Abstract
Macroalgae play a crucial role in marine ecosystems when they contribute to the global primary production in the habitats formation, providing food and shelter to a range of aquatic organisms. They have a number of interactions with bacteria and other organisms such as fouling and disease. To inhibit the settling, growing, and biofilm formation by bacteria, it has been suggested that the macroalgae influence bacterial metabolism and quorum sensing through the production of secondary metabolites with antibiotic effect. Macroalgae-bacteria interactions have been investigated for many years. These interactions can be beneficial when the bacteria assist with the normal development of macroalgae as well as reducing secondary fouling on the algal surface. On the other hand, the interactions may have a deleterious effect when the biofilm impairs the photosynthetic ability or promotes disease development. This review reports the recent advances in the understanding of bacteria-brown algae interactions, highlighting the diversity and functional role of epiphytic bacteria, including the maintenance of the health of the algae and the biological activities described from this association. Through combined bacterial culture, microscopy, and molecular biology, it has been possible to identify and establish the phylogenetic origin of different bacterial communities associated with brown algae, being predominantly the phyla Proteobacteria, Bacteroidetes, and Firmicutes. Further investigation of the bacterial communities that live on different macroalgae using new technologies are still required, mainly to evaluate the production and secretions of metabolites with biotechnological potential.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Marine macroalgae are eukaryotic, photosynthetic, sessile (usually), and multicellular organisms. They are one of the main producers at the aquatic ecosystem and they contribute to almost half of the aquatic global production (Graham and Wilcox 1999). Moreover, they serve as housing to many epibiont species and they also provide suitable substrata for microorganism attachment as well as producing many organic substances that function as nutrients for bacterial multiplication and bacterial biofilm production (Singh et al. 2013).
Bacteria associated with marine algae play an important role, directly or indirectly, in normal algal morphological development, on their growth, defense against fouling organisms, and metabolism (Goecke et al. 2010). It should be highlighted that the behavior of the macroalgae in ecological and industrial (pharmacological) environments cannot be understood without considering the interactions with the associated microbiota (Egan et al. 2013). Many studies have proposed that there is a mutualistic relationship in which the bacterial community protects the host algae against secondary biological fouling, while the host surface provides nutrients and physical protection to the associated bacteria (Penesyan et al. 2010).
Despite countless examples reporting advantages in algae-bacteria relationships, this interaction is not always beneficial, because once the bacterial communities compromise the algal tissue and algal photosynthetic capability (Hollants et al. 2013), they can induce new diseases as well as pathogens that can compromise the health of the host algae (Zozaya-Valdes et al. 2015). These observations have led to investigations of the potential of extracts and/or isolated products from different marine sources, particularly from algae, against numerous organisms, including viruses and bacteria, as possible pharmaceuticals. Furthermore, the bacteria associated with algae also represent an important potential source of new promising substances, as new bioactives and antimicrobial metabolites (Egan et al. 2008; Penesyan et al. 2009; Ismail et al. 2016).
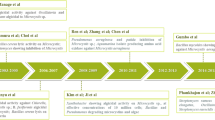
In the last decades, a great effort has been directed to the study of bacterial communities associated with algae aiming to understand the structure, succession, and dynamics of these communities in relation to ecology starting on the bacteria-algae interaction (Table 1). Most information related to algae-bacteria interactions is from studies involving green and red algae as a model for physiological and/or ecological studies (Tapia et al. 2016). These studies have demonstrated that the specific bacteria presence is necessary for morphological development and growth in green (Singh et al. 2011a; Spoerner et al. 2012; Wichard 2015; Grueneberg et al. 2016; Ghaderiardakani et al. 2017), red (Singh et al. 2011b; Fukui et al. 2014), and brown algae (Tapia et al. 2016). Bacteria are also known to induce zoospore colonization (Singh et al. 2013; Vesty et al. 2015) and spore release (Weinberger et al. 2007; Singh et al. 2015; Tapia et al. 2016). Furthermore, bacteria associated with benthic algae have ecological roles such as establishment of planktonic propagules (larvae, spores, bacteria) (Steinberg and de Nys 2002; Othmani et al. 2016; Satheesh et al. 2016) and host defense against deleterious microorganisms (Goecke et al. 2010; Singh and Reddy 2014; Campbell et al. 2015).
The identification of bacteria that inhabit macroalgae has been the object of multidisciplinary studies involving taxonomy and ecology. These have identified the phyla Proteobacteria and Firmicutes as the most abundant on macroalgal surfaces as reviewed by Hollants et al. (2013). This multidisciplinary approach is more common in research on biofilms on marine algae, where combined tools from bacteria culture, microscopy, and advanced molecular biology are used to characterize bacteria communities and explore many questions related to occurrence, distribution, persistence, and physiological and ecological functions of associated bacteria (see reviews by Steinberg et al. 2002 and Goecke et al. 2010). There is extensive literature on larvae induction and inhibition, marine algae pathogenesis, bacterial signaling molecular biology, macroalgal allelopathic chemical defenses, focusing on the general chemical structure of the colonization surface (Friedrich et al. 2001; Egan et al. 2013; Hollants et al. 2013). However, detailed knowledge of algal interaction with associated microorganisms on algae surfaces needs better understanding (Steinberg et al. 1998; Steinberg and de Nys 2002; Kubanek et al. 2003).
Brown algae (Phaeophyceae) have important ecological roles on costal ecosystems and they are one of the most diversified groups of benthic algae (Andersen 2004; Cock et al. 2011), and therefore it is of particular interest to specifically focus on the ecological roles of bacterial communities associated with these algae. Bacteria have been described in association with brown algae (Hengst et al. 2010; Lachnit et al. 2011), and some initial observations exist which connect the bacterial presence to development and growth (Pedersén 1968). To elucidate the basic aspects of brown algal biology, small filamentous species such as Ectocarpus siliculosus have been chosen as study models (Peters et al. 2004; Tapia et al. 2016). Lobophora species also have been used to investigate associated bacteria and potential induction on coral bleaching (Vieira et al. 2016).
Therefore, the aim of this review is to provide new information about (1) the diversity of bacteria associated with brown algae communities; (2) the role of biofilm on the brown algae surface; and (3) the exploration of secondary metabolite production, beginning with bacteria-brown algae interactions to discover new biological activities. To accomplish this, a literature review on the interaction of brown algae-bacteria was carried out between the years 2010 and 2018. In this search, the online databases used were Scifinder, Science direct, and Pubmed, with following keywords: “bacteria-brown algae interaction,” “biofilm and brown algae,” “biological activity and brown algae,” “algae and bacteria,” “EPS and algae,” “bacterial communities and seaweeds,” “isolation of bacteria and seaweed.” The evaluated studies were selected according to the information on the isolation and identification of bacterial communities on the surface of brown algae and on the types of interactions between brown algae and bacteria, as well as the biological activities already tested using brown seaweeds and/or bacteria. Studies addressing the isolation and identification of bacterial communities taken from water or marine sediments and work involving the transformation of heavy metals by bacteria or the association of bacteria and microalgae were excluded.
Bacterial communities associated with brown algae
Marine macroorganisms live in persistent contact with diverse microorganisms that are abundant and ubiquitous in the surrounding seawater and with biofilms on their surfaces (Wahl et al. 2012). Brown algae represent an important component of the infralittoral zone, which is present in temperate, tropical, and subtropical ecosystems (La Barre et al. 2010). Brown algae have a greater structural complexity when compared to other benthic algae. They produce chlorophyll a and c, and the carotenoids such as fucoxanthin as the most abundant photosynthetic pigments (Teixeira 2013). Macroscopic marine organisms live in persistent contact with many microorganisms that are abundant and omnipresent in the surrounding seawater (Wahl et al. 2012; Kouzuma and Watanabe 2015). Therefore, the marine algal surface provides a suitable substrate for microorganism attachment and they produce many organic substances that function as nutrients for bacterial replication and biofilm formation (Lachnit et al. 2013).
Microbial communities that live on the surface of algae are highly complex, dynamic, and consist of consortia of microorganisms, including bacteria, fungi, diatoms, protozoa, algal spores, and marine invertebrate larvae (Burke et al. 2011a; Satheesh et al. 2016). Although there is a strong pressure on the colonization by epibiont bacteria, many marine algae host microbial communities on their surfaces that differ both in quality and in quantity from the free-living bacteria in their surrounding environment (Lachnit et al. 2013).
One of the critical points involving bacterial communities is the limitation of identification techniques. Recent advances in sequencing technology have enabled researchers to characterize microbial diversity at previously unattainable scales such as the Human Microbiome Project (HMP 2012). Given the lack of a commonly accepted bacterial species concept, a phenomenological approach to categorizing microbial diversity is often chosen in practice—operational taxonomic units (OTU), defined as clusters of 16S/18S small subunit (SSU) rRNA gene similarity are used (Bondoso et al. 2013; Schmidt et al. 2014). Bondoso et al. (2013) applied denaturing gradient gel electrophoresis (DGGE) with 16S rRNA gene-specific primers for Planctomycetes to compare the communities of these organisms developing on various macroalgae. Shannon diversity indexes showed that DGGE profiles were similar in all the macroalgae. Ismail et al. (2016) studied antibacterial activities of epiphytic bacteria isolated from brown alga Padina pavonica based on 16S rRNA gene sequences. The antimicrobial activity was assessed as inhibition of growth of 12 species of pathogenic bacteria.
Epiphytic marine bacteria are intimately associated with brown algae. Between 2010 and 2018, 35 articles reported the presence of bacteria in 46 species of brown algae (Fig. 1). Among the most abundant bacterial communities are the phyla Proteobacteria, Bacteroidetes, and Firmicutes (Singh and Reddy 2014). It is suggested that the predominance of these bacteria is related to their capability to resist the effects of many stress parameters in addition to having high efficiency production system (Cray et al. 2013). Only a few studies dedicated to comprehensive assessments of total bacteria communities on algal surfaces, especially brown algae exist. However, beginning with data based on genetic sequencing, it has been revealed that the bacterial communities associated with algae are different from the planktonic bacterial communities (Burke et al. 2011b; Goecke et al. 2013). Marine macroalgae typically host diverse bacterial groups with density that varies from 102 to 107 cells cm−2, depending on the macroalgal species, thallus section, and season (Tujula et al. 2006; Bengtsson et al. 2010; Egan et al. 2013). Previous studies have reported that algae occurring in the same ecological niche have a specific bacterial community for each algae species. In contrast, macroalgae, which belong to the same species, but which occur in a different geographic location, have similar bacterial communities (Nylund et al. 2010). The specificity of bacterial communities with macroalgae may be related to three possible processes: (a) algal propagules can take specific biofilm to other areas, (b) algal chemical defenses may selectively inhibit the growth of other biofilms than that specific for host algae, and (c) algal attractants may facilitate the colonization of certain bacterial strains (Lachnit et al. 2009).
The bacterial communities associated with algae not only vary from species to species but also show temporal variation (Lachnit et al. 2011). These authors studied the epibacterial community associated with the benthic alga Fucus vesiculosus at different sampling times. They observed that among the algal bacterial community, 7–16% of sequences belonged to specific species on the host alga. For example, for F. vesiculosus, the closely related strains (Octadecabacter arcticus-Alphaproteobacteria; Granulosicoccus antarcticus-Gammaproteobacteria; Bacteroidetes-Bacteroidetes; Roseibacillus spp.-Verrucomicrobia; Planctomyces sp.-Planctomycetes) represented 16.21% of similarity between specific bacteria. In addition, other factors such as season and life cycle of the host algae can affect the associated bacterial community composition (Singh et al. 2013). Staufenberger et al. (2008) studied bacteria communities associated with the rhizoids, cauloid, meristem, and phylloid of the brown alga Laminaria saccharina (= Saccharina latíssima). They observed that the association obtained from cauloid and meristem were more specific, while the less specific associations were obtained from the more aged phylloid. Seasonal and geographic differences in the associated communities were also observed.
Clearly, there are many explanations for the host algae specificity and the temporal variations of the bacterial community associated with these algae (Singh and Reddy 2014). Epibacterial communities are sheltered in different ways (temporal and spatial on thallus distribution) on the algal surface due to biochemical and metabolite composition (Cray et al. 2013). For example, the fucoidan-degrading activity of Verrucomicrobia, members of Flavobacteriaceae and Gammaproteobacteria, suggest selective colonization on Fucus (Barbeyron et al. 2008). However, Lachnit et al. (2013) observed that F. vesiculosus carried on its surface a specific bacterial community that belongs to the phylum Proteobacteria and Bacteroidetes.
Studies focused on brown algae indicate that these bacterial communities also can act, directly or indirectly, on the morphology and reproduction (e.g., Ectocarpus sp.; Tapia et al. 2016) and on the settlement inhibition of marine biofilm bacteria and barnacle larvae (Othmani et al. 2016).
Bacterial biofilms
Biofilms are complex associations of microorganisms, immobilized on surfaces and incorporated in an extracellular biological matrix which consists of extracellular polymeric substances (EPS) secreted by cells (Silva et al. 2011). As highly complex communities in natural environments, they are characterized by the interaction with a complex of biotic communities, by genetic diversity, structural heterogeneity, and the EPS itself (Joint et al. 2007; Grossart 2010). They can grow in a high variety of surfaces, including live tissues, medical devices, industrial or potable water system pipes, and natural aquatic systems (Donlan 2002). In marine ecosystems, two bacterial populations usually exist: the planktonic, which exist freely in the water column and the sessile, as a unity bonded to a surface or at the limits of a biofilm (Egan et al. 2013).
Biofilm growth is governed by a series of biological, physical, and chemical processes, being denominated by the adherence of the binding between a cell and a substrate and cohesion, the binding between cells (Fig. 2). These mechanisms are behind the fixation forms that determine the adhesive and cohesion properties that a biofilm will exhibit (Garrett et al. 2008).
Scheme with the demonstration of the steps of bacterial biofilm formation on a host algae (Adapted from Kostakioti et al. 2013)
Among the stages of biofilm formation, at first a planktonic bacterium will interact with organic and inorganic compounds on a surface and form an initial and temporary structure. This first attachment is deemed reversible. However, with time, the attachment becomes more strongly connected to a surface and subsequently, irreversible (Kostakioti et al. 2013). Secondly, the bacteria which first colonized the substrate accumulate in the biofilm through growth and reproduction, thus changing the surface composition creating a suitable environment for colonization by other bacteria. Thirdly, planktonic bacteria and those that are bonded to each other communicate by quorum sensing (QS). This communication mechanism plays a vital role on gene expression synchronization inside the bacterial community (Garrett et al. 2008; Kostakioti et al. 2013). Therefore, bacterial biofilm forming communities provide a favorable substrate for the attachment of different microorganisms; the organic and inorganic contents of EPS provide nutrients to phytoplankton and macroalgae for their survival (Mandal et al. 2011; Singh et al. 2011c).
EPS is a matrix composed of polymeric substances, especially exopolysaccharides (40–95%) and proteins (60%), as well as nucleic acids (10%) and lipids (40%) (Flemming and Wingender 2010). These substances protect the bacterial cells from the external environment and facilitate their communication through chemical and physical signals, allowing their persistence in a favorable environment (Dang and Lovell 2000; Flemming and Wingender 2001). EPSs are also being applied as bioremediation agents in environmental management systems (Kavita et al. 2011) and in ecological studies, for example, the settlement of zoospores of algae and invertebrate larvae (Hadfield 2011; Singh et al. 2013; Othmani et al. 2016). Bacterial EPS also has the capability to emulsify organic pollutants and provide healthy environments to support algal survival (Singh et al. 2013).
However, the parameters that define the macroalgal surface environment include metabolites from the alga itself, the existing resident microbial community and secondary metabolites of microbiological origin, as well as physical-chemical conditions at the thallus surface, such as oxygen and carbon dioxide, which can modify the surface pH (Egan et al. 2013). Many of these parameters are subjected to daily (Spilling et al. 2010; Dittami et al. 2016) and/or seasonal variations (Hellio et al. 2004; Bengtsson et al. 2010). In addition, macroalgal surfaces provide a habitat rich in organic material, such as reserve substances (alginate, laminarin, mannitol, and fucoidan) that are present in brown algae (Kita et al. 2016). The algae release great amounts of organic carbon at the surrounding environment, providing nutrients to the microorganisms and unleashing bacteria chemotactic behavior (Goecke et al. 2010).
While the macroalgae represent a niche with unique and selective properties, they also experiment with a range of benefic and harmful interactions with their symbiotic bacterial community (Egan et al. 2013). Considering the ecological importance, as well as macroalgal applications, there is a growing interest in these algae-bacteria interactions. However, few studies are devoted to investigating these associations with brown algae.
Mutualism relations between brown macroalgae and bacteria
Nutrition and growing factors
Epiphytic heterotrophic bacteria mineralize organic substrates, providing carbon dioxide, minerals, and, in some cases, fixed nitrogen and plant growth regulators to the macroalgae. The algae, in turn, produce organic substances and oxygen that are used by the epiphytic bacteria (Singh et al. 2011b; Singh and Reddy 2014).
Besides nutritional benefits, it has been demonstrated that the presence of certain bacteria is needed for the normal morphological development and growth of certain brown algae (Tapia et al. 2016) such as E. siliculosus (Le Bail et al. 2010). The endogenous capacity to produce such phytohormones that determined morphogenesis in these algae has a relation with the bacterial auxin (Provasoli and Carlucci 1974), similar work with other macroalgae (Prasad et al. 2010; Spoerner et al. 2012). Similarly, catalase production by the epiphytic Pseudoalteromonas porphyrae may regulate cell growth in Saccharina japonica (Dimitrieva et al. 2006).
Life cycle and morphologic development of macroalgae
Bacteria have a positive impact on the morphological development of several macroalgae species and their life cycle (Marshall et al. 2006; Egan et al. 2013; Twigg et al. 2013). Recently, Tapia et al. (2016) isolated nine strains of epiphyte bacteria of the brown alga Ectocarpus sp. culture and evaluated its effect on the morphology, reproduction, and secreted metabolites on axenic conditions and on co-culture with bacteria. Among isolated bacteria, six strains were capable of restoring the typical branched morphology and the reproduction characteristics of Ectocarpus sp. The bacteria belonged to phylum Proteobacteria and affected significantly the metabolites released by the brown algae. Goecke et al. (2012) observed the presence of bacteria on the surface of the oogonia released from the brown algae Fucus vesiculosus. Due to the smaller size of the oogonia, bacterial degradation of unviable spores cannot be excluded. Thus, bacterial biofilms can play a role in spore release, germination, and subsequent colonization of substrates by algae. It was also observed that the bacterial biofilm plays an important role on spore germination and consequently on the colonization of new substrates by macroalgae, reporting a positive correlation between the zoospore settlement and the increase in density of the biofilm (Singh et al. 2015; Vesty et al. 2015). This fact suggests the importance of the biofilm on the recruiting of macroalgal communities in coastal environments. The impact of microorganisms on the life cycle and colonization process may be important in regulating algal populations that should be investigated. However, the effect of bacteria over algal gametes and spores remains extremely neglected. Whether or not those bacteria play a role in algal ecology is completely unknown with regard to brown algae.
Macroalgae and fouling defense
Many sessile eukaryotes are intimately associated with bacteria that enable them to expand their physiological capabilities. Associations between algae and bacteria have been described for over 100 years, and these interactions can be positive, neutral, or negative (Ainsworth et al. 2010; Hollants et al. 2013). There are many laboratory studies which demonstrate that epiphyte bacteria have inhibitor activity against fouling organisms (Rao et al., 2007; Egan et al. 2008). Recently, anti-fouling and antimicrobial properties were observed on isolated bacterial strains from brown algae species (Al-Saif et al. 2014; Susilowati et al. 2015; Othmani et al. 2016).
It is worth noting that anti-fouling and antibacterial activities are found in a range of bacterial taxonomic groups. For example, the brown alga Saccharina latissima hosts more than 100 different bacteria strains that cover the phyla Proteobacteria, Bacteroidetes, Firmicutes, and Actinobacteria (Wiese et al. 2009). In addition, Murugan et al. (2012) studied 15 isolated bacteria from the Dictyota dichotoma and Chaetomorpha linoides algae biofilm, and found that eight belonged to the genera Pseudomonas, Bacillus, Corynebacterium, Micrococcus, Vibrio, Alteromonas, Flavobacterium, and Aeromonas. Furthermore, antibacterial and anti-fouling activities demonstrated by these algae can be attributed not only to the chemical defenses inherent in them but also with contribution through symbiosis or mutualistic association by epiphytic bacterial communities (Armstrong et al. 2001). This indicates that this association could both inhibit and induce the settlement of many organisms such as invertebrate larvae (Steinberg and de Nys 2002; Dworjanyn and Pirozzi 2008; Soares et al. 2008).
Bacteria-macroalgae harmful relationships
In consideration of the relationships between macroalgae and bacteria, there are advantages and disadvantages that have been investigated for over 20 years (Hollants et al. 2013). The microorganisms increasingly are known for their role as disease etiological agents in animals, plants, and marine algae (Egan et al. 2013). This interest in microbial disease in marine ecosystems is partially boosted by concerns related to climate change that generates stress on marine habitat formers and their associated microbiota which may make them more susceptible to potential opportunistic pathogens (Gachon et al. 2010).
Although there are some beneficial aspects of the associations between macroalgae and bacteria, the formation of biofilms can be a permanent threat to macroalgae. That is because biofilms may cause an increase in the host hydrodynamic resistance, reduce buoyancy and tissue elasticity, as well as attract herbivores and thus increase tissue loss or even result in its destruction (Vairappan et al. 2008, 2010). In addition, bacteria compete for nutrients (Goecke et al. 2010). The biofilm can also inhibit the gas exchange as well as reduce the availability of light, which can reduce algal photosynthetic activity. It may also increase the attachment and growth of a variety of other fouling organisms, like diatoms, invertebrate larvae, and other epiphyte algae spores. In addition, the host macroalga can be directly damaged by the bacterial community due to toxins, digestive enzymes, inhibitors, or waste products production, resulting in algal diseases (Huggett et al. 2006; Gachon et al. 2010).
In brown algae, the enrichment of some bacteria responsible for thallus rotting disease has been observed (Gachon et al. 2010). For example, Wang et al. (2008) cultivated a large number of bacteria from the thallus of S. japonica, which exhibited symptoms of the hole-rotten disease and found abundant Pseudoalteromonas and Vibrio bacteria. Studies of Laminaria religiosa health also revealed that Alteromonas bacteria are pathogenic and that they, allied with abiotic factors, induce severe damage and bleaching to the alga (Vairappan et al. 2001).
It is very likely that some of the bacteria found in the tissue of sick macroalgae are secondary colonizers that act as potential saprophytes or decomposers (Egan et al. 2013). Therefore, certain epiphytic bacteria might be commensal; however, but under stressful conditions or macroalgae infection, they become mainly saprophitic (Zozaya-Valdes et al. 2015).
Chemical interactions between bacteria and brown algae: potential new bioactives for new drugs
The marine environment represents a still unexploited resource for the isolation of novel bacterial and/or marine algal natural products such as antimicrobials (Uzair et al. 2018; Penesyan et al. 2010, 2011; Menaa 2015). The natural products chemistry of brown algae has been widely studied and they produce many types of volatile hydrocarbons, sterols, carotenoids, polyphenols, and unique terpenes (Cavalcanti et al. 2006; Moon et al. 2011; Mesquita et al. 2015). However, marine microorganisms are also seen as good candidates for the production of new compounds as potential drugs (Penesyan et al. 2010) and it is worth noting that practically all multicellular organisms that have been collected and extracted for chemical studies include associated microorganisms and this presents questions about the real biosynthetic origin of the isolated molecules (Goecke et al. 2010).
The bacterial communities associated with algae have specific characteristics when compared to bacterial strains isolated from other marine samples and are an extremely diverse potential source of bioactive compounds (Penesyan et al. 2010, 2011; Tujula et al. 2010; Burke et al. 2011; Penesyan et al. 2015). The bacterial communities associated with algae belong to several genera such as Pseudomonas, Pseudoalteromonas, Stenotrophomonas, Vibrio, Alteromonas, Shewanella, Streptomyces, and Bacillus, and have evolved in a highly competitive environment with nutrient and host space limitations, producing allelochemicals capable of preventing secondary colonization (Egan et al. 2008; Wiese et al. 2009). Marine bioactive compounds can provide wide protection (i.e., antibacterial, antifungal, antiviral, antiparasitic, antitumor, antioxidant) to the host (i.e., marine brown algae) against other microorganisms (Horta et al. 2014; Busetti et al. 2015; Uzair et al. 2018). As algae have no immune system and are continuously exposed to a range of biotic factors, they rely on secondary chemical defenses against fouling and potentially pathogenic microorganisms (see reviews by Steinberg and de Nys 2002; Goecke et al. 2010).
From 2010 until 2018, 35 studies have reported 61 species of brown algae showing bioactivities, with antibacterial activity the main activity (Fig. 3). The emphasis on antibacterial activity is because of the increase in multidrug-resistant bacteria driving the search for new substances to combat these pathogens (Meena et al. 2015). A larger number of studies have tested only crude extracts which contain a broad spectrum of natural products, hindering the identification of the particular chemical class or compound which is responsible for each activity. Few studies have isolated and identified such substances. For example, Uzair et al. (2018) isolated a new natural antibiotic (4-[(Z)-2 phenyl ethenyl] benzoic acid; Kocumarin), extracted from a new bacterium (Kocuria marina) which is associated with the brown alga Pelvetia canaliculata. In in vitro screens, Kocumarin inhibited all pathogenic fungi and bacteria tested and represents a potential new natural antibiotic for in vivo and environmental applications. Polyketides with antibacterial properties have been isolated from Bacillus subtilis associated with the brown algae Sargassum myriocystum and Anthophycus longifolius (Chakraborty et al. 2017, 2018). These studies have also suggested an ecological and metabolic role for these compounds in algal and bacteria associations. For example, Kubanek et al. (2003) have proposed that terpenoids such as lobophoride isolated from Lobophora variegata have a role in the alga’s defense against pathogenic and saprophytic fungi.
In addition to antibacterial activities, the ecological role of secondary metabolites on bacterial surface colonization also has been investigated demonstrating that the extracts and/or the isolated products acting on bacterial biofilm formation are very specific (Lachnit et al. 2010, 2013).
Marine macroalgae communicate via the surrounding environment and defend themselves by the production of metabolites. The communication between bacteria is through quorum sensing (QS), which is a chemically mediated language system that allows bacterial behavior coordination in relation to the environment (Joint et al. 2007). This system functions by the regulation of genes in response to population density. It also takes part in many physiological process such as cell differentiation, nutrient influx, bioluminescence, induction of virulence factors on pathogens of plants and animals, antibiotic biosynthesis, and biofilm formation (Bai and Rai 2011; Hollants et al. 2013).
Gram-negative bacteria, such as Pseudomonas and Vibrio, produce N-acyl homoserine lactones as signaling substances that pass through the cell membrane and bind to regulatory proteins inside the cell (Kalia 2013). Pseudomonas spp. are also known for the production of diketopiperazines that acts as QS signals (Dickschat 2010). Signaling molecules, such as γ-butyrolactones and oligopeptides, are synthetized by Gram-positive bacteria, such as Streptomyces and Bacillus (Dobretsov et al. 2009). Kanagasabhapathy et al. (2009) have suggested that certain epiphytic bacteria of the brown alga Colpomenia sinuosa play a role on the defense mechanism and suppress the settlement of other competitive bacteria, through the production of quorum sensing inhibitors (QSI) or substances similar to QSI. Macroalgae also can control bacterial colonization by interfering with the bacterial QS system as well as by the production of reactive oxygen species, similar to what happens in terrestrial plants (Potin 2008; Dittami et al. 2011). In the last decade, it has been shown that many macroalgae are capable of stimulating, inhibiting, or activating QS by bacteria through the production of QSI or analogous molecules (Kalia and Purohit 2011; Jha et al. 2013; Carvalho et al. 2017).
Another important point is that the QSI and antimicrobial substances produced by many epiphytic bacteria work together with the secondary metabolites from marine macroalgae to protect the host surface from pathogens, fouling organisms, and herbivores (Wiese et al. 2009). Many new compounds with antibiotic activities have been identified through brown algae-bacteria interaction (Horta et al. 2014; Martin et al. 2014). With the growing need to find new drugs, the understanding of marine epiphyte associations described should provide a rich source of new biomolecules of high value with the potential economic and sustainable human benefits (Murray et al. 2013; Martin et al. 2014). Therefore, bacteria associated with algae represent an important potential source of new substances (JanakiDevi et al. 2013) and are potentially easier to use in biotechnological applications when compared to a marine algal derivative (Manilal et al. 2010).
Conclusions and perspectives
Marine benthic environments are diverse and characterized by the constant competition of organisms for light, space, and nutrients. In these habitats, many macroalgae offer a substrate rich in organic material and also a safe habitat for bacterial (and other microorganisms) colonization and reproduction. The association between brown algae and bacteria can be mutualistic, a condition where the bacterial community protects the host against biological colonization, while the host surface can provide nutrients and physical protection to the bacteria. On the other hand, other types of associations can be disadvantageous to the algae, and could involve diseases, loss of the photosynthetic capability, and the costs associated with epiphytic growth.
Many studies have shown that biofilms play an important role in the development of macroalgal communities and that the bacterial communities communicate by QS. Therefore, the capability of exploring the bacterial sensorial system contributes to the understanding of the ecological success of algae. In addition, many macroalgae are capable of stimulating or inhibiting quorum sensing in bacteria. To tolerate the fouling organisms, algae have developed defense strategies that result in a great diversity of chemical substances, making them promising organisms as a source of bioactive biochemical actives, produced by either the algae or associated microorganisms. Although there is a growing interest in the secondary metabolites of the microorganisms associated with algae as a source of new natural and antimicrobial bioactive substances, little is known about the role these metabolites play in the mediation of such biological interactions.
In the last decades, the combined utilization of new microbiological, microscopy, and molecular biology techniques have helped significantly to identify and establish the phylogenetic affiliation of the algae-associated bacterial community. However, many questions about associated bacterial occurrence, distribution, persistence, and ecological roles remain unsolved, especially in studies concerning brown algae.
Many compounds with antibacterial activity have been identified from bacteria associated with brown algae. There is an urgency in finding new antibacterial substances from different natural sources, notably from the marine environment, which has organisms capable of synthesizing many unique chemical structures providing a new mechanism of action against new or reemerging infectious diseases. Therefore, future studies should observe not only the effect of specific pathogens but also the potential probiotic pathogenic effect of algae-bacteria interactions using the advances in new technologies.
References
Abdel-Raouf N, Mohamed HM, Mostafa SS, Ibraheem IBM (2017) Controlling of microbial growth by using Cystoseira barbata extract. Egypt J Bot 57:469–477
Ainsworth TD, Thurber RV, Gates RD (2010) The future of coral reefs: a microbial perspective. Trends Ecol Evol 25:233–240
Aires T, Serrão EA, Kendrick G, Duarte CM, Arnaud-Haond S (2013) Invasion is a community affair: clandestine followers in the bacterial community associated to green algae, Caulerpara racemosa, track the invasion source. PLoS One 8(7): e68429
Akbari V, Zafari S, Yegdaneh A (2018) Anti-tuberculosis and cytotoxic evaluation of the seaweed Sargassum boveanum. Res Pharm Sci 13:30–37
Akremi N, Cappoen D, Anthonissen R, Verschaeve L, Bouraoui A (2017) Phytochemical and in vitro antimicrobial and genotoxic activity in the brown algae Dictyopteris membranacea. S Afr J Bot 108:308–314
Albakosh MA, Naidoo RK, Kirby B, Bauer R (2016) Identification of epiphytic bacterial communities associated with the brown alga Splachnidium rugosum. J Appl Phycol 28:1891–1901
Ali AIB, Bour ME, Ktari L, Bolhuis H, Ahmed M, Boudabbous A, Stal LJ (2012) Jania rubens-associated bacteria: molecular identification and antimicrobial activity. J Appl Phycol 24:525–534
Ali SS, Shaaban MT, Abomohra AEF, El-Safity K (2016) Macroalgal activity against multiple drug resistant Aeromonas hydrophila: a novel treatment study towards enhancement of fish growth performance. Microb Pathog 101:89–95
Al-Saif SSA, Raouf NA, El-Wazanani HA, Aref IA (2014) Antibacterial substances from marine algae isolated from Jeddah coast of Red Sea, Saudi Arabia. Saudi J Biol Sci 21:57–64
Alvarado P, Huang Y, Wang J, Garrido I, Leiva S (2018) Phylogeny and bioactivity of epiphytic gram-positive bacteria isolated from three co-occurring Antarctic macroalgae. Antonie Van Leeuwenhoek. https://doi.org/10.1007/s10482-018-1044-6
Alves RCC, Mercês PFF, Souza IRA, Almeida CMA, Silva APS, Lima VLM, Correia MTS, Silva MV, Silva AG (2016) Antimicrobial activity of seaweeds of Pernambuco, northeastern coast of Brazil. Afr J Microbiol Res 10:312–318
Andersen RA (2004) Biology and systematics of heterokont and haptophyte algae. Am J Bot 91:1508–1522
Armstrong E, Yan L, Boyd KG, Wright PC, Burgess JG (2001) The symbiotic role of marine microbes on living surfaces. Hydrobiologia 46:37–40
Bai AJ, Rai VR (2011) Bacterial quorum sensing and food industry. Compr Rev Food Sci Food Saf 10:183–193
Balakirev ES, Krupnova TN, Ayala FJ (2012) Symbiotic associations in the phenotypically-diverse brown alga Saccharina japonica. PLoS One 7(6):e39587
Barbeyron T, L’Haridon S, Michel G, Czjzek M (2008) Mariniflexile fucanivorans sp. nov., a marine member of the Flavobacteriaceae that degrades sulphated fucans from brown algae. Int J Syst Evol Microbiol 58:2107–2113
Barott KL, Rodriguez-Brito B, Janouskovec J, Marhaver KL, Smith JE, Keeling P, Rohwer FL (2011) Microbial diversity associated with four functional groups of benthic reef algae and the reef-building coral Montastraea annularis. Environ Microbiol 13:1192–1204
Batista D, Carvalho AP, Costa R, Coutinho R, Dobretsov S (2014) Extracts of macroalgae from the Brazilian coast inhibit bacterial quorum sensing. Bot Mar 57:441–447
Bengtsson MM, Sjøtun K, Øvreås L (2010) Seasonal dynamics of bacterial biofilms on kelp (Laminaria hyperborea). Aquat Microb Ecol 60:71–83
Bengtsson MM, Sjøtun K, Lanzén A, Øvreås L (2012) Bacterial diversity in relation to secondary production and succession on surfaces of the kelp Laminaria hyperborea. ISME J 6:2188–2198
Bogolitsyn KG, Kaplitsin PA, Dobrodeeva LK, Druzhinina AS, Ovchinnikov DV, Parshina AE, Shulgina EV (2017) Fatty acid composition and biological activity of supercritical extracts from arctic brown algae Fucus vesiculosus. Russ J Phys Chem B 11:1144–1152
Bondoso J, Balagué V, Gasol JM, Lage OM (2013) Community composition of the Planctomycetes associated with different macroalgae. FEMS Microbiol Ecol 88:445–456
Bondoso J, Godoy-Vitorino F, Balagué V, Gasol JM, Harder J, Lage OM (2017) Epiphytic Planctomycetes communities associated with three main groups of macroalgae. FEMS Microbiol Ecol 93(3):fiw255. https://doi.org/10.1093/femsec/fiw255
Burke C, Thomas T, Lewis M, Steinberg P, Kjelleberg S (2011a) Composition, uniqueness and variability of the epiphytic bacterial community of the green alga Ulva australis. ISME J 5:590–600
Burke C, Steinberg P, Rusch D, Kjelleberg S, Thomas T (2011b) Bacterial community assembly based on functional genes rather than species. Proc Natl Acad Sci U S A 108:14288–14293
Busetti A, Shaw G, Megaw J, Gorman SP, Maggs CA, Gilmore BF (2015) Marine derived quorum sensing inhibitory activities enhance the antibacterial efficacy of tobramycin against Pseudomonas aeruginosa. Mar Drugs 13:1–28
Campbell AH, Marzinelli EM, Gelber J, Steinberg PD (2015) Spatial variability of microbial assemblages associated with a dominant habitat-forming seaweed. Front Microbiol 6:230
Carvalho AP, Batista D, Dobretsov S, Coutinho R (2017) Extracts of seaweeds as potential inhibitors of quorum sensing and bacterial growth. J Appl Phycol 29:789–797
Cavalcanti DN, Rezende CM, Pinto AC, Teixeira VL (2006) Diterpenoid constituents from the brown alga Dictyota menstrualis (Dictyotaceae, Phaeophyta). Nat Prod Comm 1:609–611
Chakraborty K, Thilakan B, Chakraborty RD, Raola VK, Joy M (2017) O-heterocyclic derivatives with antibacterial properties from marine bacterium Bacillus subtilis associated with seaweed, Sargassum myriocystum. Appl Microbiol Biotechnol 101:569–583
Chakraborty K, Thilakan B, Kizhakkekalam VK (2018) Antibacterial aryl-crowned polyketide from Bacillus subtilis associated with seaweed Anthophycus longifolius. J Appl Microbiol 124:108–125
Cock JM, Peters AF, Coelho SM (2011) Brown algae. Curr Biol 21:R573–R575
Coelho-Souza SA, Jenkins SR, Casarin A, Baeta-Neves MH, Salgado LT, Guimaraes JRD, Coutinho R (2017) The effect of light on bacterial activity in a seaweed holobiont. Microb Ecol 74:868–876
Coste O, Malta EJ, López JC, Fernández-Díaz C (2015) Production of sulfated oligosaccharides from the seaweed Ulva sp. using a new ulvan-degrading enzymatic bacterial crude extract. Algal Res 10:224–231
Cox S, Abu-Ghannam N, Gupta S (2010) An assessment of the antioxidant and antimicrobial activity of six species of edible Irish seaweeds. Int Food Res J 17:205–220
Cray JA, Bell ANW, Bhaganna P, Mswaka AY, Timson DJ, Hallsworth JE (2013) The biology of habitat dominance; can microbes behave as weeds? Microb Biotechnol 6:453–492
Dang H, Lovell CR (2000) Bacterial primary colonization and early succession on surfaces in marine waters as determined by amplified rRNA gene restriction analysis and sequence analysis of 16S rRNA genes. Appl Environ Microbiol 66:467–475
De Corato U, Salimbeni R, De Pretis A, Avella N, Patruno G (2017) Antifungal activity of crude extracts from brown and red seaweeds by a supercritical carbon dioxide technique against fruit postharvest fungal diseases. Postharvest Biol Technol 131:16–30
Del Olmo A, Picon A, Nuñez M (2018) The microbiota of eight species of dehydrated edible seaweeds from north West Spain. Food Microbiol 70:224–231
Dickschat JS (2010) Quorum sensing and bacterial biofilms. Nat Prod Rep 27:343–369
Dimitrieva GY, Crawford RL, Yuksel GU (2006) The nature of plant growth-promoting effects of a pseudoalteromonad associated with the marine algae Laminaria japonica and linked to catalase excretion. J Appl Microbiol 100:1159–1169
Dittami SM, Gravot A, Renault D, Goulitquer S, Eggert A, Bouchereau A, Boyen C, Tonon T (2011) Integrative analysis of metabolite and transcript abundance during the short-term response to saline and oxidative stress in the brown alga Ectocarpus siliculosus. Plant Cell Environ 34:629–642
Dittami SM, Duboscq-Bidot L, Perennou M, Gobet A, Corre E, Boyen C, Tonon T (2016) Host–microbe interactions as a driver of acclimation to salinity gradients in brown algal cultures. ISME J 10:51–63
Dobretsov S, Teplitski M, Paul V (2009) Mini-review: quorum sensing in the marine environment and its relationship to biofouling. Biofouling 25:413–427
Dogs M, Wemheuer B, Wolter L, Bergen N, Daniel R, Simon M, Brinkhoff T (2017) Rhodobacteraceae on the marine brown alga Fucus spiralis are abundant and show physiological adaptation to an epiphytic lifestyle. Syst Appl Microbiol 40:370–382
Donlan RM (2002) Biofilms: microbial life on surfaces. Emerg Infect Dis 8:881–890
Dworjanyn SA, Pirozzi I (2008) Induction of settlement in the sea urchin tripneustes gratilla by macroalgae, biofilms and conspecifics: a role for bacteria?. Aquaculture 274:268–274
Egan S, Thomas T, Kjelleberg S (2008) Unlocking the diversity and biotechnological potential of marine surface associated microbial communities. Curr Opin Microbiol 11:219–225
Egan S, Harder T, Burke C, Steinberg P, Kjelleberg S, Thomas T (2013) The seaweed holobiont: understanding seaweed–bacteria interactions. FEMS Microbiol Rev 37:462–476
El Shafay SM, Ali SS, El-Sheekh MM (2016) Antimicrobial activity of some seaweeds species from Red Sea, against multidrug resistant bacteria. Egypt J Aquat Res 42:65–74
El-Shouny WA, Gaafar RM, Ismail GA, Elzanaty MM (2017) Antibacterial activity of some seaweed extracts against multidrug resistant urinary tract bacteria and analysis of their virulence genes. Int J Curr Microbiol Appl Sci 6:2569–2586
Fernandes N, Steinberg P, Rusch D, Kjelleberg S, Thomas T (2012) Community structure and functional gene profile of bacteria on healthy and diseased thalli of the red seaweed Delisea pulchra. PLoS One 7(12):e50854
Flemming HC, Wingender J (2001) Relevance of microbial extracellular polymeric substances (EPSs)—part I: structural and ecological aspects. Water Sci Tech 43:1–8
Flemming HC, Wingender J (2010) The biofilm matrix. Nat Rev Microbiol 8:623–633
Friedrich AB, Fischer I, Proksch P, Hacker J, Hentschel U (2001) Temporal variation of the microbial community associated with the Mediterranean sponge Aplysia aerophoba. FEMS Microbial Ecol 38:105–113
Fukui Y, Abe M, Kobayashi M, Yano Y, Satomi M (2014) Isolation of Hyphomonas strains that induce normal morphogenesis in protoplasts of the marine red alga Pyropia yezoensis. Microbiol Ecol 68:556–566
Gachon C, Sime-Ngando T, Strittmatter M, Chambouvet A, Kim GH (2010) Algal diseases: spotlight on a black box. Trends Plant Sci 15:633–640
Garrett TR, Bhakoo M, Zhang Z (2008) Bacterial adhesion and biofilms on surfaces. Prog Nat Sci 18:1049–1056
Ghaderiardakani F, Coates JC, Wichard T (2017) Bacteria-induced morphogenesis of Ulva intestinalis and Ulva mutabilis (Chlorophyta): a contribution to the lottery theory. FEMS Microbiol Ecol 93:1–12
Goecke F, Labes A, Wiese J, Imhoff JF (2010) Chemical interactions between marine macroalgae and bacteria. Mar Ecol Prog Ser 409:267–300
Goecke F, Labes A, Wiese J, Schmaljohann R, Imhoff JF (2012) Observation of bacteria over the surface of released oogonia from Fucus vesiculosus L. (Phaeophyceae). Gayana Bot 69:376–379
Goecke F, Thiel V, Wiese J, Labes A, Imhoff JF (2013) Algae as an important environment for bacteria—phylogenetic relationships among new bacterial species isolated from algae. Phycologia 52:14–24
Graham L, Wilcox L (1999) Algae. Prentice-Hall, Upper Saddle River
Graham LE, Knack JJ, Graham ME, Graham JM, Zulkifly S (2015) A metagenome for lacustrine Cladophora (Cladophorales) reveals remarkable diversity of eukaryotic epibionts and genes relevant to materials cycling. J Phycol 51:408–418
Greff S, Aires T, Serrão EA, Engelen AH, Thomas OP, Pérez T (2017) The interaction between the proliferating macroalga Asparagopsis taxiformis and the coral Astroides calycularis induces changes in microbiome and metabolomic fingerprints. Sci Rep 7:42625
Grossart HP (2010) Ecological consequences of bacterioplankton lifestyles: changes in concepts are needed. Environ Microbiol Rep 2:706–714
Grueneberg J, Engelen AH, Costa R, Wichard T (2016) Macroalgal morphogenesis induced by waterborne compounds and bacteria in coastal seawater. PLoS One 11:e0146307
Hadfield MG (2011) Biofilms and marine invertebrate larvae: what bacteria produce that larvae use to choose settlement sites. Annu Rev Mar Sci 3:453–470
Hellio C, Marechal J-P, Veron B, Bremer G, Clare A, Le Gal Y (2004) Seasonal variation of antifouling activities of marine algae from the Brittany coast (France). Mar Biotechnol 6:67–82
Hengst MB, Andrade S, González B, Correa JA (2010) Changes in epiphytic bacterial communities of intertidal seaweeds modulated by host, temporality and copper enrichment. Microbiol Ecol 60:282–290
HMP (2012) The human microbiome project consortium. Structure, function and diversity of the healthy human microbiome. Nature 486:207–214
Hollants J, Leroux O, Leliaert F, Decleyre H, De Clerck O, Willems A (2011a) Who is in there? Exploration of endophytic bacteria within the siphonous green seaweed Bryopsis (Bryopsidales, Chlorophyta). PLoS One 6:e26458
Hollants J, Decleyre H, Leliaert F, De Clerck O, Willems A (2011b) Life without a cell membrane: challenging the specificity of bacterial endophytes within Bryopsis (Bryopsidales, Chlorophyta). BMC Microbiol 11:e255
Hollants J, Leliaert F, De Clerck O, Willems A (2013) What we can learn from sushi: a review on seaweed–bacterial associations. FEMS Microbiol Ecol 83:1–16
Horincar VB, Parfene G, Tyagi AK, Gottardi D, Dinică R, Guerzoni ME, Bahrim G (2014) Extraction and characterization of volatile compounds and fatty acids from red and green macroalgae from the Romanian Black Sea in order to obtain valuable bioadditives and biopreservatives. J Appl Phycol 26:551–559
Horta A, Pinteus S, Alves C, Fino N, Silva J, Fernandez S, Rodrigues A, Pedrosa R (2014) Antioxidant and antimicrobial potential of the Bifurcaria bifurcata epiphytic bacteria. Mar Drugs 12:1676–1689
Huggett MJ, Williamson JE, de Nys R, Kjelleberg S, Steinberg PD (2006) Larval settlement of the common Australian sea urchin Heliocidaris erythrogramma in response to bacteria from the surface of coralline algae. Oecologia 149:604–619
Ibrahim HAH, Beltagy EA, El-Din NGS, Zokm GME, El-Sikaily AM, Abu-Elela GM (2015) Seaweeds agarophytes and associated epiphytic bacteria along Alexandria coastline, Egypt, with emphasis on the evaluation and extraction of agar and agarose. Rev Biol Mar Oceanogr 50:545–561
Ismail A, Ktari L, Ahmed M, Bolhuis H, Boudabbous A, Stal LJ, Cretoiu MS, El Bour M (2016) Antimicrobial activities of bacteria associated with the brown alga Padina pavonica. Front Microbiol 7:1072
JanakiDevi V, YokeshBabu M, Umarani R, Kumaraguru AK (2013) Antagonistic activity of seaweed associated bacteria against human pathogens. Int J Curr Microbiol App Sci 2:140–147
Jha B, Kavita K, Westphal J, Hartmann A, Schmitt-Kopplin P (2013) Quorum sensing inhibition by Asparagopsis taxiformis, a marine macro alga: Separation of the compound that interrupts bacterial communication. Mar Drugs 11:253–265
Joint I, Tait K, Wheeler G (2007) Cross-kingdom signaling: exploitation of bacterial quorum sensing molecules by the green seaweed Ulva. Philos Trans R Soc Lond B 362:1223–1233
Jung H, Baek G, Kim J, Shin SG, Lee C (2016) Mild-temperature thermochemical pretreatment of green macroalgal biomass: effects on solubilization, methanation, and microbial community structure. Bioresour Technol 199:326–335
Kalia VC (2013) Quorum sensing inhibitors: an overview. Biotechnol Adv 31:224–245
Kalia VC, Purohit HJ (2011) Quenching the quorum sensing system: potential antibacterial drug targets. Crit Rev Microbiol 37:121–140
Kanagasabhapathy M, Yamazaki G, Ishida A, Sasaki H, Nagata S (2009) Presence of quorum-sensing inhibitor like compounds from bacteria isolated from the brown alga Colpomenia sinuosa. Lett Appl Microbiol 49:573–579
Karthick P, Mohanraju R, Murthy KN, Ramesh CH, Mohandass C, Rajasabapathy R, Kumar SV (2015) Antimicrobial activity of Serratia sp isolated from the coralline red algae Amphiroa anceps. Indian J Geomarine Sci 44:1857–1866
Kavita K, Mishra A, Jha B (2011) Isolation and physico-chemical characterisation of extracellular polymeric substances produced by the marine bacterium Vibrio parahaemolyticus. Biofouling 27:309–317
Kientz B, Agogué H, Lavergne C, Marié P, Rosenfeld E (2013) Isolation and distribution of iridescent Cellulophaga and further iridescent marine bacteria in the Charente maritime coast, French Atlantic. coast. Syst Appl Microbiol 36:244–251
Kim JY, Park SH, Seo GY, Kim YJ, Oh DC (2015) Winogradskyella eckloniae sp. nov., a marine bacterium isolated from the brown alga Ecklonia cava. Int J Syst Evol Microbiol 65:2791–2796
Kita A, Miura T, Kawata S, Yamaguchi T, Okamura Y, Aki T, Matsumura Y, Tajima T, Kato J, Nishio N, Nakashimada Y (2016) Bacterial community structure and predicted alginate metabolic pathway in an alginate-degrading bacterial consortium. J Biosci Bioeng 121:286–292
KleinJan H, Jeanthon C, Boyen C, Dittami SM (2017) Exploring the cultivable Ectocarpus microbiome. Front Microbiol 8:2456
Kostakioti M, Hadjifrangiskou M, Hultgren SJ (2013) Bacterial biofilms: development, dispersal and therapeutic strategies in the dawn of the postantibiotic era. Cold Spring Harb Perspect Med 3:a010306
Kouzuma A, Watanabe K (2015) Exploring the potential of algae/bacteria interactions. Curr Opin Biotechnol 33:125–129
Kubanek J, Jensen PR, Keifer PA, Sullards MC, Collins DO, Fenical W (2003) Seaweed resistance to microbial attack: a targeted chemical defense against marine fungi. Proc Natl Acad Sci 100:6916–6921
Kumar V, Zozaya-Valdes E, Kjelleberg S, Thomas T, Egan S (2016) Multiple opportunistic pathogens can cause a bleaching disease in the red seaweed Delisea pulchra. Environ Microbiol 18:3962–3975
La Barre S, Potin P, Leblanc C, Delage L (2010) The halogenated metabolism of brown algae (Phaeophyta), its biological importance and its environmental significance. Mar Drugs 8:988–1010
Lachnit T, Blumel M, Imhoff JF, Wahl M (2009) Specific epibacterial communities on macroalgae: phylogeny matters more than habitat. Aquat Biol 5:181–186
Lachnit T, Wahl M, Harder T (2010) Isolated thallus-associated compounds from the macroalga Fucus vesiculosus mediate bacterial surface colonization in the field similar to that on the natural alga. Biofouling 26:247–255
Lachnit T, Meske D, Wahl M, Harder T, Schmitz R (2011) Epibacterial community patterns on marine macroalgae are host-specific but temporally variable. Environ Microbiol 13:655–665
Lachnit T, Fischer M, Kunzel S, Baines JF, Harder T (2013) Compounds associated with algal surfaces mediate epiphytic colonization of the marine macroalga Fucus vesiculosus. FEMS Microbiol Ecol 84:411–420
Le Bail A, Billoud B, Kowalczyk N, Kowalczyk M, Gicquel M, Panse SL, Stewart S, Scornet D, Cock JM, Ljung K, Charrier B (2010) Auxin metabolism and function in the multicellular brown alga Ectocarpus siliculosus. Plant Physiol 153:128–144
Lee JH, Eom SH, Lee EH, Jung YJ, Kim HJ, Jo MR, Son KT, Lee HJ, Kim JH, Lee MS, Kim YM (2014) In vitro antibacterial and synergistic effect of phlorotannins isolated from edible brown seaweed Eisenia bicyclis against acne-related bacteria. Algae 29:47–55
Liu M, Liu Y, Cao MJ, Liu GM, Chen Q, Sun L, Chen H (2017) Antibacterial activity and mechanisms of depolymerized fucoidans isolated from Laminaria japonica. Carbohydr Polym 172:294–305
Maheswari MU, Reena A, Sivaraj C (2017) GC-MS analysis, antioxidant and antibacterial activity of the brown algae, Padina tetrastromatica. Int J Pharm Sci Res 8:4014–2400
Mandal SK, Singh RP, Patel V (2011) Isolation and characterization of exopolysaccharide secreted by a toxic dinoflagellate, Amphidinium carterae Hulburt 1957 and its probable role in harmful algal blooms (HABs). Microb Ecol 62:518–527
Manilal A, Sujith S, Sabarathnam B, Kiran G, Selvin J, Shakir C, Lipton AP (2010) Bioactivity of the red alga Asparagopsis taxiformis collected from the south-western coast of India. Braz J Oceonogr 58:93–100
Manivannan K, Karthikai devi G, Anantharaman P, Balasubramanian T (2011) Antimicrobial potential of selected brown seaweeds from Vedalai coastal waters, Gulf of Mannar. Asian Pac J Trop Biomed 1:114–120
Marshall K, Joint I, Callow ME, Callow JA (2006) Effect of marine bacterial isolates on the growth and morphology of axenic plantlets of the green alga Ulva linza. Microb Ecol 52:302–310
Martin M, Portetelle D, Michel G, Vandenbol M (2014) Microorganisms living on macroalgae, diversity, interactions and biotechnological applications. Appl Microbiol Biot 98:2917–2935
Martin M, Barbeyron T, Martin R, Portetelle D, Michel G, Vandenbol M (2015) The cultivable surface microbiota of the brown alga Ascophyllum nodosum is enriched in macroalgal polysaccharide degrading bacteria. Front Microbiol 6:1487
Menaa F (2015) Tapping into deep-water reservoirs to overcome antibiotic resistance through bacteria-producing unique secondary metabolites. Pharmaceut Analyt Acta 6:e172
Meena VD, Dotaniya ML, Saha JK, Patra AK (2015) Antibiotics and antibiotic resistant bacteria in wastewater: impact on environment, soil microbial activity and human health. Afr J Microbiol Res 9:965–978
Mesquita MMF, Baddini ALQ, Netto ADP, Araujo JM, Salgueiro F, Filho EAPL, De-Paula JC, Fleury BG, Cavalcanti DN, Teixeira VL (2015) Chemical similarity between Dictyota caribaea and Dictyota menstrualis (Dictyotaceae, Phaeophyceae) from the coast of Rio de Janeiro, Brazil. Biochem Syst Ecol 58:97–101
Miranda LN, Hutchison K, Grossman AR, Brawley SH (2013) Diversity and abundance of the bacterial community of the red macroalga Porphyra umbilicalis: did bacterial farmers produce macroalgae? PLoS One 8:e58269
Moon HE, Islam N, Ahn BR, Chowdhury SS, Sohn HS, Jung HA, Choi JS (2011) Protein tyrosine phosphatase 1B and α-glucosidase inhibitory phlorotannins from edible brown algae, Ecklonia stolonifera and Eisenia bicyclis. Biosci Biotechnol Biochem 75:1472–1480
Moorthi PV, Balasubramanian C (2015) Antimicrobial properties of marine seaweed, Sargassum muticum against human pathogens. J Coast Life Med 3:122–125
Murray PM, Moane S, Collins C, Beletskaya T, Thomas OP, Duarte AW, Nobre FS, Owoyemi IO, Pagnocca FC, Sette LD, McHugh E, Causse E, Pérez-López P, Feijoo G, Moreira MT, Rubiolo J, Leirós M, Botana LM, Pinteus S, Alves C, Horta A, Pedrosa R, Jeffryes C, Agathos SN, Allewaert C, Verween A, Vyverman W, Laptev I, Sineoky S, Bisio A, Manconi R, Ledda F, Marchi M, Pronzato R, Walsh DJ (2013) Sustainable production of biologically active molecules of marine based origin. New Biotechnol 30:839–850
Murugan A, Begum MS, Ramasamy MS, Raja P (2012) Antifouling and antipredatory activity of natural products of the seaweeds Dictyota dichotoma and Chaetomorpha linoides. Nat Prod Res 26:975–978
Nylund GM, Persson F, Lindegarth M, Cervin G, Hermansson M, Pavia H (2010) The red alga Bonnemaisonia asparagoides regulates epiphytic bacterial abundance and community composition by chemical defence. FEMS Microbiol Ecol 71:84–93
Nylund GM, Enge S, Pavia H (2013) Costs and benefits of chemical defense in the red alga Bonnemaisonia hamifera. PLoS One 8(4):e61291
Ojha AK, Verma A, Pal Y, Bhatt D, Mayilraj S, Krishnamurthi S (2017) Marinomonas epiphytica sp. nov., isolated from a marine intertidal macroalga. Int J Syst Evol Microbiol 67:2746–2751
Othmani A, Bunet R, Bonnefont JL, Briand JF, Culioli G (2016) Settlement inhibition of marine biofilm bacteria and barnacle larvae by compounds isolated from the Mediterranean brown alga Taonia atomaria. J Appl Phycol 28:1975–1986
Pedersén M, (1968) Ectocarpus fasciculatus: Marine brown alga requiring kinetin. Nature 218 (5143):776–776
Penesyan A, Marshall-Jones Z, Holmstrom C, Kjelleberg S, Egan S (2009) Antimicrobial activity observed among cultured marine epiphytic bacteria reflects their potential as a source of new drugs. FEMS Microbiol Ecol 69:113–124
Penesyan A, Kjelleberg S, Egan S (2010) Development of novel drugs from marine surface associated microorganisms. Mar Drugs 8:438–459
Penesyan A, Gillings M, Paulsen IT (2015) Antibiotic discovery: combatting bacterial resistance in cells and in biofilm communities. Molecules 20:5286–5298
Penesyan A, Tebben J, Lee M, Thomas T, Kjelleberg S, Harder T, Egan S (2011) Identification of the antibacterial compound produced by the marine epiphytic bacterium pseudovibrio sp. D323 and related sponge-associated bacteria. Mar Drugs 9(8):1391–1402
Peters A, Marie D, Scornet D, Kloareg B, Cock J (2004) Proposal of Ectocarpus siliculosus (Ectocarpales, Phaeophyceae) as a model organism for brown algal genetic sand genomics. J Phycol 40:1079–1088
Potin P (2008) Oxidative burst and related responses in biotic interactions of algae. In: Amsler CH (ed) Algal chemical ecology. Springer, Berlin, pp 245–271
Prasad K, Das AK, Oza MD, Brahmbhatt H, Siddhanta AK, Meena R, Eswaran K, Rajyaguru MR, Ghosh PK (2010) Detection and quantification of some plant growth regulators in a seaweed-based foliar spray employing a mass spectrometric technique sans chromatographic separation. J Agric Food Chem 58:4594–4601
Provasoli L, Carlucci AF (1974) Vitamins and growth regulators. In: Stewart WDP (ed) Algal physiology and biochemistry. Blackwell, Oxford, pp 74–87
Rahelivao MP, Gruner M, Andriamanantoanina H, Bauer I, Knolker HJ (2015) Brown algae (Phaeophyceae) from the coast of Madagascar: preliminary bioactivity studies and isolation of natural products. Nat Prod Bioprospect 5:223–235
Rajivgandhi G, Vijayan R, Kannan M, Santhanakrishnan M, Manoharan N (2016) Molecular characterization and antibacterial effect of endophytic actinomycetes Nocardiopsis sp. GRG1 (KT235640) from brown algae against MDR strains of uropathogens. Bioactive Materials 1:140–150
Rao D, Webb JS, Holmström C, Case R, Low A, Steinberg P, Kjelleberg S (2007) Low densities of epiphytic bacteria from the marine alga Ulva australis inhibit settlement of fouling organisms. Appl Environ Microbiol 73:7844–7852
Ravisankar A, Gnanambal ME, Sundaram LR (2013) A newly isolated Pseudomonas sp., epibiotic on the seaweed, Padina tetrastromatica, off southeastern coast of India, reveals antibacterial action. Appl Biochem Biotechnol 171:1968–1985
Rizzo L, Pusceddu A, Stabili L, Alifano P, Fraschetti S (2017) Potential effects of an invasive seaweed (Caulerpa cylindracea, Sonder) on sedimentary organic matter and microbial metabolic activities. Sci Rep 7:12113. https://doi.org/10.1038/s41598-017-12556-4
Salaün S, Kervarec N, Potin P, Haras D, Piotto M, La Barre S (2010) Whole-cell spectroscopy is a convenient tool to assist molecular identification of cultivatable marine bacteria and to investigate their adaptive metabolism. Talanta 80:1758–1770
Satheesh S, Ba-akdah MA, Al-Sofyani AA (2016) Natural antifouling compound production by microbes associated with marine macroorganisms—a review. Electron J Biotechnol 21:26–35
Schmidt TSB, Rodrigues JFM, von Mering C (2014) Ecological consistency of SSU rRNA-based operational taxonomic units at a global scale. PLoS Comput Biol 10(4):e1003594
Schwartz N, Rohde S, Dobretsov S, Hiromori S, Schupp PJ (2017) The role of chemical antifouling defence in the invasion success of Sargassum muticum: a comparison of native and invasive brown algae. PLoS One 12(12):e0189761
Silva FS, Bitencourt JAP, Savergnini F, Guerra LV, Baptista-Neto JA, Crapez MAC (2011) Bioavailability of organic matter in the superficial sediment of Guanabara Bay, Rio de Janeiro, Brazil. Anuario do Instituto de Geociencias 34:52–63
Singh RP, Reddy CRK (2014) Seaweed–microbial interactions: key functions of seaweed-associated bacteria. FEMS Microbiol Ecol 88:213–230
Singh RP, Mantri VA, Reddy CRK, Jha B (2011a) Isolation of seaweed-associated bacteria and their morphogenesis inducing capability in axenic cultures of the green alga Ulva fasciata. Aquat Biol 12:13–21
Singh RP, Bijo AJ, Baghel RS, Reddy CRK, Jha B (2011b) Role of bacterial isolates in enhancing the bud induction in the industrially important red alga Gracilaria dura. FEMS Microbiol Ecol 76:381–392
Singh RP, Shukla MK, Mishra A, Kumari P, Reddy CRK, Jha B (2011c) Isolation and characterization of exopolysaccharides from seaweed-associated bacteria Bacillus licheniformis. Carbohydr Polym 84:1019–1026
Singh RP, Shukla MK, Mishra A, Reddy CRK, Jha B (2013) Bacterial extracellular polymeric substances and their effect on settlement of zoospore of Ulva fasciata. Colloids Surf B 103:223–230
Singh RP, Baghel RS, Reddy CRK, Jha B (2015) Effect of quorum sensing signals produced by seaweed-associated bacteria on carpospores liberation from Gracilaria dura. Front Plant Sci 6:117
Sousa C, Gangadhar KN, Morais TR, Conserva GA, Vizetto-Duarte C, Pereira H, Laurenti MD, Campino L, Levy D, Uemi M, Barreira L, Custódio L, Passero LF, Lago JH, Varela J (2017) Antileishmanial activity of meroditerpenoids from the macroalgae Cystoseira baccata. Exp Parasitol 174:1-9
Sneed JM, Pohnert G (2010) The green macroalga Dictyosphaeria ocellata influences the structure of the bacterioplankton community through differential effects on individual bacterial phylotypes. FEMS Microbiol Ecol 75:242–254
Sneed JM, Pohnert G (2011) The green alga Dicytosphaeria ocellata and its organic extracts alter natural bacterial biofilm communities. Biofouling 27:347–356
Soares AR, da Gama BAP, da Cunha AP, Teixeira VL, Pereira RC (2008) Induction of attachment of the mussel Perna perna by natural products from the brown seaweed Stypopodium zonale. Mar Biotechnol 10:158–165
Spavieri J, Allmendinger A, Kaiser M, Casey R, Hingley-Wilson S, Lalvani A, Guiry MD, Blunden G, Tasdemir D (2010) Antimycobacterial, antiprotozoal and cytotoxic potential of twenty-one brown algae (Phaeophyceae) from British and Irish waters. Phytother Res 24:1724–1729
Spilling K, Titelman J, Greve TM, Kühl M (2010) Microsensor measurements of the external and internal microenvironment of Fucus vesiculosus (Phaeophyceae). J Phycol 46:1350–1355
Spoerner M, Wichard T, Bachhuber T, Stratmann J, Oertel W (2012) Growth and thallus morphogenesis of Ulva mutabilis (Chlorophyta) depends on a combination of two bacterial species excreting regulatory factors. J Phycol 48:1433–1447
Staufenberger T, Thiel V, Wiese J, Imhoff JF (2008) Phylogenetic analysis of bacteria associated with Laminaria saccharina. FEMS Microbiol Ecol 64:65–77
Steinberg PD, de Nys R (2002) Chemical mediation of colonization of seaweed surfaces. J Phycol 38:621–629
Steinberg PD, de Nys R, Kjelleberg S (1998) Chemical inhibition of epibiota by Australian seaweeds. Biofouling 12:227–244
Steinberg PD, de Nys R, Kjelleberg S (2002) Chemical cues for surface colonization. J Chem Ecol 28:1935–1951
Suresh M, Iyapparaj P, Anantharaman P (2016) Antifouling activity of lipidic metabolites derived from Padina tetrastromatica. Appl Biochem Biotechnol 179:805–818
Susilowati R, Sabdono A, Widowati I (2015) Isolation and characterization of bacteria associated with brown algae Sargassum spp. from Panjang Island and their antibacterial activities. Procedia Environ Sci 23:240–246
Tapia JE, González B, Goulitquer S, Potin P, Correa JA (2016) Microbiota influences morphology and reproduction of the brown alga Ectocarpus sp. Front Microbiol 7:197
Teixeira VL (2013) Produtos naturais de algas marinhas bentônicas. Rev Virtual Quim 5:343–362
Thennarasan S, Murugesan S (2015) Antibacterial activity of crude methanolic extract of marine brown alga Lobophora variegata (J.V. Lamouroux). World J Pharm Res 4:1714–1722
Torralba MG, Franks JS, Gomez A, Yooseph S, Nelson KE, Grimes DJ (2017) Effect of Macondo Prospect 252 oil on microbiota associated with pelagic Sargassum in the northern Gulf of Mexico. Microb Ecol 73:91–100
Trias R, García-Lledó A, Sánchez N, López-Jurado JL, Hallin S, Bañeras L (2012) Abundance and composition of epiphytic bacterial and archaeal ammonia oxidizers of marine red and brown macroalgae. Appl Environ Microbiol 78:318–325
Tujula NA, Holmström C, Mußmann M, Amann R, Kjelleberg S, Crocetti GR (2006) A CARD-FISH protocol for the identification and enumeration of epiphytic bacteria on marine algae. J Microbiol Methods 65:604–607
Tujula NA, Crocetti GR, Burke C, Thomas T, Holmström C, Kjelleberg S (2010) Variability and abundance of the epiphytic bacterial community associated with a green marine ulvacean alga. ISME J 4:301–311
Twigg MS, Tait K, Williams P, Atkinson S, Cámara M (2013) Interference with the germination and growth of Ulva zoospores by quorum-sensing molecules from Ulva-associated epiphytic bacteria. Environ Microbiol 16:445–453
Uzair B, Menaa F, Khan BA, Mohammad FV, Ahmad VU, Djeribi R, Menaa B (2018) Isolation, purification, structural elucidation and antimicrobial activities of Kocumarin, a novel antibiotic isolated from actinobacterium Kocuria marina CMG S2 associated with the brown seaweed Pelvetia canaliculata. Microbiol Res 206:186–197
Vaikundamoorthy R, Krishnamoorthy V, Vilwanathan R, Rajendran R (2018) Structural characterization and anticancer activity (MCF7 and MDA-MB-231) of polysaccharides fractionated from brown seaweed Sargassum wightii. Int J Biol Macromol 111:1229–1237
Vairappan CS, Suzuki M, Motomura T, Ichimura T (2001) Pathogenic bacteria associated with lesions and thallus bleaching symptoms in the Japanese kelp Laminaria religiosa Miyabe (Laminariales, Phaeophyceae). Hydrobiologia 445:183–191
Vairappan CS, Chung CS, Hurtado AQ, Soya FE, Lhonneur GB, Critchley A (2008) Distribution and symptoms of epiphyte infection in major carrageenophyte-producing farms. J Appl Phycol 20:477–483
Vairappan CS, Anangdan SP, Tan KL, Matsunaga S (2010) Role of secondary metabolites as defense chemicals against ice-ice disease bacteria in biofouler at carrageenophyte farms. J Appl Phycol 22:305–311
Vesty EF, Kessler RW, Wichard T, Coates JC (2015) Regulation of gametogenesis and zoosporogenesis in Ulva linza (Chlorophyta): comparison with Ulva mutabilis and potential for laboratory culture. Front Plant Sci 6:1–8
Vieira C, Engelen AH, Guentas L, Aires T, Houlbreque F, Gaubert J, Serrão EA, De Clerck O, Payri CE (2016) Species specificity of bacteria associated to the brown seaweeds Lobophora (Dictyotales, Phaeophyceae) and their potential for induction of rapid coral bleaching in Acropora muricata. Front Microbiol 7:316
Wahl M, Goecke F, Labes A, Dobretsov S, Weinberger F (2012) The second skin: ecological role of epibiotic biofilms on marine organisms. Front Microbiol 3:292
Wang G, Shuai L, Li Y, Lin W, Zhao X, Duan D (2008) Phylogenetic analysis of epiphytic marine bacteria on hole-rotten diseased sporophytes of Laminaria japonica. J Appl Phycol 20:403–409
Weinberger F, Beltran J, Correa JA, Pohnert G, Kumar N, Steinberg P, Kloareg B, Potin P (2007) Spore release in Acrochaetium sp. (Rhodophyta) is bacterially controlled. J Phycol 43:235–241
Wichard T (2015) Exploring bacteria-induced growth and morphogenesis in the green macroalga order Ulvales (Chlorophyta). Front Plant Sci 6:86
Wiese J, Thiel V, Nagel K, Staufenberger T, Imhoff JF (2009) Diversity of antibiotic active bacteria associated with the brown alga Laminaria saccharina from the Baltic Sea. Mar Biotechnol 11:287–300
Yagi H, Fujise A, Itabashi N, Ohshiro T (2016) Purification and characterization of a novel alginate lyase from the marine bacterium Cobetia sp. NAP1 isolated from brown algae. Biosci Biotechnol Biochem 80:2338–2346
Zozaya-Valdes E, Egan S, Thomas T (2015) A comprehensive analysis of the microbial communities of healthy and diseased marine macroalgae and the detection of known and potential bacterial pathogens. Front Microbiol 6:146
Zozaya-Valdes E, Roth-Schulze AJ, Egan S, Thomas T (2017) Microbial community function in the bleaching disease of the marine macroalgae Delisea pulchra. Environ Microbiol 19:3012–3024
Acknowledgements
The first author is thankful to Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (Capes) for the PhD fellowship. This study was supported by Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) and Fundação de Amparo à Pesquisa do Estado do Rio de Janeiro (FAPERJ). The authors also thank the collaboration of MSc. Ana Débora Nunes Pinheiro in the review of English and suggestions for this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
de Mesquita, M.M.F., Crapez, M.A.C., Teixeira, V.L. et al. Potential interactions bacteria-brown algae. J Appl Phycol 31, 867–883 (2019). https://doi.org/10.1007/s10811-018-1573-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10811-018-1573-4