Abstract
Key message
Genome-wide analysis was carried out to identify and analyze differential expression pattern of tomato membrane bound NAC transcription factors (SlNACMTFs) during stresses. Two biotic-stress-related SlNACMTFs have been characterized to elucidate their regulatory function.
Abstract
NAC transcription factors are known regulators of stress-related gene expression. As Stresses are perceived and transmitted by membrane-bound proteins, functional characterization of membrane-associated NAC transcription factors in tomato can reveal valuable insight about membrane-mediated stress-signalling. Tomato genome encodes 13 NAC genes which have predicted transmembrane domain(s) (SlNACMTFs). mRNA of 12 SlNACMTFs were readily detected in multiple tissues, and also in polysome isolated from leaf, confirming active transcription and translation from these genes occur under normal physiological condition. Additionally, most of the SlNACMTFs were differentially regulated during stresses and stress-related transcription factor binding sites are prevalent in their promoters. SlNACMTF3 and 8 were majorly regulated in biotic and abiotic stresses. Like other MTFs, SlNACMTF3 was translocated to the plasma membrane, whereas the C-terminus truncated (ΔC) form localized in the cytoplasm and the nucleus. Accordingly, the ΔC forms significantly influenced the activity of promoters harbouring NAC binding sites (NACbs). Furthermore, the NAC domain of these transcription factors could directly interact with an NACbs, and the proteins failed to regulate a promoter lacking a crucial NACbs. Interestingly, the type of influence to an NACbs containing promoter was dependent on the context of the NACbs, as the same SlNACMTF showed an alternative mode of regulation on different promoters, as well as the same promoter activity was oppositely regulated by two different SlNACMTF. Finally, both SlNACMTFs demonstrated the differential regulatory effect on the expression of several stress-related genes by interacting with the putative NACbs in their promoter region, suggesting their direct role in plant stress response.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Several defense-related genes display vital roles when plants encounter biotic and abiotic stresses. Activation and expression of these genes are crucial for plants survival and thus, necessarily, be regulated in a controlled manner. This regulation must involve transcription factors which are activated upon a stress exposure and regulate the downstream genes that might help plants to survive during stress. The cellular membrane mainly perceives stresses, and subsequent signal transduction pathways are activated afterward. Several dormant transcription factors are located in the membrane itself which are proteolytically processed for activation upon receiving of stress signal. These transcription factors are referred as Membrane-bound Transcription Factors or MTFs, enter the nucleus and regulate downstream genes. This mode of gene regulation is considered to be a faster and more accurate when compared to the conventional mode of gene regulation (Kim et al. 2007b), as this process skips both transcriptional and translational regulation (Seo 2014). The controlled proteolytic activation of membrane-bound transcription factors is mostly mediated either by intramembrane proteases (regulated intramembrane proteolysis or RIP) or by the regulated ubiquitin/proteasome-dependent processing (RUP) (Seo et al. 2008).
As predicted by genome-wide analysis, over 10% of all Arabidopsis transcription factors are membrane-bound, suggesting, gene regulation mediated by them is a generalized process (Seo et al. 2008). Plant MTFs have been studied mostly in Arabidopsis and Rice, and both genomes contain MTFs that belong to MYB, NAC, bZIP or PHD families (Kim et al. 2010). Among these, the major focus was on bZIP or NAC MTFs. bZIP MTFs are commonly found on ER membrane and related to ER (endoplasmic reticulum) stress, and NAC MTFs are distributed on both intracellular and cellular membranes.
In Arabidopsis, AtbZIP60, a bZIP transcription factor, associated with ER stress or the unfolded protein response, was the first plant MTF to be characterized at the molecular level (Seo et al. 2008). The full length (295 aa) transcription factor undergoes proteolysis when exposed to ER stress and generate a truncated AtbZIP60 (216 aa) which in turn localizes to the nucleus and regulates genes related to ER stress response (Iwata and Koizumi 2005). Another MTF, AtbZIP17, seems to get activated during salt stress, and knock down mutants showed excessive salt sensitivity. A very recently identified membrane-bound bZIP transcription factor in maize, ZmbZIP17 triggered expression of genes responsive to ABA (abscisic acid) as well as ER stress. This MTF interacted with ABREs (ABA-responsive cis-elements) present in promoters of ABA-responsive genes (Yang et al. 2013). Arabidopsis NTM1 is a membrane-bound NAC transcription factor (NAC MTF) which mediates cytokinin signaling during cell division. Active NTM1 enters the nucleus to induce CDK inhibitor genes and represses the histone H4 gene, as a result, reduces cell division (Kim et al. 2006). Rice NAC MTFs referred to as OsNTLs, are mostly regulated under abiotic stress conditions (Kim et al. 2010). All these studies have indicated that MTFs have crucial roles in plants’ physiological signaling and stress tolerance.
NAC transcription factors have vital roles in both biotic and abiotic stress regulation (Olsen et al. 2005a; Wang and Dane 2013). NAC is one of the largest transcription factor family in a plant. The name NAC was derived from three other transcription factors, NAM (no apical meristem) of Petunia, ATAF1–2 and CUC2 (cup-shaped cotyledon) of Arabidopsis, which have the same DNA-binding domains (Aida et al. 1997; Le et al. 2011). In tomato, in a genome-wide analysis, several NAC transcription factors have been identified (Kou et al. 2014), however, few number of NAC transcription factors has been characterized till now. The most worked out NAC in tomato is SlNAC1 which is known to have multifaceted functions. SlNAC1 is induced by chilling, heat, high salinity and osmotic stresses (Ma et al. 2013). SlNAM1, another salt inducible NAC gene, expressed in flower and fruit but not in root. Both SlNAC1 and SlNAM1 are thought to play crucial roles in salt stress tolerance (Yang et al. 2011). SlNAC3, which is mainly expressed in flower, fruit, and root, is regulated under salt and drought. It is also postulated that SlNAC3 might mediate salt stress resistance through ABA signaling by functioning as a transcriptional activator (Han et al. 2012). Tomato NACs have also been implicated in biotic stress response. For example, down-regulation of SlNAC1 expression results in inhibition of viral genome replication as it interacts with Geminivirus replication enhancer (REn) protein (Selth et al. 2005). In addition, SlNAC1 is upregulated in Pseudomonas infection, indicating induced expression may result in resistance (Huang et al. 2013). Two closely related NACs of tomato JA2 and JA2L were found to regulate stomatal closure during pathogen attack, indicating that these NACs help in establishing plant’s innate immunity (Du et al. 2014). Also SlSRN1, probably an MTF, could provide resistance against Botrytis cinerea, a fungal pathogen. Induced expression of SlSRN1 was also observed when plants were treated with SA, JA or ACC, or exposed to drought conditions. However, it was also found to be a negative regulator of abiotic stress (Liu et al. 2014).
Recent studies have implicated NAC proteins in plant developmental signaling. SlNAC1 was found to be involved in fruit pigmentation and softening. Over expressed SlNAC1 can inhibit fruit ripening by interacting with the regulatory region present in the promoter of those genes related to lycopene and ethylene synthesis (Ma et al. 2014). Other newly identified tomato NACs such as SlNAM2 and Goblet genes had been found to have a role in development but so far has not been linked to stress response (Berger et al. 2009; Hendelman et al. 2013). All these studies indicate that specific NAC transcription factors are inseparably related to stress response as well as plant development in tomato, yet, not much information is known about NACMTFs in tomato (SlNACMTFs) till date.
Tomato is an important crop plant and is severely affected by different biotic and abiotic stress factors. Blights and viral infection like leaf curl disease alone can lead to 70% yield loss apart from the loss by abiotic stresses. However, cultivable resistant varieties that could ensure adequate crop production are still unavailable and membrane–originated molecular signaling related to stress tolerance remains elusive. Thus, in this study, a genome-wide analysis was performed to identify all SlNACMTFs, their regulation under different environmental and pathological challenges has been investigated, and biochemical characterization was carried out to confirm their identity and physiological significance.
Materials and methods
In silico analysis for the identification of tomato NAC genes
Amino acid sequence of the NAC domain of AtNTL6 (TAIR: AT3G49530) and AtNAC1 (TAIR: AT1G56010) of Arabidopsis were used as template for protein to protein (blastp) and protein to nucleotide blast (tblastn) in the Sol Genomics database ITAG release 2.4 (https://solgenomics.net/) and both Sol genomics and NCBI Nucleotide databases were searched with terminologies ‘NAC’ and ‘NAC transcription factor’, to identify putative NAC genes in tomato. Results were reanalyzed to check for redundancy and to eliminate pseudo genes (having partial NAC domain). The shortlisted NAC genes were further compared by consulting Plant Transcription Factor Database (http://planttfdb.cbi.pku.edu.cn) for affirmation and identification of additional NAC genes, if any. All these putative NAC proteins were subjected to domain analysis using Prosite (http://prosite.expasy.org/) to confirm the presence of NAC domain. The sequences with only full NAC domain were considered as SlNACs.
To identify SlNACMTFs, the amino acid sequence of these SlNAC genes were analyzed using TMpred (http://www.ch.embnet.org/software/TMPRED_form.html), TMHMM (http://www.cbs.dtu.dk/services/TMHMM/), PHDhtm (http://omictools.com/phdhtm-tool), HMMTOP (http://www.enzim.hu/hmmtop) and only those sequences were considered which were predicted to have transmembrane domain by at least two of these prediction tools with high confidence.
Information about exon and intron positions of SlNACMTFs was obtained from the sequence and location information available in Sol genomics database (https://solgenomics.net/), and gene map was drawn manually.
Sequence alignment and phylogenetic analysis
Sequences of Arabidopsis, rice and potato NACMTFs were collected from NCBI nucleotide and Sol Genomics Network databases. From the collected information the NAC domain sequences were extracted, and multiple sequence alignment along with SlNACMTFs was carried out using ClustalW (http://www.ch.embnet.org/software/ClustalW.html) with default parameters setting. Subsequently, the phylogenetic tree was constructed using MEGA6 by Neighbor Joining method with 1000 bootstrap replicate. Evolutionary distances were calculated using the Poisson correction method.
Analysis of promoter sequence
About 1 kb upstream sequence from the +1 site (A of ATG, predicted translation start site) of each SlNACMTFs was collected from tomato genome sequence database and analysed for identification of putative cis-elements using Matinspector (Genomatix) software. Elements which showed the core similarity and the matrix similarity parameters 1 (maximum possible = 1) and 0.9 (maximum possible = 1), respectively were only marked for subsequent analysis.
Primer designing
Primers were designed manually, and OligoAnalyzer 3.1 software from Integrated DNA Technologies (https://eu.idtdna.com/analyzer/Applications/OligoAnalyzer/) was used to determine the specificity of primers, GC content, annealing temperature, homo and heterodimer and hairpin formation tendency, etc. At least one primer of the pair used in the determination of the level of expression of SlNACMTFs was designed from an exon–exon junction to avoid amplification from the contaminating genomic DNA in the RNA preparation.
Plant material and growth conditions
Solanum lycopersicum L. cultivar Pusa Ruby (originally released by IARI, New Delhi) was used in all experiments. Seeds were initially obtained from Sutton seeds, Kolkata and propagated in house. Seeds were germinated in moistened Soilrite (Keltech, India) and seedlings were grown in green houses, maintained at 25 ± 2 °C, ~70% humidity and in natural light condition. Essential nutrients were supplied in the form of Hoagland solution, poured once in a week.
Isolation of polysomal RNA
Polysomal RNA was isolated by sucrose gradient method described by Bailey-Serres laboratory (Mustroph et al. 2009), with slight modification. Leaves from 30 day old tomato plants (both control and infected) were crushed in liquid nitrogen, and cell debris was removed by centrifugation (10,000g for 5 min). Organellar fraction including polysomes was pelleted by sucrose cushioning (1.75 M sucrose, 72,000g for 18 h, 4 °C). Polysomal fraction was further enriched by sucrose gradient centrifugation (20–60% sucrose gradient, 100,000g for 3 h, 4 °C). Polysomes were typically found in the middle of the gradient. RNA was precipitated from this fraction using guanidine-HCL and absolute ethanol. The pellet was dissolved in Trizol reagent, and RNA was isolated following standard protocol. The integrity of isolated RNA was analyzed by denaturing electrophoresis and quantity was determined by spectrophotometry. Polysomal RNA (2 µg) was reverse transcribed using 200 U RevertAid reverse transcriptase (Fermentas) and random hexamer (5 µM) primers in 20 μl reaction volume, following manufacturer’s (Fermentas) protocol.
Exposure of plants to abiotic stresses
Plants were exposed to abiotic stresses such as salt, drought, heat and cold after 20 days of germination. For salt stress, Soilrite was drenched with 200 mM NaCl (Singh et al. 2012) solution so that it replaced all the residual water with the salt solution and kept in this condition for 6 h. The mock-treated plants were watered only. In the case of drought treatment, tomato plants were uprooted from Soilrite and kept in this condition for 8 h. Mock-treated plants were kept in watered Soilrite. For heat and cold treatments, plants were exposed to 42 °C and 4 °C respectively, for 18 h. To prevent excessive water loss during heat treatment plants were kept wrapped in plastic covers. RNA was isolated after mentioned time of treatments and three biological replicates were analyzed for each case (n = 3 plants for both mock and infected).
Agroinfiltration for ToLCNDV infection
15 days old tomato plants were infected with ToLCNDV (Tomato leaf curl New Delhi virus, genomes ToLCNDV-A, GenBank: DQ629101.1 and ToLCNDV-B, GenBank: DQ169057) (Mandal et al. 2015) by Agroinfiltration technique. Agrobacterium (LBA 4404) harbouring clones of genomes A and B of ToLCNDV were grown in LB media supplemented with Kanamycin (50 µg/ml), Rifampicin (25 µg/ml) and Streptomycin (50 µg/ml) at 28 °C by continuous shaking at 180–200 rpm for 48 h. Cells were harvested afterward, and the pellet was washed and resuspended in MES buffer (10 mM MES, 10 mM MgCl2, pH 5.6). Finally, OD600 of the culture was adjusted to 0.8. Acetosyringone (100 µM) was added to this suspension before infiltration. Fully expanded true leaves of 15-day-old plants were infiltrated in the ventral surface with a 1 ml syringe. Infected plants were separately maintained in the glass house until symptoms were observed (~30 days post infiltration). Control set of plants were infiltrated with Agrobacterium LBA4404 (mock inoculated). DNA and RNA were isolated from newly formed leaves from both systemically infected and control plants. Three biological replicates were analyzed for each case (n = 3 plants for both mock and infected samples and multiple leaves for each sample were taken).
Infection with Alternaria solani
For fungal infection, 1-month-old tomato plants were infected by droplet inoculation method with mycelial suspension (106 spore forming units/10 µl in 0.01% Triton X-100) of A. solani which was grown on a potato-dextrose agar plate. Plants were kept in a humid condition before and after inoculation to facilitate the infection. Mock infected set of plants were spotted with 0.01% Triton X-100 (10 µl) solution only. Yellowing and blight symptoms were observed within 72 h of infection. Only diseased tissues, avoiding necrotic zone, were collected and processed for RNA isolation. Each sample consisted of multiple leaves and three different biological replicates (n = 3 plants for both mock and infected) were used.
Total DNA isolation and rolling circle amplification (RCA) of viral genome
Total DNA was isolated from the leaves of control and infected plants by Urea-lysis method. Newly formed systemically infected leaves were collected 30 days post infiltration for DNA isolation from infected plants. Leaf tissue was ground using liquid nitrogen, suspended in urea-lysis buffer (0.2 M Tris-HCl pH 8.0, 0.5 M NaCl, 2% SDS, 50 mM EDTA and 7 M Urea; 1 ml/mg tissues), and clarified by centrifugation at 10,000g for 5 min. DNA was extracted from the supernatant using an equal volume of Phenol:Chloroform:Isoamyl alcohol mix (25:24:1) and was precipitated from the aqueous phase by ethanol precipitation. RNase treatment was carried out to remove the contaminating RNA and further purification was done by phenol–chloroform extraction method followed by ethanol precipitation. For the screening of infected plants, PCR was done with viral coat protein and movement protein specific primers (CP-F, CP-R and MP-F, MP-R primers, Supplementary table 1) using the DNA samples. To further confirm virus replication within the infected plants, RCA reaction was also carried out with the DNA samples. The reaction mix contained 100 ng DNA, 1× reaction buffer, 1 mM dNTPs, 1.2 µM Random Hexamer primer and 10 U Φ29 DNA polymerase (Thermo Scientific) in a final reaction volume of 10 µl and it was incubated at 30 °C for 18 h. The amplified products were digested with EcoRI enzyme (New England Biolabs) to release the monomers of the viral genome (2.7 kb) from the concatemers produced by the RCA reaction. The digested products were resolved in 1% agarose gel.
RNA isolation and cDNA preparation
Total RNA was isolated from 250 mg leaf tissue of control and treated plants using RNeasy Mini kit (QIAGEN) following manufacturer’s protocol. RNA integrity was checked by resolving the denatured RNA in 1.5% agarose gels (1.5% agarose, 1× TAE) and quantity was determined by spectrophotometry. Complementary DNA was prepared from RNA samples that had distinct 28S and 18S bands showing 2:1 intensity ratio. 5 µg of the total RNA was used for cDNA synthesis following the protocol described previously.
PCR analysis
General PCRs were carried out using Taq DNA polymerase (NEB) in Master cycler Pro S (Eppendorf) machine. The primer pairs which amplified a single band were used in subsequent analysis. Quantitative real-time PCR and analysis were performed using Applied Biosystems 7500 FAST machine, software version 2.3. SYBR® Green based PCR mix (Applied Biosystems) was used in reactions (20 µl) carried out with gene-specific primers. For each sample, three biological replicates and three technical replicates were used. Data was normalized based on the expression level of EF1α transcripts. Data from the reactions which showed the single peak in melt curve analysis, first derivative of fluorescence (dF/dT) versus temperature plot, were only considered. The expression of genes, relative to the expression of mock treated or control (as described in the individual experiment), was calculated for each sample using the 2−ΔΔCT method (Schmittgen and Livak 2008).
Cloning of SlNACMTFs
Coding sequences of SlNACMTF3ΔC, 8ΔC and NAC domains of both the protein were amplified from a control plant cDNA using primers as mentioned in Supplementary table 1 and Pfu DNA polymerase (Fermentas). Amplified products of the desired size were gel purified using QIAquick gel extraction kit as per manufacturer’s protocol. Purified products were then restriction digested with appropriate enzymes. For cloning in the pET28a+ vector of ΔC and NAC domains, PCR products of SlNACMTF3 were digested with EcoRI and SalI, and SlNACMTF8 were digested with SacI and SalI. Double digested PCR products were ligated into the linearized pET28a+ vector in the same sites. To clone in the binary vector pPZPy112, the same protocol was followed but digestions were done with SmaI and SalI enzymes for both the genes. For expression in the bacterial system, Rosseta2 (DE3)pLysS host was transformed with the resulting clones. For expression in the plant system, Agrobacterium LBA4404 was transformed with pPZPy112 based clones.
Generation of GFP fusion protein and imaging
SlNACMTF3 had been predicted to have a transmembrane domain in its C-terminal region. Full length and C-terminal deleted form (SlNACMTF3 and SlNACMTF3ΔC) were cloned in gateway vector pK7WGF2 (https://gateway.psb.ugent.be/search) to generate GFP fusion tag at their N-terminal end. Clones were confirmed by sequencing (GenBank: KX151668). Agrobacterium LBA4404 was transformed with the positive clones and onion epidermal cells were infiltrated with Agrobacterium suspension in the infiltration liquid (41.65 mM d-glucose, 10 mM CaCl2,10 mM MES-KOH pH 5.6, 0.011 µM BAP, 0.05 mM MgCl2, 100 µM acetosyringone) (Xu et al. 2014). Infiltrated onions were incubated at 28 °C for 72 h in the dark. After incubation, cells were fixed with DPX mountant (SRL, India) and imaging was performed using a confocal microscope (Leica TCS SP8) to observe their subcellular localization. DAPI staining was also carried out to locate the nucleus. Agrobacterium (LBA4404) harbouring no recombinant constructs were infiltrated into onion cells to monitor the autofluorescence intensity, which typically did not produce detectable fluorescence (Fig. 6b).
Protein expression and purification
The full length and C-terminal deleted MTFs did not yield any significant amount of soluble protein in bacterial overexpression system. Thus, the NAC domains of SlNACMTF3 and SlNACMTF8 were cloned in pET28a+ vector and Rosseta™ 2(DE3)pLysS bacteria was transformed with these vectors. Bacteria were grown in Terrific Broth (TB, 24.0 g Tryptone, 42.0 g Yeast Extract, 18.6 g Potassium Phosphate dibasic, 4.4 g Potassium Phosphate monobasic in 1 l, pH 7.2) supplemented with antibiotic (Kanamycin 50 µg/ml) at 37 °C for 16 h with a constant shaking of 200–220 rpm. The culture was freshly inoculated in 500 ml TB in 1:500 dilution and allowed to grow in 37 °C for another 3 h or until the culture reached the OD600 of ~0.4. Protein expression was induced with 1 mM IPTG for 3 h in 37 °C. Induced cells were harvested by centrifugation at 8000 rpm, for 5 min, washed and resuspended in 1× binding buffer (20 mM Tris-HCl pH 7.9, 500 mM NaCl). The cells were disrupted using a French press (20 Pa, 2 pulses) and centrifuged at 12,000 rpm for 30 min to pellet the insoluble fraction. The soluble part was filtered through 0.45 micron membrane prior to the purification via Ni-NTA affinity chromatography.
Ni-NTA Agarose beads (Qiagen, USA) were used in affinity purification of His-tagged proteins. Beads were washed with 1× binding buffer (20 mM Tris-HCl pH 7.9, 500 mM NaCl) for several times prior to loading. The clarified soluble protein part was mixed with Ni-NTA beads of bed volume of 1.5 ml, incubated at 4 °C for 1 h and collected by low speed centrifugation. Beads were resuspended in 3 ml of binding buffer containing 10 mM imidazole and loaded onto a column. The column was repeatedly washed with wash buffer with increasing concentrations of imidazole (20 mM Tris-HCl pH 7.9, 500 mM NaCl and 40–100 mM Imidazole). Protein was eluted in an elution buffer (20 mM Tris-HCl pH 7.9, 500 mM NaCl and 500 mM Imidazole) in several aliquots and quality was checked by SDS-PAGE. The eluted fractions which contained the reasonably purified protein were pooled and dialyzed in a dialysis buffer (20 mM HEPES pH 7.6, 2 mM DTT, 1 mM EDTA, and 0.5 mM PMSF) for 24 h at 4 °C with three changes. Protein was concentrated using a concentrator (Vivaspin Turbo® 4, Sartorious). The concentration of the purified protein was estimated using Bradford reagent and purity was determined by SDS-PAGE analysis. About 80% pure protein could be regularly obtained.
Electrophoretic mobility shift assay
Complementary single-stranded oligonucleotide (10 nmol) of each DNA duplexes (Fig. 7a) were mixed, in a reaction mixture containing 40 mM Tris-HCl (pH 7.5), 50 mM NaCl and 20 mM MgCl2 to a final volume of 20 µl. The mixture was heated at 95 °C for 10 min and cooled down to room temperature slowly for efficient annealing. Annealed duplexes were analyzed by Polyacrylamide gel electrophoresis (7% PAGE) and band of the desired size was cut and gel purified. About 10 pmol of duplex DNA (~200 ng) was used for end-labeling with 15 U of T4 PNK (NEB) and 25 µCi γ-32P-dATP (BRIT, India) in a total volume of 15 µl according to the enzyme manufacturer’s (NEB) protocol. The labeled probes were purified using Sephadex G25 quick spin mini column (Illustra microspin G25, GE, UK) and specific activity of ~2.5 × 106 cpm/pmol was routinely obtained.
In vitro protein translation was performed with TnT T7 coupled Rabbit Reticulocyte lysate (RRL) system (Promega, USA). The proteins were expressed from pET28a+:SlNACMTF3ΔC and pET28a:SlNACMTF8ΔC in a 10 µl reaction, containing 5 µl of RRL mix, 1× reaction buffer, T7 RNA Polymerase, 1 mM methionine, 1 mM amino acid mix and 0.5 µg plasmid DNA, incubated at 30 °C for 90 min. A control reaction was performed with only pET28a+ empty vector. Protein expression from the clones was confirmed in a parallel reaction carried out with radiolabelled 35S-Methionine.
Binding reactions were done following a standard protocol (Kundu et al. 2015). A typical binding reaction contained labeled DNA (~50,000 cpm) along with varying amount of proteins. The EMSA reactions were performed with, (i) 1–4 µl of RRL-mix including the expressed protein, or (ii) purified protein (500–2000 ng) in a reaction buffer contained 5% glycerol, 20 mM HEPES (pH 7.5), 100 mM KCl, 0.2 mM EDTA, 2 mM MgCl2, 0.5 mM PMSF, 1 mM DTT and 1 µg poly (dI-dC)·poly(dI-dC) in a final reaction volume of 15 µl. In control reactions, mock translation performed with RRL-mix, or purified tag protein was added. Competitor molecules were added during the incubation stage where needed. The reaction mixes were incubated at room temperature for 30–40 min prior to resolving in a 0.5X TBE-5% Polyacrylamide gel at 140 V, 4 °C for about 1.5 h. Gels were fixed in 40% methanol and 10% acetic acid solution for 30 min, dried, and exposed to Phosphor imager screen for imaging (Typhoon, GE Healthcare).
Promoter activity assay
The full-length CaMV35S promoter located upstream to the GUS gene in pCambia 1304 vector contains two NAC binding sites (NACbs) (Fig. 8a). We have utilized this promoter and a truncated (up to –104 nt from putative transcription start site) form of the promoter, which contains one of the two NACbss (Fig. 8a), to assay the transcriptional regulatory function of NACMTFs. These two promoters should drive the expression of downstream GUS reporter gene. To understand the activities of promoters under the influence of SlNACMTFs, Agrobacterium mediated transient co-transformation assays were carried out. On fully expanded leaves of tobacco (Nicotiana tabacum, about one month old) plants Agrobacterium harboring pPZPy: SlNACMTF3ΔC or pPZPy:SlNACMTF8∆C constructs were coinfiltrated with full length or truncated CaMV35S promoter constructs. Approximately 200 µl of the bacterial suspension (as described previously) was infiltrated through the intercellular space of the abaxial surface of the leaf with the help of a 1 ml syringe. The plants were kept under controlled environment for 48 h, and then tissues were collected from the infiltration site (~2 cm2), crushed in liquid nitrogen and suspended in 250 µl of GUS extraction buffer (50 mM NaHPO4, pH 7; 10 mM 2 mercapto ethanol; 10 mM Na2EDTA; 0.1% SDS; 0.1% triton X-100). The suspension was then centrifuged at high speed to remove cellular debris, and 50 µl of the supernatant was mixed with 250 µl of GUS assay buffer (GUS extraction buffer supplemented with 2 mM MUG). The mixture was incubated at 37 °C for 1 h. 100 µl of the reaction mix was then used to measure fluorescence (excitation at 365 nm and emission at 455 nm, readings were taken for 5 s) of the product of GUS enzymatic reaction using Varioskan flash multimode plate reader (Thermo scientific, Finland). The concentration of protein for each sample was determined using Bradford reagent (Bio-Rad, CA, USA) and data was normalized using protein concentration. A standard curve for MU (4-methylumbelliferone) was also prepared and GUS activity was presented as the amount of MU produced in 1 h from 1 µg protein.
Agroinfiltration in tomato leaves and RT-PCR analysis
One month old tomato plants with fully expanded leaves were agroinfiltrated with Agrobacterium harboring pPZPy: SlNACMTF3∆C and pPZPy: SlNACMTF8∆C constructs. Plants were kept in a controlled environment for 36 h and then ~100 mg tissues were collected from the infiltration sites. RNA was isolated using RNeasy Mini kit (QIAGEN) following manufacturer’s protocol. 5 µg of the total RNA was used for cDNA synthesis. Gene specific primers were used to analyze the expression level of selected genes.
Statistics
All experiments were performed with three biological replicates. Real time PCR analyses were performed with three different RNA preparations and reactions were also done in triplicates. In Agroinfiltration assays, infiltration was done in at least 4 different spots in multiple leaves of different plants and samples were pooled whenever needed. Results are expressed as means ± standard error (SE). Two-tail Student’s t test was employed if needed, and p ≤ 0.05 considered as significant.
Results
Identification and structural derivation of tomato NAC genes
Genome-wide analysis of tomato NAC transcription factors have been reported recently (Hongyan et al. 2015; Kou et al. 2014). However, in these reports NAC MTFs are not defined. We initiated this study with identification of tomato NACMTFs. Prior to the identification of NAC MTFs we carried out an exhaustive search of tomato genomic database again for tomato NAC transcription factors using alternative approaches. This has ensured inclusion of newly described gene sequences for the identification of additional NAC TFs. We have performed, (i) extensive search of Sol Genomics database ITAG release 2.4 (https://solgenomics.net/) and NCBI Nucleotide database with terminologies ‘NAC’ and ‘NAC transcription factor’ linked to tomato, (ii) protein to protein (blastp) and protein to translated nucleotide (tblastn) BLAST analyses using the NAC domain amino acid sequences of AtNAC1 and AtNTL6, as both of them are well characterized NAC genes of Arabidopsis thaliana, and (iii) compilation of NAC TFs sequences described in PlantTFDB (http://planttfdb.cbi.pku.edu.cn). Homology search resulted in 131 genomic hits, which were further filtered to avoid redundancy and to select only highly homologous sequences (e ≤ 1–10). Each sequence was reanalyzed in Prosite to confirm the existence of NAC domain, and sequences devoid of NAC domains were eliminated. Finally, our analysis yielded 99 NAC transcription factors (Solanum lycopersicum NAC Transcription Factor or SlNACTF) genes encoded in tomato genome (Supplementary table 2). Among these, two SlNACTFs (Solyc06g083840, Solyc05g007771) were not previously reported. SlNACTFs are distributed among all 12 chromosomes but are slightly more clustered on chromosome 2, 3 and 6 (Supplementary Fig. 1). Sequence details of all SlNACTFs were obtained from Sol Genomics, and subsequent analyses revealed that these transcription factors were coded by a widely varied length of genes, could contain 1–18 exons and expressed as about 387 nt to 3305 nt long mRNAs (Supplementary table 2).
Although these SlNACTFs are extremely variable in size, they contain a highly conserved NAC domain (120–150 amino acids long), mostly (95 out of 99) full length, at their N-terminal end. However, the C-terminal regulatory domains vary, to a great extent, in size and sequence. Further analysis using MEME 4.10.2 to detect conserved motifs within the NAC domain with stringent parameters settings indicated that the NAC domain of these SlNACTFs contains five conserved motifs (Supplementary Fig. 2). These motifs are typical to the NAC domain and crucial for DNA binding as well as dimerization of the protein.
Identification of membrane-bound SlNACTFs (SlNACMTFs)
The SlNACTF genes were further analyzed using several transmembrane-domain prediction software, such as TMpred, TMHMM, PHDhtm, and HMMTOP. Only those are considered as MTFs for which at least two out of the four prediction software gave an affirmative result. These software predicted 13 SlNACMTFs (Fig. 1a) from the NACTF genes in tomato. SlNACMTFs had a wide variation in the length of a gene as well as CDS. All these MTFs contain NAC domain at their N-terminal region, whereas, the location of the transmembrane domain (TM) varied. Out of 13 SlNACMTFs in 8 the TM was found in C-terminus and in 3 at N-terminus. In two SlNACMTFs the TM overlapped with the NAC domain, and Solyc11g068750 contained double TMs (Fig. 1b). For the ease of explanation henceforth these SlNACMTFs are termed as SlNACMTF1 to 13 (Fig. 1b).
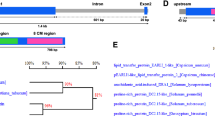
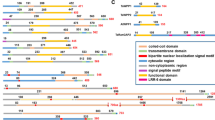
Gene structure, protein domains and phylogenetic relatedness analysis of SlNACMTFs. a Gene maps of SlNACMTFs showing exon and intron numbers and their relative length. Exon, black box and intron, thin lines are shown. b Schematic diagram of SlNACMTF proteins depicting different domains. Black and gray boxes indicate the NAC and TM domains, respectively. c Phylogenetic relationship analysis of SlNACMTFs along with Arabidopsis, rice and potato NACMTFs. Six clusters are demarcated with boxes
SlNACMTFs are evolutionarily related to NACMTFs of Arabidopsis, rice, and potato
The NAC domains of SlNACMTFs, AtNACMTFs, OsNACMTFs and StNACMTFs, as predicted by Prosite, were extracted and multiple sequence alignment was performed with ClustalW. Aligned sequences were analyzed in MEGA6 to understand the evolutionary relatedness of these NACMTFs. This analysis revealed that most of the SlNACMTFs could be categorized into four groups along with Arabidopsis, rice, and potato NACMTFs (Fig. 1c), and 3 SlNACMTFs were grouped with one of the potato NACMTFs, suggesting this group of MTFs is Solanaceae specific. Only two SlNACMTFs (SlNACMTF6 and 10) could not be clustered with other MTFs and remained as a separate group, indicating they might have originated by a slightly diverse path of evolution and are tomato specific. The apparent relatedness also suggests the conserved functionality of NACMTFs.
SlNACMTF genes undergo active transcription and translation under normal physiological condition
Transcript expressions of the predicted SlNACMTFs were analyzed in different tissues by RT-PCR. Primers were designed (Supplementary table 1) for each MTF based on the sequences available from Sol Genomics database and expression of EF1α was used for normalization. Our analysis confirmed that expressions of 12 SlNACMTFs could be detected in all three samples (root, hypocotyls, and young leaf) tested, and they show differential expression pattern in these tissues (Fig. 2a). Expression of SlNACMTF6 was detectable in root and hypocotyl, however, we could not detect its expression in leaf samples in spite of repeated trials with different primers and different reaction conditions, indicating the gene may not be ubiquitously expressed. To determine the relative level of expression of these genes in young leaf tissues (n = 3 plant samples), we performed quantitative RT-PCR experiment. In general, most of the SlNACMTFs express in relatively very low amount (0.00005 to 0.006 fold relative to EF1α), except SlNACMTF1, SlNACMTF2 and SlNACMTF 3, which showed relatively higher expressions ranging from 0.018 to 0.49 fold relative to EF1α (Fig. 2b, c).
Expression pattern of SlNACMTFs in different tissues. a Typical Agarose gel pictures of resolved bands of RT-PCR product of different SlNACMTFs, showing relative level of mRNAs in different tissue samples such as root (5 days after germination, DAG), hypocotyl (5 DAG) and young leaves (~10 DAG). b Quantitative RT-PCR data performed with RNA samples from mature leaf tissue (from 1 month old plant) also shows relative level of SlNACMTFs’ expression in normal physiological condition. The expression level of SlNACMTFs relative to the expression of EF1α is plotted
Next, we examined whether these MTFs are also translated under the normal physiological condition in leaf. We isolated polysomal RNA from tomato leaf tissues using sucrose gradient purification method (Mustroph et al. 2009), purified RNA was converted to cDNA and used for PCR-based detection of NACMTFs. In this way we could amplify 10 out of 13 SlNACMTFs (Fig. 3a), confirming that these genes are not only transcribed but also translated. Moreover, SlNACMTFs exhibited a different pattern of inclusion in polysomes compared to the pattern of mRNA expression. Purity and integrity of isolated polysomes and associated RNA were appreciable as primary miR167, or U6 RNA bands were undetectable from cDNA prepared from the isolated polysomal RNA, however, EF1α mRNA could be readily detected in the cDNA but not in mock cDNA prepared without reverse transcriptase (Fig. 3c). Concomitantly, EDTA treatment of polysomes, resulted in the loss of inclusion of mRNAs, confirming monosomal mRNA contamination was minimal in these samples (data not presented).
Detection of SlNACMTF RNAs in polysome complex. a Agarose gel photograph showing RT-PCR products of SlNACMTFs obtained from polysomal RNA. The differences in band intensities also suggest these mRNAs produce a differential amount of proteins. b Pri-miR167 or U6 snRNA could not be amplified, whereas EF1α was readily amplified from the same polysomal RNA preparation, (+RT lanes), indicating the preparation was enriched of polysomal RNA and devoid of free/monosomal mRNA. Also, no amplification product including EF1α, was observed from RT-independent PCR reactions (−RT lanes), confirming specific amplification observed only from cDNAs prepared from polysomal RNA and not from contaminating DNA
Differential regulation of SlNACMTFs occurs during stress
NAC genes are commonly regulated by stresses (Wang and Dane 2013). The role of NAC and NACMTFs in abiotic stress response is explored, however, their role in biotic stress response is appreciated recently (Nuruzzaman et al. 2013). Particularly, the role of SlNACMTFs in stress response has not been elucidated yet.
Regulation of expression of SlNACMTFs during abiotic stresses
Tomato plants were exposed to different abiotic stresses such as salt, drought, cold and heat and transcript level of SlNACMTFs were studied. To confirm that plants were indeed encountered stress, expression of some marker genes for each stress (ABA2 and NCED for salt and drought stress; Hsf3A and CBF1 for heat and cold stress, respectively) was also studied by performing a semi-quantitative RT-PCR (Fig. 4a, d inset), and it was observed that the marker genes were regulated upon exposure to different stresses. Next, semi-quantitative RT-PCR analysis was carried out using cDNA prepared from mock and stresses exposed plants’ RNA to determine which of these SlNACMTFs were regulated under the above-mentioned stresses (data not shown) and subsequently, quantitative real-time PCR was carried out for those genes only.
For salt stress, Soilrite, in which plants (~20 days old) were grown, was inundated with 200 mM NaCl and maintained in this condition for 6 h. The mock-treated plants were watered only. After 6 h, RNA was isolated from mock and salt-treated plants (n = 3 plants for both mock and infected). The result obtained (Fig. 4a) showed that SlNACMTF 2, 8 and 11 were significantly downregulated (0.33 fold, p ≤ 0.02; 0.14 fold, p ≤ 0.04 and 0.1 fold, p ≤ 3.6 × 10−5, respectively), while SlNACMTF10 and 12 exhibited 6.2 (p ≤ 5.3 × 10−7) and 1.6 (p ≤ 0.0004) folds upregulation.
Determination of SlNACMTFs’ expression level in tomato plants exposed to various abiotic stresses. Real time RT-PCR data depicting the relative level of mRNA in salt, drought, cold and heat treated plants compared to that of mock treated plants. Significant change in expression level of SlNACMTF2, 8, 10, 11 and 12 are observed in salt-treated plants when compared to mock treated plants (a). In the case of drought treatment, SlNACMTF2, 8 and 10 shows significant regulation in mRNA level (b). ABA2 and NCED used as marker genes for salt and drought stresses, respectively, show upregulation in respective stress conditions (a, inset). Cold treatment resulted in a significantly high expression level of the SlNACMTF2 transcript, while all other SlNACMTFs remained unchanged (c), and SlNACMTF1, 11, 12 are regulated by heat stress (d). Expression of marker genes for cold (CBF1) and heat (Hsf3A) are shown in the inset. Data was normalized based on the expression level of EF1α; *Indicates a significant change
To expose plants to drought stress, tomato plants (~20 days old) were uprooted from Soilrite and kept in this condition for 8 h. Mock-treated plants were kept in watered Soilrite. After 8 h, RNA was isolated (n = 3 plants for both mock and infected), and cDNA was prepared. Quantitative real-time PCR data (Fig. 4b) obtained clearly shows SlNACMTF2 and SlNACMTF8 both were significantly upregulated (1.4 fold, p ≤ 0.004 and 29 fold, p ≤ 0.0002, respectively). SlNACMTF10, on the other hand, was significantly downregulated (0.57 fold, p ≤ 0.001).
For heat and cold treatments, plants were exposed to 42 °C and 4 °C respectively, for 18 h. To prevent excessive water loss during heat treatment plants were kept wrapped in plastic covers. Results of quantitative real-time PCR evidently indicate that SlNACMTF2 expression level was upregulated (1.9 fold, p ≤ 0.02) during cold stress (Fig. 4c) whereas expressions of other MTFs remained unaltered. Heat treatment caused significant downregulation of SlNACMTF11 and 12 only (0.09 fold, p ≤ 0.04 and 0.06 fold, p ≤ 0.046, respectively) (Fig. 4d). From these observations, it is safe to conclude that different SlNACMTFs may have significant roles in alternative abiotic stresses to which plants are exposed.
Regulation of expression of SlNACMTFs during biotic stresses
Tomato plants were infected with tomato leaf curl New Delhi virus (ToLCNDV), a biotrophic pathogen, or A. solani, a necrotrophic pathogen, and gene expression was studied after infection was established. ToLCNDV infection was confirmed by (i) PCR-based amplification of viral coat protein and movement protein genes, and (ii) rolling circle amplification (RCA) of the viral genome from total DNA isolated from systemically infected leaf tissues (Fig. 5a). Mock inoculated (only LBA4404) plants did not show any stress symptoms. Alternaria infected plants developed blight symptoms at about 72 h post inoculation and mock-inoculated (inoculated with water) plants did not develop any symptom (Fig. 5e inset).
Determination of SlNACMTFs’ expression level in tomato plants infected with pathogens. (a, left panel) Photograph of mock-inoculated (infiltrated with only Agrobacterium LBA4404) and ToLCNDV infected leaves. The infected leaf is showing typical leaf curling symptoms developed in laboratory condition. (a, middle panel) Agarose gel picture showing ToLCNDV coat protein (CP) and movement protein (MP) specific PCR amplified products from systemically infected samples. (a, right panel) Digested products of RCA resolved in an agarose gel. The arrow indicates a monomer of the viral genome. b Resolved SlNACMTF specific PCR products (30 cycles) showing the relative amount of mRNAs in leaves of plants mock inoculated and systemically infected with ToLCNDV. c Real time PCR analysis data, performed with ToLCNDV-stress regulated selected SlNACMTFs, confirming differential expression in these samples. d Resolved semi-quantitative RT-PCR (30 cycles) products in 2.5% Agarose gel showing relative expression level of SlNACMTFs in mock inoculated (with only water) and Alternaria infected leaves (photograph shown in inset). Quantitative real-time RT-PCR data, performed with total RNAs from the leaf samples confirm significant fold change in mRNA expression of SlNACMTF1, 2, 5, 8 and 11 (e); and carried out with polysomal RNA (f), also confirms increased inclusion of SlNACMTF2, 3 and huge increase in the level of inclusion of SlNACMTF8 occurs during Alternaria infection. Data was normalized based on the expression level of EF1α, and *indicates a significant change
Initially, semi-quantitative RT-PCR analysis was carried out using cDNA prepared from mock and ToLCNDV infected plants’ RNA (n = 3 plants) as template and gene specific primers. The results indicated that SlNACMTF1, 2, 3, 5, 8 and 11 were differentially regulated (Fig. 5b). For further confirmation, quantitative real-time PCR was carried out for these SlNACMTFs. Data presented in Fig. 5c conclusively show that SlNACMTF3 and 11 were significantly upregulated (3.6 fold, p ≤ 0.006 and 2.4 fold, p ≤ 0.0006, respectively), whereas SlNACMTF5 showed slight but significant downregulation (0.6 fold, p ≤ 0.008).
Only infected tissues (~72 h post infection), devoid of necrotic lesions and fungus mass, from Alternaria infected plants (n = 3 plants) were taken for differential gene expression analyses. Semi-quantitative RT-PCR data indicated that SlNACMTF1, 2, 3, 5, 8, 11 and 13 were differentially expressed during the infection (Fig. 5d). For further validation, quantitative real-time RT-PCR was done only for these MTFs. Result indicated that SlNACMTF1 (twofold, p ≤ 0.002), SlNACMTF2 (2.8 fold, p ≤ 0.00000005), SlNACMTF8 (7.8 fold, and p ≤ 6 × 10−10) and SlNACMTF11 (2.3 fold, p ≤ 0.04) were significantly upregulated, and SlNACMTF5 was slightly downregulated (0.76 fold, p ≤ 0.03) during Alternaria infection (Fig. 5e). The regulation at the level of gene expression ultimately leads to the change in the level of translated protein. To further discern that the huge upregulation of expression of SlNACMTF8 also leads to its inclusion in polysomal RNA for active translation, real time RT-PCR analysis was performed with polysomal RNA isolated from mock and infected tissue samples. Data presented in Fig. 5f clearly show that SlNACMTF8 transcripts are highly abundant (216.4 fold, p ≤ 0.002) in polysomal RNAs of infected tissues. Higher amount of inclusion of SlNACMTF2 (twofold, p ≤ 0.007) and SlNACMTF3 (1.6 fold, p ≤ 0.0001) was also noticed. These results indicate that SlNACMTFs are not only transcriptionally regulated during biotic stresses, but also undergoes an elevated level of active translation, particularly during fungus stress. As expressions of SlNACMTF3 and SlNACMTF8 are majorly changed during virus and fungus stresses, respectively, and the role of SlNACMTFs in biotic stress is relatively unexplored, further characterizations of these two MTFs were carried out.
SlNACMTF3 is localized both in membrane and the nucleus
The SlNACMTFs possess at least one putative transmembrane domain. To experimentally verify that the prediction was valid and a full-length protein could be localized to the membrane, we used GFP-tagged MTF in our localization studies. A GFP tag was translationally fused at N-terminal of SlNACMTF3 and cloned into a plant expression vector. Also, the C-terminal deleted fragments were cloned in this manner, which represent the membrane detached form and should not be found exclusively in the membrane (Fig. 6a). Agrobacterium LBA4404 harboring positive clones were infiltrated in onion epidermal cells and visualized using a confocal microscope. Acquired images clearly indicated that full-length proteins were distinctly localized in the cellular membrane (Fig. 6b, c). Upon dissociation of plasma membrane from cell wall as induced by dehydration, the GFP-fusion full-length proteins remained attached to the membrane, which further proves that these proteins are exclusively localized to the membrane (Fig. 6d). On the contrary, the C-terminal deleted proteins (GFP-SlNACMTF3ΔC) were mostly found in the cytoplasm and also in the nucleus in a significant amount (Fig. 6e; Supplementary Fig. 4). These data strongly suggest that full-length SlNACMTF3 is localized to membrane under the normal physiological condition and also SlNACMTF3ΔC is capable of being translocated to the nucleus as it lacks the TM-domain.
Subcellular distribution of SlNACMTF3. N-terminal GFP-fusion proteins of full-length (GFP-SlNACMTF3) and C-terminal truncated forms (GFP-SlNACMTF3ΔC) of SlNACMTF3 gene was used (a). GFP-SlNACMTF3 is localised specifically in the membrane (b) and magnified picture also proves the same (c). Plasmolysed cells show that the localisation occurred exclusively on the membrane (red arrow) but not in the cell wall (d). e GFP-SlNACMT3ΔC localised both in the nucleus (red arrow) and cytoplasm
SlNACMTF3 and SlNACMTF8 specifically interact with a NAC binding site
The existence of a NAC domain in both of these sequences has been predicted with a confidence score of 53.8 and 53.5 using Prosite for SlNACMTF3 and SlNACMTF8, respectively. To confirm that these proteins are indeed able to interact with a consensus NAC binding site (NACbs) (Fig. 7a) we have performed EMSA analyses. For this purpose, C-terminal deleted NACMTFs (NACMTF3ΔC and NACMTF8ΔC) were used, as it is well known that the C-terminus (including the TM-domain) somehow inhibits DNA-binding ability of a NACMTF. Both NACMTF3ΔC and NACMTF8ΔC proteins were expressed in vitro using rabbit reticulocyte lysate (RRL) translation system. The translation mix was used in binding assays with proper control. Our data (Fig. 7b, c) clearly show that both MTFs in RRL mix have efficiently interacted with NACbs, however, no shifted band was visible in EMSA reactions performed with control RRL mix.
Specific interaction of SlNACMTF3 and 8 with the consensus NAC binding site (NACbs). a Sequence information of consensus NACbs (NACbs) and mutant NACbs (mutNACbs) duplexes used in this study. Core sequences are in bold font; mutated nucleotides are italicized and underlined. b EMSA assays performed with in vitro translation mix (RRL mix) containing SlNACMTF3ΔC protein, or c SlNACMTF8ΔC protein and labelled NACbs. Control reactions were performed with BSA. Arrow, shifted band. d SDS–PAGE gel stained with Coomassie brilliant blue, showing purified protein band (arrow) of NAC domain of SlNACMTF3 (NAC-MTF3). One µg and three µg proteins loaded in lane 1 and 2, respectively. The same purified protein shows a single band when denatured in the presence of DTT instead of β-mercaptoethanol (d, lane 3). e Purified protein band (arrow) of NAC domain of SlNACMTF8 (NAC-MTF8) is visible in a similar gel photograph. f Autoradiogram of EMSA reactions performed with NAC-MTF3. Lane 1, negative control, purified tag protein from pET28a backbone used in the reaction instead of the NAC-MTF3 which shows no interaction with labelled NACbs. Purified NAC-MTF3 produced prominent shifted band in a dose-dependent manner (lanes 2, 3 and 4) and was unable to bind mut-NACbs (lane 5), indicating specific interaction with NACbs occurs. Competition with unlabelled NACbs (10X) produced shifted band of lesser intensity (lanes 6), while unlabelled mut-NACbs failed to compete with labelled NACbs (lane 7), further confirming specificity of NAC-MTF3-NACbs interaction. g Autoradiogram of EMSA reactions performed with NAC-MTF8. Lane 1, negative control reaction as described above. NAC-MTF8 interacted with NACbs (lane 2) but not with mutNACbs (lane 3). Specificity of the interaction was further confirmed by competition assays in which unlabelled NACbs effectively competed with labelled NACbs in a dose-dependent manner (10X and 50X, lanes 4 and 5, respectively), however, unlabelled mutNACbs failed to compete (lane 6). Arrow indicates the position of shifted bands
Our next goal was to perform the same EMSA reactions with purified C-terminus deleted MTFs. However, repeated trials with alternative protocols failed to yield active proteins. Thus, we cloned DNA binding domain of SlNACMTF3 and 8 (NAC-MTF3 and NAC-MTF8) in bacterial overexpression vector. These clones were expressed in bacteria and purified by Ni-NTA affinity chromatography. Purified proteins (~78% for NAC-MTF3 and ~75% for NAC-MTF8, Fig. 7d, e) were further used in DNA binding assays. Control protein for these reactions was a purified polypeptide expressed from an empty expression vector (pET28a+) and a mutated NACbs sequence was used as negative control for DNA duplex. Specific interaction of NAC-MTF3 and NAC-MTF8 (arrow, Fig. 7f, g) were confirmed by (i) production of shifted band(s) resulted in reactions performed with labeled NACbs, (ii) inability of control protein to interact with the NACbs, and (iii) purified NAC-MTF3 and NAC-MTF8 did not interact with the mutated NACbs. Moreover, protein dose-dependent increment in binding was also observed when protein concentration was raised from ~20 pmol to ~80 pmol (lanes 2–4, Fig. 7f). To further confirm specific interactions, competition assays were carried out with unlabeled probes. The intensity of the shifted bands decreased when excess unlabeled NACbs oligo was added in EMSA reactions (lane 6, Fig. 7f and lanes 4–5, Fig. 7g), but not when mutated oligo was added in the reactions (lane7, Fig. 7f and lane 6, Fig. 7g). These assays confirmed that NAC domains of SlNACMTF3 and 8 could specifically interact with a consensus NACbs that has been tested.
SlNACMTF3 and SlNACMTF8 regulate transcription from NACbs-containing promoter
NAC transcription factors are known trans-acting regulators of several stress-related genes (Hu et al. 2006). To understand whether these SlNACMTFs have the ability to control the expression of a gene that has NACbs in its promoter, promoter activity assays were carried out. Full-length SlNACMTF3 and 8 should be transcriptionally inactive thus C-terminal deleted (SlNACMTF3ΔC and SlNACMTF8ΔC) fragments were cloned into a plant expression vector for transient expressions in planta. pCambia 1304 vector was employed in our assays because it contained the full-length CaMV35S promoter which possesses two NACbss (AGACGTTCCAACCACGTC and TGACGTAAGGGATGACGCAC) around −114 and −60 nt upstream of the transcription start of the promoter followed by the GFP-GUS fusion reporter gene (Fig. 8a). Besides, we constructed another promoter-construct in which only one of the NACbss (TGACGTAAGGGATGACGCAC) was present. Coinfiltration of promoter constructs along with full-length and truncated SlNACMTFs clones in tobacco leaves followed by MUG assay revealed insightful information about the regulation of genes by these NAC transcription factors. Results obtained clearly suggested that SlNACMTF3ΔC significantly downregulate the expression of GUS gene from the CaMV35S promoter containing both NACbs whereas SlNACMTF8ΔC significantly upregulate the GUS expression from the same promoter (Fig. 8b, lanes d vs. e and i vs. j). However, NAC proteins failed to regulate activities significantly from the truncated promoter (Fig. 8b, lanes b vs. c and g vs. h), although SlNACMTF8ΔC showed a trend of upregulating the truncated promoter. These results prove that C-terminal deleted (lacking the TM-domain) SlNACMTF3 and SlNACMTF8 are capable of actively regulating transcription from a promoter that contains NACbs. However, the effect is seemed to be dependent on the type of NACbs and its context.
Transcriptional regulatory function of SlNACMTFs. a Reporter constructs used for the assay of promoter activity. Types of NACbs present in full length and truncated CaMV35S promoters are indicated. NACbs, NAC binding site. b GUS activity assays performed with coinfiltrated tobacco leaf tissues. a and f are different sets of negative controls, showing only background fluorescence. Activities from truncated CaMV35S promoter (b, g); truncated CaMV35S promoter when coinfiltrated with SlNACMTF3ΔC or SlNACMTF8ΔC (c, h); CaMV35S promoter (d, i) and CaMV35S promoter coinfiltrated with SlNACMTF3ΔC or SlNACMTF8ΔC (e, j) are indicated. c, d The expression level of stress-related endogenous genes in leaves with an augmented level of SlNACMTFs. Quantitative RT-PCR data shows expression of Senescence associated protein, PR1b, Elicitor responsive protein 1 and Avr-9/Cf-9 genes in tomato leaves infiltrated with either SlNACMTF3ΔC (c) or SlNACMTF8ΔC (d). *Significantly different from control (mock infiltrated with LBA4404:pPZPY112) and data was normalized based on the expression of endogenous EF1α. Interaction of NAC-MTF3 protein with the NACbs(s) present in the promoters of Senescence associated protein (SAP1), Avr-9/Cf-9, Elicitor responsive protein 1 (Eres1) and PR1b genes (e)
SlNACMTF3 and SlNACMTF8 affect expression of stress-related genes having NACbs in their promoters
Initially, we had shortlisted some known stress-related genes for analysis of the existence of NACbs in their putative promoter region. Amongst these, four genes were selected (Senescence associated protein, PR1b, Avr9/Cf-9, Elicitor responsive protein 1) for further analysis because of the detection of NACbs in their promoter region after stringent analysis (core similarity 1 and matrix similarity 0.9, where 1 is the most stringent in both the cases) (Supplementary table 3). To understand whether these genes were indeed regulated in the presence of an active NAC transcription factors, the SlNACMTFs were ectopically expressed in tomato leaves, followed by RT-PCR based expression analyses were conducted. Agrobacterium LBA4404 harbouring SlNACMTF3 and SlNACMTF3ΔC genes in plant expression vectors were infiltrated in tomato leaves, and empty LBA4404:pPZPY112 was alone infiltrated in tomato leaves to be used as a control. After 36 h, RNA was isolated from the infiltrated tissues, cDNA was prepared, and quantitative RT-PCR was done using gene specific primers designed for the selected genes. The SlNACMTF3ΔC showed a differential effect on different genes tested (Fig. 8c). Senescence associated protein was slightly downregulated, whereas PR1b and Elicitor responsive protein 1 showed significant downregulation (0.68 fold compared to control, p ≤ 0.009 and 0.23 fold compared to control, p ≤ 7.9 × 10−6, respectively). On the contrary, the expression of Avr9/Cf9 gene was increased significantly (33.14 fold over control, p ≤ 0.006). SlNACMTF8ΔC, on the other hand, downregulated the expression of all the four above mentioned genes. PR1b, Elicitor responsive protein 1 and Avr9/Cf-9 genes were significantly downregulated to an extent of 0.3 fold (p ≤ 0.001), 0.33 fold (p ≤ 0.001) and 0.3 fold (p ≤ 0.001) respectively compared to the control. Senescence associated protein was also downregulated 0.57 fold but not significantly. Moreover, purified NAC-MTF3 protein could directly interact with the NACbs(s) located in the promoter-regulatory regions of all four genes tested (Fig. 8e).
The results obtained clearly reveal that the selected SlNACMTFs were indeed able to regulate the expression of endogenous genes that contain NACbs(s) in their promoter, and moreover, only NACMTFs devoid of their TMs were transcriptionally active. The data also suggest that overexpression of these NAC genes somehow mimic stress condition in the plant.
Discussion
Activities of a transcription factor (TF) are regulated at several steps including transcriptional, post-transcriptional, translational and secondary modifications. In addition to these, one of the recently understood mechanisms of activation is the release of a dormant form from storage. Transcription factors belonging to several groups are found tethered to either plasma membrane or endoplasmic reticulum (ER) membrane in an inactive state and are activated upon detachment from the membrane. Among these, in plants, the role of membrane-bound NAC transcription factors (NACMTFs) in stress signalling has recently been appreciated; however, in tomato, information regarding NACMTFs was largely unknown until now.
NAC transcription factor family consists of a large number of members having diversified roles in maintaining plant’s normal physiological activities such as development (Ben-Gera et al. 2012; Bergeret al., 2009; Hasson et al. 2011), as well as in defence against several stresses (Seo et al. 2010; Wang and Dane 2013). Genome-wide analysis has identified several NACs in rice, Arabidopsis, potato and soybean (Meng et al. 2007; Ooka et al. 2003; Singh et al. 2013). Also, recent publications indicated the existence of 74 and 104 NACs in tomato (Su et al. 2015; Kou et al. 2014). However, functional characterization of NACs has been limited to analysis of only a few members, and particularly tomato NACMTFs needed to be characterized. Amongst all the previous works involving tomato NACs, SlSRN1 is the only NACMTF which has been characterized (Liu et al. 2014).
We initiated identification of tomato NACMTFs by ensuring that information regarding all NAC TFs encoded in tomato genome was collected. Not surprisingly, our analysis revealed two novel NAC TFs, and we failed to confirm the existence of NAC domain in 6 genes (Solyc02g062090, Solyc02g077450, Solyc03g013190, Solyc08g068380, Solyc10g008880, and Solyc10g051070) that were previously marked as NACTFs (Su et al. 2015). While identifying SlNACMTFs, presence of at least one distinct transmembrane domain (TM) was looked for, and in 13 SlNACMTFs presence of this domain was predicted with high confidence. Thus, it is concluded that tomato genome encodes at least 13 SlNACMTFs. Further domain analysis indicated SlNACMTFs possess five distinct motifs (Supplementary Fig. 2) that are conserved for NAC (Olsen et al. 2005a; Ooka et al. 2003). High sequence similarity between NAC domains of SlNACMTFs with that of the previously reported NACMTFs of rice, Arabidopsis and potato (Fig. 1c) also confirmed their identity and suggesting these TFs probably possess similar functionality. The predicted TM domain in some of the identified SlNACMTFs is located in the N-terminus or within the NAC domain (Fig. 1b). It will be intriguing to find out whether the TM in these positions and removal of the same during membrane detachment would affect the functionality of the protein. However, in all of the characterized NAC MTFs (Kim et al. 2007b) the TM is located in the C-terminus, thus, further experimentation is required to show the effect of the TM placement in alternative positions of SlNACMTFs.
Subsequent experimental data confirmed that majority of these SlNACMTFs are always expressed, although in a varied amount in root, hypocotyl and leaf tissues. Consistent with commonly known fact, these transcription factors express in a relatively low amount in normal tissues (Fig. 2a, b). The relative level of expression of a majority of SlNACMTFs is somewhat comparable in different tissues, except the expression of SlNACMTF2, which is expressed at very high level in mature leaves compared to the young leaves (Fig. 2a, b). However, high level of SlNACMTF2 mRNA did not ensure a high level of translation, as the relative level of incorporation of the transcript in polysomal RNA was much lower compared to other SlNACMTFs (Fig. 3a). On the contrary, although SlNACMTF4, 7, 8, 9, 11 and 12 mRNAs were barely detected from total RNA, the level of these MTFs was higher in the polysomal fraction, indicating efficient translation occurs from these MTFs. Information regarding translation rate of MTFs is lacking in the literature, however, such discrepancy in mRNA expression level and translation rate is found for other genes including transcription factors (Greenbaum et al. 2003). To ascertain that these polysomal RNA fractions had minimum or very low free mRNA contamination, pri-miR167 and U6 were used as negative controls as precursor miRNA167 does not undergo translation (Wu et al. 2006) and U6 snRNA is transcribed by RNA polymerase III, thus also not translated (Kunkel et al. 1986). We could not amplify either of these sequences from the polysomal RNA preparation, hence proving that the preparation was sufficiently enriched of polysomal RNA.
NAC genes are differentially expressed depending on the type of stress the plant is enduring, and are known regulator of plant stress response (Puranik et al. 2012). Analysis of transcriptome data from many plants clearly shows that a significant number of NAC genes are involved in stress (Huang et al. 2012), however, NACMTFs were mostly studied in the context of abiotic stress response only (Shao et al. 2015). Our Study conclusively portrayed the relative level of expression of all SlNACMTFs in tomato plants exposed to alternative biotic or abiotic stress conditions. Amongst them, SlNACMTF3 and 8 were chosen for in-depth analysis to verify their membrane localization ability, DNA binding specificity, and transcriptional regulatory roles.
The role of NACMTFs in biotic stress response or characteristics of biotic stress responsive NACMTFs are much less explored. Thus we have prioritized the characterization of two biotic stress responsive SlNACMTFs. The reasons for choosing SlNACMTF3 and 8 for further characterization are as follows. These two candidates showed significant upregulation while plants are infected with ToLCNDV, a biotrophic pathogen, and A. solani, a necrotrophic pathogen, respectively. Incidentally, these two SlNACMTFs belong to two different phylogenetic groups (Fig. 1c), thus characterization of SlNACMTFs with less functional redundancy could be performed. OsNAC74, a rice NACMTF (Kimet al. 2010), known to be a homolog of OsNAC8 which is regulated in response to salt stress (Chao et al. 2005), belongs to the same group with SlNACMTF8. Whereas, SlNACMTF3 belongs to the same group in which AtNTL6 and 8 are clustered. These two NACMTFs are known for their role in biotic and abiotic stresses, respectively (Kim and Park 2007; Seo and Park 2010). Clustering in the same group with these previously reported NAC genes could also be an indication of their functional similarity.
We also questioned whether the huge upregulation of SlNACMTF8 expression in Alternaria infected tissue also leads to the increased inclusion of the mRNA in the translation machinery. In polysomal RNA fraction, high expression of SlNACMTF8 was indeed reflected, as it showed almost 216 fold upregulation in the case of fungus stressed samples (Fig. 5f). As upregulation of a gene in mRNA level does not always assure their enhanced expression, supporting information from the polysomal RNA expression data can certainly provide a validation that this gene is also highly expressed in protein level. SlNACMTF8 is also highly expressed in salt stress, which also signifies that a single NACMTF gene can act as a regulator of more than one stress. This observation is in concurrence with previously reported for Arabidopsis NAC transcription factor ATAF1 expression pattern (Wu et al. 2009).
Genes are regulated at both transcriptional and posttranscriptional stages; however, most of the regulation occurs at transcriptional initiation. Thus, we set out to uncover putative binding elements for regulatory transcription factors on SlNACMTF promoters. For this purpose, 1 kb sequences upstream to the A of predicted translation initiation site (ATG) was collected from genome sequence database and analyzed using MatInspector (Genomatix). Only elements predicted with high confidence were considered (Supplementary Fig. 3). Our analysis revealed the existence of multiple development and stress-related cis-regulatory elements, such as binding sites for WUSCHEL, NAC, Myb; as well as W-box, ABRE, Ethylene or SA-responsive elements, along with the time of day specific regulatory elements such as morning and evening elements (Supplementary Fig. 3). These findings support the view that transcriptional regulation during stress is a major determinant of their expression level. It is worth mentioning that except SlNACMTF3 all other genes possess multiple putative NAC binding sites on their promoter, which indicate that these genes might also undergo autoregulation. The promoter of SlNACMTF3 contains ABA and Auxin-responsive elements. ABRE has a substantial role in the regulation of several NACTFs during abiotic stress (Takasaki et al. 2010). Similarly, Auxin-responsive elements were also known for their role in the regulation of NAC genes while present in the promoter (Gutiérrez et al. 2009). Furthermore, some of these SlNACMTFs, including SlNACMTF8 possess W-box in their promoter. WRKY transcription factors are known regulator of biotic and abiotic stresses and works in ABA-independent mode of signalling during stress (Wang et al. 2016). The existence of W-box in the putative promoter-regulatory region of these NAC genes also strongly suggests expression of these genes will be affected by stress.
Moreover, when the coding sequences of these SlNACMTFs were analysed in psRNATarget (plantgrn.noble.org/psRNATarget/), 10 out of these 13 SlNACMTFs were predicted to be targeted by several miRNAs (Supplementary table 4), including miR164 which is a known posttranscriptional regulator of NAC gene expression (Fang et al. 2014). However, experimental verification of the SlNACMTF-miRNA interaction has yet to be conducted.
Similar to many other NACMTFs (Zhao et al. 2016), full-length SlNACMTF3 was also exclusively localized to the plasma membrane and, our analysis with plasmolysed cells confirmed that these forms did not reach the cell wall. This localization was solely dependent on the presence of the transmembrane domain, as removal of it abolished transmembrane transfer of the protein. SlNACMTF3ΔC protein was distributed in cytoplasm and the nucleus. This is in agreement with the predictions by different software (SeqNLS, cNLSmapper, NLStradamus and PSORTII) of a NLS located in the N-terminus of the protein. These observations also suggest that in full-length protein the domain responsible for transport to the nucleus is masked by the presence of the C-terminus including the TM domain.
NAC transcription factors recognise and bind to a specific sequence named as NAC binding sites (NACbs). Although the core region sequence of these NACbss are quite conserved, the flanking sequence may vary (Shamimuzzaman and Vodkin 2013). In general NAC binding sites consists of the following sequence (A/T)(T/G)(G/A)CGT(A/G)(T/C/A)(T/A/G)N1−6(C/T/A)(T/G/A)(T/C)ACG(T/C)(A/C), CGT/ACG core sequence is highlighted. Apart from the core sequence, NACbs varies greatly in their flanking sequences, and a particular NAC protein’s affinity to a NACbs varies widely with the alterations in these flanking sequences. For example, NAC recognition site for ANAC019 (TTNCGTA) and ANAC092 (TTGCGTGT) are similar but is not exactly identical (Olsen et al. 2005b). Generally, the core sequence CGT is preceded and followed by an A/G and preferably a thymine is required 2–3 bases upstream of the core sequence. Additionally, the 3′-end of the flanking sequence is preferably marked with guanine residues. The NACbs used in our study (Fig. 7a) has a likelihood of the NACbs of ANAC092. Although both SlNACMTF3 and SlNACMTF8 could specifically interact with the consensus NACbs that has been used, displayed weak affinity as the interactions did not yield intense shifted bands. This was probably not due to the quality of the purified protein utilized in these experiments or reaction conditions; because in vitro translated protein (Fig. 7b, c) and altered reaction conditions also produced a similar result. Nevertheless, SlNACMTF3 seems to have slightly more affinity towards the consensus NACbs used in this study (compare band intensity in Fig. 7f vs. g). Thus, these two different NAC proteins might interact in a slightly different manner with the same binding site, and detailed analysis is required to determine their exact preference for a NACbs sequence. Despite their apparent differences in DNA sequence selectivity, both SlNACMTF3ΔC and 8ΔC could alter expression of GUS reporter gene driven by CaMV35S promoter which contains two NACbss (Fig. 8a, b). The SlNACMTFs did not much influence promoter fragment lacking one of the NACbss, further confirming that these two SlNACMTFs can efficiently exert their effect on activities of a promoter via a particular NACbs only (Fig. 8b). We also carried out the experiment with the full-length SlNACMTF3, as expected, the protein was unable to cause any significant difference in the promoter’s activity (data not shown). This data is following observations by others that the C-terminus of NACMTFs inhibits the transcriptional regulatory function of the protein (Kim et al. 2006).
SlNACMTF3 and 8 differ in their transcriptional regulatory role depending on the type of NACbs they have encountered or the promoter context. SlNACMTF3 acted as a repressor while SlNACMTF8 activated transcription from CaMV35S promoter (Fig. 8b). NACs are known regulators of downstream genes (Fujita et al. 2004). CaMNACs or calmodulin binding NACs are found to act as repressors. These transcription factors interact with Calmodulin which in turn, controls the repressor activity (Kim et al. 2007a). We have also observed that SlNACMTF3ΔC has a putative CaM-binding site in the N-terminal region (does not overlap the NAC domain). This domain, 2KVWILVMTVTPV12, is very similar to the binding site (KRWKVVATVMVAVMVG) of Arabidopsis Calmodulin binding repressor NAC, CBNAC (NP_567986). However, further experimentation is required to link the domain with repressor activity of SlNACMTF3ΔC on CaMV35S promoter. SlNACMTF8ΔC, on the other hand, increases the activity of CaMV35S promoter significantly and thus, could be termed as an activator for the promoter. Though it interacts mildly with the consensus NACbs that we have used in our EMSA assays, it could be clearly understood from our data of promoter activity, that it was able to promote transcription efficiently from CaMV35S promoter. This information again signifies that an individual NAC protein might have a selective affinity towards different NAC binding sites. Similar behaviour of transcription factors has also been described in the literature (Olsen et al. 2005b). Although both SlNACMTF3 and 8 are potentially interacting with the same NACbs, probably their interacting partners have a crucial role in mediating transcriptional regulatory function.
There is no distinct difference in domain structure in a transcription activator and repressor, and the role that they exhibit is mainly determined by other interacting proteins called coactivators or corepressors (Singh 1998). A single transcription factor may act as either activator or repressor while regulating different genes. For example, Robatzek and Somssich (2002), have shown that AtWRKY6 protein negatively regulates its promoter while activating pathogenesis related 1 (PR1) gene expression (Robatzek and Somssich 2002). Similarly, the nuclear-localized form of SlNACMTF3 and SlNACMTF8 exhibited differential regulatory role on cellular genes which are known regulator/effector of a biotic stress response. Taken together, our data on biotic stress-mediated regulation of expressions of SlNACMTF3 and 8, and their ability to regulate biotic stress-related gene expression, clearly show that these MTFs have a potential role in mediating biotic stress signal transduction.
Tomato plants are susceptible to necrotrophic (A. solani) and biotrophic (Tomato leaf curl virus, TLCV) pathogens. Biotic stress signalling starts at the membrane, however, not much work has been done to understand membrane-initiated direct signalling for transmission of stress response for these pathogens. Our results strongly indicate that activation of NACMTFs might be crucial in this process. Understanding the cues and mechanism for activation of MTFs and subsequent regulation of a set of genes by them after mobilization into nucleus will help in the elucidation of host susceptibility pathway.
References
Aida M, Ishida T, Fukaki H, Fujisawa H, Tasaka M (1997) Genes involved in organ separation in Arabidopsis: an analysis of the cup-shaped cotyledon mutant. Plant Cell 9:841–857
Ben-Gera H, Shwartz I, Shao MR, Shani E, Estelle M, Ori N (2012) ENTIRE and GOBLET promote leaflet development in tomato by modulating auxin response. Plant J 70:903–915
Berger Y, Harpaz-Saad S, Brand A, Melnik H, Sirding N, Alvarez JP, Zinder M, Samach A, Eshed Y, Ori N (2009) The NAC-domain transcription factor GOBLET specifies leaflet boundaries in compound tomato leaves. Development 136:823–832
Chao DY, Luo YH, Shi M, Luo D, Lin HX (2005) Salt-responsive genes in rice revealed by cDNA microarray analysis. Cell Res 15:796–810
Du M, Zhai Q, Deng L, Li S, Li H, Yan L, Huang Z, Wang B, Jiang H, Huang T, Li CB, Wei J, Kang L, Li J, Li C (2014) Closely related NAC transcription factors of tomato differentially regulate stomatal closure and reopening during pathogen attack. Plant Cell 26:3167–3184
Fang Y, Xie K, Xiong L (2014) Conserved miR164-targeted NAC genes negatively regulate drought resistance in rice. J Exp Bot 65:2119–2135
Fujita M, Fujita Y, Maruyama K, Seki M, Hiratsu K, Ohme-Takagi M, Tran LS, Yamaguchi-Shinozaki K, Shinozaki K (2004) A dehydration-induced NAC protein, RD26, is involved in a novel ABA-dependent stress-signaling pathway. Plant J 39:863–876
Greenbaum D, Colangelo C, Williams K, Gerstein M (2003) Comparing protein abundance and mRNA expression levels on a genomic scale. Genome Biol 4:117
Gutiérrez J, López Núñez-Flores MJ, Gómez-Ros LV, Novo Uzal E, Esteban Carrasco A, Díaz J, Sottomayor M, Cuello J, Ros Barceló A (2009) Hormonal regulation of the basic peroxidase isoenzyme from Zinnia elegans. Planta 230:767–778
Han Q, Zhang J, Li H, Luo Z, Ziaf K, Ouyang B, Wang T, Ye Z (2012) Identification and expression pattern of one stress-responsive NAC gene from Solanum lycopersicum. Mol Biol Rep 39:1713–1720
Hasson A, Plessis A, Blein T, Adroher B, Grigg S, Tsiantis M, Boudaoud A, Damerval C, Laufs P (2011) Evolution and diverse roles of the CUP-SHAPED COTYLEDON genes in Arabidopsis leaf development. Plant Cell 23:54–68
Hendelman A, Stav R, Zemach H, Arazi T (2013) The tomato NAC transcription factor SlNAM2 is involved in flower-boundary morphogenesis. J Exp Bot 64:5497–5507
Hu H, Dai M, Yao J, Xiao B, Li X, Zhang Q, Xiong L (2006) Overexpressing a NAM, ATAF, and CUC (NAC) transcription factor enhances drought resistance and salt tolerance in rice. Proc Natl Acad Sci USA 103:12987
Huang H, Wang Y, Wang S, Wu X, Yang K, Niu Y, Dai S (2012) Transcriptome-wide survey and expression analysis of stress-responsive NAC genes in Chrysanthemum lavandulifolium. Plant Sci 193–194:18
Huang W, Miao M, Kud J, Niu X, Ouyang B, Zhang J, Ye Z, Kuhl JC, Liu Y, Xiao F (2013) SlNAC1, a stress-related transcription factor, is fine-tuned on both the transcriptional and the post-translational level. New Phytol 197:1214–1224
Iwata Y, Koizumi N (2005) An Arabidopsis transcription factor, AtbZIP60, regulates the endoplasmic reticulum stress response in a manner unique to plants. Proc Natl Acad Sci USA 102:5280–5285
Kim SG, Park CM (2007) Membrane-mediated salt stress signaling in flowering time control. Plant Signal Behav 2:517–518
Kim YS, Kim SG, Park JE, Park HY, Lim MH, Chua NH, Park CM (2006) A membrane-bound NAC transcription factor regulates cell division in Arabidopsis. Plant Cell 18:3132–3144
Kim HS, Park BO, Yoo JH, Jung MS, Lee SM, Han HJ, Kim KE, Kim SH, Lim CO, Yun DJ, Lee SY, Chung WS (2007a) Identification of a calmodulin-binding NAC protein as a transcriptional repressor in Arabidopsis. J Biol Chem 282:36292–36302
Kim SY, Kim SG, Kim YS, Seo PJ, Bae M, Yoon HK, Park CM (2007b) Exploring membrane-associated NAC transcription factors in Arabidopsis: implications for membrane biology in genome regulation. Nucleic Acids Res 35:203–213
Kim SG, Lee S, Seo PJ, Kim SK, Kim JK, Park CM (2010) Genome-scale screening and molecular characterization of membrane-bound transcription factors in Arabidopsis and rice. Genomics 95:56–65
Kou X, Wang S, Wu M, Guo R, Xue Z, Meng N, Tao X, Chen M, Zhang Y (2014) Molecular characterization and expression analysis of NAC family transcription factors in tomato. Plant Mol Biol Rep 32:501–516
Kundu P, Li M, Lu R, Stefani E, Toro L (2015) Regulation of transcriptional activation function of rat estrogen receptor α (ERα) by novel C-terminal splice inserts. Mol Cell Endocrinol 401:202–212
Kunkel GR, Maser RL, Calvet JP, Pederson T (1986) U6 small nuclear RNA is transcribed by RNA polymerase III. Proc Natl Acad Sci USA 83:8575–8579
Le DT, Nishiyama R, Watanabe Y, Mochida K, Yamaguchi-Shinozaki K, Shinozaki K, Tran LS (2011) Genome-wide survey and expression analysis of the plant-specific NAC transcription factor family in soybean during development and dehydration stress. DNA Res 18:263
Liu B, Ouyang Z, Zhang Y, Li X, Hong Y, Huang L, Liu S, Zhang H, Li D, Song F (2014) Tomato NAC transcription factor SlSRN1 positively regulates defense response against biotic stress but negatively regulates abiotic stress response. PLoS One 9:e102067
Ma NN, Zuo YQ, Liang XQ, Yin B, Wang GD, Meng QW (2013). The multiple stress-responsive transcription factor SlNAC1 improves the chilling tolerance of tomato. Physiol Plant 149:474–486
Ma N, Feng H, Meng X, Li D, Yang D, Wu C, Meng Q (2014) Overexpression of tomato SlNAC1 transcription factor alters fruit pigmentation and softening. BMC Plant Biol 14:351
Mandal A, Sarkar D, Kundu S, Kundu P (2015) Mechanism of regulation of tomato TRN1 gene expression in late infection with tomato leaf curl New Delhi virus (ToLCNDV). Plant Sci 241:221–237
Meng Q, Zhang C, Gai J, Yu D (2007) Molecular cloning, sequence characterization and tissue-specific expression of six NAC-like genes in soybean (Glycine max (L.) Merr.). J Plant Physiol 164:1002–1012
Mustroph A, Juntawong P, Bailey-Serres J (2009) Isolation of plant polysomal mRNA by differential centrifugation and ribosome immunopurification methods. Methods Mol Biol 553:109–126
Nuruzzaman M, Sharoni AM, Kikuchi S (2013) Roles of NAC transcription factors in the regulation of biotic and abiotic stress responses in plants. Front Microbiol 4:248
Olsen AN, Ernst HA, Leggio LL, Skriver K (2005a) NAC transcription factors: structurally distinct, functionally diverse. Trends Plant Sci 10:79–87
Olsen AN, Ernst HA, Lo Leggio L, Skriver K (2005b) DNA-binding specificity and molecular functions of NAC transcription factors. Plant Sci 169:785–797
Ooka H, Satoh K, Doi K, Nagata T, Otomo Y, Murakami K, Matsubara K, Osato N, Kawai J, Carninci P, Hayashizaki Y, Suzuki K, Kojima K, Takahara Y, Yamamoto K, Kikuchi S (2003) Comprehensive analysis of NAC family genes in Oryza sativa and Arabidopsis thaliana. DNA Res 10:239–247
Puranik S, Sahu PP, Srivastava PS, Prasad M (2012) NAC proteins: regulation and role in stress tolerance. Trends Plant Sci 17:369–381
Robatzek S, Somssich IE (2002) Targets of AtWRKY6 regulation during plant senescence and pathogen defense. Genes Dev 16:1139–1149
Schmittgen TD, Livak KJ (2008) Analyzing real-time PCR data by the comparative C(T) method. Nat Protoc 3:1101–1108
Selth LA, Dogra SC, Rasheed MS, Healy H, Randles JW, Rezaian MA (2005) A NAC domain protein interacts with tomato leaf curl virus replication accessory protein and enhances viral replication. Plant Cell 17:311–325
Seo PJ (2014) Recent advances in plant membrane-bound transcription factor research: emphasis on intracellular movement. J Integr Plant Biol 56:334–342
Seo PJ, Park CM (2010) A membrane-bound NAC transcription factor as an integrator of biotic and abiotic stress signals. Plant Signal Behav 5:481–483
Seo PJ, Kim SG, Park CM (2008) Membrane-bound transcription factors in plants. Trends Plant Sci 13:550–556
Seo PJ, Kim MJ, Park JY, Kim SY, Jeon J, Lee YH, Kim J, Park CM (2010) Cold activation of a plasma membrane-tethered NAC transcription factor induces a pathogen resistance response in Arabidopsis. Plant J 61:661–671
Shamimuzzaman M, Vodkin L (2013) Genome-wide identification of binding sites for NAC and YABBY transcription factors and co-regulated genes during soybean seedling development by ChIP-Seq and RNA-SEq. BMC Genomics 14:477
Shao H, Wang H, Tang X (2015) NAC transcription factors in plant multiple abiotic stress responses: progress and prospects. Front Plant Sci 6:902
Singh KB (1998) Transcriptional regulation in plants: the importance of combinatorial control. Plant Physiol 118:1111–1120
Singh J, Sastry EV, Singh V (2012) Effect of salinity on tomato (Lycopersicon esculentum Mill.) during seed germination stage. Physiol Mol Biol Plants 18:45–50
Singh AK, Sharma V, Pal AK, Acharya V, Ahuja PS (2013) Genome-wide organization and expression profiling of the NAC transcription factor family in potato (Solanum tuberosum L.). DNA Res 20:403–423
Su H, Zhang S, Yin Y, Zhu D, Han L (2015) Genome-wide analysis of NAM-ATAF1,2-CUC2 transcription factor family in Solanum lycopersicum. J Plant Biochem Biotechnol 24:176–183
Takasaki H, Maruyama K, Kidokoro S, Ito Y, Fujita Y, Shinozaki K, Yamaguchi-Shinozaki K, Nakashima K (2010) The abiotic stress-responsive NAC-type transcription factor OsNAC5 regulates stress-inducible genes and stress tolerance in rice. Mol Genet Genomics 284:173–183
Wang ZY, Dane F (2013) NAC (NAM/ATAF/CUC) transcription factors in different stresses and their signaling pathway. Acta Physiol Plant 35:1397–1408
Wang H, Shao H, Tang X (2016) Recent advances in utilizing transcription factors to improve plant abiotic stress tolerance by transgenic technology. Front Plant Sci 7:67
Wu MF, Tian Q, Reed JW (2006) Arabidopsis microRNA167 controls patterns of ARF6 and ARF8 expression, and regulates both female and male reproduction. Development 133:4211–4218
Wu Y, Deng Z, Lai J, Zhang Y, Yang C, Yin B, Zhao Q, Zhang L, Li Y, Xie Q (2009) Dual function of Arabidopsis ATAF1 in abiotic and biotic stress responses. Cell Res 19:1279–1290
Xu K, Huang X, Wu M, Wang Y, Chang Y, Liu K, Zhang J, Zhang Y, Zhang F, Yi L, Li T, Wang R, Tan G, Li C (2014) A rapid, highly efficient and economical method of Agrobacterium-mediated in planta transient transformation in living onion epidermis. PLoS One 9:e83556
Yang R, Deng C, Ouyang B, Ye Z (2011) Molecular analysis of two salt-responsive NAC-family genes and their expression analysis in tomato. Mol Biol Rep 38:857–863
Yang YG, Lv WT, Li MJ, Wang B, Sun DM, Deng X (2013) Maize membrane-bound transcription factor Zmbzip17 is a key regulator in the cross-talk of ER quality control and ABA signaling. Plant Cell Physiol 54:2020
Zhao J, Liu JS, Meng FN, Zhang ZZ, Long H, Lin WH, Luo XM, Wang ZY, Zhu SW (2016) ANAC005 is a membrane-associated transcription factor and regulates vascular development in Arabidopsis. J Integr Plant Biol 58:442
Acknowledgements
The ToLCNDV agroinfiltration constructs used in this study were generous gifts from Prof. Sunil Mukherjee, ICGEB. Primers for miR167 detection were designed by Ms. Jayanti Jodder. Authors are thankful to Drs. Debabrata Basu and Shubho Chaudhuri for their support, and Central Instrument Facility (CIF) of Bose Institute for all sequencing services. Authors are also thankful to Mr. Sougata Bhattacharya for support in bioinformatic analysis. PB is thankful to UGC, Govt. of India for fellowship. This work is supported by a project from CSIR, Govt. of India to PK.
Author contributions
PB and PK designed the research. PB carried out all experiments. PB and PK analyzed the data. RD and AM have provided some reagents. PB and PK wrote the manuscript.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Bhattacharjee, P., Das, R., Mandal, A. et al. Functional characterization of tomato membrane-bound NAC transcription factors. Plant Mol Biol 93, 511–532 (2017). https://doi.org/10.1007/s11103-016-0579-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-016-0579-z