Abstract
Lignin is the major polluting and colouring constituent present in pulp and paper mill effluent. To degrade lignin and its derivatives, bacterial enzymes can play an important role due to stability at extreme environmental conditions. This study explored the degradation of pulp and paper mill effluent by a rod-shaped Gram-positive bacterial strain RJH-1, isolated from sludge, based on its efficiency to reduce COD, colour, AOX and lignin content. This bacterial isolate was able to grow in nitrogen-free Jensen medium. Further, RJH-1 was identified as Brevibacillus agri strain after 16 s rRNA gene sequencing. Degradation potential of this isolated bacterial strain was evaluated by batch and semi-continuous reactor study. In batch study, the isolate reduced 69 % COD, 47 % colour, 37 % lignin and 39 % AOX after 5 days whereas in control flask, 40 % COD, 26 % colour, 19 % lignin and 22 % AOX reduction was observed by the indigenous bacteria present in wastewater. During semi-continuous reactor study, it reduced 62 % COD, 37 % colour, 30 % lignin and 40 % AOX of effluent at a retention time of only 32 h whereas the reduction in control reactor was 36 % COD, 21 % colour, 18 % lignin and 29 % AOX. This study confirmed that the B. agri has the potential to degrade the lignin and reduce the colour and COD of the pulp and paper mill waste water.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Pulp and paper mills are categorised as one of the largest contributor to industrial water pollution due to release of huge amount of dark coloured effluent containing recalcitrant toxic compounds which have an adverse effect on the environment (Saraswathi and Saseetharan 2010). As per the Ministry of Environment and Forest, Government of India, the pulp and paper sector is in the “Red Category” list of 17 industries having a high polluting potential owing to its serious pollution threat. It is mandatory for pulp and paper mills to comply with the appropriate standards set by Central Pollution Control Board (CPCB 2001)
The paper manufacturing process involves three steps: pulping, bleaching and finally paper making. The effluent generated at the pulping stage called black liquor is dark brown in colour due to dissolved lignin and its degradation products, hemicelluloses, resins, acids and phenols (Berryman et al. 2004). The effluent generated at bleaching stage has toxic coloured compounds, chlorinated organic compounds, adsorbable organic halides (AOX) and other such derivatives of lignin and hemicelluloses. It is estimated that 1 t of paper generates 150 m3 of effluent which is extremely toxic in nature (Pokhrel and Viraraghavan 2004). Lignin and its derivatives that are released during delignification processes of papermaking impart brown/black appearance to the effluent. Lignin is a complex heteropolymer, composed of phenylpropanoid aryl-C3 units linked together via a variety of ether and C–C bonds. Lignin is closely associated with cellulose and covalently attached to hemicelluloses. The ether and C–C linkages present in lignin are not susceptible to hydrolytic attack, and therefore, lignin is highly resistant to breakdown (Bugg et al. 2011).
The various enzymes involved in the treatment of pulp and paper mill effluent are lignin peroxidase, manganese peroxidase and laccase. Microorganisms showing good production of these enzymes have the potential to treat the effluent (Singhal and Thakur 2009). Fungal enzymes are reported for lignin solubilisation (Bucher et al. 2004; Pant and Adholeya 2007). The use of several fungal strains such as white rot fungus Phanerochaete chrysosporium has been reported for bioremediation of pulp and paper mill effluent and lignin degradation (Gomathi et al. 2012; Driessel and Christov 2001; Diwaniyan et al. 2010)
However, the low pH requirement for an optimum activity of the fungal ligninolytic enzymes (Swamy and Ramsay 1999) and the long hydraulic retention time required for complete decolourisation (Banat et al. 1996; Swamy and Ramsay 1999) are the disadvantages of using fungi. Therefore, fungal system cannot be used when pH is high and oxygen is limited as is usually common in pulp and paper mill effluents. Thus, bacterial system can play an important role under such stringent conditions. Recently, various bacterial isolates have been reported for bioremediation of pulp and paper mill effluent. Bacillus sp. has been reported for kraft lignin degradation (Chandra et al. 2007). Besides, various strains of Azotobactor, Bacillus megaterium and Serratia marcescens can also solublise lignin (Morii et al. 1995). Lignin degradation rate of the reported bacterial strains is low when compared to fungi; thus, there is a need to search for more efficient lignin degrading bacterial strains. The aim of the present study is to isolate a bacterial strain with potential ligninolytic and decolourisation efficiency for the treatment of pulp and paper mill effluent.
2 Material and Methods
2.1 Reagents
Analytical grade reagents were used in this work. All the testing was performed according to Standard Microbiological methods for the Examination of Water and Waste water (APHA)
2.2 Collection of Sludge Samples
Sludge of wastewater treatment plants is a rich source of ligninolytic bacteria. Hence, sludge was collected in sterile polythene bags from various pulp and paper mill effluent treatment plants (ETPs) near Yamuna Nagar (Haryana). The sludge samples were stored at low temperature for isolation of ligninolytic bacteria.
2.3 Enrichment of Sludge Sample for Bacterial Isolation
Five-gram sludge was added in 100-ml mineral salt medium (MSM), which contained the following (in g l−ˡ): Na2HPO4 2.4; K2HPO4 2.0; NH4NO3 0.1; MgSO4 0.01; CaCl2 0.01; D-glucose 10.0; peptone 5.0 (Chandra et al. 2007). Trace element solution (1 ml l−ˡ) as described by Pfenning and Lippert (1966) was also added, and the pH was finally adjusted to 7.5 ± 0.1. MSM was supplemented with lignin (100–400 ppm indulin or black liquor) as carbon source and incubated for 7 days at 37 °C and 150 rpm in an incubator shaker. After 7 days, 10-ml sample was retransferred in fresh 90-ml MSM supplemented with lignin and incubated for 72 h. In the same way, three more successive transfers of enriched sample were done after every 72 h. This enriched medium was used for isolation and screening of ligninolytic bacteria.
2.4 Isolation and Screening of Ligninolytic Bacteria on Lignin MSM Agar Plates
Enriched sludge samples were diluted to 10−3, 10−4 and 10−5 times in autoclaved distilled water and were spread on lignin MSM agar plates. Plates were incubated in incubator at 35 °C for 4–7 days. Phenotypically different bacterial colonies were selected and purified by repeated sub-culturing. Pure cultures were inoculated in LB broth (Luria-Bertani broth contains in g l−1 casein enzymatic hydrolysate 10, yeast extract 5, NaCl 5) media to achieve maximum growth rate and subsequently used for selection of potential ligninolytic bacterial isolates.
2.5 Tracking of Functional Diversity of Isolates Targeting Ligninolytic Enzymes by Dye Decolourisation Assay
To select bacteria capable of producing ligninolytic enzymes (laccase, lignin peroxidase and/or manganese peroxidase), a selection protocol based on their ability to decolourise synthetic dyes also called ligninolytic indicator dyes such as Azure B, Phenol red, Ramazol Brilliant Blue and Methylene Blue was used. All the indicator dyes were filter sterilised and added to autoclaved media under aseptic conditions. These dyes are structurally similar to lignin. LB agar plates containing 0.2 g/100 ml Azure B (Archibald 1992), 0.1 g/100 ml Phenol Red, 0.2 g/100 ml (Manji and Ishihara 2004) Methylene blue and 0.4 g/100 ml Ramazol Brilliant Blue (Kiiskinen et al. 2004) were streaked with bacterial isolates and incubated at 37 °C. The plates were observed for zone of decolourisation after 24–48 h. Decolourised zone that appeared around the bacterial colony indicated the presence of ligninolytic enzyme activity in the bacterial isolate; thus, bacterial isolates which decolourised any of the dye were screened as positive for ligninolytic enzyme and selected for further waste water degradation studies (Husain 2006; Pangallo et al. 2007; Ang et al. 2011; Bandounas et al. 2011; Ponting 1999).
2.6 Wastewater Characteristics
The wastewater used throughout the study was synthetic wastewater (equivalent to pulp and paper mill waste water) and was prepared manually in the lab by adding mixture of CD and EOP stage wastewater (in the ratio of 2:1), starch and then black liquor after the filtration using coarse filter paper (~40-μm pore size). The volume of these effluents was decided in such a combination so that after dilution with water, the resulting COD, colour and AOX were approximately 500 ± 25 mg O2 l−1, 1000 ± 50 Pt-Co Unit and 15 ± 1 mg l−1, respectively. The pH of synthetic wastewater was maintained 7.0 ± 0.2 with the help of NaOH and H2SO4. Throughout the study, freshly prepared, unautoclaved wastewater was used.
2.7 Evaluation of Degradative Potential of Bacterial Isolates
Each isolate screened after dye decolourisation assay was grown for 24–48 h in LB broth. Grown culture was centrifuged and washed with 1 mM phosphate buffer of pH 7. Culture pellet was inoculated in 250-ml flask containing 100-ml freshly prepared wastewater (as described above) supplemented with nutrients containing N and P to give a ratio COD/N/P = 100:5:1 referred to as standard nutrient ratio (Reddy et al. 2005). Flasks were incubated at 37 °C and 150 rpm in an incubator shaker. Control flask (wastewater without any culture inoculation) was maintained at same experimental condition. Treated wastewater sample was withdrawn after regular time interval and analyzed for reduction of colour, COD and lignin content by following standard protocols. Lignin and colour were analyzed spectrophotometrically by taking the absorbance at 280 and 465 nm, respectively, as per the standard method for the examination of wastewater colour and lignin by UV-Visible spectrophotometer (Chandra and Abhishek 2011). For the determination of colour and lignin, the treated wastewater samples were collected, and pH was adjusted to 7.4 ± 0.2 because the colour is pH dependent. Then, the samples were centrifuged at 8000 rpm for 30 min, and supernatant was collected. Supernatant was diluted 3 times, and colour reduction and lignin degradation were determined by the methods as described by Bajpai et al. (1993) and Pearl and Benson (1940). The colour was measured as platinum cobalt unit and lignin as ppm. AOX was also measured at the end of the study as per the standard protocol (APHA 1998). Based on the results of the batch study, isolates were further selected for semi-continuous reactor study.
2.8 Biochemical Characterisation of the Bacterial Isolate
The bacterial isolates were grown on LB agar plates for 48 h at 37 ± 1 °C and examined (Buchanan and Gibbsons 1974) for their cell shape, colony appearance, motility, Gram reaction, growth at different pH, temperature and % (w/v) NaCl concentration. Catalase, oxidase, Voges-Proskauer (VP) reactions, hydrolysis of casein, starch, urea, gelatin were determined as described by Barrow and Feltham (1993) and Cowan et al. (2004). Bacterial isolates were also grown in nitrogen-free Jensen medium (composition in g l−1 sucrose 20, K2HPO4 1, CaCO3 2 , MgSO4 0.5, NaCl 0.5, FeSO4 0.1, Na2MoO4 0.005) to check their ability to grow in nitrogen-deficient conditions, since pulp and paper mill wastewater is nitrogen deficient and the source of bacterial isolate was sludge collected from effluent treatment plant treating pulp and paper mill wastewater.
2.9 Identification of the Bacterial Isolate
16 s rRNA gene sequencing is widely used to identify and classify the eubacteria (Amman et al. 1995). Target sequences are amplified using universal primers to produce 1.5-kb amplicons followed by sequencing and homology generation using ribosomal DNA database. The genomic DNA from the bacterial isolate was isolated, and amplification of the 16 s rRNA gene was performed using the universal primers (Raj et al. 2007a, b). Forward primer: 5′-AGAGTTTGATCCTGGCTCAG-3′ and reverse primer: 5′-ACGGCTACCTTGTTACGACTT-3′ in a Master cycler® Thermocycler (Eppendorf, Germany). The ~1500-bp PCR product was purified to remove unincorporated dNTPS and Primers before sequencing using PCR purification kit (Norgen Biotek, Canada). Both strands of the rDNA region amplified by PCR were sequenced by automated DNA sequencer-3037xl DNA analyzer from Applied Biosystems using BigDye® Terminator v3.1 cycle sequencing Kit (Applied Biosystems). Sequence data were aligned, and dendrogram were generated using Sequence analysis software version 5.2 from Applied bio systems. The sequences obtained for plus and minus strands were aligned using DNA Baser software before performing the bioinformatics analysis. Sequences were compared to the nonredundant NCBI database by using BLASTN, with the default settings used to find the most similar sequence and were sorted by the E score. A representative sequence of 10 most similar neighbours was aligned using CLUSTAL W2 for multiple alignments with the default settings. The multiple-alignment file was then used to create phylogram using MEGA5 software (Tamura et al. 2007). The evolutionary history was inferred by using the maximum likelihood method based on the Tamura-Nei model. The tree with the highest log likelihood (−3372.5527) is shown. Initial tree(s) for the heuristic search was obtained automatically by applying neighbour-join and BioNJ algorithms to a matrix of pair wise distances estimated using the maximum composite likelihood (MCL) approach, and then selecting the topology with superior log likelihood value. Codon positions included were 1st+2nd+3rd+noncoding. All positions containing gaps and missing data were eliminated. There were a total of 1415 positions in the final dataset. Evolutionary analyses were conducted in MEGA5.
2.10 Semi-Continuous Reactor Study
To evaluate the performance of the bacterial isolates in reactors, a semi-continuous study was designed with an objective to decrease the retention time from 96 to 24 h. A fabricated bioreactor of total capacity of 2 l was used for the study (Fig. 1). Inside the bioreactor, oxygen was provided by passing air through an air pump (DO = 1 ± 0.5 mg/l), temperature was maintained to 35 ± 2 °C and agitation (200 rpm) was given by magnetic stirrer.One-litre freshly prepared wastewater was used in each reactor for lab scale study. Culture was grown in LB media, centrifuged and added to reactor containing 1-l wastewater supplemented with nitrogen and phosphorus as per standard nutrient ratio (COD/N/P = 100:5:1) given by Reddy et al. (2005). Continuous aeration and agitation were given to the reactors to prevent settling down of the microbial biomass. Acclimatisation of the culture was done under controlled temperature, aeration and agitation rate for a week. Control reactor (without any added culture) was also maintained at the same conditions. After a week, efficiency of bacterial strains was evaluated daily in terms of reduction in colour, COD and degradation of lignin by decreasing the retention time from 96 to 24 h. The fresh wastewater was added to the reactors, and treated wastewater was removed from the reactors daily. The addition and removal of wastewater to the reactors were done twice in a day by removing and adding 1/3 and 2/3 parts after 16 and 8 h, respectively, as stated in Table 1. The RT (in h) was calculated as per the formula: RT = V/Q where V is the volume of the effluent (calculated in ml) in the reactor and Q is the flow rate (calculated in ml h−1).
3 Results and Discussion
Isolation and Screening
More than 50 bacterial isolates were isolated on lignin-MSM plates from sludge samples collected from ETP’s treating pulp and paper mill wastewater. The purified bacterial isolates were further screened based on their ability to secrete ligninolytic enzymes using dye degradation method. Dye degradation is directly related to production of ligninolytic enzymes (lignin peroxidase, Mn peroxidase and Laccase); so, the pure culture of bacterial isolates was tested for their dye degradation ability (Fig. 2). A total of 27 pure cultures of bacterial isolates were screened based on their ability to degrade either of the dyes, i.e. Azure B, Phenol Red, Methylene blue, Ramazol brilliant blue. Results are conferred in Table 2. The potential isolate RJH-1 showed positive results for RBBR and Azure B.
3.1 Evaluation of Degradative Potential of Bacterial Isolates by Batch Study
Individual bacterial isolates exhibiting positive result for ligninolytic enzyme were assessed for their ability to reduce the colour, COD and degrade lignin of the wastewater by batch mode in order to check the potential of the bacterial isolates meet the discharge limit. RJH-1 was found most potent bacterium in terms of bioremediation of wastewater. During batch study, the pH of the treated wastewater increased gradually and became alkaline (8.4 ± 0.1). Initially for 24 h, the reduction in COD, colour and lignin were insignificant w.r.t control flask due to slow growth of the bacterium, but afterwards, the rate of degradation increased significantly (Figs. 3, 4 and 5). Similar findings of slow degradation during initial 24 h were reported by EL-Hanafy et al. (2008). This might be due to slow metabolism of bacteria during initial 24 h, as bacterium adapted itself to wastewater conditions and started lignin degradation afterwards. Once the bacterium started using lignin as sole carbon source, there was a significant reduction in COD and colour. Degradation of lignin and its derivatives could be associated with removal of colour and reduction in COD since lignin and related compounds account for dark colour and high COD of wastewater (Raj et al. 2014). The study was conducted for 144 h, and the potential isolate RJH-1 reduced 69 % COD, 47 % colour and 37 % lignin after 5 days whereas the control flask (with no added culture) showed 40 % COD, 26 % colour and 19 % lignin reduction. At the end of the study, AOX was also calculated in RJH-1 treated and control flask. RJH-1 showed 39 % AOX reduction while the same was reduced by 22 % in control flask. The reduction was also observed in control flask due to the presence of indigenous microflora since the wastewater used for the study was unautoclaved.
3.2 Biochemical Characterisation of Bacterium
Microscopic studies and biochemical tests showed that (Table 3) RJH-1 is Gram-positive, rod-shaped, motile bacterium. It grows on routine media such as nutrient agar, LB and also grows well on nitrogen-free Jensen medium. The colony appears translucent. It is catalase positive and oxidase negative, does not hydrolyse starch and urea but can hydrolyse casein and gelatine. The optimum temperature for growth is 28–30 °C, and maximum temperature for growth is 40 °C. It can grow in the presence of 2 % NaCl but not in 3 % NaCl. Growth occurs at pH 5.8 but not at pH 9.0; fructose, glucose and maltose are assimilated.
3.3 Molecular Characterisation of Bacterial Isolate RJH-1
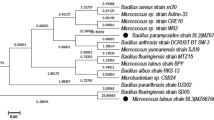
Sequence analysis of 16 s rRNA gene showed that RJH-1 is Brevibacillus agri strain. Figure 6 depicts the phylogenetic tree demonstrating the linkage distance of RJH-1 with other Brevibacillus species. The analysis showed that RJH-1 is 98 and 96 % similar to Brevibacillus agri strain DSM 6348 and Brevibacillus reuszeri, respectively. It showed 95 % similarity with Brevibacillus parabrevis strain IFO 12334, Brevibacillus choshinensis strain DSM 8552 and Brevibacillus brevis strain NBRC 15304.
3.4 Semi-Continuous Reactor Study
Reactor study was performed to evaluate the performance of the potential isolate RJH-1 (Brevibacillus agri) at different RT. The pH of the medium changed to alkaline and remained alkaline throughout the study (8.2 ± 0.4). The bacterial metabolism may be accredited for change in pH. Lignin is soluble at alkaline pH; thus, high pH aids in lignin solubilisation and degradation (Raj et al. 2014). The reduction in COD content by RJH-1-degraded sample, shown in Fig. 7, revealed that when retention time (RT) was reduced from 96 to 24 h, the percent reduction in COD was 61 ± 7 at an RT of 96 h, 61 ± 9 at an RT of 48 h, 62 ± 8 at an RT of 32 h and 43 ± 8 at an RT of 24 h. Hao and Man (2006) reported COD reduction up to 70–75 % after 3 weeks and increase to 82–88 % by subsequent aerobic treatment for 48 h by using sulphate reducing bacteria. Thus, we conclude a significant reduction in COD at a lower retention time (32 h).
The percent reduction in colour and lignin by RJH-1 was 33 ± 5, 32 ± 9, 37 ± 8, 22 ± 9 and 28 ± 5, 27 ± 5, 30 ± 8, 17 ± 6 at an RT of 96, 48, 32 and 24 h, respectively (Figs. 8 and 9). Thus, we conclude that maximum reduction by RJH-1 was obtained at 32 h RT. Singh et al. (2008) have reported potential pentachlorophenol degradation and decolourisation of pulp and paper mill effluent at 168-h incubation by a mixed culture of two bacteria. Besides, Chandra et al. (2009) have reported that PCP (pentachlorophenol) degrading bacterial strains, Bacillus cereus (ITRC-S6) and Serratia marcescens (ITRC-S7) when used for the treatment of pulp and paper mill effluent reduced colour 45–52 %, lignin 30–42 % and COD 50–60 % in 7-day incubation period. Similarly, Gupta et al. (2009) have reported that bacterial strain Aeromonas formicans reduced COD, colour and lignin in continuous reactor studies by 73, 88 and 77 %, respectively, for an 8-day retention period. Findings of Singh et al. (2011) suggest the use of tannic acid degrading bacterium Enterobacter sp. for bioremediation of pulp and paper mill effluent since the bacterium decreased the COD/BOD load to 80 %, lignin to 73 % and colour to 82 % within 16 h, using a 10 % inoculum size at 35 °C. Moreover, mixed culture of bacteria and fungus has also been acknowledged for efficient biodegradation process. Chuphal et al. (2005) mentioned significant reduction in colour (88.5 %), lignin (79.5 %), COD (87.2 %) and phenol (87.7 %) by Paecilomyces sp. and Pseudomonas syringae.
When the results of the present study were compared with previous studies, in our case the bacterial isolate reduced the COD, colour and lignin of wastewater by 62, 37 and 30 %, respectively, at 32-h retention time, i.e. the time taken to degrade lignin and reduce COD and colour was less in our case. Chandra and Bhargava (2012) and Chandra and Abhishek (2011) have also reported lignin degradation by bacteria but in the presence of glucose and peptone as additional carbon and nitrogen source, but here, we are reporting lignin degradation by the bacterial isolate without supplementing the medium with glucose and peptone so that the process remains cost-effective. Long adaptation period with lignin as sole carbon source during isolation and screening may be accredited for a considerable degradation rate at lower retention time (EL-Hanafy et al. 2007).
4 Conclusions
For the treatment of pulp and paper mill wastewater isolate, Brevibacillus agri RJH-1 can prove beneficial as it shows considerable degradation and decolourisation efficiency at a lower retention time of just 32 h. The use of nitrogen fixing Brevibacillus agri also may have added advantage of reducing the need for supplementation with nitrogenous compounds as it grows well in nitrogen-deficient conditions. Brevibacillus agri RJH-1 works efficiently using cost-effective ammonium sulphate as N and phosphoric acid as P source which are used by most of the paper mills. Thus, the ability of this bacterial isolate could be of interest for the treatment of pulp and paper mill effluent.
References
American Public Health Association. (1998). American Water Works Association and Water Environment Federation, Standard Methods for the Examination of Water and Wastewater (20th ed.). Washington: APHA.
Amman, R., Ludwig, W., & Schleifer, K. (1995). Phylogenetic identification and in situ detection of individual microbial cell without cultivation. Microbiological Reviews, 59, 143–169.
Ang, T. N., Ngoh, G. C., & Chua, A. S. (2011). A quantitative method for fungal ligninolytic enzyme screening studies. Asia-Pacific Journal of Chemical Engineering, 6, 589–595.
Archibald, F. S. (1992). A new assay for lignin-type peroxidases employing the dye Azure B. Applied and Environmental Microbiology, 58, 3110–3116.
Bajpai, P., Mehna, A., & Bajpai, P. K. (1993). Decolorization of kraft bleach plant effluent with the white rot fungus Trametes versicolor. Process Biochemistry, 28, 377–384.
Banat, I. M., Nigam, P., Singh, D., & Marchant, R. (1996). Microbial decolorization of textile dye-containing effluents: a review. Bioprocess Technology, 58, 217–227.
Bandounas, L., Wierckx, N. J. P., de Winde, J. H., & Ruijssenaars, H. J. (2011). Isolation and characterization of novel bacterial strains exhibiting ligninolytic potential. BMC Biotechnology, 11, 94.
Barrow, G. I., & Feltham, R. K. A. (1993). Cowan and Steel’s manual for the identification of medical bacteria, 3rd Edn. Cambridge University.
Berryman, D., Houde, F., Deblois, C., & O’Shea, M. (2004). Non phenolic compounds in drinking and surface waters downstream of treated textile and pulp and paper effluents: a survey and preliminary assessment of their potential effects on public health and aquatic life. Chemosphere, 56(3), 247–255.
Buchanan, R. E., & Gibbons, N. E. (1974). Bergey’s Manual of Determinative Bacteriology (8th ed., pp. 290–340). Baltimore: Williams & Wilkins.
Bucher, V. V. C., Hyde, K. D., Pointing, S. B., & Reddy, C. A. (2004). Production of wood decay enzymes, mass loss and lignin solubilization in wood by marine ascomycetes and their anamorphs. Fungal Diversity, 15, 1–14.
Bugg, T. D. H., Ahmad, M., Hardiman, M. E., & Rahmanpour, R. (2011). Pathways for degradation of lignin in bacteria and Fungi. Natural Product Reports, 28, 1883–1896.
Chandra, R., & Abhishek, A. (2011). Bacterial decolorization of black liquor in axenic and mixed condition and characterization of metabolites. Biodegradation, 22, 603–611.
Chandra, R., & Bhargava, R. (2012). Bacterial degradation of synthetic and craft lignin by axenic and mixed culture and their metabolite production. Journal of Environmental Biology, 34, 991–997.
Chandra, R., Raj, A., Purohit, H. J., & Kapley, A. (2007). Characterization and optimization of three potential aerobic bacterial strains for kraft lignin degradation from pulp paper waste. Chemosphere, 67, 839–846.
Chandra, R., Raj, A., Yadav, S., & Patel, D. (2009). Reduction of pollutants in pulp paper mill effluent treated by PCP-degrading bacterial strains. Environmental Monitoring and Assessment, 76, 547–552.
Chuphal, Y., Kumar, V., & Thakur, I. S. (2005). Biodegradation and decolorization of pulp and paper mill effluent by anaerobic and aerobic microorganisms in a sequential bioreactor. World Journal of Microbiology and Biotechnology, 21, 1439–1445.
Cowan, S. T., Steel, K. J., Barrow, G. I., Feltham, R. K. A. (2004). Cowan and Steel's manual for the identification of medical bacteria, books.google.com.
CPCB (2001). Comprehensive industry document for large pulp and paper industry. COINDS/36/2000-2001.
Diwaniyan, S., Kharab, D., Raghukumar, C., & Kuhad, R. C. (2010). Decolourization of synthetic dyes and textile effluents by basidiomycetous fungi. Water, Air and Soil Pollution, 210, 409–419.
Driessel, B. V., & Christov, L. (2001). Decolorization of bleach plant effluent by mucoralean and white-rot fungi in a rotating biological contactor reactor. Journal of Bioscience and Bioengineering, 92, 271–276.
EL-Hanafy, A. A., Abd-Elsalam, H. E., & Hafez, E. E. (2007). Fingerprinting for the lignin degrading bacteria from the soil. Journal of Applied Sciences Research, 3, 470–475.
EL-Hanafy, A. A., Abd-Elsalam, H. E., & Hafez, E. E. (2008). Molecular characterization of two native Egyptian ligninolytic bacterial strains. Journal of Applied Sciences Research, 4, 1291–1296.
Gomathi, V., Cibichakravarthy, B., Ramanathan, A., Sivaramaiah, N., Ramanjaneya, V., Mula, R., Jayasimha, R.D. (2012). Decolourization of paper mill effluent by immobilized cells of phanerochaete chrysosporium. International Journal Of Plant, Animal and Environmental Sciences, 2(1), 141–146.
Gupta, V. K., Minocha, A. K., & Jain, N. (2009). Batch and continuous studies on treatment of pulp mill waste water by Aeromonas formicans. Journal of Chemical Technology and Biotechnology, 76, 547–552.
Hao, D. T., & Man, T. D. (2006). Study on treatment of alkaline black liquor using sulphate reducing bacteria. Advances in Natural Sciences, 7, 139–144.
Husain, Q. (2006). Potential applications of the oxidoreductive enzymes in the decolorization and detoxification of textile and other synthetic dyes from polluted water: a review. Critical Reviews in Biotechnology, 26(4), 201–221.
Kiiskinen, L. L., Rättö, M., & Kruus, K. (2004). Screening for novel laccase-producing microbes. Journal of Applied Microbiology, 97, 640–646.
Manji, S., & Ishihara, A. (2004). Screening of tetrachlorodibenzo-p-dioxin-degrading fungi capable of producing extracellular peroxidases under various conditions. Applied Microbiology and Biotechnology, 63, 438–444.
Morii, H., Nakamiya, K., & Kinoshita, S. (1995). Isolation of lignin decolourising bacterium. Journal of Fermentation and Bioengineering, 80, 296–299.
Pangallo, D., ImonovicOva, A. S., Chovanova, K., & Ferianc, P. (2007). Wooden art objects and the museum environment: identification and biodegradative characteristics of isolated microflora. Letters in Applied Microbiology, 45, 87–94.
Pant, D., & Adholeya, A. (2007). Identification, ligninolytic enzyme activity and decolorization potential of two fungi isolated from a distillery effluent contaminated site. Water, Air, and Soil Pollution, 183, 165–176.
Pearl, I. A., & Benson, H. K. (1940). The determination of lignin in sulphide pulping liquor. Paper Trade Journal, 111, 35–36.
Pfenning, N., & Lippert, K. D. (1966). Uber das vitamin B-12-bedrurbins phototropher schwefelbakterien. Archives of Microbiology, 55, 245–256.
Pokhrel, D., & Viraraghavan, T. (2004). Treatment of pulp and paper mill waste water: A review. Science of the Total Environment Journal, 333, 37–58.
Ponting, S. B. (1999). Qualitative methods for determination of lignocellulolytic enzyme production by tropical fungi. Fungal Diversty, 2, 17–33.
Raj, A., Chandra, R., Reddy, M. M. K., Purohit, H. J., & Kapley, A. (2007a). Biodegradation of kraft lignin by a newly isolated bacterial strain, Aneurinibacillus aneurinilyticus from the sludge of a pulp paper mill. World Journal of Microbiology and Biotechnology, 23, 793–799.
Raj, A., Chandra, R., Reddy, M. M. K., Purohit, H. J., & Kapley, A. (2007b). Biodegradation of Kraft lignin by Bacillus sp. Isolated from sludge of pulp and paper mill. Biodegradation, 18, 783–792.
Raj, A., Kumar, S., Haq, I., & Singh, S. K. (2014). Bioremediation and toxicity reduction in pulp and paper mill effluent by newly isolated ligninolytic Paenibacillus Sp. Ecological Engineering, 71, 355–362.
Reddy, P., Pillay L.V., Kunamneni, A., Singh, S. (2005). Degradation of pulp and paper mill effluent by thermophilic microorganisms using batch systems. Water SA, 31(4), 575–580.
Saraswathi, R., & Saseetharan, M. K. (2010). Investigation on Microorganisms and their Degradation Efficiency in Paper and Pulp Mill Effluent. Journal of Water Resource and Protection, 2, 660–664.
Singh, S., Chandra, R., Patel, D. K., Reddy, M. M. K., & Rai, V. (2008). Investigation of the biotransformation of pentachlorophenol and pulp paper mill effluent decolorization by the bacterial strains in a mixed culture. Bioresource Technology, 99, 5703–5709.
Singh, Y. P., Dhall, P., Mathur, R. M., Jain, R. K., Thakur, V., Kumar, V., Kumar, R., & Kumar, A. (2011). Bioremediation of pulp and paper mill effluent by tannic acid degrading Enterobacter sp. Water, Air and Soil Pollution, 218, 693–701.
Singhal, A., & Thakur, I. S. (2009). Decolorization and detoxification of pulp and paper mill effluent by Cryptococcus sp. Biochemical Engineering Journal, 46, 21–27.
Swamy, J., & Ramsay, J. A. (1999). The evaluation of white rot fungi in the decolorization of textile dyes. Enzyme and Microbial Technology, 24, 130–137.
Tamura, K., Dudley, J., Nei, M., & Kumar, S. (2007). MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Molecular Biology and Evolution, 24, 1596–1599.
Acknowledgments
Authors thankfully acknowledge the Director, Avantha Centre for Industrial Research and Development, Yamuna Nagar for the research facilities provided during the course of investigation. The first author is thankful to Dr. Puneet Pathak for his help and support in carrying out experimental work.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hooda, R., Bhardwaj, N.K. & Singh, P. Screening and Identification of Ligninolytic Bacteria for the Treatment of Pulp and Paper Mill Effluent. Water Air Soil Pollut 226, 305 (2015). https://doi.org/10.1007/s11270-015-2535-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11270-015-2535-y