Abstract
Nitrilase is the member of carbon–nitrogen hydrogen hydrolase superfamily, which has been widely used for the hydrolysis of nitriles into corresponding carboxylic acids. But most nitrilases are plagued by product inhibition in the industrial application. In this study, a “super nitrilase mutant” of nitrilase with high activity, thermostability and improved product tolerance from Acidovorax facilis ZJB09122 was characterized. Then, an efficient process was developed by employing the whole cell of recombinant E. coli for the conversion of high concentration of 1-cyanocyclohexylacetonitrile-to-1-cyanocyclohexaneacetic acid, an important intermediate of gabapentin. Under the optimized conditions, the higher substrate concentrations such as 1.3 M, 1.5 M and 1.8 M could be hydrolyzed by 13.58 g DCW/L with outstanding productivity (> 740 g/L/day). This study developed a highly efficient bioprocess for the preparation of 1-cyanocyclohexaneacetic acid which has the great potential for industrial application.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Cyano carboxylic acids, as important building blocks, are widely used in the preparation of various pharmaceuticals and agrochemicals [1,2,3,4,5]. The selective hydrolysis of dinitrile is one of the important approaches for the preparation of cyano carboxy acids. However, the chemical method affords product in low yield and produce large amounts of unwanted byproducts and inorganic wastes. Nitrilase is a kind of important hydrolase [6,7,8,9,10,11,12,13]. Due to the high regioselectivity, nitrilase becomes attractive biocatalyst for the preparation of cyano carboxylic acids by selective hydrolysis of dinitriles in one single step.
Within the past few years, selective hydrolysis of dinitriles to cyano carboxylic acid by nitrilase has been explored and become highly appealing. The nitrilase from Acidovorax facilis 72W was utilized in the synthesis of 4-cyanopentanoic acid [14], an important precursor of 1,5-dimethyl-2-piperidone, and 1-cyanocyclohexaneacetic acid (1-CA), an intermediate of gabapentin [15]. The nitrilase bll6402 from Bradyrhizobium japonicum USDA 110 was reported to hydrolyze α,ω-dinitriles to ω-cyanocarboxylic acids [16, 17]. Plant nitrilases from Arabidopsis thaliana [18], Brassica rapa [19], Arabis alpina [20] were described to hydrolyze racemic isobutylsuccinonitrile to (3S)-3-cyano-5-methyl hexanoic acid which is the key intermediate for the preparation of Pregabalin. In addition, several nitrilases with satisfactory regioselectivity were found in natural isolates such as Rhodococcus rhodochrous and R. sphaeroides or by genome mining [5, 21,22,23,24,25].
In recent research, our group cloned a regioselective nitrilase from A. facilis ZJB09122 [4, 26]. After mutation on F168 and process optimization, the activity of the nitrilase mutant AcN (F168V) was improved greatly [26,27,28,29]. Nevertheless, similar to other nitrilases, the nitrilase AcN shared some common limitation, such as poor thermostability and product tolerance. Very recently, protein engineering was employed to modify the nitrilase AcN, and a “super nitrilase mutant” AcN-M (AcN-T201F/Q339K/Q343K) with dramatically improved thermostability was obtained [30].
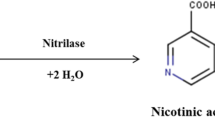
Here, we selected AcN-M as the catalyst for the hydrolysis of 1-cyanocyclohexylacetonitrile (1-CN) and its kinetic parameters were determined. After overexpressing the nitrilase in Escherichia coli (E. coli) BL21 (DE3), the recombinant E. coli was applied to the hydrolysis of 1-CN to 1-CA with high substrate loading (Scheme 1). Under optimized conditions, the conversion rate of 1-CN reached ≥ 99% with a productivity of 740.4 g/L/day, which represents the highest level so far. This work provides an advantage over all the existing biocatalysis process toward 1-CN.
Materials and methods
Chemicals
1-CN, 1-CA and 1-carboxycyclohexaneacetic acid were obtained from Zhejiang Chiral Medicine Chemicals Co., Ltd. (Hangzhou, China). Other chemicals were of analytical grade and purchased from local suppliers.
Strains, culture conditions and protein purification
The AcN and AcN-M were cloned, mutated previously and overexpressed in host E. coli BL21 (DE3) [4, 30]. These recombinant E. coli strains were grown at 37 °C in Luria–Bertani (LB) medium (10 g/L tryptone, 5 g/L yeast extract, 10 g/L NaCl) with the addition of 50 µg/mL kanamycin. The induction of the nitrilase was achieved by adding 0.1 mM of isopropyl β-d-thiogalactoside (IPTG) to the medium when OD600 was 0.6–0.8, and the strain was cultured for another 10–12 h at 28 °C and 150 rpm. The cells were harvested by centrifugation at 10,000×g for 10 min and washed by 0.9% (w/v) NaCl twice.
The resulted cells were resuspended in a 0.2 M sodium phosphate buffer (pH 7.0) and the cell extract was prepared by the ultrasonic cell-break method. The nitrilase AcN and AcN-M were purified by standard Ni–NTA affinity chromatography as previously described [30]. The concentration of purified nitrilase was determined by Bradford protein assay [31].
Kinetic measurement
About 0.25 µg purified nitrilases were assayed in 10 mL sodium phosphate buffer (0.1 M, pH 7.0) containing 0-200 mM 1-CN at 45 °C. The reaction was stopped by adding 100 µL 6.0 M HCl and the concentration of 1-CA was detected using high-performance liquid chromatography (HPLC). The inhibition constant (Ki) was determined using the sodium phosphate buffer (0.1 M, pH 7.0) containing 2–100 mM 1-CA and a series of concentrations of 1-CN (0–200 mM).
Nitrilase assay
The standard reaction was performed at 45 °C by mixing 67.9 mg DCW resting cells with 0.2 M 1-CN in 10 mL sodium phosphate buffer (0.2 M, pH 7.0). The reaction mixture was incubated in an orbital shaker at 200 rpm at 45 °C for 10 min. The aliquot (500 µL) was withdrawn and the reaction was stopped by 5µL 6.0 M HCl and centrifuged at 12,000×g for 3 min. The concentration of 1-CA in the supernatant was determined by HPLC. One unit of the nitrilase activity (U) was defined as the amounts of resting cells that produced 1 µmol of product per min under the standard assay conditions.
Product tolerance
Assay to measure the product tolerance of recombinant E. coli and cell extract was performed in an orbital shaker at 200 rpm at 45 °C. 10 mL reaction mixture contained 67.9 mg DCW cells or cell extract, 0.2 M substrate, and different concentration of 1-CA. The pH was adjusted to 7.0 with 1.0 M NaOH solution. Aliquots (500 µL) were taken after 10 min and the product concentration was determined by HPLC.
Effect of temperature and pH
The influence of temperature on the nitrilase activity of recombinant E. coli was measured in the range of 25–60 °C. To determine the thermal stability of recombinant E. coli, the resting cells were incubated in sodium phosphate buffer (0.2 M, pH 7.0) at 25–50 °C with regular time intervals, and the initial and residual activities were measured under the standard assay conditions.
The pH dependence of recombinant E. coli was studied under the standard assay condition using following buffers at 0.2 M concentration: citric acid–sodium citrate buffer, pH 4.0–5.0; sodium phosphate buffer, pH 6.0–8.0; Gly–NaOH buffer, pH 9.0–10.0. The stabilities of the recombinant E. coli under different pH values were studied by incubating them at pH 7.0 or pH 7.5 in an ice bath. The residual activities were measured under the standard assay conditions.
Effect of 1-CA concentration on cell integrity and cell-specific activity
The effect of 1-CA concentration on resting cells were monitored by incubating 13.58 g DCW/L E. coli BL21 (DE3)/pET-28b(+)-AcN-M cells in sodium phosphate buffer (0.2 M, pH 7.0) with 0.50 M or 0.75 M 1-CA at different temperature (25–45 °C). At regular time intervals, an aliquot (5 mL) was taken and centrifuged at 15,000×g for 1 min. The protein in the supernatant was analyzed by sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS–PAGE) (with 12% polyacrylamide) [32] and Bradford method. The initial and residual nitrilase activity (Ai and Ar) of cells was determined under the standard assay conditions. Half-lives were measured from plots of the natural logs of Ar/Ai versus the incubating time.
Hydrolysis of 1-CN by recombinant E. coli
Recombinant E. coli BL21 (DE3)/pET-28b(+)-AcN-M (1.358 g DCW) was added into a 250 mL flask containing 100 mL sodium phosphate buffer (0.2 M, pH 7.5) and different concentration of 1-CN (1.0–2.0 M). The reaction was carried out at different temperatures (25–45 °C) and 200 rpm. During the hydrolysis reaction, the samples were withdrawn at the required time and analyzed by HPLC for profiling the product formation. The same samples (500 µL) were extracted by 500 µL ethyl acetate and the organic phase was analyzed by gas chromatography (GC) for determining 1-CN residue.
Scaling-up the biotransformation of 1-CN
(1) In a 1-L stirred reactor, 500 mL of distilled water containing 6.79 g DCW of E. coli BL21 (DE3)/pET-28b(+)-AcN-M cells and 111 g of 1-CN were added. The mixture was incubated at 35 °C, 200 rpm for 6 h. (2) In a 1-L stirred reactor 500 mL of distilled water containing 6.79 g DCW of E. coli BL21 (DE3)/pET-28b(+)-AcN-M cells and 133 g of 1-CN were added. The mixture was incubated at 25 °C, 200 rpm for 8 h. Samples (1 mL) were withdrawn at intervals to analyze the concentrations of 1-CA and 1-CN.
Recovery of 1-CA
After the hydrolysis reaction, the cells were removed by centrifugation (12,000×g, 10 min). The supernatant was mixed with 1% (w/v) polyaluminum chloride and agitated at 400 rpm for 4 h. 1% (w/v) diatomite (median size: 19.6 µm) was added and the mixture was agitated at 400 rpm for additional 2 h. After filtration, the filtrate was acidified to a pH of 1.5–2.0 with 6.0 M HCl and mixed with an equal volume of dichloromethane under stirring. The extraction operation was repeated for 2–3 times. The organic layer was combined and distilled under vacuum. The solid obtained was dried at 52 °C.
Analytical methods
1-CA and 1-carboxycyclohexaneacetic acid were separated by HPLC (Shimadzu LC-16 with a UV detector) with a XBridge BEH C18 Column (130 Å, 5 µm, 4.6 mm × 250 mm, 1/pkg, column temperature 40 °C), 76% acetonitrile, 24% 0.58 g/L NH4H2PO4 and 1.83 g/L NaClO4 in water (pH 1.8) as solvent. The UV detector was set at a wavelength of 215 nm.
1-CN was analyzed by GC (Agilent 7890A system with a FID Detector) with Agilent J&W HP-5 Column (30 m × 0.32 mm, 0.25 µm film thickness). The inlet temperature was 320 °C, and the column temperature was 160 °C. The carrier gas was nitrogen and the detector temperature was 340 °C.
Results and discussion
Kinetic constants of mutant nitrilase
In previous work, E. coli BL21 (DE3)/pET-28b(+)-AcN was constructed and shown as a robust catalyst to hydrolyze 1-CN. But, the thermostability is relatively poor. Recently, a nitrilase mutants AcN-M (AcN-T201F/Q339K/Q343K) from AcN was isolated with improvement in thermostability and specific activity toward 1-CN [30]. To analyze the mutational effect, the kinetic parameters of nitrilase AcN and AcN-M were determined (Fig. 1). To our surprise, the mutant AcN-M has similar Km with parent nitrilase AcN, and 1-CA was bound with an affinity (Ki = 8.15 mM) that was less than that exhibited for AcN (Ki = 1.97 mM) by ca. fourfold, which indicated that AcN-M facilitated the product release without perturbing the binding of substrate. Alteration of the quaternary structure impact by the mutations on site 201 which was located on “A” surface (responsible for the dimerization and positioning of active sites) and the top of substrate-binding pocket might be the explanation for the reduction of product inhibition [33, 34].
In addition, when comparing parent and mutant nitrilases in E. coli BL21 (DE3), the recombinant E. coli BL21 (DE3)/pET-28b(+)-AcN-M showed higher specific activity (Table 1). No 1-carboxycyclohexaneacetic acid was detected in the reactions, which indicated there were no changes in their regioselectivity.
Effect of temperature and pH on the cell-specific activity
The activity of whole cells of E. coli BL21 (DE3)/pET-28b(+)-AcN-M was monitored at different temperature (25–60 °C) (Fig. 2a). The result indicated that the resting cells showed maximum activity at 55 °C and the activity declined when the temperature was above 60 °C. To analyze the thermostability, the resting cells were incubated at 25 °C, 35 °C, 45 °C and 50 °C before measuring the residual activity under the standard assay conditions. As shown in Fig. 2b, the mutant displayed high thermostability below 45 °C with a half-life time of about 198.3 h. At 50 °C, the resting cells lost activity more rapidly and only retained 20% of the initial activity after incubating for 45 h.
a Effect of temperature on the cell-specific activity of E. coli BL21 (DE3)/pET-28b(+)-AcN-M (6.79 g DCW/L biocatalyst, 0.2 M 1-CN in pH 7.0 sodium phosphate buffer (0.2 M)). b Thermostability of E. coli BL21 (DE3)/pET-28b(+)-AcN-M at different temperature. c Dependence of nitrilase activity of E. coli BL21(DE3)/pET-28b(+)-AcN-M (6.79 g DCW/L biocatalyst, 0.2 M 1-CN) on pH at 45 °C. d Stability of E. coli BL21 (DE3)/pET-28b(+)-AcN-M at different pH values
The recombinant E. coli maintained higher activities in the pH range from 7.0 to 9.0 (Fig. 2c) and Fig. 2d confirmed the excellent stability at pH 7.0 and pH 7.5. In the following experiments, pH 7.5 was selected as the optimal condition for the reactions.
Product tolerance
Generally, low product tolerance is particularly limiting factor in the enzymatic conversion of nitriles to acids under the high substrate loadings and represents a barrier to achieving the high product concentration necessary for an efficient process [35,36,37,38,39]. For example, Nocardia rhodochrous LL100-21 was inhibited by the product nicotinic acid and 0.3 M nicotinic acid caused 44% inhibition of 3-cyanopyridinase activity [40]. Nicotinic acid at concentrations greater than 0.2 M inhibited the 3-cyanopyridinase activity of nitrilase from the thermophilic bacterium Bacillus pallidus Dac521 [41].
In the nitrilase-mediated hydrolysis of 1-CN, the product tolerance to 1-CA was estimated by comparing the activities of parent and mutant nitrilases in E. coli BL21 (DE3) under inhibiting conditions to that without adding inhibitors. As shown in Fig. 3, the mutant exhibited a higher level of 1-CA resistance than the wild type. Under 0.7 M product, AcN lost ca. 82% of initial activity while AcN-M only lost ca. 45% of initial activity. Even when the concentration of 1-CA raised to 1.3 M and 1.5 M, AcN-M retained 26% and 23% of initial activity, respectively, while AcN was strongly inhibited and almost lost all of the catalytic activity, which proved the product tolerance of this mutation was improved.
Effect of product 1-CA to the activity of on the whole cells of recombinant E. coli. The ability of AcN and AcN-M to hydrolyze 1-CN was determined in the presence of 0–1.5 M 1-CA, and the reactions were carried out by standard assay. The relative activity is shown relative to the activity without adding 1-CA
Dependence of product accumulation on temperature
As described above, the mutant E. coli BL21 (DE3)/pET-28b(+)-AcN-M exhibited higher activity and great thermostability at 45 °C compared to E. coli BL21 (DE3)/pET-28b(+)-AcN (Table 1). At 45 °C, AcN-M (13.58 g DCW/L) exhibited high hydrolysis activity toward 1.0 M 1-CN and the conversion reached more than 99% within 2 h, with productivity 1886 g/L/day. But, when the concentration of 1-CN was higher than 1.3 M, substrates were not hydrolyzed completely at 45 °C. After comparing the hydrolysis reactions at different temperature (25 °C, 35 °C and 45 °C) under high substrate loading (Table 2), it showed that: (1) high temperature resulted in shorter reaction time and higher productivity under low substrate loading due to the higher cell-specific activity. At 45 °C, 13.58 g DCW/L of resting cells demonstrated faster reaction rate and higher productivity than that at low temperature (Entry 1–3). Similarly, at 35 °C, using the same amount of resting cells, almost no substrate was detected after 4 h of conversion with 1.3 M 1-CN in which the productivity reached up to 1187 g/L/day (Entry 5), when the substrate concentration increased to 1.5 M, 0.139 mol 1-CA along with the catalyst yield (gproduct/gcatalyst) of 16.4 and productivity of 974 g/L/day could be obtained (Fig. 6) (Entry 8), which was also faster than that at 25 °C (Entry 4, Entry 7). (2) The low temperature was of benefit to the accumulation of product under high substrate loading. As mentioned above, a high concentration of 1-CN (≥ 1.3 M) could not be completely converted to 1-CA at 45 °C (Entry 6, Entry 9, Entry 12). Although 1.3 M and 1.5 M substrate could be completely consumed at 35 °C, when increasing initial concentration of 1-CN to 1.8 M the conversion decreased to 89.9% (Entry 11). When the reaction run at 25 °C, a reaction time of 8 h resulted in > 99% total conversion of 1.8 M 1-CN, affording a catalyst yield of 18.9 and productivity of 767 g/L/day (Entry 10). In addition, because of high product accumulation and strong product inhibition, 2.0 M 1-CN could not be completely consumed within 12 h (Entry 13–15).
During the reaction, the product 1-CA not only acted as competition inhibitor to nitrilases, but also was hypertoxic to cells which led to cell lysis [28]. High temperature can also lead to the increased permeability of cell. To investigate the synergistic effect of 1-CA and temperature on resting cells, suspension of the resting cells in pH 7.0 sodium phosphate buffer with 0.50 M and 0.75 M 1-CA were incubated at different temperature (25 °C, 35 °C, 45 °C). As shown in Fig. 4a, the cell lysis showed a positive correlation with temperature under 0.50 M 1-CA. Under the combined effect of product and high temperature, the rate of protein accumulation in cell suspension at 45 °C and 0.50 M 1-CA was far faster than that at 35 °C and 25 °C. Similar result was obtained in the SDS–PAGE analysis of the protein in the supernatant which showed more nitrilase leaked at elevated temperature (identified based on the molecular weight) (Fig. 4b). The leakage of protein led to the inactivation of resting cells. In Fig. 5, the calculated inactivation curves and half-life t1/2 were shown. It could be seen that the inactivation of the resting cell exhibited first-order inactivation kinetics, which showed that low temperature led to a significant increase in stability of resting cells under high concentration of 1-CA. On the other hand, the activity of the leaked protein was investigated in presence of 1-CA. The product tolerance of cell extract which was equivalent of 6.79 g DCW/L fresh cells was inferior to that of whole cells, and the concentration of leaked protein which was up to 1200 parts per million (ppm) was far less than that of cell extract (ca. 4000 ppm). After diluting the cell extract of E. coli BL21 (DE3)/pET-28b(+)-AcN-M to ca. 1400 ppm, the loss of activity under 0.4 M product increased to 65%, and almost lost all of activity under 0.70 M 1-CA (data not shown), which suggested that the leak protein could not hydrolyze the substrate under a high concentration of product.
a Effect of product 1-CA and temperature on cell lysis of E. coli BL21 (DE3)/pET-28b(+)-AcN-M. 13.58 g DCW/L resting cells of E. coli BL21 (DE3)/pET-28b(+)-AcN-M were incubated under different temperature and different concentration of 1-CA. The protein concentration was determined by Bradford protein assay. b SDS–PAGE analysis of the protein leaked from free cells incubated under 0.5 M 1-CA at different temperature for 3 h. Lane 1: 25 °C; lane 2: 35 °C; lane 3: 45°C
Therefore, cell integrity was an important factor for hydrolysis reaction under high substrate loading and temperature was an essential means of regulation to balance the cell integrity and reaction rate. A temperature of 45 °C resulted in super high reaction rate and productivity which could be employed in the hydrolysis reaction with low substrate loading (≤ 1.0 M) and short reaction time (≤ 3 h). A temperature of 35 °C was optimal when the concentration of 1-CN was 1.0–1.5 M for a faster reaction rate. Meanwhile, when the hydrolysis reaction runs at higher substrate concentrations (≥ 1.5 M) or need more than 6 h to complete, a reaction temperature of 25 °C was more suitable (Fig. 6).
Scaling-up the hydrolysis of 1-CN
To explore the potential synthetic value of the AcN-M mutant in industry, the whole-cell biocatalyst was used to produce 1-CA in a 1-L stirred bioreactor. To avoid the presence of too many salts that would have to be removed during the purification steps [26], purely aqueous solution at pH 7.0 was used as a good alternative to buffers in the experiments for the preparation of 1-CA. Hydrolysis of 1.5 M and 1.8 M 1-CN by 13.58 g DCW/L whole cells was carried out in purely aqueous solution (initial pH 7.0), respectively. As shown in Fig. 7, the mutant could completely hydrolyze 1.5 M 1-CN in 6 h at 35 °C, providing a productivity of 882.9 g/L/day. The aqueous reaction mixture was extracted with dichloromethane and analyzed by HPLC to reveal an 88% yield of 1-CA. 1.8 M 1-CN could also completely convert to 1-CA at 25 °C with a productivity of 740.4 g/L/day and catalyst yield of 18.9. The yield of 1-CA separated from the reaction mixture was 86%. E. coli BL21 (DE3)/pET-28b(+)-AcN-M was demonstrated to potential in the industrial production of 1-CA.
Scaling-up the regioselective hydrolysis of 1-CN for synthesizing 1-CA in a 1-L reactor containing 0.5 L reaction mixture. The experiments were performed at 35 °C/25 °C and 200 rpm with 1.5 M/1.8 M substrate and 13.58 g DCW/L biocatalyst in purely aqueous solution. aThe productivity was defined as the 1-CA synthesized per volume; per day
Conclusion
In summary, we demonstrated that product 1-CA and temperature were crucial to sustain cell integrity and optimal biocatalyst performance. After optimization of reaction condition, the mutant E. coli BL21 (DE3)/pET-28b(+)-AcN-M was proved to be an exceedingly robust catalyst with high cell-specific activity and improved tolerance to the product. Productivity and catalyst yield have been significantly improved compared to the previous reports. This was an efficient, practical and economical enzymatic process featuring high substrate loading, outstanding catalyst yield and high productivity which has potential in industrial application.
References
Effenberger F, Oßwald S (2001) Selective hydrolysis of aliphatic dinitriles to monocarboxylic acids by a nitrilase from Arabidopsis thaliana. Synthesis 2001(12):1866–1872
Chauhan S, Wu S, Blumerman S, Fallon RD, Gavagan JE, DiCosimo R, Payne MS (2003) Purification, cloning, sequencing and over-expression in Escherichia coli of a regioselective aliphatic nitrilase from Acidovorax facilis 72W. Appl Microbiol Biotechnol 61(2):118–122
DeSantis G, Wong K, Farwell B, Chatman K, Zhu ZL, Tomlinson G, Huang HJ, Tan XQ, Bibbs L, Chen P, Kretz K, Burk MJ (2003) Creation of a productive, highly enantioselective nitrilase through gene site saturation mutagenesis (GSSM). J Am Chem Soc 125(38):11476–11477
Zhang XH, Liu ZQ, Xue YP, Zheng YG (2014) Activity improvement of a regioselective nitrilase from Acidovorax facilis and its application in the production of 1-(cyanocyclohexyl) acetic acid. Process Biochem 49(12):2141–2148
Veselá AB, Rucká L, Kaplan O, Pelantová H, Nešvera J, Pátek M, Martínková L (2015) Bringing nitrilase sequences from databases to life: the search for novel substrate specificities with a focus on dinitriles. Appl Microbiol Biotechnol 100(5):2193–2202
Brady D, Dube N, Petersen R (2006) Green chemistry: highly selective biocatalytic hydrolysis of nitrile compounds. S Afr J Sci 102(7–8):339–344
Singh R, Sharma R, Tewari N, Geetanjali, Rawat DS (2006) Nitrilase and its application as a ‘green’ catalyst. Chem Biodivers 3(12):1279–1287
Martínková L, Kren V (2010) Biotransformations with nitrilases. Curr Opin Chem Biol 14(2):130–137
Martínková L, Rucká L, Nešvera J, Pátek M (2016) Recent advances and challenges in the heterologous production of microbial nitrilases for biocatalytic applications. World J Microbiol Biotechnol 33(1):8
Bhalla TC, Kumar V, Kumar V, Thakur N, Savitri (2018) Nitrile metabolizing enzymes in biocatalysis and biotransformation. Appl Biochem Biotechnol. https://doi.org/10.1007/s12010-018-2705-7
Xue YP, Jiao B, Hua DE, Cheng F, Liu ZQ, Zheng YG (2017) Improving catalytic performance of an arylacetonitrilase by semirational engineering. Bioprocess Biosyst Eng 40(10):1565–1572
Zhang ZJ, Pan J, Li CX, Yu HL, Zheng GW, Ju X, Xu JH (2014) Efficient production of (R)-(−)-mandelic acid using glutaraldehyde cross-linked Escherichia coli cells expressing Alcaligenes sp. nitrilase. Bioprocess Biosyst Eng 37(7):1241–1248
Zhang CS, Zhang ZJ, Li CX, Yu HL, Zheng GW, Xu J-H (2012) Efficient production of (R)-O-chloromandelic acid by deracemization of O-chloromandelonitrile with a new nitrilase mined from Labrenzia aggregata. Appl Microbiol Biotechnol 95(1):91–99
Cooling FB, Fager SK, Fallon RD, Folsom PW, Gallagher FG, Gavagan JE, Hann EC, Herkes FE, Phillips RL, Sigmund A, Wagner LW, Wu W, DiCosimo R (2001) Chemoenzymatic production of 1,5-dimethyl-2-piperidone. J Mol Catal B Enzym 11(4–6):295–306
Burns MP, Wong JW (2004) Biocatalytic preparation of 1-cyanocyclohexaneacetic acid. WO2004111256 A1. https://patents.google.com/patent/WO2004111256A1
Zhu D, Mukherjee C, Biehl ER, Hua L (2007) Nitrilase-catalyzed selective hydrolysis of dinitriles and green access to the cyanocarboxylic acids of pharmaceutical importance. Adv Synth Catal 349(10):1667–1670
Duan YT, Yao PY, Ren J, Han C, Li Q, Yuan J, Feng JH, Wu QQ, Zhu DM (2014) Biocatalytic desymmetrization of 3-substituted glutaronitriles by nitrilases. A convenient chemoenzymatic access to optically active (S)-Pregabalin and (R)-Baclofen. Sci China Chem 57(8):1164–1171
Xie ZY, Feng JL, Garcia E, Bernett M, Yazbeck D, Tao JH (2006) Cloning and optimization of a nitrilase for the synthesis of (3S)-3-cyano-5-methyl hexanoic acid. J Mol Catal B Enzym 41(3–4):75–80
Ishikawa T, Okazaki K, Kuroda H, Itoh K, Mitsui T, Hori H (2007) Molecular cloning of Brassica rapa nitrilases and their expression during clubroot development. Mol Plant Pathol 8(5):623–637
Zheng RC, Zheng YG, Zhang Q, Huang YM, Li Y, Weng J, Liu T, Fan W (2015) Nitrilase from Arabis alpina, its encoding gene, vector, recombinant bacterial strain and uses thereof. US20170355976 A1. https://patents.google.com/patent/US20170355976A1/
Kobayashi M, Nagasawa T, Yamada H (1988) Regiospecific hydrolysis of dinitrile compounds by nitrilase from Rhodococcus rhodochrous J1. Appl Microbiol Biotechnol 29(2–3):231–233
Bengis-Garber C, Gutman AL (1989) Selective hydrolysis of dinitriles into cyano-carboxylic acids by Rhodococcus rhodochrous NCIB 11216. Appl Microbiol Biotechnol 32(1):11–16
Dadd MR, Claridge TDW, Walton R, Pettman AJ, Knowles CJ (2001) Regioselective biotransformation of the dinitrile compounds 2-, 3- and 4-(cyanomethyl) benzonitrile by the soil bacterium Rhodococcus rhodochrous LL100-21. Enzyme Microb Technol 29(1):20–27
Rey P, Rossi JC, Taillades J, Gros G, Nore O (2004) Hydrolysis of nitriles using an immobilized nitrilase: applications to the synthesis of methionine hydroxy analogue derivatives. J Agric Food Chem 52(26):8155–8162
Wang HL, Li GN, Li MY, Wei DZ, Wang XD (2014) A novel nitrilase from Rhodobacter sphaeroides LHS-305: cloning, heterologous expression and biochemical characterization. World J Microbiol Biotechnol 30(1):245–252
Xue YP, Wang YP, Xu Z, Liu ZQ, Shu XR, Jia DX, Zheng YG, Shen YC (2015) Chemoenzymatic synthesis of gabapentin by combining nitrilase-mediated hydrolysis with hydrogenation over Raney-nickel. Catal Commun 66:121–125
Xue YP, Zhong HJ, Zou SP, Zheng YG (2017) Efficient chemoenzymatic synthesis of gabapentin by control of immobilized biocatalyst activity in a stirred bioreactor. Biochem Eng J 125:190–195
Zou SP, Huang JW, Xue YP, Zheng YG (2018) Highly efficient production of 1-cyanocyclohexaneacetic acid by cross-linked cell aggregates (CLCAs) of recombinant E. coli harboring nitrilase gene. Process Biochem 65:93–99
Xue YP, Shu XR, Zou SP, Wang YJ, Zheng YG (2015) Efficient recovery of 1-cyanocyclohexaneacetic acid by ion-exchange process. Sep Sci Technol 50(17):2717–2725
Xu Z, Cai T, Xiong N, Zou SP, Xue YP, Zheng YG (2018) Engineering the residues on “A” surface and C-terminal region to improve thermostability of nitrilase. Enzyme Microb Technol 113:52–58
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein–dye binding. Anal Biochem 72:248–254
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680
Sewell BT, Berman MN, Meyers PR, Jandhyala D, Benedik MJ (2003) The cyanide degrading nitrilase from Pseudomonas stutzeri AK61 is a two-fold symmetric, 14-subunit spiral. Structure 11(11):1413–1422
Wu S, Fogiel AJ, Petrillo KL, Jackson RE, Parker KN, Dicosimo R, Ben-Bassat A, O’Keefe DP, Payne MS (2008) Protein engineering of nitrilase for chemoenzymatic production of glycolic acid. Biotechnol Bioeng 99(3):717–720
Xue YP, Liu ZQ, Xu M, Wang YJ, Zheng YG, Shen YC (2010) Enhanced biotransformation of (R,S)-mandelonitrile to (R)-(−)-mandelic acid with in situ production removal by addition of resin. Biochem Eng J 53(1):143–149
Yang C, Wang X, Wei D (2011) A new nitrilase-producing strain named Rhodobacter sphaeroides LHS-305: biocatalytic characterization and substrate specificity. Appl Biochem Biotechnol 165(7):1556–1567
Sharma NN, Sharma M, Bhalla TC (2011) An improved nitrilase-mediated bioprocess for synthesis of nicotinic acid from 3-cyanopyridine with hyperinduced Nocardia globerula NHB-2. J Ind Microbiol Biotechnol 38(9):1235–1243
Xue YP, Xu M, Chen HS, Liu ZQ, Wang YJ, Zheng YG (2013) A novel integrated bioprocess for efficient production of (R)-(–)-mandelic acid with immobilized Alcaligenes faecalis ZJUTB10. Org Process Res Dev 17(2):213–220
Zhu XY, Gong JS, Li H, Lu ZM, Shi JS, Xu ZH (2014) Bench-scale biosynthesis of isonicotinic acid from 4-cyanopyridine by Pseudomonas putida. Chem Pap 68(6):739–744
Vaughan PA, Knowles CJ, Cheetham PSJ (1989) Conversion of 3-cyanopyridine to nicotinic acid by Nocardia rhodochrous LL100-21. Enzyme Microb Technol 11(12):815–823
Almatawah QA, Cowan DA (1999) Thermostable nitrilase catalysed production of nicotinic acid from 3-cyanopyridine. Enzyme Microb Technol 25(8):718–724
Acknowledgements
This work was funded by the National Natural Science Foundation of China (nos. 21476210 and 21706235).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Xu, Z., Xiong, N., Zou, SP. et al. Highly efficient conversion of 1-cyanocycloalkaneacetonitrile using a “super nitrilase mutant”. Bioprocess Biosyst Eng 42, 455–463 (2019). https://doi.org/10.1007/s00449-018-2049-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00449-018-2049-2