Pseudomonas stutzeri. Braz J Microbiol 49:38–44
Sinha S, Singh R, Chaurasia AK et al (2016) Self-sustainable Chlorella pyrenoidosa strain NCIM 2738 based photobioreactor for removal of Direct Red-31 dye along with other industrial pollutants to improve the water-quality. J Hazard Mater 306:386–394
Srinivasan R, Kathiravan MN, Gopinath KP (2011) Degradation of Tectilon Yellow 2G by hybrid technique: combination of sonolysis and biodegradation using mutant Pseudomonas putida. Bioresour Technol 102:2242–2247
Stormo KE, Crawford RL (1992) Preparation of encapsulated microbial cells for environmental applications. Appl Environ Microbiol 58:727–730
Sugiura W, Miyashita T, Yokoyama T et al (1999) Isolation of azo-dye degrading microorganisms and their application to white discharge printing of fabric. J Biosci Bioeng 88:577–581
Sultan M (2017) Polyurethane for removal of organic dyes from textile wastewater. Environ Chem Lett 15:347. https://doi.org/10.1007/s10311-016-0597-8
Sumathi S, Manju B (2000) Uptake of reactive textile dyes by Aspergillus foetidus. Enzym Microb Technol 27:347–355
Surkatti R, El-Naas MH (2014) Biological treatment of wastewater contaminated with p-cresol using Pseudomonas putida immobilized in polyvinyl alcohol (PVA) gel. J Water. Process Eng 1:84–90
Suzuki Y, Yoda T, Ruhul A et al (2001) Molecular cloning and characterization of the gene encoding azoreductase from Bacillus sp. OY 1-2isolated from soil. J Biol Chem 246:9059–9065
Tamayo-Ramos JA, van Berkel WJ, de Graaff LH (2012) Biocatalytic potential of laccase-like multicopper oxidases from Aspergillus niger. Microb Cell Factories 11:165. https://doi.org/10.1186/1475-2859-11-165
Tao R, Kinnunen V, Praveenkumar R et al (2017) Comparison of Scenedesmus acuminatus and Chlorella vulgaris cultivation in liquid digestates from anaerobic digestion of pulp and paper industry and municipal wastewater treatment sludge. J Appl Phycol 29:2845–2856
Telke AA, Kalyani DC, Jadhav UU et al (2009) Purification and characterization of an extracellular laccase from a Pseudomonas sp. LBC1 and its application for the removal of bisphenol A. J Mol Cata B: Enzymatic 61:252–260
Telke AA, Kim SW, Govindwar SP (2012) Significant reduction in toxicity, BOD, and COD of textile dyes and textile industry effluent by a novel bacterium Pseudomonas sp. LBC1. Folia Microbiol 57:115–122
Thao TP, Kao HC, Juang RS et al (2013) Kinetic characteristics of biodegradation of methyl orange by Pseudomonas putida mt2 in suspended and immobilized cell systems. J Taiwan Inst Chem Eng 44:780–785
Tsai WT, Chen HR (2010) Removal of malachite green from aqueous solution using low-cost chlorella-based biomass. J Hazard Mater 175:844–849
Tuttolomondo MV, Alvarez GS, Desimone MF et al (2014) Removal of azo dyes from water by sol–gel immobilized Pseudomonas sp. J Environ Chem Eng 2:131–136
Wang B, Hu Y (2007) Comparison of four supports for adsorption of reactive dyes by immobilized Aspergillus fumigatus beads. J Environ Sci 19:451–457
Wang BE, Hu YY, Xie L et al (2008) Biosorption behavior of azo dye by inactive CMC immobilized Aspergillus fumigatus beads. Bioresour Technol 99:794–800
Wang W, Zhang Z, Ni H et al (2012) Decolorization of industrial synthetic dyes using engineered Pseudomonas putida cells with surface-immobilized bacterial laccase. Microb Cell Factories 11:75. https://doi.org/10.1186/1475-2859-11-75
Wang L, Chen X, Wang H et al (2017) Chlorella vulgaris cultivation in sludge extracts from 2,4,6-TCP wastewater treatment for toxicity removal and utilization. J Environ Manag 187:146–153
Wu J, Jung BG, Kim KS et al (2009) Isolation and characterization of Pseudomonas otitidis WL-13 and its capacity to decolorize triphenylmethane dyes. J Environ Sci 21:960–964
Xiong XJ, Meng XJ, Zheng TL (2010) Biosorption of C.I Direct Blue 199 from aqueous solution by nonviable Aspergillus niger. J Hazard Mater 175:241–246
Xu H, Miao X, Wu Q (2006) High quality biodiesel production from a microalga Chlorella protothecoides by heterotrophic growth in fermenters. J Biotechnol 126:499–507
Xu C, Wang R, Zhang YF et al (2015) Stress response of Chlorella pyrenoidosa to nitro-aromatic compounds. Environ Sci Pollut Res 22:3784–3793
Yang Y, Hu H, Wang G et al (2011a) Removal of malachite green from aqueous solution by immobilized Pseudomonas sp. DY1 with Aspergillus oryzae. Int Biodeterior Biodegrad 65:429–434
Yang Y, Jin D, Wang G et al (2011b) Competitive biosorption of Acid Blue 25 and Acid Red 337 onto unmodified and CDAB-modified biomass of Aspergillus oryzae. Bioresour Technol 102:7429–7436
Yatome C, Ogawa T, Koga D et al (1981) Biodegradability of azo and triphenylmethane dyes by Pseudomonas pseudomallei 13 NA. J Soc Dye Colour 97:166–169
Yatome C, Ogaw T, Hishida H et al (1990) Degradation of azo dyes by cell-free extract from Pseudomonas stutzeri. J Soc Dye Colour 106:280–283
Yatome C, Matsufuru H, Taguchi T et al (1993) Degradation of 4-dimethylaminoazobenzene-2-carboxylic acid by Pseudomonas stutzeri. Appl Microbiol Biotechnol 39:778–781
Yu J, Wang X, Yue PL (2001) Optimal decolorization and kinetic modeling of synthetic dyes by Pseudomonas strains. Water Res 35:3579–3586
Zabłocka-Godlewska E, Przystaś W, Grabińska-Sota E (2014) Decolourisation of different dyes by two Pseudomonas strains under various growth conditions. Water Air Soil Pollut 225:1846. https://doi.org/10.1007/s11270-013-1846-0
Zhao W, Sun H, Ren Y et al (2018) Chlorella zofingiensis as a promising strain in wastewater treatment. Bioresour Technol 268:286–291
Zheng S, He M, Sui Y et al (2017) Kelp waste extracts combined with acetate enhances the biofuel characteristics of Chlorella sorokiniana. Bioresour Technol 225:142–150
Zhipei L, Huifang Y (1991) Decolorization and biodegradation metabolism of azo dyes Pseudomonas S-42. J Environ Sci 3:89–102
Zimmermann T, Kulla GH, Leisinger T (1982) Properties of purified orange II azoreductase, the enzyme initiating azo dye degradation by Pseudomonas KF46. Eur J Biochem 29:197–203
Zope V, Kulkarni M, Chavan M (2007) Biodegradation of synthetic textile dyes Reactive Red 195 and Reactive Green 11 by Aspergillus niger grp: an alternative approach. J Sci Ind Res 66:411–414
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Pradhan, S.K., Singla, R. (2020). Potential of Thallophytes in Degradation of Dyes in Industrial Effluents. In: Arora, P. (eds) Microbial Technology for Health and Environment. Microorganisms for Sustainability, vol 22. Springer, Singapore. https://doi.org/10.1007/978-981-15-2679-4_13
Download citation
DOI: https://doi.org/10.1007/978-981-15-2679-4_13
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-2678-7
Online ISBN: 978-981-15-2679-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)