Abstract
Conventional agricultural practices rely heavily on chemical fertilizers to boost production. Among the fertilizers, phosphatic fertilizers are copiously used to ameliorate low-phosphate availability in the soil. However, phosphorus-use efficiency (PUE) for major cereals, including maize, is less than 30%; resulting in more than half of the applied phosphate being lost to the environment. Rock phosphate reserves are finite and predicted to exhaust in near future with the current rate of consumption. Thus, the dependence of modern agriculture on phosphatic fertilizers poses major food security and sustainability challenges. Strategies to optimize and improve PUE, like genetic interventions to develop high PUE cultivars, could have a major impact in this area. Here, we present the current understanding and recent advances in the biological phenomenon of phosphate uptake, translocation, and adaptive responses of plants under phosphate deficiency, with special reference to maize. Maize is one of the most important cereal crops that is cultivated globally under diverse agro-climatic conditions. It is an industrial, feed and food crop with multifarious uses and a fast-rising global demand and consumption. The interesting aspects of diversity in the root system architecture traits, the interplay between signaling pathways contributing to PUE, and an in-depth discussion on promising candidate genes for improving PUE in maize are elaborated.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Phosphorus (P), being an essential macronutrient, plays a vital role in plant growth and development. It is one of the most limiting nutrients for crop productivity. It is absorbed by the root and transported to the younger leaves and other parts via xylem and phloem tissues. P plays a vital role in cellular physiology by assimilating into key cellular constituents, viz., nucleic acids, phosphoproteins, membrane phospholipids, and energy-rich compounds like adenosine triphosphate (ATP). It participates in enzymatic reactions (e.g., phosphorylation and de-phosphorylation) and cell signaling (phosphatases). Since P is part of the building blocks for nucleic acid, therefore, an adequate supply of P is essential for cell division and plant growth. It is also essential during the reproductive phase of plants for seed formation and development. Interestingly, seeds contain a large quantity of P in the form of phytin or phytic acid. More than 60% of P in cereal grains is stored as phytin [1].
Owing to its pivotal role in plant growth and development, deficiency of P usually results in diminished crop growth and lower yield. Further, a low supply of P during the grain filling stage results in decreased seed number and less carbon deposition leading to reduced seed size, and reduction in yield [2]. P deficiency is a major constraint to achieving optimum crop yield in approximately 50% of global agricultural soils. External application of phosphate (PO4)3− based fertilizers is necessary for optimum production. P-based fertilizers are manufactured from rock phosphate reserves, which are non-renewable and confined to a few countries like Morocco, China, and the USA [3]. There is a prevailing concern that the global rock phosphate reserves may exhaust in a century or two at the current consumption rates, posing a major threat to global food security [4]. Cereals, like maize, are capable of utilizing less than 30% of the applied P fertilizer. The unutilized phosphate results in environmental degradation via eutrophication. Hence, the imminent danger of phosphate reserve depletion and low phosphorus-use efficiency are major challenges for a sustainable cropping system.
One of the strategies to improve PUE is to develop P-efficient genotypes. Although good progress has been made in the recent past in the identification and characterization of genes and miRNAs playing a pivotal role in P uptake, translocation, and homeostasis in Arabidopsis and rice, relatively lesser research has been carried out in maize in this direction. The potential candidate genes/miRNAs from Arabidopsis and rice or their maize orthologs might be utilized and explored for improving PUE in maize. The goal of the present review is to narrow down potential genetic targets and thereby, summarize the major molecular players contributing to P uptake and transport, and regulatory components playing a crucial role in maintaining P levels in plants, particularly maize. The adaptive strategies to enhance P availability under P-deficiency stress are also discussed.
P uptake and translocation
In the soil, P exists as inorganic form (Pi; usually 10–15%), organic form (20–80%), and mineral precipitates (5–7%) [5]. The inorganic phosphate (H2PO4−) form is acquired by plants through the root, while the organic form (mainly phytic acid, but depends on soil type also) often forms salts with different ions rendering them insoluble or precipitated and unavailable for uptake by plants. Inorganic phosphate (Pi) strongly reacts with oxides and hydroxides of Al and Fe in acidic soil and cations in alkaline soil (like Ca2+, Mg2+) to form insoluble precipitates, thereby further limiting its bioavailability to the plant [6]. The concentration of inorganic phosphate in soil (0.1–10 μM) is thousand fold lower than in plant cells (5–20 mM) [6] implying that Pi uptake and transport have to be done against electrochemical potential.
Pi is absorbed by roots through low- and high-affinity phosphate transport systems. The low-affinity phosphate transport system is expressed constitutively while the high-affinity phosphate transport system is down-regulated at high-P availability to avoid P toxicity or induced mainly under P-deficiency conditions i.e., regulated by Pi availability [7]. Pi acquisition from the soil is an energy-mediated process and relies primarily on the plasma membrane-localized phosphate transporters (PHT such as PHT1 family members) functioning as Pi/H+ symporters [8]. The phosphate transporters are not only involved in Pi uptake by root but also in its distribution from root to shoot. The Pht genes are mainly classified into four families, viz., Pht1, Pht2, Pht3, and Pht4 that are localized on the plasma membrane, plastidial membrane, mitochondrial membrane, and Golgi-compartment, respectively [9, 10]. The Pht1 family members express in the root-soil interface (root hairs and root epidermal cells) [11] and symbiotic root-fungus (arbuscular mycorrhiza) interface i.e., root cortical cells harboring the fungal arbuscules [12]. In several plants, including maize, root colonization by symbiotic mycorrhizal fungi plays a crucial role in Pi uptake from the soil [13]. Some Pht1 family members have also been detected in other tissues, like leaves, stems, cotyledons, phloem, anthers, pollens, flowers, and seeds, suggesting that Pht1 family genes are not only involved in direct Pi uptake but also its subsequent distribution within the plant [11, 14]. Pht1 family genes are also involved in Pi remobilization during leaf senescence (Pht1;5 gene in Arabidopsis) and fruit maturation [15]. The members of the PHT2, PHT3, and PHT4 transporters participate in Pi translocation into sub-cellular organelles, viz., plastids, mitochondria, and Golgi, respectively [9, 10, 16].
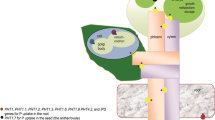
In maize, five Pht1 genes (ZmPht1;1–5) exhibit differential and diverse expression patterns under Pi-deprived conditions [17]. Subsequently, thirteen putative Pht1 genes (ZmPht1;1–1;13) have been reported [18]. Under sufficient Pi supply, only six out of thirteen ZmPht genes are expressed in at least one tissue. Among them, ZmPht1;1 expresses in roots, leaves, stems, anthers, pollen, cobs, silks, and seeds, which indicates its role in both Pi uptake and translocation. ZmPht1;3 expresses predominantly in anther and pollen [17, 18] (Fig. 1). However, under Pi-deprived conditions, twelve out of thirteen ZmPht genes are induced significantly in root tissue [18]. Among them, expression of ZmPht1;3, ZmPht1;5, ZmPht1;8 and ZmPht1;13 are up-regulated in response to low-Pi without arbuscular mycorrhizal fungi (AMF), but down-regulated in response to AMF thereby, indicating that these are involved in the direct Pi uptake pathway [18]. ZmPht1;2, 1;4, 1;6, 1;7, 1;9 and 1;11 are up-regulated in roots colonized by AMF under low-Pi conditions, indicating their probable involvement in indirect Pi uptake through AMF-colonized roots (symbiotic pathway) [17,18,19]. Among these, ZmPht1;6 is the best characterized mycorrhizal-inducible Pi transporter whose transcript level significantly and positively correlates with the level of AMF biomarkers in roots and AMF colonization rate [20]. It is implicated in Pi-uptake in symbiotic mycorrhizal maize roots and has shown to be vital for maize growth (biomass increase) and cob development under nutrient deficiency. Recently, ZmPht1;9 was characterized as a mycorrhizal induced phosphate transporter gene in maize [21]. Heterologous expression of this maize gene in Arabidopsis and rice resulted in more Pi accumulation, indicating its involvement in Pi uptake [21, 22]. Further, ZmPht1;2,1;3 and 1;6 are also expressed in old leaves and young leaves, suggesting their involvement in P re-mobilization from old leaves (source) to young leaves (sink) (Fig. 1). Besides Pht1s identification and expression analysis, studies on testing the affinities towards Pi and cellular/subcellular localization of PHT1s are lacking in maize [23]. Further research focusing on delineating molecular mechanisms of PHTs in maize might be useful in the development of alternate strategies for better P management.
Overview of phosphate (Pi) uptake, export, and re-mobilization in maize plant via PHT1 family transporters. Site-specific expression of Pht1 genes in various tissues, viz., roots, shoots, and leaves in maize are listed. Pht genes with black, red, and purple font colors represent different Phts expressed under sufficient-Pi, low-Pi, and in response to arbuscular mycorrhizal fungi (AMF) inoculation under low-Pi supply conditions, respectively (expression data utilized from [17, 18]). Maize plant adaptation under phosphorus (P)-limiting conditions is also enlisted in the box
Apart from PHT, another phosphate transporter-like SPX domain (characterized by the presence of the SPX domain at the N-terminal)-containing proteins has been implicated in phosphate sensing and transport. These proteins are grouped further into four subfamilies based on the presence or absence of a secondary domain at the C-terminal such as EXS, RING (Really Interesting New Gene), or MFS (Major Facilitator Superfamily) domain. SPX-EXS subfamily members such as Pho1 (Phosphate 1) and its homologs are implicated in Pi loading into the root xylem and transport/export from roots to shoots [10, 24]. The SPX-MFS subfamily members such as VPT1 (Vacuolar Phosphate Transporter 1, also named as Pht5;1) and OsSPX-MFS3, a low-affinity Pi transporter, have been implicated in Pi transport into the vacuole and efflux from the vacuole, respectively [25], which act as Pi storage compartment in the plant cell. Thus, the concerted action of the above-mentioned various phosphate transporters warrants proper Pi distribution to specific cells, tissues, and organelles in the plant.
Plant adaptation under Pi-limiting conditions
The availability of Pi greatly affects the plant’s growth and productivity. Maize plants grown under low-Pi conditions exhibit visual symptoms of P deficiency like purple coloration in the root, shoot, and old leaves. Further, Pi-deficiency also results in a reduction in various morphological parameters, such as plant height, root and shoot weight, biomass accumulation, etc. as compared to the Pi-sufficient conditions (Supplementary Fig. S1). To cope with Pi-deficiency, crop plants (including maize) undergo diverse morphological, biochemical, physiological, and molecular adaptations, which in turn, result in enhanced Pi availability in the rhizosphere and hence its increased uptake from the soil. These adaptations are-modifying root morphology and architecture, increased root absorptive surface area via an increasing number of root hairs, induction of secretory protein extrusion by roots [induction of acid phosphatases (APases), organic acid and ribonucleases (RNase) secretion into the rhizosphere to release available P from the soil], obtaining Pi via arbuscular mycorrhizal association, remobilization of Pi from old leaves/tissue to young leaves/tissue, anthocyanin accumulation, etc. [6, 10, 26] (Fig. 1). Low-Pi tolerant maize genotypes have a higher shoot-to-root ratio, higher anthocyanin accumulation, larger root length, root surface area, and root volume, and secrete more APases and organic acids in roots compared to sensitive ones under P-limiting conditions [26, 27].
At the molecular level, the plant responds to Pi deficiency by altering the expression levels of Pi deficiency-associated genes including transcription factors, many PHTs (mostly high-affinity transporters; discussed above), and miRNAs [28]. Under Pi limitation, some transcription factors (TFs) control the expression of a set of phosphate starvation-induced (PSI) genes by binding to cis-elements present in their promoter. The regulatory TFs implicated in phosphate starvation signaling are—phosphate starvation response 1 (PHR1), ZAT6, WRKY6, WRKY42, WRKY45, WRKY75, MYB26, and bHLH32 in Arabidopsis; OsPHR2 and OsPTF1 in rice; TaPHR1 in wheat; ZmPTF1 in maize [29,30,31,32,33,34], etc. Among these, PHR1/PTF1, a MYB domain-containing TF, is a central regulator of Pi signaling and is involved in phosphate starvation response (PSR) [16, 35]. Under low-P stress, it regulates the expression of various PSI genes such as Pht1s (high-affinity H+/Pi co-transporter; preferentially expressed in epidermal/cortical cells of the root), Pho1 (high-affinity Pi transporter expressed in root pericycle), and miR399 through binding to the P1BS sequence, GNATATNC, present in the promoter region of PSI genes [28]. PHR1 is a direct target of SIZ1, a SUMO E3 ligase, which positively regulates it post-translationally. In Arabidopsis, PSI genes are repressed in siz1 mutant even in Pi deficiency. Pho1 and some other SPX domain-containing proteins exhibit a major role in controlling PSR. Under Pi starvation, WRKY45 and WRKY75 are up-regulated while WRKY6 and WRKY42 are down-regulated. WRKY45/75 positively regulates Pht1;1 [16, 29], while WRKY6/42 negatively regulates/ repressed Pho1 expression [31]. Under P starvation conditions, degradation of WRKY42 happens, which in turn, implies lesser suppression of Pho1 [34].
A few SPX domain-containing proteins act as important feedback regulators of OsPHR2 in rice. Under Pi starvation, PHR2 activates SPX domain-containing proteins in Arabidopsis and rice [36]. Pho2, a ubiquitin E2 conjugase, is implicated in controlling the degradation of transporters by the ubiquitination process. The miRNA399 controls the expression of Pho2 and results in its post-transcriptional degradation [28]. Apart from miR399, miR827 is the other conserved small non-coding RNA family characterized in plants under P deficiency stress [37].
Acid phosphatases (APases) are also PSI genes that play a crucial role in the mobilization and utilization of organic P under Pi-deprived conditions [38]. The APases are secreted externally in the root zone and help in the release of Pi from organophosphates. Several APases have been characterized in vascular plants, like lupin, tomato, tobacco, common bean, Arabidopsis and soybean [38]. So far, 29 Purple Acid Phosphates (PAP) have been identified in Arabidopsis [39], 26 in rice [40], 35 in soybean [41], 33 in maize [42] and 25 in chickpea [43]. In Arabidopsis, 11 out of 29 members of PAP are up-regulated by Pi stress [39].
In maize, transcriptomic studies in response to the low-Pi conditions have identified a set of differentially expressed Pi-responsive genes, that are mainly involved in numerous metabolic pathways, phytohormone regulation, ion transport, redox homeostasis, transcriptional regulation, and protein synthesis and degradation [26, 44]. Among Pi-responsive genes, five maize Pht1s, eight Pho/SPX/EXS domain encoding genes, APases, phytase, peroxidase, RNase, and various TFs are up-regulated in the root tissue of P-efficient genotypes under low-phosphate stress [26, 44]. The P1BS motif—a binding site of the central regulator—is present in the promoters of 8 of 13 ZmPht1s [25]. However, it is not always necessary that P1BS containing Pht1s are involved in Pi starvation. For instance, ZmPht1;1 gene contains P1BS in its promoter but, is not induced by Pi starvation hence not involved in Pi starvation [18]. Recently, 18 ZmPhr genes were identified in the maize genome that exhibits differential and diverse expression patterns [45]. Under the low-Pi conditions, eight (ZmPhr3, ZmPhr4, ZmPhr6, ZmPhr7, ZmPhr9, ZmPhr13, ZmPhr17, and ZmPhr18) and two (ZmPhr5 and ZmPhr10) genes are up-and down-regulated in leaf, respectively, indicating their probable involvement in the regulation of Pi translocation in leaves under different Pi level. Besides Pi-starvation responsive genes, differentially expressed miRNAs that include miR156, miR166, miR169, miR393, miR395, miR398, miR399, miR528, and miR827 family members have also been identified in maize subjected to the low-Pi conditions [37, 46]. Pi starvation-dependent induction/up-regulation of miR399 family members and miR827, and down-regulation of miR827 targets (such as SPX domain-containing protein, and ubiquitin E3 ligase containing the RING and SPX domains) suggest their role in Pi homeostasis and hence Pi starvation adaptation in maize [37]. Owing to conservation in Pi-starvation response in plants, the known mechanism of Pi starvation signaling in Arabidopsis and rice, and the above-mentioned studies in maize, we propose a putative transcriptional regulatory model for Pi-starvation responses in maize (Fig. 2). In summary, plants show a remarkable adaptation response to low-Pi stress, which is manifested in terms of differential expression of genes and non-coding RNAs. Unraveling this natural response of the plant could potentially open a window for the identification of key candidate genes for improving PUE.
Transcriptional regulatory model for phosphate (Pi)-starvation responses in maize. The hypothesis is derived from the fact that the Pi deficiency response is quite conserved in plants [110]. Hence, the knowledge about the regulation of Pi starvation signaling in Arabidopsis and rice is utilized and extrapolated for maize considering transcriptomic studies in response to the low-Pi condition in maize [26, 37, 44, 111, 112]. Red-colored boxes and down arrows denote down-regulation (negative effect) while green boxes and up arrows represent up-regulation (positive effect). Redline ending with a short bar indicates inhibition. A small orange circle denotes for sumoylation (represented by the letter ‘S’) of PHR (phosphate starvation response), a transcription factor. P1BS (PHR1 binding sequence; GNATATNC) is the cis-regulatory element found in the promoter region of many Pi-starvation-induced genes, where the PHR transcription factor binds and regulates their expression. ZmPho2—Zea mays phosphate overaccumulator 2; ARF—Auxin Response Factor; ERF—ethylene response factor; ABA-REBF—ABA-responsive element binding factor; PHR—phosphate starvation response; ZmPTF1—Zea mays phosphate starvation-induced transcription factor 1; P1BS—PHR1 binding sequence on the target gene promoter
Root system architecture for PUE in maize: genetic variation
Root system architecture (RSA) is a key trait that confers better PUE to crops under Pi-deficient soils. The P-efficient cultivars can be developed through ideotype breeding via integrating a favorable combination of RSA traits for target environments. The wild relatives and landraces are the best resource to explore the RSA-associated traits as most modern cultivars have been selected through extensive selection in optimal or nutrient-rich environments [47]. The favorable RSA for imparting better PUE in maize includes a higher number of hypocotyl-borne roots and basal root whorls, greater lateral root branching density (LRBD), longer but denser root hairs, shallow root growth angles, more production of crown roots, higher axial roots, higher root cortical aerenchyma (RCA), greater cortical cell size and greater root cortical senescence [47,48,49]. Maize genotypes with greater production of crown roots exhibit better capture of P, growth, and yield in low-P soil as compared to those with fewer crown roots [50]. The greater number of hypocotyl-borne roots imparts better PUE because of their shallow growth angles and less metabolic cost [51]. Although the shallow roots and steeper angles are important to extract the water, nitrogen (N), and P during the seedling stage, deep architectural roots are essential to explore the fertile soils [47]. Densely spaced (more than 9 branches/cm), short laterals, fine roots (provides more surface area for uptake) and shallow rooting are desirable for P acquisition in maize [52, 53]. Maize genotypes possessing enhanced lateral rooting exhibit up to 100% higher P accumulation and growth rate [52].
Maize possesses ample genetic variation for RSA-associated traits related to PUE [47]. The genetic gain depends upon the genetic control of traits; complex traits are difficult to improve. Root hairs (longer, denser hairs) are preferred traits for RSA improvement due to their greater role in PUE, presence of enormous genetic variation in root-hair length and density, and simpler genetic control [54]. RCA is an important trait for PUE because it decreases the respiration and P cost of maintaining root tissue. There is a significant variation in RCA formation in maize cultivars [55]. Furthermore, SimRoot modeling in maize revealed the role of RCA in enhancing growth up to 70% under P-deficient soil stress [56]. In maize, enormous genetic variation exists for LRBD [48]. The high LRBD favors the roots to be shallower and higher root sink strength (more lateral roots), but breeders need to be cautious as sometimes high LRBD results in trade-off effects by restricting the growth rate of the lateral roots (resource limitations). Thus, the genetic diversity in RSA could be harnessed to enhance PUE in corn.
Genomic regions/QTLs mapped for RSA in maize
A few mapping studies mapped several genomic regions i.e., quantitative trait loci (QTL) governing the RSA for better PUE (Table 1). A major QTL for lateral root number (LRN) on chromosome 2 has been identified under low-P using recombinant inbred lines (RILs), in addition to QTL for plasticity of LRN on chromosome 4 [57]. Similarly, a major QTL for root hair length has been mapped on chromosome 9 [53]. Later, in the same population, one major QTL each for seminal root length (SRL) and seminal root number (SRN) was mapped on chromosomes 2 and 6, respectively, and epistatic interaction QTLs were detected [58]. Later, a stable or consistent QTL for root weight (dupssr15 locus), detected across four environments was identified [59]. Major QTLs for root diameter and surface area of fine roots on chromosomes 7 and 10, respectively in L22 × L3 based RILs were also detected [60]. Furthermore, by utilizing the multiple interval mapping approaches with multiple traits (root length, root surface area, root:shoot ratio, and P content) a major QTL has been mapped on chromosome 8 that co-localized with the ZmPSTOL candidate gene, the higher expression of this candidate gene being verified in the roots of L22.
Root exudates also play an important role in improving PUE. A major QTL for H+ secretion on chromosome 1 (bnlg2228-bnlg100) was detected [61]. Qin et al. [62] used 082 (P-efficient genotype) and Ye107 based F2 population to map a major and consistent QTL for acid phosphatase, AP1-KXNP (between bnlg1268a-umc1290a) on chromosome 1. Later, using the same population, Qiu et al. [63] mapped two stable, but minor effects QTLs for acid phosphatase activity (exudates in rhizosphere) on chromosome 1 (umc2083-umc1972) and 5 (umc2111 to dupssr10). Such inconsistency in QTL effects can be attributed to the significant influence of the environment [59, 61,62,63]. The study also identified a major and stable QTL for acid phosphatase activity in the root on chromosome 3 (bnlg1350–bnlg1449).
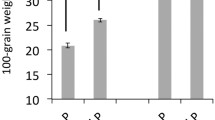
Phosphorous uptake efficiency (PUpE) has been shown as the main determinant of PUE in tropical maize under low-P soil conditions as evident from the co-localization of approximately 80% of the QTLs for PUpE with those for PUE [64]. The same study has mapped QTLs for PUE and PUpE on chromosome 1 (qPUE1 and qPAE1), flanked by root-related genes rth1 and bk213, previously associated with lateral root length [65] and with root hair elongation [66], respectively. Furthermore, the mapped region has been found to contain QTLs for root development under P-deficient conditions [53, 67]. QTLs for PUE, PUpE, and Phosphorous utilization efficiency (PUtE) exhibit dominance effects, and hence heterotic combinations can be helpful to generate P-efficient maize hybrids. QTLs related to RSA, PUE, and PUpE in RILs derived from Ye478 × Wu312 have been mapped [68]. The study identified two major QTL clusters for root system architecture, Cl-bin3.04a and Cl-bin3.04b (Table 1). The marker-assisted selection (MAS) based development of nine advanced backcross-derived lines carrying Cl-bin3.04a or Cl-bin3.04b displayed mean increases of 22–26% in PUE under low-P field evaluations. Furthermore, a line L224 pyramiding both Clbin3.04a and Cl-bin3.04b showed enhanced PUpE, mainly due to effects on changes in root morphology, rather than root physiology, under both hydroponic and field conditions. Hence, this study has demonstrated the potential of MAS in exploring the physiological and genetic contributions of RSA to maize PUE [68]. Development of multi-lines with identical shoot architecture but contrasting RSA traits can be helpful to get better performance under P-deficient, optimum and moisture stress [69]. Furthermore, the independent major QTLs can be stacked together through pyramiding to develop PUE efficient maize cultivars.
Harnessing crosstalk between signaling pathways related to other nutrients for improving PUE
As we know plants are not challenged by only one stimulus at a time i.e., plants are dealing with more than one factor at a time. Particularly, most of the plant responses to Pi deficiency are also regulated by the levels of nitrogen. Thus, it is imperative to discuss how other nutrients [e.g., nitrogen (N), and potassium (K)], interplay in signaling pathways contributing to PUE.
Crosstalk between P and N signaling pathways
In nutrient-signaling pathways, both N and P are important constituents of the protein phosphorylation process. In Arabidopsis, an E3 ligase AtNLA (Arabidopsis nitrogen limitation adaptation) has been reported as the first protein that participates during the interaction between these two nutrients. Both AtNLA and miR827 had been observed as P-uptake repressors but did not affect the N uptake [70]. The antagonistic interaction between NO3−(nitrate) and Pi can be explained by the AtPHT1’s vacuolar degradation mediated through AtNLA [71]. The TF AtHRS1 (Arabidopsis hypersensitive to low Pi-elicited primary root shortening 1) has been annotated as N and P signaling integrator and its homolog, named as HRS1 HOMOLOGUE 1 to 6 (AtHOH1 to AtHOH6), has been identified. The Arabidopsis lines overexpressing these TFs show the root phenotype of hypersensitivity to low-Pi [72]. These both kinds of TFs (AtHRS1and AtHOH) are induced in the presence of nitrate and repress the growth of the root as a P-deficiency response [73]. Both AtHRS1and AtHOH homologs 1 to 3 (known as NIGT1.1 to NIGT 1.4) are regulated by Pi starvation response TF i.e., AtPHR1 [74].
NRT1.1(Nitrate transporter 1.1)/CHL1(nitrate transceptorchlorina 1), TF that is involved in NO3− sensing in Arabidopsis, has also been reported as a component of crosstalk between N and P signaling networks whose activity is reliant on PHR1 turnover [75]. In addition to this, the role of PHO2, as an integrator of N signals and P starvation has been described. The effect of N on PSR is evident, as the majority of PSI genes such as PHT1-1, IPS1, and SPX1 are repressed in pho2 mutant. PHO2 and NRT1.1 affect the transcripts level of each other and this is well conserved in the case of wheat and rice also [75]. The downstream pathway of PHR1/PHL1 is somewhat dependent on N uptake and signals mediated by the NRT1.1 sensor. Additionally, HRS1/HHOs and NLA TFs contribute significantly as an integrator between both networks for uptake of N/P and RSA of plants [75]. Previous studies also confirmed that OSNRT1.1B (NO3− sensor) degrades the OsSPX4 (Pi signaling repressor) through an E3 ubiquitin ligase NBIP1 (NRT1.1B interacting protein 1) [76]. This OsSPX4 interacts with OsNLP3 (NO3− signaling TF) and negatively regulates the OsPHR2 (regulator of Pi signaling) in rice. Regardless of available Pi, under lower NO3− conditions, SPX protein accumulates at a higher level and represses the phosphate and nitrate responsive genes due to retention of TFs NLP (NIN-like protein) and PHR in the cytoplasm. The NRT1.1-NBIP1 mediated degradation of OsSPX4 resulted in the expression of nitrate and PSI genes due to the release of PHR and NLP from the cytoplasm to the nucleus leading to higher PUE and better RSA in the plant [76, 77]. These studies suggested that under the Pi-starvation conditions, expression of PSI genes is greatly regulated (mostly enhanced) due to NO3− induction i.e., N actively controls the PSR in plants. It can be utilized to modulate N and P concentrations precisely and to enhance PUtE under a low-Pi environment [78]. Recently, under Pi-deficient and NO3– sufficient conditions, NIGT1.1 (AtHHO2) and NIGT1.2 (AtHHO3) TFs have been shown to enhance Pi acquisition while reducing NO3− uptake by up-regulating and down-regulating the expression of Pi transporter and nitrate transporter genes, respectively [79]. In this study, a similar regulatory pathway was observed in maize also. This study has confirmed that NIGT1 TF plays a central role in maintaining P and N balance in plants during Pi starvation. Further, it should be noted that ammonium (NH4+) is also a major N source of plants. So, the regulation of N/P networks through ammonium sensing needs to be explored [80].
These kinds of responses directly represent the tight linkage between N and P signaling networks. Therefore, detailed investigations on NRT1.1-SPX mediated PHR cascade, Pi starvation-induced expression of NIGT1 resulting in higher expression of Pi transporter genes and molecular mechanism regulating NIGT1 level under various P/N supply conditions might be helpful to gain more understanding of the crosstalk between P and N signaling pathways in maize. This would lead to identifying novel targets for improving PUE in maize and other crop plants.
Cross talk between P and K signaling pathways
The crosstalk between P and K signaling pathways has been observed in plants although little is known about it. Previously, rapid induction of TFs such as MAPK, MAPKK, etc. has been reported due to changes in the concentration of external P or K in tomato [81]. Further, induction of High-affinity K uptake (HAK5)-transporter has been demonstrated under Pi-deficient conditions [82]. Recently, the cross-talk between K and P has been demonstrated in Arabidopsis and tomato as deficiency of Pi results in transcriptional repression of AKT1-type channel forming genes which results in reduced K-uptake and its translocation to shoots. This has also been observed that external higher K concentration inhibits the Pi uptake which, in turn, results in the induction of PSR (responsible TFs: PHR1 and PHL1) and phosphate-responsive genes [83]. More recently, an ionome and transcriptomic study in sorghum identified two HAKs (Potassium high-affinity transporter from KT/HAK/KUP family gene) which might be involved in root-to-shoot translocation of K under a Pi-deficient environment [84]. Although, various TFs, transporters, and nutrient-responsive genes have been identified and characterized in the recent past but P and K sensors are not identified yet. The research on the interaction between multiple nutrient signaling networks is still in its infancy stage that needs to be explored meticulously which would help target nutrient-signaling networks for better PUE.
Prime molecular targets for improving PUE in maize
PUE in maize is a complex trait-mediated by a coordinated action of a set of genes encoding for Pi transporters and regulatory components [TFs and miRNAs), whose expression alters (either induce or suppress) in response to Pi deficiency. Further, abundance, localization, and activities of these transporters are regulated at transcriptional (by TFs), post-transcriptional (by miRNAs), translational and post-translational level [by PHF1, Pho2, SPX-RING domain protein, etc.] via ubiquitination, phosphorylation, and sumoylation [16, 85]. In this context, Pi-uptake and utilization efficiency may be improved by modulating the expression level of key regulatory components and phosphate transporters [few root-specific high-affinity transporters (PHT1s), like Pho1] through a transgenic (over-expression and/or RNAi) approach which could enhance crop productivity under Pi-deprived conditions. It has been shown that many times constitutive promoter-driven over-expression of promising PHTs and regulatory factors might negatively affect transgenic plant growth under P-deficient conditions while leading to P toxicity symptoms under P-sufficient conditions [16, 86]. To minimize these unintended effects, P starvation-inducible and tissue-specific promoters such as shoot-specific, root-specific, root-hair specific, and others should be utilized while harnessing over-expression strategy using these key molecular targets for improving PUE in maize. Apart from P toxicity, sometimes over-expression of high-affinity transporter (PHT1s involved in Pi uptake in P deficiency) may not result in enhanced tolerance to P deficiency [87]. Furthermore, few studies in rice indicated that soil substrate (Pi) availability rather than high-affinity transporter activity may be the limiting step in Pi-deficient soil [88]. These findings suggest the need to explore genes encoding intracellular Pi transporters (PHT2, PHT3, PHT4 families that are involved in proper P distribution, re-mobilization, and maintaining cytosolic P homeostasis) for improving PUE in crops.
In acidic soils, Aluminium toxicity (due to the presence of solubilized ionic form i.e., Al3+) and P-deficiency (due to P-fixing/complexation by clay minerals like iron oxides and kaolinite) both stresses occur at a time. Excessive Al3+ stress inhibits root growth/elongation by destroying the cell structure of the root apex and thereby, majorly reducing immobile nutrient (i.e., Pi) uptake. Thus, in such soil types, PUE can only be improved if maize plants are also tolerant of Al toxicity. To date, few genes such as TaALMT1 and SbMATE imparting Al3+ tolerance in wheat and soybean, respectively, have been well characterized [89]. TaALMT1 encodes an anion channel located at the plasma membrane (malate transporter) that is responsible for malate efflux from root apices while SbMATE encodes a member of the Multidrug and Toxic Compound Extrusion (MATE) transporter family that facilitates citrate efflux from roots. Efflux of malate and citrate from root might result in enhanced Pi uptake by improving root growth or by enhancing the dissolution of Pi from complexes [89]. Thus, improvement of PUE in maize under acidic soil conditions can be achieved by utilizing genes conferring tolerance against high levels of toxic Al3+ along with genes having a role in imparting higher PUE.
The key useful candidates (targets) proven for improving P uptake and/or utilization in various crop plants are enlisted in Table 2. These genes [viz., ZmPTF1, ZmPHRs, Pi transporters involved in Pi mobilization and re-mobilization (ZmPht 1;2 and 1;3), miR399, and miR827, etc.] and/or their maize ortholog (such as ortholog of OsPSTOL1, AVP1, OsPHF1, OsMYB2P-1, OsPht1;6, OsPht2;1, etc.) may be potentially useful for improving PUE in maize (Table 2). Being a complex and polygenic trait, engineering multiple molecular targets simultaneously such as TFs, key high-affinity Pi transporters, and/or intracellular Pi transporters would be important. Genetic engineering approaches, like transgene stacking, knockdown via RNA interference (a negative regulator of PUE), and/or highly efficient targeted mutation(s) in the promoter(s)/gene(s) aimed to alter expression levels via genome editing methods (e.g., PHO2 ortholog, WRKY46 ortholog) might serve as a better strategy to improve PUE successfully. Apart from genetic engineering approaches, a pyramiding of multiple genes could also be achieved utilizing marker-assisted selection. The natural allelic variation within the PHT gene families and their regulatory genes (PHR1, PTF1, etc.) could also be explored to identify superior alleles with higher PUE, and thus, the same may be utilized for improving PUE in maize through a breeding approach.
Introducing phosphite metabolizing ability: an alternative strategy for improving PUE
Phosphate is the only chemical form of P that can be metabolized and assimilated by all plants and most microorganisms. However, a few species of bacteria such as Pseudomonas stutzeri WM88 strain have the natural ability to utilize phosphite (PO3−3) as a sole P source [106]. These bacteria utilize phosphite by converting /oxidizing it into phosphate with the help of the phosphite dehydrogenase (PTDH; NAD-dependent oxidoreductase class of enzyme that catalyzes the oxidation of reduced phosphite into Pi, with a consequent reduction in NAD to NADH) enzyme encoded by ptxD gene present in them. Phosphite is a reduced form of P which is easily absorbed by plants via phosphate transporters. The fertilizer use efficiency for phosphite is much higher due to its high solubility and less reactivity with soil components and soil bacteria which provide an advantage over inefficient phosphate fertilizer. However, plants cannot metabolize and assimilate phosphite which limits its use as a fertilizer. However, the introduction of the ptxD gene into the plant genome should impart phosphite metabolizing ability to them.
This novel strategy has been proved in model plants (Arabidopsis and tobacco) via heterologous expression of the ptxD gene [107]. These transgenic tobacco and Arabidopsis plants require 30–50% lesser P input with phosphite fertilizer to produce similar productivity achieved using phosphate fertilizer [107]. Recently, the ptxD gene has been introduced into the Nipponbare cultivar of Japonica rice and the transgenic rice plants were able to metabolize phosphite as the sole P source without any yield penalty [108]. As non-transgenic plants and weeds (that compete with crop plants for resources, viz., nutrients) could not metabolize phosphate, hence, it could also act as an effective pre-and post-emergent systemic herbicide having an entirely different mode of action from the currently used herbicides [109]. The robustness of the phosphate-based farming system has been demonstrated using ptxD expressing tobacco transgenic plants at two different locations having different soil types i.e., under field conditions in Argentina [89]. These recent findings indicate that heterologous expression of the codon-optimized ptxD gene can improve P utilization by plants as well as inhibit the growth of weeds. However, to avoid any deleterious effects of phosphite, it is essential to engineer maize plants for phosphite oxidation as well as subsequent metabolism; which in turn would support its sustainability as fertilizer. As a result, imparting phosphite metabolizing ability in maize might aid in the development of maize cultivars with improved PUE (phosphite as a P source) and effective weed management, resulting in sustainable maize production with a minimal environmental impact.
Conclusions and future perspectives
A comprehensive understanding of Pi-starvation signaling may lead to the development of improved maize cultivars requiring a lesser amount of external fertilizers which, in turn, would be beneficial to achieve sustainable and profitable agriculture. PUE can be effectively boosted in maize by exploiting the existing genetic diversity for RSA-associated traits. MAS can be employed to generate the PUE efficient maize ideotype by combining favorable RSA traits via stacking/pyramiding major QTLs. Further, considering the available research findings related to P uptake and translocation in plants (mainly Arabidopsis, rice, and maize) under sufficient- and low-P conditions, an array of molecular targets, viz., key regulatory components such as transcription factors (PHR1/PTF1/PHO2 homolog; these are the essential regulator of PSR and are also involved in N/P crosstalk) and miRNAs (mainly miR399, miR827 family members), crucial phosphate transporters [few transporters involved in Pi mobilization and re-mobilization (ZmPht 1;2 and 1;3), Pho1 orthologs] and ZmPSTOL have emerged as promising candidates for manipulation to improve Pi-acquisition and-use efficiency. Further, intracellular phosphate transporters and natural allelic variation within transporters and key regulators might be explored for improving PUE in maize. Being a complex trait, tissue-specific modulation of multiple targets/genes simultaneously might be more beneficial in enhancing maize productivity under phosphate-deprived conditions. For acidic soils, engineering maize plants for combined tolerance to P deficiency and Al toxicity could be a viable approach for achieving high PUE as excessive Al3+ inhibits root growth and development. On the other hand, engineering maize plants for phosphite oxidation (through heterologous expression of the ptxD gene) as well as subsequent metabolism, could potentially address the problem of low fertilizer usage efficiency and effective weed management in maize. However, utilizing a marker-assisted molecular breeding approach and/or targeted knocking out major negative regulators via genome-editing tools (so that the final edited plant should be transgene-free) to achieve high PUE might be good strategies to avoid deploying genetically modified (GM) plants in the field. Further studies focusing on decoding the molecular mechanisms of PHTs in maize and cross-talk between P and other nutrients (N, K, etc.) signaling pathways would pave the way to identify novel targets and strategies for better P management.
Data availability
Not applicable.
References
Lott JNA (1984) Accumulation of seed reserves of phosphorus and other minerals. In: Murray DR (ed) Seed physiology, vol I. Academic Press, Sydney, pp 139–166
Wang C, Ning P (2019) Post-silking phosphorus recycling and carbon partitioning in maize under low to high phosphorus inputs and their effects on grain yield. Front Plant Sci 10:784
Van Kauwenbergh SJ (2010) World Phosphate Rock Reserves and Resources. International Fertilizer Development Center, IFDC Technical Bulletin No. 75. Alabama: Muscle Shoals, 58. https://pdf.usaid.gov/pdf_docs/Pnadw835.PDF
Cordell D, White S (2011) Peak phosphorus: clarifying the key issues of a vigorous debate about long-term phosphorus security. Sustainability 3(10):2027–2049
Vu DT, Tang C, Armstrong RD (2008) Changes and availability of P fractions following 65 years of P application to a calcareous soil in a Mediterranean climate. Plant Soil 304:21–33
Raghothama KG (1999) Phosphate acquisition. Annu Rev Plant Physiol Plant Mol Biol 50:665–693. https://doi.org/10.1146/annurev.arplant.50.1.665
Lambers H, Plaxton WC (2015) Phosphorus: back to the roots. In: Plaxton WC, Lambers H (eds) Annual plant reviews volume 48: phosphorus metabolism in plants. Wiley, Hoboken, pp 1–22
Raghothama KG (2000) Phosphate transport and signaling. Curr Opin Plant Biol 3(3):182–187
Versaw WK, Harrison MJ (2002) A chloroplast phosphate transporter, PHT2;1, influences allocation of phosphate within the plant and phosphate-starvation responses. Plant Cell 14(8):1751–1766
López-Arredondo DL, Leyva-González MA, González-Morales SI, López-Bucio J, Herrera-Estrella L (2014) Phosphate nutrition: improving low-phosphate tolerance in crops. Annu Rev Plant Physiol 65:95–123
Nussaume L, Kanno S, Javot H, Marin E, Pochon N, Ayadi A et al (2011) Phosphate import in plants: focus on the PHT1 transporters. Front Plant Sci 2:83
Xu G, Chague V, Melamed B et al (2007) Functional characterization of LePT4: a phosphate transporter in tomato with mycorrhiza enhanced expression. J Exp Bot 58(10):2491–2501
Smith SE, Anderson IC, Smith FA (2015) Mycorrhizal associations and P acquisition: from cells to ecosystems. In: Plaxton WC, Lambers H (eds) Phosphorus metabolism in plants, Annual plant reviews. Wiley, Oxford, pp 409–440
Ai P, Sun S, Zhao J, Fan X, Xin W et al (2009) Two rice phosphate transporters, OsPht1;2 and OsPht1;6, have different functions and kinetic properties in uptake and translocation. Plant J 57(5):798–809
Stigter KA, Plaxton WC (2015) Molecular mechanisms of phosphorus metabolism and transport during leaf senescence. Plants (Basel) 4(4):773–798
Gu M, Chen A, Sun S, Xu G (2016) Complex regulation of plant phosphate transporters and the gap between molecular mechanisms and practical application: what is missing? Mol Plant 9(3):396–416. https://doi.org/10.1016/j.molp.2015.12.012
Nagy R, Vasconcelos MJV, Zhao S, McElver J, Bruce W et al (2006) Differential regulation of five Pht1 phosphate transporters from maize (Zea mays L.). Plant Biol 8(2):186–197. https://doi.org/10.1055/s-2005-873052
Liu F, Xu Y, Jiang H, Jiang C, Du Y, Gong C, Wang W, Zhu S, Han G, Cheng B (2016) Systematic identification, evolution and expression analysis of the Zea mays PHT1 gene family reveals several new members involved in root colonization by arbuscular mycorrhizal fungi. Int J Mol Sci 17(6):930. https://doi.org/10.3390/ijms17060930
Sawers RJ, Svane SF, Quan C, Gronlund M, Wozniak B, Gebreselassie MN, Gonzalez-Munoz E, Montes RAC, Baxter I, Goudet J et al (2017) Phosphorus acquisition efficiency in arbuscular mycorrhizal maize is correlated with the abundance of root external hyphae and the accumulation of transcripts encoding PHT1 phosphate transporters. New Phytol 214(2):632–643. https://doi.org/10.1111/nph.14403
Tian H, Drijber RA, Li X, Miller DN, Wienhold BJ (2013) Arbuscular mycorrhizal fungi differ in their ability to regulate the expression of phosphate transporters in maize (Zea mays L.). Mycorrhiza 23:507–514. https://doi.org/10.1007/s00572-013-0491-1
Liu F, Xu Y, Han G, Wang W, Li X, Cheng B (2018) Identification and functional characterization of a maize phosphate transporter induced by mycorrhiza formation. Plant Cell Physiol 59(8):1683–1694. https://doi.org/10.1093/pcp/pcy094
Xu Y, Liu F, Li X, Cheng B (2018) The mycorrhiza-induced maize ZmPt9 gene affects root development and phosphate availability in nonmycorrhizal plant. Plant Signal Behav 13(12):e1542240. https://doi.org/10.1080/15592324.2018.1542240
Roch GV, Marharajan T, Ceasar SA, Ignacimuthu S (2019) The role of PHT1 family transporters in the acquisition and redistribution of phosphorus in plants. Crit Rev Plant Sci 38(3):171–198. https://doi.org/10.1080/07352689.2019.1645402
Hamburger D, Rezzonico E, Petetot MacDonald-Comber J et al (2002) Identification and characterization of the Arabidopsis PHO1 gene involved in phosphate loading to the xylem. Plant Cell 14:889–902. https://doi.org/10.1105/tpc.000745
Wang C, Yue W, Ying Y, Wang S, Secco D, Liu Y, Whelan J, Tyerman SD, Shou H (2015) Rice SPX-Major Facility Superfamily3, a vacuolar phosphate efflux transporter, is involved in maintaining phosphate homeostasis in rice. Plant Physiol 169:2822–2831. https://doi.org/10.1104/pp.15.01005
Sun Y, Mu C, Chen Y et al (2016) Comparative transcript profiling of maize inbreds in response to long-term phosphorus deficiency stress. Plant Physiol Biochem 109:467–481. https://doi.org/10.1016/j.plaphy.2016.10.017
Gaume A, Machler F, De Leon C, Narro L, Frossard E (2001) Low-P tolerance by maize (Zea mays L) genotypes: significance of root growth, and organic acids and acid phosphatase root exudation. Plant Soil 228:253–264. https://doi.org/10.1023/A:1004824019289
Zhang Z, Liao H, Lucas WJ (2014) Molecular mechanisms underlying phosphate sensing, signaling, and adaptation in plants. J Integr Plant Biol 56(3):192–220. https://doi.org/10.1111/jipb.12163
Devaiah BN, Karthikeyan AS, Raghothama KG (2007) WRKY75 transcription factor is a modulator of phosphate acquisition and root development in Arabidopsis. Plant Physiol 143(4):1789–1801. https://doi.org/10.1104/pp.106.093971
Zhou J, Jiao FC, Wu ZC et al (2008) OsPHR2 is involved in phosphate-starvation signaling and excessive phosphate accumulation in shoots of plants. Plant Physiol 146(4):1673–1686. https://doi.org/10.1104/pp.107.111443
Chen YF, Li LQ, Xu Q, Kong YH, Wang H, Wu WH (2009) The WRKY6 transcription factor modulates PHOSPHATE1 expression in response to low Pi stress in Arabidopsis. Plant Cell 21(11):3554–3566. https://doi.org/10.1105/tpc.108.064980
Li Z, Gao Q, Liu Y, He C, Zhang X, Zhang J (2011) Overexpression of transcription factor ZmPTF1 improves low phosphate tolerance of maize by regulating carbon metabolism and root growth. Planta 233:1129–1143. https://doi.org/10.1007/s00425-011-1368-1
Wang J, Sun J, Miao J, Guo J, Shi Z, He M et al (2013) A phosphate starvation response regulator Ta-PHR1 is involved in phosphate signalling and increases grain yield in wheat. Ann Bot 111(6):1139–1153. https://doi.org/10.1093/aob/mct080
Su T, Xu Q, Zhang FC, Chen Y, Li LQ, Wu WH, Chen YF (2015) WRKY42 modulates phosphate homeostasis through regulating phosphate translocation and acquisition in Arabidopsis. Plant Physiol 167(4):1579–1591. https://doi.org/10.1104/pp.114.253799
Rubio V, Linhares F, Solano R, Martin AC, Iglesias J, Leyva A, Paz-Ares J (2001) A conserved MYB transcription factor involved in phosphate starvation signaling both in vascular plants and in unicellular algae. Genes Dev 15(16):2122–2133. https://doi.org/10.1101/gad.204401
Wang Z, Ruan W, Shi J et al (2014) Rice SPX1 and SPX2 inhibit phosphate starvation responses through interacting with PHR2 in a phosphate- dependent manner. Proc Natl Acad Sci USA 111(41):14953–14958. https://doi.org/10.1073/pnas.1404680111
Pei LM, Jin Z, Li KP, Yin HY, Wang JM, Yang AF (2013) Identification and comparative analysis of low phosphate tolerance-associated microRNAs in two maize genotypes. Plant Physiol Biochem 70:221–234. https://doi.org/10.1016/j.plaphy.2013.05.043
Tian J, Liao H (2015) The role of intracellular and secreted purple acid phosphatases in plant phosphorus scavenging and recycling. In: Plaxton WC, Lambers H (eds) Phosphorus metabolism in plants annual plant reviews, vol 48. Wiley, Hoboken, pp 265–288
Zhu H, Qian W, Lu X, Li D, Liu X, Liu K, Wang D (2005) Expression patterns of purple acid phosphatase genes in Arabidopsis organs and functional analysis of AtPAP23 predominantly transcribed in flower. Plant Mol Biol 59(4):581–594. https://doi.org/10.1007/s11103-005-0183-0
Zhang Q, Wang C, Tian J, Li K, Shou H (2011) Identification of rice purple acid phosphatases related to posphate starvation signalling. Plant Biol 13:7–15. https://doi.org/10.1111/j.1438-8677.2010.00346.x
Li C, Gui S, Yang T, Walk T, Wang X, Liao H (2012) Identification of soybean purple acid phosphatase genes and their expression responses to phosphorus availability and symbiosis. Ann Bot 109(1):275–285. https://doi.org/10.1093/aob/mcr246
Gonzalez-Munoz E, Avendano-Vazquez AO, Montes RA, Sde F, Andres-Hernandez L, Abreu-Goodger C, Sawers RJ (2015) The maize (Zea mays ssp mays var B73) genome encodes 33 members of the purple acid phosphatase family. Front Plant Sci 6:341. https://doi.org/10.3389/fpls.2015.00341
Bhadouria J, Singh AP, Mehra P, Verma L, Srivastawa R, Parida SK, Giri J (2017) Identification of purple acid phosphatases in chickpea and potential roles of CaPAP7 in seed phytate accumulation. Sci Rep 7:11012. https://doi.org/10.1038/s41598-017-11490-9
Lin HJ, Gao J, Zhang ZM, Shen YO, Lan H, Liu L et al (2013) Transcriptional responses of maize seedling root to phosphorus starvation. Mol Biol Rep 40:5359–5379. https://doi.org/10.1007/s11033-013-2636-x
Xu Y, Liu F, Han G, Cheng B (2018) Genome-wide identification and comparative analysis of phosphate starvation-responsive transcription factors in maize and three other gramineous plants. Plant Cell Rep 37:711–726. https://doi.org/10.1007/s00299-018-2262-0
Li Z, Zhang X, Liu X, Zhao Y, Wang B, Zhang J (2016) miRNA alterations are important mechanism in maize adaptations to low-phosphate environments. Plant Sci 252:103–117. https://doi.org/10.1016/j.plantsci.2016.07.009
Lynch JP (2019) Root phenotypes for improved nutrient capture: an underexploited opportunity for global agriculture. New Phytol 223(2):548–564. https://doi.org/10.1111/nph.15738
Trachsel S, Kaeppler SM, Brown KM, Lynch JP (2010) Shovelomics: high throughput phenotyping of maize (Zea mays L.) root architecture in the field. Plant Soil 341:75–87. https://doi.org/10.1007/s11104-010-0623-8
Postma JA, Dathe A, Lynch JP (2014) The optimal lateral root branching density for maize depends on nitrogen and phosphorus availability. Plant Physiol 166:590–602. https://doi.org/10.1104/pp.113.233916
Sun B, Gao Y, Lynch JP (2018) Large crown root number improves topsoil foraging and phosphorus acquisition. Plant Physiol 177(1):90–104. https://doi.org/10.1104/pp.18.00234
Walk TC, Jaramillo R, Lynch JP (2006) Architectural tradeoffs between adventitious and basal roots for phosphorus acquisition. Plant Soil 279:347–366. https://doi.org/10.1007/s11104-005-0389-6
Zhu J, Lynch JP (2004) The contribution of lateral rooting to phosphorus acquisition efficiency in maize (Zea mays L.) seedlings. Funct Plant Biol 31(10):949–958. https://doi.org/10.1071/FP04046
Zhu J, Kaeppler SM, Lynch JP (2005) Mapping of QTL controlling root hair length in maize (Zea mays L) under phosphorus deficiency. Plant Soil 270(1):299–310. https://doi.org/10.1007/s11104-004-1697-y
Gahoonia TS, Nielsen NE (2004) Root traits as tools for creating phosphorus efficient crop varieties. Plant Soil 260:47–57. https://doi.org/10.1023/B:PLSO.0000030168.53340.bc
Galindo-Castaneda T, Brown KM, Lynch JP (2018) Reduced root cortical burden improves growth and grain yield under low phosphorus availability in maize. Plant Cell Environ 41:1579–1592. https://doi.org/10.1111/pce.13197
Postma JA, Lynch JP (2010) Theoretical evidence for the functional benefit of root cortical aerenchyma in soils with low phosphorus availability. Ann Bot 107:829–841. https://doi.org/10.1093/aob/mcq199
Zhu J, Kaeppler SM, Lynch JP (2005) Mapping of QTLs for lateral root branching and length in maize (Zea mays L) under differential phosphorus supply. Theor Appl Genet 111(4):688–695. https://doi.org/10.1007/s00122-005-2051-3
Zhu J, Mickelson SM, Kaeppler SM, Lynch JP (2006) Detection of quantitative trait loci for seminal root traits in maize (Zea mays L) seedlings grown under differential phosphorus levels. Theor Appl Genet 113(1):1–10. https://doi.org/10.1007/s00122-006-0260-z
Chen J, Xu L (2011) The candidate QTLs affecting phosphorus absorption efficiency and root weight in maize (Zea mays L). Front Agric China 5(4):456–462. https://doi.org/10.1007/s11703-011-1079-1
Azevedo GC, Cheavegatti-Gianotto A, Negri BF et al (2015) Multiple interval QTL mapping and searching for PSTOL1 homologs associated with root morphology, biomass accumulation and phosphorus content in maize seedlings under low-P. BMC Plant Biol 15(1):1–7. https://doi.org/10.1186/s12870-015-0561-y
Chen J, Xu L (2011) Comparative mapping of QTLs for H+ secretion of root in maize (Zea mays L) and cross phosphorus levels on two growth stages. Front Agric China 5(3):284–290. https://doi.org/10.1007/s11703-011-1075-5
Qin H, Cai Y, Sun H, Wang J, Wang G, Liu Z (2011) QTL mapping of root exudates related to phosphorus efficiency in maize (Zea mays L). J Agric Biotechnol 19(1):93–101
Qiu H, Liu C, Yu T, Mei X, Wang G, Wang J, Cai Y (2014) Identification of QTL for acid phosphatase activity in root and rhizosphere soil of maize under low phosphorus stress. Euphytica 197(1):133–143. https://doi.org/10.1007/s10681-013-1058-0
Mendes FF, Guimarães LJ, Souza JC, Guimarães PE, Magalhaes JV et al (2014) Genetic architecture of phosphorus use efficiency in tropical maize cultivated in a low-p soil. Crop Sci 54(4):1530–1538. https://doi.org/10.2135/cropsci2013.11.0755
Brady SM, Song S, Dhugga KS, Rafalski A, Benfey PN (2006) Combining expression and comparative evolutionary analysis: the COBRA gene family. Plant Physiol 143:172–187. https://doi.org/10.1104/pp.106.087262
Wen TJ, Hochholdinger F, Sauer M, Bruce W, Schnable PS (2005) The roothairless1 gene of maize encodes a homolog of sec3, which is involved in polar exocytosis. Plant Physiol 138:1637–1643. https://doi.org/10.1104/pp.105.062174
Chen J, Xu L, Cai Y, Xu J (2008) QTL mapping of phosphorus efficiency and relative biologic characteristics in maize (Zea mays L) at two sites. Plant Soil 313(1):251–266. https://doi.org/10.1007/s11104-008-9698-x
Gu R, Chen F, Long L, Cai H, Liu Z et al (2016) Enhancing phosphorus uptake efficiency through QTL-based selection for root system architecture in maize. J Gene Genomics 43(11):663–672. https://doi.org/10.1016/j.jgg.2016.11.002
Henry A, Rosas JC, Beaver JS, Lynch JP (2010) Multiple stress response and belowground competition in multilines of common bean (Phaseolus vulgaris L). Field Crops Res 117:209–218. https://doi.org/10.1016/j.fcr.2010.03.004
Kant S, Peng M, Rothstein SJ (2011) Genetic regulation by NLA and microRNA827 for maintaining nitrate-dependent phosphate homeostasis in Arabidopsis. PLoS Genet 7:e1002021. https://doi.org/10.1371/journal.pgen.1002021
Park BS, Seo JS, Chua NH (2014) Nitrogen limitation adaptation recruits phosphate2 to target the phosphate transporter PT2 for degradation during the regulation of Arabidopsis phosphate homeostasis. Plant Cell 26(1):454–464. https://doi.org/10.1105/tpc.113.120311
Liu H, Yang H, Wu C, Feng J, Liu X, Qin H, Wang D (2009) Overexpressing HRS1 confers hypersensitivity to low phosphate-elicited inhibition of primary root growth in Arabidopsis thaliana. J Integr Plant Biol 51(4):382–392. https://doi.org/10.1111/j.1744-7909.2009.00819.x
Medici A, Marshall-Colon A, Ronzier E, Szponarski W, Wang R, Gojon A, Crawford NM et al (2015) AtNIGT1/HRS1 integrates nitrate and phosphate signals at the Arabidopsis root tip. Nat Commun 6(1):1–1
Maeda Y, Konishi M, Kiba T, Sakuraba Y, Sawaki N, Kurai T et al (2018) A NIGT1-centred transcriptional cascade regulates nitrate signalling and incorporates phosphorus starvation signals in Arabidopsis. Nat Commun 9(1):1–4
Medici A, Szponarski W, Dangeville P, Safi A, Dissanayake IM, Saenchai C et al (2019) Identification of molecular integrators shows that nitrogen actively controls the phosphate starvation response in plants. Plant Cell 31(5):1171–1184. https://doi.org/10.1105/tpc.18.00656
Hu B, Jiang Z, Wang W, Qiu Y, Zhang Z, Liu Y et al (2019) Nitrate–NRT1 1B–SPX4 cascade integrates nitrogen and phosphorus signalling networks in plants. Nat Plants 5(4):401–413
Lv Q, Zhong Y, Wang Y, Wang Z, Zhang L, Shi J et al (2014) SPX4 negatively regulates phosphate signaling and homeostasis through its interaction with PHR2 in rice. Plant Cell 26(4):1586–1597. https://doi.org/10.1105/tpc.114.123208
Sega P, Pacak A (2019) Plant PHR transcription factors: put on a map. Genes. https://doi.org/10.3390/genes10121018
Wang X, Wang HF, Chen Y, Sun MM, Wang Y, Chen YF (2020) The transcription factor NIGT1 2 modulates both phosphate uptake and nitrate influx during phosphate starvation in arabidopsis and maize. Plant Cell 32(11):3519–3534. https://doi.org/10.1105/tpc.20.00361
Hu B, Chu C (2019) Nitrogen–phosphorus interplay: old story with molecular tale. New Phytol 225(4):1455–1460. https://doi.org/10.1111/nph.16102
Wang YH, Garvin DF, Kochian LV (2002) Rapid induction of regulatory and transporter genes in response to phosphorus, potassium, and iron deficiencies in tomato roots. Evidence for cross talk and root/rhizosphere-mediated signals. Plant Physiol 130(3):1361–1370
Rubio F, Fon M, Ródenas R, Nieves-Cordones M, Alemán F, Rivero RM, Martínez V (2014) A low K+ signal is required for functional high-affinity K+ uptake through HAK5 transporters. Physiol Plant 152(3):558–570
Ródenas R, Martínez V, Nieves-Cordones M, Rubio F (2019) High external K+ concentrations impair Pi nutrition, induce the phosphate starvation response, and reduce arsenic toxicity in Arabidopsis plants. Int J Mol Sci 20(9):2237
Zhu Z, Li D, Wang P, Li J, Lu X (2020) Transcriptome and ionome analysis of nitrogen, phosphorus and potassium interactions in sorghum seedlings. Theor Exp Plant Physiol 32(4):271–285
Poirier Y, Jung JY (2015) Phosphate transporters. In: Plaxton WC, Lambers H (eds) Phosphorus metabolism in plants, annual plant reviews. Wiley, Oxford, pp 125–158
Wu P, Xu JM (2010) Does OsPHR2, central Pi-signaling regulator, regulate some unknown factors crucial for plant growth? Plant Signal Behav 5(6):712–714. https://doi.org/10.4161/psb.5.6.11645
Rae AL, Jarmey JM, Mudge SR, Smith FW (2004) Over expression of a high-affinity phosphate transporter in transgenic barley does not enhance phosphorus uptake rates. Funct Plant Biol 31:141–148. https://doi.org/10.1071/FP03159
Oono Y, Kawahara Y, Yazawa T, Kanamori H, Kuramata M, Yamagata H et al (2013) Diversity in the complexity of phosphate starvation transcriptomes among rice cultivars based on RNA-Seq profiles. Plant Mol Biol 83:523–537. https://doi.org/10.1007/s11103-013-0106-4
Heuer S, Gaxiola R, Schilling R, Herrera-Estrella L, López-Arredondo D, Wissuwa M, Delhaize E, Rouached H (2017) Improving phosphorus use efficiency: a complex trait with emerging opportunities. Plant J 90(5):868–885
Gamuyao R, Chin JH, Pariasca-Tanaka J, Pesaresi P, Dalid C, Slamet-Loedin I, Tecson-Mendoza EM, Wissuwa M, Heuer S (2012) The protein kinase OsPSTOL1 from traditional rice confers tolerance of phosphorus deficiency. Nature 488(7412):535–539. https://doi.org/10.1038/nature11346
Yang H, Knapp J, Koirala P, Rajagopal D, Peer WA, Silbart LK, Murphy A, Gaxiola RA (2007) Enhanced phosphorus nutrition in monocots and dicots over-expressing a phosphorus-responsive type I H+-pyrophosphatase. Plant Biotechnol J 5(6):735–745. https://doi.org/10.1111/j.1467-7652.2007.00281.x
Yang H, Zhang X, Gaxiola RA, Xu G, Peer WA, Murphy AS (2014) Over-expression of the Arabidopsis proton-pyrophosphatase AVP1 enhances transplant survival, root mass, and fruit development under limiting phosphorus conditions. J Exp Bot 65(12):3045–3053. https://doi.org/10.1093/jxb/eru149
Yi K, Wu Z, Zhou J, Du L, Guo L, Wu Y, Wu P (2005) OsPTF1, a novel transcription factor involved in tolerance to phosphate starvation in rice. Plant Physiol 138(4):2087–2096. https://doi.org/10.1104/pp.105.063115
Chen JY, Liu Y, Ni J, Wang YF, Bai YH, Shi J, Gan J, Wu ZC, Wu P (2011) OsPHF1 regulates the plasma membrane localization of low- and high-affinity inorganic phosphate transporters and determines inorganic phosphate uptake and translocation in rice. Plant Physiol 157(1):269–278. https://doi.org/10.1104/pp.111.181669
Wu P, Shou HX, Xu GH, Lian XM (2013) Improvement of phosphorus efficiency in rice on the basis of understanding phosphate signaling and homeostasis. Curr Opin Plant Biol 16(2):205–212. https://doi.org/10.1016/j.pbi.2013.03.002
Wang XH, Bai JR, Liu HM, Sun Y, Shi XY, Ren ZQ (2013) Overexpression of a maize transcription factor ZmPHR1 improves shoot inorganic phosphate content and growth of Arabidopsis under low-phosphate conditions. Plant Mol Biol Rep 31:665–677. https://doi.org/10.1007/s11105-012-0534-3
Guo M, Ruan W, Li C, Huang F, Zeng M, Liu Y, Yu Y, Ding X, Wu Y, Wu Z, Mao C, Yi K, Wu P, Mo X (2015) Integrative comparison of the role of the phosphate response1 subfamily in phosphate signaling and homeostasis in rice. Plant Physiol 168(4):1762–1776. https://doi.org/10.1104/pp.15.00736
Dai X, Wang Y, Yang A, Zhang WH (2012) OsMYB2P-1, an R2R3 MYB transcription factor, is involved in the regulation of phosphate-starvation responses and root architecture in rice. Plant Physiol 159(1):169–183. https://doi.org/10.1104/pp.112.194217
Ouyang X, Hong X, Zhao X, Zhang W, He X, Ma W et al (2016) Knock out of the PHOSPHATE2 gene TaPHO2-A1 improves phosphorus uptake and grain yield under low phosphorus conditions in common wheat. Sci Rep 6:29850. https://doi.org/10.1038/srep29850
Zhang F, Wu XN, Zhou HM, Wang DF, Jiang TT, Sun YF, Cao Y, Pei WX, Sun SB, Xu GH (2014) Overexpression of rice phosphate transporter gene OsPT6 enhances phosphate uptake and accumulation in transgenic rice plants. Plant Soil 384:259–270. https://doi.org/10.1007/s11104-014-2168-8
Yan W, Chen GH, Yang LF, Gai JY, Zhu YL (2014) Overexpression of the rice phosphate transporter gene OsPT6 enhances tolerance to low phosphorus stress in vegetable soybean. Sci Hortic 177:71–76. https://doi.org/10.1016/j.scienta.2014.07.037
Chang MX, Gu M, Xia YW, Dai XL, Dai CR, Zhang J, Wang SC, Qu HY, Yamaji N, Ma JF, Xu GH (2019) OsPHT1;3 mediates uptake, translocation and remobilization of phosphate under extremely low phosphate regimes. Plant Physiol 179(2):656–670. https://doi.org/10.1104/pp.18.01097
Remy E, Cabrito TR, Batista RA, Teixeira MC, Sa-Correia I, Duque P (2012) The Pht1;9 and Pht1;8 transporters mediate inorganic phosphate acquisition by the Arabidopsis thaliana root during phosphorus starvation. New Phytol 195(2):356–371. https://doi.org/10.1111/j.1469-8137.2012.04167.x
Lapis-Gaza HR, Jost R, Finnegan PM (2014) Arabidopsis phosphate transporter1 genes PHT1;8 and PHT1;9 are involved in root-to-shoot translocation of orthophosphate. BMC Plant Biol 14:334. https://doi.org/10.1186/s12870-014-0334-z
Shi SL, Wang DF, Yan Y, Zhang F, Wang HD, Gu M et al (2013) Function of phosphate transporter OsPHT2;1 in improving phosphate utilization in rice. Chin J Rice Sci 27(5):457–465. https://doi.org/10.3969/j.issn.1001-7216.2013.05.002
Costas AM, White AK, Metcalf WW (2001) Purification and characterization of a novel Phosphorus-oxidising enzyme from Pseudomonas stutzeri WM88. J Biol Chem 276(20):17429–17436. https://doi.org/10.1074/jbc.M011764200
Lopez-Arredondo DL, Herrera-Estrella L (2012) Engineering phosphorus metabolism in plants to produce a dual fertilization and weed control system. Nat Biotechnol 30:889–893. https://doi.org/10.1038/nbt.2346
Manna M, Achary MM, Islam T, Agarwal PK, Reddy MK (2016) The development of a phosphite-mediated fertilization and weed control system for rice. Sci Rep 6:24941. https://doi.org/10.1038/srep24941
Achary VMM, Ram B, Manna M, Datta D, Bhatt A, Reddy MK et al (2017) Phosphite: a novel P-fertilizer for weed management and pathogen control. Plant Biotechnol J 15(12):1493–1508. https://doi.org/10.1111/pbi.12803
Franco-Zorrilla JM, Gonzalez E, Bustos R, Linhares F, Leyva A, Paz-Ares J (2004) The transcriptional control of plant responses to phosphate limitation. J Exp Bot 55(396):285–293. https://doi.org/10.1093/jxb/erh009
Calderon-Vazquez C, Ibarra-Laclette E, Caballero-Perez J, Herrera-Estrella L (2008) Transcript profiling of Zea mays roots reveals gene responses to phosphate deficiency at the plant- and species-specific levels. J Exp Bot 59(9):2479–2497. https://doi.org/10.1093/jxb/ern115
Yadava P, Dayaman V, Agarwal A, Kumar K, Singh I, Verma R, Kaul T (2022) Fine-tuning the transcriptional regulatory model of adaptation response to phosphate stress in maize (Zea mays L). Physiol Mol Biol Plants. https://doi.org/10.1007/s12298-022-01155-x
Acknowledgements
We gratefully acknowledge the Indian Council of Agricultural Research (ICAR) and the National Agricultural Science Fund (NASF/GTR-5004/2015-16/204) for financial support. AKJ acknowledges ICAR-IIMR support in the form of YP fellowship under the in-house project (Project Code: AR:DMR:16:02).
Funding
National Agricultural Science Fund (NASF/GTR-5004/2015–16/204).
Author information
Authors and Affiliations
Contributions
KK conceptualized the idea; KK, MC, and MG collected literature and prepared the preliminary draft, which was edited and improved by PY, SHW, ZAD, BK, and SR. KK and AKJ prepared figures.
Corresponding authors
Ethics declarations
Competing interest
The authors have no financial and non-financial interests.
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kumar, K., Yadava, P., Gupta, M. et al. Narrowing down molecular targets for improving phosphorus-use efficiency in maize (Zea mays L.). Mol Biol Rep 49, 12091–12107 (2022). https://doi.org/10.1007/s11033-022-07679-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-022-07679-5