Abstract
Abiotic stresses have become a major challenge in recent years due to their pervasive nature and shocking impacts on plant growth, development, and quality. MicroRNAs (miRNAs) play a significant role in plant response to different abiotic stresses. Thus, identification of specific abiotic stress–responsive miRNAs holds immense importance in crop breeding programmes to develop cultivars resistant to abiotic stresses. In this study, we developed a machine learning–based computational model for prediction of miRNAs associated with four specific abiotic stresses such as cold, drought, heat and salt. The pseudo K-tuple nucleotide compositional features of Kmer size 1 to 5 were used to represent miRNAs in numeric form. Feature selection strategy was employed to select important features. With the selected feature sets, support vector machine (SVM) achieved the highest cross-validation accuracy in all four abiotic stress conditions. The highest cross-validated prediction accuracies in terms of area under precision-recall curve were found to be 90.15, 90.09, 87.71, and 89.25% for cold, drought, heat and salt respectively. Overall prediction accuracies for the independent dataset were respectively observed 84.57, 80.62, 80.38 and 82.78%, for the abiotic stresses. The SVM was also seen to outperform different deep learning models for prediction of abiotic stress–responsive miRNAs. To implement our method with ease, an online prediction server “ASmiR” has been established at https://iasri-sg.icar.gov.in/asmir/. The proposed computational model and the developed prediction tool are believed to supplement the existing effort for identification of specific abiotic stress–responsive miRNAs in plants.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Sustainable food production is necessary to meet the demands of the ever-increasing human population (Mochida and Shinozaki 2013). Conversely, crop plants are constantly exposed to adverse environmental perturbations that are predicted to result in a 70% yield loss in important agricultural crops (Boyer 1982; Vij and Tyagi 2007; Zurbriggen et al. 2010). Abiotic stresses, such as cold, drought, heat and salt, have been a major factor in limiting crop yield and productivity (Akpnar et al. 2013; Budak et al. 2015). The pervasiveness and startling effects of the abiotic stresses on plant growth, development and quality have made them a significant concern in recent years (Anwar and Kim 2020). In order to activate defence mechanisms in response to abiotic stress, plants activate a network of genetic regulation, which includes changed gene expression in a considerable number of genes via transcriptional and/or post-transcriptional regulation (Ku et al. 2015). The expression of protective genes is specifically increased in plants, while the expression of negative regulators is decreased. Several protein-coding genes that control how plants respond to abiotic stresses have been unearthed in recent years (Zhang and Wang 2015).
Recent findings suggest that plants use tiny (20–24 nt) endogenous RNAs called microRNAs (miRNAs) as key post-transcriptional gene-expression regulators to inhibit plant growth and development under abiotic stress (Zhang 2015). The mRNA cleavage, translational suppression, chromatin remodelling and/or DNA methylation are some of the ways that miRNAs control gene expression (Wang et al. 2019). Typically, miRNAs that are upregulated in response to abiotic stress downregulate their target mRNAs, while those that are suppressed cause positive regulators to accumulate and become active (Chinnusamy et al. 2007). Abiotic stress leads to inconsistent miRNA expression in plants, according to numerous researches. For instance, Winter and Diederichs (2011) and Iwakawa and Tomari (2013) found that through controlling important elements of complex gene networks, miRNAs have a role in plants’ response to abiotic stress. Numerous studies have been conducted to analyse the changes in plant miRNA expression in response to biotic and abiotic stresses (Noman and Aqeel 2017). The miRNA-167, miRNA-169, miRNA-171, miRNA-319, miRNA-393, miRNA-394 and miRNA-396 are a few examples of miRNAs that work in various abiotic stress–related activities (Wang et al. 2014; Gao et al. 2016).
The response of miRNAs to abiotic stresses is largely decided by genotype, stress, tissue and miRNA type (Zhang 2015). For instance, miR408 expression is downregulated in rice (Zhou et al. 2010), cotton (Xie et al. 2015) and peach (Eldem et al. 2012) during drought stress, while it is upregulated in Arabidopsis (Liu et al. 2008), Medicago (Trindade et al. 2009) and barley (Kantar et al. 2011). In terms of tissue-dependent response of miRNAs, Wang et al. (2013) discovered an altered expression profile of miRNAs in roots versus leaves in response to drought and salinity stresses in cotton. The miR169 was found to be induced by salinity treatment in Arabidopsis but inhibited by drought stress (Li et al. 2008), demonstrating that abiotic stresses induce the expression of miRNAs in a stress-dependent manner. Similar to miR169 in Arabidopsis, miR398 was activated by UVB light but was suppressed by salinity, cold and oxidative stress (Sukar et al. 2006; Jia et al. 2009). In Arabidopsis under salinity stress, the expression of miR397 was significantly induced, but that of miR398 was significantly inhibited, indicating that plant response to abiotic stresses is miRNA-dependent (Liu et al. 2008). The studies referred above indicate that miRNAs play a substantial role in how plants react to various abiotic stresses and may be exploited as genetic targets to design plants to be more resilient to such abiotic stresses. Due to the significant role of miRNAs, they have been populated in various databases, including PlantMirnaT (Rhee et al. 2015), miRPlant (An et al. 2014), PMRD (Zhang et al. 2010), miRNEST (Szcześniak et al. 2012) and miRBase (Kozomara et al. 2014). The most recent resource for abiotic stress–responsive miRNAs is PncStress (Wu et al. 2020), which comprises experimentally validated miRNA sequences linked to diverse abiotic and biotic stresses.
Techniques including RT-PCR, cloning, RNA-microarrays and northern blots have all been extensively employed to find abiotic stress–related miRNAs. These resource-intensive wet experiments also have weak analytical qualities including accuracy, linear range and limit of detection (Ku et al. 2015; Shriram et al. 2016). Although abiotic stress–responsive miRNAs have been identified using NGS and deep sequencing technologies (Tripathi et al. 2015), the sequencing methods are species-specific. Therefore, employing existing plant miRNA sequence data, machine learning–based computational approaches may be a better alternative for predicting abiotic stress–related miRNAs. To predict abiotic stress–related miRNAs from plant miRNA sequences, we have already developed a machine learning–based technique termed ASRmiRNA (Meher et al. 2022). The developed model predicts abiotic responsive miRNA from its sequence. But, predicting miRNAs for specific abiotic stress from plant miRNA sequences is still necessary. Given the significance of miRNAs in plant response to abiotic stresses and the lack of computational methods for predicting such abiotic stress–specific miRNAs, the objective of this study is to develop a machine learning–based computational model for predicting abiotic stress–specific (cold, drought, heat and salt) miRNAs using features derived from miRNA sequences. For the purpose of discovering miRNAs under certain abiotic stresses, the current study is believed to supplement wet-lab techniques and other sequencing approaches.
Materials and methods
Collection, processing and construction of datasets
On August 23, 2022, the PncStress database (Wu et al. 2020) was accessed in order to retrieve mature miRNA sequences that are particular to an abiotic stress. This database contains 4227 stress-responsive non-coding RNAs (miRNA, LncRNA and circRNA) from 114 plants that have been experimentally verified to 48 biotic and 91 abiotic stresses. We collected 2110 miRNA sequences for 4 different abiotic stresses, including drought (862), heat (241), salt (559) and cold (448). Additionally, we took into account 376 miRNA sequences that were used as a negative set in a prior study (Meher et al. 2022). We created two distinct datasets called dataset-I and dataset-II to evaluate the performance of machine learning algorithms for predicting miRNAs that are specific to a particular abiotic stress.
Dataset-I
Thirty percent of the collected abiotic stress sequences (128 sequences for cold, 267 for drought, 68 for heat, and 167 for salt) for each stress category were set aside in order to utilize them as a positive independent test set. The positive set of the training dataset was composed of the remaining miRNA sequences from each category of abiotic stress. To prevent homologous bias in the prediction accuracy, sequences with > 60% sequence homology to any other sequences within each stress set were eliminated using the CD-HIT algorithm (Huang et al. 2010). After removing redundant sequences, a total of 216, 350, 114 and 249 sequences were obtained for cold, drought, heat and salt stress, respectively which were used to build the positive training set. Homology reduction was also applied to the positive independent set, yielding a total of 79, 149, 36 and 90 sequences for cold, drought, heat and salt, respectively. For a given abiotic stress, the other three types of miRNA are taken into account equally (at random) to construct the negative training set. The independent negative set was also built in a similar way.
Dataset-II
The negative training set for each class of stress was constructed by using the same amount of observations from the collected 376 miRBase miRNA sequences, whereas the positive training set remained the same as that of dataset-I. The positive independent set remained the same as that in dataset-I, and the remaining negative sequences in each case (after excluding the negative training set) were utilized to form the negative independent test set. A balanced dataset was taken into consideration for each category to train the model in order to prevent prediction bias toward the class having a larger number of observations. Table 1 gives a summary of the positive, negative and independent datasets.
Generation of numeric features and feature selection
As the pseudo composition of nucleotides accounts for the long-range sequence order effect, we used pseudo K-tuple nucleotide compositional (PseKNC) (Guo et al. 2014; Chen et al.2014) features in this study to convert each miRNA sequence into a numeric feature vector. The PseKNC descriptor has been effectively used in several fields of computational biology, including the prediction of nucleosome placement (Guo et al. 2014), the prediction of miRNAs that are responsive to abiotic stress (Meher et al. 2022) and others (Chen et al. 2015; Yang et al. 2018). To generate the PseKNC features, it is necessary to first identify the tier of correlation (\(\lambda\)), weight factor (w) and Kmer size (K). Since the miRNA sequences are only about 20–24 nucleotides long, correlation up to 3 tiers was taken into consideration. In this work, the default weight factor w value of 0.2 was used. The number of features generated was 7, 19, 67, 259 and 1027, correspondingly, by utilizing 5 different Kmer sizes (K = 1, 2, 3, 4 and 5). Each miRNA sequence yielded a total of 1379 features. The generated features are sparse in nature because miRNAs are shorter in length. Furthermore, since the dataset is small, there is a chance that using a large number of features will lead to over prediction. However, feature selection approach aids in the removal of redundant and irrelevant features, reducing the computational burden and boosting classification accuracy (Aksu et al. 2010; Huang et al.2014).Thus, key features were chosen using the SVM-recursive feature elimination (SVM-RFE) method (Guyon et al. 2002). Pse-in-One software (Liu et al. 2015) was used to generate the PseKNC features, and the “sigFeature” R-package was used to implement the SVM-RFE approach (Das et al. 2020).
Prediction with machine learning algorithms
Machine learning approaches have been successfully used in different areas of bioinformatics, such as gene discoveries and genome annotation (Guo et al. 2017), protein class prediction (Pradhan et al. 2022), gene expression analysis (Abbas and EL-Manzalawy 2020), complex interaction modeling in biological systems (Pradhan et al. 2021) and others. In this study, we used seven different machine learning techniques, including the support vector machine (SVM) (Vapnik 1963), the extreme gradient boosting (XGB) (Chen and Guestrin 2016), the random forest (RF) (Breiman 2001), the light-gradient boosting machine (LGBM) (Ke et al. 2017), the bagging (BAG) (Breiman 1996), the adaptive boosting (ADB) (Freund and Schapire 1999) and gradient boosting decision tree (GBDT) (Friedman 2001). The R-software was implemented for execution of the learning algorithms. The R-packages used to execute the learning models and parameter configuration for different learning models are provided in Table 2.
Cross validation and performance metrics
A five-fold cross-validation approach was used to assess the performance of different learning models. Both the positive and negative datasets were randomly separated into five subgroups of equal size in order to perform the five-fold cross-validation (Jiang and Wang 2017). In each fold of the cross-validation, one randomly selected subset from each class was used as a test set, and the remaining four subsets from both classes were merged to serve as a training set. For each fold, distinct training and test sets were used during the classification process. The accuracy across all five test sets was averaged to provide the performance measures. To measure the effectiveness of the prediction models, the following metrics were used: accuracy, area under receiver operating characteristic curve (auROC) and area under precision recall curve (auPRC):
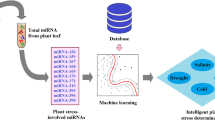
Here, TP, FP, TN and FN, respectively, represent the number of positive samples predicted to be positive, negative samples predicted to be positive, negative samples predicted to be negative and positive samples predicted to be negative. In Fig. 1, a flowchart illustrating each steps of the proposed approach is presented, and the pseudocodes for the developed algorithm are as follows:
Illustration of the brief outline of the proposed approach. The diagram depicts the overall design of the entire computational strategies followed to develop the miRNA prediction models for each abiotic stress. (A) Retrieval of experimentally validated abiotic responsive miRNA sequences from the PncStress database and processing of sequence data; (B) sequence-derived PseKNC feature generation and selection of most important features and machine learning algorithm (MLA) based on auROC and auPRC; (C) model building using machine learning techniques with selected features and assessment of cross-validation accuracy
INPUT: Cold, drought, heat and salt responsive miRNA sequences labelled as positive dataset, and equal number of non-abiotic stress responsive miRNA sequences labelled as negative dataset

Results
Analysis of discriminatory motifs
For each stress category, we conducted the discriminatory motif discovery study, which involved finding the pattern in the positive set against the negative set. Only the significant motif (p-value 0.05) was taken into consideration, and the searched length of the motif was restricted to 2 to 6 nucleotides. STREME software (Bailey 2021) was used to analyse the discriminative motifs. Figure 2 shows the discriminatory motifs that were discovered for each stress. Three cold stress motifs, including AUCMC, AUUGA and GCCGCS, were discovered to be substantially more prevalent in the positive set than the negative set. Similarly, two significant motifs were found for the drought stress, namely AAUGUU (p-value 6.810−3) and GCCGR (p-value 5.110−3). The discriminating motifs GACAGC and WGAUG were also discovered for the heat stress. While searching motifs in the positive set vs the negative set, three major conserved motifs were identified for the salt stress, including GAUUUG, AAGGAG and ASBUGC. In conclusion, different motifs were found for different stress categories, which may be important for mRNA binding.
Performance analysis of MLA with PseKNC features
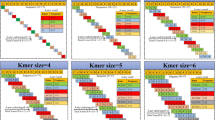
The prediction accuracy of 7 machine learning algorithms was evaluated with 5 different PseKNC feature sets using training dataset-I. With Kmer sizes 4 and 5, respectively, SVM was shown to have the highest auPRC for cold (60.4%) and drought (55.02%) (Fig. 2). When it came to heat, the PseKNC feature set with Kmer size 4 was used, and LGBM obtained the highest accuracy of 60.9% auPRC, followed by BAG (60.39%) and XGB (59.43%) (Fig. 2). With K = 5, BAG and GBDT were observed to achieve higher accuracy (56% auPRC) for salt stress; however, XGB and GBDT were seen to achieve almost similar accuracy for salt stress (55% auPRC) for Kmer size 4 (Fig. 2). The feature sets formed with Kmer sizes 4 and 5 often had higher prediction accuracies than those generated for Kmer sizes 1 to 3, which may be due to the larger size of the feature set.
Prediction analysis for training set-I using selected features
We found that using features generated with Kmer size 4 and 5 increased prediction accuracy (Fig. 3a). However, a significant portion of the features is sparse in nature due to the shorter length of the miRNA sequences (20–24nt), which may significantly create bias in the accuracy. Therefore, after integrating all of the Kmer features, the features were ranked using the SVM-RFE approach. It was discovered that different number of features was selected for each stress category to achieve the best degree of accuracy (Fig. 3b). It was also shown that when analysis was conducted using selected features, the SVM obtained the highest accuracy for all stress categories (Fig. 3b). With SVM and top 246 chosen features, the highest auPRC of 60.01% for cold stress was attained. Similarly, using 230, 310 and 240 features respectively, SVM was able to predict drought (53.54%), heat (78.34%) and salt (66.78%) stresses with the highest auPRCs (Fig. 3b). While using all of the features for prediction, the accuracy was also seen to be declining. In comparison to their highest accuracy obtained with a single PseKNC feature set, i.e. Kmer size 4 for heat and 5 for salt stress, the accuracy was shown to be enhanced by ~ 17% and ~ 10%, respectively, with the selected features (Fig. 3 a and b). Contrarily, the accuracy for cold and drought was not increased with the chosen feature sets.
Prediction with independent test set-I
The model trained with the respective training set-I was used to predict the independent dataset-I. For cold, drought, heat and salt stress, the accuracy in terms of auPRC was found to be 59.63, 66.94, 72.88 and 69.57%, respectively (Table 3). Highest prediction accuracy was observed for heat and lowest for cold, similar to cross-validation accuracy. It was also observed that, relative to their respective cross-validation accuracy, the accuracy of the independent dataset was better for drought and salt and lower for cold and heat (Table 3). For cold, drought, heat and salt, the overall accuracy was determined to be 62.02, 61.40, 77.78 and 66.67%, respectively (Table 3).
Prediction analysis with training set-II
Utilizing the chosen set of features, five-fold cross-validation prediction analysis was also carried out using SVM on the training dataset-II. For cold, drought, heat and salt, the ideal number of features to attain the maximum accuracy was 380, 272, 174 and 340, respectively (Table 4). Cross-validated prediction accuracies in terms of auPRC were found to be 90.15, 90.09, 87.71 and 89.25% with the chosen feature sets (Table 4). Prediction for the independent test set II was also done using the model learned with training set II. Overall, it was found that the prediction accuracies were 84.57, 80.62, 80.38 and 82.78%, respectively (Table 4). When compared to the training set-I and independent test set-I, respectively, the cross-validation accuracy of the training set-II and the independent test set-II were found to be significantly higher.
Analysis of selected features
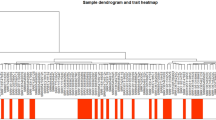
For each stress category, tSNE plots were created before and after feature selection in order to further illustrate the discriminatory feature sets. The analysis made use of the training dataset-II. The R-package Rtsne (Krijthe et al. 2017) was used to create the tSNE plot. Utilizing both the selected feature sets and all the feature sets, different tSNE plots were produced (Fig. 4a). Due to the 2-dimensional nature of the plot, it was found that the distinction between the stress and non-stress categories was not clear. When the selected features were analysed using the training dataset-II, the numbers of selected features for cold (201), drought (138) and salt (180) were greater with Kmer size 5, whereas the numbers of selected features for heat stress (62) were higher with Kmer size 4 (Fig. 4b). This might be because there were not as many observations for the heat stress, which produced more homogenous features (mainly 0s) for Kmer size 5. Additionally, it was discovered that among the selected features, 63 features were found common among the four stresses (Fig. 4c). Only 100, 51, 28 and 78 selected features were found to be independently attributed to cold, drought, heat and salt stresses respectively (Fig. 4c).
One-to-one prediction analysis
Additionally, binary classification was also done by classifying two distinct stress sets. A balanced dataset with the same number of observations from both classes was utilized to make the prediction. For the classification of cold-drought, cold-heat, cold-salt, drought-heat, drought-salt and heat-salt, the optimal number of features was 116, 80, 60, 194, 166 and 440, respectively (Fig. 5). For identifying cold-heat and drought-heat, overall cross-validation accuracy was 60.91% and 60.45%, respectively (Table 5). The classification accuracy for the remaining four combinations was found to be less than 60% (Table 5). Additionally, it was observed that with a few notable exceptions, accuracy increased up until a certain point before starting to decline (Fig. 5).
Performance analysis of deep learning models in the selected feature sets
Performance of four cutting-edge deep learning models, including one-dimensional convolutional neural networks (CNN) (Kim 2014), attention-based convolutional neural network (ABCNN) (Yin et al. 2016), long short-term memory (LSTM) (Hochreiter and Schmidhuber 1997) and Auto-encoder (AE) (Liou et al. 2014), was also compared with that of SVM. Prediction analysis was performed using the training dataset-II through five-fold cross validation, where the selected number of features (Table 4) was used for the analysis. Among the deep learning models, AE achieved higher accuracy in all four abiotic stresses (cold: 77.07%; drought: 77.57%; heat: 77.47%; salt: 79.91%) (Table 6). The ABCNN was found to be the least performer among the deep learning models (Table 6). The SVM was observed outperforming all the deep learning algorithms for predicting abiotic stress–responsive miRNA for all abiotic stresses (Table 6). Specifically, SVM achieved 5–6% higher accuracy than that of best-performing deep learning model AE.
Prediction tool ASmiR
We developed an online prediction server ASmiR (https://iasri-sg.icar.gov.in/asmir/) for prediction of abiotic stress–responsive miRNAs in cold, drought, heat and salt. The front end of the server was designed using HTML, whereas the developed R-code run at the back end with the help of PHP. The SVM model developed using the dataset-II is implemented in this server due to its high accuracy for all four abiotic stresses. The prediction can be made by using four types of abiotic stresses. The user has to paste or upload the miRNA sequences in FASTA format. The results are presented in tabular format, where the probabilities with which each miRNA sequence predicted to a specific stress category is provided.
Evaluation of ASmiR using experimentally validated dataset
For cold, drought, heat and salt stress, miRNA sequences are manually collected from available literature (Shriram et al. 2016; Begum 2022; Zhang et al. 2022) in order to further verify the effectiveness of the developed model ASmiR. Additionally, it was made sure that these sequences were not present in the positive set of the train model. We obtained 51 sequences for cold, 165 sequences for drought, 31 sequences for heat and 50 sequences for salt stress. The developed model was used to predict these sequences with respective abiotic stress, and it was found that for cold, drought, heat and salt stress respectively, 90.20, 94.54, 93.56 and 92% of the sequences were correctly predicted to their respective abiotic stress.
Discussion
Climate change, which has accelerated in recent years, is a key cause of abiotic stress, causing damage to cellular homeostasis and having a negative impact on plant growth and development (Mickelbart et al. 2015). Plant growth is impeded by abiotic stress since plants lack the ideal environmental conditions for cell division and growth. For instance, drought stress precludes plant growth because water is required for cell turgor, which promotes cell expansion (Seleiman et al. 2021); similarly, cold stress reduces plant growth because enzyme and other protein activities are limited in low temperatures (Sanghera et al. 2011).
Plants develop a variety of defence mechanisms against these abiotic stresses, among them involves using miRNAs to control the expression of abiotic stress–responsive genes. In response to various abiotic stresses, where the gene expression is controlled by translational inhibition, the miRNAs function as post-transcriptional regulators of gene expression in a sequence-specific manner (Shriram, 2016). The Argonaute proteins are recruited by the miRNA to specifically target mRNA via base-pairing in order to repress its translation and stability (Chipman and Pasquinelli 2019; Yan et al. 2018). The entire process of translational repression begins with the specific base pairing of miRNA with the target region, where the order of nucleotides in the miRNA is crucial. Targeting in particular depends on the base pairing of the miRNA’s seed region, which consists of nucleotides (nts) 2–7, to sites in the 3′UTRs of mRNA. Additionally, it has been discovered that the miRNAs’ 3′ ends play a role in controlling target specificity and regulation (Yan et al. 2018), where the degree of base pairing at the miRNA 3′ end can influence the stability of the miRNA itself (Chipman and Pasquinelli 2019). Thus, identification of abiotic stress–responsive miRNAs based on the sequence information is an important area of research as far as the plant response to different environmental stresses is concerned. In this direction, we have already developed a machine learning–based method named ASRmiRNA (Meher et al. 2022) for the first time to predict abiotic stress–related miRNA from plant miRNA sequences (Meher et al. 2022). However, this method is more generalized and cannot predict stress-specific miRNAs. Given the significance of miRNAs in plant response to specific abiotic stresses, this study focused on to develop a machine learning–based computational model for predicting abiotic stress–specific (cold, drought, heat and salt) miRNAs.
Construction of an appropriate dataset is one of the key factors determining the quality of the predictive model and is the cornerstone of machine learning algorithm learning, which directly influences the model accuracy (Sharma et al. 2021). In this study, we prepared two different datasets named as dataset-I and dataset-II for evaluation of machine learning methods for predicting abiotic stress–specific miRNAs. The accuracy was observed to be much higher for dataset-II as compared to the dataset-I. The improvement in accuracy may be due to the use of different negative sets in both datasets. As we know that same miRNA can be associated with more than one abiotic stress, the negative datasets prepared by using the observations of the rest of the stress categories may produce less accuracy. This may be the probable reason the prediction accuracy is less in case of one-to-one prediction. However, the negative sets of dataset-II were constructed from the non-abiotic stress miRNA sequences collected from miRBase (Kozomara and Griffiths-Jones 2014) which may be one of the probable reasons for higher discrimination accuracy in case of dataset-II.
Encoding of miRNAs to numeric feature vectors is essential, as machine learning algorithms can accommodate only numeric inputs (Zhang et al. 2006; Meher et al. 2018; Asefpour 2020). Sequence ordering of microRNA is important for its target recognition. It has been found that mutations in certain position may disrupt the binding of miRNAs to their original target genes (Bhattacharya and Cui 2017). Therefore, we used the pseudo K-tuple nucleotide compositional (PseKNC) features to encode miRNAs into numeric feature vectors in order to capture the sequence ordering in a miRNA. The PseKNC has also been successfully utilized in earlier studies (Guo et al. 2014; Yang et al. 2018; Meher et al. 2022) for prediction using biological sequence data.
Here, we considered Kmer size 1 to 5, and a total of 1379 numbers of features were generated. As miRNA sequences are only 20–24 nucleotides long, there is a higher probability of generated features containing large numbers of 0s, which may introduce redundancy in the feature set. In other words, because all features are derived from the PseKNC descriptor, prediction accuracy can be misleading when redundant or irrelevant features are present. Therefore, it is crucial to choose significant features from the generated features. In this study, the ideal feature set for the prediction of miRNAs specific to abiotic stress was chosen using the SVM-RFE (Wang et al. 2011). Numerous other applications, such as genomics (Tang et al. 2008), proteomics (Dao et al. 2017) and metabolomics (Lin et al. 2012), have successfully adopted the SVM-RFE method. The number of selected features was different for different stress category.
We utilised seven different machine learning methods such as SVM, RF, XGB, ADB, BAG, LGBM and GBDT for prediction of abiotic stress–responsive miRNAs. The prediction accuracies were generally found higher with the features generated with Kmer size 4 and 5, which may be due to the larger size of the feature set as compared to that of Kmer size 1 to 3. But in the selected feature sets for all abiotic stresses, SVM achieved higher accuracies over other learning algorithms in both datasets. Due to its ability to handle large and noisy data, SVM has been widely and successfully implemented in many computational studies (Brown et al. 2000; Guo et al. 2014; Chen et al. 2014). The performance of SVM was further compared with four variant of deep learning algorithms, such as CNN, ABCNN, LSTM and AE using training dataset-II with the respective selected feature sets. The SVM outperformed all four deep learning algorithms. The lower accuracies of prediction for shallow and deep learning models may be due to the features selected using SVM-RFE may not be appropriate to achieve higher accuracy with the other deep learning methods.
Conclusion
The proposed tool ASmiR (https://iasri-sg.icar.gov.in/asmir/) offers an alternative approach for predicting abiotic stress–specific (cold, drought, heat, and salt) miRNAs using features derived from miRNA sequences. Due to encouraging results, the ASmiR can be effectively used for large-scale prediction of abiotic stress–specific miRNAs by utilizing only sequence information. Given the importance of miRNAs in plant response to abiotic stresses and the lack of computational methods, it is anticipated that the proposed approach will supplement the existing experimental techniques for predicting abiotic stress–specific miRNAs.
Data Availability
All the datasets used in the study are available at https://iasri-sg.icar.gov.in/asmir/dataset.html.
References
Abbas M, EL-Manzalawy Y (2020) Machine learning based refined differential gene expression analysis of pediatric sepsis. BMC Med Genomics 13:122. https://doi.org/10.1186/s12920-020-00771-4
Akpınar BA, Lucas SJ, Budak H (2013) Genomics approaches for crop improvement against abiotic stress. Sci World J 2013:361921. https://doi.org/10.1155/2013/361921
Aksu Y, Miller DJ, Kesidis G, Yang QX (2010) Margin-maximizing feature elimination methods for linear and nonlinear kernel-based discriminant functions. IEEE Transactions on Neural Networks 21:701–717. https://doi.org/10.1109/TNN.2010.2041069
Alfaro E, Gamez M, Garcia N (2013) adabag: an R package for classification with boosting and bagging. J Stat Softw 54(2), 1-35. http://www.jstatsoft.org/v54/i02/
An J, Lai J, Sajjanhar A et al (2014) miRPlant: an integrated tool for identification of plant miRNA from RNA sequencing data. BMC Bioinformatics 15:275. https://doi.org/10.1186/1471-2105-15-275
Anwar A, Kim J-K (2020) Transgenic breeding approaches for improving abiotic stress tolerance: recent progress and future perspectives. Int J Mol Sci 21:2695. https://doi.org/10.3390/ijms21082695
AsefpourVakilian K (2020) Machine learning improves our knowledge about miRNA functions towards plant abiotic stresses. Sci Rep 10:3041. https://doi.org/10.1038/s41598-020-59981-6
Bailey TL (2021) STREME: accurate and versatile sequence motif discovery. Bioinformatics 37(18):2834–2840
Begum Y (2022) Regulatory role of microRNAs (miRNAs) in the recent development of abiotic stress tolerance of plants. Gene 821:146283. https://doi.org/10.1016/j.gene.2022.146283
Bhattacharya A, Cui Y (2017) Systematic prediction of the impacts of mutations in microRNA seed sequences. J Integr Bioinform 14:20170001, /j/jib.2017.14.issue-1/jib-2017-0001/jib-2017–0001.xml. https://doi.org/10.1515/jib-2017-0001
Boyer JS (1982) Plant productivity and environment. Science 218:443–448. https://doi.org/10.1126/science.218.4571.443
Breiman L (1996) Bagging predictors. Mach Learn 24:123–140. https://doi.org/10.1007/BF00058655
Breiman L (2001) Random forests. Machine Learning 45:5–32. https://doi.org/10.1023/A:1010933404324
Brown MP, Grundy WN, Lin D et al (2000) Knowledge-based analysis of microarray gene expression data by using support vector machines. Proc Natl Acad Sci U S A 97:262–267. https://doi.org/10.1073/pnas.97.1.262
Budak H, Kantar M, Bulut R, Akpinar BA (2015) Stress responsive miRNAs and isomiRs in cereals. Plant Sci 235:1–13. https://doi.org/10.1016/j.plantsci.2015.02.008
Chen T, Guestrin C (2016) XGBoost: a scalable tree boosting system. In: Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining. Association for Computing Machinery, New York, NY, USA, pp 785–794
Chen T, He T, Benesty M, et al (2021). xgboost: extreme gradient boosting. R package version 1.5.0.2. https://CRAN.R-project.org/package=xgboost
Chen W, Lei T-Y, Jin D-C et al (2014) PseKNC: a flexible web server for generating pseudo K-tuple nucleotide composition. Anal Biochem 456:53–60. https://doi.org/10.1016/j.ab.2014.04.001
Chen W, Lin H, Chou K-C (2015) Pseudo nucleotide composition or PseKNC: an effective formulation for analyzing genomic sequences. Mol BioSyst 11:2620–2634. https://doi.org/10.1039/C5MB00155B
Chinnusamy V, Zhu J, Zhou T, Zhu J-K (2007) Small Rnas: big role in abiotic stress tolerance of plants. In: Jenks MA, Hasegawa PM, Jain SM (eds) Advances in Molecular Breeding Toward Drought and Salt Tolerant Crops. Springer, Netherlands, Dordrecht, pp 223–260
Chipman LB, Pasquinelli AE (2019) miRNA targeting: growing beyond the seed. Trends Genet 35:215–222. https://doi.org/10.1016/j.tig.2018.12.005
Dao F-Y, Yang H, Su Z-D et al (2017) Recent advances in conotoxin classification by using machine learning methods. Molecules 22:1057. https://doi.org/10.3390/molecules22071057
Das P, Roychowdhury A, Das S et al (2020) sigFeature: novel significant feature selection method for classification of gene expression data using support vector machine and t statistic. Front Genet 11:247. https://doi.org/10.3389/fgene.2020.00247
Eldem V, Akçay UÇ, Ozhuner E, et al (2012) Genome-wide identification of miRNAs responsive to drought in peach (Prunus persica) by high-throughput deep sequencing. PLOS ONE 7:e50298. https://doi.org/10.1371/journal.pone.0050298
Freund Y, Schapire RE (1999) A short introduction to boosting. J Japan Soc Artif Intell 14(5):771–780
Friedman JH (2001) Greedy function approximation: a gradient boosting machine. Ann Stat 29:1189–1232. https://doi.org/10.1214/aos/1013203451
Gao S, Yang L, Zeng HQ et al (2016) A cotton miRNA is involved in regulation of plant response to salt stress. Sci Rep 6:19736. https://doi.org/10.1038/srep19736
Greenwell B, Boehmke B, Cunningham J, et al (2022). gbm: generalized boosted regression models. R package version 2.1.8.1. https://CRAN.R-project.org/package=gbm
Guo F-B, Dong C, Hua H-L et al (2017) Accurate prediction of human essential genes using only nucleotide composition and association information. Bioinformatics 33:1758–1764. https://doi.org/10.1093/bioinformatics/btx055
Guo S-H, Deng E-Z, Xu L-Q et al (2014) iNuc-PseKNC: a sequence-based predictor for predicting nucleosome positioning in genomes with pseudo k-tuple nucleotide composition. Bioinformatics 30:1522–1529. https://doi.org/10.1093/bioinformatics/btu083
Guyon I, Weston J, Barnhill S, Vapnik V (2002) Gene selection for cancer classification using support vector machines. Mach Learn 46:389–422. https://doi.org/10.1023/A:1012487302797
Hochreiter S, Schmidhuber J (1997) Long short-term memory. Neural Comput 9:1735–1780. https://doi.org/10.1162/neco.1997.9.8.1735
Huang M-L, Hung Y-H, Lee WM, et al (2014) SVM-RFE based feature selection and Taguchi parameters optimization for multiclass SVM classifier. ScientificWorldJournal 2014:795624. https://doi.org/10.1155/2014/795624
Huang Y, Niu B, Gao Y et al (2010) CD-HIT Suite: a web server for clustering and comparing biological sequences. Bioinformatics 26:680–682. https://doi.org/10.1093/bioinformatics/btq003
Iwakawa H, Tomari Y (2013) Molecular insights into microRNA-mediated translational repression in plants. Molecular Cell 52:591–601. https://doi.org/10.1016/j.molcel.2013.10.033
Jia X, Wang W-X, Ren L et al (2009) Differential and dynamic regulation of miR398 in response to ABA and salt stress in Populustremula and Arabidopsisthaliana. Plant Mol Biol 71:51–59. https://doi.org/10.1007/s11103-009-9508-8
Jiang G, Wang W (2017) Error estimation based on variance analysis of k-fold cross-validation. Pattern Recognition 69:94–106. https://doi.org/10.1016/j.patcog.2017.03.025
Ke G, Meng Q, Finley T, et al (2017) LightGBM: a highly efficient gradient boosting decision tree. In: Proceedings of the 31st International Conference on Neural Information Processing Systems. Curran Associates Inc., Red Hook, NY, USA, pp 3149–3157.
Kim Y (2014) Convolutional neural networks for sentence classification. In: Proceedings of the 2014 Conference on Empirical Methods in Natural Language Processing (EMNLP). Association for Computational Linguistics, Doha, Qatar, pp 1746–1751.
Kozomara A, Griffiths-Jones S (2014) miRBase: annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Res 42:D68-73. https://doi.org/10.1093/nar/gkt1181
Krijthe, J., van der Maaten, L. and Krijthe, M.J., 2017. Package ‘Rtsne’. R package version 0.13.
Ku Y-S, Wong JW-H, Mui Z et al (2015) Small RNAs in plant responses to abiotic stresses: regulatory roles and study methods. Int J Mol Sci 16:24532–24554. https://doi.org/10.3390/ijms161024532
Li W-X, Oono Y, Zhu J et al (2008) The Arabidopsis NFYA5 transcription factor is regulated transcriptionally and posttranscriptionally to promote drought resistance. Plant Cell 20:2238–2251. https://doi.org/10.1105/tpc.108.059444
Liaw A, Wiener M (2002) Classification and regression by randomForest. R News 2(3):18–22
Lin X, Yang F, Zhou L et al (2012) A support vector machine-recursive feature elimination feature selection method based on artificial contrast variables and mutual information. J Chromatogr B Analyt Technol Biomed Life Sci 910:149–155. https://doi.org/10.1016/j.jchromb.2012.05.020
Liou C-Y, Cheng W-C, Liou J-W, Liou D-R (2014) Autoencoder for words. Neurocomput 139:84–96. https://doi.org/10.1016/j.neucom.2013.09.055
Liu B, Liu F, Wang X et al (2015) Pse-in-One: a web server for generating various modes of pseudo components of DNA, RNA, and protein sequences. Nucleic Acids Res 43:W65–W71. https://doi.org/10.1093/nar/gkv458
Liu H-H, Tian X, Li Y-J et al (2008) Microarray-based analysis of stress-regulated microRNAs in Arabidopsis thaliana. RNA 14:836–843. https://doi.org/10.1261/rna.895308
Meher PK, Begam S, Sahu TK et al (2022) ASRmiRNA: abiotic stress-responsive miRNA prediction in plants by using machine learning algorithms with pseudo K-Tuple nucleotide compositional features. Int J Mol Sci 23:1612. https://doi.org/10.3390/ijms23031612
Meher PK, Sahu TK, Mohanty J et al (2018) nifPred: proteome-wide identification and categorization of nitrogen-fixation proteins of diaztrophs based on composition-transition-distribution features using support vector machine. Front Microbiol 9:1100. https://doi.org/10.3389/fmicb.2018.01100
Meyer D, Dimitriadou E, Hornik K, et al (2021) e1071: Misc Functions of the Department of Statistics, Probability Theory Group (Formerly: E1071), TU Wien. R package version 1.7-9. https://CRAN.R-project.org/package=e1071
Mickelbart MV, Hasegawa PM, Bailey-Serres J (2015) Genetic mechanisms of abiotic stress tolerance that translate to crop yield stability. Nat Rev Genet 16:237–251. https://doi.org/10.1038/nrg3901
Mochida K, Shinozaki K (2013) Unlocking Triticeae genomics to sustainably feed the future. Plant Cell Physiol 54:1931–1950. https://doi.org/10.1093/pcp/pct163
Noman A, Aqeel M (2017) miRNA-based heavy metal homeostasis and plant growth. Environ Sci Pollut Res Int 24:10068–10082. https://doi.org/10.1007/s11356-017-8593-5
Peters A, Hothorn T (2022). ipred: improved predictors. R package version 0.9-13, https://CRAN.R-project.org/package=ipred
Pradhan UK, Meher PK, Naha S, et al (2022) PlDBPred: a novel computational model for discovery of DNA binding proteins in plants. Briefings in Bioinformatics bbac483. https://doi.org/10.1093/bib/bbac483
Pradhan UK, Sharma NK, Kumar P et al (2021) miRbiom: machine-learning on Bayesian causal nets of RBP-miRNA interactions successfully predicts miRNA profiles. PLOS ONE 16:0258550. https://doi.org/10.1371/journal.pone.0258550
Rhee S, Chae H, Kim S (2015) PlantMirnaT: miRNA and mRNA integrated analysis fully utilizing characteristics of plant sequencing data. Methods 83:80–87. https://doi.org/10.1016/j.ymeth.2015.04.003
Sanghera GS, Wani SH, Hussain W, Singh NB (2011) Engineering cold stress tolerance in crop plantS. Curr Genomics 12:30–43. https://doi.org/10.2174/138920211794520178
Seleiman MF, Al-Suhaibani N, Ali N et al (2021) Drought stress impacts on plants and different approaches to alleviate its adverse effects. Plants (Basel) 10:259. https://doi.org/10.3390/plants10020259
Sharma NK, Gupta S, Kumar A et al (2021) RBPSpot: Learning on appropriate contextual information for RBP binding sites discovery. iScience 24:103381. https://doi.org/10.1016/j.isci.2021.103381
Shi Y, Ke G, Soukhavong D et al (2022) lightgbm: light gradient boosting machine. R package version 3.3.4. https://CRAN.R-project.org/package=lightgbm
Shriram V, Kumar V, Devarumath RM et al (2016) MicroRNAs as potential targets for abiotic stress tolerance in plants. Front Plant Sci 7:817. https://doi.org/10.3389/fpls.2016.00817
Sunkar R, Kapoor A, Zhu J-K (2006) Posttranscriptional induction of two Cu/Zn superoxide dismutase genes in Arabidopsis is mediated by downregulation of miR398 and important for oxidative stress tolerance. Plant Cell 18:2051–2065. https://doi.org/10.1105/tpc.106.041673
Szcześniak MW, Deorowicz S, Gapski J et al (2012) miRNEST database: an integrative approach in microRNA search and annotation. Nucleic Acids Res 40:D198-204. https://doi.org/10.1093/nar/gkr1159
Tang Y, Zhang Y-Q, Huang Z (2007) Development of two-stage SVM-RFE gene selection strategy for microarray expression data analysis. IEEE/ACM Trans Comput Biol Bioinform 4:365–381. https://doi.org/10.1109/TCBB.2007.70224
Trindade I, Capitão C, Dalmay T et al (2010) miR398 and miR408 are up-regulated in response to water deficit in Medicago truncatula. Planta 231:705–716. https://doi.org/10.1007/s00425-009-1078-0
Tripathi A, Goswami K, Sanan-Mishra N (2015) Role of bioinformatics in establishing microRNAs as modulators of abiotic stress responses: the new revolution. Front Physiol 6:286. https://doi.org/10.3389/fphys.2015.00286
Vapnik V (1963) Pattern recognition using generalized portrait method. Autom Remote Control 24:774–780
Vij S, Tyagi AK (2007) Emerging trends in the functional genomics of the abiotic stress response in crop plants. Plant Biotechnol J 5:361–380. https://doi.org/10.1111/j.1467-7652.2007.00239.x
Wang J, Shan G, Duan X, Wen B (2011) Improved SVM-RFE feature selection method for multi-SVM classifier. In: 2011 International Conference on Electrical and Control Engineering. pp 1592–1595
Wang B, Sun Y-F, Song N et al (2014) MicroRNAs involving in cold, wounding and salt stresses in Triticum aestivum L. Plant Physiol Biochem 80:90–96. https://doi.org/10.1016/j.plaphy.2014.03.020
Wang J, Mei J, Ren G (2019) Plant microRNAs: biogenesis, homeostasis, and degradation. Front Plant Sci 10:360. https://doi.org/10.3389/fpls.2019.00360
Wang M, Wang Q, Zhang B (2013) Response of miRNAs and their targets to salt and drought stresses in cotton (Gossypium hirsutum L.). Gene 530:26–32. https://doi.org/10.1016/j.gene.2013.08.009
Winter J, Diederichs S (2011) Argonaute proteins regulate microRNA stability: increased microRNA abundance by Argonaute proteins is due to microRNA stabilization. RNA Biol 8:1149–1157. https://doi.org/10.4161/rna.8.6.17665
Wu W, Wu Y, Hu D, et al (2020) PncStress: a manually curated database of experimentally validated stress-responsive non-coding RNAs in plants. Database 2020:baaa001. https://doi.org/10.1093/database/baaa001
Xie F, Wang Q, Sun R, Zhang B (2015) Deep sequencing reveals important roles of microRNAs in response to drought and salinity stress in cotton. J Exp Bot 66:789–804. https://doi.org/10.1093/jxb/eru437
Xu Q, He Q, Li S, Tian Z (2014) Molecular characterization of StNAC2 in potato and its overexpression confers drought and salt tolerance. Acta Physiol Plant 36:1841–1851. https://doi.org/10.1007/s11738-014-1558-0
Yan Y, Acevedo M, Mignacca L et al (2018) The sequence features that define efficient and specific hAGO2-dependent miRNA silencing guides. Nucleic Acids Res 46:8181–8196. https://doi.org/10.1093/nar/gky546
Yang H, Qiu W-R, Liu G et al (2018) iRSpot-Pse6NC: identifying recombination spots in Saccharomyces cerevisiae by incorporating hexamer composition into general PseKNC. International Journal of Biological Sciences 14:883–891. https://doi.org/10.7150/ijbs.24616
Yin W, Ebert S, Schütze H (2016) Attention-based convolutional neural network for machine comprehension. In: Proceedings of the Workshop on Human-Computer Question Answering. Association for Computational Linguistics, San Diego, California, pp 15–21
Zhang B (2015) MicroRNA: a new target for improving plant tolerance to abiotic stress. J Exp Bot 66:1749–1761. https://doi.org/10.1093/jxb/erv013
Zhang B, Pan X, Cannon CH et al (2006) Conservation and divergence of plant microRNA genes. Plant J 46:243–259. https://doi.org/10.1111/j.1365-313X.2006.02697.x
Zhang B, Wang Q (2015) MicroRNA-based biotechnology for plant improvement. J Cell Physiol 230:1–15. https://doi.org/10.1002/jcp.24685
Zhang F, Yang J, Zhang N et al (2022) Roles of microRNAs in abiotic stress response and characteristics regulation of plant. Front Plant Sci 13:919243. https://doi.org/10.3389/fpls.2022.919243
Zhang Z, Yu J, Li D et al (2010) PMRD: plant microRNA database. Nucleic Acids Res 38:D806-813. https://doi.org/10.1093/nar/gkp818
Zhou L, Liu Y, Liu Z et al (2010) Genome-wide identification and analysis of drought-responsive microRNAs in Oryza sativa. J Exp Bot 61:4157–4168. https://doi.org/10.1093/jxb/erq237
Zurbriggen MD, Hajirezaei M-R, Carrillo N (2010) Engineering the future. Development of transgenic plants with enhanced tolerance to adverse environments. Biotechnol Genet Eng Rev 27:33–56. https://doi.org/10.1080/02648725.2010.10648144
Acknowledgements
The authors sincerely acknowledge the Director, ICAR-IASRI, New Delhi, for providing the necessary facilities to carry out the research work. The authors also acknowledge the ASHOKA supercomputing facilities available at ICAR-IASRI, New Delhi.
Funding
This work was funded by ICAR-Indian Agricultural Statistics Research Institute, PUSA, New Delhi-110012, India.
Author information
Authors and Affiliations
Contributions
Conceptualization, PKM and UKP. Methodology, UKP, PKM, and ARR. Software, SN, PKM, SP, and UKP; validation, UK, SP and UKP; formal analysis, UKP, PKM, SN, AG, ARR, and UK.; investigation, PKM, AG, ARR, and UK.; resources, SN, UK, SP, and AG; data curation, UKP and PKM.; writing—original draft preparation, UKP and PKM.; writing—review and editing, PKM, UK, ARR, and AG; visualization, PKM, UKP, SN, SP, and UK.; supervision, PKM, UK, ARR, and AG. All the authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors. All secondary data used in the study is available at https://iasri-sg.icar.gov.in/asmir/dataset.html.
Conflict of interest
The authors have no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Pradhan, U.K., Meher, P.K., Naha, S. et al. ASmiR: a machine learning framework for prediction of abiotic stress–specific miRNAs in plants. Funct Integr Genomics 23, 92 (2023). https://doi.org/10.1007/s10142-023-01014-2
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10142-023-01014-2