Abstract
Cancer cells are embedded within the tissue and interact dynamically with its components during cancer progression. Understanding the contribution of cellular components within the tumor microenvironment is crucial for the success of therapeutic applications. Here, we reveal the presence of perivascular GFAP+/Plp1+ cells within the tumor microenvironment. Using in vivo inducible Cre/loxP mediated systems, we demonstrated that these cells derive from tissue-resident Schwann cells. Genetic ablation of endogenous Schwann cells slowed down tumor growth and angiogenesis. Schwann cell-specific depletion also induced a boost in the immune surveillance by increasing tumor-infiltrating anti-tumor lymphocytes, while reducing immune-suppressor cells. In humans, a retrospective in silico analysis of tumor biopsies revealed that increased expression of Schwann cell-related genes within melanoma was associated with improved survival. Collectively, our study suggests that Schwann cells regulate tumor progression, indicating that manipulation of Schwann cells may provide a valuable tool to improve cancer patients’ outcomes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cancer cells embed within the tissue where they reside and interact dynamically with its constituents during disease progression [1, 2]. Because such interactions are critical for cancer outcomes, dissecting the function of each component within the tumor microenvironment is crucial for the success of therapeutic applications [2,3,4,5,6]. Recently, we and others have shown that peripheral nerves also infiltrate within the tumor microenvironment of solid tumors and interact with neoplastic cells, influencing cancer initiation, progression, and spread [7,8,9,10,11,12,13,14,15,16,17,18,19,20,21,22,23]. Less is known about the role of stromal cells associated with peripheral nerves during tumor development. Schwann cells are the principal glial cells of the peripheral nervous system [24]. They ensheath and keep the integrity of peripheral axonal extensions. New genetically based technologies available to cell biologists have lately provided novel and sometimes unexpected insight into Schwann cells’ roles in distinct tissue microenvironments and have greatly expanded our knowledge. Contemporary evidence shows that in certain pathophysiologic conditions Schwann cells: (i) assume a reactive state and begin to dedifferentiate, proliferate and mediate axon regeneration [25]; (ii) contribute to proper vascular remodeling and nerve-vessel alignment [26, 27]; (iii) modulate synaptic activity [28]; (iv) participate in neuropathic pain [29,30,31]; (v) present antigens [32]; (vi) function as stem cells, forming other cell types [24, 33,34,35,36,37,38]; (vii) regulate the functioning of other stem cells [24, 39]; and drive tissue repair [40, 41]. Given this broad involvement of Schwann cells in important biological processes, it is not surprising that they may also impact cancer development. It is well-known that Schwann cells can be the cells of origin of several tumors, such as neurofibromas [42], malignant peripheral nerve sheath tumors [42] and Schwannomas [43], the most common neoplasms of the peripheral nervous system [44]. Recent data indicates that possibly some sympathoadrenal tumors [45] may also arise from Schwann cells. On the other hand, the role of Schwann cells, as stromal cells, within the microenvironment of other peripheral primary solid tumors remains little explored. The importance of Schwann cells has been more associated to the regulation of cancer cells’ spread via perineural invasion, the process in which cancer cells grow and migrate along the native nerve fibers [45,46,47,48].
To understand the role of Schwann cells in cancer most studies have relied on in vitro co-cultures or cell transplantation settings [47, 49]. However, these strategies have their disadvantages. The artificial conditions and high concentration of mitogens that characterize cell culture systems may induce some characteristics in Schwann cells as well as in the cancer cells that may not be shared by the corresponding endogenous tumor Schwann cells in vivo under pathophysiological conditions [50, 51]. Also, the cells used in grafting experiments may differ from the endogenous ones resident in the tissue. Consequently, the contribution of endogenous Schwann cells to tumor development can not be precisely accessed by these methods. Therefore, conclusions drawn from these studies may be imprecise. To circumvent these issues, endogenous Schwann cells should be studied and manipulated directly in vivo in their habitat. Recently, this approach became possible with the advent of powerful genetically based tools that precisely allow the targeting of specific cell types for tracing or elimination to study their behaviors in vivo [10, 51,52,53,54]. Here, we revealed that cells expressing glial fibrillary acidic protein (GFAP), a marker for Schwann cells, surround blood vessels within the tumor microenvironment. These cells are derived from nerve-associated tissue-resident Schwann cells. Genetic depletion of endogenous Schwann cells counteracts tumor development, by regulating tumoral growth, angiogenesis and immunosurveillance. Our results provide unequivocal evidence of the impact of endogenous Schwann cells on cancer progression.
Materials and methods
Animals
The following mice were obtained from Jackson Laboratories (Jax) (Bar Harbor, ME, USA) and bred in our animal facility: LysM-Cre [55] (stock number 004781), in which myeloid lineage cells carry Cre-recombinase; Csf1r-Cre [56] (stock number 021024), in which macrophages, dendritic cells and bone marrow derived granulocytes express Cre-recombinase; ROSA26-TdTomato [57] (stock number 007914), in which a loxP-flanked STOP cassette prevent the transcription of the red fluorescent protein tdTomato; Rosa26-mT/mG (stock number 007576) in which a cell membrane-targeted tdTomato is expressed in all cells; and NG2-DsRed (stock number 008241) in which cells expressing NG2 proteoglycan are marked by DsRed fluorescence [58,59,60,61,62,63]. Experimental animals from crosses between these animals were used as hemizygous for the transgenes.
Generation of Plp1−CreER mice (stock number 005975), in which Schwann cells express tamoxifen-inducible Cre-recombinase driven by the proteolipid protein 1 (Plp1) promoter, have been previously described [64]. ROSA26-iDTR mice (stock number 007900) present Cre-inducible expression of DTR in cells susceptible to ablation following Diphtheria Toxin (DT) treatment. To eliminate Schwann cells in vivo, Plp1−CreER mice were crossed with ROSA26-iDTR, a mouse line conditionally expressing a Cre-inducible diphtheria toxin receptor (iDTR) allele [65]. In the resulting Plp1−CreER+/iDTR+ mice, after tamoxifen treatment, upon removal of the loxP-stop-loxP cassette by Cre recombination, DTR is expressed only in Plp1+ Schwann cells. In these mice, Plp1+ cells can be ablated by systemic injection of DT, while all other cells in these mice are insensitive to DT. Plp1−CreER−/iDTR+ mice were used as controls.
C57BL/6 wild-type (WT) mice were obtained from the Central Animal Facility of the Federal University of Minas Gerais (UFMG). BALB/c nude mice were obtained from Charles Rivers. Hi-Myc mice [FVB-Tg(ARR2/Pbsn-MYC)7Key [66]] were obtained from the National Cancer Institute. Our colony of Nestin‐green fluorescent protein (GFP) transgenic mice [67, 68] were maintained for the transgene on the C57BL/6 genetic background (a kind gift from Grigori Enikolopov, Cold Spring Harbor Laboratory) [69].
All animal care and experimental procedures were approved by the Ethics Animal Care and Use Committee (CEUA) from the Federal University of Minas Gerais, in accordance with the Guide for the Care and Use of Laboratory Animals. All colonies were housed in a pathogen-free animal facility of the Department of Pathology, UFMG, under controlled light cycle (12:12-h light/dark cycle) and fed ad libitum. Age-matched 8- to 12-week-old mice were used for all experiments.
Cell cultures
Human PC-3 cells stably expressing TdTomato were cultured in Ham’s F-12 Nutrient Mix (Gibco), supplemented with 10% FBS (Gibco), 1.5 g/L sodium bicarbonate (Sigma), and 500 mg/mL G418 (Sigma) [9]. Murine TRAMP-C2 cells [70] were purchased from the American Type Culture Collection and grown in Dulbecco's Modified Eagle Medium (DMEM) (Hyclone, GE Lifesciences) supplemented with 5% fetal bovine serum (FBS) at 37 °C in a humidified 5% CO2 [71]. Murine RM1 cells were obtained from American Type Culture Collection and maintained in RPMI supplemented with 10% FBS [72]. Murine B16-F10 melanoma is a common cell line that naturally originated in melanin-producing epithelia of C57BL6 mice [10]. These cells were originally obtained from American Type Culture Collection and were used to study melanoma development in vivo. The cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10% (v/v) fetal calf serum/2 mM l-glutamine/100 U/mL penicillin/100 μg/mL streptomycin. Cells were cultured in a humidified atmosphere of 95% air and 5% (v/v) CO2 at 37 °C. All cells used in this study have been tested and found negative for mycoplasma.
Bone marrow transplantation experiments
The whole bone marrow was harvested from ROSA mT/mG mice, in which all cells express membrane-localized tdTomato (mT) fluorescence. 2 × 106 tdTomato+ cells were injected into the tail-vein of lethally irradiated (1050 Rads) wild-type C57BL6 recipient mice.
Parabiosis
ROSA mT/mG mice (8 week old), in which all cells express membrane-localized tdTomato (mT) fluorescence, were conjoined to wildtype C57BL6 mice. Parabiosis was performed as previously described [73]. Cross circulation was confirmed after 4 weeks by flow cytometry of peripheral blood.
Tumor implantation
For prostate tumor implantation, orthotopic transplantation of prostate cancer cells (PC-3, Tramp-C2 or RM1) was performed by injection of 5 × 104 cells suspended in culture cell media into the ventral prostates of 8- to 12-week-old male mice. Ventral prostates were removed for analyses at the day of sacrifice.
For melanoma transplantation, appropriate numbers of B16-F10 cells were suspended in PBS and checked for viability using trypan blue staining. Only when cell viability was higher than 90% the cell batch was considered for injection. For subcutaneous application, the skin of all mice at an age of 8–12 weeks was shaved at the site of injection. 1 × 105 cells in 100 μL were injected subcutaneously into the right flank of each animal and the growth of the tumors was monitored until sacrifice. Growth of the tumors was assessed over time with a caliper as previously reported [10, 74]. For determination of tumor volume, tumor-bearing mice were anesthetized with isoflurane in O2 by manually restraining the animal and placing its head in an in-house-built nose cone. Tumors were removed 14 days after injection and weighted. Length (L) and width (W) were measured for calculating tumor volume (V) using the formula V = 0.5 × (L × W2) [75]. Tumor area was determined using calibrated photographs of each tumor using Fiji software®, version 1.53 (National Institute of Health, Bethesda, MD).
Tamoxifen treatment
For induction of CreER recombinase activity, tamoxifen was administered intraperitoneally, diluted in sunflower seed oil [53, 76]. 200 μL/day containing 2 mg of tamoxifen were administered daily for 5 days to each animal.
DT depletion
For depletion of Plp1+ cells, tamoxifen-pre-treated Plp1−CreER+/iDTR+ and their controls, Plp1-CreER−/iDTR+ mice received intraperitoneally 2 µg of DT diluted in 1X PBS [77] for 2 days. DT binds to specific cell surface receptor (DTR) and is internalized by receptor-mediated endocytosis [65]. Then, the catalytic A fragment of the toxin is translocated to the cytosol, inhibiting protein synthesis by ADP ribosylation of elongation factor 2, which causes the death of cells expressing DTR [78], in this case Plp1+ cells.
Immunohistochemistry and microscopy
Adult mice were deeply anesthetized with isoflurane and transcardially perfused with saline followed by 4% buffered paraformaldehyde (PFA, pH 7.4). After dissection, tumors were fixed overnight at 4 °C in 4% buffered paraformaldehyde, incubated overnight at 4 °C with 30% sucrose diluted in PBS, embedded and frozen in optimal cutting temperature compound (OCT, Tissue‐Tek). Embedded tissues were stored at − 80 °C. 20 μm cryosections were cut and blocked for 2 h in 3% BSA in PBS+ 0.5% Triton and immunostained with the following antibodies: GFAP (dilution 1:400)(Abcam), PDGFRβ (a gift from Dr. W. Stallcup from Sanford-Burnham Medical Research Institute, La Jolla, CA), CD41 (clone MWReg30) (eBioscience), CD48 (clone HM48-1) (BioLegend), hematopoietic lineage cocktail (Lin) (eBioscience), NGFR (dilution 1:100)(Abcam), CD31-FITC (dilution 1:100) (BioLegend), CD31‐PE (dilution 1:100) (BioLegend), TUBB3-AlexaFluor-488 (dilution 1:100) (BioLegend), Ki67 (dilution 1:100) (BD Biosciences), and secondary antibodies conjugated with AlexaFluor-488, AlexaFluor-594 and AlexaFluor‐647 (1:1000) (Life Technologies). After this, the sections were washed with PBS containing 4',6-diamidino-2-phenylindole (DAPI, 5 μg/mL, Invitrogen) and mounted using Dako fluorescence mounting medium (Dako, Santa Clara, CA). Stained tissue sections were imaged and analyzed by confocal microscopy using an inverted Zeiss LSM 880 confocal microscope (Oberkochen, Germany) or using a ZEISS AXIO examiner D1 microscope (Zeiss) with a confocal scanner unit, CSUX1CU (Yokogawa), and reconstructed in three dimensions with Slide Book software (Intelligent Imaging Innovations). CD31 area, vessel diameter and length and number of Ki67+ cells were quantified using Fiji software®, version 1.53 (National Institute of Health). Multiple random fields of each section were used for quantification.
Tumor-infiltrating leukocytes immunophenotyping and intracellular cytokine measurement
Tumor tissues were dissociated and filtrated trough cell strainers of 40 µm (Falcon) to isolate the cells used for immunophenotyping. Cells were washed in phosphate-buffered saline (PBS), incubated with Live/Dead solution (Invitrogen), for dead cell exclusion, and with monoclonal antibodies, washed, fixed, and permeabilized (FoxP3 staining buffer set, eBioscience) according to manufacturer’s instructions. Antibodies are listed in Supplementary Table 1. Acquisition was realized on a LSR-FORTESSA. For analyses, to exclude debris, combinations of fluorochromes was done, to remove doublets a forward scatter area (FSC-A) versus forward scatter height (FSC-H) gate was used, and then cells were gated in function of time versus FSC-A to avoid a possible interference of flux interruptions. Only live leukocytes were used using a Live/Dead gate versus CD45. We gated T-cell subpopulations based on molecular markers of each subset (CD4, CD8, γδ, NKT, Treg and NK cells). In each T-cell subset, frequencies of cells expressing checkpoint inhibitors CTLA-4 and PD1 were evaluated. Tumor-infiltrating leukocytes were stimulated with autologous tumor cells for 4 h in the presence of Brefeldin A (ThermoFisher) and Monensin (ThermoFisher). Tumor-infiltrating leukocytes were stained with mAbs specific for surface proteins prior to fixation and permeabilization. Permeabilized cells were then stained with anti-IFN-γ and anti-IL-17 [79]. Ki-67 is a nuclear factor transcript in the late G1, S, G2, and M of cell cycle, therefore marks proliferating cells [71,72,73,74,75,76,77,78,79,80,81,82]. Thus, we evaluated proliferation in viable CD45 negative cells, suggesting tumoral proliferation. GraphPad Prism 8.0 software (GraphPad Software, San Diego, CA) and FlowJo V10.4.11 (TreeStar) were used for data analysis and graphic presentation.
In silico analysis
To investigate our findings in human tumors, we directly obtained gene count reads of 103 SKCM and 495 PRAD tumor samples from the TCGA (https://portal.gdc.cancer.gov/) data repository. For the remaining analyses, gene count reads (expression levels) were first normalized to transcript per million (TPM) using R. Boxplots were created using the R package ggplot2 and Wilcoxon tests between groups were performed using the R package ggpubr. To determine whether the expression levels of GFAP and PLP1 were correlated with overall survival of SKCM and PRAD patients, we stratified patients into groups with high/low expression of GFAP and PLP1 based on ROC-established optimal cutoffs. Kaplan–Meier survival curves were created in R using packages ggpubr, survminer and survival. To estimate immune infiltrated cells in SKCM and PRAD tumor samples, we applied the CIBERSORT tool [83], which uses a support vector regression method combined with prior knowledge of single-cell expression profiles (gene signatures) to produce an estimation of the abundances of immune infiltrated cells subpopulations in a RNA sequencing sample. Protein–protein interaction analysis of the manually curated set of 27 genes expressed in Schwann cells [39, 75,76,77,78,79,80,81,82,83,84,85,86,87,88,89,90,91,92,93,94,95,96] was performed in Cytoscape [97] using the STRING database [98]. To evaluate differential gene expression in SKCM versus healthy skin, we additionally obtained gene count reads of 701 healthy skin samples from the GTEx (https://gtexportal.org/) data repository. Differential gene expression was performed using DESeq2 [99] and only genes presenting an absolute log2(FoldChange) ≥ 2 and FDR adjusted p-value < 0.05 were considered as differentially expressed. To evaluate the functional role of differentially expressed genes, we performed a Gene Ontology enrichment analysis using ShinyGO [100] and REVIGO [101]. Only functional terms with a FDR < 0.01 (hypergeometric test) were considered relevant. Heatmaps were created using the R package pheatmap.
Single-cell RNA sequencing reanalysis
Publicly available scRNA -seq data for prostate tumors was downloaded from GEO with the accession number GSE141445 (PMID: 33420488) [102]. We reanalyzed 36,424 single-cells from 13 prostrate tumors using R package Seurat (version 4.0.3) (PMID: 34062119). Data normalization, scaling, transformation, clustering, dimensionality reduction, differential expression analysis and visualization was done using Seurat package. The cells were clustered by Shared nearest neighbor (SNN) at 0.8 resolution using FindClusters() function and visualized by UMAP using the top 10 principal components (PCs). Barcodes for fibroblast cells was downloaded from (http://www.pradcellatlas.com/#/), the web interface for GSE141445 dataset. Cells expressing the gene Proteolipid protein 1 (PLP1) was identified on the basis of expression > 0. Differential expression analysis was performed between PLP1+ and PLP1− cells using Wilcoxon Rank Sum test (FindMarkers()) function (Suppl. Table 2). In order to include maximum number of differentially expressed genes we used average logfc.threshold = 0 and without any percentage cutoff for features that are detected in either PLP1+ and PLP1−.
We used sequenced transcriptomes from single-cell triple-negative breast cancers (TNBC) [103] and melanoma [104] to identify the differentially expressed genes (DEGs) in cancer-associated fibroblasts (CAFs) PLP1+ and PLP1−. A total of 24,271 breast cancer cells and 4645 melanoma cells were analyzed. We obtained the expression matrix of the breast cancer and melanoma dataset from singlecell.broadinstitute.org. For the breast cancer dataset we normalized, scaled, and identified markers from populations of interest using the Seurat pipeline (Suppl. Table 3). For the melanoma dataset, available already normalized we used the limma pipeline to perform differential expression analysis (Suppl. Table 4). The identified markers should be present in more than 12.5% of the evaluated populations and present a p-value corrected for multiple tests based on bonferroni correction using all genes in the dataset ≤ 0.05.
Statistical analysis
The Shapiro–Wilk test was performed to verify the normality of the data. For parametric data unpaired Student's t-tests was used to identify differences between two independent groups. One-way repeated-measures analyses of variance (ANOVAs) was used for comparisons with multiple observations overtime of dependent groups (i.e., for the quantification of GFAP+ cells over 21 days), and two-way ANOVA was used to evaluate the interaction between different cell types and experimental groups (Plp1CreER−/iDTR+ or Plp1CreER+/iDTR+ mice) in cytokine production. When a significant F value was found, we performed Student–Newman–Keuls tests as post hoc analyses. For non-parametric data, the Mann–Whitney test (rank-sum test) was applied for comparisons across two-time points between two independent groups. The α level was set at 0.05. Data are shown as mean ± standard error (SEM). All statistical analyzes were performed using the GraphPad Prism 8.0 software (GraphPad Software, San Diego, CA). Cohen’s d effect size (ES) was also calculated. The ES allowed the assessment of the magnitude of the differences between the collected data points. For comparisons between two groups, ES was calculated by subtracting the mean value of one data point from the mean value of the other data point to which it was compared; the result was then divided by a combined SD of the data. These analyses were performed using the GPower version 3.1 (Universität Düsseldorf, Germany). The ES for ANOVAs was calculated using the equation η2 = Effect SQ/Total SQ; SQ = sum of squares. The η2 values were converted into d values [105]. ES values were classified as trivial (< 0.2), small (0.2–0.6), medium (0.6–1.2), or large (≥ 1.2) [106].
Results
Cells expressing GFAP, a Schwann cell marker, are present within the tumor microenvironment surrounding tumoral blood vessels
We and others have previously demonstrated that solid tumors are infiltrated by different nerve fibers [8,9,10,11,12,13,14,15,16, 107, 108]. In physiologic normal conditions, peripheral axons present Schwann cells attached to them [109]. In contrast, within the tumor microenvironment, we did not detect any Schwann cells attached to the innervations. Instead, we found cells expressing the GFAP, a Schwann cell marker, within the tumors. We have injected PC-3 human prostate cells into the ventral prostate of immunodeficient Balb/c nude (nu/nu) mice (Fig. 1A). Our analysis, 3 weeks after injection, has revealed the presence of GFAP+ cells infiltrating the tumor (Fig. 1B). Interestingly, most of GFAP+ cells were associated with blood vessels (86.8 ± 8.8% of GFAP+ cells; Fig. 1C), not nerves. Although nerve fibers are in a close proximity to blood vessels within the tumor microenvironment, most of intra-tumoral blood vessels are not associated with innervations (Suppl. Fig. 1). To test whether GFAP+ cells presence in the tumor microenvironment is specific to the immunosuppressed microenvironment of PC-3 tumors, we have also used a syngeneic prostate tumor mouse model. We injected Tramp-C2 mouse prostate cancer cells into the ventral prostate of immunocompetent C57BL/6 mice (Suppl. Fig. 2A). Similarly to what we found in the immunodeficient tumors, after 2 weeks, we detected the presence of blood vessel-associated GFAP+ cells in Tramp-C2 tumors (Suppl. Fig. 2B, C). We also transplanted RM1 mouse prostate cancer cells intra-prostatically (from Ras+ Myc transformed mouse prostate carcinoma) into the ventral prostate of immunocompetent C57BL/6 mice (Suppl. Fig. 2D). Similarly, after 2 weeks, we detected GFAP+ cells associated with intra-tumoral blood vessels (Suppl. Fig. 2E). Furthermore, GFAP+ cells are also present in the human prostate tumor microenvironment, as revealed by immunohistochemistry of radical prostatectomy specimens from patients with adenocarcinoma (58.71 ± 25.05 GFAP+ cells/mm2 of tumor area) (Fig. 1D).
Glial Fibrillary Acidic Protein (GFAP)+ cells are present in the prostate tumor microenvironment associated with blood vessels. A Intra-prostatic injection of TdTomato-labeled PC-3 human prostate cancer cells in nude mice, and tumor analysis after 3 weeks. B Representative photomicrographs of a prostate tumor sections 3 weeks after PC3 cells injection, showing blood vessels with GFAP+ cells attached to it. C Percentages of GFAP+ cells attached or not to blood vessels in the PC3 prostate tumor after 3 weeks (n = 3 mice) (86.8 ± 8.8% of GFAP + cells were associated to blood vessels, while 13.3 ± 8.1% were not associated to blood vessels; p = 0.0042; ES = 4.86L). BV blood vessel. D Representative image showing the presence of GFAP+ cells in human prostate adenocarcinoma (58.71 ± 25.05 GFAP+ cells/mm2 of tumor area). Statistical analysis: unpaired Student's t-tests. ES effect size; Llarge (≥ 1.2). **p < 0.01. Data are mean ± SEM. Scale bars, 10 µm
Tumor-infiltrating GFAP+ cells differ from macrophages, hematopoietic and endothelial cells, express immature Schwann cell markers and are in proliferative state
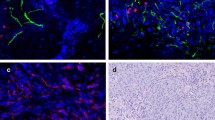
To define the identity of GFAP+ cells in the tumor microenvironment, we analyzed the expression of other molecular markers in these cells. As hematopoietic cells and tissue-resident macrophages have been shown to be located in the perivascular position under certain pathophysiologic conditions [101,102,103,104,105,106,107,108,109,110,111,112,113,114,115,116,117], we evaluated the expression of markers specific for these cells within the tumor microenvironment. By immunohistochemistry, we found that GFAP+ cells in the Tramp-C2 tumor microenvironment did not express CD31, a marker for endothelial cells. They also differed from macrophages and hematopoietic cells, as they did not express the macrophage marker F4/80 [also known as EMR1 in humans [118]], or hematopoietic lineage markers (Lin, CD41 and CD48) (Fig. 2A–D). Next, we evaluated, in the tumor microenvironment, the expression of markers characteristic of activated Schwann cells, such as p75 (NGFR) and Nestin-GFP [110,111,112,113,114,115,116,117,118,119,120,121,122,123,124] (Fig. 2E–H). In Tramp-C2 tumors, all GFAP+ cells were Nestin-GFP+ (Fig. 2G, H). By immunohistochemistry, we found that GFAP+ cells in the Tramp-C2 tumor microenvironment express p75 (NGFR) (Fig. 2E, F). Thus, our results suggest that intra-tumoral GFAP+ cells are similar to Schwann cells, differing from those in their anatomical location, attached to blood vessels instead of nerves.
GFAP+ cells differ from macrophages and hematopoietic cells, and express NGFR and Nestin-GFP in the prostate tumor microenvironment. Analysis of prostate tumor sections 2 weeks after Tramp-C2 cells injection. A Representative photomicrograph of Tramp-C2 tumor showing macrophages (F4/80+) and GFAP+ cells. B Percent of GFAP+ cells that co-express F4/80 (n = 3 mice) (100% of GFAP+ cells were negative for F4/80; p < 0.0001; ES > 1000L). C Representative prostate tumor section showing hematopoietic (Lin+CD41+CD48+) and GFAP+ cells. D Quantification of the data illustrated in C (n = 3) (99.27 ± 0.37% of GFAP+ cells were negative for Lin/CD41/CD48; p < 0.0001; ES = 266.3L). E Representative photomicrographs of a prostate tumor sections 2 weeks after Tramp-C2 cells injection, showing GFAP+ cells expressing NGFR. F Percentage of GFAP+ cells expressing NGFR in the Tramp-C2 prostate tumor after 2 weeks (n = 3 mice) (99.27 ± 0.3% of GFAP+ cells were positive for NGFR; p < 0.0001; ES = 153.2L). G Percentages of NGFR+ /GFAP+ cells attached or not to blood vessels in the Tramp-C2 prostate tumor after 2 weeks (n = 3 mice) (86.30 ± 7.0% of NGFR+/GFAP+ cells were associated to blood vessels, while 13.7 ± 7.1% were not associated to blood vessels; p = 0.0019; ES = 10.4L). BV blood vessel. H Representative image of orthotopic Tramp-C2 tumor in Nestin-GFP mice. All panels show the same area for different channels (GFAP, CD31, Nestin-GFP, and all the images merged). I Percentage of GFAP+ cells co-expressing Nestin-GFP (n = 3 mice) (99.6 ± 0.30% of GFAP+ cells were Nestin-GFP + ; p < 0.0001; ES = 172.1L). J Percentages of Nestin-GFP+/GFAP+ cells attached or not to blood vessels in the Tramp-C2 prostate tumor (n = 3 mice) (93.6 ± 3.1% of Nestin-GFP+/GFAP+ cells were associated to blood vessels, while 6.4 ± 3.1% were not associated to blood vessels; p < 0.0001; ES = 17.4L). Statistical analysis: unpaired Student's t-tests; ES effect size; Llarge (≥ 1.2). **p < 0.01; ***p < 0.001. Data are mean ± SEM. BV blood vessel. Scale bars, 10 µm
Next, to evaluate whether GFAP+ cells are infiltrating pro-actively in the tumor microenvironment or whether they are just passive cells which get surrounded by the growing tumor, we evaluated the presence of GFAP+ cells within the tumor at different time points. We discovered that after 4 days of Tramp-C2 cancer cells orthotopic transplantation, endothelial cells (CD31+) appear, while GFAP+ cells are still absent from the prostate tumor microenvironment. GFAP+ cells only start appearing in the tumor microenvironment after day 6 of cancer cells transplantation, achieving a peak in between day 7 and 14 of tumor growth (Fig. 3A–C). These data indicate that GFAP+ cells are pro-actively infiltrating within the tumor during cancer development. To explore whether GFAP+ cells are actively proliferating within the tumor microenvironment, we have analyzed human PC-3 prostate cancer xenograft mouse model (Fig. 4A). Our analysis at 3 weeks post-transplantation of PC3 cancer cells has revealed that most of intra-tumoral GFAP+ cells are proliferating, as 61.33 ± 4.7% of GFAP+ cells stained for Ki67, a marker of proliferation [80] (Fig. 4B, C). We also found that proliferating GFAP+ cells are not specific to prostate tumor microenvironment, as we found these cells also within the breast tumor microenvironment. We detected the presence of proliferating GFAP+ cells in breast cancer samples from 8-week-old BRCA1− and p53-deficient mice (52.3 ± 4.3% of GFAP+ cells were Ki67+ and 47.67 ± 4.3% were Ki67−) (Fig. 4D, E and F). Altogether, our results suggest that cells expressing Schwann cells’ markers infiltrate pro-actively within the primary tumor and proliferate during cancer development.
GFAP+ cells appearance in the prostate tumor microenvironment. A Representative photomicrographs of prostate tumor sections 4 and 7 days after Tramp-C2 cells orthotopic injection into Nestin-GFP mice. B The areas in the white boxes in A are magnified showing GFAP+ cells attached to tumoral blood vessels. C Quantification of the number of GFAP+ cells per tumor area during prostate tumor growth. Notice that, at day 4, GFAP+ cells are still not present in the tumor microenvironment; and the peak of appearance of GFAP+ cells is between 6 and 14 days (Days 4, 4.5 and 5: 0 GFAP + cells/mm2; day 6: 104.4 ± 24.3 GFAP+ cells/mm2; day 7: 186.9 ± 40.8 GFAP+ cells/mm2; day 14: 142.4 ± 8.9 GFAP + cells/mm2; and day 21: 35.6 ± 8.9 GFAP + cells/mm2; p < 0.001; ES = 4.0L). Statistical analysis: One-way repeated-measures analyses of variance [225] and post hoc Student–Newman–Keuls. ES effect size; Llarge (≥ 1.2). ***p < 0.001; **p < 0.01; *p < 0.05. n = 3 mice. Data are mean ± SEM. Scale bars, 10 µm. D Table showing tumor size related to the density of tumor-infiltrating GFAP+ cells
Tumoral GFAP+ cells proliferate within the tumor microenvironment. A Intra-prostatic injection of TdTomato-labeled PC-3 human prostate cancer cells in nude mice, and analysis of tumoral GFAP + cells after 3 weeks. B The percentage of tumoral GFAP+ cells expressing Ki67 was quantified (n = 3 mice) (61.33 ± 4.7% of GFAP+ cells were Ki67+ and 38.67 ± 4.7% were Ki67-); p = 0.027; ES = 4.8L). C Representative photomicrographs of a prostate PC-3 tumor section show GFAP+ cells labeled with Ki67 (white arrows). All panels show the same area for different channels (GFAP, Ki67, PC-3-TdTom, DAPI, and GFAP merged with Ki67). D Mammary tumors from 8-week-old BRCA1- and p53-deficient mice were surgically removed for analysis. E The percentage of tumoral GFAP + cells expressing Ki67 was quantified (n = 3) (52.3 ± 4.3% of GFAP+ cells were Ki67+ and 47.67 ± 4.3% were Ki67-; p = 0.071; ES = 1.2L). F Proliferating GFAP+ cells are present in the mammary tumor microenvironment. Representative photomicrographs of the mammary BRCA1- and p53-deficient tumor section show GFAP+ cells labeled with Ki67 (white arrow). Data are mean ± SEM. Scale bars, 10 µm. Statistical analysis: unpaired Student's t-tests. ES effect size; Llarge (≥ 1.2). *p < 0.05 Significant difference between groups
Tumor-infiltrating perivascular GFAP+ cells derive exclusively from tissue-resident Schwann cells
To evaluate the origin of intra-tumoral GFAP+ cells, we have tracked the fate of distinct cell populations. Previous studies have suggested that some cancer cells may dedifferentiate into a glial phenotype [116,117,118,119,120,121,122,123,124,125,126,127,128,129]. To test whether those tumor-infiltrating GFAP+ cells derive from cancer cells, we transplanted Tramp-C2 mouse prostate cancer cells permanently labeled with TdTomato into the ventral prostate of immunocompetent C57BL/6 mice (Fig. 5A–D). After 2 weeks, Tramp-C2 tumor cells marked with TdTomato fluorescence did not express GFAP (Fig. 5C, D), indicating that cancer cells do not originate tumor-infiltrating GFAP+ cells.
GFAP+ cells do not arise from cancer cells in the tumor microenvironment. A Generation of genetically engineered Tramp-C2 cancer cells stably expressing TdTomato. Tramp-C2 cells were transduced with lentiviral vectors encoding TdTomato under the control of the cytomegalovirus promoter. Tramp-C2 cells expressing the construct were selected by double FACS sorting. Scale bar, 20 µm. B Intra-prostatic injection of TdTomato-labeled Tramp-C2 murine prostate cancer cells into wild-type mice, and tumor analysis after 2 weeks. C Percentage of Tramp-C2 TdTom+ cells that express GFAP (n = 3 mice) (99.7 ± 0.3% of TdTom+ cells were negative for GFAP; p < 0.0001; ES = 298.3L). D Representative photomicrographs of a Tramp-C2 TdTom+ prostate tumor section, showing GFAP+ cells (TdTom-). Scale bars, 10 µm. Statistical analysis: unpaired Student's t-tests. ES effect size; Llarge (≥ 1.2). ***p < 0.001. Data are mean ± SEM
As subsets of myeloid cells have been reported to surround blood vessels within tumors [130, 131], we tested whether intra-tumoral GFAP+ cells derive from myeloid subsets. To examine whether GFAP+ cells derive from the myeloid lineage, dendritic cells or tissue-resident macrophages, we have transplanted Tramp-C2 mouse prostate cancer cells orthotopically into the ventral prostates of immunocompetent LysM-Cre/TdTomato and CSF1R-Cre/TdTomato mice, in which myeloid lineage, dendritic cells and tissue-resident macrophages are permanently labeled with TdTomato fluorescence (Fig. 6A, B, E and F). In LysM-Cre/TdTomato mice, upon removal of loxP-stop-loxP cassette by Cre recombination, TdTomato is expressed in myeloid lineage-derived cells; while, in CSF1R-Cre/TdTomato mice, with loxP-stop-loxP cassette removed by Cre recombination, TdTomato is expressed in dendritic cells, tissue-resident macrophages, and cells derived from those cells. Our analysis at 2 weeks post-injection has revealed that tumor-infiltrating GFAP+ cells do not derive from myeloid cells or from tissue-resident macrophages, as all TdTomato+ cells in those tumors were negative for GFAP (Fig. 6C, D, G and H).
Tumoral GFAP+ cells are not derived from the myeloid lineage nor from tissue-resident macrophages. A Schematic diagram of the LysM-Cre/TdTom experimental mouse model. Cre-recombinase directs the expression of TdTomato fluorofore in LysM+ cells and all cells derived from those. B Adult LysM-Cre/TdTom mice were orthotopically injected with Tramp-C2 prostate cancer cells. Tumors were surgically removed 2 weeks later for analysis. C Representative image of orthotopic Tramp-C2 tumor in LysM-Cre/TdTom mice. All panels show the same area for different channels (LysM-Cre/TdTom, GFAP, CD31, and all the images merged). D Percentage of LysM-Cre/TdTom+ cells co-expressing GFAP (n = 3 mice). (99.0 ± 0.6% of LysM-Cre/TdTom+ cells were negative for GFAP; p < 0.0001; ES = 167.3L). E Schematic diagram of the CSF1R-Cre/TdTom experimental mouse model. Cre-recombinase directs the expression of TdTomato fluorofore in CSF1R+ cells and all cells derived from those. F Adult CSF1R-Cre/TdTom mice were orthotopically injected with Tramp-C2 prostate cancer cells. Tumors were surgically removed 2 weeks later for analysis. G Representative image of orthotopic Tramp-C2 tumor in CSF1R-Cre/TdTom mice. All panels show the same area for different channels (CSF1R-Cre/TdTom, GFAP, CD31, and all the images merged). H Percentage of CSF1R-Cre/TdTom+ cells co-expressing GFAP (n = 3 mice). (99.2 ± 0.4% of CSF1R-Cre/TdTom+ cells were negative for GFAP; p < 0.0001; ES = 247.5L). Statistical analysis: unpaired Student's t-tests. ES effect size; Llarge (≥ 1.2). ***p < 0.001. Data are mean ± SEM. Scale bars, 10 µm
Since it has been proposed that bone marrow derived cells circulate and home to the tumor microenvironment [132, 133], we asked whether GFAP+ cells are derived from bone marrow cells. Thus, we transplanted irradiated C57BL/6 mice with bone marrow from mT/mG mouse in which all cells express TdTomato. After 1 month, we transplanted Tramp-C2 prostate cancer cells into the ventral prostate of the resulting chimeras (Fig. 7A). We found, at 4 weeks post-injection, that cells marked with TdTomato fluorescence did not express GFAP (Fig. 7B, C), suggesting that GFAP+ cells do not derive from bone marrow cells. While bone marrow transplantation is a standard approach to trace the fate of circulating cells, irradiation injury may affect the microenvironment of the tissue where the tumor grows. Therefore, we also utilized parabiosis, by conjoining two mice to share single blood circulation, to ask whether GFAP+ cells in the tumor microenvironment derive from circulating cells. Thus, mT/mG mice were conjoined with wild-type mice. Shared circulation was verified 4 weeks after parabiosis surgery, and Tramp-C2 cancer cells injection was performed in the prostate of the wild-type parabiont (Fig. 7D). After 4 weeks, microscopic evaluation failed to detect any GFAP+ cells that expressed TdTomato, despite robust presence of GFAP−/TdTomato+ cells and GFAP+ /TdTomato− cells in the tumor microenvironment (Fig. 7E, F). These experiments provide clear evidence that neither bone marrow derived nor circulating cells contribute to the appearance of GFAP+ cells within the tumor microenvironment, indicating that tumor-infiltrating GFAP+ cells are derived from tissue-resident cells.
Tumoral GFAP+ cells are not derived from circulating cells. A Bone marrow transplantation scheme. Bone marrow from genetically labeled mT/mG donors (in which all cells display TdTomato fluorescence at the membrane) was transplanted into lethally irradiated, unlabeled wild-type recipients. After engraftment was verified (4 weeks), recipients were injected orthotopically with Tramp-C2 prostate cancer cells. Data were assessed by confocal microscopy analysis of the prostate tumors. B, C Almost no GFAP + cells with TdTomato fluorescence were detected in the recipients’ tumors 4 weeks after cancer cells injection (n = 5 mice) (99.67 ± 0.3% of TdTom+ cells were negative for GFAP; p < 0.0001; ES = 85.4L). D Parabiosis experimental design. Genetically labeled mT/mG mice were conjoined with unlabeled wild-type mice. Shared circulation was verified from the wild-type parabiont after 4 weeks. Thereafter, the wild-type parabionts underwent orthotopic transplantation of Tramp-C2 prostate cancer cells. E, F Tumoral GFAP+ cells are not coming from the circulation. No TdTomato+ cells expressing GFAP were detected in the tumor of the wild-type parabionts, despite robust presence of TdTomato+ not expressing GFAP cells in the parabiont tumor microenvironment (n = 5 mice) (99.97 ± 0.03% of TdTom+ cells were negative for GFAP; p < 0.0001; ES = 108.4L. Statistical analysis: unpaired Student's t-tests. ES effect size; Llarge (≥ 1.2). ***p < 0.001 Significant difference between groups. Data are mean ± SEM. Scale bars, 10 µm
To understand which cells resident in the tissue give origin to tumor-infiltrating perivascular GFAP+, as these cells resemble Schwann cells, we tested whether intra-tumoral GFAP+ cells derive from nerve-associated Schwann cells. For this purpose, we transplanted Tramp-C2 mouse prostate cancer cells orthotopically into the ventral prostate of immunocompetent Plp1-CreER/TdTomato mice pre-treated with tamoxifen (Fig. 8A, B). In those mice, after tamoxifen administration, upon removal of loxP-stop-loxP cassette by Cre recombination, TdTomato is expressed only in nerve-associated Schwann cells and cells derived from Schwann cells in the peripheral tissues. Our analysis at 2 weeks post-injection has revealed that tumoral GFAP+ cells derive from tissue-resident Schwann cells (98.6 ± 0.3% of Plp1CreER/TdTom+ cells were positive for GFAP; p < 0.0001) (Fig. 8C, D). We also analyzed earlier stages of carcinogenesis in a genetically engineered mouse model of prostate cancer (Hi-Myc mice [8]) (Fig. 9). Our analysis of prostate from 20 weeks-old Hi-Myc mice has revealed the presence of GFAP+ cells at the site of disruption of the prostate acini, during tumor initiation, detaching from the nerves, but not yet attached to blood vessels (Fig. 9A, B). To evaluate whether this is specific to prostate cancer, we transplanted B16F10 mouse melanoma cells subcutaneously into immunocompetent Plp1−CreER/TdTomato mice pre-treated with tamoxifen (Fig. 8E). Our analysis at 2 weeks post-injection has revealed, similarly to what we observed in the prostate tumor model, that tumoral GFAP+ cells derive from tissue-resident Schwann cells (98.3 ± 1.1% of Plp1CreER/TdTom+ cells were positive for GFAP; p < 0.0001) (Fig. 8F, G and H).
Tumoral GFAP+ cells derive from tissue-resident Schwann cells. A Schematic diagram of the Plp1-CreER/TdTom experimental mouse model. Tamoxifen-inducible Cre-recombinase directs the expression of TdTomato fluorofore specifically to Schwann cells in those mice. After the administration of tamoxifen to those mice, all Schwann cells and cells derived from those are labelled by TdTomato. B Adult Plp1−CreER/TdTom mice 48 h after tamoxifen administration were orthotopically implanted with Tramp-C2 cancer cells. Tumors were surgically removed 2 weeks later for analysis. C Percentage of Plp1CreER/TdTomato+ cells that express GFAP within Tramp-C2 tumors (n = 3 mice) (98.6 ± 0.3% of Plp1CreER/TdTom+ cells were positive for GFAP; p < 0.0001; ES = 277.14L). D Plp1CreER/TdTomato + cells overlap with GFAP+ cells in the tumor microenvironment. E Adult Plp1-CreER/TdTom mice 48 h after tamoxifen administration were subcutaneously injected with B16F10 cancer cells. Tumors were analysed after 4 weeks. F Representative FACS plot showing the percentage of Plp1CreER/TdTom+ cells isolated from melanoma tumors grown in Plp1CreER/TdTomato mice. G Percentage of Plp1CrER/TdTomato+ cells that express GFAP within B16F10 tumors (n = 3 mice) (98.3 ± 1.1% of Plp1CreER/TdTom+ cells were positive for GFAP; p < 0.0001; ES = 97.7L). H Representative photomicrographs of a melanoma tumor section showing Plp1-TdTomato + cells expressing NGFR and GFAP. Statistical analysis: unpaired Student's t-tests. ES: effect size; Llarge (≥ 1.2). ***p < 0.001. Data are mean ± SEM. Scale bars, 10 µm
Presence of GFAP+ cells in the Hi-Myc tumor microenvironment. A Representative immunofluorescence images of prostate from 20 weeks-old Hi-Myc mice. All panels show the same area for different channels (α-SMA, GFAP, DAPI, α-SMA merged with DAPI, GFAP merged with DAPI, α-SMA merged with GFAP, and all the images merged). Note the presence of GFAP+ cells detached from their location at the site of disruption of the prostate acini, during tumor initiation (white arrows). Scale bar, 10µm. B Schematic diagram ilustrating GFAP+ cells (red arrow) at the site of disruption of the prostate acini, during tumor initiation as in (A)
As pericytes are also located in a perivascular position within tumors [23, 134,135,136,137,138,139,140,141,142,143,144], we evaluated whether perivascular glial cells were different from pericytes. We found that perivascular glial cells corresponded to 23.60 ± 4.69% of NG2 proteoglycan-expressing cells within the tumor microenvironment (Suppl. Fig. 3A, B). As NG2 proteoglycan was previously reported to be expressed also in glial cells [145], we also analyzed the expression of another pericyte marker (PDGFRβ) [146,147,148,149,150,151]. We found that all tumor-infiltrating perivascular glial cells were negative for PDGFRβ (Suppl. Fig. 3C, D), indicating that tumor-infiltrating perivascular glial cells differ from intra-tumoral pericytes. The intra-tumoral blood vessels within the melanoma microenvironment were not innervated, but presented Plp1−CreER+/TdTomato+/GFAP+ cells attached to them (Fig. 10). Thus, our data indicate that Schwann cells associated to nerves detach from the innervations during tumor progression, and associate with newly formed tumoral blood vessels (Fig. 11)
Plp1+ cells are present within the melanoma microenvironment surrounding not-innervated blood vessels. A Schematic diagram of the Plp1-CreER/TdTom experimental mouse model. Tamoxifen-inducible Cre-recombinase directs the expression of TdTomato fluorofore specifically to Schwann cells in those mice. After the administration of tamoxifen to those mice, all Schwann cells and cells derived from those are labelled by TdTomato. B Adult Plp1-CreER/TdTom mice 48 hours after tamoxifen administration were subcutaneously injected with B16F10 cancer cells. Tumors were analyzed after 4 weeks. C Representative photomicrographs of a melanoma tumor section showing Plp1CreER+/TdTomato+ cells (red) attached to CD31+ endothelial cells (pink) within the melanoma microenvironment. Notice that these intra-tumoral blood vessels are not innervated as they were negative for the pan-neuronal marker class III β tubulin (TUBB3) (green). D Percentages of Plp1CreER+/TdTomato+ cells attached or not to blood vessels in the B16F10 melanoma tumor after 4 weeks (n=3 mice) (89.7 ± 5.4 % of Plp1CreER+/TdTomato+ cells were associated to blood vessels, while 10.0 ± 5.7 % were not associated to blood vessels; p < 0.0001; ES = 8.7L). BV= blood vessel. E Representative photomicrographs of a tumor-infiltrating perivascular glial cell showing the distance to Schwann cells attached to the nerve composed by multiple innervations which stained positive for the pan-neuronal marker class III β tubulin (TUBB3). F Schematic illustration of intra-tumoral Plp1+ cells attached to blood vessels which are not innervated. Scale bars, 10µm
.
Genetic ablation of endogenous Schwann cells promotes tumor regression
After defining Plp1+ Schwann cells as the origin of tumor-infiltrating GFAP+ cells, we next sought to understand their role in cancer progression. To explore the role of endogenous Schwann cells within the tumor microenvironment, we generated mice in which Schwann cells could be postnatally targeted in an inducible manner by DT-based cell ablation.
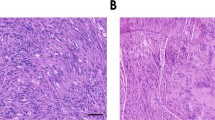
To specifically deplete endogenous Schwann cells, diphtheria toxin receptor (iDTR) floxed mice were mated with mice expressing tamoxifen-inducible Cre-recombinase driven by the proteolipid protein 1 (Plp1) promoter to generate Plp1−CreER+ /iDTR+ mice [64, 152]. In these animals, upon administration of tamoxifen and DT, DTR, expressed specifically in Schwann cells, binds to DT and promotes its endocytosis. Upon entry into the cytoplasm, DT induces apoptotic death of the targeted Schwann cell by catalyzing the inactivation of elongation factor 2, thereby halting global protein synthesis [153, 154]. DT-mediated cell ablation is highly sensitive and efficient, as a single molecule of active DT in the cytoplasm is sufficient to kill an eukaryotic cell [155] (Fig. 12A). Littermates heterozygous for iDTR but lacking the Plp1−CreER expression were used as controls (Plp1−CreER−/iDTR+mice). To evaluate the role of endogenous Schwann cells on tumor growth, we transplanted subcutaneously B16F10 melanoma cells to the lower right flank of both Schwann cell-ablated mice (Plp1−CreER+/iDTR+) and their controls (Plp1−CreER−/iDTR+). Before melanoma cells implantation, mice were treated with tamoxifen and DT to eliminate Plp1+ Schwann cells (controls were also treated with tamoxifen and DT) (Fig. 12B). These experiments revealed that after 16 days of cancer cells’ transplantation, melanoma development was decreased in the Schwann cell-depleted mice when compared to the controls (tumor weight reduced from 0.89 ± 0.23 to 0.29 ± 0.06 g; tumor weight per body weight reduced from 0.03 ± 0.01 to 0.01 ± 0.002; tumor area reduced from 2.13 ± 0.48 to 0.95 ± 0.10 cm2, the tumor volume reduced from 2.76 ± 1.07 to 0.36 ± 0.05 mm3; Fig. 12C–G). Animal weights were not affected by genetic ablation of Schwann cells in melanoma‐bearing mice (data not shown). Moreover, genetic ablation of Schwann cells led to a decrease in proliferating cells within the tumor (from 27.33 ± 6.88 to 8.00 ± 3.51% of proliferating Ki67+ cells within the tumor, analyzed by immunohistochemistry) (Fig. 12H, I), corroborated by flow cytometry analysis of CD45−cells for Ki67 expression (there was a decrease from 59.20 ± 5.50 to 31.30 ± 5.55% of CD45−/Ki67+ cells within the population of CD45−cells) (Fig. 12J). Additionally, there was a decrease in the intra‐tumoral blood vessels’ area (from 0.31 ± 0.01 to 0.20 ± 0.02 µm2 of CD31+ area/µm2 of tumor area) diameter (from 21.05 ± 1.16 to 14.09 ± 3.04 µm) and length (from 148.5 ± 18.3 to 101.8 ± 11.59 mm/µm2; Fig. 12K–N). Our data suggest that depletion of Schwann cells counteracts melanoma development.
Genetic ablation of Plp1 + Schwann cells decreases melanoma growth. A Schematic diagram of the Plp1−CreER+/iDTR+ experimental mouse model. Tamoxifen-inducible Cre-recombinase directs the expression of DTR specifically to Schwann cells in those mice. After the administration of diphteria toxin to those mice, cell death is induced specifically in Schwann cells. B Representation of the protocol for subcutaneous allograft melanoma growth. Plp1−CreER−/iDTR+ (n = 6 mice) and Plp1−CreER+/iDTR+ (n = 9 mice) mice were treated for 5 days with 2 mg/day of Tamoxifen diluted in sunflower oil. After 8 days of chase mice were treated with two consecutive doses of 2 μg of Diphteria Toxin (DT). Mice rested for 4 days before being injected with 1 × 105 B16F10 melanoma cells subcutaneously. Tumors were surgically removed for analyses after 16 days. C Representative macroscopic images of B16F10 melanoma tumors after dissection, left panel (Plp1-CreER−/iDTR +) and right panel (Plp1-CreER + /iDTR +). D Tumor weight (Plp1−CreER−/iDTR+: 0.89 ± 0.23 g; Plp1−CreER+/iDTR+: 0.29 ± 0.06 g; p = 0.012; ES = 1.4L). E Tumor weight corrected by animal body weight (Plp1−CreER−/iDTR+: 0.03 ± 0.009; Plp1−CreER+/iDTR+: 0.01 ± 0.002; p = 0.0276; ES = 1.2L). F. Tumor area (Plp1−CreER−/iDTR+: 2.13 ± 0.48 cm2; Plp1−CreER+/iDTR+: 0.95 ± 0.10 cm2; p = 0.0136; ES = 1.3L). G. Tumor volume (Plp1−CreER−/iDTR+: 2.76 ± 1.069 mm3; Plp1−CreER+/iDTR+: 0.36 ± 0.05 mm3; p = 0.0076; ES = 1.3L). H Representative immunofluorescence images of melanoma sections from Plp1−CreER−/iDTR+ and Plp1−CreER+/iDTR+ mice labelled for Ki67 (Ki67; green) to identify cell proliferation and nuclei (DAPI; blue). I Quantification of proliferation in melanomas from Plp1−CreER−/iDTR+ and Plp1−CreER+/iDTR+ animals (Plp1−CreER−/iDTR+: 27.33 ± 6.88% of Ki67 + cells in total cells; Plp1−CreER+/iDTR+: 8.00 ± 3.51% Ki67 + cells in total cells; p = 0.033; ES = 2.0L). J Quantification of proliferation (Ki67 +) by flow cytometry in CD45−cells from tumors of Plp1−CreER−/iDTR+ and Plp1−CreER+/iDTR+ mice (Plp1−CreER−/iDTR+: 59.20 ± 5.5% of Ki67 + cells in CD45−cells; Plp1−CreER+/iDTR+: 31.30 ± 5.55% of Ki67 + cells in CD45−cells; p = 0.0035; ES = 1.8L). K Representative immunofluorescence images of tumor sections from Plp1−CreER−/iDTR+ and Plp1−CreER+/iDTR+ mice labelled for endothelial cells (CD31; red) to identify blood vessels and nuclei (DAPI; blue). L–N Quantification of angiogenesis in melanomas from Plp1−CreER−/iDTR+ and Plp1−CreER+/iDTR+ animals by blood vessel area (Plp1−CreER−/iDTR+: 0.307 ± 0.012 µm2/µm2; Plp1−CreER+/iDTR+: 0.197 ± 0.02 µm2/µm2; p = 0.0095; ES = 1.3L), diameter (Plp1−CreER−/iDTR+: 21.05 ± 1.16 µm; Plp1−CreER+/iDTR+: 14.09 ± 3.04 µm; p = 0.049; ES = 3.1L), and length (Plp1−CreER−/iDTR+: 148.5 ± 18.30 mm/mm2; Plp1−CreER+/iDTR+: 101.8 ± 11.59 mm/mm2; p = 0.048; ES = 1.8L). Statistical analysis: unpaired Student's t-tests. ES effect size; Llarge (≥ 1.2). *p < 0.05 and **p < 0.01. Plp1−CreER−/iDTR+ (n = 6 mice) and Plp1−CreER+/iDTR+ (n = 9 mice). Data are shown as mean ± SEM. Scale bars, 50 µm
Genetic depletion of endogenous Schwann cells affects tumor immunosurveillance
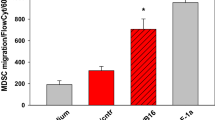
Functional studies in combination with histological analysis have demonstrated that tumor-infiltrating immune cells modulate melanoma cells’ behavior, altering cancer outcomes [156,157,158,159,160,161,162,163,164,165]. Given that Schwann cells are implicated in non-cancer disorders due, in part, to their capacity to impact and modulate immune responses [29, 166,167,168,169], we sought to probe whether the elimination of endogenous Schwann cells alters immune surveillance within the melanoma. Recent breakthroughs in tumor immunotherapy have demonstrated the remarkable capacity of the immune system to fight different types of cancers, including melanoma. The phenotypes and numbers of prevalent tumor-infiltrating lymphocytes are predictive of response to immunotherapy and key modulators of disease progression [79, 170,171,172,173]. Thus, we examined how tumor-infiltrating lymphocytes are affected by the absence of endogenous Schwann cells. We detected an increase in tumor-infiltrating CD4 + T cells (from 1.70 × 107 ± 2.77 × 106 to 4.42 × 107 ± 8.44 × 106 cells per mg of tumor), CD8 + T cells (from 3.09 × 106 ± 1.58 × 106 to 2.87 × 107 ± 6.62 × 106 cells per mg of tumor) (Fig. 13A, E), γδ T cells (from 6.20 × 107 ± 1.47 × 107 to 1.37 × 108 ± 2.54 × 107 cells per mg of tumor), NKT cells (from 1.33 × 107 ± 3.52 × 106 to 2.68 × 107 ± 2.76 × 106 cells per mg of tumor) and NK cells (from 1.16 × 107 ± 3.05 × 106 to 3.40 × 107 ± 5.20 × 106 cells per mg of tumor) (Fig. 14A, E, and I). Lymphocytes are stimulated by dendritic cells to initiate some of their anti-tumor responses within the melanoma microenvironment [174]. We observed an increase in tumor-infiltrating dendritic cells (from 2.72 × 106 ± 7.40 × 105 to 8,60 × 106 ± 2.21 × 106 cells per mg of tumor; Suppl. Fig. 4). In contrast, regulatory T cells, which mediate immunosuppression in the tumor microenvironment [175], were reduced within the tumors (from 4.25 × 106 ± 7.18 × 105 to 1.48 × 106 ± 4.20 × 105 cells per mg of tumor) (Suppl. Fig. 5).
Immune checkpoint molecules, such as cytotoxic T lymphocyte antigen 4 (CTLA-4) and programmed cell death 1 (PD-1), act fine-tuning the intense immune responses that might kill healthy cells [176,177,178]. Their expression in cytotoxic T cells may lead to dysfunction of these cells, affecting their effector function [179, 180]. We found that depletion of Schwann cells prevented the increase of immune checkpoint markers of tumor-infiltrating lymphocytes (Figs. 13 and 14). The percentage of CTLA-4-expressing CD4+ tumor-infiltrating lymphocytes decreased from 50.05 ± 8.19% in Plp1−CreER−/iDTR+ to 22.66 ± 5.21% in Plp1−CreER+ /iDTR+ animals (Fig. 13B); similarly, the percentage of PD-1-expressing CD4+ tumor-infiltrating lymphocytes decreased from 12.25 ± 2.98% in Plp1−CreER−/iDTR+ to 2.62 ± 1.52% in Plp1−CreER+ /iDTR+ mice (Fig. 13C). The percentage of PD-1-expressing CD8+ tumor-infiltrating cytotoxic lymphocytes also decreased from 21.7 ± 9.91% in Plp1−CreER−/iDTR+ to 6.51 ± 1.28% in Plp1−CreER+ /iDTR+ animals (Fig. 13G), while the expression of CTLA-4 did not vary in these cells (Fig. 13H). The percentage of PD-1-expressing γδ T tumor-infiltrating lymphocytes decreased from 27.58 ± 5.69% in Plp1−CreER−/iDTR+ to 6.52 ± 1.12% in Plp1−CreER+ /iDTR+ animals (Fig. 14C); likewise, the percentage of PD-1-expressing NKT tumor-infiltrating lymphocytes decreased from 3.75 ± 0.74% in Plp1−CreER−/iDTR+ to 2.13 ± 0.51% in Plp1−CreER+ /iDTR+ animals (Fig. 14G). The percentage of CTLA-4-expressing NK tumor-infiltrating lymphocytes decreased from 39.45 ± 4.51% in Plp1−CreER−/iDTR+ to 21.41 ± 2.66% in Plp1−CreER+ /iDTR+ animals (Fig. 14J); similarly, the percentage of PD-1-expressing NK tumor-infiltrating lymphocytes decreased from 9.20 ± 3.69% in Plp1−CreER−/iDTR+ to 1.57 ± 1.12% in Plp1−CreER+ /iDTR+ mice (Fig. 14K). Overall, our data suggest that Schwann cells ablation induces improvement of T cells effector functions within the tumor microenvironment.
It has been reported that CD4+ and CD8+ lymphocytes secreting IL-17 and interferon-γ (IFN-γ) promote melanoma regression [181, 182]. Here, we detected in response to Schwann cells depletion an increase in melanoma-infiltrating IL-17-producing CD4 + T cells (from 2.18 × 107 ± 5.29 × 106 to 6.01 × 107 ± 1.08 × 107 cells per mg of tumor) as well as in melanoma-infiltrating IL-17-producing CD8 + T cells (from 2.58 × 106 ± 1.37 × 106 to 2.54 × 107 ± 9.83 × 106 cells per mg of tumor) and in melanoma-infiltrating IFN-γ-producing CD8 + T cells (from 4.98 × 106 ± 2.17 × 106 to 1.84 × 107 ± 4.76 × 106 cells per mg of tumor) (Fig. 13I). Altogether, our data suggest that Schwann cells genetic ablation alters immune surveillance which may affect melanoma development.
Schwann cells elimination promotes CD4 + and CD8 + T-cell infiltration within the tumor. Tumor-infiltrating lymphocytes from B16F10–inoculated mice were analyzed ex vivo in Plp1-CreER−/iDTR+ (n = 6 mice) and Plp1−CreER+/iDTR+ (n = 9 mice) mice. Absolute number of CD4 + (Plp1−CreER−/iDTR+: 1.70 × 107 ± 2.77 × 106 cells per mg of tumor; Plp1-CreER+/iDTR+: 4.42 × 107 ± 8.44 × 106 cells per mg of tumor; p = 0.0106; ES = 1.5L) (A) and CD8 + (Plp1−CreER−/iDTR+: 3.09 × 106 ± 1.58 × 106 cells per mg of tumor; Plp1−CreER+/iDTR+: 2.87 × 107 ± 6.62 × 106 cells per mg of tumor; p = 0.0037; ES = 1.8L) (E) T cells from the melanomas of B16F10–inoculated mice. Column charts show proportion of CTLA-4 (B, F), PD-1 (C, G) and CTLA-4/PD-1 co-expressing (D, H) CD4 + (upper panel) and CD8 + (lower panel) T cells from tumors of B16F10–inoculated mice. B CTLA-4 + /CD4 + T cells (Plp1−CreER−/iDTR+: 50.05 ± 8.20%; Plp1−CreER+/iDTR+: 22.66 ± 5.21%; p = 0.0108; ES = 1.5L). C PD-1 + /CD4 + T cells (Plp1−CreER−/iDTR+: 12.25 ± 2.98%; Plp1−CreER+/iDTR+: 2.61 ± 1.52%; p = 0.0074; ES = 1.64L). D CTLA-4 + /PD-1 + /CD4 + T cells (Plp1−CreER−/iDTR+: 2.97 ± 0.99%; Plp1−CreER+/iDTR+: 0.18 ± 0.08%; p = 0.0138; ES = 1.5L). F CTLA-4 + /CD8 + T cells (Plp1−CreER−/iDTR+: 2.41 ± 1.04%; Plp1−CreER+ /iDTR+: 2.23 ± 0.68%; p = 0.895; ES = 0.1T). G PD-1 + /CD8 + T cells (Plp1−CreER−/iDTR+: 21.70 ± 9.90%; Plp1−CreER+ /iDTR+: 6.51 ± 1.28%; p = 0.041; ES = 1.2L). H CTLA-4 + /PD-1 + /CD8 + T cells (Plp1−CreER−/iDTR+: 0.49 ± 0.49%; Plp1−CreER+/iDTR+: 0.67 ± 0.28%; p = 0.121, ES = 0.16T). I TIL from B16F10–inoculated mice were analyzed after 4 h of culture. Column charts show absolute numbers of CD4 + and CD8 + T cells producers of IFN-γ and IL-17. Cytokines levels were measured in cells isolated from tumors of B16F10–inoculated Plp1−CreER−/iDTR+ and Plp1−CreER+/iDTR+ animals. CD4 + T cells-producing IFN-γ (Plp1−CreER−/iDTR+: 8.5 × 106 ± 2.7 × 106 per mg of tumor; Plp1-CreER+/iDTR+: 4.8 × 107 ± 2.2 × 107 per mg of tumor; p = 0.1743; ES = 0.9 M); CD4 + T cells-producing IL-17 (Plp1−CreER−/iDTR+: 2.1 × 107 ± 5.2 × 106 per mg of tumor; Plp1−CreER+/iDTR+: 6.0 × 107 ± 1.0 × 107 per mg of tumor; p = 0.017; ES = 1.5L); CD8 + T cells-producing IFN-γ (Plp1−CreER−/iDTR+: 4.9 × 106 ± 2.1 × 106 per mg of tumor; Plp1−CreER+/iDTR+: 1.8 × 107 ± 4.7 × 106 per mg of tumor; p = 0.0423; ES = 1.3L); and CD8 + T cells-producing IL-17 (Plp1−CreER−/iDTR+: 2.5 × 106 ± 1.3 × 106 per mg of tumor; Plp1−CreER+ /iDTR+: 2.5 × 107 ± 9.8 × 106 per mg of tumor; p = 0.0427; ES = 1.1M). Statistical analysis: unpaired Student's t-tests and Mann–Whitney Rank Sum Test; ES effect size; Llarge (≥ 1.2); Mmedium (0.6–1.2) and Ttrivial (< 0.2). *p < 0.05 and **p < 0.01. Data are shown as mean ± SEM
Depletion of Schwann cells promotes increase in γδT, NKT, NK-cell tumor infiltration. Tumor-infiltrating lymphocytes from B16F10–inoculated mice were analyzed ex vivo in Plp1−CreER−/iDTR+ (n = 6 mice) and Plp1−CreER+ /iDTR+ (n = 9 mice) mice. Absolute number of (A) γδ T cells (Plp1−CreER−/iDTR+: 6.20 × 107 ± 1.47 × 107 cells per mg of tumor; Plp1−CreER+/iDTR+: 1.37 × 108 ± 2.54 × 107 cells per mg of tumor; p = 0.044; ES = 1.2L), (E) NKT cells (Plp1−CreER−/iDTR+: 1.33 × 107 ± 3.52 × 106 cells per mg of tumor; Plp1−CreER+/iDTR+: 2.68 × 107 ± 2.76 × 106 cells per mg of tumor; p = 0.008; ES = 1.5L), I NK cells (Plp1−CreER−/iDTR+: 1.16 × 107 ± 3.05 × 106 cells per mg of tumor; Plp1−CreER+/iDTR+: 3.40 × 107 ± 5.20 × 106 cells per mg of tumor; p = 0.004; ES = 1.8L) from tumors of B16F10–inoculated mice. Column charts show proportion of CTLA-4 (B, F, J), PD-1 (C, G, K) and CTLA-4/PD-1 (D, H, L) co-expressing γδ T cells (upper panel), NKT cells (middle panel) and NK cells (lower panel) from tumors of B16F10–inoculated mice. B. CTLA-4 + γδ T cells (Plp1−CreER−/iDTR+: 15.11 ± 9.11%; Plp1−CreER+ /iDTR+: 13.88 ± 4.34%; p = 0.897; ES = 0.06T) C. PD-1 + γδ T cells (Plp1−CreER−/iDTR +: 27.58 ± 5.69%; Plp1−CreER+/iDTR+: 6.52 ± 1.12%; p = 0.0019; ES = 1.9L). D CTLA-4 + /PD-1 + γδ T cells (Plp1−CreER−/iDTR+: 0.42 ± 0.15%; Plp1−CreER+/iDTR+: 0.054 ± 0.03%; p = 0.0158; ES = 1.3L). F CTLA-4 + NKT cells (Plp1−CreER−/iDTR+: 21.94 ± 4.87%; Plp1−CreER+/iDTR+: 24.63 ± 2.83%; p = 0.617; ES = 0.25S). G PD-1 + NKT cells (Plp1−CreER−/iDTR+: 3.74 ± 0.73%; Plp1−CreER+/iDTR+: 2.13 ± 0.50%; p = 0.044; ES = 1.0 M). H CTLA-4 + /PD-1 + NKT cells (Plp1−CreER−/iDTR+: 0.17 ± 0.17%; Plp1−CreER+/iDTR+: 0.14 ± 0.11%; p = 0.882; ES = 0.07T). J CTLA-4 + NK cells (Plp1−CreER−/iDTR+: 39.45 ± 4.51%; Plp1−CreER+ /iDTR+: 21.41 ± 2.66%; p = 0.0028; ES = 1.9L). K PD-1 + NK cells (Plp1−CreER−/iDTR+: 9.19 ± 3.69%; Plp1−CreER+/iDTR+: 1.57 ± 1.12%); p = 0.0451; ES = 1.0 M). L CTLA-4 + /PD-1 + NK cells (Plp1−CreER−/iDTR+: 4.20 ± 1.28%; Plp1−CreER+/iDTR+: 0.74 ± 0.60%; p = 0.0198; ES = 1.3L). M TIL from B16F10–inoculated mice were analyzed after 4 h of culture. Column charts show absolute number of γδ T cells, NKT cells and NK cells-producing IFN-γ and IL-17. Cytokines levels were measured in cells isolated from tumors of B16F10–inoculated Plp1−CreER−/iDTR+ and Plp1−CreER+/iDTR+ mice. γδ T cells-producing IFN-γ (Plp1−CreER−/iDTR+: 10.7 × 106 ± 2.6 × 106 per mg of tumor; Plp1−CreER+/iDTR+: 15 × 106 ± 5 × 106 per mg of tumor; p = 0.445; ES = 0.4S); NKT cells-producing IFN-γ (Plp1−CreER−/iDTR+: 12.6 × 106 ± 3.1 × 106 per mg of tumor; Plp1−CreER+/iDTR+: 28.1 × 106 ± 7.2 × 106 per mg of tumor; p = 0.105; ES = 1.0M); NK cells-producing IFN-γ (Plp1−CreER−/iDTR+: 19.2 × 106 ± 6.2 × 106 per mg of tumor; Plp1−CreER+/iDTR+: 21.1 × 106 ± 7.9 × 106 per mg of tumor; p = 0.883; ES = 0.1 T); γδ T cells-producing IL-17 (Plp1−CreER−/iDTR+: 3.9 × 106 ± 1.1 × 106 per mg of tumor; Plp1−CreER+/iDTR+: 53.1 × 106 ± 1.1 × 107 per mg of tumor; p = 0.003; ES = 2.1L); NKT cells-producing IL-17 (Plp1−CreER−/iDTR+: 13 × 106 ± 1.7 × 106 per mg of tumor; Plp1−CreER+/iDTR+: 33.5 × 106 ± 4.7 × 106 per mg of tumor; p = 0.004; ES = 2.0L); and NK cells-producing IL-17 (Plp1−CreER−/iDTR+: 17.1 × 106 ± 6 × 106 per mg of tumor; Plp1−CreER+/iDTR+: 40.9 × 106 ± 6.4 × 106 per mg of tumor); p = 0.022; ES = 1.4L). Statistical analysis: unpaired Student's t-tests or Mann–Whitney Rank Sum Test. ES effect size; Ttrivial (< 0.2); Ssmall (0.2–0.6); Mmedium (0.6–1.2); Llarge (≥ 1.2). *p < 0.05 and **p < 0.01. Data are shown as mean ± SEM
High expression of genes related to Schwann cells correlates with worse prognosis in human melanoma patients
In order to investigate our findings also in human tumors, we performed in silico analyses using the dataset from The Cancer Genome Atlas (TCGA): 103 samples from Skin Cutaneous Melanoma (SKCM) and 495 samples Prostate Adenocarcinoma (PRAD). First, we hand-picked 27 genes that are expressed in Schwann cells (Suppl. Table 5) [39, 75,76,77,78,79,80,81,82,83,84,85,86,87,88,89,90,91,92,93,94,95,96], with which we performed a protein–protein interaction (PPI) analysis (details provided in Materials and Methods). PPI analysis is a key step for finding networks of genes that work together in cells either in normal or disease states [183,184,185]. Notably, we found 20 genes highly connected in a network (Fig. 15A). In particular, GFAP, PLP1, NES and NGFR, which we found expressed in the Schwann cells within the tumor microenvironment (Figs. 1, 2 and 8) are highly connected in the PPI network, suggesting a synergistic function among them. Next, we investigated these genes in approximately 598 human tumors (SKCM: 103 samples; PRAD: 495 samples). We observed an increased expression of two key genes in our model (GFAP and PLP1) more pronounced in SKCM tumors, but also in PRAD tumors (Fig. 15B). SKCM patients with high expression of GFAP and PLP1 show a tendency of worse overall survival (Fig. 15C). To correlate the expression of GFAP and PLP1 gene markers in SKCM and PRAD tumors with the other components of the tumor microenvironments, we performed an in silico estimation of the proportions of immune infiltrated natural killer (NK), T CD4 and T CD8 cells in these tumors. In line with our findings, for SKCM patients we found a significant enrichment of NK and T CD8 cells in tumors with lower GFAP and PLP1 expression levels (better prognosis), while only a tendency was observed for T CD4 cells (Fig. 15D). For PRAD patients, we were only able to significantly establish immune cell infiltrates for 94 from the total 495 patients analyzed (p < 0.05), and no significant differences were observed for this reduced cohort (Fig. 15D).
Overexpression of genes related to Schwann cells is associated with impaired survival. A Protein–protein interaction network of Schwann cell markers. B Expression (TPM-normalized) of GFAP and PLP1 in SKCM and PRAD samples. C Overall survival of SKCM patients presenting high versus low expression of GFAP and PLP1. D Estimated proportions of NK, T CD4 and T CD8 cell infiltrates in SKCM and PRAD tumors presenting high versus low expression of GFAP and PLP1 (Wilcoxon Test; *p < 0.05, ns not significant)
In addition, given the pronounced overexpression of GFAP and PLP1 in SKCM tumors, we decided to investigate the set of differentially expressed genes in SKCM tumors compared to healthy skin (Suppl. Table 6). To gain a better understanding of the functional role of genes up-regulated in SKCM, we performed a Gene Ontology analysis and found that these genes are enriched in biological processes related to the nervous system and immunological processes, supporting a pivotal role of nervous system cells in the SKCM tumorigenesis (FDR adjusted p-value < 6 × 10–7; Fig. 16A). Next, we checked whether genes present in Schwann cells (Suppl. Table 5) [39, 84,85,86,87,88,89,90,91,92,93,94,95,96] were within the set of differentially expressed genes in SKCM and found 10 of these genes up-regulated in cancer (SKCM) versus healthy skin (Fig. 16B and Table 1). Finally, we investigated the expression of the two key genes in our model, GFAP and PLP1, in SKCM versus healthy skin. We confirmed that GFAP and PLP1 are overexpressed in SKCM (p-value < 0.0001, Fig. 16C).
Human Skin Cutaneous Melanomas have Schwann cell-related genes overexpressed in comparison to the Healthy Skin. A Gene ontology (Biological Processes) analysis of up-regulated genes in Skin Cutaneous Melanoma (SKCM) versus healthy skin samples. Only top 10 biological processes significantly enriched are shown (FDR adjusted p-value < 6x10-7). B Heatmap showing TPM-normalized expression values of Schwann cell gene markers which are up-regulated in SKCM versus healthy skin. C TPM-normalized expression of GFAP and PLP1 in SKCM versus healthy skin samples
Importantly, we also confirmed the presence of Plp1+ cells in human tumor samples by reanalysis of single-cell RNA sequencing data from human prostate cancer, breast cancer and melanoma patients (Fig. 17). We found that Plp1+ cells were located within the population called “fibroblasts” in all three types of tumors. This may be due to the fact that markers used to define fibroblasts may be expressed in Schwann cells as well, such as vimentin [186,187,188]. In support to our findings, we found several highly expressed genes in PLP1+ cells with previously described pro-tumorigenic activity (Fig. 17B, E, H). Future studies will need to evaluate the functional role of these genes in Schwann cells within the tumor microenvironment.
Identification of Plp1+ cells in human cancer samples by Single-cell RNA transcriptomics reanalysis. A Uniform Manifold Approximation and Projection (UMAP) plot of 35,000 + single-cells from 13 prostate tumors [102]. Fibroblasts were annotated according to original authors and colored based on the expression of PLP1 gene (purple = no expression of PLP1; brown = expression of PLP1). B Differential expression analysis between PLP1+ fibroblasts and PLP1− fibroblasts. There were 614 genes significantly induced in PLP1+ fibroblasts compared with PLP1− fibroblasts (Adjusted p < 0.05). Selected genes associated with pro-tumorigenic roles were displayed in the bar graph. C TNBC cells were arranged in the first two dimensions of the UMAP dimensionality reduction (the same coordinates as in the original publication) [103], with the fibroblasts cell population shown in black and all others in grey. D Focusing on the fibroblasts’ region, PLP1+ cells are shown in orange and PLP1− cells in purple. E Differential expression analysis between PLP1+ fibroblasts and PLP1− fibroblasts. Genes shown were selected based on log fold change and prior knowledge of their roles in cancer biology (Adjusted p < 0.05). Selected genes associated with pro-tumorigenic roles were displayed in the bar graph organized by fold change. F Melanoma cells were arranged in the first two dimensions of the tSNE dimensionality reduction (the same coordinates as in the original publication) [104], with the fibroblasts population shown in black and all others in grey. G Focusing on the fibroblasts’ region, PLP1+ cells are shown in orange and PLP1− cells in purple. H Differential expression analysis between PLP1+ fibroblasts and PLP1− fibroblasts. Genes shown were selected based on log fold change and prior knowledge of their roles in cancer biology (Adjusted p < 0.05). Selected genes associated with pro-tumorigenic roles were displayed in the bar graph organized by fold change
Discussion
In the present study, we discovered the presence of cells expressing GFAP and Plp1, Schwann cell markers, within the tumor microenvironment surrounding blood vessels. Our approach, using in vivo Cre/loxP technologies in combination with tumor implantation, revealed that tumor-infiltrating GFAP+/Plp1+ cells derive from tissue-resident Schwann cells. Genetic ablation of Schwann cells induced melanoma regression with decrease in tumor growth and in new blood vessel formation, as well as a boost in the anti-tumor immune surveillance (Fig. 18). This work indicates that targeting of GFAP+/Plp1+ cells within the tumor microenvironment represents a potential new therapeutic path in the battle against cancer.
Tumors are complex organ-like structures containing a variety of cell types, including differentiated cancer cells, cancer-initiating cells, immune cells, adipocytes, fibroblasts, endothelial cells, pericytes, and others; all of which have the capacity to reciprocally interact impacting cancer disease’s outcome [189, 190]. Here, we identified intra-tumoral cells expressing some Schwann cell markers, such as GFAP and Plp1. Future studies should explore whether and how these cells interact with all other cellular components of the tumor microenvironment. The heterogeneous cell populations within the tumor are highly plastic altering their marker expression, morphology, and function in response to the tumor microenvironment milieu [191, 192]. Schwann cells can be activated by a variety of conditions [193, 194]. We found that GFAP+ cells derive from endogenous tissue-resident Schwann cells, which during tumor development infiltrate within the tumor. Although activated Schwann cells’ ability to detach from nerve fibers was previously described, as during “Wallerian degeneration” [195], in this report, we show that these cells may associate to tumoral blood vessels. Thus, similar to other cell populations that can assume this perivascular position, such as macrophages [196, 197], pericytes [136], fibroblasts [198, 199], and others [200, 201], Schwann cells also associate to blood vessels during cancer development. The molecular triggers that lead tissue-resident Schwann cells toward blood vessels and the importance of communications between different intra-tumoral perivascular cell populations remains to be explored in the years to come.
Here, we show that genetic elimination of endogenous Schwann cells counteracts tumor growth. Previous studies, using myelinating Schwann cells isolated from sciatic nerves, have suggested that transplantation of these Schwann cells stimulates tumor growth [49]. Our findings also reveal that Schwann cells’ elimination affects the immune response to the tumor. Tumor progression is affected by the complex interplay between cancer cells and different components of the immune system [202]. Cancer cells may cause disruption of the organism’s immunity to overrun and escape the immune system control [203, 204]. The role of Schwann cells in these interactions remains completely unknown. Lymphocytes are the dominant immune elements found infiltrating the tumor microenvironment. Their composition correlates with patients’ survival [164]. While CD8 + T cells, CD4 + T cells, γδ T cells, and NK cells have been shown to act against the malignant cells, regulatory T cells play pro-tumorigenic roles [164, 205,206,207,208,209,210,211]. Our data shows that Schwann cells genetic ablation induce an increase in the number of tumor-infiltrating anti-cancer lymphocytes (CD8 + T cells, CD4 + T cells, γδ T cells, and NK cells), while we detected a reduction in the number of tumor-infiltrating regulatory T cells. Thus, our overall findings suggest that elimination of Schwann cells contributes to boosting of the immune response against the tumor. Future studies will need to explore the exact molecular mechanisms involved in the interactions of Schwann cells and immune cells in the tumor microenvironment.
Schwann cells produce multiple molecular mediators [25, 29, 212,213,214,215,216,217,218,219,220,221]. Although some studies in vitro suggest that Schwann cells may induce cancer progression by the production of pro-tumorigenic molecules [222,223,224], it remains unknown which molecules are essential for endogenous Schwann cells role in the tumor microenvironment in vivo. Cell culture systems may cause alterations in the cultured Schwann cells, leading them to behave differently than the same Schwann cells in vivo. These artificial conditions and high concentration of mitogens can induce production of molecules in the cultured Schwann cells that may not be shared by the corresponding endogenous Schwann cells in vivo [50]. Therefore, future studies should thoroughly explore the molecular mechanisms by which endogenous Schwann cells promote tumor growth by genetic elimination of individual molecules specifically from Schwann cells in vivo for instance using sophisticated Cre/loxP technologies in combination with cancer mouse models.
In conclusion, this work identifies GFAP+/Plp1+ cells as important habitants of the perivascular site within the tumor microenvironment. We also demonstrate that these cells originate from tissue-resident Schwann cells, and that genetic ablation of these cells induces tumor regression.
Data availability
Data will be made available on reasonable request.
Abbreviations
- BP:
-
Biological processes
- BV:
-
Blood vessel
- CAFs:
-
Cancer-associated fibroblasts
- CD:
-
Cluster differentiation
- CEUA:
-
Ethics Animal Care and Use Committee
- CTLA-4:
-
Cytotoxic T lymphocyte Antigen-4
- DC:
-
Dendritic cells
- DEGs:
-
Differentially expressed genes
- DMEM:
-
Dulbecco's modified eagle medium
- DT:
-
Diphtheria toxin
- ES:
-
Effect size
- FBS:
-
Fetal bovine serum
- FSC-A:
-
Forward scatter area
- FSC-H:
-
Forward scatter height
- GFAP:
-
Glial fibrillary acidic protein
- GFP:
-
Green fluorescent protein
- GO:
-
Gene ontology
- iDTR:
-
Diphtheria toxin receptor
- IFN-γ:
-
Interferon gamma
- IL:
-
Interleukin
- NGFR:
-
Nerve growth factor receptor
- NG2:
-
Neuron-glial antigen 2
- NK:
-
Natural killer
- OCT:
-
Tissue-Tek
- p75:
-
Neurotrophin-75
- PBS:
-
Phosphate-buffered saline
- PD-1:
-
Programmed cell death protein 1
- PDGFRβ:
-
Platelet-derived growth factor receptor beta
- PFA:
-
Paraformaldehyde
- Plp1:
-
Proteolipid protein 1
- SC:
-
Schwann cell
- SKCM:
-
Skin cutaneous melanoma
- TCGA:
-
The cancer genome atlas
- TH:
-
Tyrosine hydroxylase
- TNBC:
-
Triple-negative breast cancer
- TUBB3:
-
Class III β tubulin
- UFMG:
-
Federal University of Minas Gerais
- UMAP:
-
Uniform Manifold Approximation and Projection
- WT:
-
Wild-type
- γδ:
-
Gamma Delta
- SEM:
-
Standard error
References
Catalano V, Turdo A, Di Franco S, Dieli F, Todaro M, Stassi G (2013) Tumor and its microenvironment: a synergistic interplay. Semin Cancer Biol 23(6 Pt B):522–532. https://doi.org/10.1016/j.semcancer.2013.08.007
Hanahan D, Weinberg RA (2011) Hallmarks of cancer: the next generation. Cell 144(5):646–674. https://doi.org/10.1016/j.cell.2011.02.013
Swartz MA, Iida N, Roberts EW, Sangaletti S, Wong MH, Yull FE, Coussens LM, DeClerck YA (2012) Tumor microenvironment complexity: emerging roles in cancer therapy. Cancer Res 72(10):2473–2480. https://doi.org/10.1158/0008-5472.CAN-12-0122
Hanahan D, Coussens LM (2012) Accessories to the crime: functions of cells recruited to the tumor microenvironment. Cancer Cell 21(3):309–322. https://doi.org/10.1016/j.ccr.2012.02.022
Hinshaw DC, Shevde LA (2019) The tumor microenvironment innately modulates cancer progression. Cancer Res 79(18):4557–4566. https://doi.org/10.1158/0008-5472.CAN-18-3962
Senthebane DA, Rowe A, Thomford NE, Shipanga H, Munro D, Mazeedi M, Almazyadi HAM, Kallmeyer K, Dandara C, Pepper MS, Parker MI, Dzobo K (2017) The role of tumor microenvironment in chemoresistance: to survive, keep your enemies closer. Int J Mol Sci. https://doi.org/10.3390/ijms18071586
Renz BW, Takahashi R, Tanaka T, Macchini M, Hayakawa Y, Dantes Z, Maurer HC, Chen X, Jiang Z, Westphalen CB, Ilmer M, Valenti G, Mohanta SK, Habenicht AJR, Middelhoff M, Chu T, Nagar K, Tailor Y, Casadei R, Di Marco M, Kleespies A, Friedman RA, Remotti H, Reichert M, Worthley DL, Neumann J, Werner J, Iuga AC, Olive KP, Wang TC (2018) beta2 adrenergic-neurotrophin feedforward loop promotes pancreatic cancer. Cancer Cell 34(5):863–867. https://doi.org/10.1016/j.ccell.2018.10.010
Magnon C, Hall SJ, Lin J, Xue X, Gerber L, Freedland SJ, Frenette PS (2013) Autonomic nerve development contributes to prostate cancer progression. Science 341(6142):1236361. https://doi.org/10.1126/science.1236361
Zahalka AH, Arnal-Estape A, Maryanovich M, Nakahara F, Cruz CD, Finley LWS, Frenette PS (2017) Adrenergic nerves activate an angio-metabolic switch in prostate cancer. Science 358(6361):321–326. https://doi.org/10.1126/science.aah5072
Prazeres P, Leonel C, Silva WN, Rocha BGS, Santos GSP, Costa AC, Picoli CC, Sena IFG, Goncalves WA, Vieira MS, Costa PAC, Campos L, Lopes MTP, Costa MR, Resende RR, Cunha TM, Mintz A, Birbrair A (2020) Ablation of sensory nerves favours melanoma progression. J Cell Mol Med. https://doi.org/10.1111/jcmm.15381
Zhao CM, Hayakawa Y, Kodama Y, Muthupalani S, Westphalen CB, Andersen GT, Flatberg A, Johannessen H, Friedman RA, Renz BW, Sandvik AK, Beisvag V, Tomita H, Hara A, Quante M, Li Z, Gershon MD, Kaneko K, Fox JG, Wang TC, Chen D (2014) Denervation suppresses gastric tumorigenesis. Sci Transl Med 6(250):250ra115. https://doi.org/10.1126/scitranslmed.3009569
Saloman JL, Albers KM, Li D, Hartman DJ, Crawford HC, Muha EA, Rhim AD, Davis BM (2016) Ablation of sensory neurons in a genetic model of pancreatic ductal adenocarcinoma slows initiation and progression of cancer. Proc Natl Acad Sci USA 113(11):3078–3083. https://doi.org/10.1073/pnas.1512603113
Renz BW, Takahashi R, Tanaka T, Macchini M, Hayakawa Y, Dantes Z, Maurer HC, Chen X, Jiang Z, Westphalen CB, Ilmer M, Valenti G, Mohanta SK, Habenicht AJR, Middelhoff M, Chu T, Nagar K, Tailor Y, Casadei R, Di Marco M, Kleespies A, Friedman RA, Remotti H, Reichert M, Worthley DL, Neumann J, Werner J, Iuga AC, Olive KP, Wang TC (2018) beta2 adrenergic-neurotrophin feed forward loop promotes pancreatic cancer. Cancer Cell 33(1):75–90. https://doi.org/10.1016/j.ccell.2017.11.007
Dubeykovskaya Z, Si Y, Chen X, Worthley DL, Renz BW, Urbanska AM, Hayakawa Y, Xu T, Westphalen CB, Dubeykovskiy A, Chen D, Friedman RA, Asfaha S, Nagar K, Tailor Y, Muthupalani S, Fox JG, Kitajewski J, Wang TC (2016) Neural innervation stimulates splenic TFF2 to arrest myeloid cell expansion and cancer. Nat Commun 7:10517. https://doi.org/10.1038/ncomms10517
Kamiya A, Hayama Y, Kato S, Shimomura A, Shimomura T, Irie K, Kaneko R, Yanagawa Y, Kobayashi K, Ochiya T (2019) Genetic manipulation of autonomic nerve fiber innervation and activity and its effect on breast cancer progression. Nat Neurosci 22(8):1289–1305. https://doi.org/10.1038/s41593-019-0430-3
Renz BW, Tanaka T, Sunagawa M, Takahashi R, Jiang Z, Macchini M, Dantes Z, Valenti G, White RA, Middelhoff MA, Ilmer M, Oberstein PE, Angele MK, Deng H, Hayakawa Y, Westphalen CB, Werner J, Remotti H, Reichert M, Tailor YH, Nagar K, Friedman RA, Iuga AC, Olive KP, Wang TC (2018) Cholinergic signaling via muscarinic receptors directly and indirectly suppresses pancreatic tumorigenesis and cancer stemness. Cancer Discov 8(11):1458–1473. https://doi.org/10.1158/2159-8290.CD-18-0046
Hayakawa Y, Sakitani K, Konishi M, Asfaha S, Niikura R, Tomita H, Renz BW, Tailor Y, Macchini M, Middelhoff M, Jiang Z, Tanaka T, Dubeykovskaya ZA, Kim W, Chen X, Urbanska AM, Nagar K, Westphalen CB, Quante M, Lin CS, Gershon MD, Hara A, Zhao CM, Chen D, Worthley DL, Koike K, Wang TC (2017) Nerve growth factor promotes gastric tumorigenesis through aberrant cholinergic signaling. Cancer Cell 31(1):21–34. https://doi.org/10.1016/j.ccell.2016.11.005
Amit M, Takahashi H, Dragomir MP, Lindemann A, Gleber-Netto FO, Pickering CR, Anfossi S, Osman AA, Cai Y, Wang R, Knutsen E, Shimizu M, Ivan C, Rao X, Wang J, Silverman DA, Tam S, Zhao M, Caulin C, Zinger A, Tasciotti E, Dougherty PM, El-Naggar A, Calin GA, Myers JN (2020) Loss of p53 drives neuron reprogramming in head and neck cancer. Nature 578(7795):449–454. https://doi.org/10.1038/s41586-020-1996-3
Madeo M, Colbert PL, Vermeer DW, Lucido CT, Cain JT, Vichaya EG, Grossberg AJ, Muirhead D, Rickel AP, Hong Z, Zhao J, Weimer JM, Spanos WC, Lee JH, Dantzer R, Vermeer PD (2018) Cancer exosomes induce tumor innervation. Nat Commun 9(1):4284. https://doi.org/10.1038/s41467-018-06640-0
Gysler SM, Drapkin R (2021) Tumor innervation: peripheral nerves take control of the tumor microenvironment. J Clin Investig. https://doi.org/10.1172/JCI147276
Peng J, Chen H, Zhang B (2022) Nerve-stem cell crosstalk in skin regeneration and diseases. Trends Mol Med. https://doi.org/10.1016/j.molmed.2022.04.005
Picoli CC, Costa AC, Rocha BGS, Silva WN, Santos GSP, Prazeres P, Costa PAC, Oropeza A, da Silva RA, Azevedo VAC, Resende RR, Cunha TM, Mintz A, Birbrair A (2021) Sensory nerves in the spotlight of the stem cell niche. Stem Cells Transl Med 10(3):346–356. https://doi.org/10.1002/sctm.20-0284
Picoli CC, Goncalves BOP, Santos GSP, Rocha BGS, Costa AC, Resende RR, Birbrair A (2021) Pericytes cross-talks within the tumor microenvironment. Biochim Biophys Acta Rev Cancer 1876 2:188608. https://doi.org/10.1016/j.bbcan.2021.188608
Lousado L, Prazeres PHDM, Andreotti JP, Paiva AE, Azevedo PO, Santos GSP, Filev R, Mintz A, Birbrair A (2017) Schwann cell precursors as a source for adrenal gland chromaffin cells. Cell Death Dis 8:e3072
Scholz J, Woolf CJ (2007) The neuropathic pain triad: neurons, immune cells and glia. Nat Neurosci 10(11):1361–1368. https://doi.org/10.1038/nn1992
Mukouyama YS, Shin D, Britsch S, Taniguchi M, Anderson DJ (2002) Sensory nerves determine the pattern of arterial differentiation and blood vessel branching in the skin. Cell 109(6):693–705
Li W, Kohara H, Uchida Y, James JM, Soneji K, Cronshaw DG, Zou YR, Nagasawa T, Mukouyama YS (2013) Peripheral nerve-derived CXCL12 and VEGF-A regulate the patterning of arterial vessel branching in developing limb skin. Dev Cell 24(4):359–371. https://doi.org/10.1016/j.devcel.2013.01.009
Colomar A, Robitaille R (2004) Glial modulation of synaptic transmission at the neuromuscular junction. Glia 47(3):284–289. https://doi.org/10.1002/glia.20086
De Logu F, Nassini R, Materazzi S, Carvalho Goncalves M, Nosi D, Rossi Degl’Innocenti D, Marone IM, Ferreira J, Li Puma S, Benemei S, Trevisan G, Monteiro S, de Araujo D, Patacchini R, Bunnett NW, Geppetti P (2017) Schwann cell TRPA1 mediates neuroinflammation that sustains macrophage-dependent neuropathic pain in mice. Nat Commun 8(1):1887. https://doi.org/10.1038/s41467-017-01739-2
Rinwa P, Calvo-Enrique L, Zhang MD, Nyengaard JR, Karlsson P, Ernfors P (2021) Demise of nociceptive Schwann cells causes nerve retraction and pain hyperalgesia. Pain 162(6):1816–1827. https://doi.org/10.1097/j.pain.0000000000002169
Wei Z, Fei Y, Su W, Chen G (2019) Emerging role of Schwann cells in neuropathic pain: receptors, glial mediators and myelination. Front Cell Neurosci 13:116. https://doi.org/10.3389/fncel.2019.00116
Wekerle H, Schwab M, Linington C, Meyermann R (1986) Antigen presentation in the peripheral nervous system: Schwann cells present endogenous myelin autoantigens to lymphocytes. Eur J Immunol 16(12):1551–1557. https://doi.org/10.1002/eji.1830161214
Martin I, Nguyen TD, Krell V, Greiner JF, Muller J, Hauser S, Heimann P, Widera D (2012) Generation of Schwann cell-derived multipotent neurospheres isolated from intact sciatic nerve. Stem Cell Rev 8(4):1178–1187. https://doi.org/10.1007/s12015-012-9387-2
Widera D, Heimann P, Zander C, Imielski Y, Heidbreder M, Heilemann M, Kaltschmidt C, Kaltschmidt B (2011) Schwann cells can be reprogrammed to multipotency by culture. Stem Cells Dev 20(12):2053–2064. https://doi.org/10.1089/scd.2010.0525
Masaki T, Qu J, Cholewa-Waclaw J, Burr K, Raaum R, Rambukkana A (2013) Reprogramming adult Schwann cells to stem cell-like cells by leprosy bacilli promotes dissemination of infection. Cell 152(1–2):51–67. https://doi.org/10.1016/j.cell.2012.12.014
Kaukua N, Shahidi MK, Konstantinidou C, Dyachuk V, Kaucka M, Furlan A, An Z, Wang L, Hultman I, Ahrlund-Richter L, Blom H, Brismar H, Lopes NA, Pachnis V, Suter U, Clevers H, Thesleff I, Sharpe P, Ernfors P, Fried K, Adameyko I (2014) Glial origin of mesenchymal stem cells in a tooth model system. Nature 513(7519):551–554. https://doi.org/10.1038/nature13536
Adameyko I, Lallemend F, Aquino JB, Pereira JA, Topilko P, Muller T, Fritz N, Beljajeva A, Mochii M, Liste I, Usoskin D, Suter U, Birchmeier C, Ernfors P (2009) Schwann cell precursors from nerve innervation are a cellular origin of melanocytes in skin. Cell 139(2):366–379. https://doi.org/10.1016/j.cell.2009.07.049
Uesaka T, Nagashimada M, Enomoto H (2015) Neuronal differentiation in schwann cell lineage underlies postnatal neurogenesis in the enteric nervous system. J Neurosci 35(27):9879–9888. https://doi.org/10.1523/JNEUROSCI.1239-15.2015
Yamazaki S, Ema H, Karlsson G, Yamaguchi T, Miyoshi H, Shioda S, Taketo MM, Karlsson S, Iwama A, Nakauchi H (2011) Nonmyelinating Schwann cells maintain hematopoietic stem cell hibernation in the bone marrow niche. Cell 147(5):1146–1158. https://doi.org/10.1016/j.cell.2011.09.053
Silva WN, Leonel C, Prazeres PHDM, Sena IFG, Guerra DAP, Heller D, Diniz IMA, Fortuna V, Mintz A, Birbrair A (2018) Role of Schwann cells in cutaneous wound healing. Wound Repair Regen 26(5):392–397. https://doi.org/10.1111/wrr.12647
Carr MJ, Johnston AP (2017) Schwann cells as drivers of tissue repair and regeneration. Curr Opin Neurobiol 47:52–57. https://doi.org/10.1016/j.conb.2017.09.003
Joseph NM, Mosher JT, Buchstaller J, Snider P, McKeever PE, Lim M, Conway SJ, Parada LF, Zhu Y, Morrison SJ (2008) The loss of Nf1 transiently promotes self-renewal but not tumorigenesis by neural crest stem cells. Cancer Cell 13(2):129–140. https://doi.org/10.1016/j.ccr.2008.01.003
Roosli C, Linthicum FH Jr, Cureoglu S, Merchant SN (2012) What is the site of origin of cochleovestibular schwannomas? Audiol Neurootol 17(2):121–125. https://doi.org/10.1159/000331394
Ostrom QT, Cioffi G, Gittleman H, Patil N, Waite K, Kruchko C, Barnholtz-Sloan JS (2019) CBTRUS statistical report: primary brain and other central nervous system tumors diagnosed in the United States in 2012–2016. Neuro Oncol 21(Suppl 5):v1–v100. https://doi.org/10.1093/neuonc/noz150
Furlan A, Dyachuk V, Kastriti ME, Calvo-Enrique L, Abdo H, Hadjab S, Chontorotzea T, Akkuratova N, Usoskin D, Kamenev D, Petersen J, Sunadome K, Memic F, Marklund U, Fried K, Topilko P, Lallemend F, Kharchenko PV, Ernfors P, Adameyko I (2017) Multipotent peripheral glial cells generate neuroendocrine cells of the adrenal medulla. Science. https://doi.org/10.1126/science.aal3753
Swanson BJ, McDermott KM, Singh PK, Eggers JP, Crocker PR, Hollingsworth MA (2007) MUC1 is a counter-receptor for myelin-associated glycoprotein (Siglec-4a) and their interaction contributes to adhesion in pancreatic cancer perineural invasion. Cancer Res 67(21):10222–10229. https://doi.org/10.1158/0008-5472.CAN-06-2483
Deborde S, Omelchenko T, Lyubchik A, Zhou Y, He S, McNamara WF, Chernichenko N, Lee SY, Barajas F, Chen CH, Bakst RL, Vakiani E, He S, Hall A, Wong RJ (2016) Schwann cells induce cancer cell dispersion and invasion. J Clin Investig 126(4):1538–1554. https://doi.org/10.1172/JCI82658
Sroka IC, Chopra H, Das L, Gard JM, Nagle RB, Cress AE (2016) Schwann cells increase prostate and pancreatic tumor cell invasion using laminin binding A6 integrin. J Cell Biochem 117(2):491–499. https://doi.org/10.1002/jcb.25300
Shurin GV, Kruglov O, Ding F, Lin Y, Hao X, Keskinov AA, You Z, Lokshin AE, LaFramboise WA, Falo LD Jr, Shurin MR, Bunimovich YL (2019) Melanoma-induced reprogramming of schwann cell signaling aids tumor growth. Cancer Res 79(10):2736–2747. https://doi.org/10.1158/0008-5472.CAN-18-3872
Snippert HJ, Clevers H (2011) Tracking adult stem cells. EMBO Rep 12(2):113–122. https://doi.org/10.1038/embor.2010.216
Birbrair A, Borges IDT, Gilson Sena IF, Almeida GG, da Silva ML, Goncalves R, Mintz A, Delbono O (2017) How plastic are pericytes? Stem Cells Dev 26(14):1013–1019. https://doi.org/10.1089/scd.2017.0044
Khan JA, Mendelson A, Kunisaki Y, Birbrair A, Kou Y, Arnal-Estape A, Pinho S, Ciero P, Nakahara F, Ma’ayan A, Bergman A, Merad M, Frenette PS (2016) Fetal liver hematopoietic stem cell niches associate with portal vessels. Science 351(6269):176–180. https://doi.org/10.1126/science.aad0084
Asada N, Kunisaki Y, Pierce H, Wang Z, Fernandez NF, Birbrair A, Ma’ayan A, Frenette PS (2017) Differential cytokine contributions of perivascular haematopoietic stem cell niches. Nat Cell Biol 19(3):214–223. https://doi.org/10.1038/ncb3475
Nobre AR, Risson E, Singh DK, Martino JD, Cheung JF, Wang J, Johnson J, Russnes HG, Bravo-Cordero JJ, Birbrair A, Naume B, Azhar M, Frenette PS, Aguirre-Ghiso JA (2021) Bone marrow NG2+/Nestin+ mesenchymal stem cells drive DTC dormancy via TGFβ2. Nat Cancer 2:327–339
Clausen BE, Burkhardt C, Reith W, Renkawitz R, Forster I (1999) Conditional gene targeting in macrophages and granulocytes using LysMcre mice. Transgenic Res 8(4):265–277. https://doi.org/10.1023/a:1008942828960
Qian BZ, Li J, Zhang H, Kitamura T, Zhang J, Campion LR, Kaiser EA, Snyder LA, Pollard JW (2011) CCL2 recruits inflammatory monocytes to facilitate breast-tumour metastasis. Nature 475(7355):222–225. https://doi.org/10.1038/nature10138
Madisen L, Zwingman TA, Sunkin SM, Oh SW, Zariwala HA, Gu H, Ng LL, Palmiter RD, Hawrylycz MJ, Jones AR, Lein ES, Zeng H (2010) A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat Neurosci 13(1):133–140. https://doi.org/10.1038/nn.2467
Birbrair A, Zhang T, Files DC, Mannava S, Smith T, Wang ZM, Messi ML, Mintz A, Delbono O (2014) Type-1 pericytes accumulate after tissue injury and produce collagen in an organ-dependent manner. Stem Cell Res Ther 5(6):122. https://doi.org/10.1186/scrt512
Birbrair A, Zhang T, Wang ZM, Messi ML, Enikolopov GN, Mintz A, Delbono O (2013) Skeletal muscle neural progenitor cells exhibit properties of NG2-glia. Exp Cell Res 319(1):45–63. https://doi.org/10.1016/j.yexcr.2012.09.008
Birbrair A, Zhang T, Wang ZM, Messi ML, Enikolopov GN, Mintz A, Delbono O (2013) Role of pericytes in skeletal muscle regeneration and fat accumulation. Stem Cells Dev 22(16):2298–2314. https://doi.org/10.1089/scd.2012.0647
Birbrair A, Zhang T, Wang ZM, Messi ML, Mintz A, Delbono O (2013) Type-1 pericytes participate in fibrous tissue deposition in aged skeletal muscle. Am J Physiol Cell Physiol 305(11):C1098–C1113. https://doi.org/10.1152/ajpcell.00171.2013
Gomes NA, do Valle IB, Gleber-Netto FO, Silva TA, Oliveira HMdC, de Oliveira RF, Ferreira LdAQ, Castilho LS, Reis PHRG, Prazeres PHDM, Menezes GB, de Magalhães CS, Mesquita RA, Marques MM, Birbrair A, Diniz IMA (2022) Nestin and NG2 transgenes reveal two populations of perivascular cells stimulated by photobiomodulation. J Cell Physiol 237:2198–2210. https://doi.org/10.1002/jcp.30680
do Valle IB, Prazeres PHDM, Mesquita RA, Silva TA, de Castro Oliveira HM, Castro PR, Freitas IDP, Oliveira SR, Gomes NA, de Oliveira RF, Marquiore LF, Macari S, do Amaral FA, Jácome-Santos H, Barcelos LS, Menezes GB, Marques MM, Birbrair A, Diniz IMA (2010) Photobiomodulation drives pericyte mobilization towards skin regeneration. Sci Rep 10(1):19257. https://doi.org/10.1038/s41598-020-76243-7
Doerflinger NH, Macklin WB, Popko B (2003) Inducible site-specific recombination in myelinating cells. Genesis 35(1):63–72. https://doi.org/10.1002/gene.10154
Buch T, Heppner FL, Tertilt C, Heinen TJ, Kremer M, Wunderlich FT, Jung S, Waisman A (2005) A Cre-inducible diphtheria toxin receptor mediates cell lineage ablation after toxin administration. Nat Methods 2(6):419–426. https://doi.org/10.1038/nmeth762
Ellwood-Yen K, Graeber TG, Wongvipat J, Iruela-Arispe ML, Zhang J, Matusik R, Thomas GV, Sawyers CL (2003) Myc-driven murine prostate cancer shares molecular features with human prostate tumors. Cancer Cell 4(3):223–238. https://doi.org/10.1016/s1535-6108(03)00197-1
Birbrair A, Sattiraju A, Zhu D, Zulato G, Batista I, Nguyen VT, Messi ML, Solingapuram Sai KK, Marini FC, Delbono O, Mintz A (2017) Novel peripherally derived neural-like stem cells as therapeutic carriers for treating glioblastomas. Stem Cells Transl Med 6(2):471–481. https://doi.org/10.5966/sctm.2016-0007
Birbrair A, Wang ZM, Messi ML, Enikolopov GN, Delbono O (2011) Nestin-GFP transgene reveals neural precursor cells in adult skeletal muscle. PLoS One 6(2):e16816. https://doi.org/10.1371/journal.pone.0016816
Mignone JL, Kukekov V, Chiang AS, Steindler D, Enikolopov G (2004) Neural stem and progenitor cells in nestin-GFP transgenic mice. J Comp Neurol 469(3):311–324. https://doi.org/10.1002/cne.10964
Foster BA, Gingrich JR, Kwon ED, Madias C, Greenberg NM (1997) Characterization of prostatic epithelial cell lines derived from transgenic adenocarcinoma of the mouse prostate (TRAMP) model. Cancer Res 57(16):3325–3330
Liu Z, Eltoum IE, Guo B, Beck BH, Cloud GA, Lopez RD (2008) Protective immunosurveillance and therapeutic antitumor activity of gammadelta T cells demonstrated in a mouse model of prostate cancer. J Immunol 180(9):6044–6053. https://doi.org/10.4049/jimmunol.180.9.6044
Yeon A, Wang Y, Su S, Lo EM, Kim HL (2020) Syngeneic murine model for prostate cancer using RM1 cells transfected with gp100. Prostate 80(5):424–431. https://doi.org/10.1002/pros.23957
Coimbra-Campos LMC, Silva WN, Baltazar LM, Costa PAC, Prazeres PHDM, Picoli CC, Costa AC, Rocha BGS, Santos GSP, Oliveira FMS, Pinto MCX, Amorim JH, Azevedo VAC, Souza DG, Russo RC, Resende RR, Mintz A, Birbrair A (2021) Circulating Nestin-GFP+ cells participate in the pathogenesis of Paracoccidioides brasiliensis in the lungs. Stem Cell Rev Rep 17:1874
Bohn T, Rapp S, Luther N, Klein M, Bruehl TJ, Kojima N, Aranda Lopez P, Hahlbrock J, Muth S, Endo S, Pektor S, Brand A, Renner K, Popp V, Gerlach K, Vogel D, Lueckel C, Arnold-Schild D, Pouyssegur J, Kreutz M, Huber M, Koenig J, Weigmann B, Probst HC, von Stebut E, Becker C, Schild H, Schmitt E, Bopp T (2018) Tumor immunoevasion via acidosis-dependent induction of regulatory tumor-associated macrophages. Nat Immunol 19(12):1319–1329. https://doi.org/10.1038/s41590-018-0226-8
Gast CE, Silk AD, Zarour L, Riegler L, Burkhart JG, Gustafson KT, Parappilly MS, Roh-Johnson M, Goodman JR, Olson B, Schmidt M, Swain JR, Davies PS, Shasthri V, Iizuka S, Flynn P, Watson S, Korkola J, Courtneidge SA, Fischer JM, Jaboin J, Billingsley KG, Lopez CD, Burchard J, Gray J, Coussens LM, Sheppard BC, Wong MH (2018) Cell fusion potentiates tumor heterogeneity and reveals circulating hybrid cells that correlate with stage and survival. Sci Adv 4(9):eaat7828. https://doi.org/10.1126/sciadv.aat7828
Herring BP, Hoggatt AM, Burlak C, Offermanns S (2014) Previously differentiated medial vascular smooth muscle cells contribute to neointima formation following vascular injury. Vasc Cell 6:21. https://doi.org/10.1186/2045-824X-6-21
Nikolakopoulou AM, Montagne A, Kisler K, Dai Z, Wang Y, Huuskonen MT, Sagare AP, Lazic D, Sweeney MD, Kong P, Wang M, Owens NC, Lawson EJ, Xie X, Zhao Z, Zlokovic BV (2019) Pericyte loss leads to circulatory failure and pleiotrophin depletion causing neuron loss. Nat Neurosci 22(7):1089–1098. https://doi.org/10.1038/s41593-019-0434-z
Cha JH, Chang MY, Richardson JA, Eidels L (2003) Transgenic mice expressing the diphtheria toxin receptor are sensitive to the toxin. Mol Microbiol 49(1):235–240. https://doi.org/10.1046/j.1365-2958.2003.03550.x
Leclerc M, Voilin E, Gros G, Corgnac S, de Montpreville V, Validire P, Bismuth G, Mami-Chouaib F (2019) Regulation of antitumour CD8 T-cell immunity and checkpoint blockade immunotherapy by Neuropilin-1. Nat Commun 10(1):3345. https://doi.org/10.1038/s41467-019-11280-z
Gerdes J, Lemke H, Baisch H, Wacker HH, Schwab U, Stein H (1984) Cell cycle analysis of a cell proliferation-associated human nuclear antigen defined by the monoclonal antibody Ki-67. J Immunol 133(4):1710–1715
Sena IFG, Rocha BGS, Picoli CC, Santos GSP, Costa AC, Goncalves BOP, Garcia APV, Soltani-Asl M, Coimbra-Campos LMC, Silva WN, Costa PAC, Pinto MCX, Amorim JH, Azevedo VAC, Resende RR, Heller D, Cassali GD, Mintz A, Birbrair A (2021) C(3)1-TAg in C57BL/6 J background as a model to study mammary tumor development. Histochem Cell Biol. https://doi.org/10.1007/s00418-021-01995-w
Sena IFG, Fernandes LL, Lorandi LL, Santana TV, Cintra L, Lima IF, Iwai LK, Kramer JM, Birbrair A, Heller D (2022) Identification of early biomarkers in saliva in genetically engineered mouse model C(3)1-TAg of breast cancer. Sci Rep. https://doi.org/10.1038/s41598-022-14514-1
Newman AM, Steen CB, Liu CL, Gentles AJ, Chaudhuri AA, Scherer F, Khodadoust MS, Esfahani MS, Luca BA, Steiner D, Diehn M, Alizadeh AA (2019) Determining cell type abundance and expression from bulk tissues with digital cytometry. Nat Biotechnol 37(7):773–782. https://doi.org/10.1038/s41587-019-0114-2
Wolbert J, Li X, Heming M, Mausberg AK, Akkermann D, Frydrychowicz C, Fledrich R, Groeneweg L, Schulz C, Stettner M, Alonso Gonzalez N, Wiendl H, Stassart R, Meyer Zu Horste G (2020) Redefining the heterogeneity of peripheral nerve cells in health and autoimmunity. Proc Natl Acad Sci USA 117(17):9466–9476. https://doi.org/10.1073/pnas.1912139117
Tasdemir-Yilmaz OE, Druckenbrod NR, Olukoya OO, Dong W, Yung AR, Bastille I, Pazyra-Murphy MF, Sitko AA, Hale EB, Vigneau S, Gimelbrant AA, Kharchenko PV, Goodrich LV, Segal RA (2021) Diversity of developing peripheral glia revealed by single-cell RNA sequencing. Dev Cell 56(17):2516–2535. https://doi.org/10.1016/j.devcel.2021.08.005
Liu Z, Jin YQ, Chen L, Wang Y, Yang X, Cheng J, Wu W, Qi Z, Shen Z (2015) Specific marker expression and cell state of Schwann cells during culture in vitro. PLoS ONE 10(4):e0123278. https://doi.org/10.1371/journal.pone.0123278
Peng K, Sant D, Andersen N, Silvera R, Camarena V, Pinero G, Graham R, Khan A, Xu XM, Wang G, Monje PV (2020) Magnetic separation of peripheral nerve-resident cells underscores key molecular features of human Schwann cells and fibroblasts: an immunochemical and transcriptomics approach. Sci Rep 10(1):18433. https://doi.org/10.1038/s41598-020-74128-3
Skelly DA, Squiers GT, McLellan MA, Bolisetty MT, Robson P, Rosenthal NA, Pinto AR (2018) Single-cell transcriptional profiling reveals cellular diversity and intercommunication in the mouse heart. Cell Rep 22(3):600–610. https://doi.org/10.1016/j.celrep.2017.12.072
Clements MP, Byrne E, Camarillo Guerrero LF, Cattin AL, Zakka L, Ashraf A, Burden JJ, Khadayate S, Lloyd AC, Marguerat S, Parrinello S (2017) The wound microenvironment reprograms Schwann cells to invasive mesenchymal-like cells to drive peripheral nerve regeneration. Neuron 96(1):98–114. https://doi.org/10.1016/j.neuron.2017.09.008
Parmantier E, Lynn B, Lawson D, Turmaine M, Namini SS, Chakrabarti L, McMahon AP, Jessen KR, Mirsky R (1999) Schwann cell-derived Desert hedgehog controls the development of peripheral nerve sheaths. Neuron 23(4):713–724. https://doi.org/10.1016/s0896-6273(01)80030-1
Woods C, Kapur RP, Bischoff A, Lovell M, Arnold M, Pena A, Flockton A, Sharkey KA, Belkind-Gerson J (2021) Neurons populating the rectal extrinsic nerves in humans express neuronal and Schwann cell markers. Neurogastroenterol Motil 33(7):e14074. https://doi.org/10.1111/nmo.14074
Frank M, Schaeren-Wiemers N, Schneider R, Schwab ME (1999) Developmental expression pattern of the myelin proteolipid MAL indicates different functions of MAL for immature Schwann cells and in a late step of CNS myelinogenesis. J Neurochem 73(2):587–597. https://doi.org/10.1046/j.1471-4159.1999.0730587.x
Torres-Mejia E, Trumbach D, Kleeberger C, Dornseifer U, Orschmann T, Backer T, Brenke JK, Hadian K, Wurst W, Lopez-Schier H, Desbordes SC (2020) Sox2 controls Schwann cell self-organization through fibronectin fibrillogenesis. Sci Rep 10(1):1984. https://doi.org/10.1038/s41598-019-56877-y
Mazzara PG, Massimino L, Pellegatta M, Ronchi G, Ricca A, Iannielli A, Giannelli SG, Cursi M, Cancellieri C, Sessa A, Del Carro U, Quattrini A, Geuna S, Gritti A, Taveggia C, Broccoli V (2017) Two factor-based reprogramming of rodent and human fibroblasts into Schwann cells. Nat Commun 8:14088. https://doi.org/10.1038/ncomms14088
Frob F, Sock E, Tamm ER, Saur AL, Hillgartner S, Williams TJ, Fujii T, Fukunaga R, Wegner M (2019) Ep400 deficiency in Schwann cells causes persistent expression of early developmental regulators and peripheral neuropathy. Nat Commun 10(1):2361. https://doi.org/10.1038/s41467-019-10287-w
Doddrell RD, Dun XP, Moate RM, Jessen KR, Mirsky R, Parkinson DB (2012) Regulation of Schwann cell differentiation and proliferation by the Pax-3 transcription factor. Glia 60(9):1269–1278. https://doi.org/10.1002/glia.22346
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13(11):2498–2504. https://doi.org/10.1101/gr.1239303
Szklarczyk D, Gable AL, Lyon D, Junge A, Wyder S, Huerta-Cepas J, Simonovic M, Doncheva NT, Morris JH, Bork P, Jensen LJ, Mering CV (2019) STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res 47(D1):D607–D613. https://doi.org/10.1093/nar/gky1131
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15(12):550. https://doi.org/10.1186/s13059-014-0550-8
Ge SX, Jung D, Yao R (2020) ShinyGO: a graphical gene-set enrichment tool for animals and plants. Bioinformatics 36(8):2628–2629. https://doi.org/10.1093/bioinformatics/btz931
Supek F, Bosnjak M, Skunca N, Smuc T (2011) REVIGO summarizes and visualizes long lists of gene ontology terms. PLoS ONE 6(7):e21800. https://doi.org/10.1371/journal.pone.0021800
Chen S, Zhu G, Yang Y, Wang F, Xiao YT, Zhang N, Bian X, Zhu Y, Yu Y, Liu F, Dong K, Mariscal J, Liu Y, Soares F, Loo Yau H, Zhang B, Chen W, Wang C, Chen D, Guo Q, Yi Z, Liu M, Fraser M, De Carvalho DD, Boutros PC, Di Vizio D, Jiang Z, van der Kwast T, Berlin A, Wu S, Wang J, He HH, Ren S (2021) Single-cell analysis reveals transcriptomic remodellings in distinct cell types that contribute to human prostate cancer progression. Nat Cell Biol 23(1):87–98. https://doi.org/10.1038/s41556-020-00613-6
Wu SZ, Roden DL, Wang C, Holliday H, Harvey K, Cazet AS, Murphy KJ, Pereira B, Al-Eryani G, Bartonicek N, Hou R, Torpy JR, Junankar S, Chan CL, Lam CE, Hui MN, Gluch L, Beith J, Parker A, Robbins E, Segara D, Mak C, Cooper C, Warrier S, Forrest A, Powell J, O’Toole S, Cox TR, Timpson P, Lim E, Liu XS, Swarbrick A (2020) Stromal cell diversity associated with immune evasion in human triple-negative breast cancer. EMBO J 39(19):e104063. https://doi.org/10.15252/embj.2019104063
Tirosh I, Izar B, Prakadan SM, Wadsworth MH 2nd, Treacy D, Trombetta JJ, Rotem A, Rodman C, Lian C, Murphy G, Fallahi-Sichani M, Dutton-Regester K, Lin JR, Cohen O, Shah P, Lu D, Genshaft AS, Hughes TK, Ziegler CG, Kazer SW, Gaillard A, Kolb KE, Villani AC, Johannessen CM, Andreev AY, Van Allen EM, Bertagnolli M, Sorger PK, Sullivan RJ, Flaherty KT, Frederick DT, Jane-Valbuena J, Yoon CH, Rozenblatt-Rosen O, Shalek AK, Regev A, Garraway LA (2016) Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352(6282):189–196. https://doi.org/10.1126/science.aad0501
Fritz CO, Morris PE, Richler JJ (2012) Effect size estimates: current use, calculations, and interpretation. J Exp Psychol Gen 141(1):2–18. https://doi.org/10.1037/a0024338
Hopkins WG, Marshall SW, Batterham AM, Hanin J (2009) Progressive statistics for studies in sports medicine and exercise science. Med Sci Sports Exerc 41(1):3–13. https://doi.org/10.1249/MSS.0b013e31818cb278
Erin N, Zhao W, Bylander J, Chase G, Clawson G (2006) Capsaicin-induced inactivation of sensory neurons promotes a more aggressive gene expression phenotype in breast cancer cells. Breast Cancer Res Treat 99(3):351–364. https://doi.org/10.1007/s10549-006-9219-7
Costa PAC, Silva WN, Prazeres P, Picoli CC, Guardia GDA, Costa AC, Oliveira MA, Guimaraes PPG, Goncalves R, Pinto MCX, Amorim JH, Azevedo VAC, Resende RR, Russo RC, Cunha TM, Galante PAF, Mintz A, Birbrair A (2021) Chemogenetic modulation of sensory neurons reveals their regulating role in melanoma progression. Acta Neuropathol Commun 9(1):183. https://doi.org/10.1186/s40478-021-01273-9
Jessen KR, Mirsky R, Lloyd AC (2015) Schwann cells: development and role in nerve repair. Cold Spring Harb Perspect Biol 7(7):a020487. https://doi.org/10.1101/cshperspect.a020487
Bechmann I, Priller J, Kovac A, Bontert M, Wehner T, Klett FF, Bohsung J, Stuschke M, Dirnagl U, Nitsch R (2001) Immune surveillance of mouse brain perivascular spaces by blood-borne macrophages. Eur J Neurosci 14(10):1651–1658
Guillemin GJ, Brew BJ (2004) Microglia, macrophages, perivascular macrophages, and pericytes: a review of function and identification. J Leukoc Biol 75(3):388–397. https://doi.org/10.1189/jlb.0303114
Honda M, Surewaard BGJ, Watanabe M, Hedrick CC, Lee WY, Brown K, McCoy KD, Kubes P (2020) Perivascular localization of macrophages in the intestinal mucosa is regulated by Nr4a1 and the microbiome. Nat Commun 11(1):1329. https://doi.org/10.1038/s41467-020-15068-4
Mai CL, Tan Z, Xu YN, Zhang JJ, Huang ZH, Wang D, Zhang H, Gui WS, Zhang J, Lin ZJ, Meng YT, Wei X, Jie YT, Grace PM, Wu LJ, Zhou LJ, Liu XG (2021) CXCL12-mediated monocyte transmigration into brain perivascular space leads to neuroinflammation and memory deficit in neuropathic pain. Theranostics 11(3):1059–1078. https://doi.org/10.7150/thno.44364
Roubeix C, Dominguez E, Raoul W, Guillonneau X, Paques M, Sahel JA, Sennlaub F (2019) Mo-derived perivascular macrophage recruitment protects against endothelial cell death in retinal vein occlusion. J Neuroinflamm 16(1):157. https://doi.org/10.1186/s12974-019-1547-8
Schnieder TP, Zhou Qin ID, Trencevska-Ivanovska I, Rosoklija G, Stankov A, Pavlovski G, Mann JJ, Dwork AJ (2019) Blood vessels and perivascular phagocytes of prefrontal white and gray matter in suicide. J Neuropathol Exp Neurol 78(1):15–30. https://doi.org/10.1093/jnen/nly103
Jolivel V, Bicker F, Biname F, Ploen R, Keller S, Gollan R, Jurek B, Birkenstock J, Poisa-Beiro L, Bruttger J, Opitz V, Thal SC, Waisman A, Bauerle T, Schafer MK, Zipp F, Schmidt MHH (2015) Perivascular microglia promote blood vessel disintegration in the ischemic penumbra. Acta Neuropathol 129(2):279–295. https://doi.org/10.1007/s00401-014-1372-1
Koizumi T, Taguchi K, Mizuta I, Toba H, Ohigashi M, Onishi O, Ikoma K, Miyata S, Nakata T, Tanaka M, Foulquier S, Steinbusch HWM, Mizuno T (2019) Transiently proliferating perivascular microglia harbor M1 type and precede cerebrovascular changes in a chronic hypertension model. J Neuroinflamm 16(1):79. https://doi.org/10.1186/s12974-019-1467-7
Hamann J, Koning N, Pouwels W, Ulfman LH, van Eijk M, Stacey M, Lin HH, Gordon S, Kwakkenbos MJ (2007) EMR1, the human homolog of F4/80, is an eosinophil-specific receptor. Eur J Immunol 37(10):2797–2802. https://doi.org/10.1002/eji.200737553
Mii S, Uehara F, Yano S, Tran B, Miwa S, Hiroshima Y, Amoh Y, Katsuoka K, Hoffman RM (2013) Nestin-expressing stem cells promote nerve growth in long-term 3-dimensional gelfoam(R)-supported histoculture. PLoS ONE 8(6):e67153. https://doi.org/10.1371/journal.pone.0067153
Kang H, Tian L, Son YJ, Zuo Y, Procaccino D, Love F, Hayworth C, Trachtenberg J, Mikesh M, Sutton L, Ponomareva O, Mignone J, Enikolopov G, Rimer M, Thompson W (2007) Regulation of the intermediate filament protein nestin at rodent neuromuscular junctions by innervation and activity. J Neurosci 27(22):5948–5957. https://doi.org/10.1523/JNEUROSCI.0621-07.2007
Kang H, Tian L, Mikesh M, Lichtman JW, Thompson WJ (2014) Terminal Schwann cells participate in neuromuscular synapse remodeling during reinnervation following nerve injury. J Neurosci 34(18):6323–6333. https://doi.org/10.1523/JNEUROSCI.4673-13.2014
Isern J, Garcia-Garcia A, Martin AM, Arranz L, Martin-Perez D, Torroja C, Sanchez-Cabo F, Mendez-Ferrer S (2014) The neural crest is a source of mesenchymal stem cells with specialized hematopoietic stem cell niche function. Elife 3:e03696. https://doi.org/10.7554/eLife.03696
Goncalves NP, Jager SE, Richner M, Murray SS, Mohseni S, Jensen TS, Vaegter CB (2020) Schwann cell p75 neurotrophin receptor modulates small fiber degeneration in diabetic neuropathy. Glia 68(12):2725–2743. https://doi.org/10.1002/glia.23881
Tomita K, Kubo T, Matsuda K, Fujiwara T, Yano K, Winograd JM, Tohyama M, Hosokawa K (2007) The neurotrophin receptor p75NTR in Schwann cells is implicated in remyelination and motor recovery after peripheral nerve injury. Glia 55(11):1199–1208. https://doi.org/10.1002/glia.20533
Mora J, Cheung NK, Juan G, Illei P, Cheung I, Akram M, Chi S, Ladanyi M, Cordon-Cardo C, Gerald WL (2001) Neuroblastic and Schwannian stromal cells of neuroblastoma are derived from a tumoral progenitor cell. Cancer Res 61(18):6892–6898
Bourdeaut F, Ribeiro A, Paris R, Pierron G, Couturier J, Peuchmaur M, Delattre O (2008) In neuroblastic tumours, Schwann cells do not harbour the genetic alterations of neuroblasts but may nevertheless share the same clonal origin. Oncogene 27(21):3066–3071. https://doi.org/10.1038/sj.onc.1210965
Iyengar B, Singh AV (2010) Patterns of neural differentiation in melanomas. J Biomed Sci 17:87. https://doi.org/10.1186/1423-0127-17-87
Banerjee SS, Eyden B (2008) Divergent differentiation in malignant melanomas: a review. Histopathology 52(2):119–129. https://doi.org/10.1111/j.1365-2559.2007.02823.x
Van Raamsdonk CD, Deo M (2013) Links between Schwann cells and melanocytes in development and disease. Pigment Cell Melanoma Res 26(5):634–645. https://doi.org/10.1111/pcmr.12134
Venneri MA, De Palma M, Ponzoni M, Pucci F, Scielzo C, Zonari E, Mazzieri R, Doglioni C, Naldini L (2007) Identification of proangiogenic TIE2-expressing monocytes (TEMs) in human peripheral blood and cancer. Blood 109(12):5276–5285. https://doi.org/10.1182/blood-2006-10-053504
Yang M, McKay D, Pollard JW, Lewis CE (2018) Diverse functions of macrophages in different tumor microenvironments. Cancer Res 78(19):5492–5503. https://doi.org/10.1158/0008-5472.CAN-18-1367
Quante M, Tu SP, Tomita H, Gonda T, Wang SS, Takashi S, Baik GH, Shibata W, Diprete B, Betz KS, Friedman R, Varro A, Tycko B, Wang TC (2011) Bone marrow-derived myofibroblasts contribute to the mesenchymal stem cell niche and promote tumor growth. Cancer Cell 19(2):257–272. https://doi.org/10.1016/j.ccr.2011.01.020
Klopp AH, Spaeth EL, Dembinski JL, Woodward WA, Munshi A, Meyn RE, Cox JD, Andreeff M, Marini FC (2007) Tumor irradiation increases the recruitment of circulating mesenchymal stem cells into the tumor microenvironment. Cancer Res 67(24):11687–11695. https://doi.org/10.1158/0008-5472.CAN-07-1406
Hosaka K, Yang Y, Seki T, Fischer C, Dubey O, Fredlund E, Hartman J, Religa P, Morikawa H, Ishii Y, Sasahara M, Larsson O, Cossu G, Cao R, Lim S, Cao Y (2016) Pericyte-fibroblast transition promotes tumor growth and metastasis. Proc Natl Acad Sci USA 113(38):E5618-5627. https://doi.org/10.1073/pnas.1608384113
Cooke VG, LeBleu VS, Keskin D, Khan Z, O’Connell JT, Teng Y, Duncan MB, Xie L, Maeda G, Vong S, Sugimoto H, Rocha RM, Damascena A, Brentani RR, Kalluri R (2012) Pericyte depletion results in hypoxia-associated epithelial-to-mesenchymal transition and metastasis mediated by met signaling pathway. Cancer Cell 21(1):66–81. https://doi.org/10.1016/j.ccr.2011.11.024
Birbrair A, Zhang T, Wang ZM, Messi ML, Olson JD, Mintz A, Delbono O (2014) Type-2 pericytes participate in normal and tumoral angiogenesis. Am J Physiol Cell Physiol 307(1):C25-38. https://doi.org/10.1152/ajpcell.00084.2014
Sinha D, Chong L, George J, Schluter H, Monchgesang S, Mills S, Li J, Parish C, Bowtell D, Kaur P, Australian Ovarian Cancer Study G (2016) Pericytes promote malignant ovarian cancer progression in mice and predict poor prognosis in serous ovarian cancer patients. Clin Cancer Res 22(7):1813–1824. https://doi.org/10.1158/1078-0432.CCR-15-1931
Eilken HM, Dieguez-Hurtado R, Schmidt I, Nakayama M, Jeong HW, Arf H, Adams S, Ferrara N, Adams RH (2017) Pericytes regulate VEGF-induced endothelial sprouting through VEGFR1. Nat Commun 8(1):1574. https://doi.org/10.1038/s41467-017-01738-3
Bernardes SS, Pinto MCX, Amorim JH, Azevedo VAC, Resende RR, Mintz A, Birbrair A (2022) Glioma pericytes promote angiogenesis by producing periostin. Cell Mol Neurobiol 42(3):557–564. https://doi.org/10.1007/s10571-020-00975-3
Birbrair A, Zhang T, Wang ZM, Messi ML, Mintz A, Delbono O (2015) Pericytes at the intersection between tissue regeneration and pathology. Clin Sci (Lond) 128(2):81–93. https://doi.org/10.1042/CS20140278
Paiva AE, Lousado L, Guerra DAP, Azevedo PO, Sena IFG, Andreotti JP, Santos GSP, Goncalves R, Mintz A, Birbrair A (2018) Pericytes in the premetastatic niche. Cancer Res 78(11):2779–2786. https://doi.org/10.1158/0008-5472.CAN-17-3883
Sena IFG, Paiva AE, Prazeres P, Azevedo PO, Lousado L, Bhutia SK, Salmina AB, Mintz A, Birbrair A (2018) Glioblastoma-activated pericytes support tumor growth via immunosuppression. Cancer Med 7(4):1232–1239. https://doi.org/10.1002/cam4.1375
Valle IB, Schuch LF, da Silva JM, Gala-Garcia A, Diniz IMA, Birbrair A, Abreu LG, Silva TA (2020) Pericyte in oral squamous cell carcinoma: a systematic review. Head Neck Pathol 14(4):1080–1091. https://doi.org/10.1007/s12105-020-01188-2
Guerra DAP, Paiva AE, Sena IFG, Azevedo PO, Silva WN, Mintz A, Birbrair A (2018) Targeting glioblastoma-derived pericytes improves chemotherapeutic outcome. Angiogenesis 21(4):667–675. https://doi.org/10.1007/s10456-018-9621-x
Zhou BO, Yue R, Murphy MM, Peyer JG, Morrison SJ (2014) Leptin-receptor-expressing mesenchymal stromal cells represent the main source of bone formed by adult bone marrow. Cell Stem Cell 15(2):154–168. https://doi.org/10.1016/j.stem.2014.06.008
Picoli CC, Coimbra-Campos LMC, Guerra DAP, Silva WN, Prazeres PHDM, Costa AC, Magno LAV, Romano-Silva MA, Mintz A, Birbrair A (2019) Pericytes act as key players in spinal cord injury. Am J Pathol 189(7):1327–1337. https://doi.org/10.1016/j.ajpath.2019.03.008
Santos GSP, Magno LAV, Romano-Silva MA, Mintz A, Birbrair A (2019) Pericyte plasticity in the brain. Neurosci Bull 35(3):551–560. https://doi.org/10.1007/s12264-018-0296-5
Almeida VM, Paiva AE, Sena IFG, Mintz A, Magno LAV, Birbrair A (2018) Pericytes make spinal cord breathless after injury. Neuroscientis 24(5):440–447. https://doi.org/10.1177/1073858417731522
Dias Moura Prazeres PH, Sena IFG, Borges IDT, de Azevedo PO, Andreotti JP, de Paiva AE, de Almeida VM, de Paula Guerra DA, Pinheiro Dos Santos GS, Mintz A, Delbono O, Birbrair A (2017) Pericytes are heterogeneous in their origin within the same tissue. Dev Biol 427(1):6–11. https://doi.org/10.1016/j.ydbio.2017.05.001
Costa MA, Paiva AE, Andreotti JP, Cardoso MV, Cardoso CD, Mintz A, Birbrair A (2018) Pericytes constrict blood vessels after myocardial ischemia. J Mol Cell Cardiol 116:1–4. https://doi.org/10.1016/j.yjmcc.2018.01.014
Birbrair A, Zhang T, Wang ZM, Messi ML, Enikolopov GN, Mintz A, Delbono O (2013) Skeletal muscle pericyte subtypes differ in their differentiation potential. Stem Cell Res 10(1):67–84. https://doi.org/10.1016/j.scr.2012.09.003
Lee Y, Morrison BM, Li Y, Lengacher S, Farah MH, Hoffman PN, Liu Y, Tsingalia A, Jin L, Zhang PW, Pellerin L, Magistretti PJ, Rothstein JD (2012) Oligodendroglia metabolically support axons and contribute to neurodegeneration. Nature 487(7408):443–448. https://doi.org/10.1038/nature11314
Collier RJ (2001) Understanding the mode of action of diphtheria toxin: a perspective on progress during the 20th century. Toxicon 39(11):1793–1803. https://doi.org/10.1016/s0041-0101(01)00165-9
Pappenheimer AM Jr, Harper AA, Moynihan M, Brockes JP (1982) Diphtheria toxin and related proteins: effect of route of injection on toxicity and the determination of cytotoxicity for various cultured cells. J Infect Dis 145(1):94–102. https://doi.org/10.1093/infdis/145.1.94
Yamaizumi M, Mekada E, Uchida T, Okada Y (1978) One molecule of diphtheria toxin fragment A introduced into a cell can kill the cell. Cell 15(1):245–250. https://doi.org/10.1016/0092-8674(78)90099-5
Klages K, Mayer CT, Lahl K, Loddenkemper C, Teng MW, Ngiow SF, Smyth MJ, Hamann A, Huehn J, Sparwasser T (2010) Selective depletion of Foxp3+ regulatory T cells improves effective therapeutic vaccination against established melanoma. Cancer Res 70(20):7788–7799. https://doi.org/10.1158/0008-5472.CAN-10-1736
Lesokhin AM, Hohl TM, Kitano S, Cortez C, Hirschhorn-Cymerman D, Avogadri F, Rizzuto GA, Lazarus JJ, Pamer EG, Houghton AN, Merghoub T, Wolchok JD (2012) Monocytic CCR2(+) myeloid-derived suppressor cells promote immune escape by limiting activated CD8 T-cell infiltration into the tumor microenvironment. Cancer Res 72(4):876–886. https://doi.org/10.1158/0008-5472.CAN-11-1792
Li X, Kostareli E, Suffner J, Garbi N, Hammerling GJ (2010) Efficient Treg depletion induces T-cell infiltration and rejection of large tumors. Eur J Immunol 40(12):3325–3335. https://doi.org/10.1002/eji.201041093
Smyth MJ, Teng MW, Swann J, Kyparissoudis K, Godfrey DI, Hayakawa Y (2006) CD4+CD25+ T regulatory cells suppress NK cell-mediated immunotherapy of cancer. J Immunol 176(3):1582–1587. https://doi.org/10.4049/jimmunol.176.3.1582
Teng MW, Ngiow SF, von Scheidt B, McLaughlin N, Sparwasser T, Smyth MJ (2010) Conditional regulatory T-cell depletion releases adaptive immunity preventing carcinogenesis and suppressing established tumor growth. Cancer Res 70(20):7800–7809. https://doi.org/10.1158/0008-5472.CAN-10-1681
Teng MW, Swann JB, von Scheidt B, Sharkey J, Zerafa N, McLaughlin N, Yamaguchi T, Sakaguchi S, Darcy PK, Smyth MJ (2010) Multiple antitumor mechanisms downstream of prophylactic regulatory T-cell depletion. Cancer Res 70(7):2665–2674. https://doi.org/10.1158/0008-5472.CAN-09-1574
Yuan J, Adamow M, Ginsberg BA, Rasalan TS, Ritter E, Gallardo HF, Xu Y, Pogoriler E, Terzulli SL, Kuk D, Panageas KS, Ritter G, Sznol M, Halaban R, Jungbluth AA, Allison JP, Old LJ, Wolchok JD, Gnjatic S (2011) Integrated NY-ESO-1 antibody and CD8+ T-cell responses correlate with clinical benefit in advanced melanoma patients treated with ipilimumab. Proc Natl Acad Sci USA 108(40):16723–16728. https://doi.org/10.1073/pnas.1110814108
Yuan J, Ginsberg B, Page D, Li Y, Rasalan T, Gallardo HF, Xu Y, Adams S, Bhardwaj N, Busam K, Old LJ, Allison JP, Jungbluth A, Wolchok JD (2011) CTLA-4 blockade increases antigen-specific CD8(+) T cells in prevaccinated patients with melanoma: three cases. Cancer Immunol Immunother 60(8):1137–1146. https://doi.org/10.1007/s00262-011-1011-9
Fridman WH, Pages F, Sautes-Fridman C, Galon J (2012) The immune contexture in human tumours: impact on clinical outcome. Nat Rev Cancer 12(4):298–306. https://doi.org/10.1038/nrc3245
Rabinovich GA, Gabrilovich D, Sotomayor EM (2007) Immunosuppressive strategies that are mediated by tumor cells. Annu Rev Immunol 25:267–296. https://doi.org/10.1146/annurev.immunol.25.022106.141609
Tzekova N, Heinen A, Kury P (2014) Molecules involved in the crosstalk between immune- and peripheral nerve Schwann cells. J Clin Immunol 34(Suppl 1):S86-104. https://doi.org/10.1007/s10875-014-0015-6
Ydens E, Lornet G, Smits V, Goethals S, Timmerman V, Janssens S (2013) The neuroinflammatory role of Schwann cells in disease. Neurobiol Dis 55:95–103. https://doi.org/10.1016/j.nbd.2013.03.005
Goncalves NP, Vaegter CB, Andersen H, Ostergaard L, Calcutt NA, Jensen TS (2017) Schwann cell interactions with axons and microvessels in diabetic neuropathy. Nat Rev Neurol 13(3):135–147. https://doi.org/10.1038/nrneurol.2016.201
Neal JW, Gasque P (2016) The role of primary infection of Schwann cells in the aetiology of infective inflammatory neuropathies. J Infect 73(5):402–418. https://doi.org/10.1016/j.jinf.2016.08.006
Gabrilovich DI, Ciernik IF, Carbone DP (1996) Dendritic cells in antitumor immune responses. I. Defective antigen presentation in tumor-bearing hosts. Cell Immunol 170(1):101–110. https://doi.org/10.1006/cimm.1996.0139
Kim KD, Choi S-C, Kim A, Choe Y-K, Choe IS, Lim J-S (2001) Dendritic cell-tumor coculturing vaccine can induce antitumor immunity through both NK and CTL interaction. Int Immunopharmacol 1(12):2117–2129
Gabrilovich DI, Nadaf S, Corak J, Berzofsky JA, Carbone DP (1996) Dendritic cells in antitumor immune responses: II. Dendritic cells grown from bone marrow precursors, but not mature DC from tumor-bearing mice, are effective antigen carriers in the therapy of established tumors. Cell Immunol 170(1):111–119
Mrass P, Takano H, Ng LG, Daxini S, Lasaro MO, Iparraguirre A, Cavanagh LL, von Andrian UH, Ertl HC, Haydon PG, Weninger W (2006) Random migration precedes stable target cell interactions of tumor-infiltrating T cells. J Exp Med 203(12):2749–2761. https://doi.org/10.1084/jem.20060710
Alvarez-Dominguez C, Calderon-Gonzalez R, Teran-Navarro H, Salcines-Cuevas D, Garcia-Castano A, Freire J, Gomez-Roman J, Rivera F (2017) Dendritic cell therapy in melanoma. Ann Transl Med 5(19):386. https://doi.org/10.21037/atm.2017.06.13
Jacobs JF, Nierkens S, Figdor CG, de Vries IJ, Adema GJ (2012) Regulatory T cells in melanoma: the final hurdle towards effective immunotherapy? Lancet Oncol 13(1):e32-42. https://doi.org/10.1016/S1470-2045(11)70155-3
Sanmamed MF, Chen L (2018) A paradigm shift in cancer immunotherapy: from enhancement to normalization. Cell 175(2):313–326. https://doi.org/10.1016/j.cell.2018.09.035
Darvin P, Toor SM, Sasidharan Nair V, Elkord E (2018) Immune checkpoint inhibitors: recent progress and potential biomarkers. Exp Mol Med 50(12):1–11. https://doi.org/10.1038/s12276-018-0191-1
Hargadon KM, Johnson CE, Williams CJ (2018) Immune checkpoint blockade therapy for cancer: an overview of FDA-approved immune checkpoint inhibitors. Int Immunopharmacol 62:29–39. https://doi.org/10.1016/j.intimp.2018.06.001
Blank CU, Haining WN, Held W, Hogan PG, Kallies A, Lugli E, Lynn RC, Philip M, Rao A, Restifo NP, Schietinger A, Schumacher TN, Schwartzberg PL, Sharpe AH, Speiser DE, Wherry EJ, Youngblood BA, Zehn D (2019) Defining “T cell exhaustion.” Nat Rev Immunol 19(11):665–674. https://doi.org/10.1038/s41577-019-0221-9
Wherry EJ, Kurachi M (2015) Molecular and cellular insights into T cell exhaustion. Nat Rev Immunol 15(8):486–499. https://doi.org/10.1038/nri3862
Muranski P, Boni A, Antony PA, Cassard L, Irvine KR, Kaiser A, Paulos CM, Palmer DC, Touloukian CE, Ptak K, Gattinoni L, Wrzesinski C, Hinrichs CS, Kerstann KW, Feigenbaum L, Chan CC, Restifo NP (2008) Tumor-specific Th17-polarized cells eradicate large established melanoma. Blood 112(2):362–373. https://doi.org/10.1182/blood-2007-11-120998
Martin-Orozco N, Muranski P, Chung Y, Yang XO, Yamazaki T, Lu S, Hwu P, Restifo NP, Overwijk WW, Dong C (2009) T helper 17 cells promote cytotoxic T cell activation in tumor immunity. Immunity 31(5):787–798. https://doi.org/10.1016/j.immuni.2009.09.014
Safari-Alighiarloo N, Taghizadeh M, Rezaei-Tavirani M, Goliaei B, Peyvandi AA (2014) Protein-protein interaction networks (PPI) and complex diseases. Gastroenterol Hepatol Bed Bench 7(1):17–31
Taylor IW, Wrana JL (2012) Protein interaction networks in medicine and disease. Proteomics 12(10):1706–1716. https://doi.org/10.1002/pmic.201100594
Chen B, Fan W, Liu J, Wu FX (2014) Identifying protein complexes and functional modules–from static PPI networks to dynamic PPI networks. Brief Bioinform 15(2):177–194. https://doi.org/10.1093/bib/bbt039
Haack H, Hynes RO (2001) Integrin receptors are required for cell survival and proliferation during development of the peripheral glial lineage. Dev Biol 233(1):38–55. https://doi.org/10.1006/dbio.2001.0213
Lefcort F, Venstrom K, McDonald JA, Reichardt LF (1992) Regulation of expression of fibronectin and its receptor, alpha 5 beta 1, during development and regeneration of peripheral nerve. Development 116(3):767–782. https://doi.org/10.1242/dev.116.3.767
Peters JH, Chen GE, Hynes RO (1996) Fibronectin isoform distribution in the mouse. II. Differential distribution of the alternatively spliced EIIIB, EIIIA, and V segments in the adult mouse. Cell Adhes Commun 4(2):127–148. https://doi.org/10.3109/15419069609010767
McAllister SS, Gifford AM, Greiner AL, Kelleher SP, Saelzler MP, Ince TA, Reinhardt F, Harris LN, Hylander BL, Repasky EA, Weinberg RA (2008) Systemic endocrine instigation of indolent tumor growth requires osteopontin. Cell 133(6):994–1005. https://doi.org/10.1016/j.cell.2008.04.045
Kaplan RN, Riba RD, Zacharoulis S, Bramley AH, Vincent L, Costa C, MacDonald DD, Jin DK, Shido K, Kerns SA, Zhu Z, Hicklin D, Wu Y, Port JL, Altorki N, Port ER, Ruggero D, Shmelkov SV, Jensen KK, Rafii S, Lyden D (2005) VEGFR1-positive haematopoietic bone marrow progenitors initiate the pre-metastatic niche. Nature 438(7069):820–827. https://doi.org/10.1038/nature04186
Johansson M, Denardo DG, Coussens LM (2008) Polarized immune responses differentially regulate cancer development. Immunol Rev 222:145–154. https://doi.org/10.1111/j.1600-065X.2008.00600.x
Gatenby RA, Gillies RJ (2008) A microenvironmental model of carcinogenesis. Nat Rev Cancer 8(1):56–61. https://doi.org/10.1038/nrc2255
Rodella U, Negro S, Scorzeto M, Bergamin E, Jalink K, Montecucco C, Yuki N, Rigoni M (2017) Schwann cells are activated by ATP released from neurons in an in vitro cellular model of Miller Fisher syndrome. Dis Model Mech 10(5):597–603. https://doi.org/10.1242/dmm.027870
Campana WM (2007) Schwann cells: activated peripheral glia and their role in neuropathic pain. Brain Behav Immun 21(5):522–527. https://doi.org/10.1016/j.bbi.2006.12.008
Yang DP, Zhang DP, Mak KS, Bonder DE, Pomeroy SL, Kim HA (2008) Schwann cell proliferation during Wallerian degeneration is not necessary for regeneration and remyelination of the peripheral nerves: axon-dependent removal of newly generated Schwann cells by apoptosis. Mol Cell Neurosci 38(1):80–88. https://doi.org/10.1016/j.mcn.2008.01.017
Kadioglu E, De Palma M (2015) Cancer metastasis: perivascular macrophages under watch. Cancer Discov 5(9):906–908. https://doi.org/10.1158/2159-8290.CD-15-0819
Harney AS, Arwert EN, Entenberg D, Wang Y, Guo P, Qian BZ, Oktay MH, Pollard JW, Jones JG, Condeelis JS (2015) Real-time imaging reveals local, transient vascular permeability, and tumor cell intravasation stimulated by TIE2hi macrophage-derived VEGFA. Cancer Discov 5(9):932–943. https://doi.org/10.1158/2159-8290.CD-15-0012
Manberg A, Skene N, Sanders F, Trusohamn M, Remnestal J, Szczepinska A, Aksoylu IS, Lonnerberg P, Ebarasi L, Wouters S, Lehmann M, Olofsson J, von Gohren AI, Domaniku A, De Schaepdryver M, De Vocht J, Poesen K, Uhlen M, Anink J, Mijnsbergen C, Vergunst-Bosch H, Hubers A, Klappe U, Rodriguez-Vieitez E, Gilthorpe JD, Hedlund E, Harris RA, Aronica E, Van Damme P, Ludolph A, Veldink J, Ingre C, Nilsson P, Lewandowski SA (2021) Altered perivascular fibroblast activity precedes ALS disease onset. Nat Med 27(4):640–646. https://doi.org/10.1038/s41591-021-01295-9
Soderblom C, Luo X, Blumenthal E, Bray E, Lyapichev K, Ramos J, Krishnan V, Lai-Hsu C, Park KK, Tsoulfas P, Lee JK (2013) Perivascular fibroblasts form the fibrotic scar after contusive spinal cord injury. J Neurosci 33(34):13882–13887. https://doi.org/10.1523/JNEUROSCI.2524-13.2013
Crisan M, Corselli M, Chen WC, Peault B (2012) Perivascular cells for regenerative medicine. J Cell Mol Med. https://doi.org/10.1111/j.1582-4934.2012.01617.x
Wanjare M, Kusuma S, Gerecht S (2013) Perivascular cells in blood vessel regeneration. Biotechnol J 8(4):434–447. https://doi.org/10.1002/biot.201200199
Bhatia A, Kumar Y (2011) Cancer-immune equilibrium: questions unanswered. Cancer Microenviron 4(2):209–217. https://doi.org/10.1007/s12307-011-0065-8
Passarelli A, Mannavola F, Stucci LS, Tucci M, Silvestris F (2017) Immune system and melanoma biology: a balance between immunosurveillance and immune escape. Oncotarget 8(62):106132–106142. https://doi.org/10.18632/oncotarget.22190
Mittal D, Gubin MM, Schreiber RD, Smyth MJ (2014) New insights into cancer immunoediting and its three component phases–elimination, equilibrium and escape. Curr Opin Immunol 27:16–25. https://doi.org/10.1016/j.coi.2014.01.004
Gooden MJ, de Bock GH, Leffers N, Daemen T, Nijman HW (2011) The prognostic influence of tumour-infiltrating lymphocytes in cancer: a systematic review with meta-analysis. Br J Cancer 105(1):93–103. https://doi.org/10.1038/bjc.2011.189
Girardi M, Oppenheim DE, Steele CR, Lewis JM, Glusac E, Filler R, Hobby P, Sutton B, Tigelaar RE, Hayday AC (2001) Regulation of cutaneous malignancy by gammadelta T cells. Science 294(5542):605–609. https://doi.org/10.1126/science.1063916
He W, Hao J, Dong S, Gao Y, Tao J, Chi H, Flavell R, O’Brien RL, Born WK, Craft J, Han J, Wang P, Zhao L, Wu J, Yin Z (2010) Naturally activated V gamma 4 gamma delta T cells play a protective role in tumor immunity through expression of eomesodermin. J Immunol 185(1):126–133. https://doi.org/10.4049/jimmunol.0903767
Rosenberg SA, Packard BS, Aebersold PM, Solomon D, Topalian SL, Toy ST, Simon P, Lotze MT, Yang JC, Seipp CA et al (1988) Use of tumor-infiltrating lymphocytes and interleukin-2 in the immunotherapy of patients with metastatic melanoma. A preliminary report. N Engl J Med 319(25):1676–1680. https://doi.org/10.1056/NEJM198812223192527
Gerber AL, Munst A, Schlapbach C, Shafighi M, Kiermeir D, Husler R, Hunger RE (2014) High expression of FOXP3 in primary melanoma is associated with tumour progression. Br J Dermatol 170(1):103–109. https://doi.org/10.1111/bjd.12641
Huntington ND, Vosshenrich CA, Di Santo JP (2007) Developmental pathways that generate natural-killer-cell diversity in mice and humans. Nat Rev Immunol 7(9):703–714. https://doi.org/10.1038/nri2154
Kobayashi N, Hiraoka N, Yamagami W, Ojima H, Kanai Y, Kosuge T, Nakajima A, Hirohashi S (2007) FOXP3+ regulatory T cells affect the development and progression of hepatocarcinogenesis. Clin Cancer Res 13(3):902–911. https://doi.org/10.1158/1078-0432.CCR-06-2363
Sacerdote P, Franchi S, Trovato AE, Valsecchi AE, Panerai AE, Colleoni M (2008) Transient early expression of TNF-alpha in sciatic nerve and dorsal root ganglia in a mouse model of painful peripheral neuropathy. Neurosci Lett 436(2):210–213. https://doi.org/10.1016/j.neulet.2008.03.023
Martucci C, Trovato AE, Costa B, Borsani E, Franchi S, Magnaghi V, Panerai AE, Rodella LF, Valsecchi AE, Sacerdote P, Colleoni M (2008) The purinergic antagonist PPADS reduces pain related behaviours and interleukin-1 beta, interleukin-6, iNOS and nNOS overproduction in central and peripheral nervous system after peripheral neuropathy in mice. Pain 137(1):81–95. https://doi.org/10.1016/j.pain.2007.08.017
Franchi S, Valsecchi AE, Borsani E, Procacci P, Ferrari D, Zaffa C, Sartori P, Rodella LF, Vescovi A, Maione S, Rossi F, Sacerdote P, Colleoni M, Panerai AE (2012) Intravenous neural stem cells abolish nociceptive hypersensitivity and trigger nerve regeneration in experimental neuropathy. Pain 153(4):850–861. https://doi.org/10.1016/j.pain.2012.01.008
Wang ZH, Zeng XY, Han SP, Fan GX, Wang JY (2012) Interleukin-10 of red nucleus plays anti-allodynia effect in neuropathic pain rats with spared nerve injury. Neurochem Res 37(8):1811–1819. https://doi.org/10.1007/s11064-012-0795-0
Keswani SC, Buldanlioglu U, Fischer A, Reed N, Polley M, Liang H, Zhou C, Jack C, Leitz GJ, Hoke A (2004) A novel endogenous erythropoietin mediated pathway prevents axonal degeneration. Ann Neurol 56(6):815–826. https://doi.org/10.1002/ana.20285
Campana WM, Li X, Shubayev VI, Angert M, Cai K, Myers RR (2006) Erythropoietin reduces Schwann cell TNF-alpha, Wallerian degeneration and pain-related behaviors after peripheral nerve injury. Eur J Neurosci 23(3):617–626. https://doi.org/10.1111/j.1460-9568.2006.04606.x
Takahashi M, Kawaguchi M, Shimada K, Konishi N, Furuya H, Nakashima T (2004) Cyclooxygenase-2 expression in Schwann cells and macrophages in the sciatic nerve after single spinal nerve injury in rats. Neurosci Lett 363(3):203–206. https://doi.org/10.1016/j.neulet.2004.03.040
Toews AD, Barrett C, Morell P (1998) Monocyte chemoattractant protein 1 is responsible for macrophage recruitment following injury to sciatic nerve. J Neurosci Res 53(2):260–267. https://doi.org/10.1002/(SICI)1097-4547(19980715)53:2%3c260::AID-JNR15%3e3.0.CO;2-A
Yajima Y, Narita M, Usui A, Kaneko C, Miyatake M, Narita M, Yamaguchi T, Tamaki H, Wachi H, Seyama Y, Suzuki T (2005) Direct evidence for the involvement of brain-derived neurotrophic factor in the development of a neuropathic pain-like state in mice. J Neurochem 93(3):584–594. https://doi.org/10.1111/j.1471-4159.2005.03045.x
Su WF, Wu F, Jin ZH, Gu Y, Chen YT, Fei Y, Chen H, Wang YX, Xing LY, Zhao YY, Yuan Y, Tang X, Chen G (2019) Overexpression of P2X4 receptor in Schwann cells promotes motor and sensory functional recovery and remyelination via BDNF secretion after nerve injury. Glia 67(1):78–90. https://doi.org/10.1002/glia.23527
Roger E, Martel S, Bertrand-Chapel A, Depollier A, Chuvin N, Pommier RM, Yacoub K, Caligaris C, Cardot-Ruffino V, Chauvet V, Aires S, Mohkam K, Mabrut JY, Adham M, Fenouil T, Hervieu V, Broutier L, Castets M, Neuzillet C, Cassier PA, Tomasini R, Sentis S, Bartholin L (2019) Schwann cells support oncogenic potential of pancreatic cancer cells through TGFbeta signaling. Cell Death Dis 10(12):886. https://doi.org/10.1038/s41419-019-2116-x
Ferdoushi A, Li X, Griffin N, Faulkner S, Jamaluddin MFB, Gao F, Jiang CC, van Helden DF, Tanwar PS, Jobling P, Hondermarck H (2020) Schwann cell stimulation of pancreatic cancer cells: a proteomic analysis. Front Oncol 10:1601. https://doi.org/10.3389/fonc.2020.01601
Salvo E, Tu NH, Scheff NN, Dubeykovskaya ZA, Chavan SA, Aouizerat BE, Ye Y (2021) TNFalpha promotes oral cancer growth, pain, and Schwann cell activation. Sci Rep 11(1):1840. https://doi.org/10.1038/s41598-021-81500-4
Ivanova E, Corona C, Eleftheriou CG, Bianchimano P, Sagdullaev BT (2021) Retina-specific targeting of pericytes reveals structural diversity and enables control of capillary blood flow. J Comp Neurol 529(6):1121–1134. https://doi.org/10.1002/cne.25011
Acknowledgements
Alexander Birbrair is supported by a research productivity fellowship from Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq-PQ2), a grant from Instituto Serrapilheira/Serra-1708-15285, a Grant from Pró-reitoria de Pesquisa/Universidade Federal de Minas Gerais (PRPq/UFMG) (Edital 05/2016); a grant from Fundação de Amparo à Pesquisa do Estado de Minas Gerais—FAPEMIG (Chamada N°01/2021—Demanda Universal, APQ-01321-21); a grant from FAPEMIG [Rede Mineira de Pesquisa Translacional em Imunobiológicos e Biofármacos no Câncer (REMITRIBIC, RED-00031-21)]; a grant from FAPEMIG [Rede Mineira de Engenharia de Tecidos e Terapia Celular (REMETTEC, RED-00570-16)]; a grant from FAPEMIG [Rede De Pesquisa Em Doenças Infecciosas Humanas E Animais Do Estado De Minas Gerais (RED-00313-16)]; and a grant from MCTIC/CNPq Nº 28/2018 (Universal/Faixa A). Akiva Mintz is supported by the National Institute of Health (1R01CA179072-01A1) and by the American Cancer Society Mentored Research Scholar grant (124443-MRSG-13-121-01-CDD). Edroaldo Lummertz da Rocha is supported by the Coordination for the Improvement of Higher Education Personnel (CAPES), the National Council of State Funding Agencies (CONFAP), the Serrapilheira Institute and the Foundation for Support of Research and Innovation of Santa Catarina (FAPESC). Marcelo Falchetti is supported by a postdoctoral fellowship from the Brazilian National Council for Scientific and Technological Development (CNPq), Brazil. Remo C. Russo is supported by a research productivity fellowship from Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq-PQ2) and a Grant from FAPEMIG (Chamada N°01/2021 – Demanda Universal, APQ-02571-21). Pedro A F Galante was supported by a research productivity fellowship from Conselho Nacional de Desenvolvimento Científico e Tecnológico, a grant from Instituto Serrapilheira, and a grant from São Paulo Research Foundation (FAPESP), grant 2012/24731-1. Gabriela D. A. Guardia was supported by São Paulo Research Foundation (FAPESP), grant 2017/19541-2, and a fellowship from Hospital Sirio-Libanês, Young Scientist initiative. Caroline C. Picoli and Alinne C. Costa are supported by doctoral fellowships from CAPES. Bryan O. P. Gonçalves is supported by a doctoral fellowship from FAPEMIG. Gabryella S.P. Santos is supported by a doctoral fellowship from CNPq. Beatriz G. S. Rocha and Walison N. Silva are supported by master fellowships from CAPES. Milla R. Almeida is supported by a scientific initiation fellowship from CNPq. Pedro A. C. Costa is supported by a postdoctoral fellowship (PNPD) from CAPES. The authors also thank CAPI (UFMG) for microscopical technical support and Laboratory of Flow Cytometry at the Instituto de Ciências Biológicas/UFMG (http://labs.icb.ufmg.br/citometria/)” for providing the equipment and technical support for experiments involving flow cytometry.
Author information
Authors and Affiliations
Contributions
AB conceived and supervised the study; BGSR, CCP, BOPG, WNS, ACC, MMM, PACC, GSPS, MRA, LMS, YS, MF, GDAG, PPGG, RCR, RRR, MCXP, JHA, VACA, AK, HIN, ELR, PAFG, AB analyzed the data and discussed the results; AB was responsible for funding; AM, PSF, AB wrote the original draft; all authors contributed to and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors indicate no potential conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Beatriz G. S. Rocha, Caroline C. Picoli and Bryan O. P. Gonçalves are co-first authors.
Supplementary Information
Below is the link to the electronic supplementary material.
10456_2022_9858_MOESM7_ESM.tif
Supplementary Figure 1. Nerves infiltrate within the tumor microenvironment. A. Adult wild-type mice were orthotopically injected with Tramp-C2 prostate cancer cells. The tumors were surgically removed 2 weeks later for analysis. B. Percentage of blood vessels with or without Peripherin+ nerve fibers attached to them in the Tramp-C2 prostate cancer microenvironment (n=3 mice) (81.25 ± 4.54 % of blood vessels were not associated to Peripherin+ nerve fibers; p < 0.0001; ES =15.7L). C and D. Representative images of sections from orthotopic Tramp-C2 tumors show nerves within the prostate tumor microenvironment. C. All panels show the same area for different channels: Peripherin (marker of peripheral nerve fibers), CD31 (marker of endothelial cells), DAPI, and all three merged. The area in the white box is magnified showing a nerve fiber attached to a blood vessel within the prostate tumor microenvironment. D. All panels show the same area for different channels: PGP9.5 (neuronal marker), CD31, DAPI, and merged. E. Percentage of blood vessels with or without PGP9.5+ nerve fibers attached to them in the Tramp-C2 prostate cancer microenvironment (n=3 mice) (79.7 ± 6.2 % of blood vessels were not associated to PGP9.5+ nerve fibers; p < 0.0001; ES =9.8L). F, G and H. Sympathetic nerves are present in the prostate tumor microenvironment. F. Intra-prostatic injection of TdTomato-labeled PC-3 human prostate cancer cells in nude mice, and tumor analysis after 3 weeks. G. Representative photomicrographs of a prostate tumor section 3 weeks after PC3 cells injection, showing blood vessels with and without attached TH+ nerve fibers. H. Percentage of blood vessels with or without attached TH+ nerve fibers in the PC3 prostate tumor microenvironment after 3 weeks (n=3 mice) (83.6 ± 4.2 % of blood vessels were not associated to TH+ nerve fibers; p < 0.0001; ES =5.4L). Data are mean ± SEM. TH, tyrosine hydroxylase, sympathetic neuron marker. Statistical analysis: unpaired Student's t-tests. ES: effect size; Llarge (≥ 1.2). ***p <0.001. Scale bars, 50µm (C and D) and 10 µm (G). (TIF 5864 kb)
10456_2022_9858_MOESM8_ESM.tif
Supplementary Figure 2. GFAP+ cells are present in the microenvironment of syngeneic C57BL6 mouse prostate tumors. A. Adult wild-type mice orthotopically injected with Tramp-C2 prostate cancer cells. B. Percentages of GFAP+ cells attached or not to blood vessels in the Tramp-C2 tumor after 2 weeks (n=3 mice) (93.7 ± 2.7 % of GFAP+ cells were associated to blood vessels, while 6.3 ± 2.7 % were not associated to blood vessels; p < 0.0001; ES =21.75L). C. Prostate Tramp-C2 tumor surgically removed after 2 weeks. D. Representative image of a section from orthotopic Tramp-C2 tumor shows blood vessel-associated GFAP+ cells in C57BL/6 mice. All panels show the same area for different channels (GFAP, CD31, and the two images merged with DAPI). E, F and G. GFAP+ cells are present within RM1 prostate tumor microenvironment associated with blood vessels. E. Intra-prostatic injection of RM1 prostate cancer cell line (from Ras+Myc transformed mouse prostate carcinoma) in wild-type mice, and tumor analysis after 2 weeks. F. Percentages of GFAP+ cells attached or not to blood vessels in the RM1 tumor after 2 weeks (n=3 mice) (91.0 ± 5.7 % of GFAP+ cells were associated to blood vessels, while 9.0 ± 5.6 % were not associated to blood vessels; p < 0.0001; ES =9.11L). G. Representative photomicrographs of a prostate tumor section 2 weeks after RM1 cells injection, showing blood vessels with GFAP+ cells attached to it. Statistical analysis: unpaired Student's t-tests. ES: effect size; Llarge (≥ 1.2). ***p <0.001. Data are mean ± SEM. Scale bars, 10µm. (TIF 3503 kb)
10456_2022_9858_MOESM9_ESM.tif
Supplementary Figure 3. Tumor-infiltrating perivascular glial cells differ from pericytes. A. Representative photomicrographs of prostate tumor sections 2 weeks after Tramp-C2 cells orthotopic injection into Nestin-GFP/NG2-DsRed mice showing GFAP+ cells (blue). B. Percentage of NG2-DsRed+ cells expressing GFAP in the Tramp-C2 prostate tumor after 2 weeks (n=5 mice) (23.60 ± 4.69 % of NG2-DsRed+ cells were positive for GFAP; p < 0.001; ES = 5.3L). C. Representative photomicrographs of melanoma tumor sections 2 weeks after B16F10 cancer cells transplantation into Plp1-CreER/TdTom mice showing perivascular Plp1CreER+/TdTomato+ cells (red) not expressing PDGFRβ (green). D. Percentage of Plp1CreER+/TdTomato+ cells not expressing PDGFRβ in the B16F10 tumor after 2 weeks (n=5 mice) (99.9 ± 0.10 % of Plp1CreER+/TdTomato+ cells were negative for PDGFRβ; p < 0.001; ES = 998.0L). Statistical analysis: unpaired Student's t-tests. ES: effect size; Llarge (≥ 1.2). ***p <0.001. Data are mean ± SEM. Scale bars, 10µm. (TIF 3434 kb)
10456_2022_9858_MOESM10_ESM.tif
Supplementary Figure 4. Ablation of Schwann cells increases the number of tumor-infiltrating dendritic cells. Dendritic cells from B16F10–inoculated mice were analyzed ex vivo in Plp1-CreER-/iDTR+ (n = 6 mice) and Plp1-CreER+/iDTR+ (n = 9 mice) mice. A. Absolute number of dendritic cells from the melanomas of B16F10–inoculated mice (Plp1-CreER-/iDTR+: 2.72x106 ± 7.40x105 cells per mg of tumor; Plp1-CreER+/iDTR+: 8.60x106 ± 2.21x106 cells per mg of tumor; p=0.039; ES = 1.2L). B. Absolute number of HLA-DR+ dendritic cells dendritic cells from the melanomas of B16F10–inoculated mice (Plp1-CreER-/iDTR+: 2.27x107 ± 2.87x106 cells per mg of tumor; Plp1-CreER+/iDTR+: 5.01x107± 9.40x106 cells per mg of tumor; p = 0.029; ES = 1.3L). Statistical analysis: unpaired Student's t-tests one-tailed. ES: effect size; Llarge (≥ 1.2). Plp1-CreER-/iDTR+ (n = 6 mice) and Plp1-CreER+/iDTR+ (n = 9 mice). * p <0.05. Data are mean ± SEM. (TIF 526 kb)
10456_2022_9858_MOESM11_ESM.tif
Supplementary Figure 5. Tumor-infiltrating Treg cells are reduced by Schwann cell ablation. Regulatory T cells from B16F10–inoculated mice were analyzed ex vivo in Plp1-CreER-/iDTR+ (n = 6 mice) and Plp1-CreER+/iDTR+ (n = 9 mice) mice. A. Absolute number of Treg cells from the melanomas of B16F10–inoculated mice (Plp1-CreER-/iDTR+: 4.25x106 ± 7.18x105 cells per mg of tumor; Plp1-CreER+/iDTR+: 1.48x106 ± 4.20x105 cells per mg of tumor, p=0.006; ES = 1.8L). Column charts show proportion of CTLA-4 (B), PD-1 (C) and CTLA-4/PD-1 co-expressing (D) Treg cells from tumors of B16F10–inoculated mice. B. CTLA-4+ Regulatory T cells (Plp1-CreER-/iDTR+: 14.35 ± 1.95%; Plp1-CreER+/iDTR+: 17.06 ± 2.03%; p = 0.363; ES = 0.5S). C. PD-1+ Regulatory T cells (Plp1-CreER-/iDTR+: 19.10 ± 3.72%; Plp1-CreER+/iDTR+: 9.24 ± 3.31%; p = 0.036; ES = 1.1M). D. and CTLA-4+/PD-1+ Regulatory T cells (Plp1-CreER-/iDTR+: 4.01 ± 0.88%; Plp1-CreER+/iDTR+: 2.67 ± 0.70%; p = 0.1305; ES = 0.7M). Statistical analysis: unpaired Student's t-tests or Mann-Whitney Rank Sum Test one-tail. ES: effect size; Ttrivial (< 0.2); Ssmall (0.2–0.6); Mmedium (0.6–1.2); Llarge (≥ 1.2). *p<0.05 and **p<0.01. Data are mean ± SEM. (TIF 553 kb)
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Rocha, B.G.S., Picoli, C.C., Gonçalves, B.O.P. et al. Tissue-resident glial cells associate with tumoral vasculature and promote cancer progression. Angiogenesis 26, 129–166 (2023). https://doi.org/10.1007/s10456-022-09858-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10456-022-09858-1