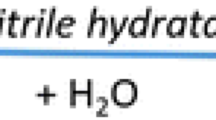Abstract
Nitrile metabolizing enzymes, i.e., aldoxime dehydratase, hydroxynitrile lyase, nitrilase, nitrile hydratase, and amidase, are the key catalysts in carbon nitrogen triple bond anabolism and catabolism. Over the past several years, these enzymes have drawn considerable attention as prominent biocatalysts in academia and industries because of their wide applications. Research on various aspects of these biocatalysts, i.e., sources, screening, function, purification, molecular cloning, structure, and mechanisms, has been conducted, and bioprocesses at various scales have been designed for the synthesis of myriads of useful compounds. This review is focused on the potential of nitrile metabolizing enzymes in the production of commercially important fine chemicals such as nitriles, carboxylic acids, and amides. A number of opportunities and challenges of nitrile metabolizing enzymes in bioprocess development for the production of bulk and fine chemicals are discussed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Nitriles are organic compounds having a carbon triple bonded to nitrogen (–C≡N) as functional group. They are widespread in the environment due to their diverse role as metabolites in large number of biological systems and industrial uses. Nitriles are produced under biotic and abiotic stress conditions in many of the biological systems, i.e., some plants, microbes, insects, and arthropods in the form of glycosides and cyanolipids, and play key role in plant microbial interaction [1]. Industrial use of nitriles as starting materials for synthesis or as reagents in several chemical processes has led to their accumulation in ecosystem [2]. Although most nitriles are highly toxic and carcinogenic due to their cyano group, yet they constitute important intermediates in the production of polyesters, polyamides, carboxylic acids, pharmaceuticals, agrochemicals, dyes, pigments, and fine chemicals [3, 4]. Enzymatic hydrolysis of these compounds is a well-known and accepted method to synthesize a range of useful amides and carboxylic acids [3]. The use of nitrile metabolizing microorganisms/enzymes for bioremediation of soil, water, and air contaminated with highly toxic nitriles/amides is also gaining importance.
Nitrile synthesis in biological systems, i.e., microbes and plants, follows two distinct pathways: (1) aldoxime dehydratase catalyzes formation of a carbon nitrogen triple bond via dehydration of aldoxime [R–CH=N–OH] to corresponding nitrile (R–C≡N) and (2) hydroxynitrile lyase or oxynitrilase-mediated transformation of aldehyde (R–CH=O) and hydrogen cyanide (H–C≡N) to cyanohydrins (R–CHOHC≡N). Cyanohydrins are immediate precursors of cyanoglycosides and cyanolipids formed in various life forms. Nitrile catabolism on the other hand also comprises two distinct pathways: (1) nitrilase-mediated conversion of nitriles (R–C≡N) to corresponding carboxylic acids (R–COOH) and ammonia (NH3) and (2) bienzymatic cascade involving nitrile hydratase and amidase, where the former catalyzes the formation of amides (R–CONH2) from nitriles and the latter subsequently converts amides to carboxylic acids and ammonia (Fig. 1).
Enzymes in nitrile metabolism: Aldoxime dehydratase catalyzes formation of a –C≡N by dehydration of aldoximes (R–CH=N–OH); formation of aldoximes from amino acids is a single or multi-step reaction represented by double arrows; hydroxynitrile lyase or oxynitrilase transforms aldehyde (R–CH=O) and hydrogen cyanide (H–C≡N) to cyanohydrins (R–CHOHC≡N); nitrilase converts nitriles (R–C≡N) to corresponding carboxylic acids (R–COOH) and ammonia (NH3); nitrile hydratase catalyzes the formation of amides (R–CONH2) from nitriles and amidase hydrolyze amides to form carboxylic acids and ammonia. Enzymatic steps involved in nitrile synthesis are represented by violet arrows, whereas its degradation steps are in red arrows
Chemical hydration, oxidation, and hydrolysis are frequently applied in both academia and industry to produce nitriles, amides, carboxylic acids, and hydroxamic acid. Unfortunately, these chemical processes are restricted to processing/production of structurally simple compounds containing no labile groups and often require harsh conditions, such as using strong acids or bases at high temperature/pressure or metal catalyst, give poor selectivity, involve multi-step reaction, and accompanied by the formation of side products [5]. On the other hand, nitrile metabolizing enzymes are capable of synthesizing various nitriles and hydrolyzing a wide range of complex nitriles and amides. These biocatalysts offer much more competitive processes compared to chemical catalysts in terms of productivity, purity, enantioselectivity, and environmental concern [4, 5]. Nitrile metabolizing enzymes have attracted the attention of the scientific community due to their immense potential to be used as industrial biocatalysts [4,5,6].
The history of nitrile metabolizing enzymes goes back to 1964, when Thimann and Mahadevan (1964) reported an enzyme from barley leaves, which catalyzed the conversion of indoleacetonitrile to indoleacetic acid. Nitrile hydratase was first discovered from the bacterium Arthrobacter sp. J1 [7], which was later identified as Rhodococcus rhodochrous J1 [8]. Aldoxime dehydratase was added to the nitrile pathways in the year 2000 [9]. Hydroxy nitrile lyase activity of almond was known since eighteenth century, but its ability to synthesize nitrile was discovered later.
Till date, around 40 microorganisms have been isolated and characterized to possess nitrilase, more than 60 microbes have been reported to have nitrile hydratase, and around hundred showed amidase activity. Aldoxime dehydratase has been reported from many bacterial and fungal genera [9]; however, its characterization is restricted to Pseudomonas chlororaphis B23 [10], Rhodococcus sp. N-771 [11], and Bacillus sp. OxB-1 [12]. Hydroxynitrile lyases were reported in 3000 species of vascular plants, ferns, and gymnosperms and well characterized in Prunus, Malus, and Sorghum. In addition to plants, these enzymes are also reported in bacteria (Chromobacterium violaceum, few species of Pseudomonas), fungi, lichens, millipedes, arthropods, and some insects [13,14,15].
The production of various acids, i.e., glycolic acid, mandelic acid, nicotinic acid, and amides, i.e., acrylamide and nicotinamide, is being carried out at industrial scale using nitrilase and nitrile hydratase, respectively. The synthesis of a number of other compounds using nitrile metabolizing enzymes is on the way from academia to industry. Amidases are employed in combination with nitrile hydratase for the production of commercially important organic acids, i.e., acrylic acid, nicotinic acid, etc. [16]. They are also utilized as industrial catalysts in effluent treatment [17], and their acyl transferase activity is exploited for the synthesis of hydroxamic acids [18,19,20]. The uses of nitrile synthesizing enzymes, i.e., aldoxime dehydratase and hydroxynitrile lyase for large-scale synthesis of nitriles, still remain a challenge. However, hydroxynitrile lyase has been used in combination with nitrilase for asymmetric synthesis of S-mandelic acid [21]. Recently, there are some studies on the synthesis of chiral cyanohydrins using hydroxynitrile lyase [22, 23] and enantioselective dehydration of racemic aldoximes to form corresponding α-branched nitriles [24, 25].
Information on basic and applied aspects of nitrile metabolizing enzymes have immensely expanded in the last several decades, and very informative and critical reviews on nitrilases [3, 4, 26,27,28,29,30], nitrile hydratases [31, 32], amidases [16], and hydroxy nitrile lyase [13,14,15] have appeared in literature. Most of these reviews focused on microbiological, biochemical, enzymological, and molecular aspects of these enzymes. This review focuses on status, challenges, and limitations of nitrile metabolizing enzymes in industrial biocatalysis and process development for synthesis of important compounds.
Nitrile Synthesizing Enzymes
Aldoxime Dehydratase
Aldoxime dehydratases (4.9.9.1) are heme containing enzymes that catalyze the formation of nitriles via dehydration of aldoxime and creation of a carbon nitrogen triple bond. Some nitriles are industrially very valuable as they are used in the production of nylon, acrylic fibers, insecticides, pharmaceuticals, and also used in organic synthesis. Although, nitrile are produced by chemical dehydration of aldoxime, but it requires several harsh conditions. Therefore, an environmentally benign process of biological dehydration by aldoximes dehydratase is a possible alternate. In nature, some microbes and plants have an aldoxime-nitrile pathway leading to the synthesis of cyanogenic glycosides and other intermediates used in energy metabolism [33, 34]. Aldoxime dehydratase is often present in cluster with other enzymes of this pathway, i.e., nitrile hydrates, amidase, acyl-CoA synthetase, etc. [34, 35] leading to the synthesis of nitriles followed by its hydrolysis. Kato et al. (2000) have explored the distribution of these enzymes in 45 genera of bacteria, 11 genera of actinomyces, 22 genera of yeasts, and 37 genera of fungi [9]. Later on, Asano’s and Kobayashi’s group have reported the presence of gene clusters of aldoxime-nitrile pathway in many microbes [10, 34,35,36]. Molecular mechanism of this heme containing enzyme has been proposed [11, 37], and genome sequence of aldoxime degrading bacterium Bacillus sp. OxB-1 has also been decoded [12]. Despite these studies, the potential of aldoxime dehydratases in nitrile synthesis has not been explored industrially. However, some efforts have been made to study and characterize the biological dehydration ability of aldoxime dehydratases (Table 1). Synthesis of phenylacetic acid from (Z)-phenylacetaldoxime via phenylacetonitrile by a combination of aldoxime dehydratase from Bacillus sp. OxB-1 and nitrilase was studied [9]. Recently, a recombinant E. coli whole-cell catalyst overexpressing the aldoxime dehydratase from Bacillus sp. OxB-1 has been used directly for enantioselective dehydration of racemic aldoximes, i.e., alkyl aryl substituted noncyclic aldoxime, aliphatic cyclic aldoxime, and heterocyclic aldoxime to form corresponding α-branched nitriles with high enantiomeric excess [24]. These investigators have proposed the use of easily accessible aldehydes as starting materials for the synthesis of aldoximes which were further transformed to chiral α- and ß-branched nitriles by a cyanide-free enantioselective approach employing aldoxime dehydratase. Their strategy is based upon a biocatalytic dehydration of racemic aldoximes with high conversion and excellent enantioselectivity [24]. This is indeed a very novel route for the synthesis of important nitriles and can be integrated with nitrilase or nitrile hydratase for the production of corresponding acids or amides. In another study by Miki and Asano (2014), the biosynthetic pathway for the production of phenylacetonitrile was constructed in E. coli utilizing enzymes from the plant glucosinolate biosynthetic pathway and bacterial aldoxime dehydratase. First step in this biosynthetic route is to produce phenylacetaldoxime from phenylalanine using recombinant E. coli expressing genes encoding for cytochrome P450 and CYP reductase from Arabidopsis thaliana. Second step is the production of phenyl acetonitrile by introducing the aldoxime dehydratase gene from Bacillus sp. OxB-1 [25]. This biocatalytic route, however, have low yield as it produced 4.9 mM phenylacetonitrile, but provided a platform for future research on exploring and improving such integrated strategies. It will be worthwhile to search or develop aldoxime dehydratases with desired specificity, activity, and selectivity for the synthesis of aldoximes through conventional isolation and screening or genome mining or directed evolution approaches. In this direction, Yamaguchi et al. (2016) discovered a novel cytochrome P71AT96 from Fallopia sachalinensis which catalyzes the conversion of aldoximes to nitriles under mild conditions [42]. Aldoxime dehydratases thus have the potential to emerge as key biocatalysts for the production of valuable nitriles. Further, the integration of this enzyme with other biocatalysts in aldoxime nitrile pathway will lead to synthesize desired acids or amides of industrial importance.
Hydroxynitrile Lyase
Hydroxynitrile lyases (E.C.4.2.1) are group of enzymes, which catalyze cleavage and synthesis of cyanohydrins and play a significant role in plant microbial interactions [13, 43]. In nature, these enzymes are found in plants [15], insects, arthropods [44], and microbes [45, 46]. These are used for the biosynthesis of various cyanoglycosides, cyanolipids, and also for their breakdown to release cyanide [15, 43, 44]. In chemical industries, hydroxynitrile lyase is used as biocatalyst for the synthesis of chiral cyanohydrins by exploiting the reversible enzymatic reaction (Table 1). Cyanohydrins are biologically active compounds used in the synthesis of various pharmaceutically and agrochemically important amino alcohols, hydroxy ketones, and hydroxy acids. Few extensive reviews on hydroxynitrile lyase sources, reaction, and biochemical and molecular properties have already been published [13, 15, 47]. Lanfranchi et al. (2013) reviewed the applications of hydroxynitrile lyases in the industry for the synthesis of some valuable nitriles. They have reviewed the hydroxynitrile lyase-mediated synthesis of (R)-2-Cl-mandelonitrile and 3-pyridinecarbaldehyde cyanohydrins and also discussed the use of this biocatalyst for the synthesis of chiral key nitrile intermediates which are used in the production of vitamin B5 and stagonolide-B. These enzymes have been used in chemo-enzymatic synthesis of some novel compounds, i.e., venlafaxine hydrochloride, stagonolide-B, and bienzymatic cascade for the production of (S)-atrolactic acid and (S)-mandelic acid [14]. Among these reactions, the asymmetric synthesis of (R)-mandelonitrile and (S)-mandelonitrile has been extensively explored. Mateoa et al. (2006) used immobilized (S)-selective HNL from Manihot esculenta and nitrilase from Pseudomonas fluorescens EBC 191 for the synthesis of enantiomerically pure (S)-mandelic acid by sequential HCN addition to benzaldehyde and then hydrolysis via nitrilase [38]. (R)-mandelonitrile was synthesized from 250 mM benzaldehyde and 900 mM acetone cyanohydrin in a biphasic system employing the HNL of Passiflora edulis resulting in 31.6% conversion and 98.6% enantiomeric excess [39]. Asif and Bhalla (2016) describe the HNL from wild apricot and were able to synthesize 8.88 mmol (1.184 g) of (R)-mandelonitrile with 89% molar conversion and 96% enantiomeric excess from benzaldehyde and cyanide [40]. This biocatalytic route of (R)-mandelonitrile synthesis is now well known and established; however, the scale-up studies still remain a challenge. An interesting study by Fuhshuku and Asano (2011) described an R-selective HNL from the noncyanogenic plant Arabidopsis thaliana which accepts nitromethane as a donor in a reaction with aromatic aldehydes to yield (R)-ß-nitro alcohols in an aqueous–organic biphasic system [41]. It has thus widened the scope of hydroxynitrile lyase in industrial synthesis. Synthesis of various chiral cyanohydrins was reported in a monophasic microaqueous reaction system using whole cells of recombinant E. coli expressing HNL of Arabidopsis thaliana [22]. Alagoz et al. reported enantioselective transformations of various aldehydes, i.e., benzaldehyde, 4-methoxybenzaldehyde, 4-methyl benzaldehyde, and 4-hydroxybenzaldehyde to corresponding cyanohydrins using immobilized preparations of HNL from Prunus dulcis. Their results showed that immobilized HNL is a powerful and cheap biocatalyst in the synthesis of (R)-mandelonitrile and can be used in combination with nitrilases to produce enantiopure mandelic acids [23]. The database mining approach has resulted in the discovery of novel HNL from Acidobacterium capsulatum ATCC 51196 which catalyzes the (R)-selective synthesis of mandelonitrile with significantly better conversion (97%) and enantioselectivity (96.7%) than other HNLs [46]. Dadashipour et al. (2015) have discovered a novel hydroxynitrile lyase from an invasive millipede, Chamberlinius hualienensis, and characterized its biocatalytic potential for the synthesis of a number of cyanohydrins from benzaldehyde and its substitutives [48]. Microbial, plant, and animal diversity needs to be explored for novel sources of HNL vis-a-vis novel HNLs. Academia and industry need to collaborate for scale-up HNL-mediated transformation reactions for large-scale synthesis of desired cyanohydrin or their derivatives.
Nitrile Degrading Enzymes
Nitrilase
Nitrilase (3.5.5.1) hydrolyzes carbon nitrogen triple bond of nitriles to form corresponding acid and liberate ammonia [26]. These enzymes play important role in nitrogen recycling and detoxification of cyanide, a defense molecule produced from cynogenic glycosides in many life forms. The high substrate specificity, enantioselectivity, and regio-selectivity make nitrilases attractive biocatalysts for the production of fine chemicals and pharmaceutical intermediates [3, 4, 29]. These are also used in the treatment of nitrile containing industrial effluent and remediation of contaminated soil [28]. Nitrilase-mediated biotransformation of various nitriles has been extensively studied and critically reviewed [4, 26, 28, 29]. A number of important compounds, i.e., nicotinic acid [49,50,51], isonicotinic acid [52, 53], mandelic acid [54,55,56,57], glycolic acid [58, 59], benzoic acid [60], hydroxybenzoic acid [61, 62], etc., have been synthesized from nitriles at laboratory scale using free, immobilized, or recombinant cells. Nitrilase-catalyzed transformations of some important compounds are discussed below and summarized in Tables 2 and 3.
Important Aromatic and Aliphatic Carboxylic Acids
A number of industrially important aromatic carboxylic acids, i.e., nicotinic acid, isonicotinic acid, benzoic acid, p-hydroxybenzoic acid, etc., have been synthesized using nitrilase as biocatalyst [49,50,51,52,53, 60,61,62]. Recently, an efficient biocatalytic process for the production of nicotinic acid with volumetric productivity of 24.6 g L−1 h−1 and 100% conversion of 1 M 3-cyanopyridine to nicotinic acid was developed using recombinant E. coli JM109 cells harboring the nitrilase gene from Alcaligenes faecalis MTCC 126 [75].
Among aliphatic and aryl aliphatic acids, acrylic acid, glycolic acid, 3-hydroxyvaleric acid, and mandelic acid have valuable applications and nitrilase-based process for their synthesis has been developed. Acrylic acid has applications in superabsorbent, adhesive, surface coating, etc. with a huge global market. Acrylic acid has been produced from acrylonitrile using nitrilase from several microorganisms [70, 71, 77]. Glycolic acid is another important acid having applications in medicine and pharmaceuticals. In the last decade, enzymatic transformation of glycolonitrile to glycolic acid has been explored extensively [58, 59, 67]. Major obstacles encountered in the synthesis of these aromatic and aliphatic acids are inhibition by position-specific substitution, substrate, and product. At higher concentration of substrate, nitrilase is inhibited; therefore, reaction is carried out in fed-batch mode [49,50,51,52, 57, 69, 81]. However, after few feeding, the accumulated product inhibits the enzyme reaction. Therefore, a high substrate/product-tolerant biocatalyst or continuous type of bioreactor is needed for economically viable synthesis of acids from nitriles at industrial scale.
Synthesis of Enatiopure Carboxylic Acids
Nitrilase is the biocatalyst of choice in asymmetric synthesis of numerous chiral compounds with high enantioselectivity. Various entiomerically pure intermediates synthesized by nitrilase includes (R)-(−)-mandelic acid, (R)-4-cyano-3-hydroxybutyric acid, (R)-o-chloromandelic acid, (R)-acetylmandelic acid, (S)-(+)-ibuprofen, etc. (R)-(−)-mandelic acid is used as chiral intermediate for the synthesis of various pharmaceutical compounds, i.e., penicillin, cephalosporin, antiobesity and antitumor agents, and agrochemicals. In the last several years, the enantioselective nitrilase-mediated route for the synthesis of (R)-(−)-mandelic acid from recemic mandelonitrile has been extensively explored [54,55,56,57, 68]. Various bacteria have been employed for the production of mandelic acid, but most of them suffer from low yields, i.e., P. putida MTCC 5110 (0.39 g g−1dcw), A. faecalis ECU0401 (3.8 g g−1dcw), and Alcaligenes sp. MTCC 10675 (3.9 g g−1dcw). Recently, some novel strategies have been designed for enhanced production of mandelic acid by enzyme immobilization, reactor designing, and medium and enzyme engineering [54,55,56,57]. Zhang et al. (2011) used immobilized nitrilase in five batches in a 2-L stirred reactor and employed the toluene–water biphasic system with a productivity of 13.8 g g−1dcw and 98.0% ee of R-(−)-mandelic acid [68]. Later, Ni et al. (2013) used nanoparticle-immobilized recombinant E. coli and ethyl acetate–water biphasic system in stirred tank reactor and achieved a high productivity of 14.9 g g−1 wet weight and hydrolyzed 1 M mandelonitrile with a final yield of 99% and 95% ee of R-(−)-mandelic acid [56]. An integrated bioprocess using immobilized cells of Alcaligenes faecalis ZJUTB10 in packed bed bioreactor coupled with anion exchange column containing resin HZ202 led to a high productivity of 8.87 mM h−1 after 16 h of reaction (550 mmol) and > 99% ee [55]. Liu et al. generated a mutant of A. faecalis using gene site saturation mutagenesis which resulted in 21.50-fold higher space–time productivity than wild-type nitrilase [82].
Among other enantioselective syntheses, (R)-4-cyano-3-hydroxybutyric acid and (R)-o-chloromandelic acid are prominent examples. (R)-4-cyano-3-hydroxybutyric acid is an intermediate for the synthesis of Lipitor, a cholesterol-lowering drug. This compound has been synthesized with 100% conversion of substrate and 99% ee of product in three steps using chemo-enzymatic strategy. Nitrilase has been also used in the hydrolysis of 3-hydroxyglutaronitrile to (R)-4-cyano-3-hydroxybutyric acid [63].
(R)-o-chloromandelic acid is the precursor for drug Clopidogrel®, a platelet aggregation inhibitor. This acid has been synthesized by the hydrolysis of o-chloromandelonitrile using a novel nitrilase from Labrenzia aggregata in toluene–water (1:9, v/v) biphasic system [66] with 96.3% ee. In an extended fed-batch reaction mode, o-chloromandelonitrile was continuously fed into the reaction containing ethanol as co-solvent (20%, v/v) and product was produced with 97.6% ee [81].
Nitrilases have immense industrial potential as evident from the abovementioned examples. Further biocatalyst improvement, medium engineering accompanied with process design will widen the scope of this biocatalyst for the synthesis of range of industrially and pharmaceutically important molecules. Chemo-enzymatic processes and biphasic system for downstream processing have also emerged as an effective and viable option in the synthesis of a number of fine and commodity chemicals.
Nitrile Hydratase
Nitrile hydratase (NHase, EC 4.2.1.84) is a key enzyme of nitrile metabolism that catalyzes the hydration reaction of nitriles into corresponding amides [32, 83]. This enzyme has been successfully used in chemical industry for the production of useful amides, i.e., acrylamide, nicotinamide, and butyramide [29]. Most of the NHases have been reported earlier but with limited substrate range, low enantioselectivity, and thermo-lability. Efforts are needed to develop robust biocatalysts for synthesis of amides. Chemical synthesis of amides is not environmental friendly; therefore, enzymatic hydration of nitrile to corresponding amide using NHase is an attractive and viable alternative. Among the NHase-producing microorganisms reported so far, only few strains of Rhodococcus and Nocardia have been extensively studied and widely used for the synthesis of amides from nitriles. In the last several years, the thrust of research on nitrile metabolizing enzymes has shifted to enantioselective catalysis of nitriles to optically active/enantiopure amides including amino amides, hydroxy-amides, α-arylaliphatic amides, and amide derivatives [5, 29, 32]. Some of the important amides synthesized beyond test tube scale using NHase are listed in Table 4.
Acrylamide and Nicotinamide
Acrylamide and its polymers are used as coagulators, stock additives in paper, leather and textile industry, flocculants, and chemical industry. Acrylamide has been synthesized from acrylonitrile using free as well as immobilized nitrile hydratase. Kim and Hyun (2002) used immobilized cells of Rhodococcus rhodochrous M33, having nitrile hydratase activity and synthesized acrylamide at 400 g L−1 with 100% conversion of substrate [84]. Raj et al. (2008) synthesized acrylamide using free and immobilized cells of R. rhodochrous PA-34. A 1 L scale bioconversion reaction of acrylonitrile to acrylamide was carried out by them which resulted in 600 g of acrylamide formation in 12 h at 10 °C. The immobilized cells completely converted 8% acrylonitrile in 3 h at 10 °C, in a partitioned fed-batch reactor, and a total 432 g L−1 acrylamide was accumulated after 1 day. There is no doubt a decrease in productivity and accumulation of acrylamide in the reaction after immobilization, but the main advantage of the process was the reusability of biocatalyst. The immobilized biocatalyst was recycled three times, and total production of 1217 g acrylamide was achieved [85]. The acrylamide produced by the immobilized cells exhibited better qualities than that produced by the free cells in terms of color, turbidity, salt content, and foam formation. Acrylamide produced by the immobilized cells contained a lower amount of proteins, salts, and other impurities.
Nicotinamide is one of the important forms of vitamin B3, used in pellagra treatment and also for animal feed supplementation. Mathew et al. (1988) employed R. rhodochrous J1 for nicotinamide production with full conversion of 3-cyanopyridine with a yield of 1465 g L−1 in 9 h [65]. Prasad et al. produced nicotinamide using free cells of R. rhodochrous PA34 in 1 L batch reaction producing 855 g nicotinamide with a productivity of 7.92 g g−1dcw h−1. Mutant of R. rhodochrous PA34 was generated by chemical mutagenesis with increased activity and improved tolerance towards substrate and product. This mutant was used in a batch reaction of 1 L scale to convert 7 M 3-cyanopyridine into nicotinamide in 3 h at 55 °C using 7 g resting cells [86]. The productivity with mutant was 40.7 g g−1dcw h−1 which was more than five times to the wild strain.
Other Important Amides
Nitrile hydratases have been explored for the conversion of heterocyclic nitriles into corresponding amides in batch mode reaction [88]. Using nitrile hydratase activity of R. rhodochrous Jl free cells, 489 g benzamide, 306 g of 2,6-difluorobenzamide, 210 g of 2-thiophenecarboxamide, 522 g of 2-furanecarboxamide, and 1045 g of 3-indoleacetamide per liter of reaction mixture were synthesized at 25 °C, with a conversion yield of 100% [88]. Butyramide is used in the synthesis of hydroxamic acids and electro-rheological fluids, and its N-substituted derivatives are used as analgesic, anticonvulsant, cardiovascular, and anti-inflammatory agent. The nitrile hydratase of R. rhodochrous PA-34 catalyzed the conversion of butyronitrile to butyramide at pH 7 and 10 °C. A yield of 597 g of butyramide (6.8 M) was obtained using 60% (v/v) butyronitrile, in a 1 L batch reaction [89].
Regio-selective nitrile hydratase of Pseudomonas chlororaphis B23 was used for the production of 5-cyanovaleramide (5-CVAM) which is an intermediate of herbicide azafenidin [90]. A chemo-enzymatic process for the preparation of levetiracetam, a drug used for the treatment of epilepsy, using nitrile hydratase has been reported [91]. A recent report has demonstrated the chemo-selective synthesis of 2,6-difluorobenzamide, which is an intermediate of pesticides [94].
A multi-enzymatic approach for the synthesis of optically pure (R)-phenylalanine from (R, S)-2-aminophenylpropionitrile via (R, S)-phenylalaninamide in one step by recombinant E. coli encoding NHase as well co-expressing amino acid amidase and mutant ACL racemase [92]. Various other α-aminonitriles were converted to (S)-α-amino acid by dynamic kinetic resolution using NHase, ACL racemase, and amino acid amidase [93]. These natural and unnatural chiral α-amino acids are used extensively in pharmaceuticals, animal feeds, and artificial sweeteners. Recently, a stereo-selective production of L-phenylglycine by using immobilized nitrile hydratase and amidase has been reported [95].
Nitrile hydratases are very good catalysts for the hydration of nitriles. Since chemical hydration usually requires harsh reaction conditions, the enzymatic hydration reduces by-product formation. Nitrile hydratases in association with amidases have great potential for the synthesis of speciality acids and treatment of wastewater containing toxic nitriles.
Amidases
Amidases (E.C. 3.5.1.4) are ubiquitous enzymes and have very wide biological functions. They possess amide hydrolytic as well as acyl transferase activity and exhibit a range of specificity, selectivity, and affinity towards different substrates. These enzymes have thus turned out to be attractive biocatalyst for organic synthesis, i.e., carboxylic acids, hydroxamic acids, hydrazides, and also for bioremediation [16]. Their hydrolytic activities are employed in combination with nitrile hydratase for the production of commercially important carboxylic acids [69, 96] and amide containing effluent treatment [17], whereas acyltransferase activity is used for the synthesis of hydroxamic acid [18, 97,98,99]. The applications of amidases in the production of various fine chemicals have been reviewed extensively [16]. Recently, acyl transferase activities of amidases were exploited mainly for the synthesis of pharmaceutically active hydroxamic acids and hydrazides. A number of important hydroxamic acids, i.e., acetohydroxamic acid [98, 100], benzohydroxamic acid [20], and nicotinyl hydroxamate [97, 99], have been synthesized using acyltransferase activity of amidases (Table 5). Hydroxamic acids (R-CONHOH) are industrially very important as they form chelates with metal ions. These chelates find applications as growth factors, food additives, antibiotics, antifungal agents, tumor inhibitors, siderophores, enzyme inhibitors, and antileukemic agents. Fournand et al. (1997) were the first to report the synthesis of hydroxamic acid, i.e., acetohydroxamic acid at 1 L scale by using purified amidase of Rhodococcus sp. R312 immobilized on Duolite A-378 resin with 61% molar conversion of acetamide [18]. During the last few years, a number of relevant hydroxamic acids have been synthesized using amidases. Pandey et al. (2011) used hyper-induced resting cells of Bacillus sp. APB-6 treated with DTT for acetohydroxamic acid synthesis with 93% molar conversion of acetamide (300 mM) in the presence of hydroxylamine (800 mM) in 1 L reaction mixture [98]. After lyophilization, 62 g containing 34% (w/w) powder containing acetohydroxamic acid was recovered. The whole-cell biocatalyst of Geobacillus pallidus BTP-5x MTCC 9225 has been used for the synthesis of acetohydroxamic acid in a batch reaction at 1 L scale that produced 0.28 M of acetohydroxamic acid in 80 min [100]. The acyl transfer activity of the amidase of Alcaligenes sp. MTCC 10674 has been used for conversion of benzamide and hydroxylamine to benzohydroxamic acid with 24.6 g L−1 h−1 productivity [20]. In another study, Bhatia et al. had synthesized 16 g of nicotinyl hydroxamic acid from nicotinamide and hydroxylamine at 1 L scale using free cells of Pseudomonas putida BR1 with 32 g L−1 h−1 volumetric productivity [97]. The Bacillus smithii IITR6b2 exhibiting acyltransferase activity has been explored for the synthesis of another important product, i.e., nicotinic acid hydroxamate [99]. To avoid substrate inhibition effect, they used a fed-batch process based on the optimized parameters with two feedings of substrates (200/200 mM) at 40-min intervals. A molar conversion of 89.4% with a productivity of 52.9 g h−1 g−1dcw was achieved at 50 mL scale. Major obstacles in hydroxamic acid synthesis are substrate/product inhibition, lower molar conversion, complex bi-substrate kinetics, stability of biocatalyst, and scale-up. To make enzymatic production of hydroxamic acid commercially viable, effort should be made to understand the bi-substrate kinetics in depth, improve enzyme properties, and explore reaction medium engineering.
Nitrile Metabolizing Enzymes in Bioremediation and Biodegradation
Most of the nitriles and few amides are toxic, and carcinogenic, therefore, harmful to human beings, animals, and plants [2, 6, 32]. In spite of toxicity, nitriles and amides are extensively used in agriculture, pharmaceuticals, cosmetics, plastics, synthetic rubbers, dye, and textile [3,4,5,6]. The major cause of nitrile/amide entry into the environment is effluent from the industrial units either engaged in nitrile production or utilization or processing, accidental spillage, and use of nitrile/amide compounds in agriculture, i.e., herbicides [32]. These toxic compounds are thus continuously increasing in our ecosystem. Therefore, their degradation is inevitable and of great significance. Chemical and physical methods are always problematic as far as the environment is concerned. On the other hand, biological route using one or more enzymes is ecofriendly. During the last decade, a number microorganisms involved in the degradation of nitriles/amides have been isolated and characterized, and their potential for bioremediation has been highlighted [3, 4, 101].
Status and Challenges for Industrial-Scale Synthesis
Nitrile hydratase-mediated synthesis of acrylamide from acrylonitrile is one of the first successful examples of production of commodity chemicals at industrial scale. In year 1996, the production of acrylamide from nitrile was estimated to be 30,000 t year−1 [31], and now, it has increased to more than 400,000 t year−1. The market of polyacrylamide (a polymer of acrylamide) was worth USD 3.95 billion in 2012, and it is expected to touch USD 6.91 billion by 2019 (http://www.tmrblog.com/2014/08/global-polyacrylamide-market-is.html). A number of important acids and amides (such as nicotinic acid, nicotinamide, and (R)-mandelic acid) have been synthesized using nitrile metabolizing enzymes by several industries like Lonza and BASF. Presently, these compounds have very good market, and reports of the surveys reflecting rapid expansion of market for these products in future are available on the internet. Mandelic acid has applications in various industries, and cosmetic industry alone had its demand to the tune of USD 465 billion in 2015, and it is estimated to reach USD 670 billion by 2023 (www.gminsights.com/industry-analysis/mandelic-acid-market). Glycolic acid had a global market of 93.3 million USD in 2011 and is expected to reach 415 million by 2024 (http://www.grandviewresearch.com/press-release/global-glycolic-acid-market). Acrylic acid market was around USD 11 billion in 2013, and it is expected to be USD 18.8 billion by 2020 (www.alliedmarketresearch.com/acrylic-acid-market). However, there may be some skepticism on these market analyses and predictions, but still, these reflect industrial importance and market trends of the products of nitrile metabolizing enzymes or the derivatives of these products. Some industries such as Prozomix Ltd. (UK) and Codexis Inc. (USA) are involved in the commercial production of nitrilase/nitrile hydratase, and these enzymes are used for the enzymatic synthesis of a number of chemicals including precursors of some important drugs, i.e., ibuprofen [102], (S)-naproxen [103], lipitor [63], clopidogrel [66], levetiracetam [90], substituted morpholines [104], taxol [105], and rosuvastatin [106]. Despite the huge commercial potential of nitrile metabolizing enzymes, their factual application in the industries mainly in the synthesis of fine and commodity chemicals is limited due to some biological, economical, and technical challenges. Major biological challenges to nitrile metabolizing enzymes in industrial biocatalysis and biotransformation are low enzyme activity in wild organism, narrow substrate spectrum, and stability concern in wide range pH, temperature and organic solvents, and substrate/product inhibition. The technological challenges include scale-up of laboratory reactions to pilot or industrial scale and downstream processing. Cost of substrate and production of biocatalysts, lower volumetric yield of products collectively makes the enzymatic synthesis invariably more expensive than chemical synthesis. Academia and industry need to collaborate to sort out these hurdles in the application and adoption of enzymes of nitrile metabolism in industry.
Future Prospects
Nitrile metabolizing enzymes are the biocatalysts of choice for the synthesis of various nitriles, amides, and carboxylic acids. Research on nitrile catabolism has undergone rapid development, and obstacles on the way would be overcome in the future. However, the enzymes of nitrile anabolism are less explored for the synthesis of nitriles. Enzymes and enzymatic processes are at the center of green chemistry; therefore, future research should focus on characterizing the enzymes involved in nitrile metabolism, so that desired product can be produced using simple substrate in a cascade reaction. Screening of novel biocatalysts, improvement in existing bioresources, and designing novel bioprocesses looks to be the thematic areas of nitrile biotransformation research in future. Extensive screening for novel biocatalysts from extreme habitats with desired properties via traditional, metagenomic, genome mining, and directed evolution approaches needs to be intensively considered. Research focus has to be on improving the properties of these enzymes such as substrate specificity, enantioselectivity, regio-specificity, stability, and function in non-conventional environments. Strategy must be developed to counter substrate and product inhibition at molecular level by directed evolution and site-directed mutagenesis. The integrated approach for synthesis of nitriles involving aldoxime dehydratases or hydroxy nitrile lyase and conversion of nitriles to amides/acids with nitrilase, nitrile hydratase, and amidase will be worthwhile to be explored further for the synthesis of fine and commodity chemicals. Nature’s way to synthesize and metabolize nitriles can be replicated in laboratory to produce desired high-value products.
References
Howden, A. J. M., & Preston, G. M. (2009). Nitrilase enzymes and their role in plant–microbe interactions. Microbial Biotechnology, 2(4), 441–451. https://doi.org/10.1111/j.1751-7915.2009.00111.x.
Bhalla, T.C., Sharma, N., & Bhatia, R.K. (2012). Microbial degradation of cyanides and nitrile. In: Satyanarayan T (Eds.), Microorganisms in Environmental Management. Springer Netherlands, pp. 569–587.
Martınkova, L., & Kren, V. (2010). Biotransformation with nitrilases. Currunt Opinion in Chemical Biology, 14(2), 130–137. https://doi.org/10.1016/j.cbpa.2009.11.018.
Gong, J. S., Lu, Z. M., Li, H., Shi, J. S., Zhou, Z. M., & Xu, Z. H. (2012). Nitrilases in nitrile biocatalysis: recent progress and forthcoming research. Microbial Cell Factory, 11(1), 142. https://doi.org/10.1186/1475-2859-11-142.
Wang, M. X. (2015). Enantioselective biotransformation of nitriles in organic synthesis. Accounts of Chemical Research, 48(3), 602–611. https://doi.org/10.1021/ar500406s.
Chen, J., Zheng, R., Zheng, Y., & Shen, Y. (2009). Microbial transformation of nitriles to high-value acids or amides. Advance in Biochemical Engineering/Biotechnology, 113, 33–77.
Thimann, K. V., & Mahadevan, S. (1964). Nitrilase I: occurrence, preparation, and general properties of the enzyme. Archives in Biochemistry and Biophysics, 105(1), 133–141. https://doi.org/10.1016/0003-9861(64)90244-9.
Asano, Y., Yasuda, T., Tani, T., & Yamada, H. (1982). A new enzymatic method of acrylamide production. Agriculture and Biological Chemistry, 46, 1183–1199.
Kato, Y., Nakamura, K., Sakiyama, H., Mayhew, S. G., & Asano, Y. (2000). Novel heme-containing lyase, phenylacetaldoxime dehydratase from Bacillus sp. strain OxB-1: purification, characterization, and molecular cloning of the gene. Biochemistry Journal, 39(4), 800–809. https://doi.org/10.1021/bi991598u.
Oinuma, K., Hashimoto, Y., Konishi, K., Goda, M., Noguchi, T., Higashibata, H., & Kobayashi, M. (2003). Novel aldoxime dehydratase involved in carbon-nitrogen triple bond synthesis of Pseudomonas chlororaphis B23. Journal of Biological Chemistry, 278(32), 29600–29608. https://doi.org/10.1074/jbc.M211832200.
Pinakoulaki, E., Koutsoupakis, C., Sawai, H., Pavlou, A., Kato, Y., Asano, Y., & Aono, S. (2011). Aldoxime dehydratase: probing the heme environment involved in the synthesis of the carbon nitrogen triple bond. Journal of Physical Chemistry B, 115(44), 13012–13018. https://doi.org/10.1021/jp205944e.
Yamaguchi, T., & Asano, Y. (2015). Complete genome sequence of an aldoxime degrader, Bacillus sp. OxB-1. Genome Announcement, 3(1), e00025–e00015. https://doi.org/10.1128/genomeA.00025-15.
Sharma, M., Sharma, N. N., & Bhalla, T. C. (2005). Hydroxynitrile lyases: at the interface of biology and chemistry. Enzyme and Microbial Technology, 37(3), 279–294. https://doi.org/10.1016/j.enzmictec.2005.04.013.
Lanfranchi, E., & Sheldon, R. A. (2013). Recent developments in hydroxynitrile lyases for industrial biotechnology. Recent Patents on Biotechnology, 7(3), 197–206. https://doi.org/10.2174/18722083113076660010.
Kassim, M. A., & Rumbold, K. (2014). HCN production and hydroxynitrile lyase: a natural activity in plants and a renewed biotechnological interest. Biotechnology Letters, 36(2), 223–228. https://doi.org/10.1007/s10529-013-1353-9.
Sharma, M., Sharma, N. N., & Bhalla, T. C. (2009). Amidases : versatile enzymes in nature. Review in Environment Science and Biotechnology, 8(4), 343–366. https://doi.org/10.1007/s11157-009-9175-x.
Chand, D., Kumar, H., Sankhian, U. D., Kumar, D., Vitzthum, F., & Bhalla, T. C. (2004). Treatment of simulated wastewater containing toxic amides by immobilized Rhodococcus rhodochrous NHB-2 using a highly 5-stage plug flow reactor. World Journal of Microbiology and Biotechnology, 20(7), 679–686. https://doi.org/10.1007/s11274-004-2158-8.
Fournand, D., Bigey, F., Ratomahenina, R., Arnaud, A., & Galzy, P. (1997). Biocatalyst mprovement for the production of short-chain hydroxamic acids. Enzyme and Microbial Technology, 20(6), 424–431. https://doi.org/10.1016/S0141-0229(96)00170-6.
Bhatia, R. K., Bhatia, S. K., Mehta, P. K., & Bhalla, T. C. (2013a). Production and characterization of acyl transfer activity of amidase from Alcaligenes sp. MTCC 10674 for synthesis of hydroxamic acids. Journal of Microbial and Biochemical Technology, 5, 1–5.
Bhatia, R. K., Bhatia, S. K., Mehta, P. K., & Bhalla, T. C. (2013b). Bench scale production of benzohydroxamic acid using acyl transfer activity of amidase from Alcaligenes sp. MTCC 10674. Journal of Industrial Microbiology and Biotechnology, 40(1), 21–27. https://doi.org/10.1007/s10295-012-1206-x.
Baum, S., van Rantwijk, F., & Stolz, A. (2012). Application of a recombinant Escherichia coli whole-cell catalyst expressing hydroxynitrile lyase and nitrilase activities in ionic liquids for the production of (S)-mandelic acid and (S)-mandeloamide. Advanced Synthesis and Catalysis, 354(1), 113–122. https://doi.org/10.1002/adsc.201100391.
Scholz, K. E., Okrob, D., Kopka, B., Grunberger, A., Pohl, M., Jaeger, K., & Krauss, U. (2012). Synthesis of chiral cyanohydrins by recombinant Escherichia coli cells in a micro-aqueous reaction system. Applied and Environment Microbiology, 78(14), 5025–5027. https://doi.org/10.1128/AEM.00582-12.
Alagoz, D., Tukel, S. S., & Yildirim, D. (2015). Enantioselective synthesis of various cyanohydrins using covalently immobilized preparations of hydroxynitrile lyase from Prunus dulcis. Applied and Environment Microbiology, 177, 1348–1363.
Metzner, R., Okazaki, S., Asano, Y., & Groger, H. (2014). Cyanide-free enantioselective synthesis of nitriles: synthetic proof of a biocatalytic concept and mechanistic insights. ChemCatChem, 6(11), 3105–3109. https://doi.org/10.1002/cctc.201402612.
Miki, Y., & Asano, Y. (2014). Biosynthetic pathway for the cyanide-free production of phenylacetonitrile in Escherichia coli by utilizing plant cytochrome P450 79A2 and bacterial aldoxime dehydratase. Applied and Environment Microbiology, 80(21), 6828–6836. https://doi.org/10.1128/AEM.01623-14.
Pace, H. C., & Brenner, C. (2001). The nitrilase superfamily: classification, structure and function. Genome Biology, 2, 1–9.
O’Reilly, C., & Turner, P. D. (2003). The nitrilase family of CN hydrolysing enzyme—a comparative study. Journal of Applied Microbiology, 95(6), 1161–1174. https://doi.org/10.1046/j.1365-2672.2003.02123.x.
Thuku, R. N., Brady, D., Benedik, M. J., & Sewell, B. T. (2009). Microbial nitrilases: versatile, spiral forming, industrial enzymes. Journal of Applied Microbiology, 106(3), 703–727. https://doi.org/10.1111/j.1365-2672.2008.03941.x.
Gong, J. S., Shi, J. S., Lu, Z. M., Zhou, Z. M., & Xu, Z. H. (2015). Nitrile converting enzymes as a tool to improve biocatalysis in organic synthesis: recent insight and promises. Critical Reviews in Biotechnology, 23, 1–13.
Martınkova, L., Rucka, L., Nesvera, J., & Patek, M. (2017). Recent advances and challenges in the heterologous production of microbial nitrilases for biocatalytic applications. World Journal of Microbiology and Biotechnology, 33(1), 8. https://doi.org/10.1007/s11274-016-2173-6.
Yamada, H., & Kobayashi, M. (1996). Nitrile hydratase and its application to industrial production of acrylamide. Bioscience Biotechnology and Biochemistry, 60(9), 1391–1400. https://doi.org/10.1271/bbb.60.1391.
Prasad, S., & Bhalla, T. C. (2010). Nitrile hydratases (NHases): at the interface of academia and industry. Biotechnology Advances, 28(6), 725–741. https://doi.org/10.1016/j.biotechadv.2010.05.020.
Mahadevan, S. (1973). Role of oximes in nitrogen metabolism in plants. Annual Review of Plant Physiology, 24(1), 69–88. https://doi.org/10.1146/annurev.pp.24.060173.000441.
Kato, Y., & Asano, Y. (2006). Molecular and enzymatic analysis of the “aldoxime–nitrile pathway” in the glutaronitrile degrader Pseudomonas sp. K-9. Applied Microbiology and Biotechnology, 70(1), 92–101. https://doi.org/10.1007/s00253-005-0044-4.
Hashimoto, Y., Hosaka, H., Oinuma, K., Goda, M., Higashibata, H., & Kobayashi, M. (2005). Nitrile pathway involving acyl-CoA synthetase. Journal of Biological Chemistry, 280, s8660–s8667.
Xie, S. X., Kato, Y., Komeda, H., Yoshida, S., & Asano, Y. (2003). A gene cluster responsible for alkylaldoxime metabolism coexisting with nitrile hydratase and amidase in Rhodococcus globerulus A-4. Biochemistry, 42(41), 12056–12066. https://doi.org/10.1021/bi035092u.
Nomura, J., Hashimoto, H., Ohtac, T., Hashimotoa, Y., Wadaa, K., Naruta, Y., Oinumaa, K., & Kobayashi, M. (2012). Crystal structure of aldoxime dehydratase and its catalytic mechanism involved in carbon-nitrogen triple-bond synthesis. PNAS, 110, 2810–2815.
Mateoa, C., Chmuraa, A., Rustlerb, S., Rantwijka, F., Stolzb, A., & Sheldona, R. A. (2006). Synthesis of enantiomerically pure (S)-mandelic acid using an oxynitrilase–nitrilase bienzymatic cascade: a nitrilase surprisingly shows nitrile hydratase activity. Tetrahedron: Asymmetry, 17(3), 320–323. https://doi.org/10.1016/j.tetasy.2006.01.020.
Ueatrongchit, T., Ohmiya, T., Kittikun, H., & Asano, Y. (2010). Hydroxynitrile lyase from Passiflora edulis : purification, characteristics and application in asymmetric synthesis of (R)-mandelonitrile. Enzyme and Microbial Technology, 46(6), 456–465. https://doi.org/10.1016/j.enzmictec.2010.02.008.
Asif, M., & Bhalla, T. C. (2016). Hydroxynitrile lyase of wild apricot (Prunus armeniaca L.): purification, characterization and application in synthesis of enantiopure mandelonitrile. Catalysis Letters, 46, 1118–1127.
Fuhshuku, K., & Asano, Y. (2011). Synthesis of (R)-β-nitro alcohols catalyzed by R-selective hydroxynitrile lyase from Arabidopsis thaliana in the aqueous–organic biphasic system. Journal of Biotechnology, 153(3-4), 153–159. https://doi.org/10.1016/j.jbiotec.2011.03.011.
Yamaguchi, T., Noge, K., & Asano, Y. (2016). Cytochrome P450 CYP71AT96 catalyses the final step of herbivore-induced phenylacetonitrile biosynthesis in the giant knotweed, Fallopia sachalinensis. Plant Molecular Biology, 91(3), 229–239. https://doi.org/10.1007/s11103-016-0459-6.
Zagrobelny, M., & Moller, L. (2011). Cynogenic glycosides in chemical warfare between plants and insects: the burnet moth-birdsfoot trefoil model system. Phytochemistry, 72(13), 1585–1592. https://doi.org/10.1016/j.phytochem.2011.02.023.
Zagrobelny, M., Bak, S., & Moller, B. L. (2008). Cyanogenesis in plants and arthropods. Journal of Phytochemistry, 69(7), 1457–1468. https://doi.org/10.1016/j.phytochem.2008.02.019.
Hussain, Z., Wiedner, R., Steiner, K., Hajek, T., Avi, M., Hecher, B., & Sessitsch, A. (2012). Characterization of two bacterial hydroxynitrile lyases with high similarity to cupin superfamily proteins. Applied and Environment Microbiology, 78(6), 2053–2055. https://doi.org/10.1128/AEM.06899-11.
Wiedner, R., Gruber-khadjawi, M., Schwab, H., & Steiner, K. (2014). Discovery of a novel (R)-selective bacterial hydroxynitrile lyase from Acidobacterium capsulatum. Computational and Structural Biotechnology Journal, 10(16), 58–62. https://doi.org/10.1016/j.csbj.2014.07.002.
Bracco, P., Busch, H., von Langermann, J., & Hanefeld, U. (2016). Enantioselective synthesis of cyanohydrins catalysed by hydroxynitrile lyases—a review. Organic & Biomolecular Chemistry, 14(27), 6375–6389. https://doi.org/10.1039/C6OB00934D.
Dadashipour, M., Ishida, Y., Yamamoto, K., & Asano, Y. (2015). Discovery and molecular and biocatalytic properties of hydroxynitrile lyase from an invasive millipede, Chamberlinius hualienensis. PNAS, 112(34), 10605–10610. https://doi.org/10.1073/pnas.1508311112.
Prasad, S., Misra, A., Jangir, V. P., Awasthi, A., Raj, J., & Bhalla, T. C. (2007). A propionitrile-induced nitrilase of Rhodococcus sp. NDB 1165 and its application in nicotinic acid synthesis. World Journal of Microbiology and Biotechnology, 23(3), 345–353. https://doi.org/10.1007/s11274-006-9230-5.
Sharma, N. N., Sharma, M., Kumar, H., & Bhalla, T. C. (2006). Nocardia globerula NHB-2: bench scale production of nicotinic acid. Process Biochemistry, 41(9), 2078–2081. https://doi.org/10.1016/j.procbio.2006.04.007.
Sharma, N. N., Monica, S., & Bhalla, T. C. (2011). An improved nitrilase-mediated bioprocess for synthesis of nicotinic acid from 3-cyanopyridine with hyperinduced Nocardia globerula NHB-2. Journal of Industrial Microbiology, 38(9), 1235–1243. https://doi.org/10.1007/s10295-010-0902-7.
Sharma, N. N., Sharma, M., & Bhalla, T. C. (2012). Nocardia globerula NHB-2 nitrilase catalyzed biotransformation of 4-cyanopyridine to isonicotinic acid. AMB Express, 2(1), 25. https://doi.org/10.1186/2191-0855-2-25.
Zhu, X. Y., Gong, J. S., Li, H., Lu, Z. M., Shi, J. S., & Xu, Z. H. (2014). Bench-scale biosynthesis of isonicotinic acid from 4-cyanopyridine by Pseudomonas putida. Chemical Papers, 68, 739–744.
Zhang, Z. J., Xu, J. H., He, Y. C., Ouyang, L. M., Liu, Y. Y., & Imanaka, T. (2010). Efficient production of (R)-(−)-mandelic acid with highly substrate/product tolerant and enatioselective nitrilase from recombinant Alcaligenes sp. Process Biochemistry, 45(6), 887–891. https://doi.org/10.1016/j.procbio.2010.02.011.
Xue, Y., Xu, M., Chen, H., Liu, Z., Wang, Y., & Zheng, Y. (2013). A novel integrated bioprocess for efficient production of (R)-(−)-mandelic acid with immobilized Alcaligenes faecalis ZJUTB10. Organic Process & Research Development, 17(2), 213–220. https://doi.org/10.1021/op3001993.
Ni, K., Wang, H., Zhao, L., Zhang, M., Zhang, S., Ren, Y., & Wei, D. (2013). Efficient production of (R)-(−)-mandelic acid in biphasic system by immobilized recombinant E. coli. Journal of Biotechnology, 167(4), 433–440. https://doi.org/10.1016/j.jbiotec.2013.07.024.
Bhatia, S. K., Mehta, P. K., Bhatia, R. K., & Bhalla, T. C. (2014). Optimization of arylacetonitrilase production from Alcaligenes sp. MTCC 10675 and its application in mandelic acid synthesis. Applied Microbiology and Biotechnology, 98(1), 83–94. https://doi.org/10.1007/s00253-013-5288-9.
He, Y. C., Xu, J. H., Su, J. H., & Zhou, L. (2009). Bioproduction of glycolic acid from glycolonitrile with a new bacterial isolate of Alcaligenes sp. ECU0401. Applied Biochemistry and Biotechnology, 160, 1428–1440.
Wu, S., Fogiel, A. J., Petrillo, K. L., Jackson, R. E., Parker, K. N., DiCosimo, R., Ben-Bassat, A., O’Keefe, D. P., & Payne, M. S. (2008). Protein engineering of nitrilase for chemoenzymatic production of glycolic acid. Biotechnology and Bioengineering, 99(3), 717–720. https://doi.org/10.1002/bit.21643.
Raj, J., Singh, N., Prasad, S., Seth, A., & Bhalla, T. C. (2007a). Bioconversion of benzonitrile to benzoic acid using free and agar entrapped cells of Nocardia globerula NHB-2. Acta Microbiologica Imunologica Hungerica, 54(1), 79–88. https://doi.org/10.1556/AMicr.54.2007.1.8.
Kumar, V., Kumar, V., Thakur, N., & Bhalla, T. C. (2015). Bench scale synthesis of p-hydroxybenzoic acid using whole cells nitrilase of Gordonia terrae mutant E9. Bioprocess and Biosystems Engineering, 38(7), 1267–1279. https://doi.org/10.1007/s00449-015-1367-x.
Kumar, V., & Bhalla, T. C. (2013). Transformation of p-hydroxybenzonitrile to p-hydroxybenzoic acid using nitrilase activity of Gordonia terrae. Biocatalysis and Biotransformation, 31(1), 42–48. https://doi.org/10.3109/10242422.2012.757761.
Bergeron, S., Chaplin, D., Edwards, J. H., Ellis, B. S. W., Hill, C. L., Holt-Tiffin, K., et al. (2006). Nitrilase-catalysed desymmetrisation of 3-hydroxyglutaronitrile: preparation of a statin side-chain intermediate. Organic Process & Research Development, 10(3), 661–665. https://doi.org/10.1021/op050257n.
Fan, H., Chen, L., Sun, H., Wang, H., Ren, Y., & Wei, D. (2017). A novel nitrilase from Ralstonia eutropha H16 and its application to nicotinic acid production. Bioprocess and Biosystems Engineering, 40(8), 1271–1281. https://doi.org/10.1007/s00449-017-1787-x.
Mathew, C. D., Nagasawa, T., Kobayashi, M., & Yamada, H. (1988). Nitrilase-catalyzed production of nicotinic acid from 3-cyanopyridine in Rhodococcus rhodochrous J1. Applied and Environment Microbiology, 54, 1030–1032.
Zhang, C. S., Zhang, Z. J., Li, C. X., Yu, H. L., Zheng, G. W., & Xu, J. H. (2012). Efficient production of (R)-o-chloromandelic acid by deracemization of o-chloromandelonitrile with a new nitrilase mined from Labrenzia aggregata. Applied Microbiology and Biotechnology, 95(1), 91–99. https://doi.org/10.1007/s00253-012-3993-4.
Wu, S., Fogiel, A. J., Petrillo, K. L., Hann, E. C., Mersinger, L. J., Dicosimo, R., et al. (2007). Protein engineering of Acidovorax facilis 72W nitrilase for bioprocess development. Biotechnology, 97, 689–693.
Zhang, Z. J., Xu, J. H., He, Y. C., Ouyang, L. M., & Liu, Y. Y. (2011). Cloning and biochemical properties of a highly thermostable and enantioselective nitrilase from Alcaligenes sp. ECU0401 and its potential for (R)-(2)-mandelic acid production. Bioprocess and Biosystems Engineering, 34(3), 315–322. https://doi.org/10.1007/s00449-010-0473-z.
Bhatia, S. K., Mehta, P. K., Bhatia, R. K., & Bhalla, T. C. (2011). An isobutyronitrile-induced bienzymatic system of Alcaligenes sp. MTCC 10674 and its application in the synthesis of α-hydroxyisobutyric acid. Bioprocess and Biosystems Engineering, 36, 613–625.
Zabaznaya, E., Kozulin, S., & Voronin, S. (1998). Selection of strains transforming acrylonitrile and acrylamide into acrylic acid. Applied Biochemistry and Microbiology, 34, 341–345.
Nagasawa, T., Nakamura, T., & Yamada, H. (1990). Production of acrylic acid and methacrylic acid using Rhodococcus rhodochrous J1 nitrilase. Applied Microbiology and Biotechnology, 34, 322–324.
Banerjee, A., Kaul, P., & Banrjee, U. C. (2006). Enhancing the catalytic potential of nitrilase from Pseudomonas putida for stereoselective nitrile hydrolysis. Applied Microbiology and Biotechnology, 72(1), 77–87. https://doi.org/10.1007/s00253-005-0255-8.
Kaul, P., Stolz, A., & Banerjee, U. C. (2007). Cross-linked amorphous nitrilase aggregates for enantioselective nitrile hydrolysis. Advanced Synthesis & Catalysis, 349(13), 2167–2176. https://doi.org/10.1002/adsc.200700125.
Almatawah, Q. A., & Cowan, D. A. (1999). Thermostable nitrilase catalysed production of nicotinic acid from 3-cyanopyridine. Enzyme and Microbial Technology, 25(8-9), 718–724. https://doi.org/10.1016/S0141-0229(99)00104-0.
Pai, O., Banoth, L., Ghosh, S., Chisti, Y., & Chand, U. (2014). Biotransformation of 3-cyanopyridine to nicotinic acid by free and immobilized cells of recombinant Escherichia coli. Process Biochemistry, 49(4), 655–659. https://doi.org/10.1016/j.procbio.2014.01.023.
Wu, Y., Gong, J. S., Lu, Z. M., Li, H., Zhu, X. Y., Li, H., Shi, J. S., et al. (2013). Isolation and characterization of Gibberella intermedia CA3-1, a novel and versatile nitrilase-producing fungus. Journal of Basic Microbiology, 53, 1–8.
Shen, M., Zheng, Y. G., & Shen, Y. C. (2009). Isolation and characterization of a novel Arthrobacter nitroguajacolicus ZJUTB06-99, capable of converting acrylonitrile to acrylic acid. Process Biochemistry, 44(7), 781–785. https://doi.org/10.1016/j.procbio.2009.03.006.
Chen, J., Zheng, Y. G., & Shen, Y. C. (2008). Biotransformation of p-methoxyphenylacetonitrile into p-methoxyphenylacetic acid by resting cells of Bacillus subtilis. Biotechnology and Applied Biochemistry, 50(3), 147–153. https://doi.org/10.1042/BA20070106.
Jin, L. Q., Li, Z. T., Liu, Z. Q., Zheng, Y. G., & Shen, Y. C. (2014). Efficient production of methionine from 2-amino-4-methylthiobutanenitrile by recombinant Escherichia coli harboring nitrilase. Journal of Industrial Microbiology and Biotechnology, 41(10), 1479–1486. https://doi.org/10.1007/s10295-014-1490-8.
Jin, L. Q., Guo, D. J., Li, Z. T., Liu, Z. Q., & Zheng, Y. G. (2016). Immobilization of nitrilase on bioinspired silica for efficient synthesis of 2-hydroxy-4-(methylthio) butanoic acid from 2-hydroxy-4-(methylthio) butanenitrile. Journal of Industrial Microbiology and Biotechnology, 43(5), 585–593. https://doi.org/10.1007/s10295-016-1747-5.
Wang, H., Sun, H., Gao, W., & Wei, D. (2014). Efficient production of (R)-o-Chloromandelic acid by recombinant Escherichia coli cells harboring nitrilase from Burkholderia cenocepacia J2315. Organic Process & Research Development, 18(6), 767–773. https://doi.org/10.1021/op400174a.
Liu, Z., Zhang, X., Xue, Y., Xu, M., & Zheng, Y. (2014). Improvement of Alcaligenes faecalis nitrilase by gene site saturation mutagenesis and its application in stereospecific biosynthesis of (R)-(−)-mandelic acid. Journal of Agriculture and Food Chemistry, 62, 4685−4694.
Martinkova, L., & Kren, V. (2002). Nitrile and amide-converting microbial enzymes: stereo-, regio- and chemoselectivity. Biocatalysis and Biotransformation, 20, 79–93.
Kim, B., & Hyun, H. (2002). Production of acrylamide using immobilized cells of Rhodococcus rhodochrous M33. Biotechnology and Bioprocess Engineering, 7(4), 194–200. https://doi.org/10.1007/BF02932969.
Raj, J., Nand, N., & Shreenath, S. (2008). Acrylamide synthesis using agar entrapped cells of Rhodococcus rhodochrous PA-34 in a partitioned fed batch reactor. Journal of Industrial Microbiology, 35(1), 35–40. https://doi.org/10.1007/s10295-007-0263-z.
Pratush, A., Seth, A., & Bhalla, T. C. (2012). Optimization of process parameters for conversion of 3-cyanopyridine to nicotinamide using resting cells of mutant 4D strain of Rhodococcus rhodochrous PA-34. International Journal of Bioautomation, 15, 151–158.
Wang, Z., Liu, Z., Cui, W., & Zhou, Z. (2017). Establishment of bioprocess for synthesis of nicotinamide by recombinant Escherichia coli expressing high-molecular-mass nitrile hydratase. Applied Biochemistry and Biotechnology, 182(4), 1458–1466. https://doi.org/10.1007/s12010-017-2410-y.
Mauger, J., Nagasawa, T., & Yamada, H. (1988). Nitrile hydratase catalyzed production of isonicotinamide, picolinamide and pyrazinamide from 4-cyanopyridine, 2-cyano- pyridine, and 3-cyanopyrazine in Rhodococcus rhodochrous J1. Journal of Biotechnology, 8(1), 87–96. https://doi.org/10.1016/0168-1656(88)90071-5.
Raj, J., Seth, A., Prasad, S., & Bhalla, T. C. (2007b). Bioconversion of butyronitrile to butyramide using whole cells of Rhodococcus rhodochrous PA-34. Applied Microbiology and Biotechnology, 74(3), 535–539. https://doi.org/10.1007/s00253-006-0693-y.
Hann, E. C., Eisenberg, A., Fager, S. K., Perkins, N. K., Gallagher, F. G., Cooper, S. M., et al. (1999). 5-Cyanovaleramide production using immobilized Pseudomonas chlororaphis B23. Bioorganic and Medical Chemistry, 7(10), 2239–2245. https://doi.org/10.1016/S0968-0896(99)00157-1.
Tucker, J.L., Xu, L., Yu, W., Scott, R., Zhao, L., & Ran, N. (2009). Chemoenzymatic processes for preparation of levetiracetam. Patent, WO 2009009117 A3.
Yasukawa, K., & Asano, Y. (2012). Enzymatic synthesis of chiral phenylalanine derivatives by a dynamic kinetic resolution of corresponding amide and nitrile substrates with a multi-enzyme system. Advanced Synthesis & Catalysis, 354(17), 3327–3332. https://doi.org/10.1002/adsc.201100923.
Yasukawa, K., Hasemi, R., & Asano, Y. (2011). Dynamic kinetic resolution of α-aminonitriles to form α-chiral amino acids. Advanced Synthesis & Catalysis, 353(13), 2328–2332. https://doi.org/10.1002/adsc.201100360.
Tang, R., Shen, Y., Wang, M., & Zhai, Y. (2017). Highly chemoselective and efficient production of 2, 6-difluorobenzamide using Rhodococcus ruber CGMCC3090 resting cells. Journal of Bioscience and Bioengineering, 124(6), 641–646. https://doi.org/10.1016/j.jbiosc.2017.07.001.
Maksimova, Y. G., Gorbunova, A. N., & Demakov, V. A. (2017). Stereoselective biotransformation of phenylglycine nitrile by heterogeneous biocatalyst based on immobilized bacterial cells and enzyme preparation. Doklady Biochemistry and Biophysics, 474(1), 183–185. https://doi.org/10.1134/S1607672917030139.
Vejvoda, V., Kaplan, O., Kubac, D., Kren, V., & Martinkova, L. (2006). Immobilization of fungal nitrilase and bacterial amidase—two enzymes working in accord. Biocatalysis and Biotransformation, 24(6), 414–418. https://doi.org/10.1080/10242420601033910.
Bhatia, R. K., Bhatia, S. K., Mehta, P. K., & Bhalla, T. C. (2014). Biotransformation of nicotinamide to nicotinyl hydroxamic acid at bench scale by amidase acyl transfer activity of Pseudomonas putida BR-1. Journal of Molecular Catalysis. B Enzymatic, 108, 89–95. https://doi.org/10.1016/j.molcatb.2014.07.001.
Pandey, D., Singh, R., & Chand, D. (2011). An improved bioprocess for synthesis of acetohydroxamic acid using DTT (dithiothreitol) treated resting cells of Bacillus sp. APB-6. Bioresource Technology, 102(11), 6579–6586. https://doi.org/10.1016/j.biortech.2011.03.071.
Agarwal, S., Gupta, M., & Choudhury, B. (2013). Bioprocess development for nicotinic acid hydroxamate synthesis by acyltransferase activity of Bacillus smithii strain IITR6b2. Journal of Industrial Microbiology and Biotechnology, 40(9), 937–946. https://doi.org/10.1007/s10295-013-1299-x.
Sharma, M., Sharma, N. N., & Bhalla, T. C. (2011). Biotransformation of acetamide to acetohydroxamic acid at bench scale using acyl transferase activity of amidase of Geobacillus pallidus BTP-5x MTCC 9225. Journal of Industrial Microbiology and Biotechnology, 52, 76–82.
Vesela, A. B., Franc, M., Pelantova, H., Kubac, D., Vejvoda, V., Sulc, M., Bhalla, T. C., et al. (2010). Hydrolysis of benzonitrile herbicides by soil actinobacteria and metabolite toxicity. Biodegradation, 21(5), 761–770. https://doi.org/10.1007/s10532-010-9341-4.
Yamamoto, K., Ueno, Y., Otsubo, K., Kawakami, K., & Komatsu, K. I. (1990). Production of S-(+)-ibuprofen from a nitrile compound by Acinetobacter sp strain-Ak226. Applied and Environmental Microbiology, 56(10), 3125–3129.
Effenberger, F., & Böhme, J. (1994). Enzyme-catalysed enantioselective hydrolysis of racemic naproxen nitrile. Bioorganic and Medical Chemistry, 2(7), 715–721. https://doi.org/10.1016/0968-0896(94)85022-4.
Ritzen, B., Hoekman, S., Verdasco, E. D., van Delft, F. L., & brutes, P. J. T. F. (2010). Enantioselective chemoenzymatic synthesis of cis and trans-2,5-disubstituted morpholines. Journal of Organic Chemistry, 75(10), 3461–3464. https://doi.org/10.1021/jo1003295.
Wilding, B., Veselá, A. B., Perry, J. J., Black, G. W., Zhang, M., Martínková, L., & Klempier, N. (2015). An investigation of nitrile transforming enzymes in the chemo-enzymatic synthesis of the taxol sidechain. Organic & Biomolecular Chemistry, 13(28), 7803–7812.
Yao, P., Li, J., Yuan, J., Han, C., Liu, X., Feng, J., Wu, Q., & Zhu, D. (2015). Enzymatic synthesis of a key intermediate for rosuvastatin by nitrilase-catalyzed hydrolysis of ethyl (R)-4-cyano-3-hydroxybutyate at high substrate concentration. ChemCatChem, 7(2), 271–275. https://doi.org/10.1002/cctc.201402877.
Acknowledgements
Authors are highly grateful to University Grants Commission (UGC), New Delhi, India, for providing financial assistance in the form of Senior Research Fellowship to Dr. Vijay Kumar. The computational facility availed at Sub-Distributed Information Centre (SDIC), Himachal Pradesh University, Shimla, is also duly acknowledged.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Bhalla, T.C., Kumar, V., Kumar, V. et al. Nitrile Metabolizing Enzymes in Biocatalysis and Biotransformation. Appl Biochem Biotechnol 185, 925–946 (2018). https://doi.org/10.1007/s12010-018-2705-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-018-2705-7