Abstract
Plant–soil microbe interactions are a key determinant of plant community composition and structure, with each plant species generating a unique soil microbiota. However, the degree of intraspecific variation in plant–soil microbe interactions in controlling plant performance has been much less investigated. We examined the strength of plant–soil microbe interactions across Solidago altissima and Schizachyrium scoparium seedlings grown in soils cultured by 24 genotypes of S. altissima in a greenhouse setting to determine the level of variation within these two target species. We also quantified leaf chemical variation across S. altissima genotypes using HPLC characterization of foliar constituents as a potential driver of below-ground processes. The impacts of soil microbe-mediated effects of genotypes ranged from negative to positive for both target species. Target species responses to the 24 soil microbial communities were positively correlated, although the strength of plant–soil microbe interactions was related to foliar chemistry for S. scoparium, but not for S. altissima. The magnitude of impacts in soils pooled from all 24 S. altissima genotypes was more negative than the average impact of genotypes tested individually, suggesting that this methodology cannot be used to assess ‘average’ effects. Our results strongly argue that intraspecific variation in plant–soil microbe interactions may be a quite large source of variation in systems dominated by clonal plants. Furthermore, simple inocula pooling obscured this variation and resulted in a marked bias toward the more antagonistic components of the soil microbial community.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Terrestrial plants interact with the biotic and abiotic components of their environment, resulting in variation in plant performance and community dynamics. Key among these interactions is the relationship between plants and their associated soil microbes, which may influence both the fitness of individual plants and their abundance within communities (van der Putten et al. 2013). Plants may condition their soil through the leaching of chemicals via litter or root exudation (Kong et al. 2008; Heinen et al. 2017; Veen et al. 2019), producing distinct soil microbial communities (Kardol et al. 2006; Pendergast et al. 2013; Orozco-Aceves et al. 2015) that may affect plant performance and coexistence (van der Putten et al. 2013; Smith-Ramesh and Reynolds 2017; Xue et al. 2018). However, associated soil microbes vary in their functional roles, generating variation in the benefits and drawbacks they provide to plant hosts (Magnoli and Lau 2020; Veen et al. 2021). Changes in soil microbial composition can impact the fitness of the individual plant itself, others of the same species (intraspecific), or individuals of other species (interspecific), leading to positive, neutral, or negative effects (Klironomos 2002; van der Putten et al. 2013; Mack and Bever 2014; Kos et al. 2015; Smith-Ramesh and Reynolds 2017). Through impacts on plant performance, interspecific variation in plant–soil microbe interactions appears to be a major driving force for plant community composition, succession, invasion, and ecosystem function (Ehrenfeld et al. 2005; Kardol et al. 2007; Miki 2012; Pendergast et al. 2013; Mack and Bever 2014; Verbeek and Kotanen 2019).
Intraspecific trait variation has long been recognized as a critical driving force in evolution that often manifests itself in population or community processes (Bolnick et al. 2011). As functional ecology has provided a more mechanistic view of plant community organization, intraspecific variation has also become a critical component (Violle et al. 2012; Hahn and Maron 2016). Intraspecific variation in plant chemistry can impact the quality of anti-predator defenses (Glassmire et al. 2016; Benedek et al. 2019), responses to interspecific competition (Jilani et al. 2008), as well as resistance to pathogens and parasites (Biere et al. 2004; Shikano et al. 2017), linking above- and below-ground processes (Van Der Putten et al. 2001; Bezemer and Van Dam 2005). Similarly, variation in plant chemical composition may result in changes to the composition of soil microbial communities (Lankau 2010; Benedek et al. 2019), with microbial biomass dependant on chemical and litter inputs and the responses of individual microbes to them (Kong et al. 2008). Litter chemical composition and quality act as an environmental filter that results in variable competitive outcomes between soil microorganisms for resources (Veen et al. 2021). Locally produced leaf litter may also provide soil legacy effects in the absence of the original host plants, affecting the soil microbiota, soil functioning, and growth of newly establishing plants (Wurst and Ohgushi, 2015; Veen et al. 2019). We should therefore expect that individual plant genotypes may influence the diversity and composition of soil microbial communities as a whole (Schweitzer et al. 2008; Bukowski and Petermann 2014; Gehring et al. 2017), as well as their reciprocal impacts on plants.
Although the preponderance of plant–microbe interaction work has focused on interspecific differences (Bukowski et al. 2018), critical intraspecific variation has been identified in a variety of systems. Genotype-specific microbial communities have been documented for both mutualistic (Gehring et al. 2017; Wang et al. 2019) and antagonistic (Liu et al. 2015; Eck et al. 2019) soil microbes, yielding population-level impacts (Bukowski et al. 2018). Overall, there may be significant variation associated with individual plant genotypes (Bukowski and Petermann 2014; Bukowski et al. 2018; Wang et al. 2019) and available microbes (Magnoli and Lau 2020), leading to a range in the strength and direction of the plant–microbe interactions generated.
Intraspecific variation in plant–microbe interactions likely varies spatially at the scale of the individual plant’s root system (Peacher and Meiners 2020). However, the effects of intraspecific plant chemical variation on soil microbes may be spatially much larger in clonal plant species, where vegetative reproduction of an individual plant can lead to a single genotype covering large areas for long periods of time (De Witte and Stöcklin 2010). Clonal reproduction is considered a competitive trait (Grime 2001), which is quite important in the dynamics of many successional plant communities (Prach et al. 1997; Wright and Fridley 2010). In systems where clonal species dominate and vary in chemistry or other key traits, the spatial scale of variation in soil microbial communities may represent an ecologically important source of interaction heterogeneity.
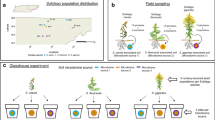
To assess intraspecific variation in plant–microbe interactions, we used a common garden of 24 genotypes of Solidago altissima, an aggressive dominant in many plant communities worldwide. We used soils collected from each individual genotype as inocula in a greenhouse study to determine the strength of plant–soil microbe interactions on two native target species, S. altissima and the bunchgrass Schizachyrium scoparium. Using these data, we addressed the following research questions: (1) Does appreciable intraspecific variation exists in the direction and magnitude of plant–soil microbe interactions across S. altissima genotypes? (2) Is this variation consistent across target species? (3) Is variation in the strength of plant–soil microbe interactions related to variation in S. altissima foliar chemistry? If appreciable variation occurs across S. altissima genotypes, this poses a methodological issue in how to deal with variation in characterizing plant–microbe interactions. Therefore, we also address: (4) Does pooling of soil microbial inocula provide a reliable estimate of the mean interaction across Solidago genotypes for both target species?
Materials and methods
Study species
Solidago altissima (syn. S. canadensis), is a chemically diverse, allelopathic perennial herb that is common in old fields and other open habitats across its native range of Eastern North America (Werner et al. 1980; Meiners et al. 2017; Kalske et al. 2019; Uesugi et al. 2019; Yip et al. 2019). Solidago altissima reproduces clonally through rhizomes and is very diverse in its chemical composition, allowing this species to become a successful invader across Europe, Japan, and Australia (Weber 1998; Uesugi et al. 2019; Yip et al. 2019). Solidago altissima has been used to document ecological impacts of chemical variation (Uesugi and Kessler 2013; Uesugi et al. 2019), impacts on soil microbial communities (Zhang et al. 2009), and variation in plant–microbe interactions (Dong et al. 2015; Howard et al. 2020; Peacher and Meiners 2020), representing a good model system to test for intraspecific variation in plant–soil microbe interactions.
Solidago altissima colonizes areas with disturbed soils through seed dispersal (Werner et al. 1980) and then begins to spread clonally. Initial populations of S. altissima have large numbers of genotypes. However, as stem density increases, genotypes interact and displace competitively inferior individuals, leading to fewer but larger clones in older populations (Hartnett and Bazzaz 1985). Therefore, S. altissima populations are expected to be more variable before clonal sorting has occurred, providing an opportunity to explore the full range of intraspecific variation in this species.
We added an additional target species, Schizachyrium scoparium, to assess the importance of intraspecific variation in interspecific plant–microbe interactions. This species is a C4 bunchgrass native to most of North America as a major component of grasslands and disturbed habitats (Pickett 1983; Huff et al. 1998; Duell et al. 2016). This second target species was added because it commonly co-occurs with S. altissima, is the dominant native grass at the garden site, responds to soil microbial communities (Bauer et al. 2017), and is inhibited by the soil microbial community of S. altissima (Peacher and Meiners 2020). The inclusion of S. scoparium provides insight into the role of intraspecific microbial variation on a common competitor of S. altissima.
Common garden
In May 2014, ramets of each of 24 genotypes of S. altissima were collected as rhizome/stem segments from the Douglas-Hart Nature Center (Mattoon, IL; 39° 29′ N; 88° 17′ W) in an area of recently restored prairie. Five rhizome segments per genotype were excavated from distinct patches within close proximity to one another (within 0.5 m) and isolated (> 5 m) from other such patches to ensure collection from different genotypes. The area had been in row-crop agriculture three years prior and S. altissima was not a part of the initial restoration seeding, so all S. altissima represented colonists from the surrounding landscape. This site age should also represent the phase before sorting of genotypes would be expected and should still retain high genetic (Hartnett and Bazzaz 1985) and potentially chemical diversity (see below).
The common garden site was a level section of land in Clark County, IL (39° 19′ N, 87° 55′ W) that had been used to grow corn the previous year using annual tillage practices. For this reason, we expect initial soil microbes and abiotic properties to be relatively homogeneous across the 12.5 × 20 m garden (Robertson et al. 1993; Röver and Kaiser 1999; Constancias et al. 2015). The 5 ramets from each S. altissima genotype were planted in a regular pattern (center and toward each corner) within a single, 1.6 × 1.6 m enclosure (one genotype per enclosure) with aluminum flashing buried 15 cm deep to prevent rhizome spread across plots. Plots were separated by 2 m and the spaces between enclosures were maintained by mowing. After planting S. altissima, other plants were allowed to colonize naturally to assess variation in competitive ability across the genotypes in a related study (B. S. Foster, Unpublished thesis). During the first two years, all S. altissima flowering heads were removed prior to seed set to prevent colonization of the plots by new genetic individuals.
Chemical analysis
High-performance liquid chromatography (HPLC) analysis was used to characterize leaf chemistry and determine whether S. altissima genotypes within the common garden varied in the amount or type of chemicals produced (Meiners et al. 2017). In the summer of 2016, five fully expanded leaves with no visible herbivore damage were collected from the top of separate stems within each enclosure. Pooled leaf samples were freeze dried with liquid nitrogen, ground, and extracted using 1 mL of HPLC-grade methanol from 100 mg of ground tissue. After centrifugation, the supernatant was filtered through a 0.22 µm filter and analyzed using a Hitachi Chromaster HPLC with a 5430 Diode Array detector. Separation was achieved using a Hypersil Gold C18 column (5 µm × 4.6 mm × 250 mm; Thermo Fisher Scientific). The mobile phase was a mixture of acetonitrile:water (v/v) at 20:80 from 0 to 5 min, a linear gradient of 20:80 to 95:5 from 5 to 45 min, 95:5 from 45 to 55 min, a linear gradient of 95:5 to 20:80 for 55–60 min, and 20:80 for 60–70 min. The flow rate was constant at 0.7 ml/min and the sample loading volume was 10 µL. Chemical constituents were separated by time of elution and the area of the peak used as an estimate of the amount present. Only peaks that were discernable from the baseline (> 75 µV s) were retained for analysis.
Chemical composition for all genotypes in the common garden was run through a non-metric multidimensional scaling (NMDS) analysis using Sorensen distance to generate independent axes of chemical variation. Peaks that occurred in 5 or fewer genotypes were omitted from analysis as uninformative, resulting in the inclusion of 117 peaks. The optimum number of dimensions for the ordination was determined by comparison to randomized data in PC-ORD 6.0 (MjM Software, Gleneden Beach, OR; McCune and Grace 2002). The three resulting axes were used to relate chemical composition to the strength and direction of plant–microbe interactions generated by each genotype (3D stress = 0.083). While above-ground chemistry may be related to plant–microbe interactions via leaching and decomposition (Veen et al. 2021), root-secreted chemicals often undergo modification in the soil, making adequate quantification difficult (Wang et al. 2020), so here we only focus on above-ground chemistry.
Greenhouse bioassay
In September 2019, the sixth growing season after planting approximately 500 ml of soil was collected from the top 10 cm within each genotype’s enclosure. Soil collection was done where S. altissima was most dense to avoid areas influenced by the roots of other species. By this time, the vast majority of biomass in most plots was contributed by S. altissima (B. S. Foster, Unpublished thesis). Collection tools were washed and sterilized with a 5% bleach solution between genotypes. Soil from each genotype was sifted to remove stones and root fragments, generating a live inoculum for each S. altissima genotype. From equal parts of each genotype, we also generated a pooled inoculum. Half of the pooled inoculum was autoclaved at 20 min at 120 °C and 0.07 MPa to produce a single control inoculum to compare to all individual genotype’s live soil inoculum. Because of the small size of the planting area and its previous use for row crops, abiotic soil conditions should have been relatively consistent across the site, justifying a single control inoculum. The remaining pooled inoculum was used live to assess whether sample pooling across genotypes would be equivalent to the mean interaction strength of the individual genotype’s microbial communities (question 4). This inoculum collection technique would contain mutualistic and antagonistic bacteria, fungi, and other soil organisms—collectively referred to here as soil microbes.
Two native species were used as targets in this experiment, Solidago altissima to detect potential direct population feedbacks and Schizachyrium scoparium to assess potential variation in microbe-mediated interspecific interactions. Seeds of S. altissima were collected from multiple individuals from the original source location of S. altissima genotypes (Douglas-Hart Nature Center) and were moist stratified for 60 d at 5 °C prior to planting. While seed collection may have included individuals originally collected to establish the common garden, clonal expansion had generated uniform cover of S. altissima, preventing isolation of any surviving individuals. Seeds of S. scoparium were purchased from Prairie Moon Nursery (Winona, MN, USA) and did not require stratification. Seeds of both species were initially grown in the greenhouse for two weeks in flats filled with soil-less potting medium (Pro-Mix BX, Premier Horticulture, Quakertown, PA, USA). This medium is not inoculated with any microbes and, while not sterile, would not possess any plant-specific microbes and was consistent across all experimental units.
Experimental plantings were done within cone-tainers (164 ml, Stuewe & Sons, Inc., Tangent, OR) filled 2/3 with the same soil-less potting medium. On top of that medium, a 10 ml layer of inoculum was added (one of 24 genotypes, live pooled, or pooled autoclaved) and mixed into the top 2 cm of potting mix. This small amount of inocula relative to growing medium (approximately 6% of volume) should minimize abiotic contributions to plant performance due to variation in soil quality across genotypes or the effects of autoclaving, but adequately represent the diversity of the soil microbial community (Howard et al. 2017). This level of inoculation should also be under the threshold for any allelopathic effects of S. altissima (Pisula and Meiners, 2010). The inoculum layer was capped with an additional 2.5 cm of soil-less potting medium to minimize potential cross-contamination. A seedling of either S. altissima or S. scoparium was transplanted into the top layer September 5, 2019. Seedlings dying within the first week were replaced; mortality following that resulted in reduced sample sizes. Initial replication was 20 for each target species/genotype combination and 40 per species for the pooled live and autoclaved control inocula. The range of final sample sizes was 15–20 (mean 19.3) for S. altissima and 14–20 (mean 17.5) for S. scoparium.
Racks of cone-tainers were placed on a single greenhouse bench and watered to saturation as necessary. After 60 d, above-ground biomass was harvested, dried at 60 °C for 4 days, and weighed. Previous work with these target species in this experimental system showed identical above- and below-ground responses to soil microbes (Peacher and Meiners 2020), and so below-ground biomass was not collected.
Data analysis
Biomass response of each target species to the soil inocula of the 24 genotypes and the autoclaved control were assessed with one-way ANOVA on inoculum identity (question 1). Genotype differences from the autoclaved control were determined from t tests of parameter estimates within the ANOVA. The overall consistency of genotype-specific soil impacts on target plant biomass of target species to inocula (24 genotypes + autoclaved control) was determined with a Pearson correlation between the means of each target species–inoculum combination (question 2). Means are used here as an estimate of the plant response to each inoculum. Similarly, the role of above-ground chemistry in influencing intraspecific variation in plant–microbe interactions was determined by relating ordination scores of S. altissima chemistry (3 NMDS axes, described above) to the average biomass of S. altissima and S. scoparium seedlings grown in each inocula using a multiple regression for each target species (question 3).
To assess whether inoculum produced from soil microbes pooled across genotypes yielded results equivalent to the average of genotypes tested individually (question 4), we conducted a two-way ANOVA of target species (two levels—S. altissima and S. scoparium) and soil handling (two levels—pooled and individual genotypes) effects on target plant biomass. For this analysis, we used the mean biomass for each target species grown in each genotypes’ soil microbiomes. Using averaged biomass instead of raw data for individual genotypes ensured that each genotype was weighed equally in the analysis (accounting for unequal sample sizes due to mortality) and resulted in a much more balanced design. All analyses were conducted in R version 3.5.1 (R Core Team 2016).
Results
There was remarkable variation in the impacts of individual genotypes’ soil microbial community on growth in both target species (question 1; Fig. 1). This variation was significant for both S. altissima (ANOVA: F24,474 = 9.63, P < 0.0001, R2 = 0.33) and S. scoparium (ANOVA: F24,426 = 4.89, P < 0.0001, R2 = 0.22). More importantly, the magnitude of soil microbial impacts of individual genotypes ranged from strongly inhibitory to being equivalent to autoclaved control inocula (neutral), to being strongly beneficial. Growth responses of S. scoparium tended to be more negative (12 genotypes) than positive (one genotype), whereas responses of S. altissima were slightly more positive (6 genotypes) than negative (three genotypes).
Variation in plant growth of Solidago altissima (top) and Schizachyrium scoparium (bottom) in response to the soil microbial communities of the 24 S. altissima genotypes. For each target species, genotypes are ordered from the smallest to largest plant mass, with closed symbols representing each of the microbial communities of the 24 S. altissima genotypes. The biomass of the control (open symbol) is indicated by the horizontal line in each panel. *Indicate genotypes with masses different from autoclaved controls. Data plotted are means ± 1 SE
The responses of S. altissima to each soil microbial community were positively related to the responses of S. scoparium (question 2; Fig. 2; r = 0.604, P = 0.0018), although there was much variation in this relationship. Part of the variation in responses between target species may be explained by differences in their relationship to foliar chemistry. Growth response to the microbial community was related to the leaf chemistry of the genotype that produced the microbial source for S. scoparium seedlings, but not for S. altissima (question 3; Table 1). In Schizachyrium scoparium, plant biomass was positively associated with two of the three axes of plant chemistry (NMDS axes 2 and 3), accounting for nearly half of its variation in growth. Foliar chemistry of genotypes with inhibitory effects on S. scoparium biomass largely overlapped with that of genotypes with neutral effects (Fig. 3). However, the one genotype’s soil microbial community that enhanced the growth of S. scoparium had a distinct foliar chemistry.
Variation in the foliar chemistry of the 24 S. altissima genotypes as determined by HPLC. Data plotted are two dimensions from an NMDS ordination of chemical composition. As only axes two and three were related to plant performance (Table 2), only those two axes are presented. Symbols represent the effects of S. altissima genotypes on S. scoparium growth, including neutral (closed circles), inhibitory (open circles), and beneficial (triangles) effects
The results of the pooled live inoculum did not reflect the average of the bioassays conducted on the genotypes individually (question 4; Table 2; Fig. 4). For both target species, pooled inocula resulted in lower plant growth than the average of each genotype’s soil microbes analyzed separately. The magnitude of the pooled inoculum’s effects on plant growth were in line with the more antagonistic soil microbial communities that were found when genotypes were tested individually.
Discussion
Our results show an appreciable level of intraspecific variation in the strength of plant–soil microbe interactions across S. altissima genotypes. Our data strongly suggest that intraspecific variation within S. altissima chemical profiles generates variation in soil microbial communities that represent a prominent source of interaction heterogeneity within communities that this species dominates.
Variation in plant chemistry and associated soil microbes is likely generated by a combination of ecological and genetic factors. For example, soil conditioning via litter inputs generate changes in soil conditions, soil microbial communities, and ecosystem function (Veen et al. 2019). Soil microbial communities of S. altissima change during succession, leading to increased herbivore resistance (Howard et al. 2020). Defensive chemistry and competitive ability in S. altissima have a strong genetic component (Genung et al. 2012; Heath et al. 2014), which can directly influence the quantity and quality of litter and soil inputs (Schweitzer et al. 2008; Uesugi and Kessler 2013). As an added level of complexity, soil microbial communities alter plant chemistry in S. altissima (Meiners et al. 2017) in ways that may additionally alter plant–plant interactions. Regardless of the underlying mechanisms, soil microbial impacts clearly varied across genotypes (Schweitzer et al. 2008; Bukowski et al. 2018; Eck et al. 2019).
The implication of genotype-specific variation in plant–microbe is that it may influence plant persistence within the community (Reynolds et al. 2003; Kardol et al. 2006; Bauer et al. 2015). Approximately half of the genotypes produced soil microbial communities that were antagonistic to S. altissima, which should promote local turnover and diversity (Liu et al. 2015; Eck et al. 2019). The soil microbial communities of 3/24 Solidago genotypes improved S. scoparium growth and were effectively neutral for another five genotypes, suggesting these S. altissima genotypes would be more invasive by other species. Variation in the plant–microbe interactions of individual genotypes may, in combination with competition and herbivore pressure, generate the successional filtering of S. altissima genotypes over time (Hartnett and Bazzaz 1985). As the individual S. altissima plants that initiated the planting were all early colonists, many of these genotypes would be displaced over time, contributing to changes in plant chemistry and soil microbial communities (Howard et al. 2020).
The correlation between the impacts of genotype’s soil microbes on plant performance in both target species suggests there are general components of the soil microbial communities that consistently impact plant species (Harrison and Bardgett 2010). As S. altissima interacts strongly with soil microbial communities (Zhang et al. 2007; Pendergast et al. 2013; Howard et al. 2020) and our genotypes varied substantially in foliar chemistry (Fig. 3), we expected variation in the resulting plant–microbe interactions. The impacts of soil microbial communities on plant performance were related to genotype foliar chemistry only for S. scoparium seedlings. This relationship appears to be due to the culturing of soil biota by the original S. altissima genotypes that generated species-specific feedbacks on S. altissima growth (Liu et al. 2015; Gehring et al. 2017; Eck et al. 2019), independent from plant chemistry. As microbes specific to S. altissima may not interact as strongly with S. scoparium, variation in soil microbes associated with foliar chemistry resulted in strong impacts of soils from most S. altissima genotypes.
While we invoke genotype-specific soil microbes as the most likely driver of our experimental results, other potential mechanisms exist. As the common garden was a small portion of a row-crop field prior to plot establishment, we expect initial soil microbial communities and abiotic conditions to be largely uniform and naïve to S. altissima when the genotypes were planted. The small size of the inoculum used in the experiment (≈ 6% by volume) should have been sufficient to transfer the soil microbial community (Howard et al. 2017), but should have contributed minimal nutrient or allelochemical inputs, well below the threshold detected in bioassays (Pisula and Meiners 2010). Therefore, the most plausible explanation of variation in target plant growth relative to autoclaved controls is the generation of genotype-specific soil microbial communities. However, this must be verified with sequencing of soil fungi and bacteria in soils cultured with S. altissima only. As genotype collections were not genetically verified, we may have collected multiple individuals from the same source genotype. However, variation in foliar chemistry shows distinct differences among most genotypes.
The variation in the strength and direction of plant–microbe interactions documented in this study presents an experimental challenge to researchers. The pooling of individual soils is often utilized in an attempt to better approximate the properties of natural, highly heterogeneous soils (Robertson et al. 1999). However, this experimental approach only works when the magnitude and direction are unchanged by sample pooling. In contrast, we found that soil pooling failed to reproduce the mean of the individual soil microbial communities’ effects on either Solidago altissima or Schizachyrium scoparium. Clearly, inocula formed by soil pooling produced markedly different impacts via plant–microbe interactions (Reinhart and Rinella 2016; Peacher and Meiners 2020). The pooling effect may be due to soil microbial beta diversity, leading to turnover among S. altissima plots and greater diversity in pooled inocula, but this effect may vary in a species-specific manner (Allen et al. 2021), generating different plant responses.
Although plant–microbe interactions for both target species ranged from positive to negative, pooled inocula generated consistently antagonistic effects on plant growth. Soil pooling may make important soil microbes present in more samples, allowing them to more frequently impact plant performance (Schnitzer et al. 2011; Peacher and Meiners 2020). In this study, pooled samples appear to have been more heavily influenced by the more antagonistic components of S. altissima soil microbial communities, yielding biased estimates of microbial impacts on both target species. Overall, plant–microbe interactions tend to be predominantly inhibitory (Kulmatiski et al. 2008), consistent with our results. However, we found variation in the strength and direction of microbial effects, representing a heterogeneous interaction landscape with potentially important implications for succession and competitive outcomes (Schnitzer et al. 2011; Pendergast et al. 2013; Mack and Bever 2014).
Ecological heterogeneity is a critical component in many plant systems, although it is often viewed as unwanted noise (Kolasa and Rollo 1991). We document biotically generated interaction heterogeneity (Pickett et al. 2000), related to inter-clonal variation in soil microbes. Interspecific differences in plant–soil microbe interactions have important impacts on population persistence and community dynamics (Van Der Heijden et al. 2008; Schnitzer et al. 2011; Liu et al. 2015; Eck et al. 2019). However, implications for intraspecific variation in plant–soil microbe interactions are difficult to predict due to the diversity of interactions between plants and soil microbes (Bukowski et al. 2018; Magnoli and Lau 2020), but may be as important as other aspects of intraspecific trait variability (Violle et al. 2012). As clonal organisms are major components of many plant communities that interact with large volumes of soil, understanding inter-clonal variation represents a critical opportunity for exploring plant–soil microbe interactions.
Data availability
All data are uploaded to https://doi.org/10.6084/m9.figshare.16783969.v1.
Code availability
This manuscript only uses standard procedures noted in the methods.
References
Allen WJ, Sapsford S, Dickie IA (2021) Soil sample pooling generates no consistent inference bias: a meta-analysis of 71 plant-soil feedback experiments. New Phytol 231:1308–1315. https://doi.org/10.1111/nph.17455
Bauer JT, Mack KML, Bever JD (2015) Plant-soil feedbacks as drivers of succession: evidence from remnant and restored tallgrass prairies. Ecosphere 6:1–12. https://doi.org/10.1890/ES14-00480.1
Bauer JT, Blumenthal N, Miller AJ, Ferguson JK, Reynolds HL (2017) Effects of between-site variation in soil microbial communities and plant-soil feedbacks on the productivity and composition of plant communities. J Appl Ecol 54:1028–1039. https://doi.org/10.1111/1365-2664.12937
Benedek K, Bálint J, Máthé I, Mara G, Felföldi T, Szabó A, Fazakas C, Albert C, Buchkowski RW, Schmitz OJ, Balog A (2019) Linking intraspecific variation in plant chemical defence with arthropod and soil bacterial community structure and N allocation. Plant Soil 444:383–397. https://doi.org/10.1007/s11104-019-04284-7
Bezemer TM, Van Dam NM (2005) Linking aboveground and belowground interactions via induced plant defenses. Trends Ecol Evol 20:617–624. https://doi.org/10.1016/j.tree.2005.08.006
Biere A, Marak HB, Van Damme JMM (2004) Plant chemical defense against herbivores and pathogens: Generalized defense or trade-offs? Oecologia 140:430–441. https://doi.org/10.1007/s00442-004-1603-6
Bolnick DI, Amarasekare P, Araújo MS, Bürger R, Levine JM, Novak M, Rudolf VHW, Schreiber SJ, Urban MC, Vasseur DA (2011) Why intraspecific trait variation matters in community ecology. Trends Ecol Evol 26:183–192
Bukowski AR, Petermann JS (2014) Intraspecific plant-soil feedback and intraspecific overyielding in Arabidopsis thaliana. Ecol Evol 4:2533–2545. https://doi.org/10.1002/ece3.1077
Bukowski AR, Schittko C, Petermann JS (2018) The strength of negative plant–soil feedback increases from the intraspecific to the interspecific and the functional group level. Ecol Evol 8:2280–2289. https://doi.org/10.1002/ece3.3755
Constancias F, Terrat S, Saby NPA, Horrigue W, Villerd J, Guillemin J-P, Biju-Duval L, Nowak V, Dequiedt S, Ranjard L, Chemidlin Prévost-Bouré N (2015) Mapping and determinism of soil microbial community distribution across an agricultural landscape. Microbiology Open 4:505–517. https://doi.org/10.1002/mbo3.255
De Witte LC, Stöcklin J (2010) Longevity of clonal plants: Why it matters and how to measure it. Ann Bot 106:859–870
Dong L-J, Sun Z-K, Gao Y, He W-M (2015) Two-year interactions between invasive Solidago canadensis and soil decrease its subsequent growth and competitive ability. J Plant Ecol. https://doi.org/10.1093/jpe/rtv003
Duell EB, Wilson GWT, Hickman KR (2016) Above- and below-ground responses of native and invasive prairie grasses to future climate scenarios. Botany 94:471–479. https://doi.org/10.1139/cjb-2015-0238
Eck JL, Stump SM, Delavaux CS, Mangan SA, Comita LS (2019) Evidence of within-species specialization by soil microbes and the implications for plant community diversity. Proc Natl Acad Sci USA 116:7371–7376. https://doi.org/10.1073/pnas.1810767116
Ehrenfeld JG, Ravit B, Elgersma K (2005) Feedback in the plant-soil system. Annu Rev Environ Resour 30:75–115. https://doi.org/10.1146/annurev.energy.30.050504.144212
Gehring CA, Sthultz CM, Flores-Rentería L, Whipple AV, Whitham TG (2017) Tree genetics defines fungal partner communities that may confer drought tolerance. Proc Natl Acad Sci USA 114:11169–11174. https://doi.org/10.1073/pnas.1704022114
Genung MA, Bailey JK, Schweitzer JA (2012) Welcome to the neighbourhood: interspecific genotype by genotype interactions in Solidago influence above- and belowground biomass and associated communities. Ecol Lett 15:65–73. https://doi.org/10.1111/j.1461-0248.2011.01710.x
Glassmire AE, Jeffrey CS, Forister ML, Parchman TL, Nice CC, Jahner JP, Wilson JS, Walla TR, Richards LA, Smilanich AM, Leonard MD, Morrison CR, Simbaña W, Salagaje LA, Dodson CD, Miller JS, Tepe EJ, Villamarin-Cortez S, Dyer LAC (2016) Intraspecific phytochemical variation shapes community and population structure for specialist caterpillars. New Phytol 212:208–219. https://doi.org/10.1111/nph.14038
Grime JP (2001) Plant strategies, vegetation processes, and ecosystem properties. Wiley, Chichester
Hahn PG, Maron JL (2016) A framework for predicting intraspecific variation in plant defense. Trends Ecol Evol 31:646–656. https://doi.org/10.1016/j.tree.2016.05.007
Harrison KA, Bardgett RD (2010) Influence of plant species and soil conditions on plant-soil feedback in mixed grassland communities. J Ecol 98:384–395. https://doi.org/10.1111/j.1365-2745.2009.01614.x
Hartnett DC, Bazzaz FA (1985) The genet and ramet population dynamics of Solidago canadensis in an abandoned field. J Ecol 74:407–413
Heath JJ, Kessler A, Woebbe E, Cipollini D, Stireman JO (2014) Exploring plant defense theory in tall goldenrod, Solidago altissima. New Phytol 202:1357–1370. https://doi.org/10.1111/nph.12755
Heinen R, van der Sluijs M, Biere A, Harvey JA, Bezemer TM (2017) Plant community composition but not plant traits determine the outcome of soil legacy effects on plants and insects. J Ecol 106:1217–1229. https://doi.org/10.1111/1365-2745.12907
Howard MM, Bell TH, Kao-Kniffin J (2017) Soil microbiome transfer method affects microbiome composition, including dominant microorganisms, in a novel environment. FEMS Microbiol Lett 364:1–8. https://doi.org/10.1093/femsle/fnx092
Howard MM, Kao-Kniffin J, Kessler A (2020) Shifts in plant-microbe interactions over community succession and their effects on plant resistance to herbivores. New Phytol. https://doi.org/10.1111/nph.16430
Huff DR, Quinn JA, Higgins B, Palazzo AJ (1998) Random amplified polymorphic DNA (RAPD) variation among native little bluestem [Schizachyrium scoparium (Michx.) Nash] populations from sites of high and low fertility in forest and grassland biomes. Mol Ecol 7:1591–1597. https://doi.org/10.1046/j.1365-294x.1998.00473.x
Jilani G, Mahmood S, Chaudhry AN, Hassan I, Akram M (2008) Allelochemicals: sources, toxicity and microbial transformation in soil—a review. Ann Microbiol 58:351–357
Kalske A, Shiojiri K, Uesugi A, Sakata Y, Morrell K, Kessler A (2019) Insect herbivory selects for volatile-mediated plant-plant communication. Curr Biol 29:3128-3133.e3. https://doi.org/10.1016/j.cub.2019.08.011
Kardol P, Martijn Bezemer T, Van Der Putten WH (2006) Temporal variation in plant-soil feedback controls succession. Ecol Lett 9:1080–1088. https://doi.org/10.1111/j.1461-0248.2006.00953.x
Kardol P, Cornips NJ, van Kempen MML, Bakx-Schotman JMT, van der Putten WH (2007) Microbe-mediated plant-soil feedback causes historical contingency effects in plant community assembly. Ecol Monogr 77:147–162. https://doi.org/10.2307/27646079
Klironomos JN (2002) Feedback with soil biota contributes to plant rarity and invasiveness in communities. Nature 417:67–70
Kolasa J, Rollo CD (1991) The heterogeneity of heterogeneity: a glossary. In: Kolasa J, Pickett STA (eds) Ecological heterogeneity. Springer, New York, pp 1–23
Kong CH, Wang P, Zhao H, Xu XH, Zhu YD (2008) Impact of allelochemical exuded from allelopathic rice on soil microbial community. Soil Biol Biochem 40:1862–1869. https://doi.org/10.1016/j.soilbio.2008.03.009
Kos M, Tuijl MAB, de Roo J, Mulder PPJ, Bezemer TM (2015) Species-specific plant-soil feedback effects on above-ground plant-insect interactions. J Ecol 103:904–914. https://doi.org/10.1111/1365-2745.12402
Kulmatiski A, Beard KH, Stevens JR, Cobbold SM (2008) Plant–soil feedbacks: a meta-analytical review. Ecol Lett 11:980–992. https://doi.org/10.1111/j.1461-0248.2008.01209.x
Lankau R (2010) Soil microbial communities alter allelopathic competition between Alliaria petiolata and a native species. Biol Invasions 12:2059–2068. https://doi.org/10.1007/s10530-009-9608-z
Liu X, Etienne RS, Liang M, Wang Y, Yu S (2015) Experimental evidence for an intraspecific Janzen-Connell effect mediated by soil biota. Ecology 96:662–671. https://doi.org/10.1890/14-0014.1
Mack KML, Bever JD (2014) Coexistence and relative abundance in plant communities are determined by feedbacks when the scale of feedback and dispersal is local. J Ecol 102:1195–1201. https://doi.org/10.1111/1365-2745.12269
Magnoli SM, Lau JA (2020) Novel plant–microbe interactions: rapid evolution of a legume–rhizobium mutualism in restored prairies. J Ecol 108:1241–1249. https://doi.org/10.1111/1365-2745.13366
McCune B, Grace JB (2002) Analysis of ecological communities. MjM Software Design, Gleneden Beach
Meiners SJ, Phipps KK, Pendergast TH, Canam T, Carson WP (2017) Soil microbial communities alter leaf chemistry and influence allelopathic potential among coexisting plant species. Oecologia 183:1155–1165. https://doi.org/10.1007/s00442-017-3833-4
Miki T (2012) Microbe-mediated plant-soil feedback and its roles in a changing world. Ecol Res 27:509–520. https://doi.org/10.1007/s11284-012-0937-5
Orozco-Aceves M, Standish RJ, Tibbett M (2015) Soil conditioning and plant-soil feedbacks in a modified forest ecosystem are soil-context dependent. Plant Soil 390:183–194. https://doi.org/10.1007/s11104-015-2390-z
Peacher MD, Meiners SJ (2020) Inoculum handling alters the strength and direction of plant-microbe interactions. Ecology. https://doi.org/10.1002/ecy.2994
Pendergast TH, Burke DJ, Carson WP (2013) Belowground biotic complexity drives aboveground dynamics: a test of the soil community feedback model. New Phytol 197:1300–1310. https://doi.org/10.1111/nph.12105
Pickett STA (1983) The absence of an Andropogon stage in old-field succession at the Hutcheson Memorial Forest. Bull Torrey Bot Club 110:533–535. https://doi.org/10.2307/2996289
Pickett STA, Cadenasso ML, Jones CG (2000) Generation of heterogeneity by organisms: creation, maintenance and transformation. In: Hutchings MJ, John EA, Stewart AJA (eds) Ecological consequences of habitat heterogeneity. Wiley, New York, pp 33–52
Pisula NL, Meiners SJ (2010) Allelopathic effects of goldenrod species on turnover in successional communities. Am Midl Nat 163:161–172
Prach K, Pyšek P, Smilauer P (1997) Changes in species traits during succession: a search for pattern. Oikos 79:201–205
R Core Team (2016) R: a language and environment for statistical computing. R Found Stat Comput, Vienna. https://www.R-project.org/
Reinhart KO, Rinella MJC (2016) A common soil handling technique can generate incorrect estimates of soil biota effects on plants. New Phytol 210:786–789. https://doi.org/10.1111/nph.13822
Reynolds HL, Packer A, Bever JD, Clay K (2003) Grassroots ecology: plant-microbe-soil interactions as drivers of plant community structure and dynamics. Ecology 84:2281–2291
Robertson GP, Crum JR, Ellis BG (1993) The spatial variability of soil resources following long-term disturbance. Oecologia 96:451–456
Robertson GP, Coleman DC, Bledsoe CS, Sollins P (eds) (1999) Standard soil methods for long-term ecological research. Oxford University Press, Oxford
Röver M, Kaiser EA (1999) Spatial heterogeneity within the plough layer: Low and moderate variability of soil properties. Soil Biol Biochem 31:175–187. https://doi.org/10.1016/S0038-0717(97)00272-1
Schnitzer SA, Klironomos JN, HilleRisLambers J, Kinkel LL, Reich PB, Xiao K, Rillig MC, Sikes BA, Callaway RM, Mangan SA, van Nes EH, Scheffer M (2011) Soil microbes drive the classic plant diversity–productivity pattern. Ecology 92:296–303. https://doi.org/10.1890/10-0773.1
Schweitzer JA, Bailey JK, Fischer DG, LeRoy CJ, Lonsdorf EV, Whitham TG, Hart SC (2008) Plant–soil–microorganism interactions: heritable relationship between plant genotype and associated soil microorganisms. Ecology 89:773–781. https://doi.org/10.1890/07-0337.1
Shikano I, Shumaker KL, Peiffer M, Felton GW, Hoover K (2017) Plant-mediated effects on an insect–pathogen interaction vary with intraspecific genetic variation in plant defences. Oecologia 183:1121–1134. https://doi.org/10.1007/s00442-017-3826-3
Smith-Ramesh LM, Reynolds HL (2017) The next frontier of plant-soil feedback research: unraveling context dependence across biotic and abiotic gradients. J Veg Sci 28:484–494. https://doi.org/10.1111/jvs.12519
Uesugi A, Kessler A (2013) Herbivore exclusion drives the evolution of plant competitiveness via increased allelopathy. New Phytol 198:916–924. https://doi.org/10.1111/nph.12172
Uesugi A, Johnson R, Kessler A (2019) Context-dependent induction of allelopathy in plants under competition. Oikos. https://doi.org/10.1111/oik.06389
Van Der Heijden MGA, Bardgett RD, Van Straalen NM (2008) The unseen majority: soil microbes as drivers of plant diversity and productivity in terrestrial ecosystems. Ecol Lett 11:296–310. https://doi.org/10.1111/j.1461-0248.2007.01139.x
Van Der Putten WH, Vet LEM, Harvey JA, Wäckers FL (2001) Linking above- and belowground multitrophic interactions of plants, herbivores, pathogens, and their antagonists. Trends Ecol Evol 16:547–554. https://doi.org/10.1016/S0169-5347(01)02265-0
van der Putten WH, Bardgett RD, Bever JD, Bezemer TM, Casper BB, Fukami T, Kardol P, Klironomos JN, Kulmatiski A, Schweitzer JA, Suding KN, Van de Voorde TFJ, Wardle DA (2013) Plant–soil feedbacks: the past, the present and future challenges. J Ecol 101:265–276. https://doi.org/10.1111/1365-2745.12054
Veen GF, Fry EL, ten Hooven FC, Kardol P, Morrien E, De Long JR (2019) The role of plant litter in driving plant–soil feedbacks. Front Environ Sci 7:1–10
Veen GF, ten Hooven FC, Weser C, Hannula SE (2021) Steering the soil microbiome by repeated litter addition. J Ecol 109:2499–2513. https://doi.org/10.1111/1365-2745.13662
Verbeek JD, Kotanen PM (2019) Soil-mediated impacts of an invasive thistle inhibit the recruitment of certain native plants. Oecologia 190:619–628. https://doi.org/10.1007/s00442-019-04435-8
Violle C, Enquist BJ, McGill BJ, Jiang L, Albert CH, Hulshof C, Jung V, Messier J (2012) The return of the variance: intraspecific variability in community ecology. Trends Ecol Evol 27:244–252. https://doi.org/10.1016/j.tree.2011.11.014
Wang XX, Hoffland E, Mommer L, Feng G, Kuyper TW (2019) Maize varieties can strengthen positive plant-soil feedback through beneficial arbuscular mycorrhizal fungal mutualists. Mycorrhiza. https://doi.org/10.1007/s00572-019-00885-3
Wang N-Q, Kong C-H, Wang P, Meiners SJ (2020) Root exudate signals in plant–plant interactions. Plant Cell Environ. https://doi.org/10.1111/pce.13892
Weber E (1998) The dynamics of plant invasions: a case study of three exotic goldenrod species (Solidago L.) in Europe. J Biogeogr 25:147–154. https://doi.org/10.1046/j.1365-2699.1998.251119.x
Werner PA, Gross RS, Bradbury IK (1980) The biology of Canadian weeds: 45. Solidago canadensis L. Can J Plant Sci 60:1393–1409. https://doi.org/10.4141/cjps80-194
Wright JP, Fridley JD (2010) Biogeographic synthesis of secondary succession rates in eastern North America. J Biogeogr 37:1584–1596. https://doi.org/10.1111/j.1365-2699.2010.02298.x
Wurst S, Ohgushi T (2015) Do plant-and soil-mediated legacy effects impact future biotic interactions? Funct Ecol 29:1373–1382
Xue W, Berendse F, Bezemer TM (2018) Spatial heterogeneity in plant–soil feedbacks alters competitive interactions between two grassland plant species. Funct Ecol 32:2085–2094. https://doi.org/10.1111/1365-2435.13124
Yip EC, Sowers RP, Helms AM, Mescher MC, De Moraes CM, Tooker JF (2019) Trade-offs between defenses against herbivores in goldenrod (Solidago altissima). Arthropod Plant Interact 13:279–287. https://doi.org/10.1007/s11829-019-09674-3
Zhang Q, Yao LJ, Yang RY, Yang XY, Tang JJ, Chen X (2007) Potential allelopathic effects of an invasive species Solidago canadensis on the mycorrhizae of native plant species. Allelopath J 20:71–78
Zhang CB, Wang J, Qian BY, Li WH (2009) Effects of the invader Solidago canadensis on soil properties. Appl Soil Ecol 43:163–169. https://doi.org/10.1016/j.apsoil.2009.07.001
Acknowledgements
Research funding was provided by the Department of Biological Sciences, Eastern Illinois University, Council on Faculty Research (SJM) and the Lewis Hanford Tiffany and Loel Zehner Tiffany Botany Graduate Research Fund (MJG). We thank Sharon N. Dubosky, Miranda Hilliard, Justin Hollinshead, Eric Micheli, Samantha Pasko, Evelin Reyes Mendez, Cole Tilford, and Jenny Trafford for assistance in the greenhouse and lab.
Funding
This work was funded by the Department of Biological Sciences, Eastern Illinois University, by a Council on Faculty Research grant to SJM, and by a Tiffany Botany Graduate Fund award to MJG.
Author information
Authors and Affiliations
Contributions
SJM conceived the project and conducted data analyses. TC and MJG performed the HPLC work. BSF, BBH, and JTC performed the greenhouse study and assembled the first draft of the manuscript. All authors edited later drafts of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare they have no conflict of interest.
Additional information
Communicated by Jaime Moyano.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Foster, B.S., Haile, B.B., Campnell, J.T. et al. Plant performance responds to intraspecific variation in soil inocula from individual Solidago clones. Plant Ecol 223, 201–212 (2022). https://doi.org/10.1007/s11258-021-01198-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11258-021-01198-2