Abstract
Purpose
While distinct soil microbiomes and individual soil microbial taxa can alter particular plant traits under highly controlled conditions, little is known about the role of particular microbial taxa and microbial functions within complex soil microbial communities for mediating plant phenotypes or if the strength of microbial mediation of plant phenotype varies among plant species or plant phenotypic traits. Examining how the plant phenotype spectrum is influenced by the taxonomic and functional composition of complex soil microbial communities allows for a more accurate understanding of the biotic environmental drivers of plant phenotype.
Methods
Using rhizosphere soil collected from field sites, we conducted a microbiome transfer glasshouse experiment to test the hypothesis that the taxonomic and functional composition of different soil microbiomes would differentially shift growth, physiological or reproductive phenotypes of three Solidago species.
Results
We found that soil microbiome inoculations influenced Solidago growth traits more than physiological and reproductive traits. We found that root growth of one of the Solidago species was negatively correlated with 77% of the indicator bacterial and fungal taxa from one of the soil microbiome treatments.
Conclusions
Soil microbial mediation of plant phenotype varies by plant traits, is not universal across plant species, and can be associated with a small number of microbial taxa. This study illustrates that specific microbial taxa within a soil microbiome are associated with shifts in plant phenotype by pinpointing important individual microbial taxa from complex field soil microbial communities.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the past 10–15 years, numerous and diverse relationships discovered between plants and the soil microbiome have shifted the long-established paradigm of plant phenotype as the sole product of interactions between a plant’s genes (G) and the abiotic environment (E) (i.e., G x E interactions; Clausen et al. 1948; Conner and Hartl 2004) to that of a ‘holobiont’ interpretation (G x G x E interactions), in which microbes at the root-soil interface serve as a reservoir of additional genes and functions for the host plant (Zilber-Rosenberg and Rosenberg 2008; Bordenstein and Theis 2015; Vandenkoornhuyse et al. 2015; Theis et al. 2016). Multiple avenues of research have informed this updated perspective. At a fine scale, single inoculation studies—mostly targeted towards crop health and production—have determined that particular individual soil microbial taxa can modify plant traits, such as growth phenotypes or flowering phenology. In isolation, bacterial genera such as Microbacterium, Pseudomonas, and Enterobacter, for example, can increase desiccation tolerance in some crop varieties by stimulating trehalose production (Vílchez et al. 2016; Niu et al. 2018). Similarly, plant growth promoting rhizobacteria assist in nutrient acquisition of nitrogen, phosphate, potassium, and iron by producing various phytohormones (Ali et al. 2009; Mishra et al. 2009; Ofek-Lalzar et al. 2014). Moreover, whole microbiome soil inoculation studies have documented microbial mediation of a diversity of plant phenotypes including physiological traits such as photosynthesis (Friesen et al. 2011; Lau and Lennon 2011; Zhu et al. 2016; Kannenberg and Phillips 2017), phenological traits such as flowering time (Wagner et al. 2014; Panke-Buisse et al. 2015) and reproductive traits such as fruit (Lau and Lennon 2011, 2012) and flower production (Lau and Lennon 2012).
At a broad scale, earlier research examining plant growth responses to bulk soil inoculations have shown that individual soil microbes collectively, referred to as a microbiome, can influence plant phenotype (Bever 1994; Bever et al. 1997; Ehrenfeld et al. 2005; Mangan et al. 2010; Putten et al. 2013). The mechanisms that underlie plant-soil feedbacks are largely attributed to the fact that phenotypically distinct plants can differentially alter both the abiotic and microbial composition of the surrounding soil, in part due to differences in organic matter turnover, and root chemical exudates that in turn favor distinct communities of microbes (Hu et al. 2018; Jones et al. 2019). The identification of core rhizosphere microbiome members for a diversity of plant taxa (Schweitzer et al. 2008; Lundberg et al. 2012; Colin et al. 2017; Yeoh et al. 2017; Fitzpatrick et al. 2018; Hugoni et al. 2018; Timm et al. 2018; Lasa et al. 2019; Singer et al. 2019) provides further evidence for the ability of different plants to recruit particular soil microbial taxa. As such, phenotypically distinct plants may, in some scenarios, alter the immediate surrounding soil (i.e., rhizosphere soil) by promoting taxonomically and/or functionally distinct microbial communities.
While findings from these research fields show that both isolated individual microbial taxa and diverse soil microbial communities can influence plant function, pinpointing the important individual microbial taxa and functions within complex soil microbial communities remains a challenge. Identifying significant individuals or functions within complex microbial communities is crucial for advancing ecology of natural ecosystems because plants in natural landscapes interact simultaneously with a multitude of beneficial, benign, and pathogenic microbes (Morris et al. 2007; Zolla et al. 2013; Putten et al. 2016). Beneficial or deleterious effects from individual taxa may be enhanced or suppressed by interactions with other nearby microbial members. Examining how the taxonomic and functional composition of soil microbial communities affects plant phenotype will allow for a more accurate understanding of the surrounding biotic environmental drivers of plant phenotype.
The goal of this study was to identify the strength of soil microbial mediation for different plant phenotypes, the consistency of these relationships among plant species, and identify specific soil microbial taxa and/or functions in complex field soil communities that are associated with particular plant phenotypes. Using field soils associated with three phenotypically distinct Solidago species, we conducted a glasshouse experiment and inoculated three Solidago species in separate treatments of each field-collected soil and microbiome. We tested the following hypotheses: 1) Soil microbiome source inoculation will differentially alter phenotypes of three Solidago species; 2) Soil microbiome source is associated with distinct taxonomic and/or functional soil microbial communities; 3) Specific microbial taxa and/or microbial functions are associated with particular Solidago phenotypes. Shifts in Solidago phenotypes between microbiome source treatments would indicate that plant traits are influenced by variation in microbial taxonomic and/or functional composition. If Solidago traits varied in response to microbiome treatments, this would indicate conditional effects of soil microbial mediation of plant phenotype. If Solidago species varied in response to microbiome treatments, this would indicate that the strength of soil microbial mediation varies by plant species. Correlations between specific microbial taxa and/or microbial functions and particular Solidago phenotypes would provide evidence for the importance of individual taxonomic or functional components within a microbiome for influencing plant phenotype.
Materials and methods
Study system
Solidago species are a model system for this study because they commonly occur across North America, with 120 species native to the United States (Semple 2016) that grow in variable habitats, with different morphologies and phenotypes. We chose to use S. caesia, S. flexicaulis, and S. gigantea in this study because they were the most abundant Solidago species found across our sampling range (northeastern TN) and vary in evolutionary history, leaf, stem, and flower morphology and habitat preference. Solidago caesia and S. flexicaulis grow in woodlands and belong to the Glomerulifloraea subgroup of Solidago (Semple 2016). Solidago gigantea grows in meadows and fields and belongs to the Triplinerviae subgroup (Semple 2016). Furthermore, previous work has found evidence for the influence of interspecific and genotypic diversity on above- and belowground biomass of S. altissima and S. gigantea (Genung et al. 2012, 2013), suggesting that some Solidago phenotypes may be mediated in part by modifications of soil biota from neighboring Solidago species.
Preliminary field surveys
To assess differences in plant phenotypes among the three Solidago species, we conducted field surveys of three geographically distinct populations of each species, all located throughout northeastern Tennessee, U.S.A. In May 2017, we measured stem height, stem base diameter, specific leaf area (SLA), and stomatal density of 15 randomly selected putative genotypes of each species (S. caesia, S. flexicaulis, and S. gigantea) in northeastern TN for a total of 45 individuals per species (Fig. 1a, Table S1). Solidago caesia and S. flexicaulis co-occur at one of the nine sites, whereas the other eight sites have single species composition (Table S1). The field survey confirmed that the three species vary in this suite of growth and physiological phenotypes (Table S2).
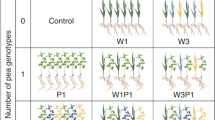
Solidago population locations, field sampling design, and experiment design for soil inoculation experiment. a Field sampling was conducted for three distinct natural field populations of Solidago gigantea, S. caesia, and S. flexicaulis in northeastern TN, U.S.A. b Growth and physiology phenotypes were measured from 15 putative genotypes at each population for each Solidago species. Rhizosphere soil (S. caesia-associated, S. flexicaulis-associated, and S. gigantea-associated) was collected from beneath each genotype and pooled at the site level. c Nursery-sourced seeds of S. caesia, S. flexicaulis, and S. gigantea and field soil were used in a 5-month glasshouse inoculation experiment. Seedlings of each species x nursery population were grown in separate treatments of soil containing inoculum from microbiome sources 1, 2, and 3 collected from the corresponding field sites. (N = 3 Solidago species × 3 seed populations × 3 microbiome sources × 3 field soil sites × 3 replicates = 243 total pots)
Soil collection and processing
To assess Hypothesis 2 that the soil microbiome sources have distinct taxonomic and/or functional microbial communities, we collected rhizosphere soil from each genotype in the field surveys by collecting soil attached to the roots of each plant (Fig. 1b). We pooled individual soil samples by field site to represent an average belowground microbiome of three soil sources (n = 3 sites per soil source). While we tried to collect soil microbes that were only associated with the rhizosphere soil of each plant species, it is likely that we also captured microbes that are representative of surrounding non-rhizosphere soil (i.e., bulk soil). As described previously, S. caesia and S. flexicaulis grow in forested habitat and S. gigantea grows in meadow/field habitat. Some climatic and edaphic soil characteristics including mean annual temperature, soil organic matter content, and soil bulk density slightly varied among the three groups of Solidago species sites (Table S3), which may be due to habitat differences. As such, we refer to the three groups of sites as soil “microbiome sources” rather than soils associated with each Solidago species. Soil samples were transported to the laboratory on ice and stored at 0 °C until analysis at the University of Tennessee, Knoxville, TN, U.S.A. A 2 g subsample of soil from each field site was stored at -80 °C for molecular analysis. We assessed the taxonomic community composition of the soils using high-throughput amplicon sequencing of the V3-V4 region of the 16S rRNA gene and the ITS2 region of the internal transcribed spacer gene regions for bacteria and fungi, respectively. Detailed molecular methods are described in Methods S1 of the Supplementary Materials.
Bioinformatics
We performed all amplicon sequence processing using the DADA2 platform. Primers were removed from the 16S and ITS sequences using the cutadapt function in conda. All further amplicon sequence processing was performed using the DADA2 pipeline (Callahan et al. 2016) Samples were normalized for sampling depth with a variance stabilizing transformation with the DESeq2 package (Love et al. 2014). We chose this method over the common practice of rarefaction because rarefaction results in loss of data by using the lowest sampling depth and it inflates variances across samples (McMurdie and Holmes 2014). Taxonomy of ASVs was assigned using the RDP (Wang et al. 2007) and UNITE (Abarenkov et al. 2010) databases for bacteria and fungi, respectively. After processing, we had 16,245 bacterial and 2,565 fungal ASVs, respectively. Additionally, we assigned fungal ASVs to functional guilds using the FUNGuild database (Nguyen et al. 2016). For analyses, we assigned taxa to one of seven broad functional guilds: arbuscular mycorrhizal fungi, ectomycorrhizal fungi, ericoid mycorrhizal fungi, endophytic fungi, plant pathogenic fungi, saprotrophic fungi, and “other.” We considered only FUNGuild assignments with a confidence ranking of “highly probable” or “probable.” Unassigned taxa were excluded from further guild-based analyses. Of the 2,565 fungal ASVs, 68% (1,741 ASVs) were assigned to a fungal guild. Of those assigned, 76% (1,328 ASVs) had a confidence ranking of “highly probable” or “probable.”
We assessed functional community composition with shotgun metagenomic sequencing, as detailed in Methods S1 of the Supplementary Materials. Sequences retrieved from shotgun metagenomic sequencing were assigned to KEGG (Kyoto Encyclopedia of Genes and Genomes) ortholog numbers using the MG-RAST online annotation tool. KEGG orthologs assign genes to microbial complexes, functional sets, and metabolic pathways and are a common tool used to describe functional attributes of microbes (Ortiz-Álvarez et al. 2018; Sorensen et al. 2019). KEGG ortholog numbers were matched to hierarchical KEGG pathways.
Glasshouse experiment
To assess Hypotheses 1 and 3 that plant phenotypes are in part mediated by the taxonomic and/or functional composition of soil microbial communities and that the strength of microbial mediation varies among plant species and phenotypic traits, we conducted a glasshouse experiment and grew S. caesia, S. flexicaulis, and S. gigantea in factorial soil inoculum treatments of each microbiome source. Seeds of each Solidago species were purchased from multiple nurseries to account for intraspecific variation in plant response to soil microbes (S. caesia: Ernst Conservation Seeds, Meadville, PA; NorthCreek Nurseries, Landenburg, PA; Michigan Wildflower Farm, Portland, MI; S. flexicaulis: Ernst Conservation Seeds, Prairie Moon Nurseries, Winona, MI; Minnesota Native Landscapes, Ostego, MN; S. gigantea: Prairie Moon Nurseries, Minnesota Native Landscapes). Seeds were refrigerated at 4 °C prior to sowing, and then were sown by population into a commercial peat moss-based, non-mycorrhizal potting mix (Premier Promix BX, containing perlite, vermiculite, and limestone). A subset of Solidago seeds did not withstand surface sterilization trials, so we did not surface sterilize the seeds used in the experiment. While it is possible that any seed-borne microbes may have impacted plant phenotype, all plants were grown in all soil treatments and exposed to the same glasshouse conditions, such that any effect of seed-borne microbes on plant phenotype should be equally distributed across treatment categories.
After approximately three weeks of growth, 54 similar-sized seedlings of each population were individually transplanted into half-gallon circular pots into soil inoculum treatments which consisted of factorial combinations of microbiome source (Microbiome source 1 vs. Microbiome source 2 vs. Microbiome source 3) (Fig. 1c). Furthermore, since soils from each field site of each microbiome source were kept separate, seeds were planted into three sites of Microbiome source 1, three sites of Microbiome source 2, and three sites of Microbiome source 3. Each pot was inoculated with approximately 5.5 g of rhizosphere soil collected from the field (< 1% of the total pot volume) to reduce effects of variation in soil nutrients on plant phenotypic responses (Troelstra et al. 2001). In total, 243 pots were established: 3 Solidago species × 3 seed populations × 3 microbiome sources (Microbiome source 1, Microbiome source 2, Microbiome source 3) × 3 field soil sites × 3 replicates = 243 total pots). Pots were randomly positioned in the glasshouse based on random number assignments. All plants were treated monthly for thrips and whiteflies throughout the experiment (0.5 tsp/gal Avid 0.15 EC insecticide, 0.5 tsp/gal AzaGuard insecticide). Plants were equally watered from above, as needed (approximately 4 days/week), and allowed to grow for 5 months in a glasshouse at the University of Tennessee.
A suite of plant phenotypes was measured during and post-experiment. Stem height and stem diameter were measured every two weeks for the first two months of growth, then at 13 weeks and at the termination of the experiment at 20 weeks. Relative growth rates were calculated from these data. For each individual plant, timing of flower bud formation (hereafter referred to as “flower bud break”) and flowering were monitored with daily surveys by recording the day of the appearance of the first distinguishable flower bud and first open flower, respectively. Prior to termination of the experiment, an average of four healthy and mature leaves were randomly selected per plant, scanned using WinFOLIA software (Regent Instruments Inc.), oven-dried at 70 °C for 72 h (Pérez-Harguindeguy et al. 2016), and weighed to calculate specific leaf area (cm2/g) (SLA). After five months of growth and regular watering, each individual was harvested and separated into shoot and root biomass and inflorescence biomass. Shoot and root tissue was weighed after 48 h of oven-drying at 60 °C. Prior to drying, roots were carefully rinsed over 2 and 0.5 mm sieves to remove lingering soil and collect all fine roots.
Statistical analyses
In the field survey, we analyzed differences in Solidago phenotypes using linear mixed-effects models with the lmer function in the lme4 package (Bates et al. 2014). We built separate mixed-effects models for each phenotype (stem height, stem diameter, SLA, and stomatal density) using Solidago species as the fixed effect and population as the random effect. When necessary, all data were transformed to conform to normality before analysis. To test Hypothesis 1 that phenotypes of each Solidago species differ when grown in soils inoculated with microbial communities associated with a different microbiome source, we built linear mixed effects models with the lmer function in the lme4 package. First, to identify traits most important to growth, physiology, and reproduction and to reduce Type I error, we tested for correlations between the ten phenotypes measured from the glasshouse experiment (relative growth rate in stem height, stem diameter at maturity, shoot biomass, root biomass, total biomass, root to shoot ratio, SLA, flower bud break, days to flower, inflorescence biomass) using the cor.test function. We chose to exclude stem diameter, total biomass, and root to shoot ratio from the analysis because they were all significantly correlated with two other growth phenotypes, shoot and root biomass (Table S6). We also chose to exclude days to flower and inflorescence biomass from the analysis because the experiment ended before the majority of S. gigantea individuals flowered. Relative growth rate, shoot biomass, root biomass, SLA, and flower bud break were included in the analysis.
Multiple models were used to assess Hypothesis 1. Separate models were built for the five phenotypes (relative growth rate, shoot biomass, root biomass, SLA, and timing of flower bud formation). When necessary, all data was transformed to conform to normality before analysis. First, to test that differences in soil microbial community composition have a general effect on plant phenotypes regardless of plant species, we built linear mixed effects models with microbiome source as a fixed effect and Solidago species, seed population, and field soil site as random effects. To individually identify how each Solidago species responded to soil microbial community composition, we separated the dataset by Solidago species and built individual linear mixed effects models for each Solidago species with microbiome source as a fixed effect and seed population and field soil site as random effects. Although the random effects in our models include fewer levels than the generally accepted minimum range of 5–6 levels, we argue that philosophically these variables (field site, Solidago seed population) are random effects and are therefore admissible in the models even with insufficient levels. For all models, we used the Anova function to calculate ANOVA tables using Type II sums of squares, with significance assessed for each fixed effect using Wald X2 statistics. If any of the fixed effects were significant, we conducted post hoc Tukey contrasts using the TukeyHSD function.
To test Hypothesis 2 that each microbiome source is associated with distinct taxonomic and/or functional soil microbial communities, we took multiple approaches. First, we assessed microbial diversity across microbiome source by calculating hill numbers based on ASV counts and unique KEGG identities using the hill_div function in the hilldiv package (Alberdi and Gilbert 2019). Hill numbers serve as effective numbers of diversity that provide more intuitive estimates of diversity compared to traditional diversity indices based on entropy (Chao et al. 2014). We calculated hill numbers for all orders of diversity at q = 0, q = 1, and q = 2, and tested for significant differences in hill numbers between microbiome source at each order of diversity using the div_test function in the hilldiv package. A diversity order q = 0 provides raw richness by weighting rare taxa the same as abundant taxa and thus not accounting for species’ abundances. A diversity order q = 1 weights ASVs by their abundance but without disproportionately favoring abundant taxa. A diversity order q = 2 overweighs abundant ASVs.
Second, we created Bray–Curtis distance matrices for microbial taxonomic and functional composition of the nine field soils. To assess variation in community composition of bacteria, fungi, and KEGGs across microbiome source, we conducted PERMANOVA analysis with 9,999 permutations using the adonis function in the vegan package (Oksanen et al. 2019). Prior to conducting PERMANOVA we confirmed homogeneity of dispersion across microbiome source with the betadisper function in the vegan package. We then performed a distance-based redundancy analysis (db-RDA) using the dbrda function in the vegan package to assign variation in composition of bacteria, fungi, and KEGGs to microbiome source and geographic location. We conducted three individual db-RDAs for bacteria, fungi, and KEGG composition. We used the anova.cca function in the vegan package to assess the cumulative significance of microbiome source and geographic location on community composition. We partitioned the variation in composition with respect to microbiome source and geographic location using the varpart function in the vegan package. To visualize composition of bacteria, fungi, and KEGGs among soil microbiome source, we used principal coordinate analysis (PCoA) for ordination based on the Bray–Curtis distance matrices.
We then performed indicator species analysis with the multipatt function in the indicspecies package (Cáceres and Legendre 2009) to identify particular bacteria, fungi, and KEGGs that are uniquely highly associated with each microbiome source. Indicator taxa were determined using a p-value threshold of 0.05. Because the FUNGuild data set contained a high amount of zero counts, we built individual zero-inflated models for each fungal guild using the glmmTMB function in the glmmTMB package (Brooks et al. 2017). We specified microbiome source as the fixed effect, count total per soil sample (i.e. field site) as the random effect, zi formula as soil microbiome source, and family as poisson. For all models, we used the Anova function in the car package (Fox et al. 2013) to calculate analysis of variance (ANOVA) tables using Type II sums of squares, with significance assessed for microbiome source using Wald X2 statistics. If the effect of microbiome source was significant, we conducted post hoc Tukey contrasts using the emmeans function in the emmeans package (Lenth et al. 2020) and the cld function in the multcomp package (Hothorn et al. 2008).
To test Hypothesis 3 that specific microbial taxa and/or functions are associated with particular Solidago phenotypes, we assessed the effect of variation in microbial indicator taxa composition on Solidago phenotypes that responded to microbiome source treatment. Since no KEGG identities were identified as indicators across the three microbiome sources, subsequent analyses were conducted only with bacterial and fungal indicator taxa. Using a db-RDA, we assigned variation in composition of bacterial and fungal indicator taxa to the three microbiome sources and geographic location. We then extracted the axes scores from the db-RDA model. For each phenotype, we built a linear model that included the two axes (CAP1, CAP2) from the db-RDA model as fixed effects. A significant relationship between db-RDA axes and plant phenotypes would indicate that differences in the community of bacterial and fungal indicator taxa associated with each microbiome source are associated with shifts in plant phenotype. To pinpoint individual bacterial and fungal indicator taxa that may be associated with particular plant phenotypes, we built linear models to test for correlations between the relative abundance of each bacterial and fungal indicator taxon and each phenotype that showed significant responses to the axes of variation from the indicator species db-RDA model.
All analyses were performed in R (R Core 2020). Boxplot, and linear regression figures were made with the ggplot2 package (Wickham 2016). Ordination figures were made with the phyloseq package (McMurdie and Holmes 2013). Heatmap figures were made with the Heatplus (Ploner 2020) and gplots (Warnes et al. 2020) packages. Individuals figures were aggregated with the patchwork package (Pedersen 2020).
Results
Plant phenotype responses to soil microbiome sources (glasshouse experiment)
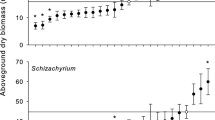
While the three Solidago species overall varied significantly in relative growth rate, shoot and root biomass, and flower bud break, only root biomass differed by microbiome source (\(\chi\) 2 = 6.14, p = 0.04) (Table 1). Among all three Solidago species, there was 29% greater root biomass production when plants were grown in inoculum from microbiome source 1 relative to microbiome source 3 (Tukey post hoc: p = 0.05) (Fig. 2a). In partial support of Hypothesis 1, the species-specific models produced marginally significant results. Phenotypic responses to microbiome source inoculum slightly varied by Solidago species and by phenotype. Solidago caesia shoot biomass differed among microbiome source treatments, whereas no S. flexicaulis or S. gigantea phenotypes differed among microbiome source treatments (Table 2). Solidago caesia produced 8.9% more shoot biomass when grown in inoculum from microbiome source 2 relative to microbiome source 3 (Tukey post hoc: p = 0.09) (Fig. 2b), indicating that different soil microbiomes can shift some plant traits.
Community composition among soil microbiome sources
Across the three soil microbiome sources, we identified over 16,000 bacterial ASVs and 2,500 fungal ASVs. Taxonomic and functional diversity of soil microbial communities did not vary by microbiome source at any order of diversity (Tables S4, S5). In partial support of Hypothesis 2, whole microbiomes did not differ in taxonomic or genetic pathway composition among the microbiome sources, but distinct indicator taxa were identified for each microbiome source. PERMANOVA analysis revealed that taxonomic and functional composition of soil microbial communities did not vary significantly by microbiome source (Bacteria: F = 1.04, p = 0.37; Fungi: F = 1.10, p = 0.18; KEGGs: F = 0.92, p = 0.55) (Fig. 3). The db-RDA revealed that microbiome source and geographic location cumulatively accounted for less than 10% of variation in bacteria, fungi, and KEGG composition (Bacteria: adj. R2 = -0.013; Fungi: adj. R2 = 0.053; KEGGs: adj. R2 = -0.068). Microbiome source accounted for less than 5% of variation in bacteria, fungi, and KEGG composition (Bacteria: adj. R2 = 0.011; Fungi: adj. R2 = 0.025; KEGGs: adj. R2 = -0.021). Similarly, geographic location also accounted for less than 5% of variation in bacteria, fungi, and KEGG composition (Bacteria: adj. R2 = 0.026; Fungi: adj. R2 = 0.033; KEGGs: adj. R2 = -0.02). The dissimilarity in microbial community composition among the S. gigantea-associated soils (Fig. 3) could be due to the fact that two of the soil sites (BRC and FB) are geographically clustered away from the other site (LPT) (Fig. 1a). Due to their proximity to one another, these two sites share more similar edaphic and climatic characteristics than with the third site, which could explain the differences in microbial community composition between LPT and the other two sites.
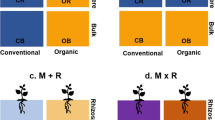
Indicator species analysis identified significant bacterial and fungal indicator taxa for each microbiome source. In total, 77 bacterial ASVs (out of 16,245 detected; 0.5%) and eight fungal ASVs (out of 2,565 detected; 0.3%) were identified as indicator taxa among the three microbiome sources (Fig. 4, Tables S7, S8). Twenty-nine bacterial ASVs were uniquely shared among microbiome sources 1 and 2, whereas microbiome source 3 uniquely shared only six bacterial ASVs with either microbiome source 1 or 2. Fungal guilds were assigned to approximately 68% of the fungal ASVs. Of those assigned to a guild, approximately 76% had a confidence ranking of “probable” or “highly probable.” Out of the five fungal guilds (arbuscular mycorrhizal fungi, ectomycorrhizal fungi, ericoid mycorrhizal fungi, endophytic fungi, and plant pathogenic fungi), ericoid mycorrhizal fungi (\(\chi\) 2 = 11.29, p = 0.004) and endophytic fungi \((\chi\) 2 = 20.14, p < 0.0001) differed significantly among the microbiome sources (Table S9). Both microbiome sources 1 and 2 had approximately 1.5- and twofold greater abundance of ericoid mycorrhizal fungi and endophytic fungi, respectively, than microbiome source 3 (Fig. 5d, e).
Heatmap of relative abundance of the 77 bacterial ASVs produced from indicator species analysis across the three microbiome sources. Rows represent individual ASVs. Columns represent soil from individual field sites. Taxa of each microbiome source with relative abundance 0.05 (5%) or greater are color coded by phylum
Mean abundance of a arbuscular mycorrhizal fungi (AMF), b ectomycorrhizal fungi (ECM), c plant pathogenic fungi, d ericoid mycorrhizal fungi, and e endophytic fungi. Emmeans are reported on the log scale. Data shown are pooled across samples (i.e. field sites). Data that do not share letters are significantly different from one another (p < 0.05)
KEGG composition overall did not differ among the three microbiome sources, indicating functional redundancy among soil microbial communities. Of the 122 pathways identified, less than a quarter accounted for more than 1% of relative abundance of all KEGGs among the three microbiome sources (Table S10). Of this subset, 70% were pathways involved in metabolism of either energy (in the form of nitrogen, methane, sulfur, and oxidative phosphorylation), amino acids, carbohydrates, or lipids. The most abundant pathways across the three microbiome sources were two pathways for ATP-binding cassette (ABC) transporters, which accounted for 20% of the relative abundance. No KEGG pathways were detected as indicators among the microbiome sources.
Correlations between soil microbiome composition and plant phenotypes
In support of Hypothesis 3, individual microbial taxa were associated with specific Solidago phenotypes. We only examined shoot biomass of S. caesia as it was the only phenotype that responded to microbiome source treatments. Axes of variation in composition of the bacteria indicator taxa were significantly correlated with S. caesia shoot biomass (Axis CAP1: F = 7.63, p = 0.03). Relative abundance of 77% (20 out of 26) of the bacterial and fungal indicator taxa of microbiome source 3 were significantly negatively correlated with S. caesia shoot biomass when S. caesia was grown in inoculum of microbiome source 3 (Fig. 6, Table S11). Although S. caesia produced slightly more shoot biomass when grown in inoculum of microbiome source 2 compared to that of microbiome source 3 (Fig. 2b), none of the eight bacterial indicator taxa or the one fungal indicator taxon of microbiome source 2 were significantly positively correlated with S. caesia shoot biomass.
Discussion
Identifying ways in which the taxonomic and functional composition of the soil microbiome influences plant phenotype is a central challenge for understanding the overall importance of complex soil microbial communities on plant function, as well as how changes to soil microbial communities may in turn affect plant function. While recent studies have explored the importance of both whole soil microbiomes and individual soil microbial taxa on particular plant phenotypes, it is also crucial to understand if and how particular taxa and functions within complex soil microbial communities influence a broad spectrum of plant phenotypes and if these relationships are consistent across multiple plant species. In this study we compared the taxonomic and functional composition of soil microbial communities associated with three phenotypically distinct and naturally occurring Solidago species and their habitats (referred to as soil microbiome source above) to understand how variation in soil microbiome composition alters plant phenotype. We subsequently conducted a microbiome transfer glasshouse experiment to test for plant phenotypic shifts in response to soil microbiome source inoculum. We found that soil microbiome taxonomic variation can shift some plant phenotypes and that this response varied by plant species, with some species more responsive to microbial taxonomic variation than others. Specifically, we identified indicator bacterial and fungal taxa associated with each microbiome source, some of which were correlated with shifts in plant growth responses. We found that microbiome source slightly altered growth traits for one of the three Solidago species. Lastly, we found that in this study plant growth traits were more likely to be influenced by variation in soil microbial communities than physiological or reproductive traits. Together, these findings show that soil microbial mediation of plant phenotype 1) varies by plant traits, 2) is not consistent across plant species, and 3) can be influenced, in part, by a small number of microbial taxa.
Soil microbial mediation of plant phenotype varies by plant phenotype and plant species
We found that soil microbial communities can shift some plant phenotypes, but that the strength of microbially-mediated phenotypic plasticity varies by plant phenotype and by plant species. Across all three Solidago species, plants grown in inocula from microbiome source 1 produced more root biomass compared to plants grown in inocula from microbiome source 3. Differences in root growth may be due to microbial effects on nutrient availability. Nutrient availability was likely very similar for all three Solidago species at the beginning of the experiment because the same amount of inoculum was used in each pot. However, over the course of the experiment, the microbial communities in the inocula may have altered nutrient availability. Specifically, microbial communities from microbiome source 1 may have consumed more nutrients than microbial communities from microbiome source 3, and as a result, plants grown in microbiome source 1 inocula may have been more nutrient limited than plants grown in microbiome source 3 inocula. This may be an explanation for why all three species overall produced more root biomass when grown in microbiome source 1 microbial communities compared to when grown with microbiome source 3 microbial communities. Alternatively, microbiome source 1 microbial communities may be more beneficial for root growth than S. gigantea microbial communities. In fact, microbial communities from microbiome sources 1 and 2 had greater amounts of endophytic and ericoid mycorrhizal fungi relative to microbiome source 3. While no S. flexicaulis or S. gigantea phenotypes responded to microbiome source treatments, S. caesia produced slightly more shoot biomass in microbiome source 2 inocula relative to microbiome source 3 inocula.
It should be noted that the volume of soil inoculum in this study accounted for < 1% of the total pot soil volume. Since increasing the amount of soil inoculum can alter the microbial composition in the resulting pot soil (Howard et al. 2017), it is possible that we only identified the plant traits that are most strongly microbially mediated, while more subtle effects on other traits were not detected. This could in part explain why our results contradict findings of other studies that multiple plant physiological and reproductive traits, including photosynthetic rates, stomatal conductance, leaf water potential, chlorophyll content, leaf nitrogen content, specific leaf area (Lau and Lennon 2011; Kannenberg and Phillips 2017; Hahn et al. 2018; Siefert et al. 2018), flowering phenology (Wagner et al. 2014; Panke-Buisse et al. 2015), numbers of flowers, seeds, and fruits (Bauer and Flory 2011; Lau and Lennon 2011, 2012; Dudenhöffer et al. 2018) and seed weight (Bauer and Flory 2011) can be responsive to soil microbial manipulations.
In all of the forementioned studies, the microbial inoculum accounted for at least 1% of the total soil volume, with the median being 3.25%. One study applied inoculum that accounted for 71% (Siefert et al. 2018). In our study we found that neither SLA nor timing of flower bud formation responded to soil microbiome source treatments. While there is a breadth of research on plant-soil biota relationships, the majority of these studies focus on plant growth associated traits. To determine if the relative strength of soil microbial mediation is similar for plant growth and non-growth traits, more studies like this one are needed that simultaneously test for effects on growth, physiological, and reproductive traits.
Soil microbial mediation of plant phenotype can be influenced in part by a small number of microbial taxa
Despite similarity in overall microbiome composition among the three soil microbiome sources, we identified specific bacterial and fungal indicator taxa of each microbiome source. In general, Proteobacteria taxa were more highly abundant in microbiome sources 1 and 2 than in microbiome source 3, whereas Acidobacteria taxa were more highly abundant in microbiome source 3 than in microbiome sources 1 and 2. Within the Proteobacteria phylum, indicator taxa of microbiome sources 1 and 2 spanned a larger diversity of taxonomic orders including Rhizobiales, Rhodospirillales, Burkholderiales, and Xanthomonadales relative to those of microbiome source 3 which comprised the orders of Rhizobiales and Myxococcales. These findings highlight the importance of taxonomic resolution when assessing the role of the soil microbiome on plant phenotype as root-associated soil is known to contain some of the highest microbial biodiversity on Earth (Curtis et al. 2002; Buée et al. 2009; Berendsen et al. 2012). In this study, we identified over 16,000 bacterial ASVs and over 2,500 fungal ASVs among the three microbiome sources. These findings suggest that identifying microbial differences among focal groups may require focusing on specific indicator taxa that have high affiliation with the plant species or field site rather than overall microbiome composition.
We found that differences in indicator taxa among separate soil microbial communities may contribute to shifts in plant phenotype even though this relationship is likely not consistent across plant species. We found that Solidago caesia produced significantly less vegetative biomass in inocula of microbiome source 3 compared to inocula of microbiome source 2, and that the relative abundance of 77% of the indicator taxa of microbiome source 3 were correlated with decreases in S. caesia shoot production. These indicator bacteria included mostly members of the Acidobacteria and Actinobacteria phyla in addition to members of the Bacteriodetes, Firmicutes, and Proteobacteria phyla. One of the indicator fungal taxa belongs to the Dothideomycetes class of the Ascomycota phylum, and various Dothideomycetes taxa have been identified as plant pathogens (Ohm et al. 2013). The other indicator fungal taxon belongs to the Chytridiomycota phylum. Despite greater shoot production in microbiome source 2 inocula, none of the microbiome source 2 indicator taxa were correlated with positive shifts of S. caesia shoot production.
Although these indicator taxa account for a very small proportion of the total microbial communities identified in this study, it is notable that out of the thousands of ASVs identified in microbiome source 2, twenty bacterial and fungal ASVs explained on average 7% of the variation in root biomass when S. caesia was grown in inocula of microbiome source 2. These results add further support for the importance of rare microbial taxa (Pedrós-Alió 2012; Shade et al. 2014; Hol et al. 2015; Jousset et al. 2017). More specifically, these results suggest that individual soil microbial taxa may be involved, in part, with mediating some plant traits.
Although terminology in microbial ecology is continually evolving and some terms, such as keystone taxa are used liberally, we note that the indicator taxa as we have identified them in this study are not necessarily keystone taxa. Microbial keystone taxa are often described as taxa that have a sizeable impact on the microbiome such that their removal can considerably alter microbiome structure and functioning (Banerjee et al. 2018). From the scope of this study, it is unknown whether these microbial indicators that are correlated to differences in vegetative biomass would have similar influence if the surrounding microbial composition differed. Moreover, some sort of removal experiment would be needed to accurately determine if these indicators are true keystone taxa for plant vegetative biomass (Röttjers and Faust 2019). What is clear is, however, is that this study builds upon considerable previous findings that influential individual microbial taxa can be identified from complex microbiomes. Given the immense and often overwhelming complexity of natural soil microbiomes, this study demonstrates the utility of this approach of examining correlations between indicator microbial taxa and plant traits. Identifying individual microbial taxa that are associated with shifts in plant traits can pinpoint particular microbial taxa to target for further experiments that test causative mechanisms of plant trait variation. While the correlative relationships we identified in this study are context-specific to these particular microbial taxa and plant species, the approach used here can be applied to any plant-microbial system.
Other factors governing soil microbial-plant interactions
While we did not assess microbial dormancy in this study, it likely plays a significant role in soil microbial mediation of plant phenotype. Dormancy, in which individuals undergo a temporary reduced state of metabolic activity, has long been hypothesized to be widespread among microorganisms because it allows them to cope with environmental variability, particularly when conditions are unfavorable (Stevenson 1977; Lau and Lennon 2011). Differentiating between active and dormant microbial taxa requires examining the active ribosomal RNA in addition to the total ribosomal DNA. Since we only used rDNA-based techniques in this study, our inferences are limited to microbial taxa that are potentially active. However, despite the fact that the indicator taxa accounted for a very small amount of the diversity of each microbiome source, evidence suggests that rare taxa may confer particular importance within a microbiome. A previous study examining proportions of rRNA to rDNA in temperate lakes found that rare taxa had a higher probability of being metabolically active than common taxa (Jones and Lennon 2010). Combined with the observation that soil microbiome diversity is primarily comprised of rare taxa (Elshahed et al. 2008), our findings and those from Jones and Lennon (2010) highlight the importance of examining how less abundant (i.e., rare) taxa within soil microbial communities may influence plant phenotype. Other microbial interactions within the soil environment also likely influenced the plant trait variation we observed, such as differences in microbial growth rates, differences in decomposition via extracellular enzymes, and changes to the microbial communities due to conditioning from the plants. However, testing these mechanisms was outside of the scope of this study.
In this study, we found high similarity in soil microbial function (i.e. KEGG pathway composition) among the three microbiome sources, suggesting functional redundancy in which the absence of one or more microbial species does not greatly affect the functioning of the whole microbial community because the same functions are fulfilled by many different taxa (Fernández et al. 1999; Louca et al. 2016, 2018). Functional redundancy is widespread in microbial systems (Tringe et al. 2005; Allison and Martiny 2008; Bezemer et al. 2010; Martiny et al. 2013; Nelson et al. 2016). The relatively small number of known functions associated with soil microbial communities indicates that much is still unknown about microbial functional genes that may translate to physiological differences among microbial taxa. While the metagenomic methods we employed in this study can be used to infer potential microbial functions, other molecular approaches such as metatranscriptomics can identify particular microbial genes that are being actively transcribed (Moran 2009; Carvalhais et al. 2012; Damon et al. 2012), and in doing so provide a more accurate representation of the microbial functions that characterize a particular soil microbiome.
Conclusions
Soil microbes represent a largely overlooked but often important biotic factor for influencing plant phenotype. This may be one of the first few studies to examine how taxonomic and functional gene composition of complex soil microbial communities influence a suite of multiple plant phenotypes across multiple plant species in natural, unmanaged ecosystems. Our study shows that soil microbiomes and specific taxa within complex soil microbial communities can alter some plant phenotypes, but that not all plant species, even those belonging to the same genus, will respond to soil microbial communities in the same manner. Thus, the belowground biotic environment is just one of a host of important biotic factors that can mediate plant phenotype, in addition to plant genetic background and abiotic environmental variation. While the findings from this study are founded in ecology theory, identifying the nuances of relationships between soil microbes and plant phenotype has wide-scale applications. Substantial efforts to engineer core rhizosphere microbiomes to optimize plant production signify the need to identify functional linkages between soil microbial communities and plants (Bakker et al. 2012; Busby et al. 2017; Wallenstein 2017; Qiu et al. 2019). While soil biota may not be a universal solution to enhance some plant phenotypes, specific microbial taxa may be harnessed to improve plant growth and plant tolerance to adverse environmental conditions. This study illustrates that it is possible to identify specific microbial taxa within a complex soil microbial community that are associated with shifts in some plant phenotypes.
Data availability
All data and accompanying analysis scripts are available on GitHub at https://github.com/kbeals2/Plant-Soil_Conditionality_of_soil_microbial_mediation.
References
Abarenkov K, Nilsson RH, Larsson K-H et al (2010) The UNITE database for molecular identification of fungi – recent updates and future perspectives. New Phytol 186:281–285. https://doi.org/10.1111/j.1469-8137.2009.03160.x
Alberdi A, Gilbert MTP (2019) hilldiv: an R package for the integral analysis of diversity based on Hill numbers. bioRxiv 545665. https://doi.org/10.1101/545665
Ali B, Sabri AN, Ljung K, Hasnain S (2009) Auxin production by plant associated bacteria: impact on endogenous IAA content and growth of Triticum aestivum L. Lett Appl Microbiol 48:542–547. https://doi.org/10.1111/j.1472-765X.2009.02565.x
Allison SD, Martiny JBH (2008) Resistance, resilience, and redundancy in microbial communities. PNAS 105:11512–11519. https://doi.org/10.1073/pnas.0801925105
Bakker MG, Manter DK, Sheflin AM et al (2012) Harnessing the rhizosphere microbiome through plant breeding and agricultural management. Plant Soil 360:1–13. https://doi.org/10.1007/s11104-012-1361-x
Banerjee S, Schlaeppi K, van der Heijden MGA (2018) Keystone taxa as drivers of microbiome structure and functioning. Nat Rev Microbiol 16:567–576. https://doi.org/10.1038/s41579-018-0024-1
Bates D, Mächler M, Bolker B, Walker S (2014) Fitting linear mixed-effects models using lme4. arXiv:14065823 [stat]
Bauer JT, Flory SL (2011) Suppression of the woodland herb senna hebecarpa by the invasive grass Microstegium vimineum. Am Midl Nat 165:105–115. https://doi.org/10.1674/0003-0031-165.1.105
Berendsen RL, Pieterse CMJ, Bakker PAHM (2012) The rhizosphere microbiome and plant health. Trends Plant Sci 17:478–486. https://doi.org/10.1016/j.tplants.2012.04.001
Bever JD (1994) Feeback between plants and their soil communities in an old field community. Ecology 75:1965–1977. https://doi.org/10.2307/1941601
Bever JD, Westover KM, Antonovics J (1997) Incorporating the Soil community into plant population dynamics: the utility of the feedback approach. J Ecol 85:561–573. https://doi.org/10.2307/2960528
Bezemer TM, Fountain MT, Barea JM et al (2010) Divergent composition but similar function of soil food webs of individual plants: plant species and community effects. Ecology 91:3027–3036. https://doi.org/10.1890/09-2198.1
Bordenstein SR, Theis KR (2015) Host biology in light of the microbiome: ten principles of holobionts and hologenomes. PLoS Biol 13:e1002226. https://doi.org/10.1371/journal.pbio.1002226
Brooks M E, Kristensen K, Benthem K J ,van, et al (2017) glmmTMB balances speed and flexibility among packages for zero-inflated generalized linear mixed modeling. R Journal 9:378. https://doi.org/10.32614/RJ-2017-066
Buée M, Reich M, Murat C et al (2009) 454 Pyrosequencing analyses of forest soils reveal an unexpectedly high fungal diversity. New Phytol 184:449–456. https://doi.org/10.1111/j.1469-8137.2009.03003.x
Busby PE, Soman C, Wagner MR et al (2017) Research priorities for harnessing plant microbiomes in sustainable agriculture. PLoS Biol 15:e2001793. https://doi.org/10.1371/journal.pbio.2001793
Cáceres MD, Legendre P (2009) Associations between species and groups of sites: indices and statistical inference. Ecology 90:3566–3574. https://doi.org/10.1890/08-1823.1
Callahan BJ, McMurdie PJ, Rosen MJ et al (2016) DADA2: high-resolution sample inference from Illumina amplicon data. Nat Methods 13:581–583. https://doi.org/10.1038/nmeth.3869
Carvalhais LC, Dennis PG, Tyson GW, Schenk PM (2012) Application of metatranscriptomics to soil environments. J Microbiol Methods 91:246–251. https://doi.org/10.1016/j.mimet.2012.08.011
Chao A, Gotelli NJ, Hsieh TC et al (2014) Rarefaction and extrapolation with Hill numbers: a framework for sampling and estimation in species diversity studies. Ecol Monogr 84:45–67. https://doi.org/10.1890/13-0133.1
Clausen J, Keck DD, Hiesey WM (1948) Experimental studies on the nature of species. III. Environresponses of climatic races of Achillea. Experimental studies on the nature of species III Environresponses of climatic races of Achillea
Colin Y, Nicolitch O, Van Nostrand JD et al (2017) Taxonomic and functional shifts in the beech rhizosphere microbiome across a natural soil toposequence. Sci Rep 7:9604. https://doi.org/10.1038/s41598-017-07639-1
Conner JK, Hartl DL (2004) A primer of ecological genetics. Volume 425. Sunderland, MA: Sinaeur Associates
Curtis TP, Sloan WT, Scannell JW (2002) Estimating prokaryotic diversity and its limits. Proc Natl Acad Sci 99:10494–10499. https://doi.org/10.1073/pnas.142680199
Damon C, Lehembre F, Oger-Desfeux C et al (2012) Metatranscriptomics reveals the diversity of genes expressed by eukaryotes in forest soils. PLoS One 7:e28967. https://doi.org/10.1371/journal.pone.0028967
Dudenhöffer J-H, Ebeling A, Klein A-M, Wagg C (2018) Beyond biomass: Soil feedbacks are transient over plant life stages and alter fitness. J Ecol 106:230–241. https://doi.org/10.1111/1365-2745.12870
Ehrenfeld JG, Ravit B, Elgersma K (2005) feedback in the plant-soil system. Annu Rev Environ Resour 30:75–115. https://doi.org/10.1146/annurev.energy.30.050504.144212
Elshahed MS, Youssef NH, Spain AM et al (2008) Novelty and uniqueness patterns of rare members of the soil biosphere. AEM 74:5422–5428. https://doi.org/10.1128/AEM.00410-08
Fernández A, Huang S, Seston S et al (1999) How stable is stable? Function versus community composition. Appl Environ Microbiol 65:3697–3704. https://doi.org/10.1128/AEM.65.8.3697-3704.1999
Fitzpatrick CR, Copeland J, Wang PW et al (2018) Assembly and ecological function of the root microbiome across angiosperm plant species. PNAS 115:E1157–E1165. https://doi.org/10.1073/pnas.1717617115
Fox J, Friendly M, Weisberg S (2013) Hypothesis tests for multivariate linear models using the car package. R Journal 5:39. https://doi.org/10.32614/RJ-2013-004
Friesen ML, Porter SS, Stark SC et al (2011) Microbially mediated plant functional traits. Annu Rev Ecol Evol Syst 42:23–46. https://doi.org/10.1146/annurev-ecolsys-102710-145039
Genung MA, Bailey JK, Schweitzer JA (2012) Welcome to the neighbourhood: interspecific genotype by genotype interactions in Solidago influence above- and belowground biomass and associated communities. Ecol Lett 15:65–73. https://doi.org/10.1111/j.1461-0248.2011.01710.x
Genung MA, Bailey JK, Schweitzer JA (2013) The afterlife of interspecific indirect genetic effects: genotype interactions alter litter quality with consequences for decomposition and nutrient dynamics. PLoS One 8. https://doi.org/10.1371/journal.pone.0053718
Hahn PG, Bullington L, Larkin B, et al (2018) Effects of short- and long-term variation in resource conditions on soil fungal communities and plant responses to soil biota. Front Plant Sci 9. https://doi.org/10.3389/fpls.2018.01605
Hol WHG, de Boer W, de Hollander M et al (2015) Context dependency and saturating effects of loss of rare soil microbes on plant productivity. Front Plant Sci. https://doi.org/10.3389/fpls.2015.00485
Hothorn T, Bretz F, Westfall P (2008) Simultaneous inference in general parametric models. Biom J 50:346–363. https://doi.org/10.1002/bimj.200810425
Howard MM, Bell TH, Kao-Kniffin J (2017) Soil microbiome transfer method affects microbiome composition, including dominant microorganisms, in a novel environment. FEMS Microbiol Lett 364. https://doi.org/10.1093/femsle/fnx092
Hu L, Robert CAM, Cadot S et al (2018) Root exudate metabolites drive plant-soil feedbacks on growth and defense by shaping the rhizosphere microbiota. Nat Commun 9:2738. https://doi.org/10.1038/s41467-018-05122-7
Hugoni M, Luis P, Guyonnet J, Haichar F el Z (2018) Plant host habitat and root exudates shape fungal diversity. Mycorrhiza 28:451–463. https://doi.org/10.1007/s00572-018-0857-5
Jones P, Garcia BJ, Furches A, et al (2019) Plant host-associated mechanisms for microbial selection. Front Plant Sci 10. https://doi.org/10.3389/fpls.2019.00862
Jones SE, Lennon JT (2010) Dormancy contributes to the maintenance of microbial diversity. Proc Natl Acad Sci 107:5881–5886. https://doi.org/10.1073/pnas.0912765107
Jousset A, Bienhold C, Chatzinotas A et al (2017) Where less may be more: how the rare biosphere pulls ecosystems strings. ISME J 11:853–862. https://doi.org/10.1038/ismej.2016.174
Kannenberg SA, Phillips RP (2017) Soil microbial communities buffer physiological responses to drought stress in three hardwood species. Oecologia 183:631–641. https://doi.org/10.1007/s00442-016-3783-2
Lasa AV, Fernández-González AJ, Villadas PJ et al (2019) Metabarcoding reveals that rhizospheric microbiota of Quercus pyrenaica is composed by a relatively small number of bacterial taxa highly abundant. Sci Rep 9:1695. https://doi.org/10.1038/s41598-018-38123-z
Lau JA, Lennon JT (2011) Evolutionary ecology of plant–microbe interactions: soil microbial structure alters selection on plant traits. New Phytol 192:215–224. https://doi.org/10.1111/j.1469-8137.2011.03790.x
Lau JA, Lennon JT (2012) Rapid responses of soil microorganisms improve plant fitness in novel environments. PNAS 109:14058–14062. https://doi.org/10.1073/pnas.1202319109
Lenth R, Buerkner P, Herve M, et al (2020) Emmeans: estimated marginal means, aka least-squares means. https://cran.r-project.org/web/packages/emmeans/index.html
Louca S, Jacques SMS, Pires APF et al (2016) High taxonomic variability despite stable functional structure across microbial communities. Nat Ecol Evol 1:1–12. https://doi.org/10.1038/s41559-016-0015
Louca S, Polz MF, Mazel F et al (2018) Function and functional redundancy in microbial systems. Nat Ecol Evol 2:936–943. https://doi.org/10.1038/s41559-018-0519-1
Love M, Anders S, Huber W (2014) Differential analysis of count data–the DESeq2 package. Genome Biol 15:10–1186
Lundberg DS, Lebeis SL, Paredes SH et al (2012) Defining the core Arabidopsis thaliana root microbiome. Nature 488:86–90. https://doi.org/10.1038/nature11237
Mangan SA, Schnitzer SA, Herre EA et al (2010) Negative plant–soil feedback predicts tree-species relative abundance in a tropical forest. Nature 466:752–755. https://doi.org/10.1038/nature09273
Martiny AC, Treseder K, Pusch G (2013) Phylogenetic conservatism of functional traits in microorganisms. ISME J 7:830–838. https://doi.org/10.1038/ismej.2012.160
McMurdie PJ, Holmes S (2013) phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS One 8:e61217. https://doi.org/10.1371/journal.pone.0061217
McMurdie PJ, Holmes S (2014) Waste not, want not: why rarefying microbiome data is inadmissible. PLoS Comput Biol 10:e1003531. https://doi.org/10.1371/journal.pcbi.1003531
Mishra PK, Mishra S, Selvakumar G et al (2009) Enhanced soybean (Glycine max L.) plant growth and nodulation by Bradyrhizobium japonicum-SB1 in presence of Bacillus thuringiensis-KR1. Acta Agric Scand Sect B Soil Plant Sci 59:189–196. https://doi.org/10.1080/09064710802040558
Moran MA (2009) Metatranscriptomics: eavesdropping on complex microbial communities. Microbe Mag 4:329–335. https://doi.org/10.1128/microbe.4.329.1
Morris WF, Hufbauer RA, Agrawal AA et al (2007) Direct and interactive effects of enemies and mutualists on plant performance: a meta-analysis. Ecology 88:1021–1029. https://doi.org/10.1890/06-0442
Nelson MB, Martiny AC, Martiny JBH (2016) Global biogeography of microbial nitrogen-cycling traits in soil. PNAS 113:8033–8040. https://doi.org/10.1073/pnas.1601070113
Nguyen NH, Song Z, Bates ST et al (2016) FUNGuild: an open annotation tool for parsing fungal community datasets by ecological guild. Fungal Ecol 20:241–248. https://doi.org/10.1016/j.funeco.2015.06.006
Niu X, Song L, Xiao Y, Ge W (2018) Drought-tolerant plant growth-promoting rhizobacteria associated with foxtail millet in a semi-arid agroecosystem and their potential in alleviating drought stress. Front Microbiol 8. https://doi.org/10.3389/fmicb.2017.02580
Ofek-Lalzar M, Sela N, Goldman-Voronov M et al (2014) Niche and host-associated functional signatures of the root surface microbiome. Nat Commun 5:1–9. https://doi.org/10.1038/ncomms5950
Ohm RA, Feau N, Henrissat B, Schoch CL, Horwitz BA, et al. (2013) Diverse lifestyles and strategies of plant pathogenesis encoded in the genomes of eighteen Dothideomycetes fungi. PLoS Pathogens 8. https://doi.org/10.1371/journal.ppat.1003037
Oksanen J, Blanchet FG, Friendly M et al (2019) (2019) vegan: Community Ecology Package. R Package Version 2:5–6
Ortiz-Álvarez R, Fierer N, de los Ríos A, et al (2018) Consistent changes in the taxonomic structure and functional attributes of bacterial communities during primary succession. ISME J 12:1658–1667. https://doi.org/10.1038/s41396-018-0076-2
Panke-Buisse K, Poole AC, Goodrich JK et al (2015) Selection on soil microbiomes reveals reproducible impacts on plant function. ISME J 9:980–989. https://doi.org/10.1038/ismej.2014.196
Pedersen TL (2020) Patchwork: the composer of plots. https://cran.rproject.org/web/packages/patchwork/index.html
Pedrós-Alió C (2012) The rare bacterial biosphere. Ann Rev Mar Sci 4:449–466
Pérez-Harguindeguy N, Díaz S, Garnier E et al (2016) Corrigendum to: new handbook for standardised measurement of plant functional traits worldwide. Aust J Bot 64:715–716. https://doi.org/10.1071/bt12225_co
Ploner A (2020) Heatplus: heatmaps with row and/or column covariates and colored clusters. https://bioconductor.org/packages/release/bioc/html/Heatplus.html
Qiu Z, Egidi E, Liu H et al (2019) New frontiers in agriculture productivity: optimised microbial inoculants and in situ microbiome engineering. Biotechnol Adv 37:107371. https://doi.org/10.1016/j.biotechadv.2019.03.010
R Core Team (2020) R: a language and environment for statistical computing, Vienna, Austria. https://www.R-project.org/
Röttjers L, Faust K (2019) Can we predict keystones? Nat Rev Microbiol 17:193–193. https://doi.org/10.1038/s41579-018-0132-y
Schweitzer JA, Bailey JK, Fischer DG et al (2008) Plant–soil–microorganism interactions: heritable relationship between plant genotype and associated soil microorganisms. Ecology 89:773–781. https://doi.org/10.1890/07-0337.1
Semple J (2016) (11) (PDF) An intuitive phylogeny and summary of chromosome number variation in the goldenrod genus Solidago (Asteraceae: Astereae). In: ResearchGate. https://www.researchgate.net/publication/303366007_An_intuitive_phylogeny_and_summary_of_chromosome_number_variation_in_the_goldenrod_genus_Solidago_Asteraceae_Astereae. Accessed 24 Mar 2020
Shade A, Jones SE, Caporaso JG, et al (2014) Conditionally rare taxa disproportionately contribute to temporal changes in microbial diversity. mBio 5:e01371–14. https://doi.org/10.1128/mBio.01371-14
Siefert A, Zillig KW, Friesen ML, Strauss SY (2018) Soil microbial communities alter conspecific and congeneric competition consistent with patterns of field coexistence in three Trifolium congeners. J Ecol 106:1876–1891. https://doi.org/10.1111/1365-2745.13042
Singer E, Bonnette J, Kenaley SC et al (2019) Plant compartment and genetic variation drive microbiome composition in switchgrass roots. Environment Microbiol Rep 11:185–195. https://doi.org/10.1111/1758-2229.12727
Sorensen JW, Dunivin TK, Tobin TC, Shade A (2019) Ecological selection for small microbial genomes along a temperate-to-thermal soil gradient. Nat Microbiol 4:55–61. https://doi.org/10.1038/s41564-018-0276-6
Stevenson LH (1977) A case for bacterial dormancy in aquatic systems. Microb Ecol 4:127–133. https://doi.org/10.1007/BF02014283
Theis KR, Dheilly NM, Klassen JL, et al (2016) Getting the hologenome concept right: an eco-evolutionary framework for hosts and their microbiomes. mSystems 1. https://doi.org/10.1128/mSystems.00028-16
Timm CM, Carter KR, Carrell AA, et al (2018) Abiotic stresses shift belowground populus-associated bacteria toward a core stress microbiome. mSystems 3. https://doi.org/10.1128/mSystems.00070-17
Tringe SG, von Mering C, Kobayashi A et al (2005) Comparative metagenomics of microbial communities. Science 308:554–557. https://doi.org/10.1126/science.1107851
Troelstra SR, Wagenaar R, Smant W, Peters B, a. M, (2001) Interpretation of bioassays in the study of interactions between soil organisms and plants: involvement of nutrient factors. New Phytol 150:697–706. https://doi.org/10.1046/j.1469-8137.2001.00133.x
van der Putten WH, Bardgett RD, Bever JD et al (2013) Plant–soil feedbacks: the past, the present and future challenges. J Ecol 101:265–276. https://doi.org/10.1111/1365-2745.12054
van der Putten WH, Bradford MA, Brinkman EP et al (2016) Where, when and how plant–soil feedback matters in a changing world. Funct Ecol 30:1109–1121. https://doi.org/10.1111/1365-2435.12657
Vandenkoornhuyse P, Quaiser A, Duhamel M et al (2015) The importance of the microbiome of the plant holobiont. New Phytol 206:1196–1206
Vílchez JI, García-Fontana C, Román-Naranjo D, et al (2016) Plant drought tolerance enhancement by Trehalose production of desiccation-tolerant microorganisms. Front Microbiol 7. https://doi.org/10.3389/fmicb.2016.01577
Wagner MR, Lundberg DS, Coleman-Derr D et al (2014) Natural soil microbes alter flowering phenology and the intensity of selection on flowering time in a wild Arabidopsis relative. Ecol Lett 17:717–726. https://doi.org/10.1111/ele.12276
Wallenstein MD (2017) Managing and manipulating the rhizosphere microbiome for plant health: a systems approach. Rhizosphere 3:230–232. https://doi.org/10.1016/j.rhisph.2017.04.004
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naïve Bayesian classifier for rapid assignment of rRNA Sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267. https://doi.org/10.1128/AEM.00062-07
Warnes G, Bolker B, Bonebakker L, et al (2020) Gplots: various R programming tools for plotting data. https://cran.r-project.org/web/packages/gplots/index.html
Wickham H (2016) Ggplot2: elegant graphics for data analysis. https://cran.r-project.org/web/packages/ggplot2/index.html
Yeoh YK, Dennis PG, Paungfoo-Lonhienne C et al (2017) Evolutionary conservation of a core root microbiome across plant phyla along a tropical soil chronosequence. Nat Commun 8:215. https://doi.org/10.1038/s41467-017-00262-8
Zhu X, Song F, Liu S, Liu F (2016) Arbuscular mycorrhiza improve growth, nitrogen uptake, and nitrogen use efficiency in wheat grown under elevated CO2. Mycorrhiza 26:133–140. https://doi.org/10.1007/s00572-015-0654-3
Zilber-Rosenberg I, Rosenberg E (2008) Role of microorganisms in the evolution of animals and plants: the hologenome theory of evolution. FEMS Microbiol Rev 32:723–735
Zolla G, Badri DV, Bakker MG et al (2013) Soil microbiomes vary in their ability to confer drought tolerance to Arabidopsis. Appl Soil Ecol 68:1–9. https://doi.org/10.1016/j.apsoil.2013.03.007
Acknowledgements
We thank members of the Schweitzer, Bailey, Lebeis, and Kivlin labs at the University of Tennessee especially Shannon Bayliss, Liam Mueller, Michael Van Nuland, Ian Ware, Alix Pfennigwerth, Rachel Wooliver, and Jessica Moore for feedback on experimental design and analysis. We thank Liam Mueller and James Fordyce in particular for feedback on statistical analysis. We thank the Schweitzer and Bailey lab undergraduate volunteers for help with data collection. We give special thanks to Aaron Floden and Terrell Carter for assistance with fieldwork, and Jeff Martin for assistance in the glasshouse. We also thank those who provided guidance and advice on microbial genomic processing and analysis including Veronica Brown, Stephanie Kivlin, Mike Lee, Robert Murdoch, Michael Van Nuland, Ian Ware, Andrew Willems, and staff of Michigan State University Genomics Core facility. Lastly, we would like to thank the two anonymous reviewers who provided critical feedback on the manuscript.
Funding
This research was supported by the Evolutionary, Ecological, or Conservation Genomics Research Award from the American Genetic Association granted to Kendall Beals and the University of Tennessee Student-Faculty Research Award granted to Kendall Beals and Jennifer Schweitzer.
Author information
Authors and Affiliations
Contributions
Kendall Beals and Jennifer Schweitzer designed the study. Joseph Bailey and Sarah Lebeis provided important feedback on the study design. Kendall Beals conducted the field, glasshouse, and lab work. Sarah Lebeis provided crucial guidance on lab work and bioinformatics analysis. Kendall Beals performed the data analysis. Kendall Beals led the writing of the manuscript and all authors contributed significantly to the drafts and gave final approval for publication.
Corresponding author
Ethics declarations
Competing interests
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Responsible Editor: Jeff R. Powell.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Beals, K.K., Lebeis, S.L., Bailey, J.K. et al. Conditionality of soil microbial mediation of Solidago plant phenotype: indicator taxa within complex microbiomes influence some, but not all Solidago traits. Plant Soil 485, 281–298 (2023). https://doi.org/10.1007/s11104-022-05828-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-022-05828-0