Abstract
Excessive use of pesticides and herbicides is a major environmental and health concern worldwide. Atrazine, a synthetic triazine herbicide commonly used to control grassy and broadleaf weeds in crops, is a major pollutant of soil and water ecosystems. Atrazine modifies the growth, enzymatic processes and photosynthesis in plants. Atrazine exerts mutagenicity, genotoxicity, defective cell division, erroneous lipid synthesis and hormonal imbalance in aquatic fauna and nontarget animals. It has threatened the sustainability of agricultural soils due to detrimental effects on resident soil microbial communities. The detection of atrazine in soil and reservoir sites is usually made by IR spectroscopy, ELISA, HPLC, UPLC, LC–MS and GC–MS techniques. HPLC/LC–MS and GC–MS techniques are considered the most effective tools, having detection limits up to ppb levels in different matrices. Biodegradation of atrazine by microbial species is increasingly being recognized as an eco-friendly, economically feasible and sustainable bioremediation strategy. This review presents the toxicity, analytical techniques, abiotic degradation and microbial metabolism of atrazine.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Atrazine (6-chloro-N-ethyl-N′-(1-methylethyl)-1,3,5-triazine-2,4-diamine is a synthetic triazine herbicide used to control grassy and broadleaf weeds in sugarcane, wheat, conifers, sorghum, nuts and corn crops (Iriel et al. 2014; Kumar et al. 2016; Zhao et al. 2017). It was first introduced in twentieth century and often used alone or in amalgamation with other herbicides for agricultural applications (Correia et al. 2007; Kadian et al. 2008; Lewis et al. 2009). It is the second most widely consumed pesticide in the world with annual consumption of about 70,000–90,000 tons (Kumar et al. 2013; Cheng et al. 2016). In India, about 340 tonnes of atrazine is consumed annually (Solomon et al. 2013a, b).
Due to its long half-life of 41–231 days (Karlsson et al. 2016), low adsorption in soils and moderate aqueous solubility, it has a sky-scraping potential to contaminate not only agricultural fields, but also ground and surface water with the highest concentration up to 30 μg/L (Table 1) (Cerejeira et al. 2003; Schwab et al. 2006; Kumar et al. 2013). It was banned in several countries like Italy (Huang et al. 2009), Denmark (Glæsner et al. 2014), Finland and Germany (Vonberg et al. 2014) in the year 1991 and European Union banned atrazine in the year 1992 (White 2016) because its metabolites/residues had the potential to persist in fields and surface water for several years (Bethsass and Colangelo 2013; Nousiainen et al. 2015) resulting in the contamination of surface and water bodies. Some studies reveal the surpass levels of atrazine 3 and 0.1 μgL−1 found in drinking water of the USA and Europe (Mahía et al. 2007). The maximum acceptable concentration (MAC) for atrazine in drinking water is 5 µg/L (Cerejeira et al. 2003). The acceptable daily intake (ADI) is derived on the basis of division of a NOAEL by appropriate uncertainty factors. It is one of the most widely used herbicide, and several times it has been reported to be at levels above the limits in water bodies (Graymore et al. 2001).
The production of atrazine increased from 0.26 metric tonnes to 0.67 metric tonnes from the year 2009 to 2012 (Ministry of fertilizers and chemicals, Govt. of India). Environment protection agency classifies atrazine in toxicity class III on a scale of I to IV (I being the most toxic). It is registered for only two crops (apple and sugar) by the central insecticides board and registration committee in India (Bhushan et al. 2013). Atrazine has been classified as an endocrine disrupting pesticide by the US Environmental Protection Agency (Morales-Pérez et al. 2016). The International Agency for Research on Cancer (IARC) has categorized atrazine in the list of carcinogenic pesticides (Mahler et al. 2017). This review covers the toxicity, analytical monitoring by using recent techniques and degradation (chemical, photochemical and microbial) aspects of atrazine.
Toxicity of atrazine
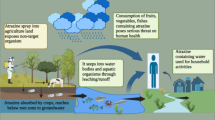
Agricultural chemicals have potential to alter species composition, decrease diversity, interfere with normal succession patterns and alter food webs as a whole (Lin et al. 2016a, b, c). The intensification of industrial and agricultural practices chiefly the utilization of pesticides has in almost every way made our natural resources miserable (Dutta et al. 2016). Various toxic effects of atrazine on humans, plants, animals and microorganisms are displayed in Fig. 1.
Effect on nontarget plants
One of the most unintentional exposures of pesticides is to the nontarget plants. Atrazine has shown to manifest complete death or stunted growth, translocation, root or shoot uptake, phenotype alteration, mutation and resistance (Burken and Schnoor 1997; Szigeti and Lehoczki 2003; Hassan and Nemat 2005; Nemat and Hassan 2006; Su and Zhu 2006). The different results of atrazine accumulation could be due to the use of different plant species, since distinct mechanisms control the accumulation of atrazine in target and nontarget plants. Concentration- and time-dependent effects of atrazine exposure have been noticed. Atrazine is absorbed by shoots and roots and transported solely by xylem (Francisco 2001; Szigeti and Lehoczki 2003). Pre-emergence herbicides are used to stop the germination of plant seeds. Atrazine is absorbed by leaves as well as roots and has postemergence as well as pre-emergence activity on weeds (Ahrens and Newton 2008). The negative effects of postemergence atrazine treatment upon peroxidase, ascorbate peroxidase and lipid peroxidation were determined in a 15-day experiment (Akbulut and Yigit 2010). Target and nontarget plants exposed to atrazine usually suffer oxidative stress caused by the generation of reactive oxygen species such as superoxide anion radical, hydroxyl radical, singlet oxygen and hydrogen peroxide. Generation of H2O2 in roots of maize exposed to atrazine was also assayed after chemical extraction in a previous study. The most important mechanism used by plants to prevent oxidative stress is through reactive oxygen species scavenging (Burken and Schnoor 1997; Hassan and Nemat 2005; Nemat and Hassan 2006; Szigeti and Lehoczki 2003; Su and Zhu 2006). It has been noticed that the exposure and accumulation of atrazine cause oxidative toxicity and antioxidant response in Zea mays (Li et al. 2012). Gao et al. (2011) indicated atrazine to be a potential threat to seagrass seedling function, and the impact is much higher for adult plants. Exposure to 10 μg/L atrazine significantly lowers the plant fresh weight and total chlorophyll concentration, and up to 86.67% mortality was recorded at the 100 μg/L concentration.
Basically plants have detoxification ability against various pesticides. Enzymes detected in plants responsible for detoxification of various pesticides are cytochrome 450, peroxidases, aryl acylamidases, esterases, lipases, proteases, amidases, oxygenases and reductases (Jiang et al. 2016). It has been noticed that the increasing antioxidant enzyme activities enable Pennisetum americanum seedlings to cope with the oxidative stress induced by moderate concentrations (20 mg kg−1 or below) of atrazine. Cytochrome P450 monooxygenase genes are known to be involved in modification and detoxification of herbicide atrazine in Oryza sativa (McGregor et al. 2008; Tana et al. 2015). Exposure to atrazine can trigger specific GT genes and enzyme activities in Oryza sativa (Lua et al. 2013). Another major detoxification mechanism in leaf tissue of maize is through glutathione conjugation (GS-atrazine). It is considered to be an important biotransformation mechanism of a atrzine in plants. The recovery of atrazine-inhibited photosynthesis is accompanied by a rapid conversation of atrazine to GS-atrazine when atrazine was directly introduced into the leaf tissue. This pathway is relatively inactive in roots and shoots (Shimabukuro et al. 1970). Another detoxification pathway for atrazine in corn corresponds to a chemical hydroxylation process. The mechanism was also described under invitro conditions. Benzoxazinones mixture (DIMBOA, DIBOA, 2-monoglucosyl DIMBOA + 2-monoglucosyl DIBOA) extracted from corn plantlets were able to transform 91% of atrazine into 2-hydroxyatrazine in 24 h. The natural concentration of benzoxazinones in the vacuolar sap of corn seedlings and the pH play a major role in high rate of atrazine chemical hydroxylation in vivo (Raveton et al. 1997). Occurrence of atrazine in water bodies can have serious detrimental effects on nontarget living organisms such as freshwater algae (Baxter et al. 2015, 2016; Bai et al. 2015; Andrus et al. 2013). Atrazine inhibited the growth of Chlamydomonas mexicana and also leads to an increase in carbohydrate content and chlorophyll a accumulation (Kabra et al. 2014). The toxicity of atrazine along with its metabolites desethylatrazine and deisopropylatrazine was evaluated on the amphipods Hyalella azteca and Diporeia sp., and the unicellular algae Pseudokirchneriella subcapitata. It was found to be the most toxic followed by desethylatrazine and deisopropylatrazine and algae being the most sensitive of all. In case of chronic exposure, Diporeia sp. was found to be sensitive as compared to H. azteca by a large magnitude (Ralston-Hooper et al. 2009). Photosynthetic process, cell division, lipid synthesis were majorly affected in green alga, Raphidocelis subcapitata on exposure to atrazine (Ma et al. 2006). According to Solomon et al. (2013a, b), atrazine was not found to cause any lethality or permanent cell damage, but it acts to inhibit photo-phosphorylation.
Effects on aquatic fauna
Atrazine exhibited significant rate of micronuclei and nuclear abnormalities in Channa punctatus (Nwani et al. 2011). Atrazine showed acute toxicity to leopard frog (Rana pipiens), American toad (Bufo americanus), rainbow trout (Onchorhynchus mykiss) and channel catfish (Ictalurus punctatus) (Orton et al. 2006). Chironomus tentans was also studied for the effect of atrazine and binary combination of atrazine with chlorpyriphos. The result depicted atrazine not acutely toxic at even higher levels but when used in combination with chlorpyriphos, methyl parathion and malathion decreases the EC50 (effective concentration) values (Belden and Lydy 2001).
Also the toxicity of active ingredients to pesticide formulations with regard to atrazine, chlorpyriphos and permethrin in glochidia and juvenile life stages of a freshwater mussel (Lampsilis siliquoidea) were compared. The atrazine formulation (Aatrez) was more toxic than technical grade atrazine in chronic tests with juvenile L. siliquoidea. For other pesticides, acute and chronic toxicity of technical grade pesticides were similar to the toxicity of pesticide formulations. Atrazine and formulations did not cause any significant acute toxicity in glochidia and juveniles (Bringolf et al. 2007). In case of freshwater fish, Rhamdia quelen, histopathological changes in liver revealed leukocyte infiltration, hepatocyte vacuolization like steatosis and necrosis areas, leading to raised lesion index levels in all tested concentrations; process of osmoregulation was disturbed and gills showed changes in pavement cells and chloride cells (Mela et al. 2013). The effect of atrazine was also evaluated on some immune parameters of red-eared slider (Trachemys scripta). Lowered serum complement and lysozyme activities, reduced leukocyte number as well as their phagocytic activity and increased neutrophil/lymphocyte ratio depicted a positive correlation between atrazine (high dose) concentration and immunosuppressive effects (Soltanian 2016). The expression of carp HSP70 and 70-kDa heat shock cognate protein (HSC70) with atrazine and chloropyriphos treatment alone or in combination was significantly up-regulated in common carp (Cyprinus carpio L.) providing new insights into the mechanisms used by fish to adapt to stressful environment (Xing et al. 2013). Increased lipid peroxidation and decline in cholesterol and total proteins in liver and muscles were observed for atrazine, glyphosate and quinclorac in tadpoles of Lithobates catesbeianus (Dornelles and Oliveira 2014).
Freshwater clam, Corbicula fluminea, was also studied to evaluate the biochemical and genotoxic effects of the herbicides atrazine and Roundup. Atrazine interfered mostly in biotransformation, while Roundup interfered mainly in antioxidant defenses leading to lipid peroxidation. Herbicides mixture caused a significant increase in the occurrence of DNA damage (Dos-Santos and Martinez 2014). Atrazine shows profound influence on the oxidative stress markers and detoxifying enzyme of the exposed zebra fish (Blahova et al. 2013). Atrazine also causes changes in the glutathione S-transferase isoenzymes (GSTs) activity and their transcription varied within each organ and among organs of common carp (Xing et al. 2012). Atrazine behaves as enzyme inhibitor, impairing hepatic metabolism, and produces genotoxic damage to different cell types as studied in Plotosus lineatus (Santos and Martinez 2012). Various concentrations of atrazine also lead to continuous decline in levels of total protein and serum albumin in grass carp, Ctenopharyngodon idella (Khan et al. 2016a, b). Atrazine is also known to show detrimental effects on the digestive gland of Crassostrea gigas, pacific oyster by modulating important molecular and biochemical parameters within relatively short time period (Lee et al. 2017). Exposure to atrazine may be associated with decreased birth weight and preterm delivery. According to Zadeh et al. (2016), hematological parameters like hemoglobin, hematocrit and RBCs were significantly decreased by chronic toxicity of atrazine in fish, Tor grypus exposed to different levels of atrazine. The increase in concentrations of lactate dehydrogenase (LDH) and decrease in concentrations of creatinine phosphokinases (CPK), serum glutamic-pyruvic transaminase (SGPT) and alkaline phosphatase indicate an adverse effect of atrazine on grass carp, Ctenopharyngodon Idella (Khan et al. 2016a, b). In Channa punctatus, the biochemical parameters such as serum total protein, glucose and cholesterol values were found to decrease, while level of urea significantly increased in all treatments suggesting anemia and hepatic damage. In a risk assessment study of atrazine in American surface waters, it was found that phytoplankton were the most sensitive organisms followed by macrophytes, benthic invertebrates, zooplanktons and fish. In estuarine crab, Neohelice granulata significant decrease in glycogen content and significant diminished content of vitellogenin proteins in ovary was detected after atrazine toxicity (Silveyra et al. 2017). In zebrafish, Danio rerio decrease in glutathione S-transferase and catalase and an increase in superoxide dismutase, glutathione peroxidase and reductase were observed indicating profound influence of atrazine on the oxidative stress markers and detoxifying enzymes (Blahova et al. 2013). Atrazine also affects gill respiration and ion regulation function of fingerlings (Caspian kutum) by damaging tissue, pavement cells and ionocytes (Khoshnood et al. 2015).
Effects on other invertebrates and vertebrates
Oluah et al. (2016) studied the toxicity and the histopathological effects of Atrazine on earthworms, Nsukkadrilus mbae, and reported significant adverse effects. Xu et al. (2006) compared atrazine and chlorotoluron toxicity on Eisenia fetida and found atrazine as more toxic to earthworms; combination showed synergistic effect. Superoxide dismutase (SOD) activity showed an increase. The exposure of chlorpyrifos to E. fetida in combination with atrazine or cyanazine was evaluated, in which the resultant toxicity was greater than the additive (Lydy and Linck 2003). Previous studies suggest that the pessimistic effects of atrazine on neuro-endocrine system occur by changing pituitary hormone levels, such as follicle-stimulating hormone (FSH) and luteinizing hormone (LH) (Cox 2001; Yang et al. 2014). Altered LH leads to prolonged prolactin secretion and subsequent stimulation of the mammary gland proliferative changes and increased incidence of mammary adenocarcinomas and fibroadenomas (Jowa and Howd 2011; Simpkins et al. 2011). Atrazine inhibits the release of gonadotropin-releasing hormone (GnRH), which decreases the secretion of LH that may lead to increased abortions in male Wistar rats (Stoker et al. 2002). Exposure to atrazine affects both germ cells as reduced motility and sperm counts in male rats (Victor-Costa et al. 2010: Pogrmic et al. 2009). Effects of atrazine were also observed as a decrease in levels of serum lipids and liver enzymes in adult mammals (Suzawa and Ingraham 2008: Hayes et al. 2002). Atrazine supposedly increases aromatase enzyme activity via inhibition of phosphodiesterase, which increases the aromatization of testosterone to estrogen (Hayes et al. 2006; Cooper et al. 2007). An increased estrogenic environment may favor altered relative sex hormone levels may affect reproduction and/or development and induction of cancers and proliferation of preexisting estrogen-dependent cancers (Oka et al. 2008). Other effects of metabolites of atrazine on the immune system, central nervous system and cardiovascular function have also been reported (Jin et al. 2010; Christin et al. 2003). In adult humans, non-Hodgkin’s lymphoma associates with the exposure of the compound (Schroeder et al. 2001). According to Gely-Pernot et al. (2015), atrazine exposure interferes with normal meiosis, which affects spermatozoa production in male mice. Oxidative stress and disruptions in calcium homeostasis play an important role in the induction of immunotoxicity in mice by atrazine as depicted by Gao et al. (2016). It is also found to elicit immunotoxic effects on murine lymphocytes, and its presence in the environment might compromise immune function in organisms (Chen et al. 2015a, b). Atrazine is known to reduce the mating rate, number of progeny and competitive fertilization ability; it also alters gene expression and production of proteins in Drosophila melanogaster (Vogel et al. 2015). Marcus and Fiumera (2016) observed reduced pupation rate, emergence rate and longevity of adult in D. Melanogaster after atrazine exposure. Atrazine was also reported to be endocrine disrupters and significant decrease in steroid levels (testosterone and 17β-estradiol), and total proteins were also noted; the histology of ovotestis showed degenerative changes including azoospermia and oocytes deformation (Omran and Salama 2016). 57% reduction in testicular volume was marked in atrazine exposed tadpoles. Also, primary spermatogonial cells and nurse cells were reduced by 70 and 74%, respectively (Tavera-Mendoza et al. 2002). Toxicity profiles of atrazine against microorganisms, aquatic lower invertebrates, higher vertebrates and humans are presented in Table 2.
The prime target of chlorinated atrazine on humans and mammals is the disruption of the endocrine system (Jin et al. 2014; Kroon et al. 2014; Weber et al. 2013). Secondly, it also induces oxidative stress by formation of reactive oxygen species resulting in the reduced semen quality sperm dysfunction and infertility of amphibians, rats and pigs (Gely-Pernot et al. 2015; Jestadi et al. 2014; Kniewald et al. 2000). In females, pesticides imbalances sexual hormones intervene androgen or estrogens receptors for improper function, irregularities of ovarian cycles, instinctive abortion, defect in developmental births, etc. (Bohn et al. 2011). Atrazine forms ROS, which cause single- and double-strand breaks in DNA and thus is genotoxic (Yang et al. 2010). The working of cardiovascular system also gets affected by atrazine exposure (Lin et al. 2016a, b, c; Cosselman et al. 2015). Atrazine is known to cause hepatic damage, as liver is the primary organ for atrazine metabolism in mammals (Campos-Pereira et al. 2012; Gojmerac and Kniewald 1989). Atrazine is also responsible for cardiotoxicity and hepatotoxicity in mice by disruption of ionic balance (Lin et al. 2016a, b).
The negative effects of atrazine on soil and aquatic biota are enormous. The above studies decipher a clear picture about the vulnerability of atrazine to terrestrial and aquatic life forms. The detection of atrazine with better accuracy and reliability is thus essential for safety of human health, biota and environment.
Analytical methods for monitoring atrazine in ecosystems
Basically, single environmental matrix can contained multiple pesticides (Kaur et al. 2017; Kumar et al. 2016, 2017; Singh et al. 2016). The extraction, cleanup and pre-treatment procedure depend upon the physiochemical nature of pesticides and environmental matrix (Kaur et al. 2017; Kumar et al. 2016, 2017). There are number of methods described for the analysis of different classes of pesticides in different matrices (AOAC 1993; Kumar et al. 2015a, b, c, d; Prasad et al. 2013; Kumar and Singh 2016; Singh et al. 2017). In day-to-day laboratory analysis, AOAC method is most authentic. Most common steps for the extractions of pesticides including atrazine have been described by Kumar et al. (2015a, b, c, d) as per the guidelines of AOAC (AOAC 1993; Kumar et al. 2015a, b, c, d; Prasad et al. 2013; Singh et al. 2016, 2017).
Highly sensitive and rapid analytical methods are essential for monitoring of residual atrazine and its metabolites in soil and water bodies (Table 3). It is monitored by using spectroscopic, immunogenic and chromatographic methods which include infrared (IR) spectroscopy, high-performance liquid chromatography (HPLC)/HPLC–MS/HPLC–MS/MS, HPLC, enzyme-linked immunosorbent assay (ELISA) and gas chromatography GC/GC–MS/GC–MS/MS (Miensah et al. 2015; Williams et al. 2014; Bonansea et al. 2013). Recently nano-based solid-based extractions ultra-performance liquid chromatography (UPLC) and GC/LC–MS methods have been developed with very good recovery and detection limits up to ppb (Table 3). The detection of atrazine in biological media is quantified using GC coupled with MS including different detectors like flame ionization detector (Haiyan and Center 2015), electron capture detector (ECD) (Miensah et al. 2015), nitrogen-phosphorus detector (NPD) (Bonansea et al. 2013) and GC coupled with mass spectrometer (Williams et al. 2014).
Quantification of atrazine was also done by a diode array (Yang et al. 2014) and a UV detector (Kong et al. 2016). Immunogenic methods are usually based on ELISA using sheep-based antibodies to atrazine (Na et al. 2012). Other immunogenic methods have been developed in which the antibody is bound to a “dipstick,” and this is used to evaluate concentrations of atrazine in water or liquid food samples (Kaur et al. 2007), while other sampling approaches have used immuno-affinity systems to concentrate atrazine prior to analysis by GC (Tran et al. 2013).
In water samples, atrazine was detected with GC equipped with nitrogen phosphate detector (NPD) and the percentage recovery is very high 96–98% (Nsibande et al. 2015). In sub-surface waters, the atrazine was quantified by both the techniques, i.e., HPLC and GC equipped with NPD, ECD and MS detectors (Barchanska et al. 2014). In aquatic plants and sediments, the concentration of atrazine was quantified by GC coupled with ECD detectors and the recovery percentage is near about 90% (Bennett et al. 2000). In water and soil samples, the solid-phase extraction method is used to concentrate the samples prior for analysis, all the three methods, i.e., GC coupled with all three detectors mentioned above, ELISA and HPLC equipped with UV detector (Hernandez et al. 2000; Lioy et al. 2000). The detection limit of atrazine in biological media moves to a greater extent as it was 0.1 µg/L in the year 2000 (Yokley et al. 2000) and it moves to 50 ng/L in the year 2008 (Gervais et al. 2008). It is all due to the new innovations in day-to-day life. In 2000, the detection was only SPE and GC based. Using styrene di-vinyl benzene sorbents, the detection limits increase to 10 ng/L (Bruzzoniti et al. 2006). Tandem mass spectroscopy combined with ultra-performance liquid chromatography detects up to 50 ng/L of the atrazine in biological media (Kuklenyik et al. 2012).
Qie et al. (2013) developed a technique direct competitive ELISA method (dcTELISA) based on thermistor enzyme for faster detection of atrazine in large-scale samples. In this method, ATZ competes for β-lactamase-labeled ATZ (ATZ–E) for the binding sites on anti-ATZ monoclonal immune response (mAb) which is covalently linked to form immunocomplexes from immune reactor with β-lactamase-labeled ATZ and atrazine. The detection limit was very good with high recovery rate (88–107%) in silage and fresh corn stalk samples. Another novel method for detection of atrazine was based on the SPR (Sepia pterin reductase) determination of P450 mRNA levels in Saccharomyces cerevisiae. The selected oligonucleotide probe that exhibits specificity for P450 mRNA was successfully immobilized on the sensor chip. The mRNA was quantified. It is a highly sensitive and rapid method that permits the detection of atrazine within 15 min (Kim et al. 2003). Kaoutit et al. (2004) also gave a simple conducting polymer-based biosensor approach. A glassy carbon electrode was prepared which is operated at open circuit and served for the immobilization of the enzyme polyphenol oxidase (PPO) during the anodic electropolymerization of polypyrrole (PPy). The concentration of atrazine in aqueous solution is attributed to its inhibitory power toward the catalytic activity of PPO. This biosensor helps in easy detection of photosynthetic inhibiting herbicide monitoring because of its analytical performance and simplicity. A protein-based conjugate method was developed for binding of atrazine with anti-atrazine monoclonal antibodies. Here immobilization was done on gold particles based on test strip method assessed with a photometric device to detect atrazine in very low limits (130 ng/mL) (Byzova et al. 2010). An immunochromatography (ICG) strip test for detection of atrazine in water samples was also developed. Monoclonal antibody (MAb) specific to atrazine was produced from the cloned hybridoma cell (AT-1-M3) and used to develop a direct competitive enzyme-linked immunosorbent assay (DCELISA) and ICG strip. The limit of detection was 3 ng/mL, and it requires only 10 min getting the results and that too in a single step. It came out to be a sensitive and accurate technique (Shim et al. 2006). A disposable immunomagnetic electrochemical sensor involving magnetic particles was developed for the detection of atrazine. The sensor was based on a magnetic monolayer of magnetic particles coated with streptavidin, formed on a gold electrode after application of a magnetic field. The atrazine interacts with biotinyl–Fab fragment K47, and the immune reactions were characterized by impedance spectroscopy. A decrease in electron transfer resistance was observed which could be attributed to rearrangements in the magnetic monolayer. The limit of detection is in the range 10–600 ng/ml; it is a sensitive approach for detection of atrazine acting as an antigen (Helali et al. 2006). Another novel immunoassay was formulated that involved direct coating of haptens on microtiter plates for detection of atrazine. The assay allows hapten-coated plates and uses affinity-purified atrazine which showed very high sensitivity. The limit of detection for atrazine is in the range 0.02–0.7 ng/mL (Suri et al. 2008). Detection of atrazine and triazines in water has also been carried out flow injection chemiluminescence analysis. The aliphatic amines in traizines react with tris(2,2′-bipyridyl)ruthenium(III) to produce chemiluminescence. The presence of natural organic matter (NOM) significantly increased the chemiluminescence, masking the signal generated by atrazine. Isolating the target analyte via solid-phase extraction (SPE) prior to analysis removed this interference and concentrated the samples. The detection limit is 14 ± 2 ng/L (Beale et al. 2009). Pardieu et al. (2009) devised an electrochemical sensor based on molecularly imprinted conducting polymer (MICP). The recognition of atrazine can be quantified by the variation of the cyclic voltammogram of MICP. It shows selectivity toward triazine family and shows a large range of detection from 10−9 to 1.5 × 10−2 mol/L. Impedance spectroscopy transduction combined with the immunosensor technology has been used for the determination of atrazine. The immunoreaction of atrazine on the attached anti-atrazine antibody leads to an increase in the charge transfer resistance which is proportional to the concentration of atrazine. Its limit of detection was 10 pg/mL (Ionescu et al. 2010). Another approach based on antibody replicas for atrazine detection was formulated by Schirhagl et al. (2011). Antibodies were used to pattern the nanoparticles for surface imprinting the polymer layer to produce replicas. Liu et al. (2011) constructed a MIP (molecularly imprinted polymers) chemosensor from a core–shell nanostructure. Vinyl-substituted zinc(II) protoporphyrin (ZnPP) was used as both fluorescent reporter and functional monomer to synthesize atrazine-imprinted polymer shell. The limit for detection is 1.8 mM. Bioluminescent reported bacteria are also utilized for detection of atrazine. Increase in bioluminescent signals is recorded for E. coli. For better interaction between insoluble atrazine and bacterial cells, centrifugation of bacterial cells and analyte dilutions can be performed (Jia et al. 2012). An electrochemical immunosensor for atrazine detection was developed by immobilization of gold nanoparticles on the gold electrode surface. The increase in surface area of work electrode leads to more anti-atrazine monoclonal antibodies capture. Ferricyanide was used as an electrochemical redoxindicator; immunosensor was characterized by cyclic voltammetry and electrochemical impedance spectroscopy. The limit for detection is as low as 0.016 ng/mL (Liu et al. 2014). A field effect transistor was developed based on network of single-walled carbon nanotubes which constitute carbon nanotubes field effect transistors and act as conductor channel for the determination of atrazine in various biological samples with detection limit up to 0.001 ng/mL (Belkhamssa et al. 2016).
Photochemical degradation of atrazine
Several chemical methods have been developed for the degradation during time to time. Konstantinou et al. (2001) have studied the photocatalytic degradation of atrazine and other s-triazine herbicides by using particulate TiO2 as photocatalyst under simulated solar light. The degradation process is highly efficient with traces of atrazine (at ppb level) being decomposed in very short times to less than 0.1 ppb. The process has been shown to lead to the formation of 2,4,6-trihydroxy-1,3,5-triazine (cyanuric acid) as the final product of the degradation process for all the investigated herbicides with several intermediates, rather than to the complete mineralization often observed for other classes of substrates. Monitoring of degradation products has been done using liquid, gas and ion chromatography, and the overall degradation process has been monitored through dissolved and particulate organic carbon measurements.
Atrazine degradation by Fenton’s reagent has been determined as a function of reagent’s concentration and ratios and pH in batch treatments (Barbusiński and Filipek 2001). The actual number and nature of oxidation products have been shown to vary with the concentration. The optimum Fenton’s reagent treatment has been achieved with 1:l molar ratios of FeSO4 and H2O2 (2.69 mM), producing two main terminal products viz, 2-chloro-4,6-diamino-s-triazine (23%) and 2-acetamido-4-amino-6-chloro-s-triazine (28%). Chloride release of 55 ± 9% indicated that dehalogenated s-triazines accounted for the balance of 14C. Atrazine degradation has been found to be pH dependent and decreases from 99% at pH 3 to 37% at pH 9.
In a modified approach, the effects of an inorganic ligand tetrapolyphosphate on the molecular oxygen activation and the subsequent aerobic atrazine degradation by Fe@Fe2O3 core–shell nanowires were investigated at a pH range of 6.0–9.0 (Wang et al. 2014a, b). It was observed that the addition of tetrapolyphosphate enhanced the rate of aerobic atrazine degradation dramatically (955 times) which was even 10 times that of the traditional organic ligand ethylenediamine tetraacetate. The rate enhancement has been attributed to enhanced reduction in Fe(III) to Fe(II) and the subsequent activation of molecular oxygen, owing to the suppressed hydrogen evolution, in the presence of tetrapolyphosphate, from the reduction in proton by Fe@Fe2O3 core–shell nanowires, making more electrons available for the reduction in Fe(III). Moreover, the complexation of tetrapolyphosphate with ferrous ions ensures enough soluble Fe(II) for Fenton reaction and also provides another route to produce more hydroxyl (·OH) radicals in the solution via the single-electron molecular oxygen reduction pathway, thus increasing the rate dramatically.
Ozone being a powerful oxidant has been used in the presence of hydrogen peroxide for degradation of atrazine. Tandem solid-phase extraction procedure was used which includes a C18 reverse-phase support and a strong cation exchanger (Nélieu et al. 2000). It was found that ammeline (2, 4-diamino-6-hydroxy-s-triazine) is the major end product (20% at pH 8) and 2-chloro-4,6-diamino-s-triazine as competitor whose ratio was dependent on the hydroxyl radical content. A number of new intermediates identified were aminoaldehydes and a carbinolamine (Nélieu et al. 2000). Triazine has also been degraded by electrochemical advanced oxidation processes such as anodic oxidation, electro-fenton and photoelectro-fenton using a small open and cylindrical cell with a boron-doped diamond anode. Anodic oxidation has been carried out either with a stainless steel cathode or an O2 diffusion cathode able to generate H2O2 formed at the boron-doped diamond surface in all electrochemical advanced oxidation processes. In the bulk form, Fenton’s reaction between added Fe2+ and electrogenerated H2O2 in electro-Fenton and photoelectron-Fenton, Hydroxyl radical (·OH) is the main oxidant (Borras et al. 2010). It has been observed that almost overall mineralization (94%) is achieved. Atrazine decay always follows a pseudo-first-order reaction, being more rapidly destroyed from ·OH in bulk than at the boron-doped diamond surface. The formation of dealkylated aromatic intermediates such as desethylatrazine and desethyldesisopropylatrazine and cyanuric acid has been revealed by reverse-phase HPLC, and short linear carboxylic acids such as formic, oxalic and oxamic have been identified and quantified by ion-exclusion HPLC. It has been observed that all initial nitrogen is transformed into NO3 − and NH4 + ions (followed by ionic chromatography) in electro-fenton and photoelectro-fenton but not in anodic oxidation, where 36% of nitrogen is released from the solution probably as volatile NO x species.
The recent results showed approximately 80% of atrazine was degraded by ozonation in the presence of hydroxylamine, while only 20% was degraded by ozonation alone. The atrazine degradation involved dealkylation, dechlorination–hydroxylation and olefination (Yang et al. 2016a, b). It has been found that at lower pH the degradation efficiency of atrazine was enhanced by UV/chlorine compared to UV or chlorine alone. The oxidation products of atrazine resulting from dealkylation, dechlorination–hydroxylation, alkylic-hydroxylation, alkylic-oxidation, alkylic-hydroxylation dehydration, deamination–hydroxylation and dechlorination–hydrogenation in UV/chlorine process were detected, which were slightly different from those formed in UV/H2O2 (Kong et al. 2016). Similar products were noticed when nitrite was added to enhance atrazine degradation during ozonation (Yang et al. 2016a, b). The experimental results of electrophotocatalytic reduction (hydroxyl radical reduction) of atrazine with an initial concentration of atrazine (100 mg/L) show that more than 99% of atrazine oxidation was obtained after 30 min of treatment and reaction kinetic constant was about 0.146/min. The analysis with liquid chromatography technique permits to identify, quantify and see the evolution of atrazine by-products which are generated by dechlorination, dealkylation and alkylic-oxidation mechanisms (Komtchou et al. 2016). The photodegradation study of atrazine was demonstrated using either Pt–TiO2 or TiO2 as a photocatalyst under 352 nm light irradiation. The Pt–TiO2-catalyzed atrazine degradation reached 76% in 3 h without adding H2O2 solution or aeration, which was more than 10% higher than the TiO2-catalyzed reaction. The decomposition product of Pt–TiO2-catalyzed atrazine degradation was mainly cyanuric acid (Chen et al. 2017). Shamsedini et al. (2017) noticed the maximum atrazine removal rate was at pH = 11 in the presence of Fe3+–TiO2 catalyst = 25 mg/L and the initial concentration of atrazine equal to 10 mg/L.
Microbial degradation of atrazine
Microorganisms are endowed with enormous and remarkable metabolic capabilities to utilize xenobiotics which are their carbon and energy source. A number of microbial species and strains have been shown to exhibit atrazine metabolism as listed in Table 4. Prokaryotic (Gram-positive and Gram-negative bacteria) and eukaryotic microbial species are involved in atrazine biodegradation both in situ and under in vitro conditions. Bacterial and fungal species usually dechlorinate the atrazine molecule leading to the formation of hyroxyatrazine, deisopropylatrazine and deethylatrazine. Xanthomonas sp. ARB2 (Sawangjit 2016); Enterobacter cloacae JS08 (Solomon et al. 2013a, b); Klebsiella sp. KB02 (Sopid 2012); Comamonas sp. A2 (Yang et al. 2010); Stenotrophomonas maltophilia, Rahnella aquatilis (Marecik et al. 2008); Chelatobacter heintzii, Aminobacter aminovorans, Stenotrophomonas maltophilia (Rousseaux et al. 2001); Pseudaminobacter C147 (Topp et al. 2000a); Bacillus subtilis HB-6 (Wang et al. 2014a, b); Arthrobacter sp. (Getenga et al. 2009; Liu et al. 2010; El Sebai et al. 2011; Zhang et al. 2011; Wang and Xie 2012; Wang et al. 2013); Bacillus licheniformis, B. megaterium (Marecik et al. 2008); Arthrobacter nicotinovorans HIM (Aislabie et al. 2005); Nocardioides sp. SP12 (Piutti et al. 2003); Arthrobacter aurescens TC1 (Strong et al. 2002); Arthrobacter crystallopoietes (Rousseaux et al. 2001); and Nocardioides sp. (Topp et al. 2000b). Some fungal species, viz. Umbelopsis isabellina, Volutella ciliate and Botrytis cinerea, were also found to be involved in degradation of atrazine (Marecik et al. 2008). Microbial species perform atrazine biodegradation by three major pathways out of which one is purely hydrolytic, while remaining two others are mixed (hydrolytic–oxidative) (Fig. 2). The first intermediate product hydroxyatrazine was first extensively converted by Pseudomonas spp. ADP consisting of three gene products atzA, atzB and atzC (Martinez et al. 2001). The dechlorination method (hydrolytic) which is catalyzed by enzyme atrazine chlorohydrolase (atzA or trzN gene product) shows dechlorination followed by elimination of N alkyl substituents to yield cyanuric acid (Solomon et al. 2013a, b). These three genes are widespread and almost found in all the atrazine-degrading strains worldwide (Rousseaux et al. 2001; Topp et al. 2000a). Usually in Gram-positive strains, atzA is replaced by trzN which belongs to hydrolase enzyme which removes several functional groups from the parent compound (Wang et al. 2005; Topp et al. 2000b). The second pathway engrosses the N-dealkylation of the atrazine into deethylatrazine or deisopropylatrazine which is further dealkylated into deisopropyldeethylatrazine or further undergoes hydrolytic to yield cyanuric acid. Rhodococcus strains N186 and 21 and SpTE1 show oxidative reactions by producing enzymes AtzA and TriA which actively deaminates the atrazine metabolites (Seffernick et al. 2001). Dechlorination of dealkylated atraizine is commonly shown by Rhodococcus corallinus NRRLB-containing hydrolase AtzB (Seffernick et al. 2002). Further, in the upper degradation pathway it gets converted into two different aminohydrolases which is encoded by atzB and atzC. Then, the final hydrolytic reaction which is encoded by trzF/atzF, trzD/atzD and trzE/atzE converts cyanuric acid into carbon dioxide (Udiković-Kolić et al. 2010; Wackett et al. 2002).
Physical and biochemical pathways involved in the microbial biodegradation of atrazine. AC atrazine chlorohydrolase, AM atrazine monooxygenase, HDEH hydroxyl dechloroatrazine ethylaminohydrolase, NEC N-ethylammeline chlorohydrolase, DEM deethylatrazine monooxygenase, NIAIA N-isopropyl ammelide Isopropyl aminohydrolase, DIHA deisopropylhydroxyatrazine aminohydrolase, NEAC N-ethylammeline chlorohydrolase, 2, 4 D6 NEA 1, 3, 5 TEH = 2, 4-dihydroxy-6-(N′-ethyl) amino-1, 3, 5-triazine ethylaminohydrolase, HDAEH hydroxydechloro atrazine ethylaminohydrolase, NIIA N-isopropylammelide isopropylaminohydrolase
Nitrogen released from atrazine metabolism serves as a nitrogen source for atrazine-degrading bacteria (Vaishampayan et al. 2007; Dutta and Singh 2013; Yang et al. 2010). Some bacteria initiate degradation of atrazine involving the enzyme atrazine chlorohydrolase through the mechanism of hydrolytic dechlorination. Aminohydrolases catalyze two hydrolytic deamination reactions that hydroxyatrazine undergoes; N-isopropylammelide (Getenga et al. 2009; Qingyan et al. 2008) or N-ethylammelide (Topp et al. 2000a) is formed as the intermediate metabolites. These ammelides are finally converted to cyanuric acid (Yang et al. 2010). Another route followed for atrazine degradation is N-dealkylation of the lateral ethyl and isopropyl chains to deethylatrazine, deisopropylatrazine and deethyldeisopropylatrazine (Zhang et al. 2011). These dealkylated atrazine metabolites undergo hydroxylation, and cyanuric acid is formed as the ultimate metabolite (Vaishampayan et al. 2007). Cyanuric acid, formed by either of the metabolic routes, is acted upon by cyanuric acid amidohydrolase, biuret amidohydrolase and allophanate hydrolase enzymes leading to the cleavage of the cyanuric acid to carbon dioxide and ammonia (El Sebai et al. 2011).
Conclusion
The demand for pesticides is on rise globally, especially in emerging economies of the world. India is the second most populated country of the world and is at the center of green revolution. It is, however, under ever-increasing demand of fulfilling the food requirements of huge population and thus relying heavily on synthetic herbicides as a weed control measure. As a consequence, environmental pollution, contamination of reservoirs, effect on food chains and life-threatening toxicities are certain to happen. Atrazine and its adverse effects are considered a highlighted threat to the environmental sustainability. Hence, an urgent need is felt to diverted resources and coordinated efforts to minimize its use, and at the same time, it is essential to monitor its impact on vertebrates, invertebrates and, most importantly, on microbial flora. Microbial biodegradation using in situ approach and use of transgenic strains having enhanced enzymatic activities and superior adaptability is considered a valid option for future studies. In future, use of biopesticides is expected to relieve our dependency on atrazine in order to minimize environmental pollution.
References
Abarikwu SO, Farombi EO (2015) Atrazine induces apoptosis of SH-SY5Y human neuroblastoma cells via the regulation of Bax/Bcl-2 ratio and caspase-3-dependent pathway. Pest Biochem Physiol 118:90–98. doi:10.1016/j.pestbp.2014.12.006
Ahel M, Evans KM, Fileman TW, Mantoura RFC (1992) Determination of atrazine and simazine in estuarine samples by high-resolution gas chromatography and nitrogen selective detection. Anal Chim Acta 268(2):195–204. doi:10.1016/0003-2670(92)85213-P
Ahrens JF, Newton M (2008) Benefits of triazine herbicides in the production of ornamentals and conifer trees. The Triazine Herbicides (Chapter 18), p 225234. ISBN: 978-0-444-51167-6
Aislabie J, Bej AK, Ryburn J, Lloyd N, Wilkins A (2005) Characterization of Arthrobacter nicotinovorans HIM an atrazine-degrading bacterium from agricultural soil New Zealand. FEMS Microbiol Ecol 52(2):279–286. doi:10.1016/j.femsec.2004
Akbulut GB, Yigit E (2010) The changes in some biochemical parameters in Zea mays cv. ‘‘Martha F1’’ treated with atrazine. Ecotoxicol Environ Saf 73:1429–1432. doi:10.1016/j.ecoenv.2010.05.023
Albanis TA, Hela DG, Sakellarides TM, Konstantinou IK (1998) Monitoring of pesticide residues and their metabolites in surface and underground waters of Imathia (N. Greece) by means of solid-phase extraction disks and gas chromatography. J Chromatogr A 823(1):59–71. doi:10.1016/S0021-9673(98)00304-5
Amistadi MK, Hall JK, Bogus ER, Mumma RO (1997) Comparison of gas chromatography and immunoassay methods for the detection of atrazine in water and soil. J Environ Sci Health B 32(6):845–860. doi:10.1080/03601239709373116
Andrus JM, Winter D, Scanlan M, Sullivan S, Bollman W, Waggoner JB, Brain RA (2013) Seasonal synchronicity of algal assemblages in three Midwestern agricultural streams having varying concentrations of atrazine, nutrients, and sediment. Sci Total Environ 458:125–139. doi:10.1016/j.scitotenv.2013.03.070
AOAC (Association of Official Analytical Chemists) (1993) Peer verified methods program. AOAC, manual on policies and procedures. AOAC, Arlington
Bai X, Sun C, Xie J, Song H, Zhu Q, Su Y, Fu Z (2015) Effects of atrazine on photosynthesis and defense response and the underlying mechanisms in Phaeodactylum tricornutum. Environ Sci Pollut Res 22(22):17499–17507. doi:10.1007/s11356-015-4923-7
Barbusiński K, Filipek K (2001) Use of Fenton’s reagent for removal of pesticides from industrial wastewater. Pol J Environ Stud 10(4):207–212
Barchanska H, Babilas B, Gluzicka K, Zralek D, Baranowska I (2014) Rapid determination of mesotrione, atrazine and its main degradation products in selected plants by MSPD–HPLC and indirect estimation of herbicides phytotoxicity by chlorophyll quantification. Int J Environ Anal Chem 94(2):99–114. doi:10.1080/03067319.2013.791979
Batra M, Pandey J, Suri CR, Jain RK (2009) Isolation and characterization of an atrazine-degrading Rhodococcus sp. strain MB-P1 from contaminated soil. Lett Appl Microbiol 49(6):721–729. doi:10.1111/j.1472-765X.2009.02724.x
Baxter L, Brain RA, Hosmer AJ, Nema M, Müller KM, Solomon KR, Hanson ML (2015) Effects of atrazine on egg masses of the yellow-spotted salamander (Ambystoma maculatum) and its endosymbiotic alga (Oophila amblystomatis). Environ Pollut 206:324–331. doi:10.1016/j.envpol.2015.07.017
Baxter L, Brain RA, Lissemore L, Solomon KR, Hanson ML, Prosser RS (2016) Influence of light, nutrients, and temperature on the toxicity of atrazine to the algal species Raphidocelis subcapitata: implications for the risk assessment of herbicides. Ecotoxicol Environ Saf 132:250–259. doi:10.1016/j.ecoenv.2016.06.022
Beale DJ, Porter NA, Roddick FA (2009) A fast screening method for the presence of atrazine and other triazines in water using flow injection with chemiluminescent detection. Talanta 78(2):342–347. doi:10.1016/j.talanta.2008.11.033
Behki RM, Khan SU (1986) Degradation of atrazine by Pseudomonas: N-dealkylation and dehalogenation of atrazine and its metabolites. J Agric Food Chem 34(4):746–749. doi:10.1021/jf00070a039
Behki RM, Khan SU (1994) Degradation of atrazine, propazine, and simazine by Rhodococcus strain B-30. J Agric Food Chem 42(5):1237–1241. doi:10.1021/jf00041a036
Behki R, Topp E, Dick W, Germon P (1993) Metabolism of the herbicide atrazine by Rhodococcus strains. Appl Environ Microbiol 59(6):1955–1959
Beilstein P, Cook AM, Hütter R (1981) Determination of seventeen s-triazine herbicides and derivatives by high-pressure liquid chromatography. J Agric Food Chem 29(6):1132–1135. doi:10.1021/jf00108a008
Belden JB, Lydy MJ (2001) Effects of atrazine on acetylcholinesterase activity in midges (Chironomus tentans) exposed to organophosphorus insecticides. Chemosphere 44(8):1685–1689. doi:10.1016/S0045-6535(00)00519-1
Belkhamssa N, Justino CI, Santos PS, Cardoso S, Lopes I, Duarte AC, Ksibi M (2016) Label-free disposable immunosensor for detection of atrazine. Talanta 146:430–434. doi:10.1016/j.talanta.2015.09.015
Bennett ER, Moore MT, Cooper CM, Smith S (2000) Method for the simultaneous extraction and analysis of two current use pesticides atrazine and lambda-cyhalothrin in sediment and aquatic plants. Bull Environ Contam Toxicol 64(6):825–833. doi:10.1007/s001280000077
Bethsass J, Colangelo A (2013) European Union bans atrazine while the United States negotiates continued use. Int J Occup Environ Health 12(3):260–267. doi:10.1179/oeh.2006.12.3.260
Bhushan C, Bhardwaj A, Misra SS (2013) State of pesticide regulations in India. Centre for Science and Environment, New Delhi, pp 1–72
Blahova J, Plhalova L, Hostovsky M, Divisova L, Dobsikova R, Mikulikova I, Svobodova Z (2013) Oxidative stress responses in zebrafish Danio rerio after subchronic exposure to atrazine. Food Chem Toxicol 61:82–85. doi:10.1016/j.fct.2013.02.041
Bohn T, Cocco E, Gourdol L, Guignard C, Hoffmann L (2011) Determination of atrazine and degradation products in Luxembourgish drinking water: origin and fate of potential endocrine-disrupting pesticides. Food Addit Contam Part A 28(8):1041–1054. doi:10.1080/19440049.2011.580012
Bonansea RI, Amé MV, Wunderlin DA (2013) Determination of priority pesticides in water samples combining SPE and SPME coupled to GC–MS. A case study: Suquía River basin (Argentina). Chemosphere 90(6):1860–1869. doi:10.1016/j.chemosphere.2012.10.007
Borras N, Oliver R, Arias C, Brillas E (2010) Degradation of atrazine by electrochemical advanced oxidation processes using a boron-doped diamond anode. J Phys Chem A 114(24):6613–6621. doi:10.1021/jp1035647
Bouquard C, Ouazzani J, Prome J, Michel-Briand Y, Plesiat P (1997) Dechlorination of atrazine by a Rhizobium sp. Isolate. Appl Environ Microbiol 63(3):862–866
Bringolf RB, Cope WG, Barnhart MC, Mosher S, Lazaro PR, Shea D (2007) Acute and chronic toxicity of pesticide formulations (atrazine chlorpyrifos and permethrin) to glochidia and juveniles of Lampsilis siliquoidea. Environ Toxicol Chem 26(10):2101–2107. doi:10.1897/06-555R.1
Bruzzoniti MC, Sarzanini C, Costantino G, Fungi M (2006) Determination of herbicides by solid phase extraction gas chromatography–mass spectrometry in drinking waters. Anal Chim Acta 578(2):241–249. doi:10.1590/S0103-50532009000500017
Brzezicki JM, Andersen ME, Cranmer BK, Tessari JD (2003) Quantitative identification of atrazine and its chlorinated metabolites in plasma. J Anal Toxicol 27(8):569–573. doi:10.1093/jat/27.8.569
Burken JG, Schnoor JL (1997) Uptake and metabolism of atrazine by poplar trees. Environ Sci Technol 31(5):1399–1406. doi:10.1021/es960629v
Byzova NA, Zherdev AV, Zvereva EA, Dzantiev BB (2010) Immunochromatographic assay with photometric detection for rapid determination of the herbicide atrazine and other triazines in foodstuffs. J AOAC Int 93(1):36–43
Cai B, Han Y, Liu B, Ren Y, Jiang S (2003) Isolation and characterization of an atrazine-degrading bacterium from industrial wastewater in China. Lett Appl Microbiol 36(5):272–276. doi:10.1046/j.1472-765X.2003.01307.x
Campos-Pereira FD, Oliveira CA, Pigoso AA, Silva-Zacarin EC, Barbieri R, Spatti EF, Marin-Morales MA, Severi-Aguiar GD (2012) Early cytotoxic and genotoxic effects of atrazine on Wistar rat liver: a morphological, immunohistochemical, biochemical, and molecular study. Ecotoxicol Environ Saf 78:170–177. doi:10.1016/j.ecoenv.2011.11.020
Caoa W, Yanga B, Qia F, Qiana L, Lib J, Luc L, Qian X (2017) Simple and sensitive determination of atrazine and its toxic metabolites in environmental water by carboxyl modified polyacrylonitrile nanofibers mat-based solid-phase extraction coupled with liquid chromatography-diode array detection. J Chromatogr A 1491:16–26. doi:10.1016/j.chroma.2017.02.035
Caux PY, Ménard L, Kent RA (1996) Comparative study of the effects of MCPA, butylate, atrazine, and cyanazine on Selenastrum capricornutum. Environ Pollut 92(2):219–225. doi:10.1016/0269-7491(95)00060-7
Cerejeira MJ, Viana P, Batista S, Pereira T, Silva E, Valério MJ, Silva-Fernandes AM (2003) Pesticides in Portuguese surface and ground waters. Water Res 37(5):1055–1063. doi:10.1016/S0043-1354(01)00462-6
Chen D, Zhang Y, Miao H, Zhao Y, Wu Y (2015a) Determination of triazine herbicides in drinking water by dispersive micro solid phase extraction with ultrahigh-performance liquid chromatography–high-resolution mass spectrometric detection. J Agric Food Chem 63:9855–9862. doi:10.1021/acs.jafc.5b03973
Chen J, Huo J, Jia Z, Song Y, Li Y, Zhang L (2015b) Effects of atrazine on the proliferation and cytotoxicity of murine lymphocytes with the use of carboxyfluorescein succinimidyl ester-based flow cytometric approaches. Food Chem Toxicol 76:61–69. doi:10.1016/j.fct.2014.11.026
Chen SM, Lu N, Chen JY, Yang CY, Yeh YP, Feng TY, Shih Y, Kokulnathan T, Chen D (2017) Enhanced photocatalytic degradation of atrazine by platinized titanium dioxide under 352 nm irradiation. Water Sci Technol 75(5):1128–1137. doi:10.2166/wst.2016.593
Cheng G, Shapir N, Sadowsky MJ, Wackett LP (2005) Allophanate hydrolase, not urease, functions in bacterial cyanuric acid metabolism. Appl Environ Microbiol 71(8):4437–4445. doi:10.1128/AEM.71.8.4437-4445
Cheng M, Zeng G, Huang D, Lai C, Xu P, Zhang C, Zhu Y (2016) Degradation of atrazine by a novel Fenton-like process and assessment the influence on the treated soil. J Hazard Mater 312:184–191. doi:10.1016/j.jhazmat.2016.03.033
Christin MS, Gendron AD, Brousseau P, Ménard L, Marcogliese DJ, Cyr D, Fournier M (2003) Effects of agricultural pesticides on the immune system of Rana pipiens and on its resistance to parasitic infection. Environ Toxicol Chem 22(5):1127–1133. doi:10.1002/etc.5620220522
Cook AM, Hutter R (1984) Deethylsimazine: bacterial dechlorination, deamination, and complete degradation. J Agric Food Chem 32(3):581–585. doi:10.1021/jf00123a040
Cooper RL, Stoker TE, Tyrey L, Goldman JM, McElroy WK (2000) Atrazine disrupts the hypothalamic control of pituitary-ovarian function. Toxicol Sci 53(2):297–307. doi:10.1093/toxsci/53.2.297
Cooper RL, Laws SC, Das PC, Narotsky MG, Goldman JM, Lee Tyrey E, Stoker TE (2007) Atrazine and reproductive function: mode and mechanism of action studies. Birth Defects Res B Dev Reprod Toxicol 80(2):98–112. doi:10.1002/bdrb.20110
Correia FV, Macrae A, Guilherme LRG, Langenbach T (2007) Atrazine sorption and fate in a Ultisol from humid tropical Brazil. Chemosphere 67(5):847–854. doi:10.1016/j.chemosphere.2006.11.03
Cosselman KE, Navas-Acien A, Kaufman JD (2015) Environmental factors in cardiovascular disease. Nat Rev Cardiol 12(11):627–642. doi:10.1038/nrcardio.2015.152
Cox C (2001) Atrazine environmental contamination and ecological effects. J Pestic Reform 21(3):1–9. doi:10.1016/j.chemosphere.2006.11.034
Dalluge J, Hankemeier T, Vreuls RJ, Udo A (1999) On-line coupling of immunoaffinity-based solid-phase extraction and gas chromatography for the determination of s-triazines in aqueous samples. J Chromatogr A 830(2):377–386. doi:10.1016/S0021-9673(98)00932-7
De Campos Ventura B, de Angelis DDF, Marin-Morales MA (2008) Mutagenic and genotoxic effects of the Atrazine herbicide in Oreochromis niloticus (Perciformes, Cichlidae) detected by the micronuclei test and the comet assay. Pestic Biochem Physiol 90(1):42–51. doi:10.1016/j.pestbp.2007.07.009
De Souza ML, Seffernick J, Martinez B, Sadowsky MJ, Wackett LP (1998) The atrazine catabolism genes atzABC are widespread and highly conserved. J Bacteriol 180(7):1951–1954
Denovan LA, Lu C, Hines CJ, Fenske RA (2000) Saliva biomonitoring of atrazine exposure among herbicide applicators. Int Arch Occup Environ Health 73(7):457–462. doi:10.1007/s004200000174
Dornelles MF, Oliveira GT (2014) Effect of atrazine, glyphosate and quinclorac on biochemical parameters, lipid peroxidation and survival in bullfrog tadpoles (Lithobates catesbeianus). Arch Environ Contam Toxicol 66(3):415–429. doi:10.1007/s00244-013-9967-4
Dos-Santos KC, Martinez CB (2014) Genotoxic and biochemical effects of atrazine and Roundup®, alone and in combination, on the Asian clam Corbicula fluminea. Ecotoxicol Environ Saf 100:7–14. doi:10.1016/j.ecoenv.2013.11.014
Dutta A, Singh N (2013) Degradation of atrazine in mineral salts medium and soil using enrichment culture. J Environ Sci Health Part B 48(10):860–868. doi:10.1080/03601234.2013.795845
Dutta A, Vasudevan V, Nain L, Singh N (2016) Characterization of bacterial diversity in an atrazine degrading enrichment culture and degradation of atrazine, cyanuric acid and biuret in industrial wastewater. J Environ Sci Health B 51(1):24–34. doi:10.1080/03601234.2015.1080487
Eaton RW, Karns JS (1991) Cloning and analysis of s-triazine catabolic genes from Pseudomonas sp. strain NRRLB-12227. J Bacteriol 173(3):1215–1222
El Sebai T, Devers-Lamrani M, Changey F, Rouard N, Martin-Laurent F (2011) Evidence of atrazine mineralization in a soil from the Nile Delta: isolation of Arthrobacter sp. TES6, an atrazine-degrading strain. Int Biodeterior Biodegrad 65(8):1249–1255. doi:10.1016/j.ibiod.2011.05.011
Fan W, Yanase T, Morinaga H, Gondo S, Okabe T, Nomura M, Nawata H (2007) Atrazine-induced aromatase expression is SF-1 dependent: implications for endocrine disruption in wildlife and reproductive cancers in humans. Environ Health Perspect 115(5):720–727. doi:10.1289/ehp.9758
Ferrari R, Nilsson T, Arena R, Arlati P, Bartolucci G, Basla R, Fungi M (1998) Inter-laboratory validation of solid-phase microextraction for the determination of triazine herbicides and their degradation products at ng/l level in water samples. J Chromatogr A 795(2):371–376. doi:10.1016/S0021-9673(97)00837-6
Francisco F (2001) Physiological mechanism of herbicide action. In: Pessarakli M (ed) Handbook of plant and crop physiology. Marcel Dekker Inc., New York, pp 773–783. doi:10.1201/9780203908426.fmatt
Gao Y, Fang J, Zhang J, Ren L, Mao Y, Li B, Zhang M, Liu D, Du M (2011) The impact of the herbicide atrazine on growth and photosynthesis of seagrass, Zostera marina (L.), seedlings. Mar Pollut Bull 62(8):1628–1631. doi:10.1016/j.marpolbul.2011.06.014
Gao S, Wang Z, Zhang C, Jia L, Zhang Y (2016) Oral exposure to atrazine induces oxidative stress and calcium homeostasis disruption in spleen of mice. Oxid Med Cellular Longevity, Article ID 7978219. doi:10.1155/2016/7978219
Gely-Pernot A, Hao C, Becker E, Stuparevic I, Kervarrec C, Chalmel F, Primig M, Jégou B, Smagulova F (2015) The epigenetic processes of meiosis in male mice are broadly affected by the widely used herbicide atrazine. BMC Genom 16(1):885. doi:10.6084/m9.figshare.c.3623711
Gervais G, Brosillon S, Laplanche A, Helen C (2008) Ultra-pressure liquid chromatography–electrospray tandem mass spectrometry for multi-residue determination of pesticides in water. J Chromatogr A 1202(2):163–172. doi:10.1016/j.foodchem.2010.10.017
Getenga Z, Dorfler U, Iwobi A, Schmid M, Schroll R (2009) Atrazine and terbuthylazine mineralization by an Arthrobacter sp, isolated from a sugarcane-cultivated soil in Kenya. Chemosphere 77(4):534–539. doi:10.1016/j,chroma,2008,07,006
Giardina MC, Giardi MT, Buffone R (1979) Soil enrichment studies with atrazine long term atrazine effects on degradation and microbiological composition. Chemosphere 8(11–12):831–834. doi:10.1016/0045-6535(79)90013-4
Giardina MC, Giardi MT, Filacchioni G (1980) 4-Amino-2-chloro-1, 3, 5-triazine: a new metabolite of atrazine by a soil bacterium. Agric Biol Chem 44(9):2067–2072. doi:10.1080/00021369.1980.10864288
Giardina MC, Giardi MT, Filacchioni G (1982) Atrazine metabolism by Nocardia: elucidation of initial pathway and synthesis of potential metabolites. Agric Biol Chem 46(6):1439–1445. doi:10.1080/00021369.1982.10865301
Giardina MC, Giardi MT, Filacchioni G (1985) Chemical and biological degradation of primary metabolites of atrazine bv a Nocardia strain. Agric Biol Chem 49(6):1551–1558. doi:10.1080/00021369.1985.10866949
Glæsner N, Bælum J, Strobel BW, Jacobsen CS (2014) Ageing of atrazine in manure amended soils assessed by bioavailability to Pseudomonas sp. strain ADP. Biodegradation 25(2):217–225. doi:10.1007/s10532-013-9654-1
Gojmerac T, Kniewald J (1989) Atrazine biodegradation in rats a model for mammalian metabolism. Bull Environ Contam Toxicol 43(2):199–206 (PMID: 2775886)
Gonzáleza NR, Gonzáleza EB, González-Castroa MJ, Mlpendurada MF (2016) On-line solid-phase extraction method for determination of triazine herbicides and degradation products in seawater by ultra-pressure liquid chromatography–tandem mass spectrometry. J Chromatogr A 1470:33–41. doi:10.1016/j.chroma.2016.10.007
González-Techera A, Zon MA, Molina PG, Fernández H, González-Sapienza G, Arévalo FJ (2015) Development of a highly sensitive noncompetitive electrochemical immunosensor for the detection of atrazine by phage anti-immunocomplex assay. Biosens Bioelectron 64:650–656. doi:10.1016/j.bios.2014.09.046
Graymore M, Stagnitti F, Allinson G (2001) Impacts of atrazine in aquatic ecosystems. Environ Int 26(7):483–495. doi:10.1016/S0160-4120(01)00031-9
Haiyan YUN, Center TEM (2015) Determination of organic phosphorous pesticides in water by LLE-GC-FPD. Shanxi Chem Ind 5:1–8
Hassan NM, Nemat AMM (2005) Oxidative stress in herbicide treated broad bean and maize plants. Acta Physiol Plant 27(4A):429–438. doi:10.1007/s11738-005-0047-x
Hayes TB, Collins A, Lee M, Mendoza M, Noriega N, Stuart AA, Vonk A (2002) Hermaphroditic demasculinized frogs after exposure to the herbicide atrazine at low ecologically relevant doses. Proc Natl Acad Sci 99(8):5476–5480. doi:10.1073/pnas.082121499
Hayes TB, Stuart AA, Mendoza M, Collins A, Noriega N, Vonk A, Johnston G, Liu R, Kpodzo D (2006) Characterization of atrazine-induced gonadal malformations in African clawed frogs (Xenopus laevis) and comparisons with effects of an androgen antagonist (cyproterone acetate) and exogenous estrogen (17beta-estradiol): support for the demasculinization/feminization hypothesis. Environ Health Perspect 114:134–141. doi:10.1289/ehp.8067
Helali S, Martelet C, Abdelghani A, Maaref MA, Jaffrezic-Renault N (2006) A disposable immunomagnetic electrochemical sensor based on functionalised magnetic beads on gold surface for the detection of atrazine. Electrochim Acta 51(24):5182–5186. doi:10.1016/j.electacta.2006.03.086
Hernandez F, Beltran J, Lopez FJ, Gaspar JV (2000) Use of solid-phase microextraction for the quantitative determination of herbicides in soil and water samples. Anal Chem 72(10):2313–2322. doi:10.1021/ac991115s
Howe GE, Gillis R, Mowbray RC (1998) Effect of chemical synergy and larval stage on the toxicity of atrazine and alachlor to amphibian larvae. Environ Toxicol Chem 17(3):519–525. doi:10.1002/etc.5620170324
Huang H, Zhang S, Wu N, Luo L, Christie P (2009) Influence of Glomus etunicatum/Zea mays mycorrhiza on atrazine degradation, soil phosphatase and dehydrogenase activities, and soil microbial community structure. Soil Biol Biochem 41(4):726–734. doi:10.1016/j.soilbio.2009.01.009
Ikonen R, Kangas J, Savolainen H (1988) Urinary atrazine metabolites as indicators for rat and human exposure to atrazine. Toxicol Lett 44(1–2):109–112. doi:10.1016/j.toxlet.2011.11.023
Inoue-Choi M, Weyer PJ, Jones RR, Booth BJ, Cantor KP, Robien K, Ward MH (2016) Atrazine in public water supplies and risk of ovarian cancer among postmenopausal women in the Iowa Women’s Health Study. Occup Environ Med 73(9):582. doi:10.1136/oemed-2016-103575
Ionescu RE, Gondran C, Bouffier L, Jaffrezic-Renault N, Martelet C, Cosnier S (2010) Label-free impedimetric immunosensor for sensitive detection of atrazine. Electrochim Acta 55(21):6228–6232. doi:10.1016/j.electacta.2009.11.029
Iriel A, Novo JM, Cordon GB, Lagorio MG (2014) Atrazine and methyl viologen effects on chlorophyll-a fluorescence revisited—implications in photosystems emission and ecotoxicity assessment. J Photochem Photobiol 90(1):107–112. doi:10.1111/php.12142
Jestadi DB, Phaniendra A, Babji U, Shanmuganathan B, Periyasamy L (2014) Effects of atrazine on reproductive health of nondiabetic and diabetic male rats. Int Scholarly Res Notices 676013:1–7. doi:10.1155/2014/676013
Jia K, Eltzov E, Toury T, Marks RS, Ionescu RE (2012) A lower limit of detection for atrazine was obtained using bioluminescent reporter bacteria via a lower incubation temperature. Ecotoxicol Environ Saf 84:221–226. doi:10.1016/j.ecoenv.2012.07.009
Jiang Z, Ma B, Erinle KO, Cao B, Liu X, Ye S, Zhang Y (2016) Enzymatic antioxidant defense in resistant plant: pennisetum americanum (L.) K. Schum during long-term atrazine exposure. Pestic Biochem Physiol 133:59–66. doi:10.1016/j.pestbp.2016.03.003
Jin Y, Zhang X, Shu L, Chen L, Sun L, Qian H, Fu Z (2010) Oxidative stress response and gene expression with atrazine exposure in adult female zebrafish (Danio rerio). Chemosphere 78(7):846–852. doi:10.1016/j.chemosphere.2009
Jin Y, Wang L, Chen G, Lin X, Miao W, Fu Z (2014) Exposure of mice to atrazine and its metabolite diaminochlorotriazine elicits oxidative stress and endocrine disruption. Environ Toxicol Pharmacol 37(2):782–790. doi:10.1016/j.etap.2014.02.014
Jowa L, Howd R (2011) Should atrazine and related chlorotriazines be considered carcinogenic for human health risk assessment. J Environ Sci Health C 29(2):91–144. doi:10.1080/10590501.2011.577681
Kabra AN, Ji MK, Choi J, Kim JR, Govindwar SP, Jeon BH (2014) Toxicity of atrazine and its bioaccumulation and biodegradation in a green microalga Chlamydomonas mexicana. Environ Sci Pollut Res 21(21):12270–12278. doi:10.1007/s11356-014-3157-4
Kadian N, Gupta A, Satya S, Mehta RK, Malik A (2008) Biodegradation of herbicide (atrazine) in contaminated soil using various bioprocessed materials. Bioresour Technol 99(11):4642–4647. doi:10.1016/j.biortech.2007.06.064
Kaoutit ME, Bouchta D, Zejli H, Izaoumen N, Temsamani KR (2004) A simple conducting polymer-based biosensor for the detection of atrazine. Anal Lett 37(8):1671–1681. doi:10.1081/AL-120037595
Karlsson AS, Weihermüller L, Tappe W, Mukherjee S, Spielvogel S (2016) Field scale boscalid residues and dissipation half-life estimation in a sandy soil. Chemosphere 145:163–173. doi:10.1016/j.chemosphere.2015.11.026
Karns JS, Eaton RW (1997) Genes encoding s-triazine degradation are plasmid-borne in Klebsiella pneumoniae strain 99. J Agric Food Chem 45(3):1017–1022. doi:10.1021/jf960464%2B
Kaur J, Singh KV, Boro R, Thampi KR, Raje M, Varshney GC, Suri CR (2007) Immunochromatographic dipstick assay format using gold nanoparticles labeled protein−hapten conjugate for the detection of atrazine. Environ Sci Technol 41(14):5028–5036. doi:10.1021/es070194j
Kaur S, Kumar V, Chawla M, Cavallo L, Poater A, Upadhyay A (2017) Pesticides curbing soil fertility: effect of complexation of free metal ions. Front Chem 5:1–9. doi:10.3389/fchem.2017.00043
Khan A, Shah N, Khan MS, Ahmad MS, Farooq M, Adnan M, Yousafzai AM (2016a) Quantitative determination of lethal concentration Lc 50 of atrazine on biochemical parameters; total protein and serum albumin of freshwater fish grass carp (Ctenopharyngodon idella). Pol J Environ Stud 25(4):1555–1561. doi:10.15244/pjoes/61849
Khan A, Yousafzai AM, Shah N, Ahmad MS, Farooq M, Aziz F, Adnan M, Rizwan M, Jawad SM (2016b) Enzymatic profile a activity of grass carp (Ctenopharyngodon idella) after exposure to the pollutant named atrazine (Herbicide). Pol J Environ Stud 25(5):2003–2008. doi:10.15244/pjoes/62821
Khilji S (2011) Toxicity testing with green alga Pseudokirchneriella subcapitata. https://ruor.uottawa.ca/bitstream/10393/20601/1/Khiji_Saadia_2011.pdf
Khoshnood Z, Jamili S, Khodabandeh S (2015) Histopathological effects of atrazine on gills of Caspian kutum Rutilus frisii kutum fingerlings. Dis Aquat Org 113(3):227–234. doi:10.3354/dao02850
Kim JY, Mulchandani A, Chen W (2003) An immunoassay for atrazine using tunable immunosorbent. Anal Biochem 322(2):251–256. doi:10.1016/j.ab.2003.08.009
Kniewald J, Jakominic M, Tomljenovic A, Simic B, Romac P, Vranesic D, Kniewald Z (2000) Disorders of male reproductive tract under the influence of atrazine. J Appl Toxicol 20:61–68. doi:10.1111/j.1349-7006.2005.00041.x
Komtchou S, Dirany A, Drogui P, Delegan N, Khakani MAE, Robert D, Lafrance P (2016) Degradation of atrazine in aqueous solution with electrophotocatalytic process using TiO2−x photoanode. Chemosphere 157:79–88. doi:10.1016/j.chemosphere.2016.05.022
Kong X, Jiang J, Ma J, Yang Y, Liu W, Liu Y (2016) Degradation of atrazine by UV/chlorine: efficiency, influencing factors, and products. Water Res 90:15–23. doi:10.1016/j.watres.2015.11.068
Konstantinou IK, Sakellarides TM, Sakkas VA, Albanis TA (2001) Photocatalytic degradation of selected s-triazine herbicides and organophosphorus insecticides over aqueous TiO2 suspensions. Environ Sci Technol 35(2):398–405. doi:10.1021/es001271c
Kroon FJ, Hook SE, Jones D, Metcalfe S, Osborn HL (2014) Effects of atrazine on endocrinology and physiology in juvenile barramundi, Lates calcarifer (Bloch). Environ Toxicol Chem 33(7):1607–1614. doi:10.1002/etc.2594
Kuklenyik Z, Panuwet P, Jayatilaka NK, Pirkle JL, Calafat AM (2012) Two-dimensional high performance liquid chromatography separation and tandem mass spectrometry detection of atrazine and its metabolic and hydrolysis products in urine. J Chromatogr B 901:1–8. doi:10.1016/j.jchromb.2012.05.028
Kumar A, Singh N (2016) Atrazine and its metabolites degradation in mineral salts medium and soil using an enrichment culture. Environ Monit Assess 188(3):1–12. doi:10.1007/s10661-016-5144-3
Kumar V, Upadhyay N, Singh S, Singh J, Kaur P (2013) Thin-layer chromatography: comparative estimation of soil’s atrazine. Curr World Environ 8(3):469–472. doi:10.12944/CWE.8.3.17
Kumar V, Upadhyay N, Kumar V, Sharma S (2015a) A review on sample preparation and chromatographic determination of acephate and methamidophos in different samples. Arab J Chem 8:624–631. doi:10.1016/j.arabjc.2014.12.007
Kumar V, Singh S, Singh J, Upadhyay N (2015b) Potential of plant growth promoting traits by bacteria isolated from heavy metal contaminated soils. Bull Environ Contam Toxicol 94:807–815. doi:10.1007/s00128-015-1523-7
Kumar V, Upadhyay N, Manhas A (2015c) Designing syntheses characterization computational study and biological activities of silver-phenothiazine metal complex. J Mol Struct 1099:135–140. doi:10.1016/j.molstruc.2015.06.055
Kumar V, Kumar V, Upadhyay N, Sharma S (2015d) Interactions of atrazine with transition metal ions in aqueous media: experimental and computational approach. 3 Biotech 5:791–798. doi:10.1007/s13205-015-0281-x
Kumar V, Kumar V, Kaur S, Singh S, Upadhyay N (2016) Unexpected formation of N-phenyl-thiophosphorohydrazidic acid O, S-dimethyl ester from acephate: chemical biotechnical and computational study. 3 Biotech 6:1–11. doi:10.1007/s13205-015-0313-6
Kumar V, Singh S, Singh R, Upadhyay N, Singh J (2017) Design, synthesis, and characterization of 2,2-bis(2,4-dinitrophenyl)-2-(phosphonatomethylamino)acetate as a herbicidal and biological active agent. J Chem Biol 11:1–12. doi:10.1007/s12154-017-0174-z
Kungolos A, Samaras P, Kipopoulou AM, Zoumboulis A, Sakellaropoulos GP (1999) Interactive toxic effects of agrochemicals on aquatic organisms. Water Sci Tech 40(1):357–364
Lang D, Criegee D, Grothusen A, Saalfrank RW, Böcker RH (1996) In vitro metabolism of atrazine, terbuthylazine, ametryne, and terbutryne in rats, pigs, and humans. Drug Metab Dispos 24(8):859–865
Lee DH, Rhee YJ, Choi KS, Nam SE, Eom HJ, Rhee JS (2017) Sublethal concentrations of atrazine promote molecular and biochemical changes in the digestive gland of the Pacific oyster Crassostrea gigas. Toxicol Environ Health Sci 9(1):50–58. doi:10.1007/s13530-017-0303-7
Lewis SE, Brodie JE, Bainbridge ZT, Rohde KW, Davis AM, Masters BL, Schaffelke B (2009) Herbicides: a new threat to the Great Barrier Reef. Environ Pollut 157(8):2470–2484. doi:10.1016/j.envpol.2009.03.006 (Epub 2009)
Li X, Wu T, Huang H, Zhang S (2012) Atrazine accumulation and toxic responses in maize (Zea mays). J Environ Sci 24(2):203–208. doi:10.1016/S1001-0742(11)60718-3
Lin J, Li HX, Qin L, Du ZH, Xia J, Li JL (2016a) A novel mechanism underlies atrazine toxicity in quails (Coturnix coturnix): triggering ionic disorder via disruption of ATPases. Oncotarget 7(51):83880. doi:10.1016/j.envpol.2017.04.015
Lin J, Li HX, Xia J, Li XN, Jiang XQ, Zhu SY, Ge J, Li JL (2016b) The chemopreventive potential of lycopene against atrazine-induced cardiotoxicity: modulation of ionic homeostasis. Sci Rep 6:24855. doi:10.1038/srep24855
Lin J, Zhao HS, Xiang LR, Xia J, Wang LL, Li XN, Li JL, Zhang Y (2016c) Lycopene protects against atrazine-induced hepatic ionic homeostasis disturbance by modulating ion-transporting ATPases. J Nutr Biochem 27:249–256. doi:10.1016/j.jnutbio.2015.09.009
Lioy PJ, Edwards RD, Freeman N, Gurunathan S, Pellizzari E, Adgate JL, Sexton K (2000) House dust levels of selected insecticides and a herbicide measured by the EL and LWW samplers and comparisons to hand rinses and urine metabolites. J Expo Sci Environ Epidemiol 10(4):327–340. doi:10.1038/sj.jea.7500099
Liu C, Yang F, Lu X, Huang F, Liu L, Yang C (2010) Isolation, identification and soil remediation of atrazine-degrading strain T3 AB1. Wei Sheng Wu Xue Bao. 50(12):1642–1650
Liu R, Guan G, Wang S, Zhang Z (2011) Core-shell nanostructured molecular imprinting fluorescent chemosensor for selective detection of atrazine herbicide. Analyst 136(1):184–190. doi:10.1039/C0AN00447B
Liu X, Li WJ, Li L, Yang Y, Mao LG, Peng Z (2014) A label-free electrochemical immunosensor based on gold nanoparticles for direct detection of atrazine. Sens Actuator B Chem 191:408–414. doi:10.1016/j.snb.2013.10.033
Lopez MA, Ortega F, Domínguez E, Katakis I (1998) Electrochemical immunosensor for the detection of atrazine. J Mol Recognit 11(1–6):178–181. doi:10.1002/(SICI)1099-1352(199812)11:1/6<178:AID-JMR417>3.0
Lua YC, Yang SN, Zhang JJ, Zhang JJ, Tana LR, Yang H (2013) A collection of glycosyltransferases from rice (Oryza sativa) exposed to atrazine. Gene 531(2):43–252. doi:10.1016/j.gene.2013.09.004
Lydy MJ, Linck SL (2003) Assessing the impact of triazine herbicides on organophosphate insecticide toxicity to the earthworm Eisenia fetida. Arch Environ Contam Toxicol 45(3):343–349. doi:10.1007/s00244-002-0218-y
Ma J, Wang S, Wang P, Ma L, Chen X, Xu R (2006) Toxicity assessment of 40 herbicides to the green alga Raphidocelis subcapitata. Ecotoxicol Environ Saf 63(3):456–462. doi:10.1016/j.ecoenv.2004.12.001
Ma K, Wu HY, Zhang B, He X, Li BX (2015) Neurotoxicity effects of atrazine-induced SH-SY5Y human dopaminergic neuroblastoma cells via microglial activation. Mol BioSyst 11(11):2915–2924. doi:10.1039/C5MB00432B
Ma L, Chen S, Yuan J, Yang P, Liu Y, Stewart K (2017) Rapid biodegradation of atrazine by Ensifer sp. strain and its degradation genes. Int Biodeterior Biodegrad 116:133–140. doi:10.1016/j.ibiod.2016.10.022
Mahía J, Martín A, Carballas T, Díaz-Raviña M (2007) Atrazine degradation and enzyme activities in an agricultural soil under two tillage systems. Sci Total Environ 378(1):187–194. doi:10.1016/j.scitotenv.2007.01.036
Mahler BJ, Van Metre PC, Burley TE, Loftin KA, Meyer MT, Nowell LH (2017) Similarities and differences in occurrence and temporal fluctuations in glyphosate and atrazine in small Midwestern streams (USA) during the 2013 growing season. Sci Total Environ 579:149–158. doi:10.1016/j.scitotenv.2016.10.236
Mandelbaum RT, Allan DL, Wackett LP (1995) Isolation and characterization of a Pseudomonas sp. that mineralizes the s-triazine herbicide atrazine. Appl Environ Microbiol 61(4):1451–1457
Marcus SR, Fiumera AC (2016) Atrazine exposure affects longevity, development time and body size in Drosophila melanogaster. J Insect Physiol 91:18–25. doi:10.1016/j.jinsphys.2016.06.006
Marecik R, Króliczak P, Czaczyk K, Białas W, Olejnik A, Cyplik P (2008) Atrazine degradation by aerobic microorganisms isolated from the rhizosphere of sweet flag (Acorus calamus L,). Biodegradation 19(2):293–301. doi:10.1007/s10532-007-9135-5
Martinez B, Tomkins J, Wackett LP, Wing R, Sadowsky MJ (2001) Complete nucleotide sequence and organization of the atrazine catabolic plasmid padp-1 from Pseudomonas sp. strain ADP. J Bacteriol 183(19):5684–5697. doi:10.1128/JB.183.19.5684-5697.2001
McGregor EB, Solomonb KR, Hanson ML (2008) Effects of planting system design on the toxicological sensitivity of Myriophyllum spicatum and Elodea canadensis to atrazine. Chemosphere 73(3):249–260. doi:10.1016/j.chemosphere.2008.06.045
Mei M, Huang X, Yang X, Luo Q (2016) Effective extraction of triazines from environmental water samples using magnetism-enhanced monolith-based in-tube solid phase microextraction. Anal Chim Acta 937:69–79. doi:10.1016/j.aca.2016.08.001
Mela M, Guiloski IC, Doria HB, Randi MAF, de Oliveira Ribeiro CA, Pereira L, de Assis HS (2013) Effects of the herbicide atrazine in neotropical catfish (Rhamdia quelen). Ecotoxicol Environ Saf 93:13–21. doi:10.1016/j.ecoenv.2013.03.026
Miensah ED, Fianko JR, Adu-Kumi S (2015) Assessment of lindane and atrazine residues in maize produced in ghana using gas chromatography-electron capture detector (GC-ECD) and gas chromatography-mass spectrometry (GC-MS). J Environ Prot Sci 6(10):1105. doi:10.4236/jep.2015.610097
Morales-Pérez AA, Arias C, Ramírez-Zamora RM (2016) Removal of atrazine from water using an iron photo catalyst supported on activated carbon. Adsorption 22(1):49–58. doi:10.1007/s10450-015-9739-8
Na Y, Sheng W, Yuan M, Li L, Liu B, Zhang Y, Wang S (2012) Enzyme-linked immunosorbent assay and immunochromatographic strip for rapid detection of atrazine in water samples. Microchim Acta 177(1–2):177–184. doi:10.1007/s00604-012-0772-y
National Institute for Occupational Safety and Health, NIOSH (1998a) Criteria for a recommended standard. Occupational exposure to noise. Revised Criteria. USDHHS, PHS, CDC, NIOSH, publication no. 98–126, Cincinnati
National Institute for Occupational Safety and Health, NIOSH (1998b) Chlorinated and organonitrogen herbicides (hand wash): method 9200, manual of analytical methods (NMAM), 4th edn
Nawaz A, Razpotnik A, Rouimi P, de Sousa G, Cravedi JP, Rahmani R (2014) Cellular impact of combinations of endosulfan, atrazine, and chlorpyrifos on human primary hepatocytes and HepaRG cells after short and chronic exposures. Cell Biol Toxicol 30(1):17–29. doi:10.1007/s10565-013-9266-x
Nélieu S, Kerhoas L, Einhorn J (2000) Degradation of atrazine into ammeline by combined ozone/hydrogen peroxide treatment in water. Environ Sci Technol 34(3):430–437. doi:10.1021/es980540k
Nemat AMM, Hassan NM (2006) Changes of antioxidants levels in two maize lines following atrazine treatments. Plant Phys Biochem 44(4):202–210. doi:10.1016/j.plaphy.2006.05.004
Nousiainen AO, Björklöf K, Sagarkar S, Nielsen JL, Kapley A, Jørgensen KS (2015) Bioremediation strategies for removal of residual atrazine in the boreal groundwater zone. Appl Microbiol Biotechnol 99(23):10249–10259. doi:10.1007/s00253-015-6828-2
Nsibande SA, Dabrowski JM, van der Walt E, Venter A, Forbes PB (2015) Validation of the AGDISP model for predicting airborne atrazine spray drift: a South African ground application case study. Chemosphere 138:454–461. doi:10.1016/j.chemosphere.2015.06.092
Nwani CD, Nagpure NS, Kumar R, Kushwaha B, Kumar P, Lakra WS (2011) Mutagenic and genotoxic assessment of atrazine-based herbicide to freshwater fish Channa punctatus (Bloch) using micronucleus test and single cell gel electrophoresis. Environ Toxicol Pharmacol 31(2):314–322. doi:10.1016/j.etap.2010.12.001
Oka T, Tooi O, Mitsui N, Miyahara M, Ohnishi Y, Takase M, Iguchi T (2008) Effect of atrazine on metamorphosis and sexual differentiation in Xenopus laevis. Aquat Toxicol 87(4):215–226. doi:10.1016/j.aquatox.2008.02.009
Oluah NS, Obiezue RNN, Ochulor AJ, Onuoha E (2016) Toxicity and histopathological effect of atrazine (herbicide) on the earthworm Nsukkadrilus mbae under laboratory conditions. Anim Res Int 7(3):1287–1293
Omotayo AE, Ilori MO, Obayori OS, Amund OO (2016) Influence of pH, temperature and nutrient addition on the degradation of atrazine by Nocardioides spp. isolated from agricultural soil in Nigeria. Malays J Microbiol. doi:10.21161/mjm.81315
Omran NE, Salama WM (2016) The endocrine disruptor effect of the herbicides atrazine and glyphosate on Biomphalaria alexandrina snails. Toxicol Ind Health 32(4):656–665. doi:10.1177/0748233713506959
Orton F, Carr JA, Handy RD (2006) Effects of nitrate and atrazine on larval development and sexual differentiation in the northern leopard frog Rana pipiens. Environ Toxicol Chem 25(1):65–71. doi:10.1897/05-136R.1
Pardieu E, Cheap H, Vedrine C, Lazerges M, Lattach Y, Garnier F, Pernelle C (2009) Molecularly imprinted conducting polymer based electrochemical sensor for detection of atrazine. Anal Chim Acta 649(2):236–245. doi:10.1016/j.aca.2009.07.029
Peighambarzadeh SZ, Safi S, Shahtaheri SJ, Javanbakht M, Forushani AR (2011) Presence of atrazine in the biological samples of cattle and its consequence adversity in human health. Iran J Public Health 40(4):112
Pensabene JW, Fiddler W, Donoghue DJ (2000) Supercritical fluid extraction of atrazine and other triazine herbicides from fortified and incurred eggs. J Agric Food Chem 48(5):1668–1672. doi:10.1021/jf990841t
Piutti S, Semon E, Landry D, Hartmann A, Dousset S, Lichtfouse E, Martin-Laurent F (2003) Isolation and characterisation of Nocardioides sp, SP12 an atrazine-degrading bacterial strain possessing the gene trzN from bulk-and maize rhizosphere soil. FEMS Microbiol Lett 221(1):111–117. doi:10.1016/S0378-1097(03)00168-X
Pogrmic K, Fa S, Dakic V, Kaisarevic S, Kovacevic R (2009) Atrazine oral exposure of peripubertal male rats downregulates steroidogenesis gene expression in Leydig cells. Toxicol Sci 111(1):189–197. doi:10.1093/toxsci/kfp135
Pommery J, Mathieu M, Mathieu D, Lhermitte M (1993) Atrazine in plasma and tissue following atrazine-aminotriazole-ethylene glycol-formaldehyde poisoning. J Toxicol Clin Toxicol 31(2):323–331. doi:10.3109/15563659309000399
Prasad R, Upadhyay N, Kumar V (2013) Simultaneous determination of seven carbamate pesticide residues in gram wheat lentil soybean fenugreek leaves and apple matrices. Microchem J 111:91–97. doi:10.1016/j.microc.2012.12.014
Qie Z, Ning B, Liu M, Bai J, Peng Y, Song N, Zhang Y (2013) Fast detection of atrazine in corn using thermometric biosensors. Analyst 138(17):5151–5156. doi:10.1039/c3an00490b
Qingyan LI, Ying LI, Xikun ZHU, Baoli CAI (2008) Isolation and characterization of atrazine-degrading Arthrobacter sp, AD26 and use of this strain in bioremediation of contaminated soil. J Environ Sci 20(10):1226–1230. doi:10.1111/lam.12584
Radosevich M, Traina SJ, Hao YL, Tuovinen OH (1995) Degradation and mineralization of atrazine by a soil bacterial isolate. Appl Environ Microbiol 61(1):297–302
Ralston-Hooper K, Hardy J, Hahn L, Ochoa-Acuña H, Lee LS, Mollenhauer R, Sepúlveda MS (2009) Acute and chronic toxicity of atrazine and its metabolites deethylatrazine and deisopropylatrazine on aquatic organisms. Ecotoxicol 18(7):899–905. doi:10.1007/s10646-009-0351-0
Raveton M, Ravanel P, Kaouadji M, Bastide J, Tissut M (1997) The chemical transformation of atrazine in corn seedlings. Pestic Biochem Physiol 58(3):199–208. doi:10.1006/pest.1997.2303
Rodriguez VM, Mendoza-Trejo MS, Hernandez-Plata I, Giordano M (2017) Behavioral effects and neuroanatomical targets of acute atrazine exposure in the male Sprague-Dawley rat. NeuroToxicol 58:161–170. doi:10.1016/j.neuro.2016.12.006
Rousseaux S, Hartmann A, Soulas G (2001) Isolation and characterisation of new Gram-negative and Gram-positive atrazine degrading bacteria from different French soils. FEMS Microbiol Ecol 36(2–3):211–222. doi:10.1111/j.1574-6941.2001.tb00842.x
Sabik H, Jeannot R (1998) Determination of organonitrogen pesticides in large volumes of surface water by liquid–liquid and solid-phase extraction using gas chromatography with nitrogen–phosphorus detection and liquid chromatography with atmospheric pressure chemical ionization mass spectrometry. J Chromatogr A 818(2):197–207. doi:10.1016/S0021-9673(98)00555-X
Santos TG, Martinez CB (2012) Atrazine promotes biochemical changes and DNA damage in a Neotropical fish species. Chemosphere 89(9):1118–1125. doi:10.1016/j.chemosphere.2012.05.096
Sawangjit S (2016) Isolation and characterization of atrazine-degrading Xanthomonas sp. ARB2 and its use in bioremediation of contaminated soils. Int J Environ Sci Technol 7(5):351. doi:10.7763/IJESD.2016.V7.798
Schipper EF, Bergevoet AJH, Kooyman RPH, Greve J (1997) New detection method for atrazine pesticides with the optical waveguide Mach-Zehnder immunosensor. Anal Chim Acta 341(2–3):171–176. doi:10.1016/S0003-2670(96)00622-8
Schirhagl R, Latif U, Dickert FL (2011) Atrazine detection based on antibody replicas. J Mater Chem 21(38):14594–14598. doi:10.1039/C1JM11576F
Schroeder JC, Olshan AF, Baric R, Dent GA, Weinberg CR, Yount B, Rothman N (2001) Agricultural risk factors for t (14; 18) subtypes of non-Hodgkin’s lymphoma. Epidemiology 12(6):701–709. doi:10.1097/00001648-200111000-00020
Schwab AP, Splichal PA, Banks MK (2006) Persistence of atrazine and alachlor in ground water aquifers and soil. Water Air Soil Pollut 171(1–4):203–235. doi:10.1007/s11270-005-9037-2
Seffernick JL, de Souza ML, Sadowsky MJ, Wackett LP (2001) Melamine deaminase and atrazine chlorohydrolase: 98 percent identical but functionally different. J Bacteriol 183(8):2405–2410. doi:10.1128/JB.183.8.2405-2410.2001
Seffernick JL, McTavish H, Osborne JP, de Souza ML, Sadowsky MJ, Wackett LP (2002) Atrazine chlorohydrolase from Pseudomonas sp, strain ADP is a metalloenzyme. Biochem 41(48):14430–14437. doi:10.1021/bi020415s
Shamsedini N, Dehghani M, Nasseri S, Baghapour MA (2017) Photocatalytic degradation of atrazine herbicide with illuminated Fe3+–TiO2 nanoparticles. J Environ Health Sci Eng 15:7. doi:10.1186/s40201-017-0270-6
Shao ZQ, Behki R (1995) Cloning of the genes for degradation of the herbicides EPTC (S-ethyl dipropylthiocarbamate) and atrazine from Rhodococcus sp. strain TE1. Appl Environ Microbiol 61(5):2061–2065. doi:10.1111/j.1472-765X.2009.02724.x
Shao ZQ, Behki R (1996) Characterization of the expression of the thcB gene, coding for a pesticide-degrading cytochrome P-450 in Rhodococcus strains. Appl Environ Microbiol 62(2):403–407
Shim WB, Yang ZY, Kim JY, Choi JG, Je JH, Kang SJ, Chung DH (2006) Immunochromatography using colloidal gold-antibody probe for the detection of atrazine in water samples. J Agric Food Chem 54(26):9728–9734. doi:10.1021/jf0620057
Shimabukuro RH, Swanson HR, Walsh WC (1970) Glutathione conjugation atrazine detoxication mechanism in corn. Plant Physiol 46(1):103–107. doi:10.1104/pp.46.1.103
Silveyra GR, Canosa IS, Rodríguez EM, Medesani DA (2017) Effects of atrazine on ovarian growth, in the estuarine crab Neohelice granulata. Comp Biochem Physiol C Toxicol Pharmacol 192:1–6. doi:10.1016/j.cbpc.2016.10.011
Simpkins JW, Swenberg JS, Weiss N, Brusick D, Eldridge JC, Stevens JT, Handa RJ, Hovey RC, Plant TM, Pastoor TP, Breckenridge CB (2011) Atrazine and breast cancer: a framework assessment of the toxicological and epidemiological evidence. Toxicol Sci 123(2):441–459. doi:10.1093/toxsci/kfr176
Singh S, Singh N, Kumar V, Datta S, Wani AB, Singh D, Singh K, Singh J (2016) Toxicity monitoring and biodegradation of the fungicide carbendazim. Environ Chem Lett 14:317–329. doi:10.1007/s10311-016-0566-2
Singh S, Kumar V, Upadhyay N, Singh J, Singla S, Datta S (2017) Efficient biodegradation of acephate by Pseudomonas pseudoalcaligenes PS-5 in the presence and absence of heavy metal ions [Cu(II) and Fe(III)], and humic acid. 3 Biotech 7:262. doi:10.1007/s13205-017-0900-9
Solomon KR, Giesy JP, LaPoint TW, Giddings JM, Richards RP (2013a) Ecological risk assessment of atrazine in North American surface waters. Environ Toxicol Chem 32(1):10–11. doi:10.1002/etc.5620150105
Solomon RDJ, Kumar A, Santhi VS (2013b) Atrazine biodegradation efficiency metabolite detection and trzD gene expression by enrichment bacterial cultures from agricultural soil. J Zhejiang Univ Sci B 14(12):1162–1172. doi:10.1631/jzus.B1300001
Soltanian S (2016) Effect of atrazine on immunocompetence of red-eared slider turtle (Trachemys scripta). J Immunotoxicol 13(6):804–809. doi:10.1080/1547691X.2016.1195463
Sopid S (2012) Characterization of novel atrazine degrading Klebsiella sp, isolated from Thai agricultural soil. World Acad Sci Eng Technol 68:1656–1658. doi:10.7763/IJESD.2016.V7.798
Stoker TE, Guidici DL, Laws SC, Cooper RL (2002) The effects of atrazine metabolites on puberty and thyroid function in the male Wistar rat. Toxicol Sci 67(2):198–206. doi:10.1093/toxsci/58.1.50
Strategic Diagnostics Inc., SDI (1998) Environmental technology verification report, immunoassay kit. EPA/600/R-98/112
Strategic Diagnostics Inc., SDI (1999) Water quality testing product profile: atrazine rapid assay. Strategic Diagnostics Inc, Newark, Delaware
Strong LC, Rosendahl C, Johnson G, Sadowsky MJ, Wackett LP (2002) Arthrobacter aurescens TC1 metabolizes diverse s-triazine ring compounds. Appl Environ Microbiol 68(12):5973–5980. doi:10.1128/AEM.68.12.5973-5980.2002
Struthers JK, Jayachandran K, Moorman TB (1998) Biodegradation of atrazine by Agrobacterium radiobacter J14a and use of this strain in bioremediation of contaminated soil. Appl Environ Microbiol 64(9):3368–3375
Su YH, Zhu YG (2006) Bioconcentration of atrazine and chlorophenols into roots and shoots of rice seedlings. Environ Pollut 139(1):32–39. doi:10.1016/j.envpol.2005.04.035
Suri CR, Kaur J, Gandhi S, Shekhawat GS (2008) Label-free ultra-sensitive detection of atrazine based on nanomechanics. Nanotech 19(23):235502. doi:10.1088/0957-4484/19/23/235502
Suzawa M, Ingraham HA (2008) The herbicide atrazine activates endocrine gene networks via non-steroidal NR5A nuclear receptors in fish and mammalian cells. PLoS ONE 3(5):e2117. doi:10.1371/journal.pone.0002117
Szigeti Z, Lehoczki E (2003) A review of physiological and biochemical aspects of resistance to atrazine and paraquat in Hungarian weeds. Pestic Manag Sci 59(4):451–458. doi:10.1002/ps.647
Tan F, Zhao C, Li L, Liu M, He X, Gao J (2015) Graphene oxide based in-tube solid-phase microextraction combined with liquid chromatography tandem mass spectrometry for the determination of triazine herbicides in water. J Sep Sci 38(13):2312–2319. doi:10.1002/jssc.201570131/full
Tana LR, Lua YC, Zhanga JJ, Luo F, Yang H (2015) A collection of cytochrome P450 monooxygenase genes involved in modification and detoxification of herbicide atrazine in rice (Oryza sativa) plants. Ecotoxicol Environ Saf 119:25–34. doi:10.1016/j.ecoenv.2015.04.035
Tavera-Mendoza L, Ruby S, Brousseau P, Fournier M, Cyr D, Marcogliese D (2002) Response of the amphibian tadpole (Xenopus laevis) to atrazine during sexual differentiation of the testis. Environ Toxicol Chem 21(3):527–531. doi:10.1002/etc.5620210309
Topp E, Mulbry WM, Zhu H, Nour SM, Cuppels D (2000a) Characterization of s-triazine herbicide metabolism by a Nocardioides sp, isolated from agricultural soils. Appl Environ Microbiol 66(8):3134–3141. doi:10.1128/AEM.66.8.3134-3141.2000
Topp E, Zhu H, Nour SM, Houot S, Lewis M, Cuppels D (2000b) Characterization of an atrazine-degrading Pseudaminobacter sp, isolated from Canadian and French agricultural soils. Appl Environ Microbiol 66(7):2773–2782. doi:10.1128/AEM.66.7.2773-2782.2000
Tortolini C, Bollella P, Antiochia R, Favero G, Mazzei F (2016) Inhibition-based biosensor for atrazine detection. Sens Actuator B Chem 224:552–558. doi:10.1016/j.snb.2015.10.095
Tran HV, Reisberg S, Piro B, Nguyen TD, Pham MC (2013) Label-free electrochemical immunoaffinity sensor based on impedimetric method for pesticide detection. Electroanalysis 25(3):664–670. doi:10.1002/elan.201200331/
Udiković-Kolić N, Hršak D, Devers M, Klepac-Ceraj V, Petrić I, Martin-Laurent F (2010) Taxonomic and functional diversity of atrazine-degrading bacterial communities enriched from agrochemical factory soil. J Appl Microbiol 109(1):355–367. doi:10.1111/j.1365-2672.2010.04700.x
Vaishampayan PA, Kanekar PP, Dhakephalkar PK (2007) Isolation and characterization of Arthrobacter sp, strain MCM B-436 an atrazine-degrading bacterium from rhizospheric soil. Int Biodeterior Biodegrad 60(4):273–278. doi:10.1016/j.ibiod.2007.05.001
Victor-Costa AB, Bandeira SMC, Oliveira AG, Mahecha GAB, Oliveira CA (2010) Changes in testicular morphology and steroidogenesis in adult rats exposed to Atrazine. Reprod Toxicol 29(3):323–331. doi:10.1016/j.reprotox.2009.12.006
Vogel A, Jocque H, Sirot LK, Fiumera AC (2015) Effects of atrazine exposure on male reproductive performance in Drosophila melanogaster. J Insect Physiol 72:14–21. doi:10.1016/j.jinsphys.2014.11.002
Vonberg D, Vanderborght J, Cremer N, Pütz T, Herbst M, Vereecken H (2014) 20 years of long-term atrazine monitoring in a shallow aquifer in western Germany. Water Res 50:294–306. doi:10.1016/j.watres.2013.10.032
Wackett L, Sadowsky M, Martinez B, Shapir N (2002) Biodegradation of atrazine and related s-triazine compounds: from enzymes to field studies. Appl Microbiol Biotechnol 58(1):39–45. doi:10.1007/s00253-001-0862-y
Wang Q, Xie S (2012) Isolation and characterization of a high-efficiency soil atrazine-degrading Arthrobacter sp, strain. Int Biodeterior Biodegrad 71:61–66. doi:10.1371/journal.pone.0107270
Wang L, Samac DA, Shapir N, Wackett LP, Vance CP, Olszewski NE, Sadowsky MJ (2005) Biodegradation of atrazine in transgenic plants expressing a modified bacterial atrazine chlorohydrolase (atzA) gene. Plant Biotech J 3(5):475–486. doi:10.1111/j.1467-7652.2005.00138.x
Wang X, Zhu L, Wang J, Xie H, Liu W, Wang Q (2006) Study on enzymatic degradation of herbicide atrazine by bacteria HB-5. J Environ Sci 26(4):579–583
Wang Q, Xie S, Hu R (2013) Bioaugmentation with Arthrobacter sp, strain DAT1 for remediation of heavily atrazine-contaminated soil. Int Biodeterior Biodegrad 77:63–67. doi:10.1016/j.ibiod.2012.11.003
Wang J, Zhu L, Wang Q, Wang J, Xie H (2014a) Isolation and characterization of atrazine mineralizing Bacillus subtilis strain HB-6. PLoS ONE 9(9):e107270. doi:10.1371/journal.pone.0107270
Wang L, Cao M, Ai Z, Zhang L (2014b) Dramatically enhanced aerobic atrazine degradation with Fe@ Fe2O3 core–shell nanowires by tetrapolyphosphate. Environ Sci Technol 48(6):3354–3362. doi:10.1021/es404741x
Wang H, Liu Y, Li J, Lin M, Hu X (2016) Biodegradation of atrazine by Arthrobacter sp. C3, isolated from the herbicide-contaminated corn field. Int J Environ Sci Technol 13(1):257–262. doi:10.1007/s13762-015-0860-8
Weber GJ, Sepúlveda MS, Peterson SM, Lewis SS, Freeman JL (2013) Transcriptome alterations following developmental atrazine exposure in zebrafish are associated with disruption of neuroendocrine and reproductive system function, cell cycle, and carcinogenesis. Toxicol Sci 132(2):458–466. doi:10.1093/toxsci/kft017
White A (2016) Atrazine-a case of discrediting science. Sci Educ News 65(1):22. ISSN: 0048-9603
Williams DBG, George MJ, Marjanovic L (2014) Rapid detection of atrazine and metolachlor in farm soils: gas chromatography-mass spectrometry-based analysis using the bubble-in-drop single drop microextraction enrichment method. J Agric Food Chem 62(31):7676–7681. doi:10.1021/jf502411t
Wolf DC, Martin JP (1975) Microbial decomposition of ring-14C atrazine, cyanuric acid, and 2-chloro-4, 6-diamino-s-triazine. J Environ Qual 4(1):134–139. doi:10.2134/jeq1975.00472425000400010032x
Wüst S, Hock B (1992) A sensitive enzyme immunoassay for the detection of atrazine based upon sheep antibodies. Anal Lett 25:1025–1037. doi:10.1080/00032719208020056
Xing H, Wang X, Sun G, Gao X, Xu S, Wang X (2012) Effects of atrazine and chlorpyrifos on activity and transcription of glutathione S-transferase in common carp (Cyprinus carpio L.). Environ Toxicol Pharmacol 33(2):233–244. doi:10.1016/j.etap.2011.12.014
Xing H, Li S, Wang X, Gao X, Xu S, Wang X (2013) Effects of atrazine and chlorpyrifos on the mRNA levels of HSP70 and HSC70 in the liver, brain, kidney and gill of common carp (Cyprinus carpio L.). Chemosphere 90(3):910–916. doi:10.1016/j.chemosphere.2012.06.028
Xu J, Zhang P, Mu H, Gao ML (2006) Toxicity effect of combined contamination of two herbicides to earthworm [J]. J Agro Environ Sci 5:018
Yang C, Li Y, Zhang K, Wang X, Ma C, Tang H, Xu P (2010) Atrazine degradation by a simple consortium of Klebsiella sp, A1 and Comamonas sp, A2 in nitrogen enriched medium. Biodegradation 21(1):97–105. doi:10.1007/s10532-009-9284-9
Yang SONG, Jia ZC, Chen JY, Hu JX, Zhang LS (2014) Toxic effects of atrazine on reproductive system of male rats. Biomed Environ Sci 27(4):281–288. doi:10.3967/bes2014.050
Yang J, Li J, Dong W, Mab J, Lia J (2016a) Influence of nitrite on the degradation of atrazine by ozonation. J Chem Technol Biotechnol 92(2):442–450. doi:10.1002/jctb.5031
Yang J, Lia J, Dong W, Ma J, Cao J, Li T, Li J, Gu J, Liu P (2016b) Study on enhanced degradation of atrazine by ozonation in thepresence of hydroxylamine. J Hazard Mat 316:110–121. doi:10.1016/j.jhazmat.2016.04.078
Yang M, Zhao X, Zheng S, Liu X, Jin B, Li H, Gan Y (2017) A new electrochemical platform for ultrasensitive detection of atrazine based on modified self-ordered Nb2O5 nanotube arrays. J Electroanal Chem 791:17–22. doi:10.1016/j.jelechem.2017.03.009
Yanze-Kontchou C, Gschwind N (1994) Mineralization of the herbicide atrazine as a carbon source by a Pseudomonas strain. Appl Environ Microbiol 60(12):4297–4302
Yaqub S, Latif U, Dickert FL (2011) Plastic antibodies as chemical sensor material for atrazine detection. Sens Actuators B Chem 160(1):227–233. doi:10.1016/j.electacta.2009.11.029
Yilmaz E, Özgür E, Bereli N, Türkmen D, Denizli A (2017) Plastic antibody based surface plasmon resonance nanosensors for selective atrazine detection. Mater Sci Eng C 73:603–610. doi:10.1016/j.msec.2016.12.090
Yokley RA, Mayer LC, Rezaaiyan R, Manuli ME, Cheung MW (2000) Analytical method for the determination of cyromazine and melamine residues in soil using LC-UV and GC-MSD. J Agric Food Chem 48(8):3352–3358. doi:10.1021/jf991231w
Zadeh AK, Sohrab AD, Alishahi M, Khazaei SH, Asgari HM (2016) Evaluation of acute and sub-lethal toxicity of herbicide, atrazine, on hematological parameters of Tor grypus. J Vet Res 71(3):295–301
Zhang Y, Jiang Z, Cao B, Hu M, Wang Z, Dong X (2011) Metabolic ability and gene characteristics of Arthrobacter sp, strain DNS10 the sole atrazine-degrading strain in a consortium isolated from black soil. Int Biodeterior Biodegrad 65(8):1140–1144. doi:10.1016/j.ibiod.2011.08.010
Zhao P, Wang L, Zhou L, Zhang F, Kang S, Pan C (2012) Multi-walled carbon nanotubes as alternative reversed-dispersive solid phase extraction materials in pesticide multi-residue analysis with QuEChERS method. J Chromatogr A 1225:17–25. doi:10.1016/j.chroma.2011.12.070
Zhao X, Wang L, Ma F, Bai S, Yang J, Qi S (2017) Pseudomonas sp, ZXY-1 a newly isolated and highly efficient atrazine-degrading bacterium and optimization of biodegradation using response surface methodology. J Environ Sci 54:52–159. doi:10.1016/j.jes.2016.06.010
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Human and animal rights
This study does not involve work
Rights and permissions
About this article
Cite this article
Singh, S., Kumar, V., Chauhan, A. et al. Toxicity, degradation and analysis of the herbicide atrazine. Environ Chem Lett 16, 211–237 (2018). https://doi.org/10.1007/s10311-017-0665-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10311-017-0665-8