Abstract
Interspecific somatic hybrids between the dihaploid Solanum tuberosum and the wild species S. pinnatisectum Dun. were produced via protoplast fusion. Protoplast isolation, electrofusion, culture of post-fusion products and regeneration of calli/shoots were undertaken following optimized protocols. Regenerants were characterized for hybridity, ploidy and resistance to Phytophthora infestans (Mont.) de Bery, causal fungal pathogen of late blight disease. From a total of 126 regenerated macrocalli, 12 somatic hybrids were confirmed by possessing species-specific diagnostic bands of their corresponding parents as revealed by RAPD, SSRs and cytoplasmic-DNA analyses. Tetraploid status of the 12 hybrids was determined using flow cytometry analysis. Intermediate phenotypes for leaf, flower, and tuber characteristics and high male fertility were observed in field-grown hybrid plants. Hybrids were highly resistant to foliage late blight based on field assessment for two seasons. In contrast, moderate level of resistance to foliage blight was observed in hybrids based on the detached leaf assay under laboratory conditions. Overall, somatic hybrids with moderate levels of resistance to foliage blight were identified, and these will be useful for in situ hybridization in potato breeding efforts.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Late blight caused by the oomycete Phytophthora infestans (Mont.) de Bery is the most devastating disease of potato throughout the world. This is the pathogen responsible for the Irish famine in the middle of the 1840’s and was the first well-described ‘plant destroyer’ (Fry 2008). This notorious disease entails global yield losses of 16% of the potato crop, estimating an annual financial loss of € 5.2 billion worldwide (Haverkort et al. 2009). This pathogen may causes losses up to 85% in the hills and 60–70% in the plains of India depending upon the disease incidence. Conditions for late blight remain conducive throughout the crop season in the hills and frequently in the plains of India, which needs to be effectively managed by growing the resistant cultivars. Unfortunately, all the present-day Indian potato cultivars have very narrow genetic base those rely upon few parents having mainly the major R genes providing vertical resistance to late blight (Gopal and Oyama 2005), which are broken down with the appearance of complex races of P. infestans. A remarkable example of the Indian potato variety is Kufri Jyoti once resistant to late blight is now categorised as highly susceptible. Henceforth, race non-specific or horizontal or durable resistance to potato late blight is now being preferred for varietal development in India.
Durable resistance to late blight has been found in several wild potato species, which needs to be incorporated into cultivated one by conventional or biotechnological approaches (Bradshaw et al. 2006). Many of the useful wild species, particularly of one endosperm balance number (EBN), cannot be hybridized in situ through conventional technique primarily because of complexities of sexual incompatibilities due to difference in ploidy and EBN (Spooner and Salas 2006). In addition to the transgenic approaches, such type of limitations can be surmounted by the application of biotechnological techniques like somatic fusion. The somatic hybridization technique of protoplast isolation and fusion can provide new opportunities for producing pre-breeding materials by transferring desirable agronomic traits into potato. In comparison to other parallel techniques of chromosomal and genetic engineering, it has a unique potential to transfer simultaneously both nuclear and cytoplasmic genomes (Zhou et al. 2001). It bypasses the gene segregation and enables to transfer both mono- and polygenic traits among the sexually incompatible species (Thieme et al. 2004). Numerous experiments had succeeded in the production of potato somatic hybrids by introgression of desirable agronomic and disease resistance traits. In recent years, wild potato species S. pinnatisectum (Greplová et al. 2008; Polzerová et al. 2011), S. cardiophyllum (Thieme et al. 2010), S. × michoacanum (Szczerbakowa et al. 2010), S. bulbocastanum (Greplová et al. 2008) and S. tarnii (Thieme et al. 2008) were used to transfer late blight resistance into cultivated potato through somatic hybridization. Protoplast fusion has also been succeeded in other crops like citrus (Grosser and Gmitter 2011), cotton (Sun et al. 2011), sweet potato (Yang et al. 2009), Brassicaceae (Ovcharenko et al. 2011). In order to facilitate potato breeding against continuous threat of P. infestans, the introduction of new resistant clones is great concern in potato. Therefore, development of more interspecific potato somatic hybrids by introducing new wild Solanum species is of growing interest. This would provide basic breeding materials to widen genetic base of late blight resistance in potato breeding through in situ hybridization.
This study focuses on the development of tetraploid potato somatic hybrids by incorporating the late blight resistance from wild species into cultivated potato. Protoplast isolation and electrofusion were performed between the diploid wild species S. pinnatisectum Dun. (2n = 2x = 24; 1 EBN) and dihaploid (2n = 2x = 24) of S. tuberosum L. (2n = 4x = 48). The S. pinnatisectum is a diploid Mexican wild tuber-bearing species, with extreme resistance to late blight (Phytophthora infestans (Mont.) de Bary) (Hawkes 1994). The results on protoplast isolation, electrofusion and regeneration; confirmation of somatic hybrids through random amplified polymorphic DNA (RAPD), simple sequence repeats (SSRs), cytoplasmic (chloroplast and mitochondria) DNA and flow cytometry (FC) analyses; and late blight resistance, phenotypic characterization and male fertility assessments are discussed here.
Materials and methods
Plant material
The androgenic dihaploid C-13 (2n = 2x = 24) of the Indian tetraploid potato (Solanum tuberosum L., 2n = 4x = 48) cv. Kufri Chipsona 2 was used for protoplast fusion with diploid wild species S. pinnatisectum Dun. (CGN No.: 17745) (2n = 2x = 24, 1 EBN). The dihaploid was developed previously at the Central Potato Research Institute, Shimla, India (Sharma et al. 2010). True potato seeds (TPS) of the wild species were obtained from the Centre for Genetic Resources, the Netherlands (CGN), Wageningen University and Research Centre, Wageningen, the Netherlands. TPS of the wild species were germinated in vitro and plantlets were multiplied by subculturing leafy nodes on MS (Murashige and Skoog 1962) medium (pH 5.8) supplemented with sucrose (20 g l−1) and solidified with gelrite (2 g l−1). Cultures were grown at 20°C under a 16-h photoperiod (light intensity 50–60 μmol m−2 s−1). The highly susceptible potato varieties Kufri Jyoti, Kufri Chandramukhi and Kufri Bahar were used as control in the late blight resistance test.
Protoplast isolation and electrofusion
Three-week-old in vitro plants were used to isolate mesophyll protoplasts following the protocol under sterile conditions: in vitro plants were cultivated in the dark (covered with dark-black muslin cloths) in a culture room at 20°C for 48 h under a 16-h photoperiod prior to protoplast isolation to integrate cell cycles (Binding et al. 1978). The enzymatic solutions of 1% cellulose ‘Onozuka’ RS (Yakult Pharmaceuticals, Tokyo) and 0.5% macerozyme R 10 (Yakult) were used for cell wall degradation. Young leaf tissues were minced in a Petri dish (Ø = 90 × 15 mm) containing protoplast digestion solution (PDS) (10 ml PDS g−1 tissue) followed by incubation in the dark at 25°C for 16 h without shaking. After incubation, 0.3 M KCl (filter sterilized) was added in a 1:1 ratio (PDS:KCl) then gentle shaking of PDS containing released protoplast. Subsequently, suspension was filtered through a 41-μm nylon sieve and collected in centrifuge tubes. Purification of protoplasts was performed as: filtrates were centrifuged at 60 RCF for 5 min and then resuspended the pellets in 9 ml of 0.6 M sucrose (filter sterilized) followed by 1 ml of 0.3 M KCl layer onto it and centrifuged as above. Live protoplasts (green) were recovered from sucrose: KCl interface, diluted with 5 ml of 0.3 M KCl, centrifuged as above and finally suspended in 200–400 μl of 0.5 M Mannitol, depending upon the isolation, to a final density of 1 × 106 protoplast ml−1. Protoplasts were counted by Hemacytometer (Hausser Scientific, Horsham, PA, USA). The protoplasts were electrofused in a 3.2-mm microslide using the BTX Electro Cell Manipulator ECM 2001 (Harvard Apparatus, Mass., USA). Purified protoplasts were aligned at 30–50 Vcm−1 (1 MHz alternating current) for 30 s and electrofused by one direct-current pulses at 850–1,250 Vcm−1 for 60 μs with a 10-s post-fusion AC field for compacting the fusion products.
Post-fusion protoplast culture and plant regeneration
Electrofused products were immobilized on thin-layer sodium alginate (2.8%) matrices and grown in VKM liquid medium (Binding and Nehls 1977) supplemented with glucose (90 mg ml−1) at 25°C in the dark. Following the cell wall development, regenerating microcalli, were transferred onto MS13K solid medium (Behnke 1975) at 20°C under a 16-h photoperiod for macrocalli and shoot development. Newly regenerated shoots from calli were cultured on MS medium for subsequent growth and multiplication. Characterization of the regenerants was based on regeneration of first shoot per callus followed by subsequent multiplication for further molecular and phenotypic studies.
Identification of somatic hybrids using molecular markers
RAPD analysis
Genomic DNA was isolated from in vitro leaf samples (~300 mg) following a modified CTAB-dichloromethane protocol of Saghai-Maroof et al. (1984). RAPD analysis of somatic hybrids was performed using random decamer primers (Operon Biotechnologies GmbH, Cologne, Germany). A total of 15 RAPD primers, 4 from OPAC series (OPAC-06, OPAC-09, OPAC-13 and OPAC-14), 5 from OPAQ series (OPAQ-02, OPAQ-14, OPAQ-15, OPAQ-16 and OAPQ-20), 3 from OPAT series (OPAT-03, OPAT-06 and OPAT-09), and one each from OPD (OPD-03), OPG (OPG-09) and OPK (OPK-06) were used for the molecular characterization. Selection of RAPD primers and PCR reactions were based on the polymorphism observed in the previous study (Sarkar et al. 2010). The polymerase chain reaction (PCR) was performed in a Mastercycler Gradient (Eppendorf, Hamburg, Germany) in a total volume of 25 μl and consisted of 50 ng DNA templates in 1× PCR buffer, 2.5 mM MgCl2, 200 μM dNTP, 0.2 μM of primer, 1 Unit Taq Polymerase (Qiagen). PCR procedure included: one cycle of 4 min at 94°C, 1 min at 36°C and 2 min at 72°C followed by 43 cycles of 1 min at 94°C, 1 min at 36°C, and 2 min at 72°C, with a final extension of 8 min at 72°C. The amplified DNA products were separated by electrophoresis on a 1.6% agarose gel staining with ethidium bromide (0.5 μg ml−1) in 0.5× TBE buffer (Tris-borate-EDTA) using horizontal gel electrophoresis system Sub-Cell GT (Bio-Rad, USA) at room temperature. The gels were visualized under Gel Doc System (Alpha Innotech, San Leandro, CA) and bands were scored using the AlphaImager 3400 software (Alpha Innotech) by comparing with 100 bp DNA ladder (Fermentas, Burlington, Canada). Somatic hybrids were confirmed on the basis of presence of species-specific diagnostic bands of the corresponding parents. Somatic hybrids profiles were looked for similarity with the PCR-amplicons of pooled parental DNA, mixed before PCR, as a control of synthetic somatic hybrid in all the DNA analyses.
SSR analysis
A total of 33 SSR markers, viz., STG0001, STG0010, STG0016, STG0025, STl0001, STI0003, STI0004, STI0012, STI0014, STI0030, STI0032, STI0033, STM0019a, STM0019b, STM0031, STM0037, STM1052, STM1053, STM1064, STM1104, STM1106, STM5114, STM5121, STM5127, STPoAc58 (Ghislain et al. 2009), POTM 1–2, STM0015, STM1043, STM1069, STM3003, STWAX-1, STWAX-2 and STWIN12G (Sharma et al. 2010) were used for the identification of somatic hybrids. Annealing temperature (Ta) of the primers was determined using Mastercycler Gradient (Eppendorf, Hamburg, Germany) and PCR was performed as described by Ghislain et al. (2009) and Sharma et al. (2010). Amplified products were resolved on 3% metaphor agarose (Lonza, Rockland, ME, USA), and 8% polyacrylamide gel electrophoresis followed by gel documentation, scoring, analysis and interpretation of results as above.
Cytoplasmic DNA analysis
Cytoplasm (chloroplast and mitochondria) type was analyzed from the genomic DNA using mitochondrial specific primers ALM_4/ALM_5 and ALM_6/ALM_7, and chloroplast specific primer ALCP_1/ALCP_3. PCR condition was followed at optimized annealing temperatures (Ta = 55°C for mitochondrial primers, and Ta = 45°C for chloroplast primer) as described by Lössl et al. (2000). Amplified products were analyzed by electrophoresis on 1.6% agarose gel in 0.5× TBE buffer stained with ethidium bromide (0.5 μg ml−1) and documented as above. Hybrids were confirmed by the presence of parental banding profiles as above. Cytoplasm type was determined by amplicons as explained by Lössl et al. (2000).
Ploidy estimation using flow cytometry
Ploidy level of the somatic hybrids was confirmed by flow cytometric analysis following the methods described by Arumuganathan and Earle (1991). Leaf samples (~100 mg) from 3- to 4-week-old in vitro plants were macerated on ice in 1 ml nuclei isolation buffer from total volume of 14.70 ml [MgSO4 buffer (MgSO4.7H2O, 10 mM; KCl, 50 mM; HEPES, 5 mM), 14.325 ml; Dithiothretol (15 mg); Triton X (375 μl)]. Macerates were filtered through a 40-μm nylon sieve followed by centrifugation at 5,000 rpm for 5 min. White pellets (nuclei) were dissolved in MgSO4 buffer and treated with 2 μl RNAse at 37°C for 15 min. Propidium iodide-stained nuclei were used to measure relative fluorescence in a FACSCalibur flow cytometer (Becton–Dickinson, San Jose, USA) using CRBC (Chicken Red Blood Cell) (2C value of DNA = 2.33 pg) as an internal standard. For each sample, there were three independent replicated measurements. The materials were analyzed for forward (FSC) versus side (SSC) scatter signals for at least 10,000 nuclei in each sample. The peak corresponding to the CRBC nuclei was adjusted to around channel 250 set on a linear scale of fluorescence intensity. The nuclear DNA amount (2C value in pg) was estimated by direct comparison of the mean position of nuclear peak of clone to that of CRBC.
Phenotypic characterization
In vitro raised somatic hybrids were evaluated for the phenotypic traits (foliage, flower and tuber) in the field grown plants. Somatic hybrids were planted in field during main crop season at Shimla, since potato plants typically flower only under long-day conditions of Shimla hills (mid-Himalayas) in northern India. Hybrid plants consisted of 5 plants per clone were planted at a spacing of 20 cm in rows 60 cm apart in a randomized complete block design (RCBD) with three replications. Fertilizers schedules (120–150 kg N : 100–150 kg P2O5 : 120 kg K2O per hectare) by combined basal application of full dose of P and K along with ½ of N at planting; and split application of N (½ at planting + ½ at earthing up) is applied. Cultural practices (irrigation and inter-cultural operations) were followed as recommended package of practices. Morphological characters were recorded as per described by Gopal et al. (2008) for general canopy characteristics (plant vigour, plant height, foliage structure, foliage color and foliage gloss), stem morphology (solidity, predominant colour, secondary colour and wings type) and leaf morphology (leaf structure, leaf length, leaf width, leaf shape and waviness of margin) were recorded at 50 days after planting. Observations on floral characteristics (days to first flowering, flower intensity, corolla size, corolla colour, anther coluor and stigma shape) were recorded after initiation of flowering. Plant maturity type was recorded at 90 days after planting, whereas tuber characters (tuber shape, tuber skin color, tuber skin type, tuber flesh colour and depth of eyes) were recorded after harvesting at 120 days after planting.
Male fertility determination
Male fertility of the somatic hybrids was determined as procedures described by Orrillo and Bonierbale (2009) by staining pollens with 1% acetocarmine (1 g carmine: 100 ml 45% glacial acetic acid). Stained viable (dark red) and non-viable (no stain) pollens were examined through microscope (Olympus IX-71, Tokyo). Counts were recorded from ten different flowers on five locations of the slide and estimated mean value was presented in the result.
Field assessment of foliage late blight resistance
Somatic hybrids were raised from in vitro plants for the field experiments. Hybrids were assessed for the foliage late blight resistance in 2 years (2009 and 2010) during the main crop season (May–August) at Shimla (31.1°N; 77.1°E; 2,205 m amsl), a highly conducive for late blight infection. During both the years foliage late blight infection was scored between 60 and 110 days after planting (DAP) i.e. from the day of appearance of first symptoms till 100% blight infection in a highly susceptible control variety Kufri Jyoti, and till existence of the hybrid plants. All the procedures for the late blight observations were followed as described in manual of International Potato Centre (CIP) (Anonymous 2007). Area under disease progressive curve (AUDPC) was calculated in percentage-days according to the midpoint rule method described in the CIP-manual. Following this method, the relative area under disease progressive curve (RAUDPC = AUDPC/[(t n − t 1) × 100] where (t n − t 1) is the duration of epidemic) was estimated to identify the disease level that ranged from 0.0 to 1.0 without any unit, compared to susceptible control (Anonymous 2007).
Detached leaf assay of late blight resistance
Phytophthora infestans isolate HP09/40 (A2, race 1–11) was used for inoculum preparation. The inoculum was multiplied using tuber slice method. Tubers of susceptible cultivar Kufri Bahar were surface sterilized with ethyl alcohol and cut into 1 cm thick slices with sterilized knife. Slices were placed in a Petri dish (Ø = 90 × 15 mm) and inoculated with P. infestans by scratching the fungal mycelium on slice surface with sterilized needle under aseptic conditions and incubated in air tight plastic boxes lined with moist foam sheet at 18 ± 1°C for a week in the dark. Tuber slices containing zoosporangia of P. infestans were dislodged in sterilized distilled water. Zoosporangial suspension was kept at 4°C for 45–60 min. for release of zoospores. The zoospore concentration was adjusted to a level of 5 × 104 (sporangia/ml) using haemocytometer for inoculation purpose (Vleeshouwers et al. 1999).
For the detached leaf assay, fourth fully developed leaves (counted from the top) were cut from net house grown plants and placed in plastic trays (lined with moist foam sheet) on perforated plastic separators. Five leaflets per compound leaf were inoculated (one drop per leaflet) by pipetting 10 μl droplet of zoospore suspension on the abaxial side (Vleeshouwers et al. 1999). The trays were incubated at 18 ± 1°C for 5 days. Observations were recorded on the 6th day of the inoculation measuring the leaf area infected. The genotypes were characterized into different disease reaction categories as: highly resistant (HR, LA ≤1.0 cm2), resistant (R, LA 1.1–2.5 cm2), moderately resistant (MR, LA 2.51–6.0 cm2) and susceptible (S, LA >6.0 cm2) following the procedures described by Kaushik et al. (2007).
Statistical analysis
Prior to univariate ANOVA analysis of the nDNA content for ploidy estimation, Levene test for homogeneity of variance, and Kolmogorov–Smirnov test for normality assumption were tested for independent variables. Accordingly, data were analyzed followed by all pairwise multiple comparisons of mean values using the post-hoc Tukey’s honestly significant difference test. The term significant has been used to indicate differences for which P ≤ 0.01 or P ≤ 0.05. Verification ability of flow cytometry and RAPD analysis was tested by McNemar Chi-square test. Statistical data were analysed using the software Windostat 8.5 (Ameerpet, Hyderabad, India).
Results
Protoplast isolation, fusion and regeneration
Mesophyll protoplasts were isolated from the in vitro grown plants and electrofused for the regeneration of somatic hybrids between the dihaploid S. tuberosum and the diploid S. pinnatisectum. Figures depicting the fusion of protoplast and rounding up of the post-fusion products are included in the Appendix Fig. 4. Details of number of protoplast fusion attempted, regenerated shoots and identified hybrids are presented in the Appendix Table 4. In total 142 fusion experiments were performed and 126 macrocalli were regenerated from the post-fusion products. Subsequently, first shoot primordia were observed after 6 months of macrocalli development. Four–six months later, 63 of the microshoots produced elongated shoots and 23 of in vitro shoots were detached for rooting. Of these, the most vigorously growing rooted plants were further propagated in vitro, while the rest were discarded because of various growth abnormalities and contaminations. Fourteen hybrids regenerants were finally multiplied and 12 somatic hybrids were confirmed through the molecular and phenotypic characterizations.
Identification of somatic hybrids
RAPD analysis
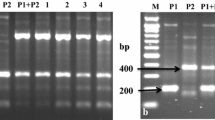
Somatic hybrids were identified by using 15 selected RAPD primers that revealed species-specific diagnostic bands of their corresponding parents in the hybrids. Figure 1a shows RAPD profiles of somatic hybrids and parents generated by one primer OPAQ-02, where somatic hybrids showed five diagnostic bands—three (753, 1,433 and 1,647 bp) of parent C-13, and two (885 and 1,270 bp) of S. pinnatisectum. Similar trends of species-specific diagnostic band profiles were revealed by the other 14 RAPD primers and confirmed 12 somatic hybrids (P1–P10, P12 and P13). Whereas, two regenerants (P11 and P14) were not somatic hybrids, since they showed only S. pinnatisectum-specific bands. DNA profiles of the hybrids using random primers are documented in the Appendix Table 5.
Profiles of a RAPD primer OPAQ-02, b SSRs primer STI0012, and c mitochondrial primer pairs ALM_6/ALM_7 of regenerants between the dihaploid Solanum tuberosum and the diploid S. pinnatisectum. M = 100 bp (a), 50 bp (b and c) ladder; P 1 = C-13, P 2 = S. pinnatisectum, P 1 + P 2 = Pooled parental DNA; Identified somatic hybrids (1–10, 12 and 13)
SSR analysis
Somatic hybrids were also confirmed by using 33 SSR markers, out of all only 7 primers STl0001, STI0012, STM0019, STM1104, POTM 1–2, STWING12G and STWAX-1 were detected somatic hybrids possessing species-specific diagnostic bands. SSR primers STI0012 produced combinations of 186 and 191 bp of C-13, and 201 bp of the wild parent in the somatic hybrids (Fig. 1b). Whereas, SSR primers STI0001 produced amplification products viz. 190, 194 and 212 bp of the parent C-13, and 200, 212 and 248 bp of the parent S. pinnatisectum. Somatic hybrids were identified based on the combination of the presence of DNA fragments of both the parents in the hybrids’ profile. As a result of SSRs profiling these somatic hybrid P1–P10, P12, P13 were confirmed like random primers except P11 and P14. In the Fig. 1b, c, regenerant P14 was not included in SSRs analysis because of failure of its hybridity revealed by random primers and phenotypic assessments. But simultaneously P11 was included in analysis as a negative control (i.e. non-somatic hybrid). The primers amplified with some common bands (data not shown) that could not be characterized as species-specific diagnostic bands were failed to characterize the hybrids. Details of PCR amplified products amplified by SSRs primers are presented in the Appendix Table 6.
Cytoplasm type analysis
Cytoplasmic genome was examined by mitochondrial and chloroplast DNA specific primers as described by Lössl et al. (2000). Amplification with mitochondrial specific primers ALM_4/ALM_5 and ALM_6/ALM_7; and chloroplast specific primers ALCP_1/ALCP_3 showed good polymorphism in parental and somatic hybrids’ profiles. Mitochondrial primers ALM_6/ALM_7 confirmed the hybridity by the combined amplification products of 400 bp of C-13, and 480 and 1,200 bp of S. pinnatisectum in the somatic hybrids DNA profiles’ (Fig. 1c). Primers ALM_4/ALM_5 revealed distinct PCR products of 2,400 bp of C-13 and 1,600 bp of S. pinnatisectum, while somatic hybrids were confirmed by the presence of both 1,600 and 2,400 bp products. Primers ALCP_1/ALCP_3 amplified with 622 bp in parent C-13 and somatic hybrids, whereas no amplicons were observed in wild parent. The amplicons of 622, 1,200 and 2,400 bp indicated dominance of W/α type cytoplasm in the somatic hybrids as same as described by Lössl et al. (2000).
Flow cytometric analysis
Somatic hybrids were evaluated for the ploidy level by flow cytometric analysis based on nDNA content estimation as outlined in the Table 1. The co-efficient of variation of the nDNA contents varied between 1.47 and 3.81%. Figure 2 illustrates the nDNA peaks vis-a-vis fluorescence intensities associated with somatic hybrids and internal standard. Flow cytometry analysis revealed variations in the ploidy level of somatic hybrids. The nDNA values were analysed for normality assumption using Kolmogorov–Smirnov test. Levene test for homogeneity of variance of nDNA content for group I (diploids) and group II (tetraploids) showed that population is identically distributed but variances are not same for the groups I and II. Univariate F test for the nDNA content and T test between the groups I and II was highly significant at P ≤ 0.01, since they represent different values of diploid and tetraploid groups. Pair-wise multiple comparisons statistics between individuals of group I and group II was highly significant at P ≤ 0.01 according to Tukey’s honestly significant difference (HSD) test. Moreover, according to McNemar Chi-square test no difference was existed between flow cytometry and RAPD analysis for the confirmation of somatic hybrids. Finally, 12 somatic hybrids (P1–P10, P12 and P13) exhibited tetraploid, whereas two regenerants P11 and P14 were found diploid.
Phenotypic assessment
Distinct phenotypic variations were observed in the field-grown somatic hybrid plants. General and canopy characteristics stem, leaf, floral and tuber attributes of the hybrids and their parents are documented in the Table 2 and displayed in the Fig. 3. Somatic hybrids displayed intermediate morphology of their parents, except two regenerants P11 and P14, which showed wild parent type phenotypes. Interestingly, leaf shape of the somatic hybrids was ovate lanceolate, whereas ovate in C-13 and narrow lanceolate in S. pinnatisectum. Tuber skin colour of the somatic hybrids was dark purple, whereas brown-green of C-13, and brown-purple of S. pinnatisectum. Intermediate leaf structure and medium waviness of margin were appeared in the somatic hybrids. Unlike parents, floral characters of the somatic hybrids were also observed different. Flowers of somatic hybrids and S. pinnatisectum were white with purple shade colour, whereas C-13 had white with yellow stripes. However, corolla size of somatic hybrids was larger (3.87 cm) than that of C-13 (2.38 cm) and S. pinnatisectum (3.14 cm). All the somatic hybrid plants were found late maturing type and survived till 120 days after planting in the field.
Phenotypes of a Leaves (A), flowers (B) and tubers (C) of the parents C-13, S. pinnatisectum and somatic hybrid, respectively (from left to right), b Plant morphology of C-13 (D), S. pinnatisectum (E) and somatic hybrid P7 (F), and c Late blight susceptible cultivar (control) Kufri Jyoti (G) and resistant somatic hybrid P7 (H)
Male fertility determination
Flower intensity was high in somatic hybrids grown in the natural field at Shimla conditions. Following the acetocarmine staining, mean value of the male fertility of the hybrids was estimated as 82.48%. In parents C-13 and S. pinnatisectum male fertility was estimated 81.73 and 76.52%, respectively (Table 2).
Field assessment of late blight resistance
Field assessment of somatic hybrids showed a high degree of late blight resistance to foliage under the Shimla conditions (Fig. 3c) for two seasons. Table 3 represents the AUDPC and RAUDPC values of the late blight scoring of the hybrids. However, somatic hybrids showed a minimal presence of late blight symptoms in both seasons where AUDPC values ranged 0.17–2.13 in the foliage, compared to susceptible control (AUDPC = 1,910.45 and 1,915.30 in the years 2009 and 2010, respectively). It appeared that few blight symptoms were recorded after 90 days of planting i.e. delayed expression of P. infestans by that time susceptible control was almost fully infected with blight. In dihaploid (C-13) parent late blight symptoms were observed and AUDPC values were score as: 54.05 and 117.5 in the years 2009 and 2010, respectively. Wild parent was found highly resistance during both the seasons and negligible symptoms were recorded (AUDPC = 0.08 and 0.11 in the years 2009 and 2010, respectively). Whereas, late blight symptoms were severe in the experimental plot of control i.e. Kufri Jyoti, a highly susceptible variety and RAUDPC values were scored as: 0.45 and 0.46 in the years 2009 and 2010, respectively. As a consequence, somatic hybrids were characterized as highly resistant.
Detached leaf assay of late blight resistance
Somatic hybrids were also assessed by detached leaf test for foliage blight resistance under highly congenial conditions in the laboratory. Infected detached leaf area of each of the somatic hybrid, parent and susceptible control is presented in the Table 3. In contrast to field test, based on the detached leaf area test, where infected lesion size varied in between 0.25 and 17.30 cm2 in the tested plants. Accordingly, somatic hybrids P5 (5 cm2), P7 (4.80 cm2), P12 (5.10 cm2) and P13 (4.81 cm2) were categorised into moderate resistant. Besides, minor P. infestans sporulations were noticed on few leaflets of the somatic hybrids P5, P12 and P13, whereas no sporulations were observed on either of the leaflets in the somatic hybrid P7. Other hybrids P1–P4, P6, P8–P10 were categorised into susceptible type based on the infected lesion area. Lastly, result demonstrates that somatic hybrid P7 could be better source of late blight resistance in potato breeding.
Discussion
In the present study, protoplast isolation and electrofusion was performed to produce interspecific somatic hybrids between wild and cultivated potato. Our optimized procedures of protoplast isolation, electrofusion and regeneration of hybrid regenerants were proved quite efficient. We obtained an average frequency of regeneration of about 17% in vitro shoots over protoplast fusion attempted. It was demonstrated previously that electrofusion is a widely used technique and is more efficient than chemical fusion (Pehu et al. 1989). Serraf et al. (1991) reported a higher (~30%) average frequency of electrofusion success. Our lower regeneration potential of somatic hybrids were due to the first time experiment that took time for optimization of regeneration protocol for the protoplast isolation, electrofusion and post-fusion products from the exotic wild species and indigenous dihaploid potato. As a result of efficient somatic fusions and cytoplasmic and nuclear recombinations, we regenerated 126 calli, out of them 63 were excised and regenerated into shoots. From them 23 were able to develop roots and only 12 were confirmed somatic hybrids of S. tuberosum and S. pinnatisectum. Low regeneration ability was corresponded to the somatic hybrids of S. pinnatisectum and dihaploid H-1805 produced by Szczerbakowa et al. (2005). It was believed that the low frequency of regeneration of somatic hybrids could be a result of both low yield and lack of calli differentiation capacity of the mesophyll protoplast in parents (Szczerbakowa et al. 2005).
Somatic hybrids were identified at molecular and morphological levels and ploidy level by flow cytometry estimation. Morphology of the 12 somatic hybrids exhibited intermediate phenotype, whereas two regenerants were not somatic hybrids. Probably failure of two regenerants was due to high regeneration potential of individual calli and shoot development consisted of one parent only. Pollen fertility assessment of the field-grown hybrid plants showed higher fertility to be utilized for in situ hybridization. In flow cytometric analysis, 12 somatic hybrids were identified as expected ploidy: tetraploid, from the fusion of dihaploid S. tuberosum C-13 (+) diploid S. pinnatisectum. Variability in the morphology and ploidy estimation of hybrids originated from any fusion partners proved more common in Solanum (Szczerbakowa et al. 2011; Polzerová et al. 2011; Greplová et al. 2008), and in other crops as well like passion fruit (Pinto et al. 2010), cactus (Lema-Rumińska 2011) and Devil’s claw (Stancheva et al. 2011). RAPD analysis was used as first method to identify somatic hybrids followed by SSR and cytoplasm type analyses. Application of molecular markers like RAPD (Greplová et al. 2008), SSRs and cytoplasm types (Polzerová et al. 2011; Tiwari et al. 2010) were used successfully for the hybrids confirmation and cytoplasmic characterizations. However, the most common problem with RAPD technique is its low reproducibility. Even though, to check the minor differences in amplified products, it can be optimized with PCR reaction to obtain reproducible and interpretable results (Caetano-Anolles et al. 1992). We obtained good random amplification with optimized PCR reaction conditions for the identification of hybrid plants at lower annealing temperature (Ta = 36°C). RAPD clearly revealed complementary banding patterns in the somatic hybrids, indicating presence of genetic materials from each parent. Furthermore, out of 33 SSR, only 7 markers identified the 12 somatic hybrids having variability in the fingerprinting patterns that corresponded to profiles described by Ghislain et al. (2009) and Sharma et al. (2010). Other SSR markers did not confirm the hybridity due to lack of complementary banding profiles in the somatic hybrids of their corresponding parents. Our study on microsatellite analysis is comparable to the variable fingerprints of somatic hybrids of S. tuberosum and S. pinnatisectum as demonstrated by Polzerová et al. (2011).
Cytoplasmic analyses conducted in the present study focused on the identification of mitochondrial and chloroplast types in the somatic hybrids. Coexistence of mitochondrial parental types was observed in the somatic hybrids of S. tuberosum and S. pinnatiestum, however, cosegregation of chloroplasts was not found in these somatic hybrids. It was reported that the chloroplast DNA does not recombine during protoplast fusion (Wenzel 1994). Since, plastomes from the two parents are normally not mixed in one cell and as a result the post-fused products contain plastome of either of the fusion parents only (Wenzel 1994). There are previous reports of chloroplast genome about random segregation (Bastia et al. 2000), sporadic cosegregation (Liu et al. 2005) and recombination of plastid genomes (Bidani et al. 2007). In contrast, chondriomes may recombine, since mitochondrial DNA rarely eliminated from plastomes (Polzerová et al. 2011). Our study showed dominance of cytoplasm type W/α than W/β in the somatic hybrids in consistent with Chimote et al. (2008).
Our late blight resistance tests explained that there were less inoculum dose and less favourable environments for blight infections in the field tests, compared to highly congenial conditions in the detached leaf test under the laboratory conditions. Hybrid plants were performed well in the natural conditions and few traces of blight infection were recorded at the later stage of crop growth i.e. after 90 DAP to a minimal extent by that time susceptible control variety was almost finished. Unlike laboratory conditions, potato crop are grown naturally in the field and infection varied from the low to high degree of blight incidence. Indeed, our experiments envisaged that somatic hybrid P7 has moderate degree of resistance as well as delayed expression of few late blight symptoms in the field. Our result corresponds to the reduce resistance level somatic hybrids of S. pinnatisectum (+) S. tuberosum that could be associated with the genome-doses effects, chromosome instability and preferential elimination of some chromosome of the wild species (Thieme et al. 1997), somaclonal variations (Sree Ramulu 1986) and/or presence of negative gene interaction between parents (Thieme et al. 1997). At the end, our knowledge of the inheritance of late blight resistance is insufficient and more detailed studies on the genetics and inheritance of this disease are required in the hybrids.
We have demonstrated that late blight resistance was expressed to moderate degree and delayed blight expressions in the somatic hybrids. Since, at present in India we target currently on the producing early bulker potato cultivars that mature in between 75 and 90 DAP with at least some degree of resistance to late blight. Thus, this study provides an example of successful somatic hybridization of sexually incompatible wild potato species and transfer of late blight resistance trait into cultivated potato. The hybrids offer new insight into the transfer of resistance into cultivated tetraploid potato through in situ hybridization. Our future research investigations are progressing on the in situ hybridization of the somatic hybrids, characterization and validation of resistance genes to accelerate MAS in the potato breeding.
Abbreviations
- AUDPC:
-
Area under the disease progress curve
- CRBC:
-
Chicken red blood cell
- CTAB:
-
Cetyltrimethylammonium bromide
- DAP:
-
Days after planting
- nDNA:
-
Nuclear DNA
- EBN:
-
Endosperm balance number
- FC:
-
Flow cytometry
- HEPES:
-
4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid
- MS:
-
Murashige and Skoog
- MAS:
-
Marker assisted selection
- PDS:
-
Protoplast digestion solution
- RAPD:
-
Random amplified polymorphic DNA
- RAUDPC:
-
Relative area under the disease progress curve
- RCBD:
-
Randomized complete block design
- SSR:
-
Simple sequence repeat
- TPS:
-
True potato seed
References
Anonymous (2007) Procedures for standard evaluation trials of advanced potato clones. An International Cooperators’ Guide. International Potato Center, Lima, pp 41–53
Arumuganathan K, Earle ED (1991) Estimation of nuclear DNA content of plants by flow cytometry. Plant Mol Biol Rep 9:229–241
Bastia T, Carotenuto N, Basile B, Zonia A, Cardi T (2000) Induction of novel organelle DNA variation and transfer of resistance to frost and Verticillium wilt in Solanum tuberosum through somatic hybridization with 1 EBN S. commersonii. Euphytica 116:1–10
Behnke M (1975) Regeneration in Gewebekulturen einiger dihaplider Solanum tuberosum-Klone. Z Pflanzenziicht 75:262–265
Bidani A, Nouri-Ellouz O, Lakhoua L, Sihachakr D, Cheniclet C, Mahjoub A, Drira N, Gargouri-Bouzid R (2007) Interspecific potato somatic hybrids between Solanum berthaultii and Solanum tuberosum L. showed recombinant plastome and improved tolerance to salinity. Plant Cell Tiss Organ Cult 91:179–189
Binding H, Nehls R (1977) Regeneration of isolated protoplast to plant Solanum dulcamara L. Z Pflanzenphysiol 85:279–280
Binding H, Nehls R, Schieder O, Sopory SK, Wenzel C (1978) Regeneration of mesophyll protoplast isolated from dihaploid clones of Solanum tuberosum L. Plant Physiol 43:52–54
Bradshaw JE, Bryan GJ, Ramsay G (2006) Genetic resources (including wild and cultivated Solanum species) and progress in their utilisation in potato breeding. Potato Res 49:49–65
Caetano-Anolles G, Bassam BJ, Gresshoff PM (1992) Primer template interactions during DNA amplification fingerprinting with single arbitrary oligonucleotides. Mol Gen Genet 235:157–165
Chimote VP, Chakrabarti SK, Pattanayak D, Pandey SK, Naik PS (2008) Molecular analysis of cytoplasm type in Indian potato varieties. Euphytica 162:69–80
Fry W (2008) Phytophthora infestans: the plant (and R gene) destroyer. Mol Plant Pathol 9:385–402
Ghislain M, Núñez J, Herrera MR, Pignataro J, Guzman F, Bonierbale M, Spooner DM (2009) Robust and highly informative microsatellite-based genetic identity kit for potato. Mol Breed 23:377–388
Gopal J, Oyama K (2005) Genetic base of Indian potato selections as revealed by pedigree analysis. Euphytica 142:23–31
Gopal J, Pandey SK, Kumar V, Kumar R, Pandey PC, Singh SV (2008) Morphological descriptors for DUS testing of potato varieties. Plant Gen Res New 154:40–47
Greplová M, Polzerová H, Vlastníková H (2008) Electrofusion of protoplasts from Solanum tuberosum, S. bulbocastanum and S. pinnatisectum. Acta Physiol Plant 30:787–796
Grosser JW, Gmitter FG Jr (2011) Protoplast fusion for production of tetraploids and triploids: applications for scion and rootstock breeding in citrus. Plant Cell Tiss Organ Cult 104:343–357
Haverkort A, Struik P, Visser R, Jacobsen E (2009) Applied biotechnology to combat late blight in potato caused by Phytophthora infestans. Potato Res 52:249–264
Hawkes JG (1994) Origins of cultivated potatoes and species relationships. In: Bradshaw JE, Mackay GR (eds) Potato genetics. CAB International, Wallingford, pp 3–42
Kaushik SK, Bhardwaj V, Singh PH, Singh BP (2007) Evaluation of potato germplasm for adaptability and resistance to late blight. Potato J 34:443–444
Lema-Rumińska J (2011) Flow cytometric analysis of somatic embryos, shoots, and calli of the cactus Copiapoa tenuissima Ritt. forma monstruosa. Plant Cell Tiss Organ Cult. doi:10.1007/s11240-011-9941-7
Liu J, Xu X, Deng X (2005) Intergeneric somatic hybridization and its application to crop generic improvement. Plant Cell Tiss Organ Cult 82:19–44
Lössl A, Götz M, Braun A, Wenzel G (2000) Molecular markers for cytoplasm in potato: male sterility and contribution of different plastid-mitochondrial configurations to starch production. Euphytica 116:221–230
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue culture. Physiol Plant 15:473–497
Orrillo M, Bonierbale M (2009) Biología reproductiva y citogenética de la papa. International Potato Center, Lima, pp 9–17
Ovcharenko O, Momot V, Cherep N, Sheludko Y, Komarnitsky I, Rudas V, Kuchuk N (2011) Transfer of transformed Lesquerella fendleri (Gray) Wats. chloroplasts into Orychophragmus violaceus (L.) O.E. Schulz by protoplast fusion. Plant Cell Tiss Organ Cult 105:21–27
Pehu E, Karp A, Moore K, Steele S, Dunckley R, Jones MGK (1989) Molecular, cytogenetic and morphological characterization of somatic hybrids of dihaploid Solanum tuberosum and diploid S. brevidens. Theor Appl Genet 78:696–704
Pinto DLP, Barros BA, Viccini LF, de Campos JMS, da Silva ML, Otoni WC (2010) Ploidy stability of somatic embryogenesis-derived Passiflora cincinnata Mast. plants as assessed by flow cytometry. Plant Cell Tiss Organ Cult 103:71–79
Polzerová H, Patzak J, Greplová M (2011) Early characterization of somatic hybrids from symmetric protoplast electrofusion of Solanum pinnatisectum Dun. and Solanum tuberosum L. Plant Cell Tiss Organ Cult 104:163–170
Saghai-Maroof MA, Soliman KM, Jorgensen RA, Allard RW (1984) Ribosomal DNA spacer-length polymorphisms in barley: Mendelian inheritance, chromosomal locations, and population dynamics. Proc Natl Acad Sci 81:8014–8018
Sarkar D, Sharma S, Chandel P, Pandey SK (2010) Evidence for gametoclonal variation in potato (Solanum tuberosum L.). Plant Growth Regul 61:109–117
Serraf I, Sihachakr D, Ducreux G, Brown SC, Allot M, Barghi N, Rossignol L (1991) Interspecific somatic hybridization in potato by protoplast electrofusion. Plant Sci 76:115–126
Sharma S, Sarkar D, Pandey SK (2010) Phenotypic characterization and nuclear microsatellite analysis reveal genomic changes and rearrangements underlying androgenesis in tetraploid potatoes (Solanum tuberosum L.). Euphytica 171:313–326
Spooner DM, Salas A (2006) Structure, biosystematics, and genetic resources. In: Gopal J, Khurana SMP (eds) Handbook of potato production, improvement and postharvest management. Food Product Press, New York, pp 1–40
Sree Ramulu K (1986) Case histories of genetic variability in vitro: potato. In: Vasil IK (ed) Cell, culture and somatic cell genetics of plants. Academic Press, Minnesota, pp 449–473
Stancheva N, Weber J, Schulze J, Alipieva K, Ludwig-Müller J, Haas C, Georgiev V, Bley T, Georgiev M (2011) Phytochemical and flow cytometric analyses of Devil’s claw cell cultures. Plant Cell Tiss Organ Cult 105:79–84
Sun Y, Liu S, Wang Y, Jones BJ, Wang H, Zhu S (2011) An interspecific somatic hybrid between upland cotton (G. hirsutum L. cv. ZDM-3) and wild diploid cotton (G. klotzschianum A.). Plant Cell Tiss Organ Cult. doi:10.1007/s11240-011-9939-1
Szczerbakowa A, Boltowicz D, Lebecka R, Radomski P, Wielgat B (2005) Characteristics of the interspecific somatic hybrids Solanum pinnatisectum (+) S. tuberosum H-8105. Acta Physiol Plant 3:265–273
Szczerbakowa A, Tarwacka J, Oskiera M, Jakuczun H, Wielgat B (2010) Somatic hybridization between the diploids of S. × michoacanum and S. tuberosum. Acta Physiol Plant 32:867–873
Szczerbakowa A, Tarwacka J, Sliwinska E, Wielgat B (2011) Nuclear DNA content and chromosomal number in somatic hybrid allopolyploid of Solanum. Plant Cell Tiss Organ Cult. doi:10.1007/s11240-011-9932-8
Thieme R, Darsow U, Gavrilenko T, Dorokhov D, Tiemann H (1997) Production of somatic hybrids between S. tuberosum L. and late blight resistant Mexican wild potato species. Euphytica 97:189–200
Thieme R, Darsow U, Rakosy-Tican L, Kang Z, Gavrilenko T, Antonova O, Heimbach U, Thieme T (2004) Use of somatic hybridization to transfer resistance to late blight and potato virus Y (PVY) into cultivated potato. Plant Breed Seed Sci 50:113–118
Thieme R, Rakosy-Tican E, Gavrilenko T, Antonova O, Schubert J, Nachtigall M, Heimbach U, Thieme T (2008) Novel somatic hybrids (Solanum tuberosum L. + Solanum tarnii) and their fertile BC1 progenies express extreme resistance to potato virus Y and late blight. Theor Appl Genet 116:691–700
Thieme R, Rakosy-Tican E, Nachtigall M, Schubert J, Hammann T, Antonova O, Gavrilenko T, Heimbach U, Thieme T (2010) Characterization of the multiple resistance traits of somatic hybrids between Solanum cardiophyllum Lindl. and two commercial potato cultivars. Plant Cell Rep 29:1187–1201
Tiwari JK, Poonam B, Sarkar D, Pandey SK, Gopal J, Kumar SR (2010) Molecular and morphological characterization of somatic hybrids between Solanum tuberosum L. and S. etuberosum Lindl. Plant Cell Tiss Organ Cult 103:175–187
Vleeshouwers VGAA, van Dooijeweert W, Keizer LCP, Sijpkes L, Govers F, Colon LT (1999) A laboratory assay for Phytophthora infestans resistance in various Solanum species reflects the field situation. Eur J Plant Pathol 105:241–250
Wenzel G (1994) Tissue culture. In: Bradshaw JE, Mackey GR (eds) Potato genetics. CAB International, Wallingford, pp 173–195
Yang Y, Guan S, Zhai H, He S, Liu Q (2009) Development and evaluation of a storage root-bearing sweetpotato somatic hybrid between Ipomoea batatas (L.) Lam. and I. triloba L. Plant Cell Tiss Organ Cult 99:83–89
Zhou A, Xia G, Zhang X, Chen H, Hu H (2001) Analysis of chromosomal and organellar DNA of somatic hybrids between Triticum aestivum and Haynaldia villosa Schur. Mol Genet Genomics 265:387–393
Acknowledgments
Authors are grateful to Dr. PS Ahuja, Director, Institute of Himalayan Bioresource and Technology (IHBT), Palampur, Himachal Pradesh, India for the facility provided for flow cytometry analysis. Authors thank to Mr. Sheeshram Thakur for the in vitro multiplication and field management. The financial assistance received in the form of an ad-hoc research project (F. No. 8-45/2004-Hort. II) from the Indian Council of Agricultural Research (ICAR), New Delhi is also acknowledged. Useful comments and suggestions on the manuscript from the editor, associate editor and two anonymous reviewers are gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding author
Appendix
Appendix
See Tables 4, 5, 6 and Fig. 4.
Rights and permissions
About this article
Cite this article
Sarkar, D., Tiwari, J.K., Sharma, S. et al. Production and characterization of somatic hybrids between Solanum tuberosum L. and S. pinnatisectum Dun.. Plant Cell Tiss Organ Cult 107, 427–440 (2011). https://doi.org/10.1007/s11240-011-9993-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-011-9993-8