Abstract
Vanilloids, including capsaicin and eugenol, are ligands of transient receptor potential channel vanilloid subfamily member 1 (TRPV1). Prolonged treatment with vanilloids triggered the desensitization of TRPV1, leading to analgesic or antinociceptive effects. Caenorhabditis elegans (C. elegans) is a model organism expressing vanilloid receptor orthologs (e.g., OSM-9 and OCR-2) that are associated with behavioral and physiological processes, including sensory transduction. We have shown that capsaicin and eugenol hamper the nocifensive response to noxious heat in C. elegans. The objective of this study was to perform proteomics to identify proteins and pathways responsible for the induced phenotype and to identify capsaicin and eugenol targets using a thermal proteome profiling (TPP) strategy. The results indicated hierarchical differences following Reactome Pathway enrichment analyses between capsaicin- and eugenol-treated nematodes. However, both treated groups were associated mainly with signal transduction pathways, energy generation, biosynthesis and structural processes. Wnt signaling, a specific signal transduction pathway, is involved following treatment with both molecules. Wnt signaling pathway is noticeably associated with pain. The TPP results show that capsaicin and eugenol target OCR-2 but not OSM-9. Further protein–protein interaction (PPI) analyses showed other targets associated with enzymatic catalysis and calcium ion binding activity. The resulting data help to better understand the broad-spectrum pharmacological activity of vanilloids.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
All organisms rely on a set of regulatory and protective behaviors to ensure survival. Following exposure to a noxious mechanical, thermal or chemical stimulus, animals perform a protective withdrawal reflex to ensure organismal integrity and prevent cellular damage [1,2,3,4,5,6]. In mammals, the presence of tissue-damaging stimuli is sensed by primary afferent nociceptors with transient receptor potential (TRP) channels, including TRPV1, which is widely associated with the generation of nociceptive and painful responses. TRP channels are involved in the transduction of polymodal stimuli, including temperature, mechanical and osmotic changes, electrical charge, light, hypotonic swelling and chemical stimuli, including xenobiotics and endogenous lipids. Capsaicin, the pungent ingredient in the chili pepper, is a well-known molecule that activates TRPV1 [7,8,9]. Other vanilloids displayed similar properties, including eugenol [10,11,12]. Upon sustained exposition, TRPV1 agonists elicit receptor desensitization, leading to antinociceptive effects and alleviation of pain [7, 13,14,15].
Caenorhabditis elegans (C. elegans) is a powerful animal model system to study the dynamics of molecular networks and diseases [16,17,18,19]. Adult C. elegans consists of 959 cells, of which 302 are neurons, making this model attractive to study nociception at the physiological and molecular levels. C. elegans is particularly useful for the study of nociception because it exhibits a well-defined and reproducible nocifensive behavior, involving a reversal and change in direction away from the noxious stimuli (i.e., noxious heat), as we have already described [6, 20, 21]. Several C. elegans genes encode TRP ion channel proteins with important sequence homologies to mammalian TRP channels, including TRPVs [22, 23]. Specifically, several TRPV analogs (e.g., OSM-9 and OCR-1-4) were identified. Further studies have demonstrated that TRPV analog channels share similar activation and regulatory mechanisms with their mammalian counterparts [1, 23,24,25]. Recently, we revealed that capsaicin [26] and eugenol [27] impeded the nocifensive response of C. elegans to noxious heat (i.e., 32–35 °C) following sustained exposure, and the antinociceptive effect was reversed 6 h post-exposure. Additionally, we have shown that capsaicin’s target was the C. elegans transient receptor potential channel OCR-2, but the results for eugenol were not conclusive. However, very little is known regarding the molecular or cellular processes that are modulated after C. elegans exposure to capsaicin or eugenol. Network biology has become a promising approach, particularly using proteomics and bioinformatics. Using this approach, drug efficacy and toxicity can be assessed across various scales of complexity, including the molecular, pathway, cellular and organism levels. Proteomics is instrumental in deciphering the architecture and dynamics of molecular networks to better understand diseases and drug effects [28,29,30]. Additionally, thermal proteome profiling (TPP) is an unbiased method allowing the quantification of drug-target engagement. The identification of the protein targets of drugs is essential to decipher biochemical or biophysical activities in cells or organisms and is a key step in drug discovery. While proteomics is focused on measuring relative protein abundance for functional analysis, TPP provides a novel approach to gather complementary information on target engagement that is not available in classical differential proteomic experiments [28, 29, 31]. We believe that combining these proteomic and bioinformatic approaches will allow us to systematically analyze interactions among proteins, and the phenotypic consequences can drive the development of innovative therapeutic strategies to alleviate pain. The objective of this study was to (1) perform proteomics to identify proteins and pathways modulated following C. elegans exposure to capsaicin or eugenol and (2) use TPP to further characterize the modes of action of capsaicin and eugenol in C. elegans.
Materials and Methods
Chemicals and Reagents
All chemicals and reagents were obtained from Fisher Scientific (Fair Lawn, NJ, USA) or Millipore Sigma (St. Louis, MO, USA). Capsaicin (Cap) and eugenol (Eug) were purchased from Toronto Research Chemicals (North York, ON, CAN).
C. elegans Strains
The N2 (Bristol) isolate of C. elegans was used as a reference strain. N2 (Bristol) was obtained from the Caenorhabditis Genetics Center (CGC), University of Minnesota (Minneapolis, MN, USA). Strains were maintained and manipulated under standard conditions as described [32, 33]. Nematodes were grown and kept on nematode growth medium (NGM) agar at 22 °C in a Thermo Scientific Heratherm refrigerated incubator. Experiments and analyses were performed at room temperature unless otherwise noted.
C. elegans Pharmacological Manipulations
Cap or Eug was dissolved in Type 1 Ultrapure Water at a concentration of 25 µM. The solution was warmed for brief periods combined with vortexing and sonication for several min to completely dissolve the compound. C. elegans were isolated and washed according to the protocol outlined by Margie et al. [32]. After 72 h of feeding and growing on 92 × 16 mm petri dishes with NGM, the nematodes were exposed to Cap or Eug solution (25 µM). An aliquot of 7 mL of Cap or Eug solution was added to produce a 2–3 mm solution film (the solution was partly absorbed by NGM) so that the nematodes were swimming in solution. C. elegans were exposed to Cap or Eug for 60 min. Additionally, one experimental group included nematodes exposed to elevated temperature (33 °C) for 1 h in absence of Cap or Eug. The temperature selection was based on previous electrophysiological study of OSM-9 and OCR-2 [34]. Three independent biological replicates (n = 3) per condition was performed.
Protein Extraction from C. elegans
Nematodes (with or without Cap or Eug exposure) were collected in liquid medium, centrifuged at 1000×g for 10 min, collected and thoroughly washed. Nematodes were resuspended in 8 M urea/100 mM TRIS–HCl buffer (pH 8) containing cOmplete™ protease inhibitor cocktail (Roche), and aliquots were transferred to reinforced 1.5 mL homogenizer tubes containing 500 μm glass bead homogenizer tubes containing 50 mg glass beads. The samples were homogenized using a Bead Mill Homogenizer (Fisherbrand) with 3 bursts of 60 s at a speed of 5 m/s. The homogenates were centrifuged at 12,000×g for 10 min. The protein concentration for each homogenate was determined using a Bradford assay. Two hundred micrograms of protein were extracted using ice-cold acetone precipitation (1:5, v/v). The protein pellet was dissolved in 100 µL of 50 mM TRIS–HCl buffer (pH 8), and the solution was mixed with a Disruptor Genie at maximum speed (2800 rpm) for 15 min and sonicated to improve the protein dissolution yield. Proteins were reduced with 20 mM dithiothreitol (DTT), and the reaction was performed at 90 °C for 15 min. Then, proteins were alkylated with 40 mM iodoacetamide (IAA), and the reaction was allowed to proceed while protected from light at room temperature for 30 min. Then, 5 µg of proteomic-grade trypsin was added, and the reaction was allowed to proceed at 37 °C for 24 h. Protein digestion was quenched by adding 10 µL of a 1% trifluoroacetic acid (TFA) solution. Samples were centrifuged at 12,000×g for 10 min, and 100 µL of the supernatant was transferred into injection vials for analysis. The protein extraction and further analyses were performed using 3 independent biological replicates (n = 3) per condition.
Thermal Proteome Profiling to Identify Targets
Thermal proteome profiling (TTP) using a range of ligand concentrations (TTP-CCR) allows proteome-wide studies of drug-protein interaction [31]. Specifically, it allows us to study target engagement. The principles behind TPP are that proteins can be thermally stabilized following interaction with a ligand, leading to a higher apparent solubility following thermal stress. Nematodes were collected in liquid medium, centrifuged at 1000×g for 10 min, isolated and thoroughly washed. The nematodes were then resuspended in 50 mM phosphate buffer (pH 7.4) containing cOmplete™ protease inhibitor cocktail (Roche), and aliquots were transferred to reinforced 1.5 mL homogenizer tubes containing 50 mg glass beads (500 μm). The samples were homogenized using a Bead Mill Homogenizer (Fisherbrand) with 3 bursts of 60 s at a speed of 5 m/s. The homogenates were centrifuged at 12,000×g for 10 min. The protein concentration in each homogenate was determined using a Bradford assay. Two hundred micrograms of protein from the lysate was exposed to a series of Cap or Eug concentrations (i.e., 0, 1, 2, 5 and 10 µM) and then heated at 60 °C for 5 min using a heated block. Samples were centrifuged at 12,000×g for 10 min, and 100 µL aliquots of the supernatant (e.g., soluble fraction) were transferred into 1.5 mL microtubes. Reduction, alkylation and digestion were performed as described in the previous section. Three independent biological replicates (n = 3) per condition was performed.
Proteomic Analysis
The Ultra-High-Performance Liquid Chromatography (UHPLC) system was a Thermo Scientific Vanquish FLEX UHPLC system (San Jose, CA, USA). Chromatography was performed using a gradient elution along with a Thermo Biobasic C18 150 × 1 mm microbore column with a particle size of 5 μm. The initial mobile phase conditions consisted of acetonitrile and water (both fortified with 0.1% formic acid) at a ratio of 5:95. From 0 to 3 min, the ratio was maintained at 5:95. From 3 to 63 min, a linear gradient was applied up to a ratio of 40:60, which was maintained for 2 min. The mobile phase composition ratio was reverted to the initial conditions, and the column was allowed to re-equilibrate for 25 min. The flow rate was fixed at 50 µL/min, and 5 µL samples were injected. A Thermo Scientific Q Exactive Plus Orbitrap Mass Spectrometer (San Jose, CA, USA) was interfaced with the UHPLC system using a pneumatic assisted heated electrospray ion source. Nitrogen was used for the sheath and auxiliary gases, which were set at 10 and 5 arbitrary units, respectively. The heated ESI probe was set to 4000 V, and the ion transfer tube temperature was set to 200 °C. Mass spectrometry (MS) detection was performed in positive ion mode and operating in TOP-10 Data Dependent Acquisition (DDA) mode. A DDA cycle entailed one MS1 survey scan (m/z 400–1500) acquired at 70,000 resolution (full width at half maximum; FWHM) and precursor ions meeting user-defined criteria for charge state (i.e., z = 2, 3 or 4), monoisotopic precursor intensity (dynamic acquisition of MS2-based TOP-10 most intense ions with a minimum 1 × 104 intensity threshold). Precursor ions were isolated using the quadrupole (1.5 Da isolation width), activated by Higher-energy Collisional Dissociation (HCD) at 28 Normalized Collision Energy (NCE) and fragment ions were detected in the Orbitrap at a resolution of 17,500 (FWHM). Data were processed using Thermo Proteome Discoverer (version 2.4) in conjunction with the SEQUEST HT engine using the default settings unless otherwise specified. The identification of peptides and proteins with SEQUEST HT was performed based on the reference proteome extracted from UniProt (C. elegans taxon identifier 6239) as FASTA sequences. Parameters were set as follows: MS1 tolerance of 10 ppm; MS2 mass tolerance of 0.02 Da for Orbitrap detection; enzyme specificity was set as trypsin with two missed cleavages allowed; carbamidomethylation of cysteine was set as a fixed modification; and oxidation of methionine and acetylation of the protein N-terminus were treated as variable modifications. The minimum peptide length was set to six amino acids. Datasets were further analyzed with Percolator. Peptide-spectrum matches (PSMs) were filtered at a 1% false discovery rate (FDR) threshold. For protein quantification and comparative analysis, we used the peak integration feature of the Proteome Discoverer 2.4 software. For each identified protein, the average ion intensity of unique peptides was used for protein abundance.
Bioinformatics
The abundance ratio (log2): (experimental group)/(N2 control), abundance ratio adjusted p-value: (experimental group)/(N2 control) and accession columns were extracted from the datasets generated by Proteome Discoverer 2.4. The adjusted p-values were determined using an ANOVA followed by a Tukey's HSD post hoc test. Volcano plots were generated using all identified and quantified proteins with both a log2 ratio and adjusted p-value. Proteins with an adjusted p-value ≥ 0.05 were not used for further analysis. Additionally, only proteins with an absolute log2 ratio ≥ 1.0 were used for bioinformatics analysis. Venn diagram analysis was used to determine the degree of overlaps between experimental groups. Reactome enrichment analyses were performed using Metascape [35], in which all differentially expressed proteins (DEPs) were used (absolute log2 ratio ≥ 1.0; adjusted p-value ≤ 0.05). Further pathway analysis was performed using ClueGO [36] and Cytoscape [37] using the Reactome pathway databases. For TTP-CCR experiments, Kyoto Encyclopedia of Genes and Genomes (KEGG) and Reactome enrichment analyses were performed using Metascape and ClueGO for proteins with a fold change ≥ 4 compared to 0 µM (control samples). The statistical test used to determine the enrichment score for KEGG and Reactome enrichment analyses was based on a right-sided hypergeometric distribution with multiple testing correction (Benjamini & Hochberg (BH) method) [38].
Results and Discussion
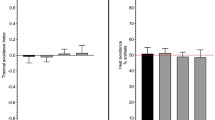
As shown in our previous studies [26, 27, 39], capsaicin, eugenol and other vanilloids displayed noteworthy antinociceptive effects in C. elegans following controlled and prolonged exposure. As displayed in Figure S1 (supplementary file), the data revealed a significant antinociceptive effect after a 1 h exposure to Cap or Eug compared with the WT (CTL) group. These molecules effectively hamper the C. elegans nocifensive response to noxious heat. However, the molecular mechanism and targets still need to be better described. Therefore, we used mass spectrometry-based proteomics, network biology as well as TPP for the identification of drug–target interactions in C. elegans to decipher the molecular mechanism associated with the antinociceptive effects observed. Label-free proteomic investigations were performed on C. elegans exposed to heat (33 °C), Cap (25 µM) and Eug (25 µM) for 1 h. Figure 1 shows volcano plots to illustrate the differential abundances of proteins, with the x-axis representing the log2 ratio and the y-axis plotting − log10 (adjusted p-value). The boxes represent a twofold change and adjusted p-value ≤ 0.05. Several DEPs were identified, including 45 upregulated and 100 downregulated proteins following noxious heat exposure (Fig. 1A), 93 upregulated and 203 downregulated proteins following Cap exposure (Fig. 1B) and 75 upregulated and 187 downregulated proteins following Eug exposure (Fig. 1C). Table S1, S2 and S3 (supplementary file) unveils the fold changes and adjusted p-values for all DEP’s identified. Proteins with a single high-scoring peptide hit were not excluded based on recommendations [40].
Visualization of proteomic results using volcano plots. Volcano plots illustrate the differential abundances of proteins, with the x-axis showing the log2 ratio with respect to the control group (N2 maintained at 22 °C) and the y-axis representing − 1 × log10 (p value). The boxes represent log twofold change and above with adjusted p-value ≤ 0.05. The adjusted p-values were determined by a one-way ANOVA with post-hoc Tukey HSD. A C. elegans exposed for 1 h to 33 °C were compared to individuals maintained at 22 °C. B C. elegans exposed for 1 h to 25 µM Cap were compared to individuals maintained at 22 °C. C. C. elegans exposed for 1 h to 25 µM Eug were compared to individuals maintained at 22 °C. D. Venn diagram representing the degree of DEP overlap between the three experimental conditions under investigation
Analyzing lists of DEPs using Venn diagrams (Fig. 1D) reveal several specific DEPs for each experimental groups but also an important degree of overlapping, specially between the C. elegans exposed to Cap and Eug. This was expected since they both interact with vanilloid receptor orthologs. Although Cap appears to target more specifically OCR-2 according to our previous study based on phenotyping [26, 27]. Thus, leading potentially to differential biological and molecular effects.
Reactome pathway enrichment analyses are presented in Figs. 2, 3 and 4. Hierarchical clustering of Reactome pathways shows distant proximity between the Cap- and Eug-treated nematodes. It is already known that Cap is a more pungent vanilloid than Eug. In mammals, initial exposure to Cap produces burning and irritant effects. Cap produces pain, an effect not reported for Eug [10]. This differential effect is associated with the pungent nature of Cap evoking a significant increase in ion influx, such as calcium, increasing neural activity. However, interestingly, abundant literature reports have shown Cap can relieve pain [41,42,43,44,45]. Although the molecular mechanisms are very different [46], this paradox is also observed with opioids, a well-established analgesic drug class, which also induces hyperalgesia [47, 48]. Thus, the differences noted between Cap- and Eug-triggered responses may help in understanding their respective therapeutic uses and highlight the importance of understanding their respective mechanisms of action. As reveal in Figs. 2A and 3A, only nematodes exposed to noxious heat (33˚C) showed enrichment for Heat shock factor protein 1 (HSF1)-dependent transactivation (R-CEL-3371571), a pathway directly associated with Cellular response to heat stress (R-CEL-3371556) as shown in Fig. 4A. Remarkably, Cap and Eug exposed nematodes revealed enrichment for Signaling by WNT (CEL-195721) closely related to Signal transduction (R-CEL-162582) as shown in Fig. 3B and 3C. Remarkably, our previous study suggested that Wnt signaling is triggered by the interaction of Resiniferatoxin (RTX) with C. elegans vanilloid receptors [39]. RTX is a very potent capsaicin analog. Recent evidence has shown that Wnt signaling underlies the pathogenesis of neuropathic pain [49] and it is a potential therapeutic target for the treatment of chronic pain [50]. It is important to note, following Cap treatment, β-catenin phosphorylation cascade (R-CEL-196299) is significantly enriched. Wnt/β-catenin signaling is central component in the development of chronic and neuropathic pain [51].
Reactome enriched terms. A. Analysis was performed with all DEPs (∣FC∣ ≥ 2 and adjusted p-value ≤ 0.05). B. Analysis was performed using only the specific DEPs for each experimental condition under investigation (∣FC∣ ≥ 2 and adjusted p-value ≤ 0.05). Enrichment analyses were performed using Metascape, and color densities were derived from − log10 (adjusted p-value)
Parent node to root node analysis of enriched Reactome terms derived from only specific DEPs for three experimental conditions under investigation. A C. elegans exposed for 1 h to 33 °C; B C. elegans exposed for 1 h to 25 µM Cap; C C. elegans exposed for 1 h to 25 µM Eug. Node analyses were performed using ClueGO and CluePedia
Another process significantly enriched following Cap treatment is G beta:gamma signaling through CDC42 (R-CEL-8964616). A recent study shows protein 42 homolog (CDC42) regulates Schwann cell proliferation and migration through Wnt/β-catenin and p38 MAPK signaling pathway following the induction of sciatic nerve injury leading to neuropathic pain [52]. Furthermore, CDC42 is associated with the dysregulation of the innate immune system [53]. Inflammageing and chonic or neuropathic pain are strongly associated [54]. Besides, one strongly enriched process for Cap-stimulated nematodes is the L13a-mediated translational silencing of Ceruloplasmin expression (R-CEL-156827). Despite this process is still not well characterized, it has been suggested to be involved in the translational control of effectors playing an important role in disbalance of pro-inflammatory cytokines leading to disturbance of pro-inflammatory signaling pathways [55]. Interestingly, our recent study demonstrates that nerve injury has a long-lasting impact on systemic inflammation contributing to the development of neuropathic pain [54]. Another interesting process specific to Cap exposure is Axon guidance (R-CEL-422474) since it is related to chemotaxis. This is most likely related to the initial pungent effect of Cap. Interestingly, it is a process related to Robo receptors [56]. Slit-Robo signals are involved in synaptic plasticity contributing to the development of pain [57].
Venn diagrams analysis (Fig. 1D) permitted to isolate specific DEPs for each treatment and further Reactome enrichment analysis was performed as display in Fig. 2B. The Table S4 (supplementary file) lost all proteins expressed differentially specific to each treatment. Once more, hierarchical clustering of Reactome pathways shows distant proximity between the Cap- and Eug-treated nematodes. One striking enrich pathway for Eug treated nematodes is Signaling by VEGF (R-CEL-194138). Vascular Epithelial Growth Factors (VEGF) is one of the most important mediators participating in inflammatory pain [58]. Consequently, alterations in the VEGF system, characterized by expression changes of its mediators, could modulate the sensitivity of the primary nociceptors including TRP channels [59]. Interestingly, Cap and Eug treatments leading to antinociceptive effect activate Rho signaling but from different mediators as illustrated in Fig. 2B, Fig. 4B and Fig. 4C. Additionally, it is also involved in noxious heat perception as shown in Fig. 2B and Fig. 4A. Rho signaling pathway plays an important role in neuronal growth inhibition associated with neuronal plasticity. The phenomenon of neuroplasticity is related to changes at the nervous tissue leading to the amplification of pain signal transmission to the brain. Thus, Rho signaling pathway could be a potential drug target for the treatment of chronic or neuropathic pain.
Further experiments were conducted to identify the primary targets for Cap and Eug. A TPP method using the concentration range of a ligand (TTP-CR) was used to specifically identify Cap and Eug targets. The 60˚C challenge temperature for the TTP-CR experiments was selected following the determination of protein’s melting temperature (Tm) as shown in Figure S2 (supplementary file). As an initial objective, we used a non-targeted data acquisition for target analysis (nDATA) of C. elegans vanilloid receptors using a FASTA database including only these specific proteins. As shown in Fig. 5A and B, using a nDATA strategy for targeted analysis of vanilloid receptors (i.e., OSM-9, OCR-1 to 4) showed that the primary target for Cap and Eug is OCR-2. The Table S5 (supplementary file) reveals the observed fold changes and adjusted p-values. Likewise, our previous studies using specific mutants have identified OCR-2 for Cap [26], but the assay sensitivity was inadequate to identify Eug targets [27]. OSM‑9 and OCR‑2 vanilloid receptor channels are associated with temperature sensing in C. elegans [34]. The TTP-CR method serves to identify target engagement, and off-target effects may help to explain some of the adverse effects [28, 29, 31]. Shotgun proteomics was used to identify and quantify proteins in the soluble fraction following drug treatment and heat denaturation. As revealed in Fig. 6A and C, over a thousand proteins were identified and quantified in the soluble fraction. However, only a few proteins emerge with a shift in the apparent solubility following interaction with the ligand. Figure 6B and D illustrate that Cap and Eug have 37 and 41 targets, respectively, other than vanilloid receptors. All these proteins had at least a fourfold increase in concentration within the soluble fraction after heat denaturation (60 °C) for 5 min. Interestingly, as shown in Fig. 7A, 33 proteins were shared between both ligands, which is reasonable since they share the same pharmacophore. Table S6 (supplementary file) reveals the list of proteins for each Venn diagram compartment. Protein–protein interaction (PPI) analyses were performed using the MCODE node. As shown in Fig. 7B, 3 PPI clusters were identified, but only two MCODEs were identified using the KEGG and Reactome databases for the 33 shared proteins. No PPIs were found for the limited number of proteins in the Cap or Eug groups. As shown in the parent-to-root node analysis displayed in Fig. 7C, the processes using the 33 shared protein targets are coherent with the results presented in Figs. 2 and 3. PPI analysis reveals associations with biological processes in the categories of catabolism, energy generation, biosynthesis and structural processes. At this stage, we cannot be sure all these targets contribute to the antinociceptive effect observed in C. elegans, and most likely not all do. Small interfering RNA (siRNA) strategies could be used to test the phenotypes of selected Cap and Eug targets. Specific targets for Cap (5 proteins) or Eug (9 proteins) are implicated in enzymatic catalysis and calcium ion binding activity, or they are ribosomal subunits. We have already discussed the possible implication of enzymatic catalysis and ribosomal subunits, but calcium ion binding activity is very important. Vanilloid receptors are non-selective cation channels involved in temperature transduction. Very little is reported for protein CELE_ZK1307.8 or tax-6 other than sequence comparisons and identification of conserved domain architectures that appear to be related to calcium ion binding activity.
Heatmap representation of the general thermal stability of C. elegans soluble TRPV mammalian orthologs after exposure to specific concentrations of Cap or Eug. Thermal proteome profiling using the Cap (A) or Eug (B) concentration range to identify targets. The color range depicts the relative protein abundance of the soluble fractions at different concentrations ranging from 1 to 10 µM. This representation estimates the potency of the tested ligand against each of the targets. OCR-2 appears to be the primary target for both compounds
Heatmap representation of the general thermal stability of C. elegans soluble proteins following exposure to specific concentrations of Cap or Eug. The color range depicts the relative protein abundance of the soluble fractions at different concentrations ranging from 1 to 10 µM. Thermal proteome profiling resulting from Cap (A) exposure and topmost significant target engagement observed with fold change ≥ 4 (B). Thermal proteome profiling resulting from Eug (C) exposure and topmost significant target engagement observed with fold change ≥ 4 (D). The data suggest that Cap and Eug have 37 and 41 targets, respectively, excluding TRPV mammalian orthologs
Protein‒protein interaction (PPI) enrichment analyses. A Venn diagram including both treatments was used to demonstrate the degree of overlap. B PPI network and MCODE components identified from the overlapping protein lists. C Functional analysis of enriched PPI terms from the parent node to the root node. No enrichment was found with proteins exclusively found following Cap or Eug exposure
Conclusion
Shotgun proteomics and TPP-CR experiments provided new insights into the mechanisms of action leading to the antinociceptive effects of Cap and Eug in C. elegans. We noted some hierarchical differences following Reactome pathway enrichment analyses between Cap- and Eug-treated nematodes. However, both treated groups were associated mainly with signal transduction pathways, energy generation, biosynthesis and structural processes. Interestingly, Wnt signaling is enriched following both treatments a pathway associated with pain. The TPP-CR experiment allowed us to identify the vanilloid receptor targeted by Cap and Eug, OCR-2, and not OSM-9. Further PPI analyses showed other targets implicated in enzymatic catalysis and calcium ion binding activity or showed that they were ribosomal subunits. The resulting data help to better understand the broad-spectrum pharmacological activity of vanilloids, specifically Cap and Eug.
Data Availability
The data that support the findings from this study are available from the corresponding author upon reasonable request. Proteome discoverer 2.4 and Metascape datasets can be found online at Mendeley Data (https://doi.org/10.17632/4kv68ds8p5.1).
References
Glauser DA, Chen WC, Agin R et al (2011) Heat avoidance is regulated by transient receptor potential (TRP) channels and a neuropeptide signaling pathway in Caenorhabditis elegans. Genetics 188:91–103. https://doi.org/10.1534/genetics.111.127100
Goodman MB, Sengupta P (2019) How Caenorhabditis elegans senses mechanical stress, temperature, and other physical stimuli. Genetics 212:25–51. https://doi.org/10.1534/genetics.118.300241
Aoki I, Mori I (2015) Molecular biology of thermosensory transduction in C. elegans. Curr Opin Neurobiol 34:117–124. https://doi.org/10.1016/j.conb.2015.03.011
Schafer WR (2006) Proprioception: a channel for body sense in the worm. Curr Biol 16:R509–R511. https://doi.org/10.1016/j.cub.2006.06.012
Calahorro F, Izquierdo PG (2018) The presynaptic machinery at the synapse of C. elegans. Invertebr neurosci 18:4. https://doi.org/10.1007/s10158-018-0207-5
Wittenburg N, Baumeister R (1999) Thermal avoidance in Caenorhabditis elegans: an approach to the study of nociception. Proc Natl Acad Sci U S A 96:10477–10482
Wortley MA, Birrell MA, Belvisi MG (2017) Drugs affecting TRP channels. Handb Exp Pharmacol 237:213–241. https://doi.org/10.1007/164_2016_63
Martins D, Silva M, Tavares I (2017) TRPV1 in pain control from the brain. Oncotarget 8:16101–16102. https://doi.org/10.18632/oncotarget.13316
Caterina MJ, Schumacher MA, Tominaga M et al (1997) The capsaicin receptor: a heat-activated ion channel in the pain pathway. Nature 389:816–824. https://doi.org/10.1038/39807
Yang BH, Piao ZG, Kim Y-B et al (2016) Activation of vanilloid receptor 1 (VR1) by eugenol. J Dent Res 82:781–785. https://doi.org/10.1177/154405910308201004
Yang BH, Piao ZG, Kim Y-B et al (2003) Activation of vanilloid receptor 1 (VR1) by eugenol. J Dent Res 82:781–785
Ohkubo T, Shibata M (1997) The selective capsaicin antagonist capsazepine abolishes the antinociceptive action of eugenol and guaiacol. J Dent Res 76:848–851. https://doi.org/10.1177/00220345970760040501
Jancsó G, Dux M, Oszlács O, Sántha P (2008) Activation of the transient receptor potential vanilloid-1 (TRPV1) channel opens the gate for pain relief. Brit J Pharmacol 155:1139–1141. https://doi.org/10.1038/bjp.2008.375
Fernández-Ballester G, Ferrer-Montiel A (2008) Molecular modeling of the full-length human TRPV1 channel in closed and desensitized states. J Membr Biol 223:161–172. https://doi.org/10.1007/s00232-008-9123-7
Elokely K, Velisetty P, Delemotte L et al (2016) Understanding TRPV1 activation by ligands: Insights from the binding modes of capsaicin and resiniferatoxin. Proc Natl Acad Sci 113:E137–E145. https://doi.org/10.1073/pnas.1517288113
Ikenaka K, Tsukada Y, Giles AC et al (2019) A behavior-based drug screening system using a Caenorhabditis elegans model of motor neuron disease. Sci Rep 9:10104. https://doi.org/10.1038/s41598-019-46642-6
Martinez BA, Caldwell KA, Caldwell GA (2017) C. elegans as a model system to accelerate discovery for Parkinson disease. Curr Opin Genet Dev 44:102–109. https://doi.org/10.1016/j.gde.2017.02.011
Markaki M, Tavernarakis N (2020) Caenorhabditis elegans as a model system for human diseases. Curr Opin Biotechnol 63:118–125. https://doi.org/10.1016/j.copbio.2019.12.011
Schmeisser K, Parker JA (2018) Worms on the spectrum - C. elegans models in autism research. Exp Neurol 299:199–206. https://doi.org/10.1016/j.expneurol.2017.04.007
Mohammadi A, Rodgers JB, Kotera I, Ryu WS (2013) Behavioral response of Caenorhabditis elegans to localized thermal stimuli. BMC Neurosci 14:66. https://doi.org/10.1186/1471-2202-14-66
Srinivasan J, Durak O, Sternberg PW (2008) Evolution of a polymodal sensory response network. BMC Biol 6:52. https://doi.org/10.1186/1741-7007-6-52
Lindy AS, Parekh PK, Zhu R et al (2014) TRPV channel-mediated calcium transients in nociceptor neurons are dispensable for avoidance behaviour. Nat Commun 5:4734. https://doi.org/10.1038/ncomms5734
Liedtke WB, Heller S, Sze JY (2007) The TRPV channel in C. elegans serotonergic neurons. CRC Press, Boca Raton
Kahn-Kirby AH, Bargmann CI (2006) TRP channels in C. elegans. Annu Rev Physiol 68:719–736. https://doi.org/10.1146/annurev.physiol.68.040204.100715
Montell C (2003) The venerable inveterate invertebrate TRP channels. Cell Calcium 33:409–417
Nkambeu B, Salem JB, Beaudry F (2020) Capsaicin and its analogues impede nocifensive response of caenorhabditis elegans to noxious heat. Neurochem Res. https://doi.org/10.1007/s11064-020-03049-4
Nkambeu B, Salem JB, Beaudry F (2021) Eugenol and other vanilloids hamper caenorhabditis elegans response to noxious heat. Neurochem Res 46:252–264. https://doi.org/10.1007/s11064-020-03159-z
Perrin J, Werner T, Kurzawa N et al (2020) Identifying drug targets in tissues and whole blood with thermal-shift profiling. Nat Biotechnol 38:303–308. https://doi.org/10.1038/s41587-019-0388-4
Hong KT, Lee J-S (2020) Label-free proteome profiling as a quantitative target identification technique for bioactive small molecules. Biochemistry-us 59:213–215. https://doi.org/10.1021/acs.biochem.9b00975
Kaur U, Meng H, Lui F et al (2018) Proteome-wide structural biology: an emerging field for the structural analysis of proteins on the proteomic scale. J Proteome Res 17:3614–3627. https://doi.org/10.1021/acs.jproteome.8b00341
Mateus A, Määttä TA, Savitski MM (2017) Thermal proteome profiling: unbiased assessment of protein state through heat-induced stability changes. Proteome sci. https://doi.org/10.1186/s12953-017-0122-4
Margie O, Palmer C, Chin-Sang I (2013) Chemotaxis assay. J vis exp. https://doi.org/10.3791/50069
Brenner S (1974) The genetics of caenorhabditis elegans. Genetics 77:71–94. https://doi.org/10.1093/genetics/77.1.71
Ohnishi K, Saito S, Miura T et al (2020) OSM-9 and OCR-2 TRPV channels are accessorial warm receptors in Caenorhabditis elegans temperature acclimatisation. Sci Rep-uk 10:18566. https://doi.org/10.1038/s41598-020-75302-3
Zhou Y, Zhou B, Pache L et al (2019) Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat Commun 10:1523. https://doi.org/10.1038/s41467-019-09234-6
Bindea G, Mlecnik B, Hackl H et al (2009) ClueGO: a Cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics 25:1091–1093. https://doi.org/10.1093/bioinformatics/btp101
Shannon P, Markiel A, Ozier O et al (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504. https://doi.org/10.1101/gr.1239303
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J Royal Stat Soc Ser B Methodol 57:289–300. https://doi.org/10.1111/j.2517-6161.1995.tb02031.x
Salem JB, Nkambeu B, Arvanitis DN, Beaudry F (2022) Resiniferatoxin hampers the nocifensive response of caenorhabditis elegans to noxious heat, and pathway analysis revealed that the Wnt signaling pathway is involved. Neurochem Res 47:622–633. https://doi.org/10.1007/s11064-021-03471-2
Gupta N, Pevzner PA (2009) False discovery rates of protein identifications: a strike against the two-peptide rule. J Proteome Res 8:4173–4181. https://doi.org/10.1021/pr9004794
Iftinca M, Defaye M, Altier C (2021) TRPV1-targeted drugs in development for human pain conditions. Drugs 81:7–27. https://doi.org/10.1007/s40265-020-01429-2
Sultana A, Singla RK, He X et al (2021) Topical capsaicin for the treatment of neuropathic pain. Curr Drug Metab 22:198–207. https://doi.org/10.2174/1389200221999201116143701
Leavell Y, Simpson DM (2022) The role of the capsaicin 8% patch in the treatment of painful diabetic peripheral neuropathy. Pain Manag 12:595–609. https://doi.org/10.2217/pmt-2021-0025
Giaccari LG, Aurilio C, Coppolino F et al (2021) Capsaicin 8% patch and chronic postsurgical neuropathic pain. J Pers Med 11:960. https://doi.org/10.3390/jpm11100960
Ausín-Crespo MD, Castro EM, Roldán-Cuartero J et al (2022) Capsaicin 8% dermal patch for neuropathic pain in a pain unit. Pain Manag Nurs. https://doi.org/10.1016/j.pmn.2021.12.005
Roeckel L-A, Coz G-ML, Gavériaux-Ruff C, Simonin F (2016) Opioid-induced hyperalgesia: cellular and molecular mechanisms. Neuroscience 338:160–182. https://doi.org/10.1016/j.neuroscience.2016.06.029
Guichard L, Hirve A, Demiri M, Martinez V (2021) Opioid-induced hyperalgesia in patients with chronic pain: a systematic review of published cases. Clin J Pain 38:49–57. https://doi.org/10.1097/ajp.0000000000000994
Svensson CK (2022) Opioid-induced hyperalgesia: is it a clinically relevant phenomenon? Int J Pharm Pract. https://doi.org/10.1093/ijpp/riac031
Zhang Y-K, Huang Z-J, Liu S et al (2013) WNT signaling underlies the pathogenesis of neuropathic pain in rodents. J Clin Invest 123:2268–2286. https://doi.org/10.1172/jci65364
Zhou Y-Q, Tian X-B, Tian Y-K et al (2022) Wnt signaling: a prospective therapeutic target for chronic pain. Pharmacol Therapeut 231:107984. https://doi.org/10.1016/j.pharmthera.2021.107984
Itokazu T, Hayano Y, Takahashi R, Yamashita T (2014) Involvement of Wnt/β-catenin signaling in the development of neuropathic pain. Neurosci Res 79:34–40. https://doi.org/10.1016/j.neures.2013.12.002
Han B, Zhao J, Wang W et al (2017) Cdc42 promotes schwann cell proliferation and migration through Wnt/β-catenin and p38 MAPK signaling pathway after sciatic nerve injury. Neurochem Res 42:1317–1324. https://doi.org/10.1007/s11064-017-2175-2
Gernez Y, de Jesus AA, Alsaleem H et al (2019) Severe autoinflammation in 4 patients with C-terminal variants in cell division control protein 42 homolog (CDC42) successfully treated with IL-1β inhibition. J Allergy Clin Immun 144:1122-1125.e6. https://doi.org/10.1016/j.jaci.2019.06.017
Zhou WBS, Shi XQ, Liu Y et al (2022) Unbiased proteomic analysis detects painful systemic inflammatory profile in the serum of nerve injured mice. Pain. https://doi.org/10.1097/j.pain.0000000000002695
Mazumder B, Sampath P, Seshadri V et al (2003) Regulated release of L13a from the 60S ribosomal subunit as a mechanism of transcript-specific translational control. Cell 115:187–198. https://doi.org/10.1016/s0092-8674(03)00773-6
Chédotal A (2019) Roles of axon guidance molecules in neuronal wiring in the developing spinal cord. Nat Rev Neurosci 20:380–396. https://doi.org/10.1038/s41583-019-0168-7
Ke C, Gao F, Tian X et al (2017) Slit2/Robo1 mediation of synaptic plasticity contributes to bone cancer pain. Mol Neurobiol 54:295–307. https://doi.org/10.1007/s12035-015-9564-9
Llorián-Salvador M, González-Rodríguez S (2018) Painful Understanding of VEGF. Front Pharmacol 9:1267. https://doi.org/10.3389/fphar.2018.01267
den Eynde CV, Vriens J, Clercq KD (2021) Transient receptor potential channel regulation by growth factors. Biochimica Et Biophysica Acta Bba - Mol Cell Res 1868:118950. https://doi.org/10.1016/j.bbamcr.2021.118950
Acknowledgements
This project was funded by the National Sciences and Engineering Research Council of Canada (F. Beaudry discovery Grant Number RGPIN-2020-05228). Laboratory equipment was funded by the Canadian Foundation for Innovation (CFI) and the Fonds de Recherche du Québec (FRQ), the Government of Quebec (F. Beaudry CFI John R. Evans Leaders Grant Number 36706). F. Beaudry is the holder of the Canada Research Chair in metrology of bioactive molecule and target discovery (Grant Number CRC-2021-00160). This research was undertaken partly thanks to funding from the Canada Research Chairs Program. A Ph.D. scholarship was awarded to J. Ben Salem from the Fonds de recherche du Québec—Santé (FRQS).
Funding
This study was supported by Natural Sciences and Engineering Research Council of Canada, RGPIN-2020-05228, Canadian Foundation for Innovation, 36706, Canada Research Chairs, CRC-2021-00160.
Author information
Authors and Affiliations
Contributions
BN: Planning and Execution of experiments, Writing - original draf. JBS: Execution of experiments, Writing - review & editing. FB: Conceptualization, Funding, Organisation, Writing - review & editing.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Nkambeu, B., Salem, J.B. & Beaudry, F. Antinociceptive Activity of Vanilloids in Caenorhabditis elegans is Mediated by the Desensitization of the TRPV Channel OCR-2 and Specific Signal Transduction Pathways. Neurochem Res 48, 1900–1911 (2023). https://doi.org/10.1007/s11064-023-03876-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11064-023-03876-1