Abstract
The aim of this study was to detect the association between the microRNA-10b (miR-10b) expression level and prognosis in glioma patients. Quantitative real-time polymerase chain reaction (qRT-PCR) analysis was used to measure the expression of miR-10b levels in different-grade glioma tissues and normal brain tissues. The relationship between miR-10b expression levels and clinical pathological characteristics was statistically analyzed. The influence of miR-10b on survival of glioma patients was also analyzed. As a result, miR-10b expression levels in glioma tissues were significantly increased compared to those of normal brain tissues (p < 0.001). And the increased expression levels were associated with the advanced glioma grade (p < 0.001) and larger tumor size (p < 0.001). Moreover, the results of the Kaplan–Meier survival curve indicated that overall survival (OS) and progression-free survival (PFS) were significantly poorer in the high-expression-level group than those in the low-expression-level group (p < 0.001). Finally, the results of the multivariate Cox regression model indicated that the miR-10b expression level was an independent prognostic factor for glioma patients. Taken together, these findings offer convincing evidence that increased miR-10b expression may be an independent marker to predict poor prognosis in patients with gliomas.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Human gliomas are the most common primary intracranial tumors worldwide [1]. Approximately 20,000 new cases of gliomas are diagnosed in the USA every year [2]. Gliomas include four pathologic grades based on the degree of malignancy according to World Health Organization (WHO) classification: pilocytic astrocytoma (PA, WHO grade I), diffuse astrocytoma (DA, WHO grade II), anaplastic astrocytoma (AA, WHO grade III), and glioblastoma (GBM, WHO grade IV) [3]. Despite considerable progression having been made in the diagnosis and treatment of gliomas, the prognosis of glioma patients, especially high-grade glioma patients, remains poor because gliomas are highly proliferative and invasive [4]. Therefore, it is indispensable to further explore the molecular mechanisms of gliomas in order to improve the therapeutic effect.
MicroRNAs are a class of short, noncoding RNA molecules that regulate the expression of multiple target genes, which are involved in modulation of cell apoptosis, metabolism, differentiation, and stem cell maintenance [5–7]. Accumulated evidences show that microRNAs may take part in tumor angiogenesis, invasion, and metastasis. They act as oncogenes or anti-oncogenes depending on the their target genes, which may provide insight into the diagnosis and prognosis of human cancer [8, 9]. The abnormal expression of many microRNAs has been detected in human gliomas. Besides, they may affect the prognosis of glioma patients. For example, the downregulation of miR-124 [10], miR-200b [11], and miR-205 [12] and the upregulation of miR-224 [13], miR-650 [14], and miR-17 [15] are related to poorer prognosis of glioma patients. All these researches demonstrate that the microRNA expression pattern might be used as an effective marker for analyzing molecular pathogenesis and classifying subtypes of human gliomas. A recent study has revealed that miR-128 also appears to be one of the most promising treatment targets in human gliomas by suppressing the proliferation and enhancing differentiation of glioma-initiating cells [16].
MicroRNA-10b (miR-10b) is located on human chromosome 2q31 [17], which is found to be associated with tumor invasive potential in many types of cancers, such as in hepatic cancer [18], pancreatic cancer [19], and esophageal cancer [20]. Meanwhile, miR-10b was previously reported to be upregulated in glioblastoma tissues [21]. Researches about the correlation between elevated miR-10b expression and poor prognosis in gastric cancer [22], renal cancer [23], colorectal cancer [24], and breast cancer [25] have been recently published. But the studies about the correlation between the miR-10b expression level and the clinicopathologic characteristics and prognosis of glioma patients were not found in the literatures. The aim of the present study was to investigate the clinical significance of miR-10b expression in gliomas.
Methods and materials
Patients and tissue samples
This study was approved by the Research Ethics Committee of Yuquan Hospital, Tsinghua University. Written informed consent was obtained from all of the patients. All specimens were handled and made anonymous according to the ethical and legal standards.
Ninety-five fresh glioma tissue samples for quantitative real-time polymerase chain reaction (qRT-PCR) were obtained from the Department of Neurosurgery, Yuquan Hospital, Tsinghua University, between 2004 and 2013. None of the patients had received chemotherapy or radiotherapy prior to surgery. Resected specimens were snap frozen in liquid nitrogen and stored at −80 °C for qRT-PCR. The histopathology of all specimens was diagnosed by the Department of Pathology, Yuquan Hospital, Tsinghua University. A total of 47 males and 48 females were enrolled in this research, and the median age was 47 years (range, 11–76 years). According to WHO grade criteria, 28 of the 95 glioma tissues were classified as low grade (11 WHO grade I and 17 WHO grade II), and 67 were classified as high-grade gliomas (29 WHO grade III and 38 WHO grade IV). Twenty normal brain tissue samples used as controls were taken from patients who underwent surgical operation for cerebral trauma and cerebral hemorrhage.
Clinical follow-up was available for all patients, and the median follow-up was 31 months (10–60 months). During the follow-up, overall survival time was measured from diagnosis to death or last follow-up. Patients’ clinicopathological information, such as sex, age, Karnofsky performance status score (KPS), and WHO grade, is indicated in Table 1. Follow-up was conducted by telephone visit. The patients who died of diseases not directly related to their gliomas or due to unexpected events were excluded from this study.
RNA isolation and quantitative real-time PCR for miRNA detection
Total RNA was isolated from tissues using TRIzol reagent (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s instructions. RNA concentration and purity were measured using the NanoDrop ND-1000 spectrophotometer (NanoDrop Technologies, Houston, TX, USA). The isolated RNA was reverse transcribed and amplified using the TaqMan MicroRNA Assays (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s protocol. Real-time PCR was performed using an Applied Biosystems 7500 real-time PCR system according to the manufacturer’s instructions. The primers for miR-10b and U6B internal control were synthesized by Sagon Biotech (Shanghai). Relative quantification of target microRNA expression was analyzed using the comparative cycle threshold (CT) method. Each sample was examined in triplicate, and the relative quantities of the PCR products produced were normalized to U6B which served as internal control.
Statistical analysis
All computations were carried out using the Statistical Package of SPSS version 13.0 for Windows (SPSS Inc, IL, USA) and GraphPad Prism 5 (GraphPad Software Inc, CA, USA). Data were expressed as mean ± SD (standard deviation), and independent sample t test and the chi-square test were used to analyze the relationship between miR-10b expression and the clinicopathological parameters. The life curves of the patients were determined according to the Kaplan–Meier method. The Cox proportional hazard model was estimated using the multivariate Cox regression analysis to evaluate the effect of multiple independent prognostic factors on survival outcomes. Differences were considered statistically significant when p < 0.05.
Results
miR-10b upregulation in human glioma tissues
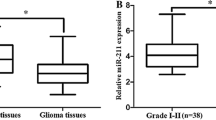
The expression levels of miR-10b were detected in 95 glioma and 20 normal brain tissues. As shown in Fig. 1, the expression level of miR-10b was significantly increased in tumor tissues compared with normal brain tissues (66.62 ± 5.44 vs 1.24 ± 0.06, p < 0.001). Importantly, we found that the expression levels of miR-10b in glioma grades I, II, III, and IV increased in a stepwise manner (9.51 ± 0.37 vs 20.29 ± 0.82, 20.29 ± 0.82 vs 45.17 ± 1.31, 45.17 ± 1.31 vs 120.20 ± 6.93, both p < 0.001). As shown in Fig. 2.
Association between miR-10b expression and clinicopathological parameters of human gliomas
We then analyzed the association between miR-10b expression and clinicopathological parameters in gliomas. Fifty-six specimens were assigned to the high-expression group, and 39 specimens were assigned to the low-expression group according to the median expression level of miR-10b. Table 2 summarizes the association between miR-10b expression and clinicopathological parameters in gliomas, indicating that the expression level of miR-10b in glioma tissues increased gradually with the advanced WHO grade (from grade I to IV) (p < 0.001). Moreover, the high expression level of miR-10b was significantly associated with larger tumor size and advanced WHO grade (both p < 0.001). No statistically significant association was found between miR-10b expression and other clinicopathological factors including gender, age, and KPS (both p > 0.05).
Relationship between miR-10b expression and overall glioma patient survival
In order to evaluate the prognostic value of miR-10b expression in human gliomas, we reviewed all the clinical information of patients in our database. The results of Kaplan–Meier survival analysis in the glioma patients with WHO grade I–IV are shown in Figs. 3 and 4. According to the log-rank test, the overall survival (OS) rate and progression-free survival (PFS) rate of patients with high miR-10b expression were significantly lower than those with low miR-10b expression (p < 0.001).
In multivariate analysis, the Cox proportional hazard model involving the expression level of miR-10b and various clinical parameters identified miR-10b upregulation (p < 0.001) as an independent prognostic factor for glioma patients. Statistical values of the expression of miR-10b and other clinical parameters are indicated in Table 3, which revealed that the miR-10b expression level (hazard ratio HR = 4.71, p < 0.001) and WHO grade (HR = 3.24, p < 0.001) were independently associated with the OS and PFS.
Discussion
Biomarker screening is an emerging field for gliomas, and several clinicopathologic characteristics have been used as prognostic factors for gliomas, such as histopathologic grades and KPS score. Unfortunately, it is difficult to evaluate the prognosis of glioma patients exactly in clinical practice because of patients’ heterogeneity [26]. Therefore, it is of great significance to identify specific prognostic indicators that can predict the clinical outcomes.
Abnormal expressions of microRNAs are involved in the pathogenesis of human gliomas. Prior to the present research, increased expression of miR-10b in malignant gliomas was confirmed by many researchers [21, 27]. Consistent with these previous studies, in the present study, we detected the miR-10b expression level in different WHO grades of glioma tissues. The results indicated that the expression levels of miR-10b were much higher in high-grade WHO glioma than those in low-grade WHO glioma and nonneoplastic brain tissues. Meanwhile, we also found the expression level of miR-10b was increased gradually from WHO grade I to grade IV gliomas. These findings indicated that increased miR-10b expression might play an important part in the progression of human gliomas. To verify this possibility, we analyzed the association between miR-10b expression and the clinicopathologic features of human gliomas. The results showed increased expression of miR-10b in glioma tissues was significantly correlated with an advanced pathological grade and a larger tumor.
Regarding its clinical significance, the results of Kaplan–Meier analysis showed that glioma patients with a high miR-10b expression level had distinctly shorter OS and PFS than those with low miR-10b expression. Furthermore, multivariate analyses showed that the miR-10b expression level was an independent prognostic parameter of glioma patients. Therefore, we present evidence here which shows that miR-10b is a potential prognostic indicator of human gliomas.
Upregulation of the expression of miR-10b may have potential value for predicting the prognosis of glioma patients, suggesting that miR-10b is an important candidate oncogene. Increasing evidences suggest miR-10b may play an important role in glioma development and progression, but the mechanism that lies behind may be complex. It is now accepted that miR-10b may act as an oncogene according to the roles of their target genes. For example, miR-10b can regulate the expression of metastasis-related genes, such as RhoC and urokinase-type plasminogen activator [27]. RhoC has been verified as an especially important player in the metastatic spread of gastric cancer [28]. The effect of miR-10b in cancer metastasis has been also correlated with the metastasis-promoting transcription factor Twist, which plays a vital role in tumor metastasis [17]. In addition, it is reported that miR-10b modulates breast cancer metastasis by targeting E-cadherin [29]. Similarly, miR-10b downregulates syndecan-1, which induces the expressions of transcription factors (AML1/RUNX1 and ATF2), resulting in cell cycle progression and metastasis [30]. In gliomas, miR-10b regulates several key cell cycle inhibitors and proapoptotic genes such as Bim, TFAP2C, p16, and p21 [31], and induces glioma cell invasion by modulating MMP-14 and uPAR expression through the tumor suppressor HOXD10 [32], which can modulate many genes that promote invasion, migration, and tumor progression [33]. So in our study, the increased expression of miR-10b was significantly more common in larger glioma tissues.
In conclusion, we proved that the increased expression of miR-10b might be associated with advanced tumor grade and poor prognosis in patients with gliomas, and miR-10b might function as a novel and valuable marker for predicting the clinical outcomes of patients with gliomas.
References
Omuro A, DeAngelis LM. Glioblastoma and other malignant gliomas: a clinical review. JAMA. 2013;310:1842–50.
Davis FG, McCarthy BJ. Current epidemiological trends and surveillance issues in brain tumors. Expert Rev Anticancer Ther. 2001;1:395–401.
Louis DN, Ohgaki H, Wiestler OD, Cavenee WK, Burger PC, Jouvet A, et al. The 2007 WHO classification of tumours of the central nervous system. Acta Neuropathol. 2007;114:97–109.
Caruso C, Carcaterra M, Donato V. Role of radiotherapy for high grade gliomas management. J Neurosurg Sci. 2013;57:163–9.
Yates LA, Norbury CJ, Gilbert RJ. The long and short of microRNA. Cell. 2013;153:516–9.
Schickel R, Boyerinas B, Park SM, Peter ME. MicroRNAs: key players in the immune system, differentiation, tumorigenesis and cell death. Oncogene. 2008;27:5959–74.
Lee HK, Finniss S, Cazacu S, Bucris E, Ziv-Av A, Xiang C, et al. Mesenchymal stem cells deliver synthetic microRNA mimics to glioma cells and glioma stem cells and inhibit their cell migration and self-renewal. Oncotarget. 2013;4:346–61.
Dela CF, Matushansky I. MicroRNAs in chromosomal translocation-associated solid tumors: learning from sarcomas. Discov Med. 2011;12:307–17.
Lovat F, Valeri N, Croce CM. MicroRNAs in the pathogenesis of cancer. Semin Oncol. 2011;38:724–33.
Chen T, Wang X, Li C, Xu S. Downregulation of microRNA-124 predicts poor prognosis in glioma patients. Neurol Sci. 2015;36:131–5.
Men D, Liang Y, Chen L. Decreased expression of microRNA-200b is an independent unfavorable prognostic factor for glioma patients. Cancer Epidemiol. 2014;38:152–6.
Hou S, Ding B, Li H, Wang L, Xia F, Du F, et al. Identification of microRNA-205 as a potential prognostic indicator for human glioma. J Clin Neurosci. 2013;20:933–7.
Lu S, Wang S, Geng S, Ma S, Liang Z, Jiao B. Upregulation of microRNA-224 confers a poor prognosis in glioma patients. Clin Transl Oncol. 2013;15:569–74.
Sun B, Pu B, Chu D, Chu X, Li W, Wei D. MicroRNA-650 expression in glioma is associated with prognosis of patients. J Neuro-Oncol. 2013;115:375–80.
Lu S, Wang S, Geng S, Ma S, Liang Z, Jiao B. Increased expression of microRNA-17 predicts poor prognosis in human glioma. J Biomed Biotechnol. 2012;2012:970761.
Verhaak RGW, Hoadley KA, Purdom E, Wang V, Qi Y, Wilkerson MD, et al. Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell. 2010;17:98–110.
Ma L, Teruya-Feldstein J, Weinberg RA. Tumour invasion and metastasis initiated by microRNA-10b in breast cancer. Nature. 2007;449:682–8.
Ladeiro Y, Couchy G, Balabaud C, Bioulac-Sage P, Pelletier L, Rebouissou S, et al. MicroRNA profiling in hepatocellular tumors is associated with clinical features and oncogene/tumor suppressor gene mutations. Hepatology. 2008;47:1955–63.
Bloomston M, Frankel WL, Petrocca F, Volinia S, Alder H, Hagan JP, et al. MicroRNA expression patterns to differentiate pancreatic adenocarcinoma from normal pancreas and chronic pancreatitis. JAMA. 2007;297:1901–8.
Tian Y, Luo A, Cai Y, Su Q, Ding F, Chen H, et al. MicroRNA-10b promotes migration and invasion through klf4 in human esophageal cancer cell lines. J Biol Chem. 2010;285:7986–94.
Ciafre SA, Galardi S, Mangiola A, Ferracin M, Liu CG, Sabatino G, et al. Extensive modulation of a set of microRNAs in primary glioblastoma. Biochem Biophys Res Commun. 2005;334:1351–8.
Li X, Zhang Y, Zhang Y, Ding J, Wu K, Fan D. Survival prediction of gastric cancer by a seven-microRNA signature. Gut. 2010;59:579–85.
Heinzelmann J, Henning B, Sanjmyatav J, Posorski N, Steiner T, Wunderlich H, et al. Specific miRNA signatures are associated with metastasis and poor prognosis in clear cell renal cell carcinoma. World J Urol. 2011;29:367–73.
Vickers MM, Bar J, Gorn-Hondermann I, Yarom N, Daneshmand M, Hanson JE, et al. Stage-dependent differential expression of microRNAs in colorectal cancer: potential role as markers of metastatic disease. Clin Exp Metastasis. 2012;29:123–32.
Parrella P, Barbano R, Pasculli B, Fontana A, Copetti M, Valori VM, et al. Evaluation of microRNA-10b prognostic significance in a prospective cohort of breast cancer patients. Mol Cancer. 2014;13:142.
Chu SH, Ma YB, Feng DF, Zhang H, Zhu ZA, Li ZQ, et al. Correlation of low SLC22a18 expression with poor prognosis in patients with glioma. J Clin Neurosci. 2012;19:95–8.
Sasayama T, Nishihara M, Kondoh T, Hosoda K, Kohmura E. MicroRNA-10b is overexpressed in malignant glioma and associated with tumor invasive factors, UPAR and RHOC. Int J Cancer. 2009;125:1407–13.
Liu N, Zhang G, Bi F, Pan Y, Xue Y, Shi Y, et al. RHOC is essential for the metastasis of gastric cancer. J Mol Med (Berl). 2007;85:1149–56.
Zhang J, Xu L, Yang Z, Lu H, Hu D, Li W, Zhang Z, Liu B, Ma S. MicroRNA-10b indicates a poor prognosis of non-small cell lung cancer and targets e-cadherin. Clin Transl Oncol. 2014.
Ibrahim SA, Yip GW, Stock C, Pan JW, Neubauer C, Poeter M, et al. Targeting of syndecan-1 by microRNA mir-10b promotes breast cancer cell motility and invasiveness via a Rho-GTPase- and E-cadherin-dependent mechanism. Int J Cancer. 2012;131:E884–96.
Gabriely G, Yi M, Narayan RS, Niers JM, Wurdinger T, Imitola J, et al. Human glioma growth is controlled by microRNA-10b. Cancer Res. 2011;71:3563–72.
Sun L, Yan W, Wang Y, Sun G, Luo H, Zhang J, et al. MicroRNA-10b induces glioma cell invasion by modulating MMP-14 and UPAR expression via HOXD10. Brain Res. 2011;1389:9–18.
Myers C, Charboneau A, Cheung I, Hanks D, Boudreau N. Sustained expression of homeobox d10 inhibits angiogenesis. Am J Pathol. 2002;161:2099–109.
Conflicts of interest
None
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Ji, Y., Wei, Y., Wang, J. et al. Correlation of microRNA-10b upregulation and poor prognosis in human gliomas. Tumor Biol. 36, 6249–6254 (2015). https://doi.org/10.1007/s13277-015-3310-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-015-3310-9