Abstract
Amides are an important type of synthetic intermediate used in the chemical, agrochemical, pharmaceutical, and nutraceutical industries. The traditional chemical process of converting nitriles into the corresponding amides is feasible but is restricted because of the harsh conditions required. In recent decades, nitrile hydratase (NHase, EC 4.2.1.84) has attracted considerable attention because of its application in nitrile transformation as a prominent biocatalyst. In this review, we provide a comprehensive survey of recent advances in NHase research in terms of natural distribution, enzyme screening, and molecular modification on the basis of its characteristics and catalytic mechanism. Additionally, industrial applications and recent significant biotechnology advances in NHase bioengineering and immobilization techniques are systematically summarized. Moreover, the current challenges and future perspectives for its further development in industrial applications for green chemistry were also discussed. This study contributes to the current state-of-the-art, providing important technical information for new NHase applications in manufacturing industries.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Compared with traditional chemical catalysts, biocatalysts are of great interest because of their significant advantages in terms of reaction conditions, such as their high efficiency, excellent selectivity (regioselectivity, chemoselectivity and enantioselectivity), and eco-friendly reaction conditions. Thus, “green” biocatalysts have gained much attention, providing another route for the industrial production of bulk chemicals and pharmaceuticals (de Carvalho 2011; Du et al. 2011; Patel 2011; Wang et al. 2012; Lee et al. 2019).
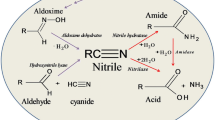
Nitriles, organic compounds widely present in nature, have attracted much attention in the chemical market for the synthesis of important amides (acrylamide (Fleming et al. 2010), nicotinamide (Nikas et al. 2020), cyanoverlamide (Wang et al. 2020b), and drug intermediates (Banerjee et al. 2016). Nitrile hydratase (NHase), which is a key enzyme in the bienzymatic pathway of nitrile degradation, catalyzes the conversion of nitriles to the corresponding amides (Jiao et al. 2020; Cui et al. 2014). NHase has been found in a variety of microbes belonging to various species of diverse genera, including Actinobacteria, Proteobacteria, Cyanobacteria and Firmicutes, since it was initially identified in the bacterium Arthrobacter sp. J1 (now known as Rhodococcus rhodochrous J1) in 1980 (Asano et al. 1980). With the increasing demand for NHase, many studies have focused on screening and modifying NHase enzymes for green industrial chemistry (Ma et al. 2024a; Wang et al. 2022; Guo et al. 2024; Zhang et al. 2023). Yamada et al. for the first time achieved large-scale industrial production of acrylamide in three NHase producing strains: R. rhodochrous J1, Rhodococcus sp. N-774 and Pseudomonas chlororaphis B23 (Yamada et al. 1996). To date, the third-generation industrial strain R. rhodochrous J1 has dominated in the industrial production of amides, especially acrylamide and nicotinamide. In China, Shen et al. used Nocardia sp. to industrialize acrylamide and nicotinamide production (Wang et al. 2007). Additionally, NHase also plays an important role in the textile industry; e.g., it can improve the properties of polyacrylonitrile (PAN) fibres as a synthetic block (Tauber et al. 2000; Guebitz and Cavaco-Paulo 2008). In addition, on the basis of great achievements in synthetic biology, the industrial production of amides could be improved by the use of engineered strains harbour robust NHase genes.
Like many other scientific fields, the history of NHase since its discovery to date is the journey from academia to industry, which has entered a new era to create more possibilities by revealing the secrets of ecological, microbiological, molecular, protein chemistry and bioremediation areas for nearly three decades. Most of the previous NHase reviews invariably covered NHase cloning, structural, and molecular characteristics, mechanisms and applications (Prasad and Bhalla 2010; Supreetha et al. 2019; Cheng et al. 2020b). This review summarizes recent NHase research progress with respect to its natural distribution, enzyme screening, molecular modification, industrial application and recent significant biotechnology. Finally, we briefly discuss the challenges, opportunities, and future prospects for its further development in industrial applications for green chemistry. This review provides useful information and insight for basic research and the industrial application of NHase.
Natural distribution of NHase
In nature, it has been reported that the distribution of NHase-producing microorganisms is very widespread, with bacteria accounting for the majority of producing microorganisms, mainly Actinomycetes and Proteobacteria. However, the majority of NHases are obtained from various species of Rhodococcus (Prasad and Bhalla 2010). Recently, NHase genes have been found in the genomes of some eukaryotes, such as Monosiga brevicollis (Tanii 2017). Foerstner et al. explored the NHase gene cluster through sequence-based metagenomic screening method and reported an unusual NHase structure consisting of two usually separated NHase subunits fused in one protein, which might open a new way to study the structure and function of eukaryotic NHases further (Foerstner et al. 2008). However, there is no related research report on gene function identification of NHase in the eukaryote. Therefore, bacteria are still the main source of NHase. Recently, Zhou et al. explored an archaeal NHase from halophilic archaeon A07HB70, which exhibits high tolerance to 3-cyanopyridine and nicotinamide, further broadening our understanding of NHase (Guo et al. 2024).
Screening methods for NHase
Traditional enrichment cultivation
Nitrile compounds, which are metabolic products of biological systems, are ubiquitous in the natural environment and exist in a variety of forms (Legras et al. 1990). It is estimated that there are hundreds of millions of microorganisms in each gram of soil. In order to screen microorganisms harboring NHase, an effective and feasible screening model is required. Conventional screening of NHase has been carried out by enrichment cultivation using selective cultures with nitriles as the sole C/N source. In recent years, owing to the rapid development of gene sequencing technology, the amount of genomic data has increased rapidly, and the researches on screening of metagenomic libraries and genome mining have also become more and more prosperous.
Function- and sequence-based screening of metagenomic libraries
The natural environment contains abundant microbial resources and is an important natural repository for biocatalysts (Kimura and Nobutada 2006). NHase is mainly distributed in microorganisms in natural environment, of which bacteria occupy the majority (Prasad and Bhalla 2010). The strains harbouring NHase isolated and screened thus far have been obtained from environments (such as wastewater, soil, and farmland) through traditional isolation and culture techniques, but less than 1% of the microbial resources in the environment can be cultivated. Metagenomics is the genomic analysis of microbial communities through expression-based or sequence-based methods without culturing; a large amount of genetic information can be obtained without cultivation (Ye et al. 2019). Owing to the use of metagenomic technology, the diversity of the obtained microbial genetic information can be greatly improved, which is conducive to the discovery of many unknown biocatalysts. In particular, the application of high-throughput screening (HTS) greatly enhances screening efficiency and increases the application scope of metagenomic technology. The screening of unknown biocatalysts via metagenomic technology can generally be divided into the following steps: sample collection, DNA extraction, library construction, screening for NHase activity, subcloning and expression, identification and sequencing (Gong et al. 2013).
Genome mining based on the conserved amino acid sequence
The traditional screening methods for NHase mostly involve isolationed from basal medium with nitrile compounds as the only nitrogen source, which is not only time-consuming and labour intensive, but also suffer from low screening efficiency. Gene mining, a network technology, allows the search and screening of homologous sequences with similar functions in the database using the nucleotide or amino acid sequences of known enzyme proteins as probes (Zhao et al. 2023). With this novel method, researchers can design primers according to the known gene sequence, utilize polymerase chain reaction to amplify the target enzyme gene and then perform functional expression of the gene in the host cell (Gong et al. 2013).
In the postgenome era, although gene resources are very abundant, a large number of gene sequences do not have corresponding functional annotations or clear biological functions (Seffernick et al. 2009). The nucleotide sequences of the subunits can be searched in NCBI, which are valuable resources for NHase studying. Genome mining provides technical support for obtaining novel NHase genes.
Molecular modification of NHase based on its characteristics and catalytic mechanism.
NHase characteristics
NHase consists of two allogenic α- and β-subunits (Fig. 1), which are usually present in equimolar amounts and generally have similar molecular weights, but NHases from different sources are structurally different. Structural analysis of NHases, revealed that the VC(T/S) LCSC(Y/T) in the α- subunit region is highly conserved, and this special site is a metal-binding domain (catalytic domain), that can coordinate with metal ions (Hashimoto et al. 2002; Miyanaga et al. 2004). According to the type of metal ions at the active site, NHases are divided into two categories: (a) Fe-containing NHase and (b) Co-containing NHase (Miller et al. 2024). Fe-NHase generally interacts with the small nitrile compounds, while the Co-NHase is considered to act more strongly on aromatic halogenated molecules (Desai and Zimmer 2004). Furthermore, Co-NHase is more robust and has wider substrate specificity compared with Fe-NHase. These metals are constitutional components of functional NHase, which fulfill a significant role in folding, stability and catalysis of the NHase polypeptide chains (Komeda et al. 1996). It is noteworthy that the nitrile substrate can only be catalysed in the interior of NHase, because the catalytic domain of NHase (the Fe−/Co-ion centre at the active site) is deeply buried in protein scaffold (Prasad and Bhalla 2010).In addition, according to the different protein molecular weight, NHase is divided into two types: low-molecular-weight (L-NHase) and high-molecular-weight NHase (H-NHase). Owing to its excellent thermal stability and organic solvent tolerance, H-NHases have been widely used in the acrylamide and nicotinamide industrial production (Miyanaga et al. 2004; Lan et al. 2017).
Crystal structure of nitrile hydratase (NHase) from Pseudonocardia thermophila JCM 3095 (PDB 1ugp). NHase consists of two allogenic α- and β- subunits. The α- subunits and β-subunits are represented in gray and purple, respectively. The square box represents the active-site center as the channel used for entering and exiting of substrate and product molecules
The characteristics of the previously reported NHasea are summarized in Table 1. The optimal pH value of the NHase reported to date is 6.5–8.5, and the optimal temperature is 20–35 °C, except for those isolated from thermophilic bacteria, e.g. Bacillus RAPc8 (60 °C) (Pereira et al., 1998), Bacillus pallidus Dac521 (50 °C) (Cramp and Cowan, 1999), and Pseudonocardia (60 °C) (Yamaki et al., 1997). The reaction involving amidase is the main rate-limiting factor for amide production, due to the existing of original amidase, which can further hydrolyse the formed amides into the corresponding carboxylic acids and ammonia. Thus, significant strategies have been developed to overcome this obstacle, for instance, the reaction can be carried out at low temperature (< 25 °C) for reducing amidase activity to negligible levels (Prasad and Bhalla 2010). Besides, cloning and expression of NHases in a heterologous host lacking amidase activity is a widely used method. Furthermore, ceasing the amidase activity through knock-out or interferring the amidase gene in parent strain is another feasible strategy (Ma et al. 2010).
Catalytic mechanism of NHase
In order to illustrate the the complete mechanism of NHase, many studies have focused on experimental and theoretical studies, and some plausible mechanisms have been proposed. To date, four catalytic mechanisms have been proposed for NHase catalysis, as shown in Fig. 2. The inner-sphere mechanism indicates that nitriles initially bind to metal ions, after which the binding group is hydrolysed by water molecules (Fig. 2A). Sugiura and Kuwahara et al. used electron spin resonance spectroscopy (ESR) to analyse the status of Fe ions in P. chlororaphis B23 NHase, and reported that the spectra shifted when the acrylic nitrile was added as the optimal substrate. However, when isobutyronitrile (not the catalytic substrate) and the product proacrylamide were added, no change in the spectra occurred, suggesting that the substrate may be directly connected to the metal ion (Fe3+) (Sugiura et al., 1988). The outer-sphere mechanism demonstrated that the hydroxide ion liberated from water molecule coordinated with metal ion and catalysed the nitrile substrate hydrolysis reaction (Prasad and Bhalla 2010) (Fig. 2B). This catalytic mechanism has been confirmed by many studies. Peplowski et al. used a computer-aided molecular docking technique to analyse the conformation of a Co-type NHase from P. thermophila JCM 3095 with different substrates and products, and the results supported the outer-sphere mechanism (Peplowski et al., 2007). Kubiak and Nowak et al. used molecular dynamics simulation technology to analyse the action mechanism of NHase derived from Rhodococcus sp. N-771, and the results also showed that the water molecules connected with metal ions directly attacked the cyanide carbon atoms in nitrile compounds (Kubiak and Nowak 2008). Yu et al. analysed the Co-type NHase of P. hermophila JCM 3095 using a more accurate semiempirical quantum mechanical calculation method, and the results also supported the catalytic mechanism (Yu et al. 2008). Another recently proposed outer-sphere mechanism indicated that the metal-bound hydroxide would activate another free water molecule from the second coordination shell, and that the second water molecule would catalyse the hydrolysis of nitriles (Mitra and Holz 2007; Yu et al. 2008; Yamanaka et al. 2010) (Fig. 2C). More recent research has further demonstrated that posttranslational sulfonate (which acts as a nucleophile) initially attacks nitriles and that the source of the product carboxamide oxygen is the protein (Nelp et al. 2016) (Fig. 2D).
Molecular modifications of NHase
Mining novel NHases from nature not only is one way to obtain NHases with high activity and superior stability, but also provides new ideas for the molecular modification of NHases on the basis of the characteristics of novel enzymes. Molecular modification has become one of the most powerful and widespread tools for engineering improved or novel functions in proteins (Ma et al. 2024b). The main molecular modification strategies to improve the robustness of NHase include stabilizing the subunit terminus, stabilizing the mesophilic zone, redesigning the active pocket, and enhancing the hydrophobic network between subunits (Fig. 3). Yokota et al. reported that thermophilins contain more polar amino acid residues and more easily form salt bridges compared with mesophilins (Yokota et al., 2006). With the increase of the total salt bridges and the proportion of the salt bridge network, the heat resistance of the protein is significantly enhanced, indicating that the salt bridge is one of the direct factors affecting the protein stability (Gurry et al. 2010). Directed evolution is an effective modification method, that can modify enzyme-encodinge genes in vitro by simulating natural evolution, error-prone PCR or chemical and physical mutagenesis (Reetz and Carballeira 2007). Zhou et al. developed a high-throughput automatic in vivo screening platform based on a niacin biosensor (NAsensor) for evolving nitrile metabolism-related enzymes (nitrilase, amidase, and NHase) (Han et al. 2022). Homologous recombination is a type of genetic recombination in which nucleotide sequences are exchanged between two similar or identical molecules of DNA.
Using this technology, researchers have integrated homologous protein fragments from different sources into a protein body to design highly variable but still naturally folded chimeric proteins (Carbone and Arnold 2007). NHases with high stability/activity can be screened by constructing a library of hybrid protein mutants.Moreover, the fusion of α- and β- subunits can effectively increase protein stability (Azzam et al. 2012; Xia et al. 2016). Additional molecular modifications of NHase are summarized in Table 2.
NHase applications
Synthesis of fine chemicals
NHase, well-known for its great impact on the revolution wave of acrylamide biosynthesis, has undergone 40 years of academic and industrial utilizationand is one of the most successful cases of biocatalysis in biology. Acrylamide, a synthetic monomer, has attracted much attention in the industrial applications of the leather industry, water treatment, enhanced oil recovery, and many other fields (Taeymans et al. 2004; Jiao et al. 2020). NHase in R. rhodochrous N-774, developed by Japan Nitto chemical industry, was the first biocatalyst for the production of acrylamide. Besides, P. chlororaphis B23 and R. rhodochrous J1, as the new generation of NHase-producing strains, have also been employed as a vehicle for acrylamide industrial production (Yamada et al. 1996). Shen et al. screened a strain Nocardia sp. 86–163 with high production of NHase in 1986, which was successfully applied in the industrial production of acrylamide (Asano 2002; Sahu et al. 2022).
What’s more, NHase also serves as an important biocatalyst for nicotinamide industrial production. Nicotinamide is an important vitamin with a wide range of industrial applications in the pharmaceutical, nutraceutical, cosmetic, and feed industries, among other fields (Prasad et al. 2007). Nicotinamide has potential as a safe, well-tolerated, and cost-effective agent to be used in cancer chemoprevention and therapy, such as laryngeal and urinary bladder cancers (Nikas et al. 2020). Among the NHases reported thus far, the NHases from R. rhodochrous J1 and R. rhodochrous PA-34 exhibit high activity towards 3-cyanopyridine (Pratush et al. 2013). Recently, the potential applications of NHase in the synthesis of other valuable amides, such as 2-(1H-indol-2-yl)-acetamide, butyramide, 5-cyanovaleramide, isonicotinamide, picolinamide, indole-3-acetamide, pyrazine-2-carboxamide, lactamide, 2H-thiopyran-6-carboxamide, 2,6-difluorobenzamide, benzamide and adipamide have been disclosed (Table 3).
Environmental bioremediation
NHase is not only widely used in the manufacture of amides, but also fulfills a role in environmental protection. Synthetic nitriles can currently be employed for organic synthesis as starting materials and intermediates. However, most of the nitriles are neurotoxic and belong to mutagenic, teratogenic and carcinogenic compounds in nature, which are continuously released as effluents by industries (Supreetha et al. 2019; Tanii 2017). Thus, removal of nitrile from industrial contaminated soil and water is urgently needed. NHase in combination with amidase or nitrilase is considered promising for potential application in nitrile biotransformation and degradation, which has made significant contributions to the environmental bioremediation in recent decades (Table 1). For instance, Kohyama et al. degraded the acetonitrile wastewater by using the dual-bacteria (R. pyridinovorans S85-2 and Brevundimonas diminuta AM10-C) coupling process, and 90% of the acetonitrile was degraded to acetic acid within 10 h (Kohyama et al. 2006). Wyatt et al. degraded highly toxic wastewater containing acrylonitrile and other compounds by mixed microorganism producing nitrile converting enzymes (i.e., NHase, nitrilase and amidase). The results revealed that the chemical oxygen demand (COD) decreased by 75%, and 99% of the COD was metabolized by cells as a result of the nitrile compounds (Wyatt and Knowles 1995). Hansen et al. realized the biodegradation of cyanide in a gold tailings environment via nitrilase, NHase and thiocyanate (Welman-Purchase et al. 2024).
Advances in NHase engineering
Whole-cell biocatalysis engineering driven by synthetic biology
NHase has been universally found in a variety of microbes belonging to various species. Compared with other hosts, Rhodococcus strains (e.g., Rhodococcus sp. M8, R. rhodochrous J1 and R. ruber TH) have become increasingly attractive for value-added amides synthesis due to their outstanding characteristics, e.g., high NHase activity and superior stability to high temperature and organic solvents, enabling their wide application in whole-cell biocatalysis (Jiao et al. 2020). Recent advances in synthetic biology have revolutionized the technology to engineer microbial hosts for the production of a wide variety of valuable amides. Thus, it is significant to highlight the achievements in Rhodococcus strains as platform organisms for amides production.
In the past several decades, basic genetic elements, including the development of different promoters, ribosome binding sites (RBSs), reporter genes, and selection markers, have been developed in engineered Rhodococcus sp. to facilitate whole-cell biocatalysis applications (Liang and Yu 2021). What’s more, to realize the large-scale amides production at an industrial level, various strategies have been explored in Rhodococcus, ranging from random mutagenesis to precise genome editing, including traditional homologous recombination, bacteriophage recombinase-assisted recombineering and the CRISPR/Cas9 system (Liang and Yu 2021). In particular, with respect to the CRISPR/Cas9 system, the development and application of CRISPR/Cas9 in Rhodococcusare important. The first CRISPR/Cas9 system in R. ruber for gene deletion, mutation, and insertion was successfully developed by introducing recombinases Che9c60 and Che9c61 in the study of Liang et al., and the editing efficiency reached 75% (Liang et al. 2020).
NHase immobilization
Although biotransformation mediated by free cells or soluble enzymes has been successful, immobilized cells or enzymes have other advantages compared with free enzymes or cells. In particular, immobilization promotes biocatalyst retention and by-product removal, simplifying the process of catalyst separation and product purification (Rangraz et al. 2024). Immobilization can also improve the reusability and stability of biocatalyst, and immobilized cells have also been reported to catalyse a wider range of substrates than free cells do (Dias et al. 2001).
The most mature technology in industry using NHase to convert nitriles into amides is to catalyze acrylonitrile to acrylamide and catalyze nicotinonitrile to nicotinamide by immobilized Rhodococcus (Raj et al. 2010; Wang et al. 2020a). The Swiss company Lonza and Japan’s Mitsubishi Corporation have successfully used these two processes to operate on a thousand-ton scale for over a decade.
It is relatively mature to immobilize NHase and cells harboring NHase using traditional matrices (such as calcium alginate, agar, and polyvinyl alcohol) and commercial immobilization materials. In recent years, the application of metal–organic frameworks and biomimetic mineralization in immobilizing NHases is emerging. Common immobilization methods such as covalent binding, embedding, cross-linking, and physical adsorption have also been reported, of which embedding is often used to immobilize the cells containing NHase (Table 4).
Commonly, immobilization of NHase has the advantages mentioned above, but also introduces a series of issues, such as unstable properties of the immobilizing supports under extreme conditions, partial loss of catalytic activity of biocatalyst, the mismatch of immobilization carrier size (Velankar et al. 2010). We hope the future research will make a breakthrough in improving the applicability and stability of the immobilized carrier.
Conclusion and Future Perspectives
The history of NHase since its discovery to date is the journey from academia to industry, and researchers have witnessed the rapid progress of NHase in aspects of industrial application, natural distribution, enzyme screening, molecular modification and significant biotechnology in amide production. NHase, as a green biocatalyst, is launching a revolutionary wave at the forefront of green biomanufacturing, e.g., the agricultural, pharmaceutical, material, and textile industries, along with the fields of chemical engineering and environmentalstudies. What is exciting that NHase has been successfully used for the industrial production of acrylamide, nicotinamide, etc. Although rapid progress has been made in the past decade, the performance of most NHases cannot achieve the goals required for large-scale industrial production because of the notable instability, unsatisfactory catalytic activity, unwanted byproduct formation, etc. In spite of the efforts discussed herein to exploit NHase for various applications, a great deal of work is still necessary to achieve the goals of “Green and Sustainable Chemistry” which are being faced by the scientific community, in both academia and industry. Emerging biological tools and strategies in synthetic biology, protein engineering, and bioinformatics have been developed to not only improve the properties of NHase but also generate novel process technologies, which will be promising for the improvement of the NHase as a robust biocatalyst. Especially, as -omics and other high throughput technologies have been rapidly developed, the promise of applying machine learning (ML) techniques in NHase design has started to become a reality.
Data availability
Data will be made available on request.
References
Asano Y (2002) Overview of screening for new microbial catalysts and their uses in organic synthesis-selection and optimization of biocatalysts. J Biotechnol 94(1):65–72. https://doi.org/10.1016/S0168-1656(01)00419-9
Asano Y, Tani Y, Yamada H (1980) A new enzyme “Nitrile Hydratase” which degrades acetonitrile in combination with amidase. Agric Biol Chem 44:2251–2252
Azzam N, Bar-Shalom R, Fares F (2012) Conversion of TSH heterodimer to a single polypeptide chain increases bioactivity and longevity. Endocrinology 153(2):954–960. https://doi.org/10.1210/en.2011-1856
Banerjee A, Sharma R (2016) The nitrile-degrading enzymes: current status and future prospects (Retraction of Vol 60, Pg 33. Appl Microbiol Biotechnol 100(16):7359–7359. https://doi.org/10.1007/s00253-016-7708-0
Carbone MN, Arnold FH (2007) Engineering by homologous recombination: exploring sequence and function within a conserved fold. Curr Opin Struct Biol 17(4):454–459. https://doi.org/10.1016/j.sbi.2007.08.005
Chen CY, Chen SC, Fingas M, Kao CM (2010) Biodegradation of propionitrile by Klebsiella oxytoca immobilized in alginate and cellulose triacetate gel. J Hazard Mater 177(1–3):856–863. https://doi.org/10.1016/j.jhazmat.2009.12.112
Chen J, Yu HM, Liu CC, Liu J, Shen ZY (2013) Improving stability of nitrile hydratase by bridging the salt-bridges in specific thermal-sensitive regions. J Biotechnol 164(2):354–362. https://doi.org/10.1016/j.jbiotec.2013.01.021
Cheng ZY, Cui WJ, Liu ZM, Zhou L, Wang M, Kobayashi M, Zhou ZM (2016) A switch in a substrate tunnel for directing regioselectivity of nitrile hydratases towards α, ω-dinitriles. Catal Sci Technol 6(5):1292–1296. https://doi.org/10.1039/c5cy01997d
Cheng Z, Cui W, Xia Y, Peplowski L, Zhou Z (2017) Modulation of nitrile hydratase regioselectivity towards dinitriles by tailoring the substrate binding pocket residues. Chem Cat Chem 10:449–458
Cheng Z, Lan Y, Guo J, Ma D, Jiang S, Lai Q, Zhou Z, Peplowski L (2020) Computational design of nitrile hydratase from pseudonocardia thermophila JCM3095 for improved thermostability. Molecules. https://doi.org/10.3390/molecules25204806
Cheng ZY, Xia YY, Zhou ZM (2020) Recent advances and promises in nitrile hydratase: from mechanism to industrial applications. Front Bioeng Biotechnol. https://doi.org/10.3389/fbioe.2020.00352
Cheng Z, Jiang S, Zhou Z (2021) Substrate access tunnel engineering for improving the catalytic activity of a thermophilic nitrile hydratase toward pyridine and pyrazine nitriles. Biochem Biophys Res Commun 575:8–13. https://doi.org/10.1016/j.bbrc.2021.08.059
Cui YT, Cui WJ, Liu ZM, Zhou L, Kobayashi M, Zhou ZM (2014) Improvement of stability of nitrile hydratase via protein fragment swapping. Biochem Biophys Res Commun 450(1):401–408. https://doi.org/10.1016/j.bbrc.2014.05.127
de Carvalho CCCR (2011) Enzymatic and whole cell catalysis: finding new strategies for old processes. Biotechnol Adv 29(1):75–83. https://doi.org/10.1016/j.biotechadv.2010.09.001
Desai LV, Zimmer M (2004) Substrate selectivity and conformational space available to bromoxynil and acrylonitrile in iron nitrile hydratase. Dalton T 6:872–877. https://doi.org/10.1039/b313380j
Dias JCT, Rezende RP, Linardi VR (2001) Bioconversion of nitriles by CCT 7207 cells immobilized in barium alginate. Appl Microbiol Biotechnol 56(5–6):757–761. https://doi.org/10.1007/s002530100681
Du J, Shao ZY, Zhao HM (2011) Engineering microbial factories for synthesis of value-added products. J Ind Microbiol Biotechnol 38(8):873–890. https://doi.org/10.1007/s10295-011-0970-3
Endo T, Watanabe I (1989) Nitrile hydratase of Rhodococcus sp. N-774 Purification and amino acid sequences. FEBS Lett 243(1):61–64
Fleming FF, Yao LH, Ravikumar PC, Funk L, Shook BC (2010) Nitrile-containing pharmaceuticals: efficacious roles of the nitrile pharmacophore. J Med Chem 53(22):7902–7917. https://doi.org/10.1021/jm100762r
Foerstner KU, Doerks T, Muller J, Raes J, Bork P (2008) A Nitrile hydratase in the eukaryote. Plos One. https://doi.org/10.1371/journal.pone.0003976
Gong JS, Lu ZM, Li H, Zhou ZM, Shi JS, Xu ZH (2013) Metagenomic technology and genome mining: emerging areas for exploring novel nitrilases. Appl Microbiol Biotechnol 97(15):6603–6611. https://doi.org/10.1007/s00253-013-4932-8
Guebitz GM, Cavaco-Paulo A (2008) Enzymes go big: surface hydrolysis and functionalisation of synthetic polymers. Trends Biotechnol 26(1):32–38. https://doi.org/10.1016/j.tibtech.2007.10.003
Guo JL, Cheng ZY, Zhou ZM (2024) An archaeal nitrile hydratase from the halophilic archaeon A07HB70 exhibits high tolerance to 3-cyanopyridine and nicotinamide. Protein Expr Purif 214:106390. https://doi.org/10.1016/j.pep.2023.106390
Gurry T, Nerenberg PS, Stultz CM (2010) The Contribution of interchain salt bridges to triple-helical stability in collagen. Biophys J 98(11):2634–2643. https://doi.org/10.1016/j.bpj.2010.01.065
Han L, Liu X, Cheng Z, Cui W, Guo J, Yin J, Zhou Z (2022) Construction and application of a high-throughput in vivo screening platform for the evolution of nitrile metabolism-related enzymes based on a desensitized repressive biosensor. Acs Synth Biol 11(4):1577–1587. https://doi.org/10.1021/acssynbio.1c00642
Hann EC, Eisenberg A, Fager SK, Perkins NE, Gallagher FG, Cooper SM, Gavagan JE, Stieglitz B, Hennessey SM, DiCosimo R (1999) 5-Cyanovaleramide production using immobilized Pseudomonas chlororaphis B23. Bioorg Med Chem 7(10):2239–2245. https://doi.org/10.1016/s0968-0896(99)00157-1
Hashimoto Y, Nishiyama M, Ikehata O (1991) Cloning and characterization of an amidase gene from Rhodococcus species N-774 and its expression in Escherichia coli. Biochimica et Biophysica Acta (BBA) - Gene Struct Expr 1088(2):225–233
Hashimoto Y, Sasaki S, Herai S, Oinuma KI, Shimizu S, Kobayashi M (2002) Site-directed mutagenesis for cysteine residues of cobalt-containing nitrile hydratase. J Inorg Biochem 91(1):70–77. https://doi.org/10.1016/S0162-0134(02)00373-2
Jiao S, Li FL, Yu HM, Shen ZY (2020) Advances in acrylamide bioproduction catalyzed with cells harboring nitrile hydratase. Appl Microbiol Biotechnol 104(3):1001–1012. https://doi.org/10.1007/s00253-019-10284-5
Kimura N (2006) Metagenomics: Access to Unculturable Microbes in the Environment. Microbes Environ 21(4):201–215
Kohyama E, Yoshimura A, Aoshima D, Yoshida T, Kawamoto H, Nagasawa T (2006) Convenient treatment of acetonitrile-containing wastes using the tandem combination of nitrile hydratase and amidase-producing microorganisms. Appl Microbiol Biotechnol 72(3):600–606. https://doi.org/10.1007/s00253-005-0298-x
Komeda H, Kobayashi M, Shimizu S (1996) Characterization of the gene cluster of high-molecular-mass nitrile hydratase (HNHase) induced by its reaction product in Rhodococcus rhodochrous J1. Proc Natl Acad Sci 93(9):4267
Kubac D, Cejková A, Masák J, Jirku V, Lemaire M, Gallienne E, Bolte J, Stloukal R, Martínková L (2006) Biotransformation of nitriles by A4 immobilized in LentiKats®. J Mol Catal B-Enzym 39(1–4):59–61. https://doi.org/10.1016/j.molcatb.2006.01.004
Kubác D, Kaplan O, Elisáková V, Pátek M, Vejvoda V, Slámová K, Tóthová A, Lemaire M, Gallienne E, Lutz-Wahl S, Fischer L, Kuzma M, Pelantová H, van Pelt S, Bolte J, Kren V, Martinková L (2008) Biotransformation of nitriles to amides using soluble and immobilized nitrile hydratase from. J Mol Catal B-Enzym 50(2–4):107–113. https://doi.org/10.1016/j.molcatb.2007.09.007
Kubiak K, Nowak W (2008) Molecular dynamics simulations of the photoactive protein nitrile hydratase. Biophys J 94(10):3824–3838. https://doi.org/10.1529/biophysj.107.116665
Lan Y, Zhang XH, Liu ZM, Zhou L, Shen RH, Zhong XP, Cui WJ, Zhou ZM (2017) Overexpression and characterization of two types of nitrile hydratases from J1. Plos One 12(6):e0179833. https://doi.org/10.1371/journal.pone.0179833
Legras JL, Chuzel G, Arnaud A (1990) Natural nitriles and their metabolism. World J Microbiol Biotechnology 6(2):83–108
Lee CY, Choi SK, Chang HN (1993) Bench-scale production of acrylamide using the resting cells of Brevibacterium sp. CH2 in a fed-batch reactor. Enzyme Microb Technol. https://doi.org/10.1016/0141-0229(93)90175-2
Lee SY, Kim HU, Chae TU, Cho JS, Kim JW, Shin JH, Kim DI, Ko YS, Jang WD (2019) Jang YS (2019) A comprehensive metabolic map for production of bio-based chemicals (vol 2, pg 18. Nat Catal 2(10):942–944. https://doi.org/10.1038/s41929-019-0358-8
Liang YX, Yu HM (2021) Genetic toolkits for engineering species with versatile applications. Biotechnol Adv 49:107748. https://doi.org/10.1016/j.biotechadv.2021.107748
Liang YX, Jiao S, Wang MM, Yu HM, Shen ZY (2020) A CRISPR/Cas9-based genome editing system for RTH. Metab Eng 57:13–22. https://doi.org/10.1016/j.ymben.2019.10.003
Ma YC, Yu HM, Pan WY, Liu CC, Zhang SL, Shen ZY (2010) Identification of nitrile hydratase-producing TH and characterization of an -negative mutant. Bioresour Technol 101(1):285–291. https://doi.org/10.1016/j.biortech.2009.07.057
Ma D, Cheng Z, Han L, Guo J, Peplowski L, Zhou Z (2024a) Structure-oriented engineering of nitrile hydratase: reshaping of substrate access tunnel and binding pocket for efficient synthesis of cinnamamide. Int J Biol Macromol 254(Pt 2):127800. https://doi.org/10.1016/j.ijbiomac.2023.127800
Ma D, Cheng ZY, Han LC, Guo JL, Peplowski L, Zhou ZM (2024) Structure-oriented engineering of nitrile hydratase: reshaping of substrate access tunnel and binding pocket for efficient synthesis of cinnamamide. Int J Biol Macromol 254:127800. https://doi.org/10.1016/j.ijbiomac.2023.127800
Maksimov A, Maksimova IuG, Kuznetsova MV, Olontsev VF, Demakov VA (2007) Immobilization of Rhodococcus ruber strain gt1, possessing nitrile hydratase activity, on carbon sorbents. Prikl Biokhim Mikrobiol 43(2):193–198
Mauger J, Nagasawa T, Yamada H (1989) Synthesis of various aromatic amide derivatives using nitrile hydratase of Rhodococcus-Rhodochrous J1. Tetrahedron 45(5):1347–1354. https://doi.org/10.1016/0040-4020(89)80133-4
Miller C, Huntoon D, Kaley N, Ogutu I, Fiedler AT, Bennett B, Liu D, Holz R (2024) Role of second-sphere arginine residues in metal binding and metallocentre assembly in nitrile hydratases. J Inorg Biochem 256:112565. https://doi.org/10.1016/j.jinorgbio.2024.112565
Mitra S, Holz RC (2007) Unraveling the catalytic mechanism of nitrile hydratases. J Biol Chem 282(10):7397–7404. https://doi.org/10.1074/jbc.M604117200
Miyanaga A, Fushinobu S, Ito K, Shoun H, Wakagi T (2004) Mutational and structural analysis of cobalt-containing nitrile hydratase on substrate and metal binding. Eur J Biochem 271(2):429–438. https://doi.org/10.1046/j.1432-1033.2003.03943.x
Nagasawa T, Nanba H, Ryuno K, Takeuchi K, Yamada H (1987) Nitrile hydratase of pseudomonas chlororaphis B23 purification and characterization. Eur J Biochem 162(3):691–698. https://doi.org/10.1111/j.1432-1033.1987.tb10692.x
Nagasawa T, Mathew CD, Mauger J, Yamada H (1988) Nitrile hydratase-catalyzed production of nicotinamide from 3-cyanopyridine in Rhodococcus rhodochrous J1. Appl Environ Microbiol 54(7):1766–1769. https://doi.org/10.1128/aem.54.7.1766-1769.1988
Nagasawa T, Takeuchi K, Yamada H (1991) Characterization of a new cobalt-containing nitrile hydratase purified from urea-induced cells of Rhodococcus rhodochrous J1. Eur J Biochem 196(3):581–589. https://doi.org/10.1111/j.1432-1033.1991.tb15853.x
Nelp MT, Song Y, Wysocki VH, Bandarian V (2016) A Protein-derived oxygen is the source of the amide oxygen of nitrile hydratases. J Biol Chem 291(15):7822–7829. https://doi.org/10.1074/jbc.M115.704791
Nikas IP, Paschou SA, Ryu HS (2020) The Role of nicotinamide in cancer chemoprevention and therapy. Biomolecules 10(3):477. https://doi.org/10.3390/biom10030477
Patel RN (2011) Biocatalysis: synthesis of key intermediates for development of pharmaceuticals. Acs Catal 1(9):1056–1074. https://doi.org/10.1021/cs200219b
Pawar SV, Yadav GD (2014) PVA/chitosan-glutaraldehyde cross-linked nitrile hydratase as reusable biocatalyst for conversion of nitriles to amides. J Mol Catal B-Enzym 101:115–121. https://doi.org/10.1016/j.molcatb.2014.01.005
Peplowski L, Kubiak K, Nowak W (2007) Insights into catalytic activity of industrial enzyme Co-nitrile hydratase. Docking studies of nitriles and amides. J Mol Model 13(6):725–730
Pereira RA, Graham D, Rainey FA, Cowan DA (1998) A novel thermostable nitrile hydratase. Extremophiles 2:347–57
Pei XL, Wu YF, Wang JP, Chen ZJ, Liu W, Su WK, Liu FM (2020) Biomimetic mineralization of nitrile hydratase into a mesoporous cobalt-based metal-organic framework for efficient biocatalysis. Nanoscale 12(2):967–972. https://doi.org/10.1039/c9nr06470b
Prasad S, Bhalla TC (2010) Nitrile hydratases (NHases): at the interface of academia and industry. Biotechnol Adv 28(6):725–741. https://doi.org/10.1016/j.biotechadv.2010.05.020
Prasad S, Raj J, Bhalla TC (2007) Bench scale conversion of 3-cyanopyidine to nicotinamide using resting cells of Rhodococcus rhodochrous PA-34. Indian J Microbiol 47(1):34–41. https://doi.org/10.1007/s12088-007-0007-9
Prasad JRS, Sharma NN, Bhalla TC (2010) Bioconversion of acrylonitrile to acrylamide using polyacrylamide entrapped cells of PA-34. Folia Microbiol 55(5):442–446. https://doi.org/10.1007/s12223-010-0074-x
Pratush A, Seth A, Bhalla TC (2013) Purification and characterization of nitrile hydratase of mutant 4D of PA-34. 3 Biotech 3:165–171
Raj J, Seth A, Prasad S, Bhalla TC (2007a) Bioconversion of butyronitrile to butyramide using whole cells of PA-34. Appl Microbiol Biotechnol 74(3):535–539. https://doi.org/10.1007/s00253-006-0693-y
Raj J, Seth A, Prasad S, Bhalla TC (2007b) Bioconversion of butyronitrile to butyramide using whole cells of Rhodococcus rhodochrous PA-34. Appl Microbiol Biotechnol 74(3):535–539. https://doi.org/10.1007/s00253-006-0693-y
Raj J, Prasad S, Sharma NN, Bhalla TC (2010) Bioconversion of acrylonitrile to acrylamide using polyacrylamide entrapped cells of Rhodococcus rhodochrous PA-34. Folia Microbiol (Praha) 55(5):442–446. https://doi.org/10.1007/s12223-010-0074-x
Rangraz Z, Amini MM, Habibi Z (2024) One-pot synthesis of 1,3,5-trisubstitued Pyrazoles via immobilized Thermomyces lanuginosus Lipase (TLL) on a metal-organic framework. ACS Omega 9(17):19089–19098. https://doi.org/10.1021/acsomega.3c09875
Reetz MT, Carballeira JD (2007) Iterative saturation mutagenesis (ISM) for rapid directed evolution of functional enzymes. Nat Protoc 2(4):891–903. https://doi.org/10.1038/nprot.2007.72
Sahu R, Meghavarnam AK, Janakiraman S (2022) Evaluation of acrylamide production by (RS-6) cells immobilized in agar matrix. J Appl Microbiol 132(3):1978–1989. https://doi.org/10.1111/jam.15303
Seffernick JL, Samanta SK, Louie TM, Wackett LP, Subramanian M (2009) Investigative mining of sequence data for novel enzymes: a case study with nitrilases. J Biotechnol 143(1):17–26. https://doi.org/10.1016/j.jbiotec.2009.06.004
Singh R, Pandey D, Devi N, Chand D (2018) Bench scale production of butyramide using free and immobilized cells of Bacillus sp. APB-6. Bioprocess Biosyst Eng 41:1225–1232. https://doi.org/10.1007/s00449-018-1951-y
Singh P, Kumari A, Chauhan K, Attri C, Seth A (2020) Nitrile hydratase mediated green synthesis of lactamide by immobilizing NIT-36 cells on N, N′-Methylene bis-acrylamide activated chitosan. Int J Biol Macromol 161:168–176. https://doi.org/10.1016/j.ijbiomac.2020.06.004
Sugiura Y, Kuwahara J, Nagasawa T, Yamada H (1988) Significant interaction between low-spin iron (III) site and pyrroloquinoline quinone in active center of nitrile hydratase. Biochem Biophys Res Commun 154:522–528. https://doi.org/10.1016/0006-291x(88)90171-4
Sun XD, Shi Y, Yu HM, Shen ZY (2004) Bioconversion of acrylnitrile to acrylamide using hollow-fiber membrane bioreactor system. Biochem Eng J 18(3):239–243. https://doi.org/10.1016/j.bej.2003.09.001
Sun WF, Zhu LB, Chen XG, Wu LJ, Zhou ZM, Liu Y (2016) The Stability enhancement of nitrile hydratase from by swapping the c-terminal domain of β subunit. Appl Biochem Biotech 178(8):1481–1487. https://doi.org/10.1007/s12010-015-1961-z
Supreetha K, Rao SN, Srividya D, Anil HS, Kiran S (2019) Advances in cloning, structural and bioremediation aspects of nitrile hydratases. Mol Biol Rep 46(4):4661–4673. https://doi.org/10.1007/s11033-019-04811-w
Taeymans D, Wood J, Ashby P, Blank I, Studer A, Stadler RH, Gondé P, Van Eijck P, Lalljie S, Lingnert H, Lindblom M, Matissek R, Müller D, Tallmadge D, O’Brien J, Thompson S, Silvani D, Whitmore T (2004) A review of acrylamide: An industry perspective on research, analysis, formation and control. Crit Rev Food Sci 44(5):323–347. https://doi.org/10.1080/10408690490478082
Tanii H (2017) Allyl nitrile: toxicity and health effects. J Occup Health 59(2):104–111. https://doi.org/10.1539/joh.16-0147-RA
Tauber MM, Cavaco-Paulo A, Robra KH, Gübitz GM (2000) Nitrile hydratase and amidase from hydrolyze acrylic fibers and granular polyacrylonitriles. Appl Environ Microbiol 66(4):1634–1638. https://doi.org/10.1128/Aem.66.4.1634-1638.2000
van Pelt S, Quignard S, Kubác D, Dimitry YSB, van Rantwijk F, Sheldon RA (2008) Nitrile hydratase CLEAs: The immobilization and stabilization of an industrially important enzyme. Green Chem 10(4):395–400. https://doi.org/10.1039/b714258g
Velankar H, Clarke KG, du Preez R, Cowan DA, Burton SG (2010) Developments in nitrile and amide biotransformation processes. Trends Biotechnol 28(11):561–569. https://doi.org/10.1016/j.tibtech.2010.08.004
Wang Y, Zheng Y, Xue J (2007) Characterization of nitrile hydratation catalysed by Nocardiasp. World J Microbiol Biotechnology 23(3):355–362
Wang M, Si T, Zhao HM (2012) Biocatalyst development by directed evolution. Bioresour Technol 115:117–125. https://doi.org/10.1016/j.biortech.2012.01.054
Wang L, Guan S, Bai J, Jiang Y, Song Y, Zheng X, Gao J (2020a) Enzyme immobilized in BioMOFs: Facile synthesis and improved catalytic performance. Int J Biol Macromol 144:19–28. https://doi.org/10.1016/j.ijbiomac.2019.12.054
Wang L, Liu SX, Du WJ, Dou TY, Liang CH (2020b) High regioselectivity production of 5-cyanovaleramide from adiponitrile by a novel nitrile hydratase derived from CCM2595. ACS Omega 5(29):18397–18402. https://doi.org/10.1021/acsomega.0c02188
Wang L, Cui B, Qiu K, Huang J, Liang C (2022) Modification of nitrile hydratase from Rhodococcus erythropolis CCM2595 by semirational design to enhance its substrate affinity. Biointerphases 17(6):061007. https://doi.org/10.1116/6.0002061
Welman-Purchase MD, Castillo J, Gomez-Arias A, Matu A, Hansen RN (2024) First insight into the natural biodegradation of cyanide in a gold tailings environment enriched in cyanide compounds. Sci Total Environ 906:167174. https://doi.org/10.1016/j.scitotenv.2023.167174
Wyatt JM, Knowles CJ (1995) Microbial degradation of acrylonitrile waste effluents: the degradation of effluents and condensates from the manufacture of acrylonitrile. Int Biodeterior Biodegrad 35(1-3):227–248
Xia YY, Cui WJ, Liu ZM, Zhou L, Cui YT, Kobayashi M, Zhou ZM (2016) Construction of a subunit-fusion nitrile hydratase and discovery of an innovative metal ion transfer pattern. Sci Rep Uk 6:19183. https://doi.org/10.1038/srep19183
Yokota K, Satou K, Ohki S (2006) Comparative analysis of protein thermostability: Differences in amino acid content and substitution at the surfaces and in the core regions of thermophilic and mesophilic proteins. Sci Technol Adv Mater 7(3):255–262
Yamaki T, Oikawa T, Ito K (1997) Cloning and sequencing of a nitrile hydratase gene from Pseudonocardia thermophila. J Ferment Bioeng 83(5):474–477
Yamada H, Kobayashi M (1996) Nitrile hydratase and its application to industrial production of acrylamide. Biosci Biotechnol Biochem 60:1391–1400. https://doi.org/10.1271/bbb.60.1391
Yamanaka Y, Hashimoto K, Ohtaki A, Noguchi K, Yohda M, Odaka M (2010) Kinetic and structural studies on roles of the serine ligand and a strictly conserved tyrosine residue in nitrile hydratase. J Biol Inorg Chem 15(5):655–665. https://doi.org/10.1007/s00775-010-0632-3
Yang ZF, Pei XL, Xu G, Wu JP, Yang LR (2019) Efficient production of 2,6-difluorobenzamide by recombinant expressing the nitrile hydratase. Appl Biochem Biotechnol 187(2):439–448. https://doi.org/10.1007/s12010-018-2823-2
Ye SH, Siddle KJ, Park DJ, Sabeti PC (2019) Benchmarking Metagenomics Tools for Taxonomic Classification. Cell 178(4):779–794. https://doi.org/10.1016/j.cell.2019.07.010
Yu HM, Liu J, Shen ZY (2008) Modeling catalytic mechanism of nitrile hydratase by semi-empirical quantum mechanical calculation. J Mol Graph Model 27(4):522–528. https://doi.org/10.1016/j.jmgm.2008.09.003
Zhang LY, Zhao SY, Chang C, Wang JA, Yang C, Cheng ZY (2023) N-terminal loops at the tetramer interface of nitrile hydratase act as “hooks” determining resistance to high amide concentrations. Int J Biol Macromol 245(12553):1. https://doi.org/10.1016/j.ijbiomac.2023.125531
Zhao YX, Guo LL, Sun SL, Guo JJ, Dai YJ (2020) Bioconversion of indole-3-acetonitrile by the N-fixing bacterium CGMCC 7333 and its -expressed nitrile hydratase. Int Microbiol 23(2):225–232. https://doi.org/10.1007/s10123-019-00094-0
Zhao F, Sun C, Liu Z, Cabrera A, Escobar M, Huang S, Yuan Q, Nie Q, Luo KL, Lin A, Vanegas JA, Zhu T, Hilton IB, Gao X (2023) Multiplex base-editing enables combinatorial epigenetic regulation for genome mining of fungal natural products. J Am Chem Soc 145(1):413–421. https://doi.org/10.1021/jacs.2c10211
Funding
This work was supported by the Zhejiang Traditional Chinese Medicine Administration [grant numbers 2024ZR060].
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Writing-original draft, visualization and investigation were performed by Chao Feng. Writing-review & editing were performed by Jing Chen and Wenxin Ye. Writing-review & editing and supervision were performed by Zhanshi Wang. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Feng, C., Chen, J., Ye, W. et al. Nitrile hydratase as a promising biocatalyst: recent advances and future prospects. Biotechnol Lett (2024). https://doi.org/10.1007/s10529-024-03530-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10529-024-03530-y