Abstract
The effect of grazing on the abundance, composition, and methane (CH4) uptake of methanotrophs in grasslands has been well documented in the past few decades, but the dominant communities of active methanotrophs responsible for CH4 oxidation activity in grazed soils are still poorly understood. In this study, we characterized the metabolically active, aerobic methanotrophs in grasslands with three different levels of grazing (light, medium, and heavy) by combining DNA-stable isotope probing (SIP) and quantitative PCR (qPCR) for methane monooxygenase (pmoA) gene– and 16S rRNA gene–based amplicon sequencing. The CH4 oxidation potential was as low as 0.51 μmol g dry weight−1 day−1 in the ungrazed control, while it decreased as grazing intensity increased in grazed fields, ranging from 2.25 μmol g dry weight−1 day−1 in light grazed fields to 1.59 in heavily grazed fields. Increased CH4 oxidation activity was paralleled by twofold increases in abundance of pmoA genes and relative abundance of methanotroph-affiliated 16S rRNA genes in the total microbial community in grazed soils. SIP and sequencing revealed that the genera Methylobacter and Methylosarcina (type I; Gammaproteobacteria) and Methylocystis (type II; Alphaproteobacteria) were active methanotrophs responsible for CH4 oxidation in grazed soils. Light and intermediate grazing stimulated the growth and activity of methanotrophs, while heavy grazing decreased the abundance and diversity of the active methanotrophs in the typical steppe. Redundancy and correlation analysis further indicated that the variation of bulk density and soil C and N induced by grazing determined the abundance, diversity of active methanotrophs, and methane oxidation activity in the long-term grazed grassland soil.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Methane (CH4) is the second most important anthropogenic greenhouse gas and is about 28 times more efficient at absorbing infrared radiation than CO2 (Stocker et al. 2013). Methane emissions can be attenuated by aerobic methane-oxidizing bacteria known as methanotrophs. These bacteria have the ability to utilize methane as their source of C and energy. Methanotrophs can act either as a bio-filter in wetland agricultural soils (paddy field) or as a methane sink in well-aerated soils (Bridgham et al. 2013; Fan et al. 2019a, b; Kolb 2009; Wei et al. 2019). Although methane can also be oxidized anaerobically by microbes using nitrite, nitrate, ferric iron, and sulfate as electron acceptors, methanotrophs are ubiquitous in nature and represent the only oxic biogenic sink for the greenhouse gas methane (Fan et al. 2019b; Hanson and Hanson 1996; Shen et al. 2019; Trotsenko and Murrell 2008).
Historically, aerobic methanotrophs have been classified as types I, II, and X, based on physiological, biochemical, and phenotypical characteristics (Veraart et al. 2015). Nowadays, they are preferentially classified based on phylogeny, either belonging to the classes of Gammaproteobacteria (referred to as type I or X methanotrophs), Alphaproteobacteria (referred to as type II methanotrophs), or to the phyla Verrucomicrobia (referred to as type III methanotrophs) and NC10 (Knief 2015; Stein et al. 2012). More specifically, type I methanotrophs belong to the Methylococcaceae (type Ia and type Ib) and Methylothermaceae (type Ic) families, while type II methanotrophs are divided into type IIa (Methylocystaceae) and type IIb (Beijerinckiaceae) (Lüke and Frenzel 2011). Converting CH4 to methanol is the key step in the CH4 oxidation pathway, which is catalyzed by the enzyme methane monooxygenase (Hanson and Hanson 1996). The pmoA gene encodes the β-subunit of the particulate methane monooxygenase enzyme (pMMO) and is the most commonly used functional marker for identifying methanotrophs in environmental samples (Dumont 2014).
Grasslands are one of the largest terrestrial ecosystems in the world, covering approximately 20–40% of the earth’s surface and accounting for up to 40% of the total land area in China (Nan 2005). Grassland soils are one of the largest terrestrial methane sinks with the capacity for methane oxidation (Pachauri et al. 2014). As the most popular management of grasslands, animal grazing may lead to different soil physical conditions (e.g., bulk density and aeration status) and chemical properties (e.g., pH and organic matter content) (Li et al. 2008; Steffens et al. 2008). Soil bulk density and moisture are considered to be critical factors for CH4 oxidation activity in soil (Serrano-Silva et al. 2014). Animal grazing alters methanotroph composition and methane uptake activities by reducing soil water content and soil aeration through herbage removing and tramping, respectively (Leriche et al. 2001; Zhou et al. 2008). Heavy grazing has been reported to significantly reduce annual CH4 uptake, while light-to-moderate grazing has been shown to have either a considerable positive impact on CH4 uptake or to not significantly change CH4 uptake (Chen et al. 2011; Ma et al. 2018). Previous studies either measured the abundance and/or composition of methanotroph in grassland soils or try to link CH4 uptake with the abundance of the pmoA gene. However, mere presence of pmoA genes under in situ soil conditions may not necessarily reflect the functional activity of CH4 oxidation (Nannipieri et al. 2019). To our knowledge, there are little studies that link CH4 oxidation with functional active methanotrophs in grazed grassland soils.
The objective of this study was to obtain information on the presence of aerobic methanotrophs affected by grazing gradients in the field and to identify the active microorganisms responsible for CH4 oxidation in a typical steppe grazed soils. We applied quantitative PCR (qPCR) of the pmoA gene and Miseq sequencing coupled with DNA-based stable isotope probing (SIP) to study the active methanotroph communities from field soils. Based on previous studies, we hypothesized that different levels of grazing would alter the abundance and communities of active methanotrophs and change the methane oxidation activity.
Materials and methods
Setup of the experiment
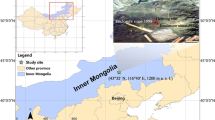
Four different stocking rates were established in 2005, with zero, three, nine, and fifteen sheep per two hectares, at the Inner Mongolia Grassland Ecosystem Research Station (IMGERS, 43° 37′ N, 116° 43′ E) on the Xilingol steppe of the Xilin River basin. The dominant plant species in the study area is Leymus chinensis with some Stipa grandis and Cleistogenes squarrosa. The four stocking rates were classified as ungrazed control (CK), light grazing (G1), moderate grazing (G2), and heavy grazing (G3). Each year, sheep are in the field from June to September (~ 95 days), in accordance with the local summer grazing season. The different stocking rate treatments were arranged in a randomized block design with three replicates separated by fences, and the plot size for each block was 2 ha. Composite soil samples of each replicate were collected from the upper 10-cm layer from 5 random locations using a 5 cm in diameter soil auger in August 2015 and transported to the laboratory with a cold chain. After passing through a 2-mm sieve, soil samples of each replicate were separated into three subsamples for DNA extraction, physicochemical property analysis, and SIP incubation experiments, respectively. Soil physicochemical properties were determined according to the protocols of the Handbook of Soil Analysis (Pansu and Gautheyrou 2007). The soil type is dark chestnut (calcic Chernozem according to the ISSS Working Group RB (1998)).
Microcosm incubations for SIP and CH4 oxidation potential were conducted in sterile 120-mL glass serum vials in triplicate, containing 10 g (dry weight) of soils, and sealed with butyl rubber stoppers. Each microcosm was injected with either 13CH4 (99 atom % 13C, Sigma-Aldrich, USA) or CH4 (99.5% pure, as control) to a final concentration of 8% (v/v) in the headspace and incubated at 25 °C in dark conditions for 28 days. CH4 oxidation potential was assessed from the zero-order decrease in CH4 concentration in the headspace of the serum vials within 12 h and measured with gas chromatography (Shimadzu GC12-A, Japan) (Kightley et al. 1995). After more than 90% of the CH4 was consumed, the headspace was flushed with pressurized synthetic air (20% O2, 80% N2) for 1 min to maintain oxic conditions and, after that, the labelled or unlabelled CH4 was renewed. The atmosphere of the microcosms was renewed three times with pressurized synthetic air and labelled or unlabelled CH4 during the incubation period. Destructive sampling was performed in triplicate for further soil analysis after incubation of SIP microcosms for 28 days.
Nucleic acid extraction and SIP fractionation
DNA was carefully extracted from 0.5 g of soil using a FastDNA SPIN kit for soil (MP Biomedicals; Solon, OH, USA) according to the manufacturer’s instructions. Negative control without soil was not tested as this study focuses on the active methanotrophs which were labelled with 13C (Vestergaard et al. 2017). The concentrations and quality of DNA were estimated by a Nanodrop® ND-2000 UV-Vis spectrophotometer (NanoDrop Technologies, Wilmington, DE, USA) and gel electrophoresis.
Density gradient centrifugation was performed on bulk DNA extracted from the 13CH4 and CH4 treatments as described by Liu et al. (2019a) and Zhang et al. (2019), with a minor modification in scale: 2.5 μg of DNA were mixed with gradient buffer/CsCl solution in a 5.1-mL Beckman ultracentrifuge tube and DNA was fractionated into 14 equal fractions after centrifuged at 177,000gav for 44 h at 20 °C in a Vti65.2 vertical rotor (Beckman Coulter, Palo Alto, CA, USA) (Jia et al. 2019). The fractionated DNA was purified and dissolved in 30 μL of TE buffer.
Quantitative PCR of the pmoA genes
Quantitative PCR analysis of the pmoA gene in bulk DNA and in each DNA gradient fraction was performed to determine the growth and efficiency of 13C incorporation into the genomic DNA of methanotroph communities, respectively. The primer pair A189f and mb661r (Costello and Lidstrom 1999; Holmes et al. 1995) was used for the qPCR of the pmoA gene as described previously (Zheng et al. 2014). qPCR runs were carried out in a LightCycler® 480II (Roche, Germany). Efficiencies of 89–105% were obtained for all gene amplifications, with R2 values ranging between 0.992 and 0.999.
MiSeq sequencing and phylogenetic analysis
Methanotroph-affiliated 16S rRNA gene was amplified in both 13CH4 and CH4-control microcosms in bulk DNA and in heavy fractions of each gradient with standard PCR conditions and quantified DNA as suggested (Schöler et al. 2017). Methanotroph-affiliated 16S rRNA libraries were constructed and sequenced using an Illumina® MiSeq sequencer (Illumina, San Diego, CA, USA) by Majorbio Bio-pharm Technology Co., Ltd. (Shanghai, China) with a universal 515F-907R primer assay as reported previously (Daebeler et al. 2014; Zheng et al. 2014). Raw fastq files were quality-filtered by Trimmomatic and merged by FLASH with the following criteria: (i) The reads were truncated at any site receiving an average quality score < 20 over a 50-bp sliding window. (ii) Sequences with overlap being longer than 10 bp were merged according to their overlap with mismatch no more than 2 bp. (iii) Sequences of each sample were separated according to barcodes (exactly matching) and primers (allowing 2 nucleotide mismatching), and reads containing ambiguous bases were removed. We obtained a total of 1,809,594 high-quality sequences with an average of 37,940 for each sample. Rarefying may bring out some problems but is still a popular method in the study of microbial ecology (Delgado-Baquerizo et al. 2018). To avoid potential bias caused by sequencing depth, all sequence data were rarefied to 17,454 sequences per sample for the downstream analyses. Operational taxonomic units (OTUs) were clustered with 97% similarity cutoff using UPARSE (version 7.1 http://drive5.com/uparse/) with a novel “greedy” algorithm that performs chimera filtering and OTU clustering simultaneously. The taxonomy of each 16S rRNA gene sequence was analyzed by the RDP Classifier algorithm (http://rdp.cme.msu.edu/) against the Silva (SSU123) 16S rRNA database using a confidence threshold of 97%.
Distribution of the methanotroph communities in situ and in the incubation experiment among different grazing intensities was evaluated by principal component analysis (PCA) in R using the vegan package. Redundancy analysis (RDA) was also performed to identify the abiotic factors (bulk density, total C, Olsen P, and NO3−-N) that are most important in shaping active methanotroph communities in the grazed grassland soils. The neighbor-joining tree was constructed by MEGA 7 with 1000-fold bootstrap support (Kumar et al. 2016).
Statistical analysis and sequencing data deposition
Significant differences of CH4 oxidation potential, pmoA, and methanotroph-affiliated 16S rRNA genes relative abundance among different treatments were assessed by one-way analysis of variance (ANOVA) followed by Tukey’s post hoc test. All analyses were conducted by SPSS version 20 (IBM Co., Armonk, NY, USA).
The reads for the 16S rRNA genes of the in situ and incubated soil samples were deposited in the National Center for Biotechnology Information (NCBI) Sequence Read Archive (SRA) under accession number PRJNA432864.
Results
Soil physicochemical properties
The general physicochemical characteristics for the soils from different stocking rates (no grazing (CK), light grazing (G1), moderate grazing (G2), and heavy grazing (G3)) are summarized in Table 1. Soil pH was significantly lower in the G2 (6.89) soil compared with other grazed soils (G1: 8.12; G3: 7.69) and the ungrazed control (CK: 7.62). Soil bulk density showed a significant positive relationship with increasing stocking rates (r = 0.96, p < 0.001). The contents of total soil C and nutrients, including SOM, total C, total N, Olsen P, and K, were highest in the G1 soils compared with other samples. Grazing led to significantly less exchangeable NH4+-N in the grazed soils of G1, G2, and G3, compared with the ungrazed control.
Abundance and diversity of methanotrophs in situ
The abundance of methanotrophs was estimated in soil samples in situ by qPCR targeting the pmoA gene (Fig. 1a). The results showed that pmoA gene abundance ranged from 7.74 × 105 ± 2.13 × 104 copies g−1 soil in G1 soil to 6.59 × 105 ± 0.95 × 105 copies g−1 soil in G3 soil. The abundance of pmoA gene increased with increasing grazing intensities. The light grazing (G1) had a significantly lower abundance of the pmoA gene compared with the ungrazed control (CK).
Abundance of pmoA gene (a) revealed by quantitative PCR and principal component analysis (PCA) of methanotroph-affiliated 16S rRNA reads (b) by MiSeq amplicon sequencing in grazed grassland soil in situ. Bars represent standard error of triplicate samples. The different letters above the columns indicate a significant difference (p < 0.05) based on the analysis of variance with Tukey’s post hoc test
PCA showed that over 67% methanotroph variations were explained by the first two axes, with PC1 and PC2 explaining 48.9% and 17.7% of the total variance, respectively (Fig. 1b).
Methane oxidation potential and abundance of methanotrophs
The CH4 oxidation potential under a headspace of 8% (v/v) CH4 concentration ranged from 1.59 ± 0.07 to 2.25 ± 0.07 μmol g dry weight−1 day−1 for the grazed soils (Fig. 2). No significant differences in the CH4 oxidation potential were observed between 13CH4 labelled and CH4 control microcosms. The CH4 oxidation potential was as low as 0.51 ± 0.05 μmol g dry weight−1 day−1 in the ungrazed control, which was significantly lower than the grazed soils. Compared with the light and intermediate grazing, the heavy grazing significantly decreased the CH4 oxidation potential.
The community size of methane-oxidizing bacteria was not significantly different for the ungrazed control after 28 days of incubation (Fig. 3a). The copy number of pmoA genes increased significantly from 1.38 × 107 ± 3.21 × 105 at day 0 to 3.39 × 107 ± 0.5 × 106 in microcosms of the G3 soil after 28 days of incubation. Significantly higher abundances of the pmoA gene were observed in the G1 and G2 soils, representing 12- and 6-fold increases, respectively (Fig. 3a). Similar results were observed from the relative abundance of methanotrophs reads to total 16S rRNA reads by MiSeq amplicon sequencing (Fig. 3b). Methanotrophs reads showed a significant increase after 28 days of incubation in the grazed soils but not in the ungrazed control. Correlation analysis showed a strong correlation between the potential CH4 oxidation rates and pmoA quantities (r = 0.92, p < 0.01) during incubation, but not with pmoA quantities in situ (r = − 0.517, p = 0.085).
Abundances of pmoA gene by quantitative PCR (a) and relative abundance of methanotrophic reads to total 16S rRNA gene reads by MiSeq amplicon sequencing (b) in soil microcosms over an incubation period of 28 days. The error bars represent the standard errors of triplicate microcosms. The different letters above the columns indicate a significant difference (p < 0.05) based on the analysis of variance with Tukey’s post hoc test
Active methanotrophs in soils
The relative proportion of pmoA across CsCl gradients was similar among the three soils despite the differences in the intensity of grazing (Fig. 4a). The maximum relative proportion of pmoA was initially detected in the light fractions (around a buoyant density of 1.723 g mL−1) in the CH4 microcosms but shifted to the heavy fractions (around a buoyant density of 1.745 g mL−1) in the 13CH4 microcosms after the 28-day incubation (Fig. 4a). This indicated that methanotrophs grew by assimilating CH4 during active methane oxidation. By contrast, no significant shifts in the relative proportion of pmoA were observed in the CH4 and 13CH4 incubations of the ungrazed control soil (Fig. 4a). Miseq sequencing of methanotroph-affiliated 16S rRNA gene distributed across the CsCl gradient also indicated the assimilation of CH4 in the three grazed soils but not in the ungrazed control during the active methane oxidation (Fig. 4b).
Quantitative distribution of the pmoA gene based on qPCR across the entire buoyant density gradient of the DNA fractions (a) and percentage distribution of methanotroph-affiliated 16S rRNA reads by MiSeq amplicon sequencing for the heavy DNA (fraction as 4–7) (b) from soil microcosms incubated with 12CH4 or 13CH4 for 28 days. The normalized data are the ratios of the gene copy number in each DNA gradient to the maximum quantities from each treatment
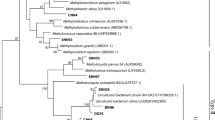
Phylogenetic analysis revealed that active methanotrophs in the 13C-DNA from the three grazed soils were most closely affiliated with Methylobacter and Methylosarcina of type Ia and Methylocystis of type II methanotrophs (Fig. 5; Supplementary fig. 1). Methylosarcina was enriched in the heavy fractions in grazed soils G1 and G2 after incubation, while Methylocystis was only enriched in G1 soil (Fig. 5).
Correlating soil properties with active methanotroph communities
Bulk density, total C, and Olsen P in combination with NO3−-N explained the highest percentage of the variance of active methanotroph communities (Fig. 6). The soil bulk density and NO3−-N content were significantly correlated with the first axis (explaining 77.3% of the total variance).
Correlation analysis showed that the soil bulk density was negatively correlated with the increase of pmoA gene abundance (r = − 0.767, p < 0.05) and methane oxidation potential (r = − 0.782, p < 0.05) during the incubation in the grazed grassland soils (Supplementary fig. 3). The ungrazed control was excluded for the correlation analysis because of no detectable growth of methanotrophs and extremely low CH4 oxidation potential in the ungrazed control soil (Figs. 2 and 3).
Discussion
Effect of grazing on methanotroph community in situ
The abundance and community composition of methanotrophic bacteria were evaluated for three field samples all exhibiting various levels of grazing. The abundance of a key gene in the methane oxidation pathway, the pmoA, was used as a proxy for the population of methanotrophs in the steppe. Compared with the ungrazed control, the abundance of pmoA gene significantly decreased in G1 soil, while increased in G3 soil (Fig. 1a). Abell et al. (2009) found that the abundance of the predominant type I methanotrophs was positively affected by long-term cattle grazing in an alpine meadow soil. In contrast, no significant change of methanotroph abundance with grazing was observed in an alpine meadow (Zheng et al. 2012). On the other hand, grazing was reported to impact the composition of the methanotrophic community in a typical grassland, while no impact was apparent in an alpine meadow (Zhou et al. 2008; Zheng et al. 2012). In this study, distribution of methanotrophs among the soils analyzed shown by PCA indicated a significant impact of grazing on the methanotroph communities (Fig. 1b). In addition, we observed a significantly lower abundance of methanotrophs in G1 soil, even though the concentrations of SOM and TN were significantly higher than others (Table 1). Some factors like the availability of N, cross-feeding, and other C sources, apart from the CH4 availability, have also been proposed to regulate the population size of methanotrophs in upland soils (Li et al. 2018; Malghani et al. 2016). In this study, the pmoA abundance was strongly correlated with SOM (r = − 0.758, p < 0.01), TN (r = − 0.584, p < 0.05), and moisture content (r = − 0.611, p < 0.05). The low abundance of methanotrophs in G1 soil was mainly ascribed to the competition between methanotrophs, which represent only a small fraction of the total bacterial community, and heterotrophs, which was markedly stimulated by the higher nutrients in G1 soil. These results indicate that long-term grazing changes not only the soil properties but also the abundance and composition of functional microbes like methanotrophs in situ.
Effect of grazing on CH4 oxidation potential
The CH4 oxidation potential measurements are useful for comparing the relative activities of the methanotroph populations within samples from different environmental conditions (Kightley et al. 1995). The potential CH4 oxidation rates were variable between the grassland samples, ranging from 0.51 ± 0.05 to 2.25 ± 0.07 μmol g−1 d−1 (Fig. 2). While the potential CH4 oxidation rates were within ranges seen in wetlands (0.17 to 80 μmol CH4 g−1 d−1) (Graef et al. 2011) and geothermal environments (1.0–141 μmol CH4 g−1 d−1) (Sharp et al. 2014) with similar elevated CH4 concentrations (> 5% v/v), they were higher than those in upland soils (up to 74.64 nmol CH4 g−1 d−1) reported by Knief et al. (2003), and in grassland soils (2.42 to 21.54 nmol CH4 g−1 d−1) reported by Kou et al. (2017) with lower concentrations of CH4 (< 0.1% v/v). The strong correlation (r = 0.92, p < 0.01) between the potential CH4 oxidation rates and pmoA quantities suggests that there are a constant activity and a consistent abundance of methanotrophs throughout the 28-day incubation period. A similar trend was observed from the relative abundance of methanotroph reads to total 16S rRNA gene reads by MiSeq amplicon sequencing (Fig. 3b), further proving the activity of methanotrophs during incubation. However, the CH4 oxidation potential was as low as 0.51 ± 0.05 μmol g−1 dry weight day−1 in the ungrazed control. In agreeing with this, no growth of methanotrophs was detected in the ungrazed control using qPCR of pmoA genes and relative abundance of methanotroph reads to total 16S rRNA gene reads by MiSeq amplicon sequencing (Fig. 3). Also, methanotrophs were not labelled by 13C-CH4 confirming our expectation of low CH4 oxidation activity in the site and negligible growth of methanotrophs (Fig. 4). It is well known that N fertilizers inhibit CH4 oxidation by ammonia, which competes with CH4 for the methane monooxygenases in methanotrophs (Bédard and Knowles 1989). Even though the affinity of MMO for CH4 is 600–1300-fold higher than its affinity for ammonia, high concentrations of ammonium (40 mg NH4+-N kg−1) are known to substantially inhibit CH4 oxidation (Alam and Jia 2012). Grazing exclusion has been reported to increase plant biomass, root biomass, root exudate, and available soil N (Wang et al. 2018). The concentrations of exchangeable NH4+-N were significantly higher in the ungrazed control than the grazed soils, which could partly explain the low CH4 oxidation in the ungrazed control soil. Ho et al. (2019) suggest that “high-affinity” methanotrophs predominate CH4 oxidation in native upland soils, while canonical methanotrophs predominate in the anthropogenic-impacted upland soils. Another possible explanation for the low CH4 oxidation potential in the ungrazed control could be due to the methanotrophs in this site predominantly composed of the putative “high-affinity” methanotrophs. Moreover, no correlation (r = − 0.517, p = 0.085) between the potential CH4 oxidation rates and pmoA abundance in situ was observed, further proving that the mere presence of pmoA genes under in situ soil conditions may not necessarily reflect the functional activity of CH4 oxidation (Nannipieri et al. 2019).
It is widely accepted that heavy grazing would cause a decline in CH4 oxidation rate (Chen et al. 2011). Previous studies have indicated that an increase in stocking rate induced a reduction in CH4 uptake (Holst et al. 2008; Wang et al. 2012). Simulating the effects of grazing management with the PaSim model, Soussana et al. (2004) have suggested that a decline in the greenhouse gas sink activity of managed steppes occurs with increased stocking intensity. In this study, heavy grazing significantly decreased the potential CH4 oxidation rate in G3 soil compared with light and intermediate grazing in G1 and G2 soils (Fig. 2). Heavy grazing could significantly increase soil bulk density and directly affect the air permeability incurred from sheep trampling (Table 1) (Ball et al. 2012; Pan et al. 2018a). Furthermore, heavy grazing would decrease aboveground plant and litter biomass and consequently increase water stress, potentially inhibiting the activities of methanotrophs indirectly (Chen et al. 2011; Cui et al. 2018; Liu et al. 2007). Heavy grazing has been previously reported to inhibit the growth of ammonia-oxidizing bacteria, reducing the nitrification activity in grazed grassland soils (Pan et al. 2018b). In contrast to these negative impacts on plants, soil properties, and microbes by heavy grazing, light grazing could lead to a greater diversity of plant species, and the dense fibrous rooting systems of plants would benefit soil organic matter formation and soil C sequestration (Reeder and Schuman 2002). N returned in animal excreta and/or modification of N uptake and C exudation by frequently defoliated plants could also promote soil fertility and enhance microbial activities (Le Roux et al. 2003; Luo et al. 2019; Pan et al. 2018a; Zhou et al. 2010; Zhu et al. 2018). This study showed that grazing alters soil functional traits with light and intermediate grazing stimulating the growth and activity of methanotrophs, while heavy grazing significantly decreased the abundance of methanotrophs and the methane oxidation potential.
Active methanotrophs in grazed soils
Overall, the methanotroph communities present before and after incubation were similar (Supplementary fig. 1). Communities of the active 13C-labelled methanotrophs were more diverse in the light and intermediate grazed soils than in the heavily grazed soil (Fig. 5). The active methanotroph community included members of Methylobacter and Methylosarcina (type I methanotrophs) and Methylocystis (type II methanotrophs) (Supplementary fig. 2). Most of the active methanotrophs (> 90%) were closely related to the type I methanotroph, Methylobacter luteus (Fig. 5), a species originally isolated from a sewage (Bowman et al. 1993; Romanovskaia et al. 1978). Methylobacter-related type I methanotrophs have been reported to be responsible for the majority of methane oxidation in a long-term grazing site in Austria and also in six grazed grassland soils across New Zealand (Abell et al. 2009; Di et al. 2010). The mean annual temperature of 0.3 °C and maximum monthly mean temperatures of 19 °C in the studied field favored the growth of Methylobacter-related type I methanotrophs, as Methylobacter species have been reported to prefer cold environments such as the active layer of Arctic permafrost (Liebner et al. 2009), high Arctic wetlands (Graef et al. 2011), lake sediments (He et al. 2012), plateau wetlands (Deng and Dumont 2016), and rice fields from cold regions (Sultana et al. 2019). No active methanotrophs in this study were affiliated with the proposed atmospheric methane-oxidizing lineages USCα and USCγ (Knief et al. 2003). Moreover, Methylosarcina of type I methanotrophs, which have shown a transient ability to oxidize methane at atmospheric levels and also possible support “high-affinity” methane oxidation activity in paddy soil (Cai et al. 2016), were detected during CH4 oxidation in G1 and G2 soils. Recently, canonical methanotrophs have been suggested to predominate CH4 oxidation as high-affinity methane-oxidizers in anthropogenically-impacted upland soils (Ho et al. 2019). Furthermore, Methylocystis of type II methanotrophs were only detected in the G1 soil. The significantly higher content of SOM in the G1 soil might partly explain the result, as some Methylocystis species are known to oxidize and grow on acetate and ethanol in addition to methane (Belova et al. 2011; Im et al. 2011). Another reason for it should be the significantly higher Olsen P in the G1 soil, because the abundance of type II methanotrophs are positively related to phosphorus and adopt a competitor-ruderal lifestyle (Ho et al. 2013). It thus suggests that the diverse active methanotrophs and higher abundance of pmoA genes in the light and intermediate grazed soils enabled the significantly higher CH4 oxidation potential. These results indicate that light and intermediate grazing stimulate the growth and activity of diverse methanotrophs, while heavy grazing significantly decreases the abundance and diversity of active methanotrophs in this typical steppe.
Methanotroph activity
Animal grazing alters soil water and energy balance by reducing vegetation, increasing soil compaction, or reducing soil aeration by trampling and also soil chemical properties (e.g., pH and organic matter content), which would subsequently induce variation of microbial communities and activity (Li et al. 2019; Liu et al. 2019b; Lu et al. 2019; Pan et al. 2018c; Saggar et al. 2004; Steffens et al. 2008; Yu et al. 2018). Numerous studies have estimated the impact of grazing on either methanotroph communities or methanotroph activity (CH4 uptake), even though the abundance and composition of methanotrophs may not necessarily reflect their activity (Abell et al. 2009; Savian et al. 2014; Van den Pol-van Dasselaar et al. 1999; Zheng et al. 2012). In this study, DNA-SIP was used to link the identity and function of methanotrophs in grazed grassland soils. RDA showed that soil bulk density and NO3−-N were significantly correlated with the distribution of active methanotrophs in grazed grassland soils (Fig. 6). Methane diffusion, which is determined by the soil bulk density and moisture, is considered the limiting factor for CH4 oxidation in soil (Serrano-Silva et al. 2014; Walkiewicz et al. 2018). The single quantification of target genes may not necessarily reflect their functional activity (Nannipieri et al. 2019). In this study, no correlation (r = − 0.517, p = 0.085) between the potential CH4 oxidation rates and pmoA abundance in situ was observed. We linked the potential CH4 oxidation rates with the increase of pmoA gene abundance during the incubation. The significant correlation between the soil bulk density and the increases of pmoA gene abundance (r = − 0.767, p < 0.05), and also methane oxidation potential (r = − 0.782, p < 0.05) during the incubation in the grazed grassland soils further indicated the important impact of bulk density on the methanotroph communities and functional activity (Supplementary fig. 3). These results indicate that grazing induced variation of bulk density and soil C and N altering the abundance and communities of active methanotrophs and subsequently changes the CH4 oxidation activity.
SIP studies often require in vitro incubations and only partially reflect conditions in situ. This method may distort the relative abundance of organisms active in a particular process (Chen and Murrell 2010; McDonald et al. 2005). Fairly high CH4 concentrations, which probably do not reflect on in situ methane levels (typically atmospheric methane levels), were selected in our incubations in order to ensure the labelling of active microbial communities during the CH4 oxidation. The often occurring cross-feeding effect for SIP experiments is not a problem in this study because methanotrophs prefer to utilize CH4 as their source of C and energy even with the existence of microbial metabolites or microbial residues from labelled methanotrophs (Bao et al. 2019). No significant changes in the methanotroph communities before and after incubation were found in this study, which indicate that SIP results might largely reflect the functional process of methane oxidation under field conditions (Supplementary fig. 1). The lack of labelled methanotrophs in the ungrazed control could mainly be due to little microbial growth during incubation, as DNA-SIP relies on cell proliferation. The lowest CH4 oxidation potential rate observed in the ungrazed control further supports the negligible activity of methanotrophs, even though the activity might have resulted from the activation of dormant microbial populations rather than by their growth (Ho et al. 2015). It is worth mentioning that phospholipid fatty acid analysis (PLFA)–based SIP could ensure labelling methanotrophs at low methane concentrations, however, which would enable the detection of microbial groups but not on microbial genera or species as DNA-SIP (Ho et al. 2019).
Conclusions
Overall, this study showed that high abundance and diversity of methanotroph communities under in situ soil conditions may not necessarily reflect high functional activity of CH4 oxidation. Light and intermediate grazing stimulated the growth and activity of active methanotrophs, while heavy grazing significantly decreased the abundance and diversity of active methanotrophs. Phylogenetic analysis of the 13C-enriched DNA fractions from the DNA-SIP microcosms revealed that the active methanotrophs were dominated by the genus Methylobacter of type I. This study also showed that soil physicochemical properties, bulk density, and soil C and N are key factors determining the abundance and composition of active methanotrophs and subsequently the CH4 oxidation activity in the long-term grazed grassland soil.
References
Abell GC, Stralis-Pavese N, Sessitsch A, Bodrossy L (2009) Grazing affects methanotroph activity and diversity in an alpine meadow soil. Environ Microbiol Rep 1:457–465
Alam MS, Jia Z (2012) Inhibition of methane oxidation by nitrogenous fertilizers in a paddy soil. Front Microbiol 3:246
Ball BC, Cameron KC, Di HJ, Moore S (2012) Effects of trampling of a wet dairy pasture soil on soil porosity and on mitigation of nitrous oxide emissions by a nitrification inhibitor dicyandiamide. Soil Use Manag 28:194–201
Bao Y, Dolfing J, Wang B, Chen R, Huang M, Li Z, Lin X, Feng Y (2019) Bacterial communities involved directly or indirectly in the anaerobic degradation of cellulose. Biol Fertil Soils 55:201–211
Bédard C, Knowles R (1989) Physiology biochemistry and specific inhibitors of CH4 NH4+ and CO oxidation by methanotrophs and nitrifiers. Microbiol Mol Biol Rev 53:68–84
Belova SE, Baani M, Suzina NE, Bodelier PL, Liesack W, Dedysh SN (2011) Acetate utilization as a survival strategy of peat-inhabiting Methylocystis spp. Environ Microbiol Rep 3:36–46
Bowman JP, SLY LI, Nichols PD, Hayward AC (1993) Revised taxonomy of the methanotrophs: description of Methylobacter gen. nov. emendation of Methylococcus validation of Methylosinus and Methylocystis species and a proposal that the family Methylococcaceae includes only the group I methanotrophs. Int J Syst Evol Microbiol 43:735–753
Bridgham SD, Cadillo-Quiroz H, Keller JK, Zhuang Q (2013) Methane emissions from wetlands: biogeochemical microbial and modeling perspectives from local to global scales. Glob Chang Biol 19:1325–1346
Cai Y, Zheng Y, Bodelier PL, Conrad R, Jia Z (2016) Conventional methanotrophs are responsible for atmospheric methane oxidation in paddy soils. Nat Commun 7:11728
Chen Y, Murrell JC (2010) When metagenomics meets stable-isotope probing: progress and perspectives. Trends Microbiol 18:157–163
Chen W, Wolf B, Zheng X, Yao Z, Butterbach-Bahl K, Brüggemann N, Liu C, Han S, Han X (2011) Annual methane uptake by temperate semiarid steppes as regulated by stocking rates aboveground plant biomass and topsoil air permeability. Glob Chang Biol 17:2803–2816
Costello AM, Lidstrom ME (1999) Molecular characterization of functional and phylogenetic genes from natural populations of methanotrophs in lake sediments. Appl Environ Microbiol 65:5066–5074
Cui Y, Fang L, Guo X, Wang X, Zhang Y, Li P, Zhang X (2018) Ecoenzymatic stoichiometry and microbial nutrient limitation in rhizosphere soil in the arid area of the northern Loess Plateau China. Soil Biol Biochem 116:11–21
Daebeler A, Bodelier PL, Yan Z, Hefting MM, Jia Z, Laanbroek HJ (2014) Interactions between Thaumarchaea Nitrospira and methanotrophs modulate autotrophic nitrification in volcanic grassland soil. ISME J 8:2397–2410
Delgado-Baquerizo M, Oliverio AM, Brewer TE, Benavent-González A, Eldridge DJ, Bardgett RD, Maestre FT, Singh BK, Fierer N (2018) A global atlas of the dominant bacteria found in soil. Science 359:320–325
Deng YC, Dumont MG (2016) Identification of active aerobic methanotrophs in plateau wetlands using DNA stable isotope probing. FEMS Microbiol Lett 363:fnw168
Di HJ, Cameron KC, Shen JP, Winefield CS, O’Callaghan M, Bowatte S, He JZ (2010) Methanotroph abundance not affected by applications of animal urine and a nitrification inhibitor dicyandiamide in six grazed grassland soils. J Soils Sediments 11:432–439
Dumont MG (2014) Primers: functional marker genes for methylotrophs and methanotrophs hydrocarbon and lipid microbiology protocols. In: McGenity T, Timmis K, Nogales B (eds) Hydrocarbon and lipid microbiology protocols. Springer Protocols Handbooks. Springer, Berlin, pp 57–77
Fan LC, Shahbaz M, Ge T, Wu J, Kuzyakov Y, Dorodnikov M (2019a) To shake or not to shake: silicone tube approach for incubation studies on CH4 oxidation in submerged soils. Sci Total Environ 657:893–901.538
Fan LC, Shahbaz M, Ge T, Wu J, Dippold M, Thiel V, Kuzyakov Y, Dorodnikov M (2019b) To shake or not to shake: 13C-based evidence on anaerobic methane oxidation in paddy soil. Soil Biol Biochem 133:146–154
Graef C, Hestnes AG, Svenning MM, Frenzel P (2011) The active methanotrophic community in a wetland from the High Arctic. Environ Microbiol Rep 3:466–472
Hanson RS, Hanson TE (1996) Methanotrophic bacteria. Microbiol Mol Biol Rev 60:439–471
He R, Wooller MJ, Pohlman JW, Quensen J, Tiedje JM, Leigh MB (2012) Diversity of active aerobic methanotrophs along depth profiles of arctic and subarctic lake water column and sediments. ISME J 6:1937–1948
Ho A, Kerckhof FM, Luke C, Reim A, Krause S, Boon N, Bodelier PL (2013) Conceptualizing functional traits and ecological characteristics of methane-oxidizing bacteria as life strategies. Env Microbiol Rep 5:335–345
Ho A, van den Brink E, Reim A, Krause SM, Bodelier PL (2015) Recurrence and frequency of disturbance have cumulative effect on methanotrophic activity abundance and community structure. Front Microbiol 6:1493
Ho A, Lee HJ, Reumer M, Meima-Franke M, Raaijmakers C, Zweers H, Boer W, Putten WH, Bodelier PL (2019) Unexpected role of canonical aerobic methanotrophs in upland agricultural soils. Soil Biol Biochem 131:1–8
Holmes AJ, Costello A, Lidstrom ME, Murrell JC (1995) Evidence that participate methane monooxygenase and ammonia monooxygenase may be evolutionarily related. FEMS Microbiol Lett 132:203–208
Holst J, Liu C, Yao Z, Brüggemann N, Zheng X, Giese M, Butterbach-Bahl K (2008) Fluxes of nitrous oxide methane and carbon dioxide during freezing–thawing cycles in an Inner Mongolian steppe. Plant Soil 308:105–117
Im J, Lee SW, Yoon S, Dispirito AA, Semrau JD (2011) Characterization of a novel facultative Methylocystis species capable of growth on methane acetate and ethanol. Environ Microbiol Rep 3:174–181
ISSS Working Group RB (1998) World reference base for soil resources. In: Schulte A, Ruhiyat D (eds) Soils of tropical forest ecosystems. Springer, Berlin, Heidelberg
Jia Z, Cao W, Hernández M (2019) DNA-based stable isotope probing. In: Dumont M, Hernández García M (eds) Stable isotope probing. Methods in molecular biology. Springer Nature, New York, pp 17–29
Kightley D, Nedwell DB, Cooper MJ (1995) Capacity for methane oxidation in landfill cover soils measured in laboratory-scale soil microcosms. Appl Environ Microbiol 1:592–601
Knief C (2015) Diversity and habitat preferences of cultivated and uncultivated aerobic methanotrophic bacteria evaluated based on pmoA as molecular marker. Front Microbiol 6:1346
Knief C, Lipski A, Dunfield PF (2003) Diversity and activity of methanotrophic bacteria in different upland soils. Appl Environ Microbiol 69:6703–6714
Kolb S (2009) The quest for atmospheric methane oxidizers in forest soils. Environ Microbiol Rep 1:336–346
Kou Y, Li J, Wang Y, Li C, Tu B, Yao M, Li X (2017) Scale-dependent key drivers controlling methane oxidation potential in Chinese grassland soils. Soil Biol Biochem 111:104–114
Kumar S, Stecher G, Tamura KJ (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Le Roux X, Bardy M, Loiseau P, Louault F (2003) Stimulation of soil nitrification and denitrification by grazing in grasslands: do changes in plant species composition matter? Oecologia 137:417–425
Leriche H, LeRoux X, Gignoux J, Tuzet A, Fritz H, Abbadie L, Loreau MJ (2001) Which functional processes control the short-term effect of grazing on net primary production in grasslands? Oecologia 129:114–124
Li C, Hao X, Zhao M, Han G, Willms WD (2008) Influence of historic sheep grazing on vegetation and soil properties of a desert steppe in Inner Mongolia. Agric Ecosyst Environ 128:109–116
Li YF, Hu SD, Chen JH, Müller K, Li YC, Fu WJ, Lin ZW, Wang HL (2018) Effects of biochar application in forest ecosystems on soil properties and greenhouse gas emissions: a review. J Soils Sediments 18:546–563
Li Y, Liu H, Pan H, Zhu X, Liu C, Zhang Q, Luo Y, Di H, Xu J (2019) T4-type viruses: important impacts on shaping bacterial community along a chronosequence of 2000-year old paddy soils. Soil Biol Biochem 128:89–99
Liebner S, Rublack K, Stuehrmann T, Wagner D (2009) Diversity of aerobic methanotrophic bacteria in a permafrost active layer soil of the Lena Delta Siberia. Microb Ecol 57:25–35
Liu C, Holst J, Brüggemann N, Butterbach-Bahl K, Yao Z, Yue J, Han S, Han X, Krümmelbein J, Horn R, Zheng X (2007) Winter-grazing reduces methane uptake by soils of a typical semi-arid steppe in Inner Mongolia China. Atmos Environ 41:5948–5958
Liu H, Pan H, Hu H, Zhang Q, Liu Y, Jia Z, Xu J, Di H, Li Y (2019a) Archaeal nitrification is preferentially stimulated by rice callus mineralization in a paddy soil. Plant Soil 445:39–53
Liu H, Ding Y, Zhang Q, Liu X, Xu J, Li Y, Di H (2019b) Heterotrophic nitrification and denitrification are the main sources of nitrous oxide in two paddy soils. Plant Soil 445:55–69
Lu XH, Li YF, Wang HL, Singh BP, Hu SD, Luo Y, Li JW, Xiao YH, Cai XQ, Li YC (2019) Responses of soil greenhouse gas emissions to different application rates of biochar in a subtropical Chinese chestnut plantation. Agric For Meteorol 271:168–179
Lüke C, Frenzel PJ (2011) Potential of pmoA amplicon pyrosequencing for methanotroph diversity studies. Appl Environ Microbiol 7:6305–6309
Luo Y, Zhu Z, Liu S, Peng P, Xu J, Brookes P, Ge T, Wu J (2019) Nitrogen fertilization increases rice rhizodeposition and its stabilization in soil aggregates and the humus fraction. Plant Soil 445:125–135
Ma L, Zhong M, Zhu Y, Yang H, Johnson DA, Rong Y (2018) Annual methane budgets of sheep grazing systems were regulated by grazing intensities in the temperate continental steppe: a two-year case study. Atmos Environ 174:66–75
Malghani S, Reim A, von Fischer J, Conrad R, Kuebler K, Trumbore SE (2016) Soil methanotroph abundance and community composition are not influenced by substrate availability in laboratory incubations. Soil Biol Biochem 101:184–194
McDonald R, Radajewski S, Murrell JC (2005) Stable isotope probing of nucleic acids in methanotrophs and methylotrophs: a review. Org Geochem 36:779–787
Nan Z (2005) The grassland farming system and sustainable agricultural development in China. Jpn J Grassl Sci 51:15–19
Nannipieri P, Penton CR, Purahong W, Schloter M, van Elsas JD (2019) Recommendations for soil microbiome analyses. Biol Fertil Soils 55:765–766
Pachauri RK, Allen MR, Barros VR, Broome J, Cramer W, Christ R, Church JA, Clarke L, Dahe Q, Dasgupta P (2014) Climate change 2014: synthesis report. Contribution of Working Groups I II and III to the fifth assessment report of the Intergovernmental Panel on Climate Change. IPCC, Geneva
Pan H, Liu H, Liu Y, Zhang Q, Luo Y, Liu X, Liu Y, Xu J, Di H, Li Y (2018a) Understanding the relationships between grazing intensity and the distribution of nitrifying communities in grassland soils. Sci Total Environ 634:1157–1164
Pan H, Xie K, Zhang Q, Jia Z, Xu J, Di H, Li Y (2018b) Archaea and bacteria respectively dominate nitrification in lightly and heavily grazed soil in a grassland system. Biol Fertil Soils 54:41–54
Pan H, Ying S, Li Y, Zeng L, Zhang Q, Xu J, Di H (2018c) Microbial pathways for nitrous oxide emissions from sheep urine and dung in a typical steppe grassland. Biol Fertil Soils 54:717–730
Pansu M, Gautheyrou J (2007) Handbook of soil analysis: mineralogical organic and inorganic methods. Springer, Heidelberg
Reeder JD, Schuman GE (2002) Influence of livestock grazing on C sequestration in semi-arid mixed-grass and short-grass rangelands. Environ Pollut 116:457–463
Romanovskaia V, Malashenko I, Bogachenko V (1978) Refinement of the diagnosis of the genera and species of methane-using bacteria. Mikrobiologiia 47:120–130
Saggar S, Bolan N, Bhandral R, Hedley C, Luo JJ (2004) A review of emissions of methane ammonia and nitrous oxide from animal excreta deposition and farm effluent application in grazed pastures. New Zeal J Agres 47:513–544
Savian JV, Neto AB, David DB, Bremm C, Schons RM, Genro TC, Amaral GA, Gere J, McManus CM, Bayer C, de Faccio Carvalho PC (2014) Grazing intensity and stocking methods on animal production and methane emission by grazing sheep: implications for integrated crop–livestock system. Agric Ecosyst Environ 190:112–119
Schöler A, Jacquiod S, Vestergaard G, Schulz S, Schloter M (2017) Analysis of soil microbial communities based on amplicon sequencing of marker genes. Biol Fertil Soils 53:485–498
Serrano-Silva N, Sarria-Guzmán Y, Dendooven L, Luna-Guido MJ (2014) Methanogenesis and methanotrophy in soil: a review. Pedosphere 24:291–307
Sharp CE, Martinez-Lorenzo A, Brady AL, Grasby SE, Dunfield PF (2014) Methanotrophic bacteria in warm geothermal spring sediments identified using stable-isotope probing. FEMS Microbiol Ecol 90:92–102
Shen LD, Ouyang L, Zhu Y, Trimmer M (2019) Active pathways of anaerobic methane oxidation across contrasting riverbeds. ISME J 13:752–766
Soussana JF, Loiseau P, Vuichard N, Ceschia E, Balesdent J, Chevallier T (2004) Carbon cycling and sequestration opportunities in temperate grasslands. Soil Use Manag 20:219–230
Steffens M, Kölbl A, Totsche KU, Kögel-Knabner I (2008) Grazing effects on soil chemical and physical properties in a semiarid steppe of Inner Mongolia (PR China). Geoderma 143:63–72
Stein LY, Roy R, Dunfield PF (2012) Aerobic methanotrophy and nitrification: processes and connections. In: Battista J (ed) Encyclopedia of life sciences (ELS). Wiley, Chichester, pp 1–11
Stocker T, Qin D, Plattner G, Tignor M, Allen S, Boschung J, Nauels A, Xia Y, Bex V, Midgley P (2013) IPCC 2013: summary for policymakers in climate change 2013: the physical science basis contribution of working group I to the fifth assessment report of the intergovernmental panel on climate change. Cambridge University Press Cambridge, New York
Sultana N, Zhao J, Zheng Y, Cai Y, Faheem M, Peng X, Wang W, Jia Z (2019) Stable isotope probing of active methane oxidizers in rice field soils from cold regions. Biol Fertil Soils 55:243–250
Trotsenko YA, Murrell JC (2008) Metabolic aspects of aerobic obligate methanotrophy. Adv Appl Microbiol 63:183–229
Van den Pol-van Dasselaar A, Van Beusichem M, Oenema O (1999) Effects of nitrogen input and grazing on methane fluxes of extensively and intensively managed grasslands in the Netherlands. Biol Fertil Soils 29:24–30
Veraart AJ, Steenbergh AK, Ho A, Kim SY, Bodelier PL (2015) Beyond nitrogen: the importance of phosphorus for CH4 oxidation in soils and sediments. Geoderma 259:337–346
Vestergaard G, Schulz S, Schöler A, Schloter M (2017) Making big data smart—how to use metagenomics to understand soil quality. Biol Fertil Soils 53:479–484
Walkiewicz A, Brzezińska M, Bieganowski A (2018) Methanotrophs are favored under hypoxia in ammonium-fertilized soils. Biol Fertil Soils 54:861–870
Wang CJ, Tang SM, Wilkes A, Jiang YY, Han GD, Huang D (2012) Effect of stocking rate on soil-atmosphere CH4 flux during spring freeze-thaw cycles in a northern desert steppe China. PLoS One 7:e36794
Wang L, Gan Y, Wiesmeier M, Zhao G, Zhang R, Han G, Siddique K, Hou F (2018) Grazing exclusion—an effective approach for naturally restoring degraded grasslands in northern China. Land Degrad Dev 29:4439–4456
Wei XM, Zhu ZK, Wei L, Wu JS, Ge TD (2019) Biogeochemical cycles of key elements in the paddy-rice rhizosphere: microbial mechanisms and coupling processes. Rhizosphere 10:100145
Yu Z, Chen L, Pan S, Li Y, Kuzyakov Y, Xu J, Brookes PC, Luo Y (2018) Feedstock determines biochar-induced soil priming effects by stimulating the activity of specific microorganisms. Eur J Soil Sci 69:521–534
Zhang Q, Li Y, He Y, Liu H, Dumont MG, Brookes PC, Xu J (2019) Nitrosospira cluster 3-like bacterial ammonia oxidizers and Nitrospira-like nitrite oxidizers dominate nitrification activity in acidic terrace paddy soils. Soil Biol Biochem 131:229–237
Zheng Y, Yang W, Sun X, Wang SP, Rui YC, Luo CY, Guo LD (2012) Methanotrophic community structure and activity under warming and grazing of alpine meadow on the Tibetan plateau. Appl Microbiol Biotechnol 93:2193–2203
Zheng Y, Huang R, Wang BZ, Bodelier PE, Jia ZJ (2014) Competitive interactions between methane- and ammonia-oxidizing bacteria modulate carbon and nitrogen cycling in paddy soil. Biogeosciences 11:3353–3368
Zhou XQ, Wang YF, Huang XZ, Tian JQ, Hao YB (2008) Effect of grazing intensities on the activity and community structure of methane-oxidizing bacteria of grassland soil in Inner Mongolia. Nutr Cycl Agroecosys 80:145–152
Zhou X, Wang J, Hao Y, Wang Y (2010) Intermediate grazing intensities by sheep increase soil bacterial diversities in an Inner Mongolian steppe. Biol Fertil Soils 46:817–824
Zhu Z, Ge T, Liu S, Hu Y, Ye R, Xiao M, Tong C, Kuzyakov Y, Wu J (2018) Rice rhizodeposits affect organic matter priming in paddy soil: the role of N fertilization and plant growth for enzyme activities CO2 and CH4 emissions. Soil Biol Biochem 116:369–377
Acknowledgments
We thank the Inner Mongolia Grassland Ecosystem Research Station, Institute of Botany, Chinese Academy of Sciences for assistance in soil sampling.
Funding
This work was financially supported by the Natural Science Foundation of China (41671249 and 41721001), Fundamental Research Funds for the Central Universities (2019QNA6011), and National Key Basic Research Program of China (2014CB138801). Yong Li extends his thanks to the Pao Yu-Kong and Pao Zhao-Long Scholarships for financial support.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PPTX 533 kb)
Rights and permissions
About this article
Cite this article
Li, Y., Liu, Y., Pan, H. et al. Impact of grazing on shaping abundance and composition of active methanotrophs and methane oxidation activity in a grassland soil. Biol Fertil Soils 56, 799–810 (2020). https://doi.org/10.1007/s00374-020-01461-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00374-020-01461-0