Abstract
Abiotic stresses are detrimental to plant growth and development and have a major negative impact on crop yields. A growing body of evidence indicates that a large number of long non-coding RNAs (lncRNAs) are key to many abiotic stress responses. Thus, identifying abiotic stress-responsive lncRNAs is essential in crop breeding programs in order to develop crop cultivars resistant to abiotic stresses. In this study, we have developed the first machine learning-based computational model for predicting abiotic stress-responsive lncRNAs. The lncRNA sequences which were responsive and non-responsive to abiotic stresses served as the two classes of the dataset for binary classification using the machine learning algorithms. The training dataset was created using 263 stress-responsive and 263 non-stress-responsive sequences, whereas the independent test set consists of 101 sequences from both classes. As the machine learning model can adopt only the numeric data, the Kmer features ranging from sizes 1 to 6 were utilized to represent lncRNAs in numeric form. To select important features, four different feature selection strategies were utilized. Among the seven learning algorithms, the support vector machine (SVM) achieved the highest cross-validation accuracy with the selected feature sets. The observed 5-fold cross-validation accuracy, AU-ROC, and AU-PRC were found to be 68.84, 72.78, and 75.86%, respectively. Furthermore, the robustness of the developed model (SVM with the selected feature) was evaluated using an independent test dataset, where the overall accuracy, AU-ROC, and AU-PRC were found to be 76.23, 87.71, and 88.49%, respectively. The developed computational approach was also implemented in an online prediction tool ASLncR accessible at https://iasri-sg.icar.gov.in/aslncr/. The proposed computational model and the developed prediction tool are believed to supplement the existing effort for the identification of abiotic stress-responsive lncRNAs in plants.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Due to the growing world population, demand is going to be increased in global food consumption, and by 2050, that demand is expected to be doubled (Tilman et al. 2011). Abiotic stresses, on the other hand, present a substantial challenge to agriculture and the ecosystem due to changing climatic conditions, resulting in significant crop yield loss (Saeed et al., 2023; Wani et al., 2016). In order to adapt to challenging environmental conditions, plants modify the expression of several genes at the transcriptional, post-transcriptional, and epigenome levels in response to different abiotic stresses (Liu et al. 2022a; Choudhury et al. 2021; Zhu et al., 2022). The functional elucidation of many genes at the transcription, post-transcriptional, post-translational, and epigenetic levels has been significantly improved with the advancement in genome sequencing technology, especially next-generation sequencing (NGS) (Li et al. 2018). The NGS technologies have led to the identification of novel non-coding RNAs (ncRNAs) (Öztürk Gökçe et al., 2021; Bhogireddy et al., 2021; Yu et al. 2019) and their roles in the regulation of multiple biological processes, including plant response to various abiotic stresses (Yang et al. 2023;Yu et al. 2019).
The long non-coding RNAs (lncRNAs) are a group of ncRNAs which are more than 200 bp long and not be translated into a protein (Quan et al. 2015). Transcriptional, post-transcriptional, and epigenetic regulations of gene expression are three ways that lncRNA acts as a gene regulatory factor (Quan et al. 2015). The lncRNAs are reported to be important modulators of various biological processes (Mercer et al., 2009). Their involvement in controlling transcription through enhancers and providing regulatory binding sites has been well documented (Wang and Chekanova, 2017). These are also said to act as miRNA sponges, suppressing miRNA function by causing deflection to their potential target (Wang et al. 2010). The lncRNAs are also found in the nucleus, where they serve as major components of nuclear speckles (Hutchinson et al. 2007). In the cytoplasm, lncRNAs interact with a variety of RNA-binding proteins (RBPs) to monitor and control their regulatory dynamics (Glisovic et al. 2008).
Plant lncRNAs make up around 80% of all ncRNAs and are involved in a wide range of biological processes, including abiotic stress response (Wang et al. 2021). The first lncRNA reported in plants was ENOD40 in Soybean (Yang et al. 1993). Despite the fact that the plant genomes are more complicated than animal genomes, the number of experimentally identified lncRNAs in plants are much less than that reported for animals. Several lncRNAs that respond to abiotic stresses have been reported to be present in a wide range of plant species. Table 1 contains a list of recently identified lncRNAs reported to be involved in various abiotic stresses. Due to the discovery of abiotic stress-responsive lncRNAs and their target genes in a range of plant species, we now have a better understanding of the molecular mechanism underlying these stress adaptations. For example, in drought conditions of Arabidopsis thaliana, lncRNA lincRNA340 is induced to repress miR169, relieving nuclear factor Y (NF-Y) gene expression to improve stress tolerance (Qin et al. 2017). Further, lncRNA973 functions as a positive regulator of salt-responsive genes in ROS (reactive oxygen species), enhancing salinity tolerance in cotton (Zhang et al., 2019). Similarly, GhDNA1, which targets AAAG DNA double strands to regulate drought-responsive genes in trans, was discovered to be associated with drought tolerance in cotton (Tao et al., 2021). These findings support the idea that lncRNAs can be induced or suppressed in response to abiotic stress. Furthermore, these abiotic stress-responsive lncRNAs have been linked to phytohormone signal transduction, secondary metabolite biosynthesis, and sucrose metabolism pathways, each of which has been reportedly engaged in plant abiotic stress response (Ding et al. 2019; Yang et al., 2022; Lamin-Samu et al., 2022).
The studies cited above indicate that lncRNAs may be exploited as genetic targets to develop crop cultivars that are resistant to abiotic stresses. However, the lncRNAs are needed to be identified first before using them as genetic targets. To date, techniques such as serial expression of gene expression (SAGE), the expressed sequence tag (EST), whole-genome tiling arrays, lncRNA microarray, RNA capture sequencing (RNA CaptureSeq), and RNA-sequencing (RNA-seq) have all been employed to identify abiotic stress-related lncRNAs. However, the wet-lab experiments consume a lot of resources (Lee and Kikyo 2012). Furthermore, the advanced sequencing techniques are species-specific. Thus, there is a need to develop a computational method for predicting abiotic stress-responsive lncRNAs using lncRNA sequence data. In other words, the development of machine learning-based computational methods may be a better alternative for predicting lncRNAs associated with abiotic stress. Considering the above facts, the present study is devoted to develop the first machine learning-based computational model for predicting abiotic stress-responsive lncRNAs using sequence-derived features. The proposed approach is expected to supplement wet-lab methods and other sequencing techniques for identifying abiotic stress-responsive lncRNAs in plants.
Materials and methods
Collection of abiotic stress-responsive lncRNA sequence data
The PncStress database (Wu et al., 2020) is the most recent source for abiotic stress-responsive lncRNAs. It contains experimentally validated ncRNA sequences linked to a variety of abiotic and biotic stresses. With 114 species responding to 48 abiotic and 91 biotic stresses, PncStress now has 4227 entries, including 2523 miRNAs, 444 lncRNAs, and 52 circRNAs validated by different experimental methods. The PncStress database (Wu et al., 2020) was accessed on July 30, 2022, in order to retrieve lncRNA sequences relevant to abiotic stresses. A total of 444 abiotic stress-responsive lncRNA sequences, representing 27 different abiotic stress categories, were obtained from 24 plant species.
Construction of positive and negative dataset
The abiotic stress-responsive lncRNA sequences obtained from the PncStress database were used to construct the positive set. On the other hand, 238,226 lncRNA sequences retrieved from the PLncDB V2.0 database (accessed on August 05, 2022) (Jin et al., 2021) were used to construct the negative set. To prevent homologous bias in the prediction accuracy, the homology reduction at 50% sequence identity was applied to both positive and negative datasets using the CD-HIT method (Huang et al., 2010). After the redundancy sequences were removed, the positive and negative sets produced 364 and 97,654 lncRNA sequences, respectively. To avoid prediction bias toward the non-abiotic stress class having a larger number of sequences, a balanced dataset with an equal number of abiotic stress and non-abiotic stress-responsive lncRNA sequences was taken into consideration. In other words, 364 non-abiotic stress sequences were chosen at random from the pool of 97,654 sequences to prepare a balanced training dataset that comprises an equal number of sequences from both classes. Out of the 364 sequences in each class, 101 lncRNA sequences were kept aside to prepare the independent dataset. The remaining 263 stress-responsive lncRNAs and 263 non-stress-responsive lncRNAs were used as positive and negative sets for the training dataset.
Numeric feature generation
In this study, we generated Kmer features to transform each lncRNA sequence into a numeric feature vector. The Kmer features are represented as the occurrence frequencies of K neighboring nucleic acids (Lee et al. 2011), which has been successfully used in several computational studies including lncRNA prediction (Sun et al. 2013). The numeric value for the Kmer size k can be calculated as
where Nk(t) is the number of Kmer type t of size k, and N is the length of the nucleotide sequence. For example, for an RNA sequence ‘CUGACUGACUGACUGUA’, \({f}_1(C)=\frac{4}{17}\), \({f}_2(CU)=\frac{4}{16}\), \({f}_3(CUG)=\frac{4}{15},\) \({f}_4(CUGA)=\frac{3}{14}\), \({f}_5(CUGAC)=\frac{3}{13}\), and \({f}_6(CUGACU)=\frac{3}{12}\). A brief representation of the Kmer feature is shown in Fig. 1. The number of Kmer features of size k is 4k. In this study, we have considered Kmer sizes 1 to 6 to generate the features for each sequence. Thus, for Kmer sizes 1, 2, 3, 4, 5, and 6, the number of features generated was 4, 16, 64, 256, 1024, and 4096, respectively. The Kmer sizes 1 to 6 were denoted as K1, K2, K3, K4, K5, and K6. In total, 5460 features were generated for each lncRNA sequence.
Prediction algorithms
Several bioinformatics fields have effectively applied machine learning techniques for prediction purposes (Guo et al. 2017, Pradhan et al. 2022, Abbas and EL-Manzalawy 2020, Pradhan et al. 2021). The support vector machine (SVM; Vapnik 1963), extreme gradient boosting (XGB; Chen and Guestrin 2016), random forest (RF; Breiman 2001), light-gradient boosting machine (LGBM; Ke et al. 2017), bagging (BAG; Breiman 1996), adaptive boosting (ADB; Freund and Schapire 1999), and gradient boosting decision trees (GBDT; Friedman 2001) were the seven machine learning techniques we used in this study. Table 2 lists the R-packages used to implement the learning models and the parameter settings for each learning model.
Feature selection approach
By eliminating duplicate and irrelevant features, feature selection reduces the computational burden while increasing classification accuracy (Pradhan et al., 2022). The support vector machine recursive feature elimination (SVM-RFE; Guyon et al., 2002), random forest variable importance measure (RF-VIM; Daz-Uriarte and Alvarez de Andrés, 2006), XGB variable importance (XGB-VIM; Sandri and Zuccolotto, 2008), and LGBM variable importance measure (LGB-VIM; Ke et al., 2017) were used to select important and relevant features. According to past studies (Guyon et al., 2002; Pradhan et al., 2022), the top features in this study that led to a classifier with the best classification accuracy was chosen. The sigFeature R-package was used to implement the SVM-RFE technique (Das et al., 2020). The R-packages randomForest (Liaw and Wiener 2002), xgboost (Chen et al., 2021b), and lightgbm (Shi et al. 2022) were used to implement the RF-VIM, XGB-VIM, and LGB-VIM methods, respectively.
Cross-validation and performance metrics
A five-fold cross-validation approach was used to assess the performance of the prediction models. Both the positive and negative datasets were randomly separated into five subgroups of equal size to perform the five-fold cross-validation (Jiang and Wang, 2017). In each fold of the cross-validation, one randomly selected subset from each class served as the test set, while the remaining four subsets from both classes were pooled to serve as the training set. With distinct training and test sets for each fold, the experiment was carried out five times, and the accuracy over the five folds was recorded. The different steps involved to develop the proposed approach are shown in Fig. 2. The following metrics were used to evaluate the performance of the prediction models: sensitivity, specificity, accuracy, precision, area under receiver operating characteristic curve (AU-ROC; Fawcett, 2006), and area under precision recall curve (AU-PRC; Boyd et al., 2013). In the following formulae, TP and FP respectively represent the number correctly and wrongly predicted positive samples, whereas TN and FN respectively represent the number correctly and wrongly predicted negative samples.
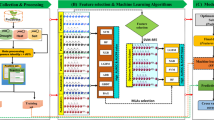
Illustration of the brief outline of the proposed computational approach. The diagram depicts the overall workflow of the entire computational strategies followed to develop the abiotic stress-responsive lncRNA prediction models. (A) Retrieval of experimentally validated abiotic responsive and non-responsive lncRNA sequences from the PncStress and PLncDB V2.0 database and processing of sequence data; (B) sequence-derived Kmer feature generation and selection of most important features and machine learning algorithm (MLA) based on AU-ROC and AU-PRC; (C) model building using machine learning technique and cross-validation with selected features and assessment of model in the independent test dataset
Results
Performance analysis of MLAs with independent Kmer feature set
The performance of each machine learning method was evaluated independently with each Kmer feature set (K1 to K6). The highest sensitivity of 69.05% was achieved with LGBM for K4, followed by the BAG (67.59%) with K2 (Fig. 3). In comparison to the other combinations of Kmer size and learning algorithm, BAG also achieved the highest specificity (72.68%) with K4. The BAG algorithm also achieved the highest precision of 68.10% for K4 (Fig. 3). As far as overall accuracy is concerned, RF achieved the highest value of 61.79% with tri-nucleotide compositional features (K3), followed by XGB (61.95%) and GBDT (61.21%) with dinucleotide (K2) and tri-nucleotide (K3) features, respectively (Fig. 3). With K3, RF also achieved the highest AU-ROC (70.70%) and AU-PRC (70.69%). In comparison to the remaining learning algorithms, XGB with K2 was found to produce higher AU-ROC (70.32%) and AU-PRC (69.51%) (Fig. 3). Because the features generated with large Kmer sizes are sparse, the accuracy obtained with K5 and K6 may be worse than with K1, K2, K3, and K4, similar to the present study.
Performance analysis of MLAs with combined Kmer feature set
In addition to evaluating the accuracy of each Kmer feature set separately, the performance of machine learning algorithms was evaluated using combined Kmer feature sets such as K12 (K1+K2), K123 (K1+K2+K3), K1234 (K1+K2+K3+K4), K12345 (K1+K2+K3+K4+K5), and K123456 (K1+K2+K3+K4+K5+K6). The highest sensitivity (79.98%) was achieved by SVM with K12 features, whereas the BAG method achieved the highest sensitivity for the rest of the feature combinations (Fig. 4). The highest specificity (66.17%) and precision (62.82%) was achieved by GBDT with K123, followed by RF (65.36%, 61.91%) with K12 features. When XGB was used, the highest accuracy was found to be 62.16% with K12 features, followed by GBDT (62.15%) with K123 and RF (62.14) with K12 features (Fig. 4). Barring a few exceptions, the accuracies were seen to be declining with an additional increase in the Kmer features (Fig. 4). The RF achieved the highest AU-ROC (69.4%) with K123, followed by XGB (69.37%) with K12 features (Fig. 4). The highest AU-ROC with K123 features was seen to be less than that obtained with RF for K3 (70.70%). When RF was employed as the classifier, K12 produced the highest AU-PRC (70.18%), which was also lower than the AU-PRC of RF achieved with K3 (70.70%) (Fig. 4).
Performance analysis MLAs with selected Kmer features
In order to improve prediction accuracy further, four different feature selection procedures (SVM-RFE, RF-VIM, XGB-VIM, and LGB-VIM) were employed to identify relevant and non-redundant features. The features were ranked in order of relevance, with the first being the most significant and the final being the least important. The prediction accuracy of learning algorithms was further evaluated in terms of AU-PRC by adding 10 top features at a time (Fig. 5). The BAG method was observed to achieve the highest AU-PRC of 65.08% using the top 70 XGB-VIM features (Table 3). Similarly, BAG achieved the highest AU-PRC of 65.66% with 590 top-selected features of LGB-VIM. SVM was found to achieve the maximum accuracy (72.66%) among the considered models with 100 top features chosen by RF-VIM (Table 3). Furthermore, SVM was observed to achieve the highest AU-PRC of 76.16% using the top 530 SVM-RFE features (Table 3). The prediction accuracy of the learning algorithms was observed to be improved when compared to the performance with all 5460 features. The SVM was found to be the best performer, followed by the RF when the prediction was done using the selected features of SVM-RFE and RF-VIM (Fig. 5). The BAG method was found to be the better achiever when it came to prediction using the chosen features of XGB-VIM and LGB-VIM in comparison to the other methods (Fig. 5).
Analysis of cross-validation and independent test set prediction
Since the SVM was found to achieve the highest accuracy with 530 top-selected features of SVM-RFE, the same combination was employed for cross-validation performance analysis. As far as cross-validation analysis is concerned, the sensitivity, specificity, overall accuracy, precision, AU-ROC, and AU-PRC were observed to be 73.03, 64.61, 68.84, 67.58, 73.98, and 75.54%, respectively (Table 4). The model trained with SVM using 530 selected features was also employed to predict the independent test set (101 positive and 101 negative sequences). For the independent test set, the sensitivity, specificity, overall accuracy, precision, AU-ROC, and AU-PRC were found to be 91.08, 61.38, 76.23 and 70.22, 87.71, and 88.49%, respectively (Table 4). The higher degree of sequence similarity with the training dataset may be attributed to the higher accuracy of the independent test set when compared to the cross-validation accuracy.
Development of an online prediction tool
In order to predict the abiotic stress-responsive lncRNAs, we further developed an online prediction tool called ASLncR (https://iasri-sg.icar.gov.in/aslncr/). The front end of the server was designed using HTML, while its back end uses PHP to execute the developed in-house R-code. This server implemented the SVM model using the 530 chosen features. For prediction, the user has to either paste or upload the lncRNA sequences in FASTA format. The results are displayed in tabular format, where the probability of each lncRNA being associated with stress is provided.
Performance analysis of ASLncR with experimentally validated dataset
To further confirm the efficiency of the developed tool ASLncR, lncRNA sequences for various abiotic stresses were manually collected from published literature (Jha et al. 2020; Urquiaga et al. 2020; Patra et al. 2023). For 9 different plant species, a total of 190 sequences were collected for the abiotic stresses cold, heat, light, salt, drought, flood, and others. We were left with 138 sequences for the evaluation using our model after eliminating the sequences that were present in the positive set of training and independent test dataset. The abiotic stress responsiveness of the sequences was predicted using the ASLncR server, and it was discovered that 81.88% (113 out of 138) of the sequences were correctly identified.
Discussion
Abiotic stresses brought about by climate change pose a serious challenge to crop production and productivity. Therefore, it is necessary to develop abiotic stress-tolerant crop cultivars to meet the food security demand. In the last decade, a considerable amount of research has focussed to understand the different regulatory roles of lncRNAs in plant response to abiotic stresses and their indispensable roles in environmental adaptation (Chen et al., 2023; Yang et al., 2022; Liu et al., 2022b; Zhang et al., 2022; Tian et al., 2023; Ye et al., 2022; Chen et al., 2022). To put it another way, lncRNAs are multifaceted regulatory components that are essential for controlling cellular stress in response to various abiotic stimuli. For instance, Eom et al. (2019) revealed that lncRNAs co-express with mRNA in tomatoes in response to drought stress. Network analysis of the interactions between lncRNA and miRNA in Brassica juncea reveals a target for regulating drought tolerance (Bhatia et al., 2020). In order to understand how plants respond to various environmental stresses, it is crucial to identify abiotic stress-responsive lncRNAs. However, due to intricate genomic architecture, wet-lab experiments for lncRNA identification are costly and time-consuming. Thus, we developed a machine learning-based computational model for predicting abiotic stress-responsive lncRNAs based on the sequence-derived features.
Though several tools are available for plant lncRNA prediction, no single tool is available for predicting abiotic stress-responsive lncRNAs. It has been shown that lncRNAs with related functions share comparable K-mer profiles (Kirk et al., 2018). Additionally, the K-mer features have been successfully utilized to establish relationships between sequence and function among lncRNAs (Kirk et al. 2018; Kirk et al. 2021). In order to capture the abundance of short motifs in an lncRNA, in the present study, the K-mer features were used to encode lncRNAs into numeric feature vectors. The Kmer features have also been successfully applied in other areas of bioinformatics such as sequence assembly (Li et al. 2010), metagenomics (Dubinkina et al. 2016), DNA barcoding (Meher et al. 2016), and lncRNA prediction (Sun et al. 2013). We considered Kmer sizes 1 to 6, where the accuracy obtained with individual Kmer features was found to be higher than the accuracy obtained by combining all 5460 Kmer features. Shorter K-mers are more common, and their relative frequencies are more strongly cross-correlated than for longer K-mers (Klapproth et al. 2021), which could be a probable reason for the low accuracy with higher K-mer features.
It was seen that while all the 5460 features were utilized, the prediction accuracy was low. Thus, in order to improve prediction accuracy, significant and non-redundant features were selected by employing feature selection methods. To choose important features, four distinct feature selection strategies, including SVM-RFE, RF-VIM, XGB-VIM, and LGB-VIM, were adopted. As compared to all the 5460 features, BAG achieved the highest accuracy with 70 and 590 features selected using XGB-VIM and LGB-VIM methods, respectively. Similarly, SVM achieved the highest accuracy with 100 and 530 features selected using RF-VIM and SVM-RFE methods, respectively. Compared to the other three approaches, SVM-RFE ranking features had greater accuracy. Furthermore, it was discovered that prediction with selected features improved the accuracy of learning algorithms. When using the 530 top-ranked features of SVM-RFE, SVM had the highest accuracy among the learning algorithms, despite being the least effective when the prediction was done with individual or combined Kmer features.
The robustness of the proposed approach was also assessed using an independent dataset. The higher accuracy with the independent dataset as compared to the cross-validation accuracy may be attributed to a higher degree of sequence similarity between the training and independent test dataset. For easy implementation of our computational approach to predict abiotic stress-responsive lncRNA, we have established an online prediction tool ASLncR. Furthermore, to check the effectiveness of ASLncR, 138 experimentally confirmed abiotic stress-related lncRNAs were revalidated. The accuracy obtained from the cross-validation, independent test set validation, and the revalidation of ASLncR supports the applicability of the proposed model for predicting abiotic stress-responsive lncRNA in a plant.
Conclusion
Intensifying evidence from various plant species signifies that lncRNAs play critical roles in abiotic stress responses. Compared to humans, the application of lncRNAs in plant breeding is still in its initial phases. Despite the fact that lncRNAs mediate plant regulation in response to abiotic stresses in many species, their potential as valuable genomic resources in plant molecular breeding or as indicators have yet to be confirmed. Studies of lncRNAs in a wider range of plant species will aid in understanding the evolution and diversity of their roles in environmental adaptation. Due to the dearth of wet-lab as well as computational approaches, potential applications of lncRNAs in plant abiotic stress are currently lacking. The present work provides one of the first computational methods, ASLncR (https://iasri-sg.icar.gov.in/aslncr/), for predicting lncRNAs that are responsive to abiotic stress. The ASLncR can be successfully employed for large-scale prediction of abiotic stress-responsive lncRNAs using only sequence information. The suggested strategy is expected to supplement the current experimental approaches for predicting abiotic stress-related lncRNAs, given the significance of lncRNAs in plant response to abiotic challenges.
References
Abbas M, El-Manzalawy Y (2020) Machine learning based refined differential gene expression analysis of pediatric sepsis. BMC Med Genet 13:122. https://doi.org/10.1186/s12920-020-00771-4
Alfaro E, Gamez M, Garcia N (2013) adabag: an R package for classification with boosting and bagging. J Stat Softw 54(2):1–35 http://www.jstatsoft.org/v54/i02/
Bhatia G, Singh A, Verma D et al (2020) Genome-wide investigation of regulatory roles of lncRNAs in response to heat and drought stress in Brassica juncea (Indian mustard). Environ Exp Bot 171:103922. https://doi.org/10.1016/j.envexpbot.2019.103922
Bhogireddy S, Mangrauthia SK, Kumar R et al (2021) Regulatory non-coding RNAs: a new frontier in regulation of plant biology. Funct Integr Genom 21:313–330. https://doi.org/10.1007/s10142-021-00787-8
Boyd K, Eng KH, Page CD (2013) Area under the precision-recall curve: point estimates and confidence intervals. In: Blockeel H, Kersting K, Nijssen S, Železný F (eds) Machine Learning and Knowledge Discovery in Databases. Springer, Berlin, Heidelberg, pp 451–466
Breiman L (1996) Bagging predictors. Mach Learn 24:123–140. https://doi.org/10.1007/BF00058655
Breiman L (2001) Random forests. Mach Learn 45:5–32. https://doi.org/10.1023/A:1010933404324
Cao Z, Zhao T, Wang L et al (2021) The lincRNA XH123 is involved in cotton cold-stress regulation. Plant Mol Biol 106:521–531. https://doi.org/10.1007/s11103-021-01169-1
Chen J, Zhong Y, Qi X (2021a) LncRNA TCONS_00021861 is functionally associated with drought tolerance in rice (Oryza sativa L.) via competing endogenous RNA regulation. BMC Plant Biol 21:410. https://doi.org/10.1186/s12870-021-03195-z
Chen L, Shi S, Jiang N et al (2018) Genome-wide analysis of long non-coding RNAs affecting roots development at an early stage in the rice response to cadmium stress. BMC Genomics 19:460. https://doi.org/10.1186/s12864-018-4807-6
Chen P, Song Y, Liu X et al (2022) LncRNA PMAT–PtoMYB46 module represses PtoMATE and PtoARF2 promoting Pb2+ uptake and plant growth in poplar. J Hazard Mater 433:128769. https://doi.org/10.1016/j.jhazmat.2022.128769
Chen T, Guestrin C (2016) XGBoost: a scalable tree boosting system. In: Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining. Association for Computing Machinery, New York, pp 785–794
Chen X, Jiang X, Niu F et al (2023) Overexpression of lncRNA77580 regulates drought and salinity stress responses in soybean. Plants 12:181. https://doi.org/10.3390/plants12010181
Choudhury S, Mansi MSK et al (2021) Genome-wide identification of Ran GTPase family genes from wheat (T. aestivum) and their expression profile during developmental stages and abiotic stress conditions. Funct Integr Genom 21:239–250. https://doi.org/10.1007/s10142-021-00773-0
Das P, Roychowdhury A, Das S et al (2020) sigFeature: novel significant feature selection method for classification of gene expression data using support vector machine and t statistic. Front Genet 11:247. https://doi.org/10.3389/fgene.2020.00247
Díaz-Uriarte R, Alvarez de Andrés S (2006) Gene selection and classification of microarray data using random forest. BMC Bioinform 7:3. https://doi.org/10.1186/1471-2105-7-3
Ding Z, Tie W, Fu L et al (2019) Strand-specific RNA-seq based identification and functional prediction of drought-responsive lncRNAs in cassava. BMC Genomics 20:214. https://doi.org/10.1186/s12864-019-5585-5
Dubinkina VB, Ischenko DS, Ulyantsev VI et al (2016) Assessment of k-mer spectrum applicability for metagenomic dissimilarity analysis. BMC Bioinform 17:38. https://doi.org/10.1186/s12859-015-0875-7
Eom SH, Lee HJ, Lee JH et al (2019) Identification and functional prediction of drought-responsive long non-coding RNA in tomato. Agronomy 9:629. https://doi.org/10.3390/agronomy9100629
Fawcett T (2006) An introduction to ROC analysis. Pattern Recognition Letters, ROC Analysis in Pattern Recognition 27:861–874. https://doi.org/10.1016/j.patrec.2005.10.010
Freund Y, Schapire RE (1999) A short introduction to boosting. Jpn Soc Artif Intell 14(5):771–780
Friedman JH (2001) Greedy function approximation: a gradient boosting machine. Ann Stat 29:1189–1232. https://doi.org/10.1214/aos/1013203451
Glisovic T, Bachorik JL, Yong J, Dreyfuss G (2008) RNA-binding proteins and post-transcriptional gene regulation. FEBS Lett 582:1977–1986. https://doi.org/10.1016/j.febslet.2008.03.004
Greenwell B, Boehmke B, Cunningham J, et al (2022). gbm: generalized boosted regression models. R package version 2.1.8.1. https://CRAN.R-project.org/package=gbm
Guo F-B, Dong C, Hua H-L et al (2017) Accurate prediction of human essential genes using only nucleotide composition and association information. Bioinformatics 33:1758–1764. https://doi.org/10.1093/bioinformatics/btx055
Guyon I, Weston J, Barnhill S, Vapnik V (2002) Gene selection for cancer classification using support vector machines. Mach Learn 46:389–422. https://doi.org/10.1023/A:1012487302797
He X, Guo S, Wang Y et al (2020) Systematic identification and analysis of heat-stress-responsive lncRNAs, circRNAs and miRNAs with associated co-expression and ceRNA networks in cucumber (Cucumis sativus L.). Physiol Plant 168:736–754. https://doi.org/10.1111/ppl.12997
Huang Y, Niu B, Gao Y et al (2010) CD-HIT suite: a web server for clustering and comparing biological sequences. Bioinformatics 26:680–682. https://doi.org/10.1093/bioinformatics/btq003
Hutchinson JN, Ensminger AW, Clemson CM et al (2007) A screen for nuclear transcripts identifies two linked noncoding RNAs associated with SC35 splicing domains. BMC Genomics 8:39. https://doi.org/10.1186/1471-2164-8-39
Jha UC, Nayyar H, Jha R et al (2020) Long non-coding RNAs: emerging players regulating plant abiotic stress response and adaptation. BMC Plant Biol 20:466. https://doi.org/10.1186/s12870-020-02595-x
Jiang G, Wang W (2017) Error estimation based on variance analysis of k-fold cross-validation. Pattern Recogn 69:94–106. https://doi.org/10.1016/j.patcog.2017.03.025
Jin J, Lu P, Xu Y et al (2021) PLncDB V2.0: a comprehensive encyclopedia of plant long noncoding RNAs. Nucleic Acids Res 49:D1489–D1495. https://doi.org/10.1093/nar/gkaa910
Ke G, Meng Q, Finley T et al (2017) LightGBM: a highly efficient gradient boosting decision tree. In: Proceedings of the 31st International Conference on Neural Information Processing Systems. Curran Associates Inc., Red Hook, pp 3149–3157
Kirk JM, Kim SO, Inoue K et al (2018) Functional classification of long non-coding RNAs by kmer content. Nat Genet 50:1474–1482. https://doi.org/10.1038/s41588-018-0207-8
Kirk JM, Sprague D, Calabrese JM (2021) Classification of long noncoding RNAs by k-mer content. Methods Mol Biol 2254:41–60. https://doi.org/10.1007/978-1-0716-1158-6_4
Klapproth C, Sen R, Stadler PF et al (2021) Common features in lncRNA annotation and classification: a survey. Non-Coding RNA 7:77. https://doi.org/10.3390/ncrna7040077
Lamin-Samu AT, Zhuo S, Ali M, Lu G (2022) Long non-coding RNA transcriptome landscape of anthers at different developmental stages in response to drought stress in tomato. Genomics 114:110383. https://doi.org/10.1016/j.ygeno.2022.110383
Lee D, Karchin R, Beer MA (2011) Discriminative prediction of mammalian enhancers from DNA sequence. Genome Res 21:2167–2180. https://doi.org/10.1101/gr.121905.111
Lee C, Kikyo N (2012) Strategies to identify long noncoding RNAs involved in gene regulation. Cell Biosci 2:37. https://doi.org/10.1186/2045-3701-2-37
Li C, Nong W, Zhao S et al (2022b) Differential microRNA expression, microRNA arm switching, and microRNA:long noncoding RNA interaction in response to salinity stress in soybean. BMC Genomics 23:65. https://doi.org/10.1186/s12864-022-08308-y
Li J-R, Liu C-C, Sun C-H, Chen Y-T (2018) Plant stress RNA-seq nexus: a stress-specific transcriptome database in plant cells. BMC Genomics 19:966. https://doi.org/10.1186/s12864-018-5367-5
Li R, Zhu H, Ruan J et al (2010) De novo assembly of human genomes with massively parallel short read sequencing. Genome Res 20:265–272. https://doi.org/10.1101/gr.097261.109
Li S, Cheng Z, Dong S et al (2022a) Global identification of full-length cassava lncRNAs unveils the role of cold-responsive intergenic lncRNA 1 in cold stress response. Plant Cell Environ 45:412–426. https://doi.org/10.1111/pce.14236
Liaw A, Wiener M (2002) Classification and regression by randomForest. R News 2(3):18–22
Liu G, Liu F, Wang Y, Liu X (2022b) A novel long noncoding RNA CIL1 enhances cold stress tolerance in Arabidopsis. Plant Sci 323:111370. https://doi.org/10.1016/j.plantsci.2022.111370
Liu P, Zhang Y, Zou C et al (2022a) Integrated analysis of long non-coding RNAs and mRNAs reveals the regulatory network of maize seedling root responding to salt stress. BMC Genomics 23:50. https://doi.org/10.1186/s12864-021-08286-7
Meher PK, Sahu TK, Rao AR (2016) Identification of species based on DNA barcode using k-mer feature vector and random forest classifier. Gene 592:316–324. https://doi.org/10.1016/j.gene.2016.07.010
Mercer TR, Dinger ME, Mattick JS (2009) Long non-coding RNAs: insights into functions. Nat Rev Genet 10:155–159. https://doi.org/10.1038/nrg2521
Meyer D, Dimitriadou E, Hornik K et al (2021) e1071: Misc functions of the department of statistics, probability theory group (formerly: E1071), TU Wien. R package version 1:7–9 https://CRAN.R-project.org/package=e1071
Öztürk Gökçe ZN, Aksoy E, Bakhsh A et al (2021) Combined drought and heat stresses trigger different sets of miRNAs in contrasting potato cultivars. Funct Integr Genom 21:489–502. https://doi.org/10.1007/s10142-021-00793-w
Patra GK, Gupta D, Rout GR, Panda SK (2023) Role of long non coding RNA in plants under abiotic and biotic stresses. Plant Physiol Biochem 194:96–110. https://doi.org/10.1016/j.plaphy.2022.10.030
Pradhan UK, Sharma NK, Kumar P et al (2021) miRbiom: machine-learning on Bayesian causal nets of RBP-miRNA interactions successfully predicts miRNA profiles. PLoS ONE 16:e0258550. https://doi.org/10.1371/journal.pone.0258550
Qin T, Zhao H, Cui P et al (2017) A nucleus-localized long non-coding RNA enhances drought and salt stress tolerance. Plant Physiol 175:1321–1336. https://doi.org/10.1104/pp.17.00574
Quan M, Chen J, Zhang D (2015) Exploring the secrets of long noncoding RNAs. Int J Mol Sci 16:5467–5496. https://doi.org/10.3390/ijms16035467
Quan M, Liu X, Xiao L et al (2021) Transcriptome analysis and association mapping reveal the genetic regulatory network response to cadmium stress in Populus tomentosa. J Exp Bot 72:576–591. https://doi.org/10.1093/jxb/eraa434
Ramírez Gonzales L, Shi L, Bergonzi SB et al (2021) Potato cycling DOF factor 1 and its lncRNA counterpart StFLORE link tuber development and drought response. Plant J 105:855–869. https://doi.org/10.1111/tpj.15093
Ren J, Jiang C, Zhang H et al (2022) LncRNA-mediated ceRNA networks provide novel potential biomarkers for peanut drought tolerance. Physiol Plant 174:e13610. https://doi.org/10.1111/ppl.13610
Rutley N, Poidevin L, Doniger T et al (2021) Characterization of novel pollen-expressed transcripts reveals their potential roles in pollen heat stress response in Arabidopsis thaliana. Plant Reprod 34:61–78. https://doi.org/10.1007/s00497-020-00400-1
Saeed F, Chaudhry UK, Raza A et al (2023) Developing future heat-resilient vegetable crops. Funct Integr Genom 23:47. https://doi.org/10.1007/s10142-023-00967-8
Sandri M, Zuccolotto P (2008) A bias correction algorithm for the gini variable importance measure in classification trees. J Comput Graph Stat 17:611–628. https://doi.org/10.1198/106186008X344522
Shi Y, Ke G, Soukhavong D et al (2022) lightgbm: light gradient boosting machine. R package version 3(3):4 https://CRAN.R-project.org/package=lightgbm
Suksamran R, Saithong T, Thammarongtham C, Kalapanulak S (2020) Genomic and transcriptomic analysis identified novel putative cassava lncRNAs involved in cold and drought stress. Genes 11:366. https://doi.org/10.3390/genes11040366
Sun L, Luo H, Bu D et al (2013) Utilizing sequence intrinsic composition to classify protein-coding and long non-coding transcripts. Nucleic Acids Res 41:e166. https://doi.org/10.1093/nar/gkt646
Tan X, Li S, Hu L, Zhang C (2020) Genome-wide analysis of long non-coding RNAs (lncRNAs) in two contrasting rapeseed (Brassica napus L.) genotypes subjected to drought stress and re-watering. BMC Plant Biol 20:81. https://doi.org/10.1186/s12870-020-2286-9
Tao X, Li M, Zhao T et al (2021) Neofunctionalization of a polyploidization-activated cotton long intergenic non-coding RNA DAN1 during drought stress regulation. Plant Physiol 186:2152–2168. https://doi.org/10.1093/plphys/kiab179
Tian R, Sun X, Liu C et al (2023) A Medicago truncatula lncRNA MtCIR1 negatively regulates response to salt stress. Planta 257:32. https://doi.org/10.1007/s00425-022-04064-1
Tilman D, Balzer C, Hill J, Befort BL (2011) Global food demand and the sustainable intensification of agriculture. Proc Natl Acad Sci 108:20260–20264. https://doi.org/10.1073/pnas.1116437108
Urquiaga MCO, Thiebaut F, Hemerly AS, Ferreira PCG (2020) From trash to luxury: the potential role of plant LncRNA in DNA methylation during abiotic stress. Front Plant Sci 11:603246. https://doi.org/10.3389/fpls.2020.603246
Vapnik V (1963) Pattern recognition using generalized portrait method. Autom Remote Control 24:774–780
Wang H-LV, Chekanova JA (2017) Long noncoding RNAs in plants. In: Rao MRS (ed) Long Non Coding RNA Biology. Springer, Singapore, pp 133–154
Wang J, Liu X, Wu H et al (2010) CREB up-regulates long non-coding RNA, HULC expression through interaction with microRNA-372 in liver cancer. Nucleic Acids Res 38:5366–5383. https://doi.org/10.1093/nar/gkq285
Wang J, Chen Q, Wu W et al (2021) Genome-wide analysis of long non-coding RNAs responsive to multiple nutrient stresses in Arabidopsis thaliana. Funct Integr Genomics 21:17–30. https://doi.org/10.1007/s10142-020-00758-5
Wani SH, Kumar V, Shriram V, Sah SK (2016) Phytohormones and their metabolic engineering for abiotic stress tolerance in crop plants. Crop J 4:162–176. https://doi.org/10.1016/j.cj.2016.01.010
Wen X, Ding Y, Tan Z et al (2020) Identification and characterization of cadmium stress-related LncRNAs from Betula platyphylla. Plant Sci 299:110601. https://doi.org/10.1016/j.plantsci.2020.110601
Wu W, Wu Y, Hu D et al (2020) PncStress: a manually curated database of experimentally validated stress-responsive non-coding RNAs in plants. Database 2020:baaa001. https://doi.org/10.1093/database/baaa001
Xu S, Dong Q, Deng M et al (2021) The vernalization-induced long non-coding RNA VAS functions with the transcription factor TaRF2b to promote TaVRN1 expression for flowering in hexaploid wheat. Mol Plant 14:1525–1538. https://doi.org/10.1016/j.molp.2021.05.026
Yang H, Cui Y, Feng Y et al (2023) Long non-coding RNAs of Plants in response to abiotic stresses and their regulating roles in promoting environmental adaption. Cells 12:729. https://doi.org/10.3390/cells12050729
Yang W-C, Katinakis P, Hendriks P et al (1993) Characterization of GmENOD40, a gene showing novel patterns of cell-specific expression during soybean nodule development. Plant J 3:573–585. https://doi.org/10.1046/j.1365-313X.1993.03040573.x
Yang X, Liu C, Niu X et al (2022) Research on lncRNA related to drought resistance of Shanlan upland rice. BMC Genomics 23:336. https://doi.org/10.1186/s12864-022-08546-0
Ye X, Wang S, Zhao X et al (2022) Role of lncRNAs in cis- and trans-regulatory responses to salt in Populus trichocarpa. Plant J 110:978–993. https://doi.org/10.1111/tpj.15714
Yu F, Tan Z, Fang T et al (2020) A comprehensive transcriptomics analysis reveals long non-coding RNA to be involved in the key metabolic pathway in response to waterlogging stress in maize. Genes 11:267. https://doi.org/10.3390/genes11030267
Yu Y, Zhang Y, Chen X, Chen Y (2019) Plant noncoding RNAs: hidden players in development and stress responses. Annu Rev Cell Dev Biol 35:407–431. https://doi.org/10.1146/annurev-cellbio-100818-125218
Zhang X, Dong J, Deng F et al (2019) The long non-coding RNA lncRNA973 is involved in cotton response to salt stress. BMC Plant Biol 19:459. https://doi.org/10.1186/s12870-019-2088-0
Zhang X, Shen J, Xu Q et al (2021) Long noncoding RNA lncRNA354 functions as a competing endogenous RNA of miR160b to regulate ARF genes in response to salt stress in upland cotton. Plant Cell Environ 44:3302–3321. https://doi.org/10.1111/pce.14133
Zhang Z, Zhong H, Nan B, Xiao B (2022) Global identification and integrated analysis of heat-responsive long non-coding RNAs in contrasting rice cultivars. Theor Appl Genet 135:833–852. https://doi.org/10.1007/s00122-021-04001-y
Zhu L, Wang X, Tian J et al (2022) Genome-wide analysis of VPE family in four Gossypium species and transcriptional expression of VPEs in the upland cotton seedlings under abiotic stresses. Funct Integr Genom 22:179–192. https://doi.org/10.1007/s10142-021-00818-4
Chen T, He T, Benesty M, et al (2021b). xgboost: extreme gradient boosting. R package version 1.5.0.2. https://CRAN.R-project.org/package=xgboost
Peters A, Hothorn T, Ripley BD, et al (2023) ipred: improved predictors. https://cran.r-project.org/package=ipred
Pradhan UK, Meher PK, Naha S et al (2022) PlDBPred: a novel computational model for discovery of DNA binding proteins in plants. Brief Bioinform:bbac483. https://doi.org/10.1093/bib/bbac483
Acknowledgments
The authors sincerely acknowledge the Director, ICAR-IASRI, New Delhi, for providing the necessary facilities to carry out the research work. The authors also acknowledge the ASHOKA supercomputing facilities available at ICAR-IASRI, New Delhi.
Funding
This work was funded by the ICAR-Indian Agricultural Statistics Research Institute, PUSA, New Delhi-110012, India.
Author information
Authors and Affiliations
Contributions
Conceptualization: PKM and UKP. Methodology: UKP, PKM, and ARR. Software: SN, PKM, and UKP; validation: UKP; formal analysis: UKP, PKM, SN, and AG; investigation: PKM, AG, and ARR; resources: SN and AG; data curation: UKP and PKM; writing—original draft preparation: UKP and PKM; writing—review and editing: PKM, UK, ARR, and AG; visualization: PKM, UKP, and SN; supervision: PKM, ARR, and AG. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors. All secondary data used in the study are available at https://iasri-sg.icar.gov.in/aslncr/dataset.php
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Pradhan, U.K., Meher, P.K., Naha, S. et al. ASLncR: a novel computational tool for prediction of abiotic stress-responsive long non-coding RNAs in plants. Funct Integr Genomics 23, 113 (2023). https://doi.org/10.1007/s10142-023-01040-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10142-023-01040-0