Abstract
The molecular and physiological work related to cold hardiness in Eucalyptus globulus and the cold-responsive dehydrins is reported. The identification and full-length gene sequence of three dehydrins of 10, 20, and 30 kDa and the comparison of their promoters regarding to potential stress and hormone response elements in E. globulus are shown. The categorization of cold-responsive proteins as dehydrin was based on the similarity in amino acid composition with selected sequenced peptides from chilling-responsive dehydrin reported for other woody plants and the increasing of gene expression level during cold acclimation. The transcript accumulation for these three dehydrin genes increased with cold acclimation and decreased with deacclimation in leaf and stem tissues, being higher in a freezing-resistant genotype of E. globulus compared to a sensitive genotype. By western blot, five dehydrin peptides were identified which increased their expression, under cold stress in leaf and stem tissues. These results provide valuable information about cold acclimation and gene regulation in eucalypt genotypes that differ in their ability to tolerate frost temperature.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Eucalyptus globulus (blue gum) is the main species for commercial forestry in temperate climates free of severe frosts and is studied around the world due to its exceptional wood quality combined with fast growth (Teulieres and Marque 2007). Considerable effort has been directed towards understanding how E. globulus responds and adapts to low temperature due to its low freezing resistance (Travert et al. 1997; Moraga et al. 2006; Costa e Silva et al. 2008; Fernández et al. 2006; Rasmussen-Poblete et al. 2008). The majority of plants in temperate climates are capable of developing a certain degree of frost resistance when exposed for some days to low but nonfreezing temperatures (Chinusamy et al. 2007). This phenomenon is called cold acclimation or hardening and involves changes in gene expression and cell metabolism (Thomashow 1999). During cold acclimation the transcriptional activation and repression of genes by low temperature are of central importance (Thomashow 1999). A large-scale EST sequencing data set is available from cold stress studies (Dhanaraj et al. 2004; Wei et al. 2005; Joosen et al. 2006; Bassett et al. 2006; Keller et al. 2009; Wisniewski et al. 2008). All these studies revealed enrichment of dehydrin ESTs in response to cold exposure under field or controlled environments.
Dehydrins/DHNs, a subgroup of the late embryogenesis abundant protein family, are commonly induced by abiotic stresses that involve cellular desiccation (Raynal et al. 1990; Close 1996; Campbell and Close 1997; Bomal et al. 2002; Kosová et al. 2007; Tunnacliffe and Wise 2007; Layton et al. 2010). Proteins immunologically related to DHNs accumulate in several woody species (birch, pistachio, blueberry, rhododendron, and peach) in autumn, reaching maximal levels in winter coinciding with their maximum freezing tolerance and their levels decreasing in spring (Arora and Wisniewski 1994; Artlip and Wisniewski 1997; Muthalif and Rowland 1994; Close et al. 1997; Golan-Goldhirsh and Shachak 1998; Rinne et al. 1999; Marian et al. 2004, Welling et al. 2004). Although, the role of DHN in freeze stress tolerance is still not fully understood, their hydrophilic properties (Close 1996) indicate they are involved in conferring tolerance to freeze desiccation. Rinne et al. (1999) suggested that dehydrins hold water necessary for α-amylase activity in birch, providing starch degradation products, and thus improving winter hardiness by protecting cells from freeze desiccation stress. Bravo et al. (2003) reported that low temperature-induced DHNs have cryoprotective or antifreeze activity in barley. Peng et al. (2008) recently supported these observations and demonstrated that a rhododendron DHN (RcDhn5) rescued enzyme activity from dehydration effects in vitro and enhanced freezing tolerance in Arabidopsis overexpressing a RcDhn5. Wisniewski et al. (1999) and Reyes et al. (2008) have demonstrated cryoprotective properties of peach and rhododendron DHNs, respectively, using a lactate dehydrogenase freeze-thaw assay and accordingly speculated that these DHNs may prevent denaturation of proteins when exposed to freeze desiccation. It has been proposed that the short amphipathic K segments of DHN polypeptides interact with solvent-exposed hydrophobic patches on proteins undergoing partial denaturation and thereby interfere with protein aggregate formation (Close 1996). Furthermore, Koag et al. (2003) suggested that DHNs may stabilize cellular membranes via conformational changes in the K segment, under stress conditions.
Based on their abundance in an expression library of Eucalyptus gunnii and their known involvement in cold response for other plant species, Keller et al. (2009) proposed a dehydrin (EguDHN1) as a candidate gene for investigating the molecular basis of cold tolerance in woody plants. The transcripts level for EguDHN1 quantified by RT-polymerase chain reaction (PCR) was significantly induced by cold treatment. Recently, Navarro et al. (2010) reported a DHN2 gene in E. gunnii as the most strongly regulated gene in a cold-acclimated EguCBF1-overexpressing line. The present study describes the molecular and physiological work related to cold hardiness in E. globulus and cold-responsive DHNs. We report the identification of three dehydrin genes from two EST libraries of E. globulus subjected to low temperature conditions. Also, we describe the expression of these DHN at the transcript and protein levels, under cold acclimation, in leaf and stem tissues of two genotypes, which differ in their cold tolerance.
Materials and methods
Plant materials and cold treatments
Four genotypes of E. globulus were obtained from Forestal Mininco S.A., Chile. These genotypes have diverse freezing resistance under field conditions, two presented a marked sensitivity to freezing temperatures (S1, S2) while R1 and R2 were resistant. The company provided 6-month-old rooted plants (35 cm of height) with at least eight expanded leaves. Ramets were put in an aislapool box covered with vermiculite to maintain substrate humidity and placed in a growth chamber with controlled temperature and photoperiod. Two growth chamber assays were established. Temperature and photoperiod were changed according to the growth chamber assay reported by Fernández et al. (2010). In the first growth chamber assay, four genotypes were studied. Forty ramets per genotype were placed in the growth chamber and subjected to four treatments. For treatment 1 nonacclimated (NA) genotypes were maintained under a 14-h day length at 20/12°C day/night temperature; on day 10, leaves were collected and the condition of the growth chamber was changed. For treatment 2, cold-acclimated genotypes before night frosts of −2°C (CABF), i.e., short days (10 h day) at 8/4°C, and leaves were collected on day 14. Treatment 3 comprised of cold acclimated after night frosts of −2°C (CAAF); after 1 week of CABF, four night frosts of −2°C (one per night) with a decrease of 2°C/h were applied, and leaves were collected after the last frost (at 8°C). For treatment 4, deacclimated (DA), genotypes were exposed to long day length (14 h day length) and 12/6°C day/night temperature for 1 week, and leaves were collected on day 28. Leaves from three ramets per genotype were sampled at NA and CAAF for RNA extraction and qRT-PCR for exploratory analysis of DHN gene expression. A night frost of −6°C during DA treatment was applied to determine the survival and percent leaf damage for each genotype to assess their tolerance to a simulated late-spring frost. Twenty DA ramets were exposed to the night frost and survival (percentage of live ramets), and leaf damage (percent necrotic area per ramet) was visually evaluated after 10 days (Table 1).
The temperature causing injury to 50% of the leaf tissues (LT50) was estimated using ramets from each genotype at the four treatments. Ramets were cut 1 cm above the soil and labeled. The aerial portions with at least five pairs of expanded leaves were introduced into steel boxes, which were sealed and exposed for 2 h in a cooling bath to −3°C, −5°C, −7°C, −9°C, and −12°C (decreasing at 2°C/h). After the freezing treatment, ramets were stored in hermetic bags and allowed to thaw overnight at 4°C. Each leaf per ramet was observed against the light, and the percentage of necrotic leaf area was estimated for each ramet. The first growth chamber experiment was arranged as a completely randomized design with 10 replications (ramets) per genotype for LT50 at each of the four treatments, and 3 replications (ramets) per genotype each at NA and CAAF treatments as an exploratory assay for dehydrin gene expression.
A second cold chamber assay was carried out to analyze gene expression from two different tissues of E. globulus (leaf and stem tissue) at four treatments NA, CABF, CAAF, DA. In this assay only two (S2, R1) of the four genotypes were selected according to their freeze tolerance after the first cold chamber assay. Thirty ramets per genotype were placed in the cold chamber. Leaves and stem from three ramets per genotype at four treatments were collected, frozen in liquid nitrogen, and stored at −80°C until RNA extraction. The second growth chamber experiment was arranged as a completely randomized design with three replications (ramets) per genotype at each NA, CABF, CAAF, and DA treatment for gene expression analysis.
Screening of cDNA library and DNA sequence analysis
The analysis of a cDNA library (available at NCBI) constructed from E. globulus seedlings subjected to low temperature conditions (Rasmussen-Poblete et al. 2008) and a 454 cDNA library constructed from two genotypes of E. globulus subjected to low temperature conditions (data not published) allowed the identification of three putative dehydrins. Three E. globulus ESTs sequences were found in the NCBI database (supplementary Table S1), two of which were named EuglDHN1 and EuglDHN10. The coding regions and the putative amino acid sequences were compared with other dehydrin sequences available in NCBI, which confirmed that these sequences indeed correspond to dehydrin genes. The composition of putative peptides was analyzed using the software Geneious pro 5.4.5 A third putative dehydrin, named EuglDHN2, was partially represented in the NCBI database by two ESTs. To complete the coding sequence, Firtschoice® RNA ligase-mediated rapid amplification of cDNA ends (RLM-RACE) PCR was used. For sequencing the three EuglDHN genes, oligo primers were designed using Primer Express 2.0 software (PE; Applied Biosystems, Foster City, CA, USA) and the sequencing was conducted under the BigDye terminator cycling condition (Macrogen, Korea). Nucleotide sequences for each dehydrin gene were aligned and assembled using the software Geneious pro 5.4.5.
DNA isolation and southern blot analysis
DNA was extracted from leaf tissues according to Doyle and Doyle (1987). Ten micrograms of genomic DNA was digested overnight with SphI and EcoRV and electrophoresed on 0.8% agarose gels. DNA was transferred to a positively charged nylon membrane (Roche, USA) by vacuum blotting using the salt transfer protocol as described by the manufacturer and crosslinked by UV irradiation. Dig-labeled probes for EuglDhn1, EuglDhn2, and EuglDhn10 were prepared by PCR using Taq DNA polymerase and incorporating DIG-11-dUDP as described in Roche Applied Science DIG System User's Guide for Filter Hybridization. The 898-bp EuglDhn1, 643-bp Eug l Dhn2, 352-bp EuglDhn10 gene probes were amplified from 50 ng DNA of the S2 genotype (E. globulus) using the PCR DIG Probe Synthesis Kit™ (Roche). The PCR conditions were: 1 cycle of 2 min at 95°C; 10 cycles of 10 s at 95°C, 30 s at 60°C, and 2 min at 72°C; and 20 cycles of 10 s at 95°C, 30 s at 60°C, 2 min plus additional 20 s for each successive cycle at 72°C, and 7 min at 72°C for final elongation. The PCR products were analyzed on a 2% agarose gel. The chemioluminescent detection was performed using the CSPD (ready to use, Roche), and the membrane was exposed to an X-ray film for 15 min.
RNA extraction, RT-PCR, and real-time qRT-PCR
RNA was extracted from leaves and stems using the CTAB method (Chang et al. 1993). Concentration of each RNA sample was measured using a NanoDrop ND-1000 spectrophotometer™. The integrity of RNA was checked by electrophoresis on 1% agarose gel. For each sample, 1 μg of total RNA was treated with DNaseI (Fermentas, Life Sciences, USA) to remove any contaminating DNA. First-strand cDNA was synthesized by reverse transcription using the High-Capacity cDNA Archive Kit (Applied Biosystems, USA) according to the manufacturer's instructions. The PCR mixture, qRT-PCRs conditions, and two housekeeping genes (UBC and a-TUB) were used as described by Fernández et al. (2010). Forward and reverse primer pairs used to amplify UBC, α-TUB, and EuglDHN genes, respectively, were: UBC-F/R: 5′-GACGGACAGGAACAAGTATGAGAC-3′ and 5′-CCCTCCACGGAATAATGATCGC-3′; α-TUB-F/R: 5′-GGAAGATGCTGCCAACAACTTTGC-3′ and 5′-AACCAGTGCCTCCACCAACAG-3′; EuglDHN1-F/R: 5′-GGCGGTGGCGGAATGCTC-3′ and 5′-TGCTGCTGCTGCTCATCGTC-3′; EuglDHN2-F/R: 5′-CCTGTGGAGAAGTGCGACGAG-3′ and 5′-GGCGGCGGAGGAGAATAGAC-3′; and EuglDHN10-F/R: 5′-ACAGCAGCAGCAGCGACAG-3′ and 5′-ATGATTAGAGTGGACGAGGAGGAC-3′. The amplicon length was between 93 and 157 bp, and the specificity was verified by checking the melting curve (60°C to 95°C) after 40 cycles. Three biological replicates for each sample were used, and three technical replicates were analyzed for each biological replicate. For each gene, PCR efficiency was determined by measuring the C T to a specific threshold for a serial dilution of bulked cDNA. All PCRs displayed efficiencies between 95% and 100%.
5′ RLM-RACE
The RLM-RACE was carried out with total RNA extracted from genotype R2 and was used to determine the transcription start points and size of EuglDHN2 mRNA. 5′ RACE was carried out using a FirstChoice RLM-RACE kit (Ambion) according to the manufacturer's instructions. The nested PCR conditions for 5′ outer PCR were with 0.4 μM gene-specific outer primer EuglDHN2-5′-Outer (5′-GCAGAGACATATACAAGCACATAGC-3′), 1.25 U of Ambion's SuperTaq polymerase, 0.4 μM 5′ RACE outer primer (5′-GCTGATGGCGATGAATGAACACTG-3′; Ambion), 1× SuperTaq PCR buffer (Ambion), 100 μM deoxynucleoside triphosphates, 1 ng∙μl−1 first-strand cDNA reaction, and H2O nuclease free to 50 μl. The PCR conditions were as follows: 4 min at 94°C, 35 cycles of 30 s at 94°C, 30 s at 60°C, 60 s at 72°C, and 1 cycle of 7 min at 72°C. The 5′ inner PCR was carried out with 0.4 μM gene-specific inner primer EuglDHN2-5′-Inner (5′-GGCGGCGGAGGAGAATAGAC-3′) and 0.4 μM 5′ RACE inner primer (5′-CGCGGATCCGAACACTGCGTTTGCTGGCTTTGATG; Ambion) using the same conditions as for the 5′ outer PCR. PCR products were observed on 2% agarose gels, purified and sequenced.
Cloning of the 5′ flanking regions
The putative promoter sequences of EuglDHN1, EuglDHN2, and EuglDHN10 were obtained using the method described in the GenomeWalker™ Kit (BD Biosciences). Genomic DNA was digested with the restriction enzymes DraI, StuI, EcoRV, and PvuII. Each digested product was ligated with the GenomeWalker adaptor. The primary PCR used the AP1 primer (provided with the kit) and EuglDHN1P1 (5′-TGTCCATCATCCCCTTCTTC-3′), EuglDHN2P1 (5′-GGGCTCGTCGCACTTCTCC-3′), or EuglDHN10P1 (5′-ATGATTAGAGTGGACGAGGAGGAC-3′) primers. The thermal cycling program was 7 cycles of 94°C for 25 s, 72°C for 3 min, then 32 cycles of 94°C for 25 s, 67°C for 3 min, followed by 67°C for 7 min. The secondary nested PCR used AP2 GenomeWalker primer and EuglDHN1P2 (5′-CGTAGCTGTCAGTTCCAGCA-3′) or EuglDHN2P2 (5′-CCTCCACCACGCACTCTTCC-3′) or EuglDHN10P2 (5′-GTCGCTACCGCCGTGAATC-3′) primers. The thermal cycling conditions were 5 cycles of 94°C for 25 s, 72°C for 3 min, 25 cycles of 94°C for 25 s, 67°C for 3 min, followed by 67°C for 7 min. The nested PCR products were purified and sequenced. In order to identify regulatory motifs that are represented among dehydrin genes, a database of previously characterized motifs was used and searched for presence of these motifs in the promoter regions of the genes. This database contained merged motifs from the PLACE (Higo et al. 1999) and plantCARE (Lescot et al. 2002) databases as well as motifs obtained from the literature. Only perfect matches to the motif and those with more than four IUPAC letters were considered.
Protein extraction, SDS-PAGE, and immunoblot analysis
Leaf proteins were extracted from 0.15 g of tissue and stem proteins from 0.20 g of tissue according to the protocol of Wang et al. (2006) standardized for high-quality protein extraction from recalcitrant plant tissues, with one additional TCA/acetone wash. Pellets were rinsed with acetone, air-dried, and solubilized in 75 μl of solubilization buffer [62.5 mM Tris–HCl, pH 6.8, 2% SDS, 5% (v:v) β-mercaptoethanol, 10% (v:v) glycerol, 0.001% (w:v) bromophenol blue] (Laemmli 1970). Samples were boiled for 7 min, cooled to room temperature, and centrifuged at 16,000×g. The protein concentration of the supernatant containing the thermostable fraction of proteins was measured using the method of Esen (1978). Ten micrograms of proteins was loaded to a 12.5% SDS-PAGE with a Mini Protean II Cell (Bio-Rad). Gels were stained with Coomassie brilliant blue R-250 or transferred onto a 0.45-μm nitrocellulose membrane (Bio-Rad) using the Mini Trans Blot electrophoretic transfer cell (Bio-Rad). Polyclonal antibody against dehydrin (kindly supplied by Dr. TJ Close, University of California, Riverside) was used at 1:500 dilution in Tris-buffered saline Tween 20 (TBST) buffer (10 mM Tris pH 8, 150 mM NaCl, Tween 20). After three washes in TBST, membranes were incubated in the secondary antibody anti-rabbit IgG-AP. Membranes were washed three times in TBST and incubated in developing buffer with NBT and BCIP to detect alkaline phosphatase activity. Membranes were photographed and analyzed with the E-capt software (Vilber Lourmat, France).
Statistical analysis
The gene expression and LT50 were subjected to analysis of variance to test the effect of cold treatments and genotypes. Before ANOVA, data were checked for normality and homogeneity of variances. Differences between the values (P < 0.05) were determined by a two-way ANOVA considering treatment and genotype. A Tukey test was used to identify those values with significant differences. SAS 9.00 was used for both analyses.
Results
Isolation and characterization of EuglDHNs
Identification of EuglDHNs was carried out from an EST library constructed from seedlings subjected to low temperature conditions. After filtering the redundant and partial sequences of 8,737 ESTs (NCBI; Rasmussen-Poblete et al. 2008) and 13,662 ESTs from our 454 cDNA library, five cDNAs representing three unique transcripts containing typical DHN domains were identified as DHN genes (supplementary Table S1, Fig. 1). 5′-RLM-RACE was used to obtain the full-length sequence of one of the partial-length cDNA clones of EuglDHN2. The sequenced DNA segment of EuglDHN1 (JN052208) is 1,594 bp long. It contains one intron of 225 bp and a coding sequence of 627 bases corresponding to 209 amino acids with a deduced molecular mass of ~20 kDa and a calculated isoelectric point of 6.52; this DHN belongs to the SK group. The sequenced DNA segment of EuglDHN2 (JN052209) is 2,130 bp long; it contains one intron of 113 bp and a coding sequence of 774 bases corresponding to 258 amino acids with a deduced molecular mass of ~30 kDa and a calculated isoelectric point of 4.82, and belongs to SK2 group. The sequenced DNA segment of EuglDHN10 (JN052210) contains an intronless open reading frame of 294 bases corresponding to 98 amino acids with a deduced molecular mass of ~11 kDa and a calculated isoelectric point of 6.87; this DHN belongs to KS group.
Alignment of the predicted protein sequences of three dehydrin genes isolated from E. globulus. Sequences were compared using ClustalW. The numbers on top of the sequences represent the corresponding amino acid residues for each DHN. A consensus sequence, comprised of all amino acid residues found in the same positions of the alignment, is shown above the sequences
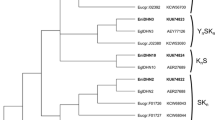
EuglDHN1 has 47% and 48% sequence similarity with two Coffea canephora dehydrins CcDH1a (ABC55670) and CcDH1b (ACJ65691), respectively (Fig. 2a). EuglDHN2 shows 51% and 53% similarity with two poplar DHNs, Populus maximowiczii dehydrin (ABS12346) and Populus alba × Populus tremula var. glandulosa dehydrin (ABH11546), respectively (Fig. 2b). And EuglDHN10 presents 62% similarity with a DHN10 from Solanum commersonii (ACJ26759, Fig. 2c).
The optimized alignment of three E. globulus dehydrins with DHNs homologues. This alignment was generated using the ClustalW program in the Geneious pro 5.4.5 software and then further optimized manually. Shaded blocks indicate identical amino acids. The rectangles with broken lines demarcate the S segments and the solid dark rectangles demarcate the K segments. a EuglDHN1 b EuglDHN2 c EuglDHN10
Southern blot data for EuglDHNs indicate only one fragment for EuglDHN1 and EuglDHN2 (Fig. 3a, b), while two bands were detected for EuglDHN10 (Fig. 3c). This suggests that EuglDHN1 and EuglDHN2 are encoded by a single gene in the E. globulus genome, as also observed for CcDH2 DHN from C. canephora (Hinniger et al. 2006) and CsDHN from Citrus sinensis (Porat et al. 2004). The KS-type dehydrin EuglDHN10 occurs as a multicopy gene similar as cor15 in the citrus genome (Porat et al. 2002).
Several potential regulatory elements involved in the regulation of gene expression during cold and osmotic stress including consensus sequences for the binding of transcription factors, such as as Myb, and several light-, heat stress-, low temperature-, gibberellin-, and salicylic acid-responsive elements were identified in the 5′ upstream region of the EuglDHN genes (supplementary Table S2, Fig. 4). One cis-acting element similar to that of the Arabidopsis thaliana dehydrin, cor15a (GGCCGACAT), that confers cold-, drought-, and ABA-regulated gene expression (Iwasaki et al. 1995) was found in the EuglDHN2 promoter region. An LT-response element (LTR; CCGAAA), previously identified in the promoter of barley (Hordeum vulgare) blt4.9 gene (Dunn et al. 1998), was found in EuglDHN10. Multiple ACGT-containing ABA-responsive elements (ABRE) were found in EuglDHN1 and EuglDHN2 (supplementary Table S2). Two putative regulatory elements responsive to photoperiod changes, Sp1 and G-box, were found in the promoter region of all three Eug1DHNs; additionally, while a Box4 element was present in the EuglDHN1, an ACE element (CTAACGTATT) was present only in EuglDHN2. Furthermore, one circadian rhythm-related element (CAANNNNATC) was observed in the EuglDHN2 (supplementary Table S2; Fig. 4).
Location of cis-elements associated with the three dehydrins promoter region. Negative numbers indicate the position of nucleotides relative to the translation start site. Grey oval TATA box, white triangle Sp1 light-responsive element, black triangle ACE light-responsive element, grey triangle Box 4 motif, inverted white triangle CE3 ABA responsiveness, inverted grey triangle G-box light-responsive element, white circle heat stress-responsive element, black circle regulatory element involved in circadian control, white diamond Myb binding site, white square CRT low temperature response motif, black square LTR low temperature response element, black trapezoid CAAT-BOX enhancer region, white rectangle TCA element involved in salicylic acid responsiveness, black rectangle ABRE motif involved in abscisic acid responsiveness, grey rectangle regulatory element involved in MeJA responsiveness, striped rectangles GARE-motif gibberellin-responsive element
Gene expression of EuglDHN1, EuglDHN2, and EuglDHN10 in response to cold acclimation
To elucidate the role of these E. globulus DHN genes in cold acclimation, EuglDHN1, EuglDHN2, and EuglDHN10 transcript abundance was measured by qRT-PCR in leaf tissues of four genotypes of blue gum used in the first growth chamber assay (Fig. 5). All DHNs showed upregulation from nonacclimated to acclimated state.
Quantification of three DHN genes in the first cold chamber assay, using two internal controls UBC and α-TUB, calibrator sample corresponds to one ramet of S1 genotype at NA. Different lowercase letters on top of the bars (mean+SE) indicate statistically significant differences between acclimation treatment and genotype evaluated with Tukey test (n = 3; P ≤ 0.05). Asterisks indicate significant differences evaluated with a Student's t test (n = 5; P ≤ 0.05) on four genotypes comparing NA treatment with CABF, CAAF, and DA treatments. a EuglDHN1. b EuglDHN2. c EuglDHN10. d Level of freezing tolerance expressed as lethal temperature (LT 50 )
Genotypes S1, S2, and R2 showed a transcript accumulation from 2- to 6.6-fold at CAAF compared to NA for EuglDHN1. The resistant genotype R1 showed the highest transcript accumulation at CAAF compared to all genotypes (Fig. 5a). Genotypes S1 and R1 showed a significant transcript accumulation after CAAF. EuglDHN2 transcripts level (Fig. 5b) increased significantly in genotypes S1 and R1 at CAAF compared to NA, while in R1 it reached the maximal accumulation at CAAF (4.3-fold) compared to NA treatment. For EuglDHN10 (Fig. 5c) all genotypes increased their gene expression at CAAF, but only resistant genotype R1 showed a significant transcript accumulation (threefold) after CAAF compared to NA and reached the highest level compared to other genotypes. Figure 5d shows the response to cold acclimation of four genotypes.
The relative quantification of three DHNs for the second growth chamber study using four treatments and two tissues (leaf and stem) in the two most contrasting genotypes in their tolerance was assessed (Fig. 6). The gene expression for EuglDHN1 showed similar results compared with those obtained in the first growth chamber study. Genotype R1 showed the highest transcript accumulation in leaf and increased 1.8-fold at CABF, 6-fold at CAAF, and decreased 0.7-fold at DA compared to NA treatment (Fig. 6a); there were significant differences between the transcript accumulation at CAAF compared to NA and DA for both genotypes. In stem, the S2 genotype showed a higher transcript accumulation compared to R1 at CABF treatment, however both genotypes showed significant differences compared to NA (Fig. 6b). Transcript accumulation reached the highest level at CAAF for both genotypes, and R1 exhibited a higher accumulation compared to S2 (Fig. 6b). For both genotypes EuglDHN1 expression decreased at DA treatment and genotype R1 had similar expression as NA. For genotype R1, EuglDHN2 expression increased significantly at CABF and CAAF compared to NA in leaf tissue. Transcript accumulation reached a higher level at CAAF compared to genotype S1; this accumulation was significantly different compared to NA and CABF and decreased at DA treatment (Fig. 6c). Sensitive genotype S2 showed significant differences at CAAF compared to NA, and the transcript accumulation decreased at DA to the level in NA. In stem tissues both genotypes showed similar upregulation at CABF, however genotype R1 had significantly higher transcript accumulation at CAAF compared to genotype S1 (Fig. 6d). In leaf tissues (Fig. 6e), the R1 genotype at CABF and CAAF treatment showed a significant higher transcript accumulation of EuglDHN10 than S2. In both genotypes the transcript level decreased at DA treatment. The EuglDHN10 expression showed a similar accumulation patterns at CABF for genotypes S2 and R1 in stem tissues (Fig. 6f), however at CAAF, R1 had a significantly higher transcript accumulation compared to S2.
Quantification of three DHN genes for the second cold chamber assay in leaf and stem tissue, using two internal controls UBC and α-TUB, calibrator sample corresponds to one ramet of S2 genotype at NA. Different lowercase letters on top of the bars (mean+SE) indicate statistically significant differences between acclimation treatment and genotype evaluated with Tukey test (n = 3; P ≤ 0.05). a EuglDHN1 leaf tissue. b EuglDHN1 stem tissue. c EuglDHN2 leaf tissue. d Level EuglDHN2 stem tissue. e EuglDHN10 leaf tissue. f EuglDHN10 stem tissue
Proteins and western blot analysis
Several quantitative changes were observed in leaf and stem protein profiles during the cold acclimation and deacclimation of R1 (cold resistant) and S2 (cold sensitive) genotypes (supplementary Fig. S1). Four of these polypeptide bands were recognized by the anti-dehydrin antibody. A 60, 45, 27, and 23-kDa DHN accumulated in the leaves and stems during cold acclimation (Fig. 7). Additionally, a 20-kDa DHN that was not visible on the SDS-PAGE profiles was detected on the immunoblots of leaf proteins but not in the stem tissues (Fig. 7a, b). Unlike on western blots, it was not possible to clearly discern DHN peptides from SDS-PAGE profiles (except for a 23-kDa DHN), as has also been observed by others (Lim et al. 1999).
Expression profile of five DHN peptides from freezing-resistant and -sensitive genotypes during nonacclimated (NA), cold acclimated before night frost (CABF), cold acclimated after night frost (CAAF), and deacclimated (DA), calibrator sample corresponds to S1 genotype at NA. a Expression profile of five DHN peptides from leaf tissue, left panel shows the immunoblotting using polyclonal antiserum raised against a DHN antibody for the consensus K segment. The right panel shows the expression estimated by image analysis densitometry. b Expression profile of five DHN peptides from stem tissue, left panel shows the immunoblotting and right panel the expression estimated by image analysis densitometry
In leaf tissues, the 23-kDa peptide showed a higher expression at NA compared to CABF and CAAF for genotype R1. In contrast to transcript accumulation where all DHNs were downregulated at DA, the 60, 27, 23, and 20-kDa DHN peptides did not show any reduced accumulation. In stem tissues for genotype R1, the 23 and 27-kDa peptides decreased in abundance at DA compared to CAAF, but genotype S2 showed no such decline.
The low-molecular weight 20-kDa DHN was not detectable in the stem tissues (Fig. 7b). In leaf tissues (Fig. 7a), however, it was apparent as a weak band at NA, CABF, and CAAF but was clearly detectable in the DA samples. Its accumulation appears to be higher at CABF and CAAF compared to NA levels (genotype S2).
Discussion
Seasonal accumulation of DHN transcripts and polypeptides in the fall and winter have been observed in several temperate woody plant species (e.g., Arora et al. 1992; Arora and Wisniewski 1994; Wisniewski et al. 1996; Levi et al. 1999; Artlip and Wisniewski 1997; Sarnighausen et al. 2004; Welling et al. 2004; Yakubov et al. 2005; Bassett et al. 2009; Wachowiak et al. 2009). Our results indicated that three EuglDHNs showed upregulation from a nonacclimated to acclimated state, while all three were downregulated during a DA treatment. In 2010, analyzing two of the four genotypes (R1 and S1) with greatest differences in freeze tolerance both under field and controlled conditions, it was possible to determine a clear clinal variation in the expression of three DHNs, during cold hardiness transitions in the leaf and stem tissues (Fig. 6) as Welling et al. (2004) reported for an SK-type DHN from Betula pendula and Artlip and Wisniewski (1997) in an evergreen peach, where transcript level was low during the autumn and started to increase in winter. The resistant genotype (R1) exhibited the maximal expression for the three genes at acclimated condition (CAAF), and differences in the expression of EuglDHN1, EuglDHN2, and EuglDHN10 showed significant differences between resistant and sensitive genotypes after CAAF. These results suggest that the difference in EuglDHNs' expression patterns in S1 and R1 are attributable to their divergent cold acclimation capacity/freeze tolerance.
Gene expression data were corroborated, in general, by the protein accumulation patterns. Different DHN proteins were observed in the acclimation–deacclimation treatments, confirming the presence of a family of dehydrins in E. globulus and their significance in plant cold hardiness. It is important to note that we first tried the protein extraction method used for Eucalyptus leaves as reported by Gaspar et al. (1997). However, it proved unsuccessful because it was not possible to see immunoreactive bands clearly on the western blots and the SDS-PAGE profiles were also of poor quality (data not shown). We therefore used the extraction protocol described by Wang et al. (2006) which yielded satisfactory results (Fig. 7). In general, five DHN peptides were increased in their expression after cold treatment reaching a relatively higher level in most cases after a night frost of −2°C (CAAF). The treatment also provided higher freeze tolerance than the other cold acclimation exposure, i.e. without the night frost (CABF). This result is consistent with our gene expression data where the highest transcript accumulation was also at CAAF (Fig. 6).
A decrease at DA of the 60-kDa DHN protein in the leaf tissues (genotype R1), 45-kDa DHN in the leaf (genotypes S2 and R1) and stem tissues (genotype R1), and of 27 and 23-kDa DHNs in stem tissues (genotype R1) was observed. However, particularly for the S2 genotype, the accumulation remained similar as CAAF even after DA, for the 60-, 45-, and 27-kDa DHNs in stem tissues, a rather unexpected observation. Although the transcript levels of EuglDHN1 and EuglDHN10 decreased after DA in the stem tissues, it was higher compared to NA treatment. These observations indicate a lag between the transcript changes and the resultant protein changes. Ohno et al. (2003) made a similar observation that a DHN mRNA in wheat increased in response to low temperature peaking at days 3 to 5, while the protein accumulation lagged behind, and that protein continued to increase even as the transcript level became barely detectable after 10 days of cold acclimation. Moreover, despite a rapid decline in the transcript level of this wheat dehydrin during deacclimation, the protein remained at considerably high levels at least up to 3 days. It appears, therefore, that the 60-, 45-, and 27-kDa DHN proteins from E. globulus were relatively stable during the 7 days of DA. It is also possible that the deacclimation treatment used in the present study was, presumably, not sufficient to allow a full deacclimation (complete loss of acquired freeze tolerance during cold acclimation). We believe that in future, the simulated acclimation–deacclimation program for E. globulus should consider a longer deacclimation treatment to resolve this question.
It was not possible to associate the identities of three EuglDHNs cDNAs with the DHN peptide bands observed on immunoblots. Only a 20-kDa DHN on the SDS-PAGE could, potentially, represent the characterized EuglDHN1 based solely on the proximity of molecular mass (~20.5 kDa as deduced from the cDNA sequence); however, no confirmation was attempted to test this notion. In this regard, reports by several authors are noteworthy in that the relative migrations of some DHNs do not necessarily match the putative peptide length deduced from cDNAs due to posttranslational modifications, e.g., glycosylation (Levi et al. 1999; Rowland et al. 2004).
It has been proposed that DHNs, via conformational changes in their K segment, may stabilize cellular membranes under stress conditions (Koag et al. 2003) and that they are highly evolved proteins, selected to maintain high configurational flexibility and to resist unspecific collapse and aggregation (Mouillon et al. 2006; Koag et al. 2009). The acidic nature and the absence of the glycine-rich repeat in EuglDHN2 suggest that it belongs to the so-called acidic dehydrin subfamily (Danyluk et al. 1998). It has been shown in wheat to preferentially associate with the plasma membranes in the sensitive vascular transition area where freeze-induced dehydration is likely to be most severe. Danyluk et al. (1998) proposed that this acidic DHN might play a role in preventing destabilization of the plasma membrane under dehydrative conditions. Analysis of the regulatory elements in the promoter region of EuglDHN2 (supplementary Table S2) suggests that it could be involved in the freeze- and/or dehydration-tolerance mechanisms. Furthermore, the hydrophilic nature of EuglDHN2 and EuglDHN1 polypeptides is well suited to replace water and stabilize membranes through polar interactions during dehydration.
The only specific physiological role of the KS-type DHNs identified so far is the iron-binding activity of a castor bean (Ricinus communis) DHN (Krüger et al. 2002) and a Citrus unshiu DHN, which are enriched in His residues (Hara et al. 2005). In general, His residues are rare in proteins, with their proportion being approximately 2% of the amino acid content (Ueda et al. 2003). Analysis of the EuglDHN10 peptide residues, in the present study, revealed a presence of His residues at 10.2% of the total amino acids. Moreover, EuglDNH10 also had relatively high transcript abundance in the stem tissues. EuglDNH10 protein might be involved in the iron transport in phloem-mediated long-distance transport as it has been reported for castor bean DHN (Krüger et al. 2002). This phenomenon has also been reported for Solanum sogarandinum and Solanum tuberosum, supporting the putative role of His-rich EuglDHN10 as an iron transport protein and possibly a component of protection mechanism against oxidative stress (Rorat et al. 2004) during cold stress.
The promoter analysis of EuglDHNs is consistent with their expression response. The analysis of their promoter regions suggests that the three DHNs are stress inducible. Related studies on the promoter region of a salt-induced dehydrin from Eucalyptus species revealed several stress-related motifs, such as Myb and C-box (Ishige et al. 2004), and using the dehydrin promoter and a DREB1A gene, these authors demonstrated transgenic Eucalyptus lines to have improved salt and drought tolerance. Based on these results, the three EuglDHNs merit further analysis to define their specific role(s) during cold acclimation. Welling et al. (2004) have reported the differential expression of two birch (Betula pubescens) dehydrins by photoperiod and temperature independently, while Wisniewski et al. (2006) examined the response of PpDhn2 to short days, low temperature, and water deficit, determining that this dehydrin was strongly induced by water deficit but not by low temperature or short days. Further analysis is needed to demonstrate specific stress induction of EuglDHNs in E. globulus. One approach to confirm the significance of the putative cis-acting elements in the promoters of three E. globulus dehydrin genes is to perform extensive deletion studies and gene expression analysis, under cold acclimation regimes involving controlled photoperiod and temperature changes.
References
Arora R, Wisniewski ME (1994) Cold acclimation in genetically related (sibling) deciduous and evergreen peach (Prunus persica (L.) Batsch). II. A 60-kilodalton bark protein in cold-acclimated tissues of peach are heat stable and related to the dehydrin family of proteins. Plant Physiol 105:95–101
Arora R, Wisniewski ME, Scorza R (1992) Cold acclimation in genetically related (sibling) deciduous and evergreen peach (Prunus persica [L.] Batsch). I. Seasonal changes in cold hardiness and polypeptides of bark and xylem tissues. Plant Physiol 99:1562–1568
Artlip T, Wisniewski ME (1997) Tissue-specific expression of a dehydrin gene in one-year-old ‘Rio Oso Gem’ peach trees. J Am Soc Hortic Sci 122:784–787
Bassett CL, Wisniewski ME, Artlip TS, Norelli JL, Renaut J, Farell RE Jr (2006) Global analysis of genes regulated by low temperature and photoperiod in peach bark. J Am Soc Hortic Sci 131:551–563
Bassett CL, Wisniewski ME, Artlip TS, Richart G, Norelli JL, Farrell RE Jr (2009) Comparative expression and transcript initiation of three peach dehydrin genes. Planta 230:107–118
Bomal C, van Quy Le, Tremblay FM (2002) Induction of tolerance to fast desiccation in black spruce (Picea mariana) somatic embryos: relationship between partial water loss, sugars, and dehydrins. Physiol Plant 115:523–530
Bravo LA, Gallardo J, Navarrete A, Olave N, Martínez J, Alberdi M, Close TJ, Corcuera LJ (2003) Cryoprotective activity of a cold induced dehydrin purified from barley. Physiol Plant 118:262–269
Campbell SA, Close TJ (1997) Dehydrins: genes, proteins, and associations with phenotypic traits. New Phytol 137:61–74
Chang S, Puryear J, Cairney J (1993) A simple method for isolating RNA from pine trees. Plant Mol Biol Rep 11:113–116
Chinusamy V, Zhu J, Zhu JK (2007) Cold stress regulation of gene expression in plants. Trends Plant Sci 10:444–451
Close TJ (1996) Dehydrins: emergence of a biochemical role of a family of plant dehydration proteins. Physiol Plant 97:795–803
Close TJ, Artlip TS, Callahan AM, Bassett CL, Wisniewski ME (1997) Seasonal expression of a dehydrin gene in sibling deciduous and evergreen genotypes of peach (Prunus persica (L.) Batsch). Plant Mol Biol 33:61–70
Costa e Silva F, Shvaleva A, Broetto F, Ortuño MF, Rodrigues ML, Almeida MH, Chaves MM, Pereira JS (2008) Acclimation to short-term low temperatures in two Eucalyptus globulus clones with contrasting drought resistance. Tree Physiol 29:77–86
Danyluk J, Perron A, Houde M, Limin A, Fowler B, Benhamou N, Sarhan F (1998) Accumulation of an acidic dehydrin in the vicinity of the plasma membrane during cold acclimation of wheat. Plant Cell 10:623–638
Dhanaraj AL, Slovin JP, Rowland LJ (2004) Analysis of gene expression associated with cold acclimation in blueberry floral buds using expressed sequence tags. Plant Sci 166:863–872
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure from small quantities of fresh leaf tissues. Phytochem Bull 19:11–15
Dunn MA, White AJ, Vural S, Hughes MA (1998) Identification of promoter elements in a low-temperature-responsive gene (blt4.9) from barley (Hordeum vulgare L.). Plant Mol Biol 38:551–564
Esen A (1978) A simple method for quantitative, semi-quantitative and qualitative assay of protein. Anal Biochem 80:264–273
Fernández M, Valenzuela S, Balocchi C (2006) RAPD and freezing resistance in Eucalyptus globulus. Electron J Biotechnol 9:303–309
Fernández M, Villarroel C, Balbontín C, Valenzuela S (2010) Validation of reference genes for real-time qRT-PCR normalization during cold acclimation in Eucalyptus globulus. Trees 24:1109–1116
Gaspar MM, Ferreira RB, Chaves MM, Teixeira AR (1997) Improved method for the extraction of proteins from Eucalyptus leaves. Application in leaf response to temperature. Phytochem Anal 8(279):285
Golan-Goldhirsh A, Shachak A (1998) Immunological cross-reaction between bud and bark proteins of dormant deciduous trees. Sci Hortic 73:165–173
Hara M, Fujinaga M, Kuboi T (2005) Metal binding by citrus dehydrin with histidine-rich domains. J Exp Bot 56:2695–2703
Higo K, Ugawa Y, Iwamoto M, Korenaga T (1999) Plant cis-acting regulatory DNA elements (PLACE) database: 1999. Nucleic Acids Res 27:297–300
Hinniger C, Caillet V, Michoux F, Ben Amor M, Tanksley S, Lin C, McCarthy J (2006) Isolation and characterization of cDNA encoding three dehydrins expressed during Coffea canephora (Robusta) grain development. Ann Bot 97:755–765
Ishige N, Kondo K, Furujyo A, Hibino T (2004) Genetic improvement for environmental stress resistance in Eucalyptus. In: Proc Plant and Animal Genome 12th Conf, San Diego, California, 10–14 January, http://www.intl-pag.org/12/abstracts
Iwasaki T, Yamaguchi-Shinozaki K, Shinozaki K (1995) Identification of a cis-regulatory region of a gene in Arabidopsis thaliana whose induction by dehydration is mediated by abscisic acid and requires protein synthesis. Mol Gen Genet 247:391–398
Joosen RVL, Lammers M, Balk PA, Brønnum P, Konings MC, Perks M, Stattin E, Van Wordragen MF, Van der Geest AL (2006) Correlating gene expression programs to physiological parameters and environmental conditions during cold acclimation of pine (Pinus sylvestris). Tree Physiol 26:1297–1313
Keller G, Marchal T, SanClemente H, Navarro M, Ladouce N, Wincker P, Couloux A, Teulières C, Marque C (2009) Development and functional annotation of an 11,303-EST collection from Eucalyptus for studies of cold stress. Tree Genet Genomes 5:317–327
Koag MC, Fenton RD, Wilens S, Close TJ (2003) The binding of maize DHN1 to lipid vesicles. Gain of structure and lipid specificity. Plant Physiol 131:309–316
Koag MC, Wilkens S, Fenton RD, Resnik J, Vo E, Close TJ (2009) The K-segment of maize DHN1 mediates binding to anionic phospholipid vesicles and concomitant structural changes. Plant Physiol 150:1503–1514
Kosová K, Vítámvás P, Práŝil IT (2007) The role of dehydrins in plant response to cold. Biol Plant 51:601–617
Krüger C, Berkowitz O, Stephan UW, Hell R (2002) A metal-binding member of the late embryogenesis abundant protein family transports iron in the phloem of Ricinus communis L. J Biol Chem 277:25062–25069
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Layton B, Boyd M, Tripepi M, Bitonti B, Norman M, Dollahon R, Balsamo R (2010) Dehydration-induced expression of a 31-kDa dehydrin in Polypodium polypodioides (Polypodiaceae) may enable large, reversible deformation of cell walls. Am J Bot 97:535–544
Lescot M, Déhais P, Moreau Y, De Moor B, Rouzé P, Rombauts S (2002) PlantCARE: a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res 30:325–327
Levi A, Panta GR, Parmentier CM, Muthalif MM, Arora R, Shanker S, Rowland LJ (1999) Complementary DNA cloning, sequencing and expression of an unusual dehydrin from blueberry floral buds. Physiol Plant 107:98–109
Lim CC, Krebs SL, Arora R (1999) A 25-kDa dehydrin associated with genotype- and age-dependent leaf freezing-tolerance in Rhododendron: a genetic marker for cold hardiness? Theor Appl Genet 99:912–928
Marian CO, Krebs SL, Arora R (2004) Dehydrin variability among Rhododendron species: a 25-kDa dehydrin is conserved and associated with cold acclimation across diverse species. New Phytol 161:773–780
Moraga P, Escobar R, Valenzuela S (2006) Resistance to freezing in three Eucalyptus globulus Labill subspecies. Electron J Biotechnol 9:310–314
Mouillon J-M, Gustafsson P, Harryson P (2006) Structural investigation of disordered stress proteins. Comparison of full-length dehydrins with isolated peptides of their conserved segments. Plant Physiol 141:638–650
Muthalif MM, Rowland LJ (1994) Identification of dehydrin-like proteins responsive to chilling in floral buds of blueberry (Vaccinium, section Cyanococcus). Plant Physiol 104:1439–1447
Navarro M, Ayax C, Martinez Y, Laur J, Kayal WE, Marque C, Teulieres C (2010) Two EguCBF1 genes overexpressed in Eucalyptus display a different impact on stress tolerance and plant development. Plant Biotechnol J 9:50–63
Ohno R, Takumi S, Nakamura C (2003) Kinetics of transcript and protein accumulation of a low-molecular-weight wheat LEA D-11 dehydrin in response to low temperature. J Plant Physiol 160:193–200
Peng Y, Reyes JL, Wei H, Yang Y, Karlson D, Covarrubias AA, Krebs SL, Fessehaie A, Arora R (2008) RcDhn5, a cold acclimation- responsive dehydrin from Rhododendron catawbiense rescues enzyme activity from dehydration effects in vitro and enhances freezing tolerance in RcDhn5-overexpressing Arabidopsis plants. Physiol Plant 134:583–597
Porat R, Pavoncello D, Lurie S, McCollum GT (2002) Identification of a grapefruit cDNA belonging to a unique class of citrus dehydrins and characterization of its expression patterns under temperature stress conditions. Physiol Plant 115:598–603
Porat R, Pasentsis K, Rozentzvieg D, Gerasopoulos D, Falara V, Samach A, Lurie S, Kanellis AK (2004) Isolation of a dehydrin cDNA from orange and grapefruit citrus fruit that is specifically induced by the combination of heat followed by chilling temperatures. Physiol Plant 120:256–264
Rasmussen-Poblete S, Valdés J, Gamboa M, Valenzuela P, Krauskopf E (2008) Generation and analysis of an Eucalyptus globulus cDNA library constructed from seedlings subjected to low temperature conditions. Electron J Biotechnol 11:2
Raynal M, Gaubier P, Grellet F, Delseny M (1990) Nucleotide sequence of a radish cDNA clone coding for a late embryogenesis abundant (LEA) protein. Nucleic Acids Res 18:6132
Reyes JL, Campos F, Wei H, Arora R, Yang Y, Karlson DT, Covarrubias AA (2008) Functional dissection of hydrophilins during in vitro freeze protection. Plant Cell Environ 31:1781–1790
Rinne PL, Kaikuranta PL, van der Plas LH, van der Schoot C (1999) Dehydrins in cold-acclimated apices of birch (Betula pubescens Ehrh.): production, localization and potential role in rescuing enzyme function during dehydration. Planta 209:377–388
Rorat T, Grygorowicz WJ, Irzykowski W, Rey P (2004) Expression of KS-type dehydrins is primarily regulated by factors related to organ type and leaf developmental stage during vegetative growth. Planta 218:878–885
Rowland LJ, Panta GR, Mehra S, Parmentier-Line C (2004) Molecular genetic and physiological analysis of the cold-responsive dehydrins of blueberry. J Crop Improv 10:53–76
Sarnighausen E, Karlson DT, Zeng Y, Goldsvrough PB, Raghothama KG, Ashworth EN (2004) Characterization of novel YnSKn class of dehydrin-like cDNAs from cold acclimated red-osier dogwood (Cornus sericea L.) xylem. In: Arora R (ed) Adaptations and responses of woody plants to environmental stresses. The Haworth Press, Binghamton, pp 17–35
Teulieres C, Marque C (2007) Eucalyptus. Biotechnol Agric For 60(Section II):387–406
Thomashow MF (1999) Plant cold acclimation: freezing tolerance genes and regulatory mechanisms. Annu Rev Plant Physiol Plant Mol Biol 50:571–599
Travert S, Valeria L, Fourasté I, Boudet AM, Teuliéres C (1997) Enrichment in specific soluble sugars of two Eucalyptus cell suspension cultures by various treatments enhances their frost tolerance via a non colligative mechanism. Plant Physiol 114:1433–1442
Tunnacliffe A, Wise MJ (2007) The continuing conundrum of the LEA proteins. Naturwissenschaften 94:791–812
Ueda EKM, Gout PW, Morganti L (2003) Current and prospective applications of metal ion-protein binding. J Chromatogr 988:1–23
Wachowiak W, Balk P, Savolainen O (2009) Search for nucleotide diversity patterns of local adaptation in dehydrins and other cold-related candidate genes in Scots pine (Pinus sylvestris L.). Tree Genet Genomes 5:117–132
Wang W, Vignani R, Scali M, Cresti M (2006) A universal and rapid protocol for protein extraction from recalcitrant plant tissues for proteomic analysis. Electrophoresis 27:2782–2786
Wei H, Dhanaraj AL, Rowland LJ, Fu Y, Krebs SL, Arora R (2005) Comparative analysis of expressed sequence tags (ESTs) from cold-acclimated and non-acclimated leaves of Rhododendron catawbiense Michx. Planta 221:406–416
Welling A, Rinne P, Vihera-Aarnio A, Kontunen-Soppela S, Heino P, Palva ET (2004) Photoperiod and temperature differentially regulate the expression of two dehydrin genes during overwintering of birch (Betula pubescens Ehrh.). J Exp Bot 55:507–516
Wisniewski M, Close TJ, Artlip T, Arora R (1996) Seasonal patterns of dehydrins and 70-kDa heat-shock proteins in bark tissues of eight species of woody plants. Physiol Plant 96:496–505
Wisniewski ME, Webb R, Balsamo R, Close TJ, Yu-Xiao M, Griffith M (1999) Purification, immunolocalization, cryoprotective, and antifreeze activity of PCA60: a dehydrin from peach (Prunus persica). Physiol Plant 105:600–608
Wisniewski ME, Bassett C, Renaut J et al (2006) Differential regulation of two dehydrin genes from peach (Prunus persica) by photoperiod, low temperature and water deficit. Tree Physiol 26:575–584
Wisniewski M, Bassett C, Norelli J, Macarisin D, Artlip T, Gasic K, Korban S (2008) Expressed sequence tag analysis of the response of apple (Malus × domestica ‘Royal Gala’) to low temperature and water deficit. Physiol Plant 133:298–317
Yakubov B, Barazani O, Shachack A, Rowland LJ, Shoseyov O, Golan-Goldhirsh A (2005) Cloning and expression of a dehydrin-like protein from Pistacia vera L. Trees 19:224–230
Acknowledgments
The authors would like to thank Dr. Leon Bravo, Valeria Neira, Claudia Flores, and Daniela Salgado for their assistance in the establishment of growth chambers assays. The authors would also like to express their appreciation to Forestal Mininco S.A for providing the plant material, a grant from MECESUP, and a scholarship from Conicyt for the financial support to MF, including a research visit to Dr. Rajeev Arora's lab at Iowa State University. This research was supported by Genomica Forestal, grant from CORFO 05CTE04-03 and partially by Conicyt (grant no. 24100220).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by R. Sederoff
Electronic supplementary material
Below is the link to the electronic supplementary material.
Table S1
Description of three DHN sequences found in the cDNA library prepared from plants of E. globulus subjected to low temperature (nonfreezing) condition. Accession numbers of EuglDHNs nucleotide sequences are listed. The number of deduced amino acids residue, the protein pI, and the molecular weight of each EuglDHN protein are shown (DOC 31 kb)
Table S2
The collection of cis-regulatory elements in the promoter of three dehydrin genes of E. globulus (DOC 33.0 kb)
Supplementary Fig. S1
SDS-PAGE (12.5%) of protein extract from freezing-resistant and -sensitive genotypes during NA nonacclimated, CABF cold acclimated before night frost, CAAF cold acclimated after night frost, and DA deacclimated. Arrows indicate the relative migration of five peptides identified by western blot analysis. a Protein extract from leaf tissue in the upper panel and stem. b Protein extract from stem tissue in the lower panel (DOC 226 kb)
Rights and permissions
About this article
Cite this article
Fernandez, M., Valenzuela Águila, S., Arora, R. et al. Isolation and characterization of three cold acclimation-responsive dehydrin genes from Eucalyptus globulus . Tree Genetics & Genomes 8, 149–162 (2012). https://doi.org/10.1007/s11295-011-0429-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11295-011-0429-8